

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
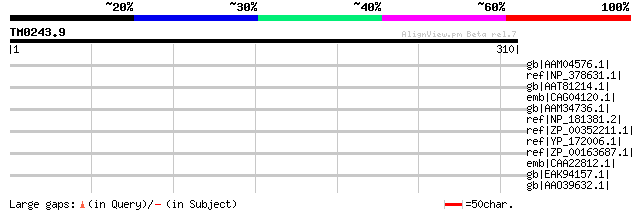
Query= TM0243.9
(310 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM04576.1| predicted protein [Methanosarcina acetivorans str... 39 0.25
ref|NP_378631.1| hypothetical protein ST2622 [Sulfolobus tokodai... 37 0.74
gb|AAT81214.1| endo-chitinase [Microbulbifer hydrolyticus] 36 1.3
emb|CAG04120.1| unnamed protein product [Tetraodon nigroviridis] 35 2.8
gb|AAM34736.1| WRKY transcription factor 33 [Arabidopsis thalian... 34 4.8
ref|NP_181381.2| WRKY family transcription factor [Arabidopsis t... 34 4.8
ref|ZP_00352211.1| hypothetical protein Krad07004920 [Kineococcu... 34 4.8
ref|YP_172006.1| putative zinc-binding oxidoreductase [Synechoco... 34 6.2
ref|ZP_00163687.1| COG0604: NADPH:quinone reductase and related ... 34 6.2
emb|CAA22812.1| SPBC646.08c [Schizosaccharomyces pombe] gi|19112... 33 8.2
gb|EAK94157.1| hypothetical protein CaO19.9926 [Candida albicans... 33 8.2
gb|AAO39632.1| AT28824p [Drosophila melanogaster] 33 8.2
>gb|AAM04576.1| predicted protein [Methanosarcina acetivorans str. C2A]
gi|20090021|ref|NP_616096.1| hypothetical protein MA1155
[Methanosarcina acetivorans C2A]
Length = 1381
Score = 38.5 bits (88), Expect = 0.25
Identities = 29/103 (28%), Positives = 43/103 (41%), Gaps = 14/103 (13%)
Query: 155 IREKGVLEGAFQTLHSGYFRLFPFSFFSLPLSPPLTSTISSSTGTISSSSTTKKKTSRGV 214
+ EKG+ GAF ++ SG P + F P S + G IS+ T
Sbjct: 943 LEEKGLSTGAFDSVTSG-----PMTLFDNPAMEKAASALGQDMGIISTIKVT-------- 989
Query: 215 GRSEELQIWCESKGRVPDLVRQWLY-GSDPALNGEEQWWRNLG 256
G+ E W E+ + D +LY D L GE +W ++G
Sbjct: 990 GQPESKYSWTENLTLIVDQKPNYLYHDPDFDLRGEYEWADSMG 1032
>ref|NP_378631.1| hypothetical protein ST2622 [Sulfolobus tokodaii str. 7]
gi|15623753|dbj|BAB67740.1| 702aa long hypothetical
protein [Sulfolobus tokodaii str. 7]
Length = 702
Score = 37.0 bits (84), Expect = 0.74
Identities = 19/60 (31%), Positives = 33/60 (54%)
Query: 172 YFRLFPFSFFSLPLSPPLTSTISSSTGTISSSSTTKKKTSRGVGRSEELQIWCESKGRVP 231
Y +++ SF +L +PP++S ++ T +SSSTT TS + QI+ ++ VP
Sbjct: 613 YNQIYAISFNNLISTPPISSNTTTIPTTTTSSSTTFTYTSTSISSKTTTQIYSSTESAVP 672
>gb|AAT81214.1| endo-chitinase [Microbulbifer hydrolyticus]
Length = 590
Score = 36.2 bits (82), Expect = 1.3
Identities = 26/85 (30%), Positives = 37/85 (42%), Gaps = 13/85 (15%)
Query: 190 TSTISSSTGTISSSSTTKKKTSRGVGRSEELQIW----------CESKGRVPDLVRQWLY 239
+S+ SSS+ + SSSS++ +S G L +W V + W
Sbjct: 107 SSSSSSSSSSSSSSSSSSGGSSSGSTSCNNLPVWDATTVYVGGNAVQHSSVKYTAKWWTQ 166
Query: 240 GSDPALNGEEQWWRNLGVSGSVSGS 264
G +P+ GE WRN GS SGS
Sbjct: 167 GDNPSQGGEYGVWRN---DGSCSGS 188
>emb|CAG04120.1| unnamed protein product [Tetraodon nigroviridis]
Length = 427
Score = 35.0 bits (79), Expect = 2.8
Identities = 29/84 (34%), Positives = 37/84 (43%), Gaps = 5/84 (5%)
Query: 207 KKKTSRGVGRSEELQIWCESKGRVPDLVRQWLYGSDPALNGEEQWWRNLGVSGSVSGSLL 266
+K +R R EE Q+ +G P R+ G D AL+G E R GV G V L
