

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0258a.2
(167 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
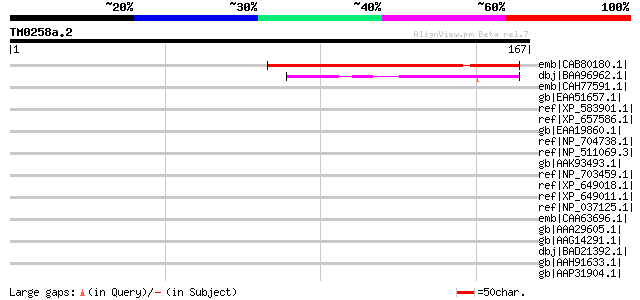
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB80180.1| putative protein [Arabidopsis thaliana] gi|30969... 80 2e-14
dbj|BAA96962.1| unnamed protein product [Arabidopsis thaliana] g... 46 4e-04
emb|CAH77591.1| conserved hypothetical protein [Plasmodium chaba... 43 0.003
gb|EAA51657.1| hypothetical protein MG03252.4 [Magnaporthe grise... 43 0.004
ref|XP_583901.1| PREDICTED: similar to nucleolin-related protein... 42 0.005
ref|XP_657586.1| hypothetical protein 1.t00028.part_1 [Entamoeba... 42 0.006
gb|EAA19860.1| Drosophila melanogaster LD33051p [Plasmodium yoel... 42 0.006
ref|NP_704738.1| hypothetical protein [Plasmodium falciparum 3D7... 42 0.008
ref|NP_511069.3| CG9653-PA [Drosophila melanogaster] gi|45446835... 41 0.011
gb|AAK93493.1| SD02279p [Drosophila melanogaster] 41 0.011
ref|NP_703459.1| hypothetical protein [Plasmodium falciparum 3D7... 41 0.014
ref|XP_649018.1| hypothetical protein 354.t00010 [Entamoeba hist... 41 0.014
ref|XP_649011.1| hypothetical protein 354.t00008 [Entamoeba hist... 41 0.014
ref|NP_037125.1| n-arginine dibasic convertase 1 [Rattus norvegi... 40 0.019
emb|CAA63696.1| NRD2 convertase [Rattus sp.] 40 0.019
gb|AAA29605.1| glutamic acid-rich protein [Plasmodium falciparum... 40 0.019
gb|AAG14291.1| glutamic acid-rich protein [Plasmodium falciparum] 40 0.019
dbj|BAD21392.1| mFLJ00158 protein [Mus musculus] 40 0.024
gb|AAH91633.1| Unknown (protein for MGC:98482) [Xenopus laevis] 40 0.024
gb|AAP31904.1| histidine-rich calcium binding protein [Rattus no... 40 0.024
>emb|CAB80180.1| putative protein [Arabidopsis thaliana] gi|3096932|emb|CAA18842.1|
putative protein [Arabidopsis thaliana]
gi|21554188|gb|AAM63267.1| unknown [Arabidopsis
thaliana] gi|22136348|gb|AAM91252.1| putative protein
[Arabidopsis thaliana] gi|20466402|gb|AAM20518.1|
putative protein [Arabidopsis thaliana]
gi|15236167|ref|NP_195189.1| expressed protein
[Arabidopsis thaliana] gi|7487661|pir||T05283
hypothetical protein T4L20.210 - Arabidopsis thaliana
Length = 199
Score = 80.5 bits (197), Expect = 2e-14
Identities = 42/81 (51%), Positives = 54/81 (65%), Gaps = 2/81 (2%)
Query: 84 YLHGYAYEDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKNSK 143
Y G Y D +++ + DD DLD+ELVP V K+GRQRM+KLGKRA +K+ SK
Sbjct: 118 YSVGITYGDSVNDDGEEDREYDDSYDLDEELVPRSVSKKVGRQRMRKLGKRAIAKVYASK 177
Query: 144 RSPYLFVRPGCVGGKHGLGLK 164
SP F++PG V GKHGLG+K
Sbjct: 178 ISP--FLKPGIVRGKHGLGMK 196
>dbj|BAA96962.1| unnamed protein product [Arabidopsis thaliana]
gi|15238968|ref|NP_199653.1| hypothetical protein
