

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0247.8
(170 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
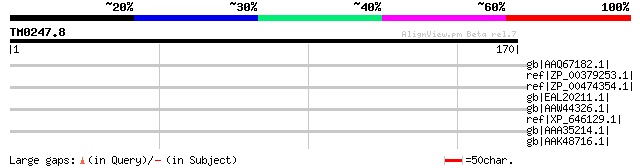
Score E
Sequences producing significant alignments: (bits) Value
gb|AAQ67182.1| CAD [Leptopeza sp. NCSU-99071981] 33 2.4
ref|ZP_00379253.1| COG0654: 2-polyprenyl-6-methoxyphenol hydroxy... 33 3.1
ref|ZP_00474354.1| Thiolase [Chromohalobacter salexigens DSM 304... 33 4.0
gb|EAL20211.1| hypothetical protein CNBF0230 [Cryptococcus neofo... 33 4.0
gb|AAW44326.1| conserved hypothetical protein [Cryptococcus neof... 33 4.0
ref|XP_646129.1| putative Nek family protein kinase [Dictyosteli... 33 4.0
gb|AAA35214.1| protein kinase 32 9.0
gb|AAK48716.1| dissimilatory sulfite reductase alpha subunit [su... 32 9.0
>gb|AAQ67182.1| CAD [Leptopeza sp. NCSU-99071981]
Length = 1289
Score = 33.5 bits (75), Expect = 2.4
Identities = 21/69 (30%), Positives = 36/69 (51%), Gaps = 2/69 (2%)
Query: 94 SGVSSIHKVSRLSEASRLIPWMANKLLNERRSTNILRTYQTPQHLDPTTLLVFPKLWFDF 153
+ + + + V +L E +++ PW NK+ N N+L T +L+ TTLL KL F
Sbjct: 765 AAIKANYTVEKLYELTKIDPWFLNKMKNIIDYLNLLEV--TGNNLNRTTLLEAKKLGFSD 822
Query: 154 QCLPSRLLS 162
+ + S + S
Sbjct: 823 RQIASAIKS 831
>ref|ZP_00379253.1| COG0654: 2-polyprenyl-6-methoxyphenol hydroxylase and related
FAD-dependent oxidoreductases [Brevibacterium linens
BL2]
Length = 625
Score = 33.1 bits (74), Expect = 3.1
Identities = 19/61 (31%), Positives = 28/61 (45%), Gaps = 3/61 (4%)
Query: 112 IPWMANKLLNERRSTNILRTYQTPQHLDPTTLLVFPKLWFDFQCLPSRLL---SLLKDFE 168
I W ++L+ R S +LRTY + + L+ F K W P L + L+DF
Sbjct: 363 ISWKLGQVLSGRSSQELLRTYSAERQVTAKNLIDFDKEWSTLMATPQEDLPDETYLQDFY 422
Query: 169 V 169
V
Sbjct: 423 V 423
>ref|ZP_00474354.1| Thiolase [Chromohalobacter salexigens DSM 3043]
gi|67518355|gb|EAM22330.1| Thiolase [Chromohalobacter
salexigens DSM 3043]
Length = 401
Score = 32.7 bits (73), Expect = 4.0
Identities = 17/62 (27%), Positives = 29/62 (46%)
Query: 50 VQPLTMTLRVINTLTKSHASVKFHALTRSHASVEFHAFHKVSRLSGVSSIHKVSRLSEAS 109
++ T+ R +N L K+H V+ T + + +FH + V S HK R ++
Sbjct: 140 IEDTTIGWRFVNPLMKNHYGVESMPETAENVAEQFHVSREDQDAFAVRSQHKTERAQQSG 199
Query: 110 RL 111
RL
Sbjct: 200 RL 201
>gb|EAL20211.1| hypothetical protein CNBF0230 [Cryptococcus neoformans var.
