

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.2
(583 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
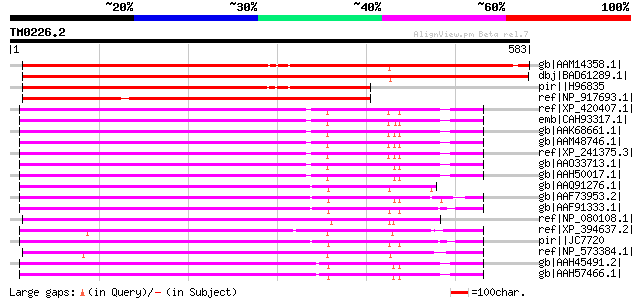
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM14358.1| putative N-terminal acetyltransferase [Arabidopsi... 806 0.0
dbj|BAD61289.1| acetyltransferase 1-like [Oryza sativa (japonica... 787 0.0
pir||H96835 hypothetical protein T21F11.26 [imported] - Arabidop... 623 e-177
ref|NP_917693.1| P0686E09.16 [Oryza sativa (japonica cultivar-gr... 603 e-171
ref|XP_420407.1| PREDICTED: similar to transcriptional coactivat... 370 e-101
emb|CAH93317.1| hypothetical protein [Pongo pygmaeus] 368 e-100
gb|AAK68661.1| gastric cancer antigen Ga19 [Homo sapiens] gi|145... 367 e-100
gb|AAM48746.1| transcriptional coactivator tubedown-100 [Homo sa... 367 e-100
ref|XP_241375.3| PREDICTED: similar to N-terminal aceyltransfera... 365 2e-99
gb|AAO33713.1| N-terminal aceyltransferase 1 [Mus musculus] gi|2... 365 3e-99
gb|AAH50017.1| NMDA receptor-regulated gene 1 [Mus musculus] gi|... 363 6e-99
gb|AAQ91276.1| transcriptional coactivator tubedown-100 [Danio r... 363 1e-98
gb|AAF73953.2| acetyltransferase Tubedown-1 [Mus musculus] 362 2e-98
gb|AAF91333.1| putative N-terminal acetyltransferase [Xenopus la... 361 3e-98
ref|NP_080108.1| NMDA receptor regulated 1-like [Mus musculus] g... 358 3e-97
ref|XP_394637.2| PREDICTED: similar to ENSANGP00000006226 [Apis ... 357 4e-97
pir||JC7720 acetyltransferase (EC 2.3.1.-) 1, Xat-1 - African cl... 357 6e-97
ref|NP_573384.1| CG12202-PA [Drosophila melanogaster] gi|7293585... 356 1e-96
gb|AAH45491.2| NMDA receptor-regulated gene 1 [Danio rerio] 354 4e-96
gb|AAH57466.1| NMDA receptor-regulated gene 1 [Danio rerio] gi|4... 354 5e-96
>gb|AAM14358.1| putative N-terminal acetyltransferase [Arabidopsis thaliana]
gi|17381118|gb|AAL36371.1| putative N-terminal
acetyltransferase [Arabidopsis thaliana]
gi|26451312|dbj|BAC42757.1| putative N-terminal
acetyltransferase [Arabidopsis thaliana]
gi|22330770|ref|NP_178157.2| acetyltransferase-related
[Arabidopsis thaliana]
Length = 897
Score = 806 bits (2083), Expect = 0.0
Identities = 414/601 (68%), Positives = 484/601 (79%), Gaps = 39/601 (6%)
Query: 15 QRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLEHSI 74
+RIPLDFLQ + F+EA YI+PLLTKGVPSLFSDLSSLY+HP K DILEQL+++++HSI
Sbjct: 302 KRIPLDFLQDENFKEAVAKYIKPLLTKGVPSLFSDLSSLYDHPRKPDILEQLVVEMKHSI 361
Query: 75 RTSGQYPGSMEKEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDLYSVKS 134
T+G +PGS KEPPSTL+WTLF LAQHYDRRGQY++A+ KIDEAI HTPTVIDLYSVKS
Sbjct: 362 GTTGSFPGSDVKEPPSTLLWTLFFLAQHYDRRGQYDVALCKIDEAIAHTPTVIDLYSVKS 421
Query: 135 RILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTKEGDQH 194
RI+KHAGD +AAAA ADEAR MDLADRY+NSECVKRMLQADQV LAEKTAVLFTKEGDQ
Sbjct: 422 RIMKHAGDLTAAAALADEARGMDLADRYINSECVKRMLQADQVPLAEKTAVLFTKEGDQL 481
Query: 195 NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCLRKMTLR 254
NNLHDMQCMWY+LASG+S+FRQGDLGRALKKFL VEKHYADI+EDQFDFHSYCLRKMTLR