[Arabidopsis thaliana]
Length = 142
Score = 45.8 bits (107), Expect = 4e-04
Identities = 31/77 (40%), Positives = 40/77 (51%), Gaps = 14/77 (18%)
Query: 90 YEDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKNSKRSPYLF 149
Y+ DEE D +D DD D ELV QR++K GKR K K+ Y +
Sbjct: 75 YDTDEEDDFDYYDDLDD----DHELVKI--------QRIRKTGKRTFWKNVTLKKKKYSY 122
Query: 150 --VRPGCVGGKHGLGLK 164
++PGCV GKHGL +K
Sbjct: 123 SQLKPGCVHGKHGLDIK 139
>emb|CAH77591.1| conserved hypothetical protein [Plasmodium chabaudi]
Length = 755
Score = 43.1 bits (100), Expect = 0.003
Identities = 26/86 (30%), Positives = 47/86 (54%), Gaps = 9/86 (10%)
Query: 33 VDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAYE- 91
++ +EE+E+E D +++D EEI++ + D + VE D ++ + E
Sbjct: 38 IEVDEEEEEEEDDDEED----EEIEVDEEEEEEEEDDEEDEEVEADEEEEEEIDDDGEEE 93
Query: 92 --DDEEQVVDDEE--DDDDRCDLDDE 113
DDEE++ +DEE DDD+ + D+E
Sbjct: 94 ADDDEEEIEEDEEEVDDDEEIEEDEE 119
Score = 38.5 bits (88), Expect = 0.071
Identities = 24/94 (25%), Positives = 49/94 (51%), Gaps = 10/94 (10%)
Query: 20 QDDSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDR 79
++D + E ++V + EEE+E++ +++++ A EE + I D D + E +
Sbjct: 47 EEDDDEEDEEIEVDEEEEEEEEDDEEDEEVEADEEEEEEIDDDGEEEADDDEEEIEEDEE 106
Query: 80 QQYHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDE 113
+ DD+E++ +DEE+ DD D ++E
Sbjct: 107 EV----------DDDEEIEEDEEEMDDDDDAEEE 130
Score = 36.6 bits (83), Expect = 0.27
Identities = 21/83 (25%), Positives = 41/83 (49%), Gaps = 12/83 (14%)
Query: 31 DVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAY 90
D + EE+ E++ ++ + D EEI++ + D + +E+D
Sbjct: 13 DDEEDEEDVEEDDEEEEDDDEEDEEIEVDEEEEEEEDDDEEDEEIEVD------------ 60
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
E++EE+ DDEED++ D ++E
Sbjct: 61 EEEEEEEEDDEEDEEVEADEEEE 83
Score = 34.3 bits (77), Expect = 1.3
Identities = 24/102 (23%), Positives = 48/102 (46%), Gaps = 32/102 (31%)
Query: 12 EQVQFLKVQDDSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDA 71
E+++ + +++ E + V+A+EE+E+E D + ++ A +E +I
Sbjct: 55 EEIEVDEEEEEEEEDDEEDEEVEADEEEEEEIDDDGEEEADDDEEEI------------- 101
Query: 72 VAVVEIDRQQYHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDE 113
E+DEE+V DDEE ++D ++DD+
Sbjct: 102 -------------------EEDEEEVDDDEEIEEDEEEMDDD 124
Score = 33.9 bits (76), Expect = 1.7
Identities = 26/96 (27%), Positives = 44/96 (45%), Gaps = 25/96 (26%)
Query: 20 QDDSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDR 79
+DD + + D D EE+ E+E D +++D EEI++ + D
Sbjct: 6 EDDEDDEDDEEDEEDVEEDDEEEEDDDEED----EEIEVDEEEEEEEDDD---------- 51
Query: 80 QQYHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDELV 115
E+DEE VD+EE++++ D +DE V