Sbjct: 157 RKNEARPGRRGEERQV---HRGAGPARARRQRDGRDAALHGTETGER--GVKGGVQPRSL 211
Query: 267 HLLHRLCFSLSESGAWLRKEGCDG 290
+ R S S G W +EG G
Sbjct: 212 FVCSRTLKSQSCCGRWRPEEGSSG 235
>gb|AAM34736.1| WRKY transcription factor 33 [Arabidopsis thaliana]
gi|20197246|gb|AAM14994.1| putative WRKY-type DNA
binding protein [Arabidopsis thaliana]
gi|29839571|sp|Q8S8P5|WRK33_ARATH Probable WRKY
transcription factor 33 (WRKY DNA-binding protein 33)
Length = 512
Score = 34.3 bits (77), Expect = 4.8
Identities = 29/103 (28%), Positives = 52/103 (50%), Gaps = 10/103 (9%)
Query: 157 EKGVLEGAFQTLHSGYFRLFPFSFF--SLPLSPPLTSTISSSTGTISSSSTTKKKTSRGV 214
+KG+ EG ++ F LF FSF S +S P T+T +++T T ++SS + + +
Sbjct: 91 QKGINEG--DKSNNNNFNLFDFSFHTQSSGVSAPTTTTTTTTTTTTTNSSIFQSQEQQKK 148
Query: 215 GRSEELQIWCESKGRVPDLVRQWLYGSDPALNGEEQW-WRNLG 256
+SE+ W +++ R + + Y GE+ + WR G
Sbjct: 149 NQSEQ---WSQTETRPNN--QAVSYNGREQRKGEDGYNWRKYG 186
>ref|NP_181381.2| WRKY family transcription factor [Arabidopsis thaliana]
Length = 519
Score = 34.3 bits (77), Expect = 4.8
Identities = 29/103 (28%), Positives = 52/103 (50%), Gaps = 10/103 (9%)
Query: 157 EKGVLEGAFQTLHSGYFRLFPFSFF--SLPLSPPLTSTISSSTGTISSSSTTKKKTSRGV 214
+KG+ EG ++ F LF FSF S +S P T+T +++T T ++SS + + +
Sbjct: 98 QKGINEG--DKSNNNNFNLFDFSFHTQSSGVSAPTTTTTTTTTTTTTNSSIFQSQEQQKK 155
Query: 215 GRSEELQIWCESKGRVPDLVRQWLYGSDPALNGEEQW-WRNLG 256
+SE+ W +++ R + + Y GE+ + WR G
Sbjct: 156 NQSEQ---WSQTETRPNN--QAVSYNGREQRKGEDGYNWRKYG 193
>ref|ZP_00352211.1| hypothetical protein Krad07004920 [Kineococcus radiotolerans
SRS30216]
Length = 397
Score = 34.3 bits (77), Expect = 4.8
Identities = 24/70 (34%), Positives = 34/70 (48%), Gaps = 7/70 (10%)
Query: 92 GDVQGKVEEADSRARRSGRSSRWFASHQFHIAVGDAIKDVMVLEGERRRGGGRIPPARRP 151
G V G+V E R RR ++ H + G D+ V+ G+RR GGGR+ P R
Sbjct: 114 GQVGGRVHER--RQRRGALGAQ--VGHDGDVRPGG---DLHVVRGDRRLGGGRVRPGRVD 166
Query: 152 GCFIREKGVL 161
G + +G L
Sbjct: 167 GEVVAARGQL 176
>ref|YP_172006.1| putative zinc-binding oxidoreductase [Synechococcus elongatus PCC
6301] gi|56686264|dbj|BAD79486.1| putative zinc-binding
oxidoreductase [Synechococcus elongatus PCC 6301]
Length = 338
Score = 33.9 bits (76), Expect = 6.2
Identities = 29/103 (28%), Positives = 43/103 (41%), Gaps = 11/103 (10%)
Query: 53 LLERLKFIVALCSSLDFIFKPLATHTNWKFFD------CLSIFFTGDVQGKVEEADSRAR 106
+LE+ V L L I P A TNWK L + T +Q E + +
Sbjct: 231 VLEQTFPAVRLYGDLVTILAPDAA-TNWKVARDRNLRISLELMLTPQLQRVTEARQHQTQ 289
Query: 107 RSGRSSRWFASHQFHIAVGDAIKDVMVLEG----ERRRGGGRI 145
+SRWF Q HI +G + + + ER++G G+I
Sbjct: 290 ILTTASRWFDQEQLHIHIGQVLPISEISQAHQFLERQQGAGKI 332
>ref|ZP_00163687.1| COG0604: NADPH:quinone reductase and related Zn-dependent
oxidoreductases [Synechococcus elongatus PCC 7942]
Length = 338
Score = 33.9 bits (76), Expect = 6.2
Identities = 29/103 (28%), Positives = 43/103 (41%), Gaps = 11/103 (10%)
Query: 53 LLERLKFIVALCSSLDFIFKPLATHTNWKFFD------CLSIFFTGDVQGKVEEADSRAR 106
+LE+ V L L I P A TNWK L + T +Q E + +