Sbjct: 482 NNLHDMQCMWYDLASGDSYFRQGDLGRALKKFLAVEKHYADISEDQFDFHSYCLRKMTLR 541
Query: 255 TYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQ 314
+Y++MLKFQD+LHS YFHKAA AIRCY+KLHDSP KSTA EDE MSKL PAQKKK++
Sbjct: 542 SYVDMLKFQDRLHSFPYFHKAAIRAIRCYLKLHDSP-KSTAGEDE-MSKLAPAQKKKIK- 598
Query: 315 KQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHGEKLLQVDDPLSEAIKYLK 374
KQ+KAEARAKK AE K+EE +ASG SKSGKR+VKPVDPDPHG+KL+QV++P++EA KYL+
Sbjct: 599 KQKKAEARAKKEAESKSEESTASGASKSGKRNVKPVDPDPHGQKLIQVEEPMAEASKYLR 658
Query: 375 LLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEHPDSHRCLV--------- 425
LLQK+SP+SLETHLLSFE+ RKQK LL FQAVKQLL+L AE+PDSHR LV
Sbjct: 659 LLQKHSPNSLETHLLSFEVNMRKQKFLLAFQAVKQLLKLGAENPDSHRSLVKFFLMTESI 718
Query: 426 ----------------------CQLHEKTLFEANNSFLEKHKDSLMHRAAFAETLYILDP 463
QL K+L EAN FL +H+DSL+HRAA+AE LYILDP
Sbjct: 719 SAPTTEAEKLRWRVLEAERPSISQLQNKSLMEANKEFLGRHEDSLVHRAAYAEMLYILDP 778
Query: 464 NRKSEAVKLIEESTNNIVPRNGALGPIREWKLKDCIAVHKLLGTVLLDQDAALRWKVRCA 523
++K+EA+K+IE+STN +V N ALG REWKLKDCIAVH LL TVLLD AA RWK RCA
Sbjct: 779 SKKTEAIKIIEDSTNKVVQTNEALGQAREWKLKDCIAVHTLLDTVLLDSQAASRWKSRCA 838
Query: 524 EYFPYSRYFEGSRSSASSNTALKQLSKNSEN-ETLNHSVCTQNVGSITSNGKLEAFKQLA 582
EYFP S +FEG S ++ K++EN +T NH + + S+G+LEAFK L+
Sbjct: 839 EYFPCSTHFEGKHCSLMPDSVYNSSRKSNENGDTPNHPMGQTEL----SDGQLEAFKSLS 894
Query: 583 I 583
+
Sbjct: 895 V 895
>dbj|BAD61289.1| acetyltransferase 1-like [Oryza sativa (japonica cultivar-group)]
Length = 909
Score = 787 bits (2033), Expect = 0.0
Identities = 398/603 (66%), Positives = 471/603 (78%), Gaps = 35/603 (5%)
Query: 15 QRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLEHSI 74
+RIPLDFL+G+KF+EAADNY+RPLLTKGVPSLFSDLS LY PGKA+ILE+L L LE SI
Sbjct: 302 KRIPLDFLEGEKFQEAADNYVRPLLTKGVPSLFSDLSPLYEQPGKANILEELFLKLERSI 361
Query: 75 RTSGQYPGSMEKEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDLYSVKS 134
RTSG +PGS EPPSTL+WTLFL++QHYDRRGQY++A+ KIDEAI HTPTVIDLYS+K
Sbjct: 362 RTSGCFPGSSHTEPPSTLLWTLFLISQHYDRRGQYDIALDKIDEAISHTPTVIDLYSIKG 421
Query: 135 RILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTKEGDQH 194
+IL+HAG+FSAAAA ADEAR MDLADRY+NSECV +MLQADQV LAEKTAVLFTK+GDQH
Sbjct: 422 KILQHAGNFSAAAALADEARSMDLADRYLNSECVMQMLQADQVGLAEKTAVLFTKDGDQH 481
Query: 195 NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCLRKMTLR 254
NNLHDMQCMWYELASGES++RQGDLGRALK FL VEKHYAD+ EDQFDFHSYCLRKMTLR
Sbjct: 482 NNLHDMQCMWYELASGESYYRQGDLGRALKNFLAVEKHYADMTEDQFDFHSYCLRKMTLR 541
Query: 255 TYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQ 314
Y+ MLKFQD+LH+H YFHKAAAGAIRCY+KLHDSP KS+ EE++EMSKL PAQ+KK+RQ
Sbjct: 542 AYVSMLKFQDRLHAHEYFHKAAAGAIRCYMKLHDSPSKSSTEENDEMSKLPPAQRKKLRQ 601
Query: 315 KQRKAEARAKKGAEEKNE-ELSASGVSKSGKR-HVKPVDPDPHGEKLLQVDDPLSEAIKY 372
KQ+KAEARAK+ AEEK E E ++S SKSGK+ + +PVD DPHGEKL+Q+++PL+E KY
Sbjct: 602 KQKKAEARAKREAEEKQEDETTSSHTSKSGKKQNARPVDLDPHGEKLVQIENPLAEGTKY 661
Query: 373 LKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEHPDSHRCLVC------ 426