Sbjct: 52 -----------EEDEEIEVDEEEEEEEEDDEEDEEV 76
>gb|EAA51657.1| hypothetical protein MG03252.4 [Magnaporthe grisea 70-15]
gi|39942344|ref|XP_360709.1| hypothetical protein
MG03252.4 [Magnaporthe grisea 70-15]
Length = 4048
Score = 42.7 bits (99), Expect = 0.004
Identities = 26/113 (23%), Positives = 54/113 (47%), Gaps = 3/113 (2%)
Query: 19 VQDDSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEID 78
V+DD +WE D E++ EQ+ + A ++E ++ H + P + V
Sbjct: 2520 VEDDGASEWESESENDDEDDDEQDEIDYEDGAQDIDEARL-HGLEPGQLSLENYRDVVRA 2578
Query: 79 RQQYHYLHGYAYEDDE--EQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMK 129
++ + DD E+ D++D+D+ D+D+++ ++ GN +G R +
Sbjct: 2579 AMGTDFIDDFPAIDDRFGEEDEADDDDEDEEDDIDEDVFVYDHGNMVGGHRQR 2631
Score = 35.4 bits (80), Expect = 0.60
Identities = 24/84 (28%), Positives = 40/84 (47%), Gaps = 13/84 (15%)
Query: 31 DVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAY 90
D ++ +E+ Q+ ++N D +EI + + + D V V+ D
Sbjct: 2427 DEMEYDEDMSQDDEENVSDDD--DEIDGMGHVEGLPGDAGVVEVIMGDD----------- 2473
Query: 91 EDDEEQVVDDEEDDDDRCDLDDEL 114
+DD+E + DDEED D D DDE+
Sbjct: 2474 DDDDESMDDDEEDTSDEDDEDDEM 2497
>ref|XP_583901.1| PREDICTED: similar to nucleolin-related protein, partial [Bos
taurus]
Length = 675
Score = 42.4 bits (98), Expect = 0.005
Identities = 29/113 (25%), Positives = 51/113 (44%), Gaps = 4/113 (3%)
Query: 34 DAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAYEDD 93
D EEE+E E D+ +D + E ++I+ P V V + + +DD
Sbjct: 148 DDEEEEEDEEDEEDEDDSEEEPMEIV----PAKGKKAPVKAVPVKAKSTAEDEDEEDDDD 203
Query: 94 EEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKNSKRSP 146
++ ++EEDDDD D D+E E + + +K+ + +M K +P
Sbjct: 204 DDDEEEEEEDDDDEDDDDEEEEEEEEEEEEEEEPVKEAPGKRKKEMAKQKAAP 256
Score = 34.7 bits (78), Expect = 1.0
Identities = 24/96 (25%), Positives = 43/96 (44%), Gaps = 17/96 (17%)
Query: 31 DVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAY 90
D D ++ +E++ D ++D V + P D D E D
Sbjct: 101 DEEDEDDSEEEDDDDEEEDEPAVRKAAAAFGAAPATDDEDD----EDDD----------- 145
Query: 91 EDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQ 126
+DD+E+ +DEED++D D ++E P E+ G++
Sbjct: 146 DDDDEEEEEDEEDEEDEDDSEEE--PMEIVPAKGKK 179
>ref|XP_657586.1| hypothetical protein 1.t00028.part_1 [Entamoeba histolytica
HM-1:IMSS] gi|56474845|gb|EAL52200.1| hypothetical
protein 1.t00028.part_1 [Entamoeba histolytica
HM-1:IMSS]
Length = 208
Score = 42.0 bits (97), Expect = 0.006
Identities = 28/93 (30%), Positives = 45/93 (48%), Gaps = 25/93 (26%)
Query: 21 DDSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQ 80
D++ ++EL D D EEE E+E D+ ++D D D + E D +
Sbjct: 37 DNNEDEYELEDEDDNEEEDEEEDDELEEDD----------------DDDDEYELEEEDEE 80
