

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.7
(399 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
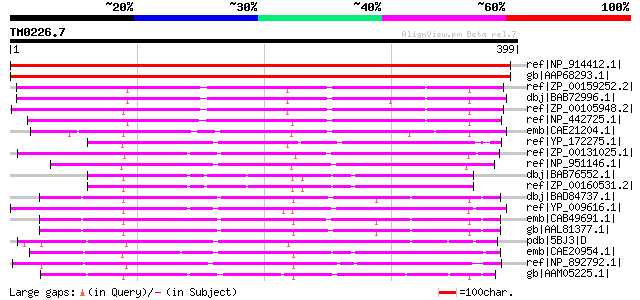
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_914412.1| putative aspartate aminotransferase [Oryza sati... 618 e-175
gb|AAP68293.1| At1g80360 [Arabidopsis thaliana] gi|20260520|gb|A... 585 e-166
ref|ZP_00159252.2| COG0436: Aspartate/tyrosine/aromatic aminotra... 225 2e-57
dbj|BAB72996.1| alr1039 [Nostoc sp. PCC 7120] gi|17228534|ref|NP... 220 5e-56
ref|ZP_00105948.2| COG0436: Aspartate/tyrosine/aromatic aminotra... 218 2e-55
ref|NP_442725.1| aspartate aminotransferase [Synechocystis sp. P... 199 2e-49
emb|CAE21204.1| Aminotransferases class-I [Prochlorococcus marin... 189 2e-46
ref|YP_172275.1| aspartate aminotransferase [Synechococcus elong... 154 4e-36
ref|ZP_00131025.1| COG0436: Aspartate/tyrosine/aromatic aminotra... 151 4e-35
ref|NP_951146.1| aminotransferase, classes I and II [Geobacter s... 147 4e-34
dbj|BAB76552.1| aspartate aminotransferase [Nostoc sp. PCC 7120]... 147 6e-34
ref|ZP_00160531.2| COG0436: Aspartate/tyrosine/aromatic aminotra... 146 1e-33
dbj|BAD84737.1| aromatic aminotransferase [Thermococcus kodakare... 144 4e-33
ref|YP_009616.1| aromatic aminotransferase [Desulfovibrio vulgar... 144 6e-33
emb|CAB49691.1| aspC-1 aspartate aminotransferase (EC 2.6.1.1) [... 143 8e-33
gb|AAL81377.1| aspartate transaminase [Pyrococcus furiosus DSM 3... 143 1e-32
pdb|5BJ3|D Chain D, Thermus Thermophilus Aspartate Aminotransfer... 142 1e-32
emb|CAE20954.1| Aminotransferases class-I [Prochlorococcus marin... 142 2e-32
ref|NP_892792.1| Aminotransferases class-I [Prochlorococcus mari... 141 3e-32
gb|AAM05225.1| aspartate aminotransferase [Methanosarcina acetiv... 140 7e-32
>ref|NP_914412.1| putative aspartate aminotransferase [Oryza sativa (japonica
cultivar-group)] gi|15289767|dbj|BAB63467.1| putative
aspartate aminotransferase [Oryza sativa (japonica
cultivar-group)]
Length = 394
Score = 618 bits (1593), Expect = e-175
Identities = 285/394 (72%), Positives = 355/394 (89%)
Query: 1 MGSYANLARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPS 60
MGS+ LARRA+ETE P+MV+MQE+LRG K+ +SLAQGVVYWQPP+ A++K+KE+VWEPS
Sbjct: 1 MGSFGRLARRAVETEAPVMVKMQELLRGNKDVMSLAQGVVYWQPPEAAMNKIKEIVWEPS 60
Query: 61 VSRYGADEGLPELRAALVKKLRQENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFA 120
+S+YG+D+GLPELR AL++KLR+EN L KSS+MVT+GANQAFVN+VLTLCDAGD+VVMFA
Sbjct: 61 ISKYGSDDGLPELREALLEKLRRENKLTKSSIMVTSGANQAFVNVVLTLCDAGDAVVMFA 120
Query: 121 PYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGT 180
PYYFN+YMSFQMTG+T+I VG +P+TL+PDVDWLE++L E+ P+PKLV+VVNPGNP+G
Sbjct: 121 PYYFNSYMSFQMTGVTDILVGASNPETLHPDVDWLEKVLQENNPIPKLVSVVNPGNPSGA 180
Query: 181 YIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCVEGNHIVNIFSFSKAYGMMGW 240
+IP+ +L+RI+ LC+NAG+WL+VDNTYEYFM+DG++H C+EGNHIVN+FSFSKAYGMMGW
Sbjct: 181 FIPKPMLERISELCRNAGAWLVVDNTYEYFMYDGMEHYCLEGNHIVNLFSFSKAYGMMGW 240
Query: 241 RVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNRE 300
RVGYIA+P+E +GL AQLLKVQDNIPICASII Q LAL++LE GPEW+RERV+ L KNRE
Sbjct: 241 RVGYIAHPNEADGLHAQLLKVQDNIPICASIIGQRLALYALEAGPEWIRERVRDLVKNRE 300
Query: 301 IVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN 360
++ EA+SPLG+ SVKGGEGAIYL+AKLP+ DDF+VVRWLAN+HG+AVIPGSA G G
Sbjct: 301 LLMEAMSPLGKDSVKGGEGAIYLWAKLPEKCSDDFEVVRWLANKHGVAVIPGSASGGPGY 360
Query: 361 LRISFGGLTESDCRAAAERLKKGLEELVANGLVQ 394
+R+SFGGL ESD R AAERL++GL+ELV G+VQ
Sbjct: 361 IRVSFGGLKESDTRLAAERLRRGLQELVTEGMVQ 394
>gb|AAP68293.1| At1g80360 [Arabidopsis thaliana] gi|20260520|gb|AAM13158.1|
putative aspartate aminotransferase [Arabidopsis
thaliana] gi|15220119|ref|NP_178152.1| aminotransferase
class I and II family protein [Arabidopsis thaliana]
gi|12324981|gb|AAG52437.1| putative aspartate
aminotransferase; 38163-36256 [Arabidopsis thaliana]
gi|25406657|pir||C96835 hypothetical protein F5I6.11
[imported] - Arabidopsis thaliana
Length = 394
Score = 585 bits (1508), Expect = e-166
Identities = 274/394 (69%), Positives = 335/394 (84%)
Query: 1 MGSYANLARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPS 60
MGS+ L+RR L T+MP+M Q++ ++ N +SLAQGVV+WQPP+ AL+KVKELVW+P
Sbjct: 1 MGSFGMLSRRTLGTDMPVMAQIRSLMAELTNPMSLAQGVVHWQPPQKALEKVKELVWDPI 60
Query: 61 VSRYGADEGLPELRAALVKKLRQENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFA 120
+S YG DEGLPELR AL+KKLR+EN L S VMVTAGANQAFVNLV+TLCDAGDSVVMF