LKLLQ NS DSLETH LSFEL RKQK+LL FQAVKQL++LD PDSHRCL+
Sbjct: 662 LKLLQNNSSDSLETHTLSFELNMRKQKILLAFQAVKQLIKLDENSPDSHRCLIRFFHKIN 721
Query: 427 -------------------------QLHEKTLFEANNSFLEKHKDSLMHRAAFAETLYIL 461
QLH K+L E N SFLEKH SL HRAA AE +Y+L
Sbjct: 722 NLPSPGTDSEKLIWNVLEAERPDLRQLHGKSLVEVNRSFLEKHSASLTHRAAAAEMMYLL 781
Query: 462 DPNRKSEAVKLIEESTNNIVPRNGALGPIREWKLKDCIAVHKLLGTVLLDQDAALRWKVR 521
+P++K EA+KLIE+S N+ N LGP+ EWK+ DCI VHKLL T+ DQD A WK R
Sbjct: 782 EPDKKLEAIKLIEDSVNSTASGNSVLGPVNEWKILDCIDVHKLLETIFGDQDVANSWKAR 841
Query: 522 CAEYFPYSRYFEGSRS-SASSNTALKQLSKNSENETL-NHSVCTQNVGSITSNGKLEAFK 579
CAEYFPYS YFEG +S SA+ + L +SEN + N + + + + T NG +
Sbjct: 842 CAEYFPYSTYFEGIKSASAAYCSVANSLEDSSENGIVANAQMKSADGETCTLNGTVHIVD 901
Query: 580 QLA 582
+L+
Sbjct: 902 ELS 904
>pir||H96835 hypothetical protein T21F11.26 [imported] - Arabidopsis thaliana
gi|6730746|gb|AAF27136.1| putative N-terminal
acetyltransferase; 84330-89402 [Arabidopsis thaliana]
Length = 683
Score = 623 bits (1607), Expect = e-177
Identities = 309/391 (79%), Positives = 353/391 (90%), Gaps = 3/391 (0%)
Query: 15 QRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLEHSI 74
+RIPLDFLQ + F+EA YI+PLLTKGVPSLFSDLSSLY+HP K DILEQL+++++HSI
Sbjct: 287 KRIPLDFLQDENFKEAVAKYIKPLLTKGVPSLFSDLSSLYDHPRKPDILEQLVVEMKHSI 346
Query: 75 RTSGQYPGSMEKEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDLYSVKS 134
T+G +PGS KEPPSTL+WTLF LAQHYDRRGQY++A+ KIDEAI HTPTVIDLYSVKS
Sbjct: 347 GTTGSFPGSDVKEPPSTLLWTLFFLAQHYDRRGQYDVALCKIDEAIAHTPTVIDLYSVKS 406
Query: 135 RILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTKEGDQH 194
RI+KHAGD +AAAA ADEAR MDLADRY+NSECVKRMLQADQV LAEKTAVLFTKEGDQ
Sbjct: 407 RIMKHAGDLTAAAALADEARGMDLADRYINSECVKRMLQADQVPLAEKTAVLFTKEGDQL 466
Query: 195 NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCLRKMTLR 254
NNLHDMQCMWY+LASG+S+FRQGDLGRALKKFL VEKHYADI+EDQFDFHSYCLRKMTLR
Sbjct: 467 NNLHDMQCMWYDLASGDSYFRQGDLGRALKKFLAVEKHYADISEDQFDFHSYCLRKMTLR 526
Query: 255 TYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQ 314
+Y++MLKFQD+LHS YFHKAA AIRCY+KLHDS PKSTA ED EMSKL PAQKKK++
Sbjct: 527 SYVDMLKFQDRLHSFPYFHKAAIRAIRCYLKLHDS-PKSTAGED-EMSKLAPAQKKKIK- 583
Query: 315 KQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHGEKLLQVDDPLSEAIKYLK 374
KQ+KAEARAKK AE K+EE +ASG SKSGKR+VKPVDPDPHG+KL+QV++P++EA KYL+
Sbjct: 584 KQKKAEARAKKEAESKSEESTASGASKSGKRNVKPVDPDPHGQKLIQVEEPMAEASKYLR 643
Query: 375 LLQKNSPDSLETHLLSFELYTRKQKVLLTFQ 405
LLQK+SP+SLETHLLSFE+ RKQK LL FQ
Sbjct: 644 LLQKHSPNSLETHLLSFEVNMRKQKFLLAFQ 674
>ref|NP_917693.1| P0686E09.16 [Oryza sativa (japonica cultivar-group)]
Length = 686
Score = 603 bits (1555), Expect = e-171
Identities = 297/393 (75%), Positives = 342/393 (86%), Gaps = 10/393 (2%)
Query: 15 QRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLEHSI 74
+RIPLDFL+G+KF+EAADNY+RPLLTKGVPSLFSDLS LY PGKA+ILE+L L LE SI
Sbjct: 294 KRIPLDFLEGEKFQEAADNYVRPLLTKGVPSLFSDLSPLYEQPGKANILEELFLKLERSI 353
Query: 75 RTSGQYPGSMEKEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDLYSVKS 134
RTSG +PGS EPPSTL+WTLFL++QHYDRRGQY++A+ KIDEAI HTPT
Sbjct: 354 RTSGCFPGSSHTEPPSTLLWTLFLISQHYDRRGQYDIALDKIDEAISHTPT--------G 405
Query: 135 RILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTKEGDQH 194
+IL+HAG+FSAAAA ADEAR MDLADRY+NSECV +MLQADQV LAEKTAVLFTK+GDQH
Sbjct: 406 KILQHAGNFSAAAALADEARSMDLADRYLNSECVMQMLQADQVGLAEKTAVLFTKDGDQH 465
Query: 195 NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCLRKMTLR 254
NNLHDMQCMWYELASGES++RQGDLGRALK FL VEKHYAD+ EDQFDFHSYCLRKMTLR
Sbjct: 466 NNLHDMQCMWYELASGESYYRQGDLGRALKNFLAVEKHYADMTEDQFDFHSYCLRKMTLR 525
Query: 255 TYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQ 314
Y+ MLKFQD+LH+H YFHKAAAGAIRCY+KLHDSP KS+ EE++EMSKL PAQ+KK+RQ
Sbjct: 526 AYVSMLKFQDRLHAHEYFHKAAAGAIRCYMKLHDSPSKSSTEENDEMSKLPPAQRKKLRQ 585
Query: 315 KQRKAEARAKKGAEEKNE-ELSASGVSKSGKR-HVKPVDPDPHGEKLLQVDDPLSEAIKY 372
KQ+KAEARAK+ AEEK E E ++S SKSGK+ + +PVD DPHGEKL+Q+++PL+E KY
Sbjct: 586 KQKKAEARAKREAEEKQEDETTSSHTSKSGKKQNARPVDLDPHGEKLVQIENPLAEGTKY 645
Query: 373 LKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQ 405
LKLLQ NS DSLETH LSFEL RKQK+LL FQ
Sbjct: 646 LKLLQNNSSDSLETHTLSFELNMRKQKILLAFQ 678
>ref|XP_420407.1| PREDICTED: similar to transcriptional coactivator tubedown-100
isoform 1; putative N-acetyltransferase; gastric cancer
antigen Ga19 [Gallus gallus]
Length = 867
Score = 370 bits (949), Expect = e-101
Identities = 210/562 (37%), Positives = 327/562 (57%), Gaps = 56/562 (9%)
Query: 12 LVCQRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLE 71
LV +R+PL+FL G+KF+E D ++R +KG P +F+ L SLY K I+E+L++ E
Sbjct: 293 LVPRRLPLNFLSGEKFKECLDKFLRMNFSKGCPPVFNTLRSLYKDKEKVAIIEELVVGYE 352
Query: 72 HSIRTSGQYPGSME--KEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDL 129
S+R+ + + + +EPP+TL+W + LAQHYD+ GQ LA+ I+ AIE TPT+I+L
Sbjct: 353 TSLRSCRLFNPNDDGKEEPPTTLLWVQYYLAQHYDKIGQPSLALEYINAAIESTPTLIEL 412
Query: 130 YSVKSRILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTK 189
+ VK++I KHAG+ AA + DEA+ +D ADR++NS+C K ML+A+ + AE+ FT+
Sbjct: 413 FLVKAKIYKHAGNIKEAARWMDEAQALDTADRFINSKCAKYMLKANFIKEAEEMCSKFTR 472
Query: 190 EGDQH-NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCL 248
EG NL++MQCMW++ +++ G ALKK +E+H+ +I +DQFDFH+YC+
Sbjct: 473 EGTSAVENLNEMQCMWFQTECAQAYKAMNKFGEALKKCHEIERHFVEITDDQFDFHTYCM 532
Query: 249 RKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSP-PKSTAEEDEEMSKLLPA 307
RK+TLR+Y+++LK +D L H ++ KAA AI Y+KLHD+P E + + + +
Sbjct: 533 RKITLRSYVDLLKLEDVLRQHPFYFKAARIAIEIYLKLHDNPLTDENKEHEADTANMSDK 592
Query: 308 QKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPH----------GE 357
+ KK+R KQR+A+ +A+ E+KN E K + K D D E
Sbjct: 593 ELKKLRNKQRRAQKKAQLEEEKKNAE-----KEKQQRNQKKKKDDDDEEIGGPKEELIPE 647
Query: 358 KLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEH 417
KL +V+ PL EAIK+L L+ + +ETHL +FE+Y RK+K LL Q+VK+ +D+ H
Sbjct: 648 KLAKVEAPLEEAIKFLTPLKNLVKNKIETHLFAFEIYFRKEKFLLMLQSVKRAFAIDSSH 707