Query: 81 QYHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDE 113
+ +DDE++ DD+EDDD+ D DD+
Sbjct: 81 E---------DDDEDEEEDDDEDDDEDDDEDDD 104
>gb|EAA19860.1| Drosophila melanogaster LD33051p [Plasmodium yoelii yoelii]
Length = 939
Score = 42.0 bits (97), Expect = 0.006
Identities = 30/110 (27%), Positives = 52/110 (47%), Gaps = 4/110 (3%)
Query: 33 VDAEEEQEQEPDQNQQDAAPVEEIKI-IHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAYE 91
VD EE E+E D+ + D V+E ++ ++ D + V E+D ++ A
Sbjct: 674 VDEEEVDEEEVDEEEVDEEEVDEEEVDEEEVDEEEVDEEEVDEEEVDEEEADAEEEEADA 733
Query: 92 DDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKN 141
++EE ++EE D++ D ++ E G K+ R K R K+KN
Sbjct: 734 EEEEADAEEEEADEEEVDEEEAEAEAEAGGKINGSRKK---ARREKKIKN 780
Score = 37.4 bits (85), Expect = 0.16
Identities = 28/111 (25%), Positives = 49/111 (43%), Gaps = 9/111 (8%)
Query: 33 VDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAYED 92
VD EE E+E D+ + D V+E ++ D + V E D ++ D
Sbjct: 689 VDEEEVDEEEVDEEEVDEEEVDEEEV---------DEEEVDEEEADAEEEEADAEEEEAD 739
Query: 93 DEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKNSK 143
EE+ D+EE D++ + + E G++ +R KK+ + KN +
Sbjct: 740 AEEEEADEEEVDEEEAEAEAEAGGKINGSRKKARREKKIKNELMIQKKNEE 790
>ref|NP_704738.1| hypothetical protein [Plasmodium falciparum 3D7]
gi|23505099|emb|CAD51881.1| hypothetical protein
[Plasmodium falciparum 3D7]
Length = 3381
Score = 41.6 bits (96), Expect = 0.008
Identities = 25/95 (26%), Positives = 48/95 (50%), Gaps = 2/95 (2%)
Query: 19 VQDDSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEID 78
+++D+ V + D + + E+ ++QD ++ + IR + + + VVE+
Sbjct: 804 MKNDTFNNKNNVTINDEQNDSEEYIYNSKQDDVSCDDSSVNKDIRK--NNYEGIPVVEVQ 861
Query: 79 RQQYHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDE 113
+Y+ + +DD++ DD EDDDD D DDE
Sbjct: 862 DDEYYEENEQDNDDDDDDDEDDGEDDDDDEDDDDE 896
>ref|NP_511069.3| CG9653-PA [Drosophila melanogaster] gi|45446835|gb|AAF46251.3|
CG9653-PA [Drosophila melanogaster]
gi|4530532|gb|AAD22080.1| putative transcription factor
[Drosophila melanogaster] gi|4587308|dbj|BAA76710.1| Brk
[Drosophila melanogaster]
Length = 704
Score = 41.2 bits (95), Expect = 0.011
Identities = 42/156 (26%), Positives = 73/156 (45%), Gaps = 26/156 (16%)
Query: 4 QPMQFLLLEQVQFLKVQDDSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIR 63
QP + LE+V +K + ++ + E V+V D E EQ +E P +++K+ +
Sbjct: 415 QPDKISKLEEV--IKKEPETETENEDVEV-DVETEQPEE------HKLPSKQVKLF---K 462
Query: 64 PVLPDPDAVAVVEIDRQQYHYLHGYAYEDDEEQVVDDEEDDDD--RCDLDDELVPWEVGN 121
P L D D ++ +H+ H + ED +E ++E+DD++ R DDE+ E +
Sbjct: 463 PYLLDDDE------EQDHHHHHHHHRQEDLDEGAAEEEQDDEEESRYADDDEVDSKEAAD 516
Query: 122 KLGRQRMKKLGKRACSKMKNSKRSPYLFVRPGCVGG 157
K R+ KK N +R P ++ GG