Sbjct: 61 ISSYGPDEGLPELRQALLKKLREENKLTNSQVMVTAGANQAFVNLVITLCDAGDSVVMFE 120
Query: 121 PYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGT 180
PYYFN+YM+FQMTG+TNI VGPG DTLYPD DWLER L+ESKP PK+VTVVNPGNP+GT
Sbjct: 121 PYYFNSYMAFQMTGVTNIIVGPGQSDTLYPDADWLERTLSESKPTPKVVTVVNPGNPSGT 180
Query: 181 YIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCVEGNHIVNIFSFSKAYGMMGW 240
Y+PE LL+RIA +CK+AG WL+VDNTYEYFM+DGLKH CVEG+HIVN+FSFSK YGMMGW
Sbjct: 181 YVPEPLLKRIAQICKDAGCWLIVDNTYEYFMYDGLKHCCVEGDHIVNVFSFSKTYGMMGW 240
Query: 241 RVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNRE 300
R+GYIAY ++G + +L+K+QDNIPICA+IISQ LA+++LE G W+ ERV++L KNR+
Sbjct: 241 RLGYIAYSERLDGFATELVKIQDNIPICAAIISQRLAVYALEEGSGWITERVKSLVKNRD 300
Query: 301 IVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN 360
IV EAL PLG+ +VKGGEGAIYL+AKLP+G DDF VVRWLA+RHG+ VIPG A G+ G
Sbjct: 301 IVKEALEPLGKENVKGGEGAIYLWAKLPEGHRDDFKVVRWLAHRHGVVVIPGCASGSPGY 360
Query: 361 LRISFGGLTESDCRAAAERLKKGLEELVANGLVQ 394
LR+SFGGL E + RAAA RL+KG+EEL+ +G+V+
Sbjct: 361 LRVSFGGLQEVEMRAAAARLRKGIEELLHHGMVE 394
>ref|ZP_00159252.2| COG0436: Aspartate/tyrosine/aromatic aminotransferase [Anabaena
variabilis ATCC 29413]
Length = 391
Score = 225 bits (574), Expect = 2e-57
Identities = 130/394 (32%), Positives = 214/394 (53%), Gaps = 20/394 (5%)
Query: 6 NLARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYG 65
N R + PI+ + E+++ + +SL QGVV + PP A++ + + + +P+ + Y
Sbjct: 3 NFTSRMQAVQSPIIPVVGELIQNSPGTISLGQGVVSYSPPPEAIELLPKFLADPANNLYK 62
Query: 66 ADEGLPELRAALVKKLRQENNLHKSS---VMVTAGANQAFVNLVLTLCDAGDSVVMFAPY 122
A EG+P L AL KL N + ++ ++VTAG+N AF+N +L + GD +++ PY
Sbjct: 63 AVEGIPPLLNALTDKLSTFNKMEITTDNCIVVTAGSNMAFMNAILAITSPGDEIILNTPY 122
Query: 123 YFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYI 182
YFN M+ M G + V L P E I P + V ++P NPTG
Sbjct: 123 YFNHEMAIAMAGCRAVLVETDENYQLRP-----EAIAQAITPKTRAVVTISPNNPTGVVY 177
Query: 183 PESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKH-----SCVEGNHIVNIFSFSKAYGM 237
E LL+ + +C N G + + D YEYF +DG+KH + ++++S SKAYG
Sbjct: 178 CEDLLRNVNQICANCGIYHISDEAYEYFTYDGVKHVSPASFAGSSEYTISLYSLSKAYGF 237
Query: 238 MGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAK 297
WR+GY+ P L + KVQD I IC ++SQ+ AL +L+ PE++++ +Q +A+
Sbjct: 238 ASWRIGYMVIPKH---LLVAIKKVQDTILICPPVVSQYAALGALQAKPEYLQDHIQAIAQ 294
Query: 298 NREIVSEALSPL-GEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACG 356
R+IV + L+ L G ++ +GA Y+F K+ D F +V+ L + +AVIPG+ G
Sbjct: 295 VRKIVFDYLNQLQGLCTITPADGAFYVFLKV-HTQIDAFTLVKQLIQEYKVAVIPGTTFG 353
Query: 357 AAGN--LRISFGGLTESDCRAAAERLKKGLEELV 388
LR+++G L + + ERL +GL+ ++
Sbjct: 354 MESGCYLRVAYGALQKDTVQEGIERLVQGLKTIL 387
>dbj|BAB72996.1| alr1039 [Nostoc sp. PCC 7120] gi|17228534|ref|NP_485082.1|
hypothetical protein alr1039 [Nostoc sp. PCC 7120]
gi|25315786|pir||AD1936 hypothetical protein alr1039
[imported] - Nostoc sp. (strain PCC 7120)
Length = 398
Score = 220 bits (561), Expect = 5e-56
Identities = 132/404 (32%), Positives = 214/404 (52%), Gaps = 27/404 (6%)
Query: 6 NLARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYG 65
N R + PI+ + E+++ + +SL QGVV + PP A++ + + +P+ + Y
Sbjct: 3 NFTSRMQAVQSPIIPVVGELIQNSPGTISLGQGVVSYSPPPEAIELLPRFLADPANNLYK 62
Query: 66 ADEGLPELRAALVKKLRQENNLHKSS---VMVTAGANQAFVNLVLTLCDAGDSVVMFAPY 122
A EG+P L AL +KL NN+ ++ ++VTAG+N AF+N +L + GD +++ PY
Sbjct: 63 AVEGIPPLLNALTEKLSTFNNIEITTDNCIVVTAGSNMAFMNAILAITSPGDEIILNTPY 122
Query: 123 YFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYI 182
YFN M+ M G + V L P E I P + V ++P NPTG
Sbjct: 123 YFNHEMAITMAGCRAVLVETDENYQLCP-----EAIAQAITPKTRAVVTISPNNPTGVVY 177
Query: 183 PESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKH-----SCVEGNHIVNIFSFSKAYGM 237
E LL+ + +C N G + + D YEYF +DG+KH + ++++S SKAYG
Sbjct: 178 CEDLLRNVNQICANYGIYHISDEAYEYFTYDGVKHVSPASFAGSSEYTISLYSLSKAYGF 237
Query: 238 MGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAK 297
WR+GY+ P L + KVQD I IC ++SQ+ AL +L+ PE++++ + LA+
Sbjct: 238 ASWRIGYMVIPKH---LLVAIKKVQDTILICPPVVSQYAALGALQAKPEYLQDHIGALAQ 294
Query: 298 N-------REIVSEALSPL-GEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAV 349
R+IV + L L G ++ +GA Y+F K+ D F +V+ L + +AV
Sbjct: 295 PAVGIAQVRQIVFDYLKQLQGLCNITPADGAFYVFLKV-HTQIDAFALVKQLIQEYKVAV 353
Query: 350 IPGSACGAAGN--LRISFGGLTESDCRAAAERLKKGLEELVANG 391
IPG+ G LR+++G L + + ERL +GL+ ++ G
Sbjct: 354 IPGTTFGMENGCYLRVAYGALQKDTVKEGIERLVQGLKTILDTG 397