neoformans B-3501A]
Length = 2094
Score = 32.7 bits (73), Expect = 4.0
Identities = 30/120 (25%), Positives = 54/120 (45%), Gaps = 11/120 (9%)
Query: 6 AVISQTTSPTSLDTSPRVLAKLMSLKAYPKVKSILRQNTIIIKRVQPLTMTLRVINTLTK 65
A +S T+PT D R AK +++ P T+ +K ++ L ++ +
Sbjct: 1753 ADLSDVTAPTRPDVPERP-AKATAIEKGP-------MGTVAVKVIEQLNDEVQRLKDALA 1804
Query: 66 SHASVKFHALTRSHASVEFHAFHKVSRLSGVSSIHKVSRLSEASRLIPWMANKLLNERRS 125
++A TR+ +VEFHA + L S+ S++S+ RLI + + +R S
Sbjct: 1805 ANAEYHQQLRTRNAGTVEFHA---LVNLVEQQSVIPFSQISQVQRLIDELYPTVTKQRDS 1861
>gb|AAW44326.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58268954|ref|XP_571633.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 2094
Score = 32.7 bits (73), Expect = 4.0
Identities = 30/120 (25%), Positives = 54/120 (45%), Gaps = 11/120 (9%)
Query: 6 AVISQTTSPTSLDTSPRVLAKLMSLKAYPKVKSILRQNTIIIKRVQPLTMTLRVINTLTK 65
A +S T+PT D R AK +++ P T+ +K ++ L ++ +
Sbjct: 1753 ADLSDVTAPTRPDVPERP-AKATAIEKGP-------MGTVAVKVIEQLNDEVQRLKDALA 1804
Query: 66 SHASVKFHALTRSHASVEFHAFHKVSRLSGVSSIHKVSRLSEASRLIPWMANKLLNERRS 125
++A TR+ +VEFHA + L S+ S++S+ RLI + + +R S
Sbjct: 1805 ANAEYHQQLRTRNAGTVEFHA---LVNLVEQQSVIPFSQISQVQRLIDELYPTVTKQRDS 1861
>ref|XP_646129.1| putative Nek family protein kinase [Dictyostelium discoideum]
gi|60474227|gb|EAL72164.1| putative protein
serine/threonine kinase [Dictyostelium discoideum]
Length = 737
Score = 32.7 bits (73), Expect = 4.0
Identities = 28/114 (24%), Positives = 46/114 (39%), Gaps = 11/114 (9%)
Query: 9 SQTTSPTSLDTSPRVLAKLMSLKAYPKVKSILRQNTIIIKRVQPLTMTL-------RVIN 61
S +SP S P + S K+ K K L++ I+K LT +
Sbjct: 221 SSPSSPLSPQQHPVTSPQRKSSKSERKKKCSLKERKCIVKMTVDLTKKIFKKDNSNHTTA 280
Query: 62 TLTKSHASVKFHALTRSHASVEFHAFHKVSRLSGVSSIHKVSRLSEASRLIPWM 115
T + + + + S+ ++E H H GV SI+ L E ++LI W+
Sbjct: 281 ATTTTTTNTTHSSSSSSNLNIEEHVIHSNEIKKGVDSIY----LIEQTQLIEWL 330
>gb|AAA35214.1| protein kinase
Length = 1455
Score = 31.6 bits (70), Expect = 9.0
Identities = 20/98 (20%), Positives = 49/98 (49%), Gaps = 3/98 (3%)
Query: 36 VKSILRQNTIIIKRVQPLTMTLRVINTLTKSHASVKFHALTRSHASVEFHAFHKVSRLSG 95
++SI+ T +K ++ L + + ++ K V + + + A ++ +
Sbjct: 433 IRSIVSTATKPVKNLELLAVFAQFVSDENKIDRVVPYFVCCFEDSDQDVQALSLLTLIQV 492
Query: 96 VSSIHKVSRLSE---ASRLIPWMANKLLNERRSTNILR 130
++S+ K+++L+E L+P + L++ R++TN LR
Sbjct: 493 LTSVRKLNQLNENIFVDYLLPRLKRLLISNRQNTNYLR 530
>gb|AAK48716.1| dissimilatory sulfite reductase alpha subunit [sulfate-reducing
bacterium AK-01]
Length = 351
Score = 31.6 bits (70), Expect = 9.0
Identities = 26/108 (24%), Positives = 44/108 (40%), Gaps = 1/108 (0%)
Query: 4 CNAVISQTTSPTSLDTSPRVLAKLMSLKAYPKVKSILRQNTIIIKRVQPLTMTLRVINTL 63
C A +S+T+S +SL +P + + P + + +I R PLT+ + T
Sbjct: 143 CTARLSRTSSSSSLTAAPTAAWLPLPVPTCPSSEPGATTSALIRMR-SPLTLAANIRPTP 201
Query: 64 TKSHASVKFHALTRSHASVEFHAFHKVSRLSGVSSIHKVSRLSEASRL 111
+ + H +R +S H H+V S S+ AS L
Sbjct: 202 ALTKTATGEHLTSRRKSSTCAHRLHEVQAASWKSTTKSAPAACIASTL 249
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.322 0.131 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 236,958,073
Number of Sequences: 2540612
Number of extensions: 7664151
Number of successful extensions: 19429
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 19427
Number of HSP's gapped (non-prelim): 9
length of query: 170
length of database: 863,360,394
effective HSP length: 119
effective length of query: 51
effective length of database: 561,027,566
effective search space: 28612405866
effective search space used: 28612405866
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0247.8