Sbjct: 517 KKQRRLKKK------PSAINEQREPIIWSNHPYPGG 546
>gb|AAK93493.1| SD02279p [Drosophila melanogaster]
Length = 502
Score = 41.2 bits (95), Expect = 0.011
Identities = 42/156 (26%), Positives = 73/156 (45%), Gaps = 26/156 (16%)
Query: 4 QPMQFLLLEQVQFLKVQDDSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIR 63
QP + LE+V +K + ++ + E V+V D E EQ +E P +++K+ +
Sbjct: 213 QPDKISKLEEV--IKKEPETETENEDVEV-DVETEQPEE------HKLPSKQVKLF---K 260
Query: 64 PVLPDPDAVAVVEIDRQQYHYLHGYAYEDDEEQVVDDEEDDDD--RCDLDDELVPWEVGN 121
P L D D ++ +H+ H + ED +E ++E+DD++ R DDE+ E +
Sbjct: 261 PYLLDDDE------EQDHHHHHHHHRQEDLDEGAAEEEQDDEEESRYADDDEVDSKEAAD 314
Query: 122 KLGRQRMKKLGKRACSKMKNSKRSPYLFVRPGCVGG 157
K R+ KK N +R P ++ GG
Sbjct: 315 KKQRRLKKK------PSAINEQREPIIWSNHPYPGG 344
>ref|NP_703459.1| hypothetical protein [Plasmodium falciparum 3D7]
gi|23504600|emb|CAD51479.1| hypothetical protein
[Plasmodium falciparum 3D7]
Length = 10061
Score = 40.8 bits (94), Expect = 0.014
Identities = 34/162 (20%), Positives = 76/162 (45%), Gaps = 35/162 (21%)
Query: 7 QFLLLEQVQFLKVQDDSLLQWELVDVV----------------DAEEEQEQEPDQNQQDA 50
Q ++E + L +D+ +L + DV+ D E+E+E+E D++++D
Sbjct: 7570 QAKIIEVGKKLTTKDEDMLHSKKTDVIQHGDEEEDDEEDDEEDDEEDEEEEEEDEDEEDV 7629
Query: 51 APVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAYEDD-------EEQVVDDEED 103
VE+I+ + + + + V+ D+ + +Y +DD +E + ++E++
Sbjct: 7630 EDVEDIEDVEDVEDIEDN-----YVDDDQYEDNYDDDNDNDDDDEYDHDYDEHINEEEQE 7684
Query: 104 DDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKNSKRS 145
DDD+ + + +E G + G ++ +K+K K+S
Sbjct: 7685 DDDKKNNVNINDSYEEGEEEGDGKLN-------TKIKKDKKS 7719
Score = 32.7 bits (73), Expect = 3.9
Identities = 13/22 (59%), Positives = 14/22 (63%)
Query: 92 DDEEQVVDDEEDDDDRCDLDDE 113
DDE V+DD DDD CD D E
Sbjct: 1139 DDESSVIDDNYIDDDSCDCDSE 1160
Score = 31.6 bits (70), Expect = 8.6
Identities = 13/23 (56%), Positives = 17/23 (73%)
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
+DDE+ DDE+DDDD D DD+
Sbjct: 8326 DDDEDDDEDDEDDDDDDDDDDDD 8348
Score = 31.6 bits (70), Expect = 8.6
Identities = 13/23 (56%), Positives = 17/23 (73%)
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
EDD+E DD++DDDD D DD+
Sbjct: 8329 EDDDEDDEDDDDDDDDDDDDDDD 8351
>ref|XP_649018.1| hypothetical protein 354.t00010 [Entamoeba histolytica HM-1:IMSS]
gi|56465359|gb|EAL43632.1| hypothetical protein
354.t00010 [Entamoeba histolytica HM-1:IMSS]
Length = 277
Score = 40.8 bits (94), Expect = 0.014
Identities = 25/85 (29%), Positives = 43/85 (50%), Gaps = 14/85 (16%)
Query: 31 DVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAY 90
D D E+E+++E +++++D E+ + + + D DAVA
Sbjct: 137 DEEDEEDEEDEEDEEDEEDEEDEEDEEDEYDLEEEEEDDDAVA------------ESLFA 184