Query: 418 PDSHRCL-------------------VCQLHEKTLFEA------NNSFLEKHKDSLMHRA 452
P H C+ V LF A N +FL+K+ DSL HR
Sbjct: 708 PWLHECMIHLFSSVSESKDLPDAVRTVLNQEMNRLFGATNPKNFNEAFLKKNYDSLPHRL 767
Query: 453 AFAETLYILDPNRKSEAVKLIEESTNNIVPRNGALGPIREWKLKDCIAVHKLL--GTVLL 510
+ A+ +Y LDP+ + AV+L +++ RN L+ C+ V + L G++
Sbjct: 768 SAAKMMYYLDPSSQKRAVELAMTLDESLINRN----------LQTCMEVLEALCDGSLGD 817
Query: 511 DQDAALRWKVRCAEYFPYSRYF 532
++A+ ++ C + FPY+ F
Sbjct: 818 CKEASETYRANCHKLFPYALAF 839
>emb|CAH93317.1| hypothetical protein [Pongo pygmaeus]
Length = 866
Score = 368 bits (945), Expect = e-100
Identities = 208/563 (36%), Positives = 330/563 (57%), Gaps = 57/563 (10%)
Query: 12 LVCQRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLE 71
LV +R+PL+FL G+KF+E D ++R +KG P +F+ L SLY K I+E+L++ E
Sbjct: 291 LVPRRLPLNFLSGEKFKECLDKFLRMNFSKGCPPVFNTLRSLYKDKEKVAIIEELVVGYE 350
Query: 72 HSIRTSGQYPGSME--KEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDL 129
S+++ + + + +EPP+TL+W + LAQHYD+ GQ +A+ I+ AIE TPT+I+L
Sbjct: 351 TSLKSCRLFNPNDDGKEEPPTTLLWVQYYLAQHYDKIGQPSIALEYINTAIESTPTLIEL 410
Query: 130 YSVKSRILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTK 189
+ VK++I KHAG+ AA + DEA+ +D ADR++NS+C K ML+A+ + AE+ FT+
Sbjct: 411 FLVKAKIYKHAGNIKEAARWMDEAQALDTADRFINSKCAKYMLKANLIKEAEEMCSKFTR 470
Query: 190 EGDQH-NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCL 248
EG NL++MQCMW++ +++ G ALKK +E+H+ +I +DQFDFH+YC+
Sbjct: 471 EGTSAVENLNEMQCMWFQTECAQAYKAMNKFGEALKKCYEIERHFIEITDDQFDFHTYCM 530
Query: 249 RKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSP-PKSTAEEDEEMSKLLPA 307
RK+TLR+Y+++LK +D L H ++ KAA AI Y+KLHD+P E + + + +
Sbjct: 531 RKITLRSYVDLLKLEDVLRQHPFYFKAARIAIEIYLKLHDNPLTDENKEHEADTANMSDK 590
Query: 308 QKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPH----------GE 357
+ KK+R KQR+A+ +A+ E+KN E K + K D D E
Sbjct: 591 ELKKLRNKQRRAQKKAQIEEEKKNAE-----KEKQQRNQKKKKDDDDEEIGGPKEELIPE 645
Query: 358 KLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEH 417
KL +V+ PL EAIK+L L+ + +ETHL +FE+Y RK+K LL Q+VK+ +D+ H
Sbjct: 646 KLAKVETPLEEAIKFLTPLKNLVKNKIETHLFAFEIYFRKEKFLLMLQSVKRAFAIDSSH 705
Query: 418 PDSHRCL-------VCQLHEKT-------------LFEA------NNSFLEKHKDSLMHR 451
P H C+ VC+ + + LF A N +FL+++ DSL HR
Sbjct: 706 PWLHECMIRLFNTAVCESKDLSDTVRTVLKQEMHRLFGATNPKNFNETFLKRNSDSLPHR 765
Query: 452 AAFAETLYILDPNRKSEAVKLIEESTNNIVPRNGALGPIREWKLKDCIAVHKLL--GTVL 509
+ A+ +Y LDP+ + A++L ++ RN L+ C+ V + L G++
Sbjct: 766 LSAAKMVYYLDPSSQKRAIELATTLDESLTNRN----------LQTCMEVLETLYDGSLG 815
Query: 510 LDQDAALRWKVRCAEYFPYSRYF 532
++AA ++ C + FPY+ F
Sbjct: 816 DCKEAAEIYRANCHKLFPYALAF 838
>gb|AAK68661.1| gastric cancer antigen Ga19 [Homo sapiens]
gi|14589342|emb|CAC43228.1| putative N-acetyltransferase
[Homo sapiens] gi|57012969|sp|Q9BXJ9|NARG1_HUMAN NMDA
receptor regulated protein 1 (N-terminal
acetyltransferase) (Tubedown-1 protein) (Tbdn100)
(Gastric cancer antigen Ga19) gi|13195460|gb|AAK15707.1|
putative acetyltransferase [Homo sapiens]
gi|17149828|ref|NP_476516.1| NMDA receptor regulated 1