Sbjct: 231 VLEQTFPAVRLYGDLVTILAPDAA-TNWKVARDRNLRISLELMLTPQLQRVTEARQHQTQ 289
Query: 107 RSGRSSRWFASHQFHIAVGDAIKDVMVLEG----ERRRGGGRI 145
+SRWF Q HI +G + + + ER++G G+I
Sbjct: 290 ILTTASRWFDQEQLHIHIGQVLPISEISQAHQFLERQQGAGKI 332
>emb|CAA22812.1| SPBC646.08c [Schizosaccharomyces pombe]
gi|19112158|ref|NP_595366.1| oxysterol binding protein
[Schizosaccharomyces pombe 972h-] gi|7492679|pir||T40584
probable involvement in ergosterol biosynthesis -
fission yeast (Schizosaccharomyces pombe)
Length = 516
Score = 33.5 bits (75), Expect = 8.2
Identities = 51/192 (26%), Positives = 83/192 (42%), Gaps = 16/192 (8%)
Query: 37 EYYQKLDGTKIWVVLMLLER---LKFIVALCSSLDF-IFKPLATHTNWKFFDCLSIFFTG 92
E YQK +G K +VL +L++ +K I +L SL + +P+ W + D F
Sbjct: 31 EGYQKEEG-KFKLVLSILKQCIGVKDIASLRFSLPAQLLEPVGNLEYWNYVDRPDYFAV- 88
Query: 93 DVQGKVEEADSRARRSGRSSRWFASHQFHIAVGDAIKDVMVLEGERRRGGGRI--PPARR 150
+ ++D R RW+ + G +K + GE R + P R
Sbjct: 89 -----MGDSDDELERMLGVLRWWFTKDLRFVRGRVVKPYNSVLGEFFRCKWVVTDPTVRE 143
Query: 151 PGCFIREKGVLEGAFQTLHSGYFRLFPFSFFSLPLSPPLTSTIS-SSTGTISSSSTTKKK 209
+ L ++T +S + FP P + TS+ S +ST T SS+ T+KKK
Sbjct: 144 DHTLDPDSSQLP-TYKTEYSETTK-FPLGKSYRPKASRTTSSQSVASTMTKSSTKTSKKK 201
Query: 210 TSRGVGRSEELQ 221
+S+ +SE Q
Sbjct: 202 SSKKNSKSESNQ 213
>gb|EAK94157.1| hypothetical protein CaO19.9926 [Candida albicans SC5314]
gi|46434704|gb|EAK94106.1| hypothetical protein
CaO19.2390 [Candida albicans SC5314]
Length = 125
Score = 33.5 bits (75), Expect = 8.2
Identities = 24/71 (33%), Positives = 34/71 (47%), Gaps = 10/71 (14%)
Query: 45 TKIWVVLMLLERLKFIVALCSSL-------DFIFKPLATHTNWKFFDCLSI---FFTGDV 94
T + ++L+LL + C L + IF+PL TN FFD + + FFTGD
Sbjct: 17 TVLLLLLLLLLWESGVCGFCRCLLRYDGVSNTIFRPLIGVTNSLFFDLVGVSIMFFTGDC 76
Query: 95 QGKVEEADSRA 105
G + AD A
Sbjct: 77 GGGIANADGDA 87
>gb|AAO39632.1| AT28824p [Drosophila melanogaster]
Length = 642
Score = 33.5 bits (75), Expect = 8.2
Identities = 31/112 (27%), Positives = 47/112 (41%), Gaps = 12/112 (10%)
Query: 203 SSTTKKKTSRGVGRSEELQIWCESKGRVPDLVRQWLYGSDPALNGEEQWWRNLGVSGSVS 262
SS K +R G+ + C+S G++ ++ DP L+GE + W S +
Sbjct: 1 SSKFNIKLNRSFGQKTSATM-CQSGGQMLCEIKNMALSGDPELDGEIRKWIRWDKCASTA 59
Query: 263 GSLLH---------LLHRLCFSLSESGAWLRKEGCDGDGFESRLELFVVFVA 305
++ L RLC +S A LR C GF+S EL V+ A
Sbjct: 60 CQIMDAVRAKDWDTLRKRLCTRISFGTAGLR--ACMRAGFDSMNELVVIQTA 109
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.323 0.139 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 579,258,150
Number of Sequences: 2540612
Number of extensions: 24536901
Number of successful extensions: 84740
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 84729
Number of HSP's gapped (non-prelim): 18
length of query: 310
length of database: 863,360,394
effective HSP length: 127
effective length of query: 183
effective length of database: 540,702,670
effective search space: 98948588610
effective search space used: 98948588610
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0243.9