>ref|ZP_00105948.2| COG0436: Aspartate/tyrosine/aromatic aminotransferase [Nostoc
punctiforme PCC 73102]
Length = 437
Score = 218 bits (556), Expect = 2e-55
Identities = 123/397 (30%), Positives = 217/397 (53%), Gaps = 19/397 (4%)
Query: 2 GSYANLARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSV 61
G+ +L R + + PI+ + E+++ + +SL QGVV++ PP A++ + + + +P+
Sbjct: 48 GNMESLTSRMQDVQSPIIPVVGELIKNSPGTISLGQGVVHYNPPPEAIEFLSKFLADPAN 107
Query: 62 SRYGADEGLPELRAALVKKLRQENNLH---KSSVMVTAGANQAFVNLVLTLCDAGDSVVM 118
+ Y + EG+P L A+ KL+ N + ++ ++VTAG+N F+N +L + + GD +++
Sbjct: 108 NLYKSVEGIPPLLTAIAGKLQAFNGIEINGENCIVVTAGSNMGFMNAILAITNPGDEIIL 167
Query: 119 FAPYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPT 178
PYYFN M+ + G + V L P E I P + + ++P NPT
Sbjct: 168 NTPYYFNHEMAIAIAGCRAVLVATDENYQLRP-----EAIAQAITPKTRAIVTISPNNPT 222
Query: 179 GTYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKH----SCVEGNHIVNIFSFSKA 234
G E+ L+++ +C + + D Y+YF ++G+KH + + + +++FS SKA
Sbjct: 223 GVVYSEAALRQVNQICDTHRIYHISDEAYQYFTYNGVKHVSPGAFGKSKYTISLFSLSKA 282
Query: 235 YGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQT 294
YG WR+GY+ P L + KVQD I IC +ISQ+ AL +L+ E+++ +
Sbjct: 283 YGFASWRIGYMVIPKH---LFVAIKKVQDTILICPPVISQYAALGALQAKEEYLQSNIGA 339
Query: 295 LAKNREIVSEALSPL-GEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGS 353
+ + R++V ++L+ L G S+ GA Y F K+ D F++V+ L H +AVIPG+
Sbjct: 340 IFQVRQLVLDSLNRLEGLCSITPANGAFYFFLKV-NTQMDAFELVKRLIQEHKVAVIPGT 398
Query: 354 ACGAAGN--LRISFGGLTESDCRAAAERLKKGLEELV 388
G LR+++G L + + ERL +GLE +V
Sbjct: 399 TFGMDDGCYLRVAYGALQKETAKEGIERLVQGLETIV 435
>ref|NP_442725.1| aspartate aminotransferase [Synechocystis sp. PCC 6803]
gi|1001309|dbj|BAA10796.1| aspartate aminotransferase
[Synechocystis sp. PCC 6803] gi|7447812|pir||S75949
probable aspartate transaminase (EC 2.6.1.1) aspC
slr0036 [similarity] - Synechocystis sp. (strain PCC
6803)
Length = 389
Score = 199 bits (505), Expect = 2e-49
Identities = 122/384 (31%), Positives = 204/384 (52%), Gaps = 20/384 (5%)
Query: 15 EMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELR 74
+ P++ + + + + +SL QGV ++ PP V+E + + + +Y G+P L
Sbjct: 14 QSPVIPVVGQWIVDSPGTISLGQGVAFYPPPNEVAVAVRESLEQTPLHQYQPVAGIPSLI 73
Query: 75 AALVKKLRQENNLHKSS---VMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQ 131
+AL +KLR++N+++ SS V+VTAGAN F+N VL + + GD +++ PYYFN M+ +
Sbjct: 74 SALTEKLRRDNDINLSSDQAVVVTAGANMGFLNAVLAITEVGDEIILNTPYYFNHEMAVR 133
Query: 132 MTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIA 191
+ G + V L L+ I P + V ++P NPTG PE+ L+ +
Sbjct: 134 IAGCQPVLVPTDDQYQLQ-----LDLIAQAIAPRTRAVVTISPNNPTGAIYPEADLRAVN 188
Query: 192 NLCKNAGSWLLVDNTYEYFMFDGLKHSCVE-----GNHIVNIFSFSKAYGMMGWRVGYIA 246
LC+ G + + D Y+YF +D E G H ++++SFSKAYGM GWRVGY+
Sbjct: 189 QLCQERGIYHIHDEAYDYFAYDQTPIFSPEAMGDSGGHTISLYSFSKAYGMAGWRVGYMV 248
Query: 247 YPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEAL 306
P E L + K+QD IC ++SQ+ AL L +G + + + +A R+ + E L
Sbjct: 249 IPLE---LLLAVKKIQDTNLICPVVVSQYAALACLRVGKNYSAQFLPEMAACRQQLLETL 305
Query: 307 SPLGE-GSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN--LRI 363
L + + +GA Y ++ D ++V+ L + +AV+PGS G LRI
Sbjct: 306 GQLSDYCRLVVPQGAFYCLLEV-NSPLTDLELVKRLIDEFKVAVLPGSTFGVDSGCYLRI 364
Query: 364 SFGGLTESDCRAAAERLKKGLEEL 387
++G L ++ A RL++GL+ +
Sbjct: 365 AYGALRQATASVAIARLEQGLKSI 388
>emb|CAE21204.1| Aminotransferases class-I [Prochlorococcus marinus str. MIT 9313]
gi|33863300|ref|NP_894860.1| Aminotransferases class-I
[Prochlorococcus marinus str. MIT 9313]
Length = 404
Score = 189 bits (479), Expect = 2e-46
Identities = 130/391 (33%), Positives = 198/391 (50%), Gaps = 28/391 (7%)
Query: 17 PIMVQMQEVLRGAKNCVSLAQGVVYWQPP---KPALDKVKELVWEPSVSRYGADEGLPEL 73
P++ + E++ +SLAQG+V W PP K A++ L E S++RYG G P+L
Sbjct: 22 PVIPALNELVAKTPGTLSLAQGMVNWPPPIAVKLAMNNAL-LNQESSLNRYGPARGDPDL 80
Query: 74 RAALVKKLRQENNLH--KSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQ 131
+ +KL +N L +S VMVTAG+N AF + LCD GD V++ PYYFN +M+ Q
Sbjct: 81 LELIKQKLMMQNGLDLAESMVMVTAGSNMAFHAIAQVLCDPGDEVILPLPYYFNHFMAIQ 140
Query: 132 MTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIA 191
+ G + V G L P+ +E +T+ + + ++P NP+G P++LL I
Sbjct: 141 LAGGVPVPVNAG----LIPNPGLIEAAITKR---TRAIVTISPNNPSGIVFPQTLLAAIN 193
Query: 192 NLCKNAGSWLLVDNTYEYFMFDGLKHSCV-----EGNHIVNIFSFSKAYGMMGWRVGYIA 246
+C G + D YE F+F + H GNH V+++SFSKAYGM GWR+GY++
Sbjct: 194 RICAQHGLLHISDEAYEDFVFGDVPHWSPGSLPGAGNHTVSLYSFSKAYGMAGWRLGYMS 253