[Homo sapiens] gi|63992246|gb|AAY40950.1| unknown [Homo
sapiens] gi|62739615|gb|AAH93928.1| NMDA receptor
regulated 1 [Homo sapiens]
Length = 866
Score = 367 bits (943), Expect = e-100
Identities = 208/563 (36%), Positives = 329/563 (57%), Gaps = 57/563 (10%)
Query: 12 LVCQRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLE 71
LV +R+PL+FL G+KF+E D ++R +KG P +F+ L SLY K I+E+L++ E
Sbjct: 291 LVPRRLPLNFLSGEKFKECLDKFLRMNFSKGCPPVFNTLRSLYKDKEKVAIIEELVVGYE 350
Query: 72 HSIRTSGQYPGSME--KEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDL 129
S+++ + + + +EPP+TL+W + LAQHYD+ GQ +A+ I+ AIE TPT+I+L
Sbjct: 351 TSLKSCRLFNPNDDGKEEPPTTLLWVQYYLAQHYDKIGQPSIALEYINTAIESTPTLIEL 410
Query: 130 YSVKSRILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTK 189
+ VK++I KHAG+ AA + DEA+ +D ADR++NS+C K ML+A+ + AE+ FT+
Sbjct: 411 FLVKAKIYKHAGNIKEAARWMDEAQALDTADRFINSKCAKYMLKANLIKEAEEMCSKFTR 470
Query: 190 EGDQH-NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCL 248
EG NL++MQCMW++ +++ G ALKK +E+H+ +I +DQFDFH+YC+
Sbjct: 471 EGTSAVENLNEMQCMWFQTECAQAYKAMNKFGEALKKCHEIERHFIEITDDQFDFHTYCM 530
Query: 249 RKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSP-PKSTAEEDEEMSKLLPA 307
RK+TLR+Y+++LK +D L H ++ KAA AI Y+KLHD+P E + + + +
Sbjct: 531 RKITLRSYVDLLKLEDVLRQHPFYFKAARIAIEIYLKLHDNPLTDENKEHEADTANMSDK 590
Query: 308 QKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPH----------GE 357
+ KK+R KQR+A+ +A+ E+KN E K + K D D E
Sbjct: 591 ELKKLRNKQRRAQKKAQIEEEKKNAE-----KEKQQRNQKKKKDDDDEEIGGPKEELIPE 645
Query: 358 KLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEH 417
KL +V+ PL EAIK+L L+ + +ETHL +FE+Y RK+K LL Q+VK+ +D+ H
Sbjct: 646 KLAKVETPLEEAIKFLTPLKNLVKNKIETHLFAFEIYFRKEKFLLMLQSVKRAFAIDSSH 705
Query: 418 PDSHRCL-------VCQLHE-------------KTLFEA------NNSFLEKHKDSLMHR 451
P H C+ VC+ + LF A N +FL+++ DSL HR
Sbjct: 706 PWLHECMIRLFNTAVCESKDLSDTVRTVLKQEMNRLFGATNPKNFNETFLKRNSDSLPHR 765
Query: 452 AAFAETLYILDPNRKSEAVKLIEESTNNIVPRNGALGPIREWKLKDCIAVHKLL--GTVL 509
+ A+ +Y LDP+ + A++L ++ RN L+ C+ V + L G++
Sbjct: 766 LSAAKMVYYLDPSSQKRAIELATTLDESLTNRN----------LQTCMEVLEALYDGSLG 815
Query: 510 LDQDAALRWKVRCAEYFPYSRYF 532
++AA ++ C + FPY+ F
Sbjct: 816 DCKEAAEIYRANCHKLFPYALAF 838
>gb|AAM48746.1| transcriptional coactivator tubedown-100 [Homo sapiens]
Length = 866
Score = 367 bits (942), Expect = e-100
Identities = 208/563 (36%), Positives = 329/563 (57%), Gaps = 57/563 (10%)
Query: 12 LVCQRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLE 71
LV +R+PL+FL G+KF+E D ++R +KG P +F+ L SLY K I+E+L++ E
Sbjct: 291 LVPRRLPLNFLSGEKFKECLDKFLRMNFSKGCPPVFNTLRSLYKDKEKVAIIEELVVGYE 350
Query: 72 HSIRTSGQYPGSME--KEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDL 129
S+++ + + + +EPP+TL+W + LAQHYD+ GQ +A+ I+ AIE TPT+I+L
Sbjct: 351 TSLKSCRLFNPNDDGKEEPPTTLLWVQYYLAQHYDKIGQPSIALEYINTAIESTPTLIEL 410
Query: 130 YSVKSRILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTK 189
+ VK++I KHAG+ AA + DEA+ +D ADR++NS+C K ML+A+ + AE+ FT+