Query: 91 EDDEEQVVDDEE--DDDDRCDLDDE 113
E D+E ++D +E DDDD DLDD+
Sbjct: 185 EFDDEDLLDSDEGLDDDDELDLDDD 209
Score = 33.9 bits (76), Expect = 1.7
Identities = 26/104 (25%), Positives = 49/104 (47%), Gaps = 7/104 (6%)
Query: 12 EQVQFLKVQDDSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDA 71
++ Q + +DD + + D D E+E+++E +++++D E+ + + D
Sbjct: 46 DEFQLDEEEDDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEED- 104
Query: 72 VAVVEIDRQQYHYLHGYAYEDDEEQVVD--DEEDDDDRCDLDDE 113
E D + E+DEE D DEED++D D +DE
Sbjct: 105 ----EEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDE 144
Score = 33.5 bits (75), Expect = 2.3
Identities = 23/85 (27%), Positives = 41/85 (48%), Gaps = 7/85 (8%)
Query: 31 DVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAY 90
D D E+E+++E +++++D E+ + + D E D +
Sbjct: 92 DEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEED-----EEDEEDEEDEEDEED 146
Query: 91 EDDEEQVVD--DEEDDDDRCDLDDE 113
E+DEE D DEED++D DL++E
Sbjct: 147 EEDEEDEEDEEDEEDEEDEYDLEEE 171
>ref|XP_649011.1| hypothetical protein 354.t00008 [Entamoeba histolytica HM-1:IMSS]
gi|56465355|gb|EAL43629.1| hypothetical protein
354.t00008 [Entamoeba histolytica HM-1:IMSS]
Length = 274
Score = 40.8 bits (94), Expect = 0.014
Identities = 25/85 (29%), Positives = 43/85 (50%), Gaps = 14/85 (16%)
Query: 31 DVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAY 90
D D E+E+++E +++++D E+ + + + D DAVA
Sbjct: 134 DEEDEEDEEDEEDEEDEEDEEDEEDEEDEYDLEEEEEDDDAVA------------ESLFA 181
Query: 91 EDDEEQVVDDEE--DDDDRCDLDDE 113
E D+E ++D +E DDDD DLDD+
Sbjct: 182 EFDDEDLLDSDEGLDDDDELDLDDD 206
Score = 33.9 bits (76), Expect = 1.7
Identities = 26/104 (25%), Positives = 49/104 (47%), Gaps = 7/104 (6%)
Query: 12 EQVQFLKVQDDSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDA 71
++ Q + +DD + + D D E+E+++E +++++D E+ + + D
Sbjct: 46 DEFQLDEEEDDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEED- 104
Query: 72 VAVVEIDRQQYHYLHGYAYEDDEEQVVD--DEEDDDDRCDLDDE 113
E D + E+DEE D DEED++D D +DE
Sbjct: 105 ----EEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDE 144
Score = 33.5 bits (75), Expect = 2.3
Identities = 23/85 (27%), Positives = 41/85 (48%), Gaps = 7/85 (8%)
Query: 31 DVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAY 90
D D E+E+++E +++++D E+ + + D E D +
Sbjct: 89 DEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEEDEED-----EEDEEDEEDEEDEED 143
Query: 91 EDDEEQVVD--DEEDDDDRCDLDDE 113
E+DEE D DEED++D DL++E
Sbjct: 144 EEDEEDEEDEEDEEDEEDEYDLEEE 168
>ref|NP_037125.1| n-arginine dibasic convertase 1 [Rattus norvegicus]
gi|1352519|sp|P47245|NRDC_RAT Nardilysin precursor
(N-arginine dibasic convertase) (NRD convertase) (NRD-C)