Query: 247 YPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEAL 306
P S L KVQD + IC Q A+ +L G +WVR+ V L +++ +
Sbjct: 254 VPIV---WSKALEKVQDTVLICPPRFCQRAAIAALLDGSDWVRQNVSQLMSRYQLLLKRF 310
Query: 307 SPLGEGS---VKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN--- 360
+ + + GA Y ++ G D ++R L + +A I G + G
Sbjct: 311 AASNDRPWRFLHEPNGAFYCLLEVDCGCNGD-TLMRQLVRDYRVATIGGCSFGFKNESCV 369
Query: 361 LRISFGGLTESDCRAAAERLKKGLEELVANG 391
LRIS G L ++ A RL+ G+ V G
Sbjct: 370 LRISVGMLEGAELIDAFNRLEAGMLNAVQQG 400
>ref|YP_172275.1| aspartate aminotransferase [Synechococcus elongatus PCC 6301]
gi|56686533|dbj|BAD79755.1| aspartate aminotransferase
[Synechococcus elongatus PCC 6301]
gi|46130575|ref|ZP_00165505.2| COG0436:
Aspartate/tyrosine/aromatic aminotransferase
[Synechococcus elongatus PCC 7942]
Length = 392
Score = 154 bits (390), Expect = 4e-36
Identities = 105/336 (31%), Positives = 168/336 (49%), Gaps = 26/336 (7%)
Query: 62 SRYGADEGLPELRAALVKKLRQENNL--HKSSVMVTAGANQAFVNLVLTLCDAGDSVVMF 119
+RYG G P+LR A+ +KLR +N L ++++VT G Q+ NL+ L D GD V++
Sbjct: 61 TRYGPAAGEPDLREAIAQKLRADNGLDYQAANILVTNGGKQSLYNLMQVLLDPGDEVIIP 120
Query: 120 APYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTG 179
APY+ + ++ G + V + D L +T P +L+ + +P NPTG
Sbjct: 121 APYWLSYPEMVKLAGGVPVIVETFASDGFKLQPQQLAGAIT---PRTRLLVLNSPSNPTG 177
Query: 180 TYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKH--------SCVEGNHIVNIFSF 231
L+ IA + + W++ D YE ++DG H +C E I N F
Sbjct: 178 MVYSRQELEAIAPIIEAHDFWVVSDEIYEKILYDGADHHSIGSLSPACFERTLISN--GF 235
Query: 232 SKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRER 291
+KAY M GWRVGY+A PSE+ +A L Q + +Q+ A+ +L+ + V E
Sbjct: 236 AKAYSMTGWRVGYLAGPSELIAAAASL---QSHSTSNVCTFAQYGAIAALQGPQDCVAEM 292
Query: 292 VQTLAKNREIVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIP 351
+ + R+++ L+ + S EGA Y+F + K G D R L ++H +A IP
Sbjct: 293 LAAFTERRQLILNGLNQIAGLSCPIPEGAFYVFVDISKTGLDSMTYCRQLLDQHQVAAIP 352
Query: 352 GSACGAAGNLRISFGGLTESDCRAAAERLKKGLEEL 387
G A G ++R+S+ +DC + ++KGLE L
Sbjct: 353 GIAFGDDRSIRLSYA----TDC----QTIEKGLERL 380
>ref|ZP_00131025.1| COG0436: Aspartate/tyrosine/aromatic aminotransferase
[Desulfovibrio desulfuricans G20]
Length = 392
Score = 151 bits (381), Expect = 4e-35
Identities = 118/392 (30%), Positives = 182/392 (46%), Gaps = 21/392 (5%)
Query: 7 LARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKV-KELVWEPSVSRYG 65
+ARR + + M + CVSL QGV + P +D V + L +PS +Y
Sbjct: 7 VARRVQQIRISATKLMPMLAARIGGCVSLGQGVPSFSTPPHIVDAVCRVLRDDPSAGKYS 66
Query: 66 ADEGLPELRAALVKKLRQENNLHK---SSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPY 122
G+P LR A+ + L S V VT GA +A + +LT+ D GD V++ +P
Sbjct: 67 LQPGMPALRQAIAEHLTAAKGFAADPDSEVAVTVGAMEALLMALLTVVDRGDEVIIPSPG 126
Query: 123 YFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYI 182
Y + M +HV P P DV+ + +TE + V V NPGNPTGT
Sbjct: 127 YASHAEQVLMAEGVPVHV-PLRPADWGLDVEAVRAAVTERT---RAVIVCNPGNPTGTVY 182
Query: 183 PESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCVEGN------HIVNIFSFSKAYG 236
+ ++ + L L+ D TY+Y +F G + +++ I SFSK Y
Sbjct: 183 ADEDVRALCRLALEHNLVLISDETYDYMVFGGHAAPLSPASLPEMRENVIVINSFSKKYA 242
Query: 237 MMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLA 296
+ GWRVGY A + G ++LKV D ICA +SQH AL +L + V + L
Sbjct: 243 LTGWRVGYAAASA---GWMGEMLKVHDATAICAPTVSQHAALAALTGPQDCVEQMCAALE 299
Query: 297 KNREIVSEALSPLGEG-SVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSAC 355
K R + + ++ L + GA Y+ A+ +V R + + +PG++
Sbjct: 300 KRRSLTMDRMAALAPAFTCVPPRGAFYVMARYSCTPEPSPEVARRILEEARVITVPGASF 359
Query: 356 GAAG--NLRISFGGLTESDCRAAAERLKKGLE 385
G AG +LR+SFG E++ +RL++ E
Sbjct: 360 GPAGENHLRLSFGA-DEAELTECFDRLQRWAE 390
>ref|NP_951146.1| aminotransferase, classes I and II [Geobacter sulfurreducens PCA]
gi|39981957|gb|AAR33419.1| aminotransferase, classes I
and II [Geobacter sulfurreducens PCA]
Length = 391
Score = 147 bits (372), Expect = 4e-34
Identities = 98/358 (27%), Positives = 168/358 (46%), Gaps = 15/358 (4%)
Query: 33 VSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQENN--LHKS 90
V L Q V + P + D + L+ +P VS+Y DEGLPE+R + + + ++
Sbjct: 36 VDLCQAVPDYPPARQLTDYLAALLDDPLVSKYSPDEGLPEVREGVCARYGRVYGAAMNPD 95
Query: 91 SVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHVGPGSPDTLYP 150
+ +T GA+QAF ++TLC AGD V++ P YF+ M+ + G+ +++ P
Sbjct: 96 QLCLTIGASQAFWLAMVTLCRAGDEVIVPLPAYFDHPMALDILGVRPVYLPFDEERGGVP 155