Sbjct: 411 FLVKAKIYKHAGNIREAARWMDEAQALDTADRFINSKCAKYMLKANLIKEAEEMCSKFTR 470
Query: 190 EGDQH-NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCL 248
EG NL++MQCMW++ +++ G ALKK +E+H+ +I +DQFDFH+YC+
Sbjct: 471 EGTSAVENLNEMQCMWFQTECAQAYKAMNKFGEALKKCHEIERHFIEITDDQFDFHTYCM 530
Query: 249 RKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSP-PKSTAEEDEEMSKLLPA 307
RK+TLR+Y+++LK +D L H ++ KAA AI Y+KLHD+P E + + + +
Sbjct: 531 RKITLRSYVDLLKLEDVLRQHPFYFKAARIAIEIYLKLHDNPLTDENKEHEADTANMSDK 590
Query: 308 QKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPH----------GE 357
+ KK+R KQR+A+ +A+ E+KN E K + K D D E
Sbjct: 591 ELKKLRNKQRRAQKKAQIEEEKKNAE-----KEKQQRNQKKKKDDDDEEIGGPKEELIPE 645
Query: 358 KLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEH 417
KL +V+ PL EAIK+L L+ + +ETHL +FE+Y RK+K LL Q+VK+ +D+ H
Sbjct: 646 KLAKVETPLEEAIKFLTPLKNLVKNKIETHLFAFEIYFRKEKFLLMLQSVKRAFAIDSSH 705
Query: 418 PDSHRCL-------VCQLHE-------------KTLFEA------NNSFLEKHKDSLMHR 451
P H C+ VC+ + LF A N +FL+++ DSL HR
Sbjct: 706 PWLHECMIRLFNTAVCESKDLSDTVRTVLKQEMNRLFGATNPKNFNETFLKRNSDSLPHR 765
Query: 452 AAFAETLYILDPNRKSEAVKLIEESTNNIVPRNGALGPIREWKLKDCIAVHKLL--GTVL 509
+ A+ +Y LDP+ + A++L ++ RN L+ C+ V + L G++
Sbjct: 766 LSAAKMVYYLDPSSQKRAIELATTLDESLTNRN----------LQTCMEVLEALYDGSLG 815
Query: 510 LDQDAALRWKVRCAEYFPYSRYF 532
++AA ++ C + FPY+ F
Sbjct: 816 DCKEAAEIYRANCHKLFPYALAF 838
>ref|XP_241375.3| PREDICTED: similar to N-terminal aceyltransferase 1 [Rattus
norvegicus]
Length = 865
Score = 365 bits (937), Expect = 2e-99
Identities = 210/562 (37%), Positives = 327/562 (57%), Gaps = 56/562 (9%)
Query: 12 LVCQRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLE 71
LV +R+PL+FL G+KF+E D ++R +KG P +F+ L SLY K I+E+L++ E
Sbjct: 291 LVPRRLPLNFLSGEKFKECLDRFLRMNFSKGCPPVFNTLRSLYRDKEKVAIVEELVVGYE 350
Query: 72 HSIRTSGQYPGSME--KEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDL 129
S ++ + + + +EPP+TL+W + LAQHYD+ GQ +A+ I+ AIE TPT+I+L
Sbjct: 351 TSPKSCRLFNPNDDGKEEPPTTLLWVQYYLAQHYDKIGQPSIALEYINTAIESTPTLIEL 410
Query: 130 YSVKSRILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTK 189
+ VK++I KHAG+ AA + DEA+ +D ADR++NS+C K ML+A+ + AE+ FT+
Sbjct: 411 FLVKAKIYKHAGNIKEAARWMDEAQALDTADRFINSKCAKYMLKANLIKEAEEMCSKFTR 470
Query: 190 EGDQH-NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCL 248
EG NL++MQCMW++ +++ G ALKK +E+H+ +I +DQFDFH+YC+
Sbjct: 471 EGTSAVENLNEMQCMWFQTECAQAYKAMNKFGEALKKCHEIERHFIEITDDQFDFHTYCM 530
Query: 249 RKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSP-PKSTAEEDEEMSKLLPA 307
RK+TLR+Y+++LK +D L H ++ KAA AI Y+KLHD+P E + + + +
Sbjct: 531 RKITLRSYVDLLKLEDVLRQHPFYFKAARIAIEIYLKLHDNPLTDENKEHEADTANMSDK 590
Query: 308 QKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPH----------GE 357
+ KK+R KQR+A+ +A+ E+KN E K + K D D E
Sbjct: 591 ELKKLRNKQRRAQKKAQIEEEKKNAE-----KEKQQRNQKKKKDDDDEEIGGPKEELIPE 645
Query: 358 KLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEH 417
KL +V+ PL EAIK+L L+ + +ETHL +FE+Y RK+K LL Q+VK+ +D+ H