gi|529592|gb|AAA21818.1| NRD convertase
Length = 1161
Score = 40.4 bits (93), Expect = 0.019
Identities = 29/101 (28%), Positives = 53/101 (51%), Gaps = 15/101 (14%)
Query: 14 VQFLKVQDDSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVA 73
+Q L + D S ++ + + D EEE+E+E ++ +++ E+ D D+ A
Sbjct: 119 LQALLISDLSNVEGKTGNATDEEEEEEEEEEEGEEEEEEEEDDDDDD-------DEDSGA 171
Query: 74 VVEIDRQQYHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDEL 114
++ D ++ G+ DDEE+ DDE DDDD + ++EL
Sbjct: 172 EIQDDDEE-----GF---DDEEEFDDDEHDDDDLDNEENEL 204
>emb|CAA63696.1| NRD2 convertase [Rattus sp.]
Length = 1229
Score = 40.4 bits (93), Expect = 0.019
Identities = 29/101 (28%), Positives = 53/101 (51%), Gaps = 15/101 (14%)
Query: 14 VQFLKVQDDSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVA 73
+Q L + D S ++ + + D EEE+E+E ++ +++ E+ D D+ A
Sbjct: 119 LQALLISDLSNVEGKTGNATDEEEEEEEEEEEGEEEEEEEEDDDDDD-------DEDSGA 171
Query: 74 VVEIDRQQYHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDEL 114
++ D ++ G+ DDEE+ DDE DDDD + ++EL
Sbjct: 172 EIQDDDEE-----GF---DDEEEFDDDEHDDDDLDNEENEL 204
>gb|AAA29605.1| glutamic acid-rich protein [Plasmodium falciparum]
gi|120943|sp|P13816|GARP_PLAFF Glutamic acid-rich
protein precursor gi|627054|pir||A54514 glutamic
acid-rich protein precursor - malaria parasite
(Plasmodium falciparum)
Length = 678
Score = 40.4 bits (93), Expect = 0.019
Identities = 30/97 (30%), Positives = 51/97 (51%), Gaps = 2/97 (2%)
Query: 18 KVQDDSL-LQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVE 76
+VQ++S +Q + +V + EEE+E+E ++ +++ EE + + D + E
Sbjct: 557 EVQEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEDEDEEDEDDAE 616
Query: 77 IDRQQYHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDE 113
D A EDD+E+ DDEEDDD+ D D+E
Sbjct: 617 EDEDDAEEDEDDAEEDDDEED-DDEEDDDEDEDEDEE 652
Score = 31.6 bits (70), Expect = 8.6
Identities = 18/81 (22%), Positives = 42/81 (51%), Gaps = 12/81 (14%)
Query: 33 VDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAYED 92
VD EE++++E + Q+++ V+E D + V E + ++ E+
Sbjct: 545 VDKEEDKKEESKEVQEESKEVQE------------DEEEVEEDEEEEEEEEEEEEEEEEE 592
Query: 93 DEEQVVDDEEDDDDRCDLDDE 113
+EE+ ++EE+++D + D++
Sbjct: 593 EEEEEEEEEEEEEDEDEEDED 613
>gb|AAG14291.1| glutamic acid-rich protein [Plasmodium falciparum]
Length = 682
Score = 40.4 bits (93), Expect = 0.019
Identities = 31/96 (32%), Positives = 46/96 (47%), Gaps = 16/96 (16%)
Query: 18 KVQDDSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEI 77
+VQ+D E V+ + EEE+E+E ++ +++ VEE D D E
Sbjct: 568 EVQEDE----EEVEEDEEEEEEEEEEEEEEEEEDEVEE------------DEDDAEEDED 611
Query: 78 DRQQYHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDE 113
D ++ G +D EE D EEDDDD + DDE