Query: 151 DVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYEYF 210
D +ER++T P + + +V P NPTG P +Q + + + G L++D TY F
Sbjct: 156 DPAAVERLIT---PRTRAILLVTPSNPTGVVTPPETIQELHGVARRRGIALVLDETYADF 212
Query: 211 MFDGLKHSCV-----EGNHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNI 265
+ G + + G+H++++ SF K Y + G+R G +A E G LK QD +
Sbjct: 213 IPGGERPHDLFLDPRWGDHLIHLMSFGKTYALTGYRAGCLAASKEFIG---HALKAQDTM 269
Query: 266 PICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIYLFA 325
+C I+Q+ L+ + WV E + + ++ + G G + +
Sbjct: 270 AVCQPRITQYAVLYGVSHLDGWVEENRLMMTRRHDLFRSLFTRPGNPFSLVASGTFFAWV 329
Query: 326 KLPKGGYDDFDVVRWLANRHGIAVIPGSACGAA--GNLRISFGGLTESDCRAAAERLK 381
+ P + R LA GI +PG G LR++FG + + A ER +
Sbjct: 330 RHPLQEGTGREAARRLAVEAGIICLPGEVFGPGLEPYLRLAFGNIRDEAIPGAVERFR 387
>dbj|BAB76552.1| aspartate aminotransferase [Nostoc sp. PCC 7120]
gi|17232345|ref|NP_488893.1| aspartate aminotransferase
[Nostoc sp. PCC 7120] gi|25315788|pir||AE2412 aspartate
aminotransferase [imported] - Nostoc sp. (strain PCC
7120)
Length = 388
Score = 147 bits (371), Expect = 6e-34
Identities = 100/313 (31%), Positives = 164/313 (51%), Gaps = 16/313 (5%)
Query: 62 SRYGADEGLPELRAALVKKLRQENNLH--KSSVMVTAGANQAFVNLVLTLCDAGDSVVMF 119
++YGA G P+LR A+ +KL+++N+L +V+VT G + NL++ L D GD V++
Sbjct: 61 TKYGAAAGEPKLREAIARKLQKDNHLDYKPENVIVTNGGKHSLYNLIVALIDPGDEVIIP 120
Query: 120 APYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTG 179
APY+ + + G ++ V P T Y E++ P KL + +P NPTG
Sbjct: 121 APYWLSYPEMVTLVGGKSVIV-PTDASTGYKITP--EQLRKAITPKTKLFVLNSPSNPTG 177
Query: 180 -TYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCVE--GNHIVNIF----SFS 232
Y PE + + +A + +A +++ D YE ++DG +H + G I N F+
Sbjct: 178 MVYTPEEI-KALAQVVVDADIYVVSDEIYEKILYDGAQHISIGSLGKEIFNRTLISNGFA 236
Query: 233 KAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERV 292
KAY M GWR+GY+A P ++ +A ++ +C +Q+ A+ +LE + V E
Sbjct: 237 KAYSMTGWRLGYLAGPVDIIK-AASSIQGHSTSNVCT--FAQYGAIAALEDSQDCVEEMR 293
Query: 293 QTLAKNREIVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPG 352
Q AK R+++ + L+ + S +GA YLF + K G + L H +AVIPG
Sbjct: 294 QAFAKRRQVMLDRLNAIPGLSTAKPDGAFYLFPDISKTGLKSLEFCDALIEEHKVAVIPG 353
Query: 353 SACGAAGNLRISF 365
A GA N+R+S+
Sbjct: 354 IAFGADDNIRLSY 366
>ref|ZP_00160531.2| COG0436: Aspartate/tyrosine/aromatic aminotransferase [Anabaena
variabilis ATCC 29413]
Length = 388
Score = 146 bits (368), Expect = 1e-33
Identities = 100/313 (31%), Positives = 163/313 (51%), Gaps = 16/313 (5%)
Query: 62 SRYGADEGLPELRAALVKKLRQENNLH--KSSVMVTAGANQAFVNLVLTLCDAGDSVVMF 119
++YGA G P+LR A+ KL+++N+L +V+VT G + NL++ L D GD V++
Sbjct: 61 TKYGAAAGEPKLREAIAHKLQKDNHLDYKPENVIVTNGGKHSLYNLIVALIDPGDEVIIP 120
Query: 120 APYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTG 179
APY+ + + G ++ V P T Y E++ P KL + +P NPTG
Sbjct: 121 APYWLSYPEMVTLVGGKSVIV-PTDASTGYKITP--EQLRKAITPKTKLFVLNSPSNPTG 177
Query: 180 -TYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCVE--GNHIVNIF----SFS 232
Y PE + + +A + +A +++ D YE ++DG +H + G I N F+
Sbjct: 178 MVYTPEEI-KALAQVVVDADIYVVSDEIYEKILYDGAQHISIGSLGKEIFNRTLISNGFA 236
Query: 233 KAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERV 292
KAY M GWR+GY+A P ++ +A ++ +C +Q+ A+ +LE + V E
Sbjct: 237 KAYSMTGWRLGYLAGPVDIIK-AASSIQGHSTSNVCT--FAQYGAIAALEDSQDCVEEMR 293
Query: 293 QTLAKNREIVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPG 352
Q AK R+++ + L+ + S +GA YLF + K G + L H +AVIPG
Sbjct: 294 QAFAKRRQVMLDRLNAIPGLSTAKPDGAFYLFPDISKTGLKSLEFCDALIEEHKVAVIPG 353
Query: 353 SACGAAGNLRISF 365
A GA N+R+S+
Sbjct: 354 IAFGADDNIRLSY 366
>dbj|BAD84737.1| aromatic aminotransferase [Thermococcus kodakarensis KOD1]
gi|57640483|ref|YP_182961.1| aromatic aminotransferase
[Thermococcus kodakaraensis KOD1]
Length = 389
Score = 144 bits (364), Expect = 4e-33
Identities = 103/375 (27%), Positives = 186/375 (49%), Gaps = 21/375 (5%)
Query: 24 EVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQ 83
++ G ++ +SL G + P+ KE + + + YG + GLP LR A+ KKL++
Sbjct: 20 DLAAGMQDVISLGIGEPDFDTPEHIKGYAKEAL-DKGYTHYGPNAGLPMLREAVAKKLKE 78
Query: 84 ENNLH---KSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHV 140
+N + K+ VM+ GANQAF+ + T G+ V++ +P + + + + G + V
Sbjct: 79 QNGIEADPKTEVMILTGANQAFLMGLATFLKDGEEVLIPSPMFVSYAPAVILAGGKPVEV 138
Query: 141 GPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSW 200
+ VD LE+ +T+ + + + P NPTG+ + + ++ IA+
Sbjct: 139 PTYEENEFRLSVDDLEKHVTDKT---RALIINTPSNPTGSVLTKKDIEEIADFAVEHDLM 195
Query: 201 LLVDNTYEYFMFDGLKHSCVEG-----NHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLS 255
++ D YE+F++DG+K+ + + + FSK + M GWR+G++A P+ V
Sbjct: 196 VISDEVYEHFVYDGVKNHSIASLDGMFERTITVNGFSKTFAMTGWRLGFVAAPAWV---I 252
Query: 256 AQLLKVQDNIPICASIISQHLALHSLEMGPEW--VRERVQTLAKNREIVSEALSPLGEGS 313
++ + Q C +Q+ A +LE W V E + + R +V + L+ +G +
Sbjct: 253 EKMTRFQMYNSTCPVTFAQYAAAKALEDPRSWKAVEEMRKEYDRRRNLVWKRLNEMGLPT 312
Query: 314 VKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN--LRISFGGLTES 371
VK +GA Y+F ++ G D + +AV+PGSA G AG +RIS+ E
Sbjct: 313 VK-PKGAFYIFPRIRDTGLTDHQFSELMLKEARVAVVPGSAFGKAGEGYIRISYATAYEK 371
Query: 372 DCRAAAERLKKGLEE 386
A +R++K L+E
Sbjct: 372 -LEEAMDRMEKVLKE 385
>ref|YP_009616.1| aromatic aminotransferase [Desulfovibrio vulgaris subsp. vulgaris
str. Hildenborough] gi|46448220|gb|AAS94875.1| aromatic
aminotransferase [Desulfovibrio vulgaris subsp. vulgaris
str. Hildenborough]
Length = 399
Score = 144 bits (362), Expect = 6e-33
Identities = 118/403 (29%), Positives = 185/403 (45%), Gaps = 20/403 (4%)
Query: 1 MGSYANLARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKV-KELVWEP 59
M ++RR + + M + CVSL QGV ++ P+ ++ V + L +
Sbjct: 1 MSDVFRVSRRVQQIRISATKLMPMLAARVGGCVSLGQGVPSFRTPEHIVEAVCRALRDKA 60
Query: 60 SVSRYGADEGLPELRAALVKKLRQENNLH---KSSVMVTAGANQAFVNLVLTLCDAGDSV 116
RY G+P LR A+ L S V VT GA +A + +LT+ D GD V
Sbjct: 61 DAGRYTLQPGMPALREAIAADLAARKGYMVNPDSEVGVTVGAMEALLMALLTVVDRGDEV 120
Query: 117 VMFAPYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGN 176
++ +P Y + M +HV P D DVD + +T P + V V NPGN
Sbjct: 121 IIPSPGYASHAEQVLMAEGVPVHV-PLRADDWGLDVDAIRAAVT---PRTRAVIVCNPGN 176
Query: 177 PTGTYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDG---LKHSCVE--GNHIVNIFSF 231
PTGT ++ ++ + L L+ D TY+Y ++ G L + + H++ + SF
Sbjct: 177 PTGTVYDDADVRALCELALERNIMLISDETYDYMVYGGGEPLSPASLPEMRRHVIVVNSF 236
Query: 232 SKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRER 291
SK Y + GWRVGY A + G +LLKV D ICA +SQ+ AL +L + V +
Sbjct: 237 SKKYALTGWRVGYCAADAAWMG---ELLKVHDAAAICAPAVSQYAALAALTGPQDCVDDM 293
Query: 292 VQTLAKNREIVSEALSPLG-EGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVI 350
L+ R + L + GA Y+ A+ V R L + +
Sbjct: 294 RAALSARRNLACARLDAMAPHFDYVQPRGAFYIMARYTFTDAPSDMVARRLLEEGRVITV 353
Query: 351 PGSACGAAG--NLRISFGGLTESDCRAAAERLKKGLEELVANG 391
PG++ G G +LR+SF G+ E++ A +R+ + +VA+G
Sbjct: 354 PGASFGPTGERHLRLSF-GMEEAELDEAFDRMAAWTQRVVASG 395
>emb|CAB49691.1| aspC-1 aspartate aminotransferase (EC 2.6.1.1) [Pyrococcus abyssi
GE5] gi|14520985|ref|NP_126460.1| aspartate
aminotransferase [Pyrococcus abyssi GE5]
gi|7447827|pir||B75122 probable aromatic-amino-acid
transaminase (EC 2.6.1.57) PAB0525 [similarity] -
Pyrococcus abyssi (strain Orsay)
gi|14285338|sp|Q9V0L2|AAT_PYRAB Aspartate
aminotransferase (Transaminase A) (AspAT)
Length = 389
Score = 143 bits (361), Expect = 8e-33
Identities = 104/375 (27%), Positives = 186/375 (48%), Gaps = 21/375 (5%)
Query: 24 EVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQ 83
++ G K+ +SL G + P+ + KE + + ++ YG + GLPELR A+ +KL++
Sbjct: 20 DIAAGMKDVISLGIGEPDFDTPQHIKEYAKEAL-DMGLTHYGPNIGLPELREAIAEKLKK 78
Query: 84 ENNLH---KSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHV 140
+NN+ +MV GANQAF+ + G+ V++ P + + + + G + V
Sbjct: 79 QNNIEADPNKEIMVLVGANQAFLMGLSAFLKDGEEVLIPTPAFVSYAPAVILAGGKPVEV 138
Query: 141 GPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSW 200
+ +VD L++ +TE K + + +P NPTG+ + + L+ IA+
Sbjct: 139 PTYEENEFRLNVDELKKYVTEKT---KALIINSPCNPTGSVLKKKDLEEIADFAVEHDLI 195
Query: 201 LLVDNTYEYFMFDGLKHSCVEG-----NHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLS 255
++ D YE+F++D +KH + + + FSK + M GWR+G++A PS +
Sbjct: 196 VISDEVYEHFIYDDVKHYSIASLDGMFERTITVNGFSKTFAMTGWRLGFVAAPS---WII 252
Query: 256 AQLLKVQDNIPICASIISQHLALHSLEMGPEW--VRERVQTLAKNREIVSEALSPLGEGS 313
+++K Q C Q+ A +L W V E + + R++V + L+ +G +
Sbjct: 253 EKMVKFQMYNATCPVTFIQYAAAKALRDERSWKAVEEMRKEYDRRRKLVWKRLNEMGLPT 312
Query: 314 VKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN--LRISFGGLTES 371
VK +GA Y+F ++ G + + +AV+PGSA G AG +RIS+ E
Sbjct: 313 VK-PKGAFYIFPRIKDTGLTSKEFSELMLMEAKVAVVPGSAFGKAGEGYVRISYATAYEK 371
Query: 372 DCRAAAERLKKGLEE 386
A +R++K L E
Sbjct: 372 -LEEAMDRMEKVLRE 385
>gb|AAL81377.1| aspartate transaminase [Pyrococcus furiosus DSM 3638]
gi|18977625|ref|NP_578982.1| aspartate transaminase
[Pyrococcus furiosus DSM 3638]
Length = 389