Sbjct: 612 DAEEDEDDAGEDEDDAEEDEDDAEEDDDDAEEDDDE 647
Score = 37.7 bits (86), Expect = 0.12
Identities = 26/84 (30%), Positives = 40/84 (46%), Gaps = 9/84 (10%)
Query: 31 DVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAY 90
D + EEE+E+E ++ ++D +E D D E D ++
Sbjct: 579 DEEEEEEEEEEEEEEEEEDEVEEDEDDAEEDEDDAEEDEDDAGEDEDDAEED-------- 630
Query: 91 EDDEEQVVDD-EEDDDDRCDLDDE 113
EDD E+ DD EEDDD+ D++DE
Sbjct: 631 EDDAEEDDDDAEEDDDEEEDVEDE 654
>dbj|BAD21392.1| mFLJ00158 protein [Mus musculus]
Length = 236
Score = 40.0 bits (92), Expect = 0.024
Identities = 29/85 (34%), Positives = 44/85 (51%), Gaps = 14/85 (16%)
Query: 36 EEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAYEDDEE 95
EEE+EQE +++Q++ EE + + + V D V E D + +EE
Sbjct: 40 EEEEEQEQEEHQEEEQEEEEEE---ENQKVFQDQMEADVEESDDVE-----------EEE 85
Query: 96 QVVDDEEDDDDRCDLDDELVPWEVG 120
+V D+EED+DD D D+E EVG
Sbjct: 86 EVDDEEEDEDDDYDDDEEEDRMEVG 110
>gb|AAH91633.1| Unknown (protein for MGC:98482) [Xenopus laevis]
Length = 946
Score = 40.0 bits (92), Expect = 0.024
Identities = 29/85 (34%), Positives = 44/85 (51%), Gaps = 14/85 (16%)
Query: 36 EEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAYEDDEE 95
EEE+EQE +++Q++ EE + + + V D V E D + +EE
Sbjct: 750 EEEEEQEQEEHQEEEQEEEEEE---ENQKVFQDQMEADVEESDDVE-----------EEE 795
Query: 96 QVVDDEEDDDDRCDLDDELVPWEVG 120
+V D+EED+DD D D+E EVG
Sbjct: 796 EVDDEEEDEDDDYDDDEEEDRMEVG 820
>gb|AAP31904.1| histidine-rich calcium binding protein [Rattus norvegicus]
gi|31077132|ref|NP_852034.1| histidine rich calcium
binding protein [Rattus norvegicus]
Length = 755
Score = 40.0 bits (92), Expect = 0.024
Identities = 25/86 (29%), Positives = 41/86 (47%)
Query: 21 DDSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQ 80
DD + E + + EEE+E+E ++ + D E + HQ + D E D
Sbjct: 290 DDDDEEEEEEEEEEEEEEEEEEEEEEEDDDDDSTEHRHRHQTQGHRKGKDEDESDEDDHV 349
Query: 81 QYHYLHGYAYEDDEEQVVDDEEDDDD 106
H G+ EDD++ DD++DD+D
Sbjct: 350 TRHGRRGHEDEDDDDDDDDDDDDDND 375
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 314,404,262
Number of Sequences: 2540612
Number of extensions: 14054969
Number of successful extensions: 123641
Number of sequences better than 10.0: 856
Number of HSP's better than 10.0 without gapping: 285
Number of HSP's successfully gapped in prelim test: 607
Number of HSP's that attempted gapping in prelim test: 109031
Number of HSP's gapped (non-prelim): 5623
length of query: 167
length of database: 863,360,394
effective HSP length: 118
effective length of query: 49
effective length of database: 563,568,178
effective search space: 27614840722
effective search space used: 27614840722
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0258a.2