Score = 143 bits (360), Expect = 1e-32
Identities = 103/375 (27%), Positives = 184/375 (48%), Gaps = 21/375 (5%)
Query: 24 EVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQ 83
++ G K+ +SL G + P + KE + + ++ YG + GLPELR A+ +KL++
Sbjct: 20 DLAAGMKDVISLGIGEPDFDTPAHIKEYAKEGL-DKGLTHYGPNVGLPELREAIAEKLKK 78
Query: 84 ENNLH---KSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHV 140
+N + S +MV GANQAF+ + T G+ V++ +P + + + + G + V
Sbjct: 79 QNGIEADPNSEIMVLVGANQAFLMSLATFLKDGEEVLIPSPMFVSYAPAVILAGGKPVEV 138
Query: 141 GPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSW 200
+ +VD L++ ++E + + + +P NPTG + + L+ IA+
Sbjct: 139 PTYEDNEFRINVDDLKKHVSEKT---RALIINSPNNPTGAVLTKKDLEEIADFANEHDLM 195
Query: 201 LLVDNTYEYFMFDGLKHSCVEG-----NHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLS 255
++ D YE+F++DG KH + + + FSK + M GWR+G++ PS V
Sbjct: 196 IISDEVYEHFIYDGAKHYSIAALDGMFGRTITVNGFSKTFAMTGWRLGFVVAPSWV---I 252
Query: 256 AQLLKVQDNIPICASIISQHLALHSLEMGPEW--VRERVQTLAKNREIVSEALSPLGEGS 313
+++K Q C Q+ A +L W V E + + R +V + L+ +G +
Sbjct: 253 EKMVKFQMYNATCPVTFIQYAAAKALRDERSWKAVEEMRKEYERRRNLVWKRLNEMGLPT 312
Query: 314 VKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN--LRISFGGLTES 371
V+ +GA Y+F ++ G + + +AV+PGSA G AG +RIS+ E
Sbjct: 313 VE-PKGAFYIFPRIKDTGLSSKEFSELMLMEAKVAVVPGSAFGRAGEGYIRISYATAYEK 371
Query: 372 DCRAAAERLKKGLEE 386
A R++K L+E
Sbjct: 372 -LEEAMNRMEKVLKE 385
>pdb|5BJ3|D Chain D, Thermus Thermophilus Aspartate Aminotransferase Tetra
Mutant 1 gi|34810108|pdb|5BJ3|C Chain C, Thermus
Thermophilus Aspartate Aminotransferase Tetra Mutant 1
gi|34810107|pdb|5BJ3|B Chain B, Thermus Thermophilus
Aspartate Aminotransferase Tetra Mutant 1
gi|34810106|pdb|5BJ3|A Chain A, Thermus Thermophilus
Aspartate Aminotransferase Tetra Mutant 1
Length = 385
Score = 142 bits (359), Expect = 1e-32
Identities = 107/386 (27%), Positives = 185/386 (47%), Gaps = 15/386 (3%)
Query: 7 LARR--ALETEMPIMVQMQ--EVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVS 62
L+RR A++ + P+ V + E+ R + V+L G + P+ + + + + +
Sbjct: 4 LSRRVQAMKPDAPVAVNAKALELRRQGVDLVALTAGEPDFDTPEHVKEAARRALAQGK-T 62
Query: 63 RYGADEGLPELRAALVKKLRQENNLHKS--SVMVTAGANQAFVNLVLTLCDAGDSVVMFA 120
+Y G+PELR AL +K R+EN L + +VT G +QA NL + D GD V++ +
Sbjct: 63 KYAPPAGIPELREALAEKFRRENGLSVTPEETIVTVGGSQALFNLFQAILDPGDEVIVLS 122
Query: 121 PYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGT 180
PY+ + + G + V + PD + + R +T P K + V +P NPTG
Sbjct: 123 PYWVSYPEMVRFAGGVVVEVETLPEEGFVPDPERVRRAIT---PRTKALVVNSPNNPTGA 179
Query: 181 YIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHS--CVEGNHIVNIFSFSKAYGMM 238
P+ +L+ +A L +L+ D YE+ +++G S V H + + +KA+ M
Sbjct: 180 VYPKEVLEALARLAVEHDFYLVSDEIYEHLLYEGEHFSPGRVAPEHTLTVNGAAKAFAMT 239
Query: 239 GWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKN 298
GWR+GY P EV A + + P + + AL + E +V + +
Sbjct: 240 GWRIGYACGPKEVIKAMASVSRQSTTSPDTIAQWATLEALTNQEASRAFVEMAREAYRRR 299
Query: 299 REIVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAA 358
R+++ E L+ LG +V+ GA Y+ D+ L G+AV+PG+ A
Sbjct: 300 RDLLLEGLTALGLKAVR-PSGAFYVLMDTSPIAPDEVRAAERLLEA-GVAVVPGTDFAAF 357
Query: 359 GNLRISFGGLTESDCRAAAERLKKGL 384
G++R+S+ +E + R A ER + L
Sbjct: 358 GHVRLSY-ATSEENLRKALERFARVL 382
>emb|CAE20954.1| Aminotransferases class-I [Prochlorococcus marinus str. MIT 9313]
gi|33863051|ref|NP_894611.1| Aminotransferases class-I
[Prochlorococcus marinus str. MIT 9313]
Length = 392
Score = 142 bits (358), Expect = 2e-32
Identities = 106/381 (27%), Positives = 192/381 (49%), Gaps = 19/381 (4%)
Query: 16 MPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRA 75
+ I + + + + ++ SL+ G + P+ +D + + + ++RYG G PELR
Sbjct: 20 LAISARAKALQQEGRDICSLSAGEPDFNTPEFIIDATVKALRD-GITRYGPAAGDPELRE 78
Query: 76 ALVKKLRQENNL--HKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMT 133
A+ KL +EN + + V+VT G QA NL + + GD V++ APY+ + ++
Sbjct: 79 AIATKLSKENTVPTNAEQVLVTNGGKQAIFNLFQVILNPGDEVLIPAPYWLSYPEMARLA 138
Query: 134 GITNIHVGPGSPDTLYP-DVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIAN 192
G + P +P+ + D++ LE + P +L+ + +PGNPTG + L+ +A+
Sbjct: 139 G-AKVTTLPSTPENGFCLDLNNLEASIG---PKTRLLLLNSPGNPTGRVMARKELETLAD 194
Query: 193 LCKNAGSWLLV-DNTYEYFMFDGLKHSCVEG------NHIVNIFSFSKAYGMMGWRVGYI 245
L ++ L++ D YE+ + +G +H N + F+K + M GWR+GY+
Sbjct: 195 LLRDHPQILVMSDEIYEFILEEGQQHYSFSAIAPDLSNRTFIVNGFAKGWAMTGWRLGYL 254
Query: 246 AYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEA 305
A P++ +A L+ Q +C+ +Q AL +L+ E V++ V++ RE+++
Sbjct: 255 AGPADAVK-AATALQSQSTSNVCS--FAQRGALAALQGSRECVKKMVKSYNTRRELLTSG 311
Query: 306 LSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGNLRISF 365
L L S+ +GA Y F KLP D HG+A++PG+A G +R++
Sbjct: 312 LLSLEGMSLVPPKGAFYAFPKLPPESLDSVSFCEQALENHGLAMVPGAAFGDDSCVRLTC 371
Query: 366 GGLTESDCRAAAERLKKGLEE 386
E+ C ERL+K L++
Sbjct: 372 AVSPETIC-DGLERLRKALKQ 391
>ref|NP_892792.1| Aminotransferases class-I [Prochlorococcus marinus subsp. pastoris
str. CCMP1986] gi|33639963|emb|CAE19133.1|
Aminotransferases class-I [Prochlorococcus marinus
subsp. pastoris str. CCMP1986]
Length = 393
Score = 141 bits (356), Expect = 3e-32
Identities = 109/398 (27%), Positives = 188/398 (46%), Gaps = 28/398 (7%)
Query: 3 SYANLARRALETEMPIMVQMQ----EVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWE 58
S NL++RAL E + +Q+ ++ + K+ +L+ G + P L E +++
Sbjct: 2 SEVNLSKRALSIEPSLTLQISAKANQLAKEGKDICNLSAGEPDFDAPNEILKATSEAIFD 61
Query: 59 PSVSRYGADEGLPELRAALVKKLRQENNLHKS--SVMVTAGANQAFVNLVLTLCDAGDSV 116
++YG G ELR A+ +KL+ +NNL+ +VMVT GA Q+ NL L + GD V
Sbjct: 62 -GYTKYGPASGDLELRKAIAEKLQTQNNLNVEYENVMVTNGAKQSIYNLFQVLLNDGDEV 120
Query: 117 VMFAPYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGN 176
++ APY+ + ++ G + + + D ++ L+ ++ P K + + +P N
Sbjct: 121 IIPAPYWLSYPQMVRLAGGKPVFLNSSAEDGFKINIQDLK---SKISPKTKFIIINSPNN 177
Query: 177 PTGTYIPESLLQRIANLCK-NAGSWLLVDNTYEYFMFDGLKHSCVEG------NHIVNIF 229
PTG +P+ L +IA L + N +L D YE + KH + I I
Sbjct: 178 PTGRVMPKEELLQIAELVRENKNINILSDEIYELILKKEFKHLSLASLATDLKERIFIIN 237
Query: 230 SFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVR 289
F+K + M GWRVGY+ +V S+ L + Q +C+ + Q AL +L++ E+
Sbjct: 238 GFAKGWAMTGWRVGYLVGQKDVIKASSAL-QSQSTSNVCSFV--QRGALEALKINREFFL 294
Query: 290 ERVQTLAKNREIVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAV 349
E RE++ + L + + GA Y F +LP + + + N +G+ V
Sbjct: 295 EINSHYDLRREVLYKGLKNIEGLFISPPNGAFYAFPRLPNNSMTSVEFCKRILNDYGLVV 354
Query: 350 IPGSACGAAGNLRISFGGLTESDCRAAAERLKKGLEEL 387
+PG G +RIS C ++ E++ GL L
Sbjct: 355 VPGKPFGDDQCIRIS--------CASSKEKILDGLTRL 384
>gb|AAM05225.1| aspartate aminotransferase [Methanosarcina acetivorans str. C2A]
gi|20090670|ref|NP_616745.1| aspartate aminotransferase
[Methanosarcina acetivorans C2A]
Length = 380
Score = 140 bits (353), Expect = 7e-32
Identities = 109/367 (29%), Positives = 182/367 (48%), Gaps = 22/367 (5%)
Query: 25 VLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQE 84
+++ + ++ + G + PK D + ++E + Y G+PELRAA+ +KL+ E
Sbjct: 24 MIKEGTDVINFSLGEPDFDTPKNICDAAAKAMYEGK-THYAPSAGIPELRAAIAEKLKTE 82
Query: 85 NNLH--KSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHVGP 142
N+L + V+VT GA QA +++ D GD ++F P + + +G + V P
Sbjct: 83 NHLEVTEKDVLVTPGAKQAIFEIMMGALDDGDRALLFDPAWVTYDACIRFSGANTVWV-P 141
Query: 143 GSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLL 202
P+ + ++ E I ++K L+ V +PGNPTG + LQ IA+L + ++
Sbjct: 142 TVPERGFLPDNFAEYINDKTK----LIVVNSPGNPTGGVFGKKTLQCIADLAIDHDLLVV 197
Query: 203 VDNTYEYFMFDGLKHSCVEG-----NHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQ 257
D YE ++D +H + + + + FSKAY M GWR+GY+ P E+ L
Sbjct: 198 SDEIYEKIIYDR-EHISIGSFDGMQDRTITVNGFSKAYAMTGWRLGYLTAPPEIFKL--- 253
Query: 258 LLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGG 317
L K+Q + A+ Q+ L +L+ + V+ V R+I+ + L+ +G K
Sbjct: 254 LQKIQSHSVSSATTFVQYGGLEALQGPQDGVKAMVDRFKMRRDILIDGLNKIGI-ECKKP 312
Query: 318 EGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGN--LRISFGGLTESDCRA 375
+GA Y FA + + G R L H +AV PG A GA+G +RIS+ + R
Sbjct: 313 DGAFYAFANVSEYGNGTEVAERLLKEAH-VAVTPGIAFGASGEDFIRISYATSIDR-IRE 370
Query: 376 AAERLKK 382
A ERL+K
Sbjct: 371 ALERLEK 377
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.136 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 710,080,851
Number of Sequences: 2540612
Number of extensions: 30702819
Number of successful extensions: 76768
Number of sequences better than 10.0: 2822
Number of HSP's better than 10.0 without gapping: 760
Number of HSP's successfully gapped in prelim test: 2062
Number of HSP's that attempted gapping in prelim test: 71916
Number of HSP's gapped (non-prelim): 2993
length of query: 399
length of database: 863,360,394
effective HSP length: 130
effective length of query: 269
effective length of database: 533,080,834
effective search space: 143398744346
effective search space used: 143398744346
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0226.7