Sbjct: 646 KLAKVETPLEEAIKFLTPLKNLVKNKIETHLFAFEIYFRKEKFLLMLQSVKRAFAIDSSH 705
Query: 418 PDSHRCL------VCQLHE-------------KTLFEA------NNSFLEKHKDSLMHRA 452
P H C+ VC+ + LF A N +FL+++ DSL HR
Sbjct: 706 PWLHECMIRLFNSVCESKDLPEAVRTVLKQEMNRLFGATNPKNFNETFLKRNSDSLPHRL 765
Query: 453 AFAETLYILDPNRKSEAVKLIEESTNNIVPRNGALGPIREWKLKDCIAV-HKLLGTVLLD 511
+ A+ +Y LD + + A++L ++ RN L+ C+ V L G L D
Sbjct: 766 SAAKMIYYLDSSSQKRAIELATTLDGSLTNRN----------LQTCMEVLEALCGGSLGD 815
Query: 512 -QDAALRWKVRCAEYFPYSRYF 532
++AA ++V C + FPY+ F
Sbjct: 816 CKEAAEAYRVSCHKLFPYALAF 837
>gb|AAO33713.1| N-terminal aceyltransferase 1 [Mus musculus]
gi|28269697|ref|NP_444319.2| NMDA receptor-regulated
gene 1 [Mus musculus]
Length = 865
Score = 365 bits (936), Expect = 3e-99
Identities = 205/562 (36%), Positives = 326/562 (57%), Gaps = 56/562 (9%)
Query: 12 LVCQRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLE 71
LV +++PL+FL G+KF+E D ++R +KG P +F+ L SLY K I+E+L++ E
Sbjct: 291 LVPRKLPLNFLSGEKFKECLDRFLRMNFSKGCPPVFNTLRSLYRDKEKVAIVEELVVGYE 350
Query: 72 HSIRTSGQYPGSME--KEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDL 129
S+++ + + + +EPP+TL+W + LAQHYD+ GQ +A+ I+ AIE TPT+I+L
Sbjct: 351 TSLKSCRLFNPNDDGKEEPPTTLLWVQYYLAQHYDKIGQPSIALEYINTAIESTPTLIEL 410
Query: 130 YSVKSRILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTK 189
+ VK++I KHAG+ AA + DEA+ +D ADR++NS+C K ML+A+ + AE+ FT+
Sbjct: 411 FLVKAKIYKHAGNIKEAARWMDEAQALDTADRFINSKCAKYMLKANLIKEAEEMCSKFTR 470
Query: 190 EGDQH-NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCL 248
EG NL++MQCMW++ +++ G ALKK +E+H+ +I +DQFDFH+YC+
Sbjct: 471 EGTSAVENLNEMQCMWFQTECAQAYKAMNKFGEALKKCHEIERHFIEITDDQFDFHTYCM 530
Query: 249 RKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSP-PKSTAEEDEEMSKLLPA 307
RK+TLR+Y+++LK +D L H ++ KAA AI Y+KLHD+P E + + + +
Sbjct: 531 RKITLRSYVDLLKLEDVLRQHPFYFKAARIAIEIYLKLHDNPLTDENKEHEADTANMSDK 590
Query: 308 QKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPH----------GE 357
+ KK+R KQR+A+ +A+ E+KN E K + K D D E
Sbjct: 591 ELKKLRNKQRRAQKKAQIEEEKKNAE-----KEKQQRNQKKKKDDDDEEIGGPKEELIPE 645
Query: 358 KLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEH 417
KL +V+ PL EAIK+L L+ + +ETHL +FE+Y RK+K LL Q+VK+ +D+ H
Sbjct: 646 KLAKVETPLEEAIKFLTPLKNLVKNKIETHLFAFEIYFRKEKFLLMLQSVKRAFAIDSSH 705
Query: 418 PDSHRCLVCQLHE-------------------KTLFEA------NNSFLEKHKDSLMHRA 452
P H C++ H LF A N +FL+++ DSL HR
Sbjct: 706 PWLHECMIRLFHSVCESKDLPETVRTVLKQEMNRLFGATNPKNFNETFLKRNSDSLPHRL 765
Query: 453 AFAETLYILDPNRKSEAVKLIEESTNNIVPRNGALGPIREWKLKDCIAVHKLL--GTVLL 510
+ A+ +Y LD + + A++L ++ RN L+ C+ V + L G++
Sbjct: 766 SAAKMVYYLDSSSQKRAIELATTLDGSLTNRN----------LQTCMEVLEALCDGSLGD 815
Query: 511 DQDAALRWKVRCAEYFPYSRYF 532
++AA ++ C + FPY+ F
Sbjct: 816 CKEAAEAYRASCHKLFPYALAF 837
>gb|AAH50017.1| NMDA receptor-regulated gene 1 [Mus musculus]
gi|57012950|sp|Q80UM3|NARG1_MOUSE NMDA receptor
regulated protein 1 (N-terminal aceyltransferase 1)
(Tubedown-1 protein)
Length = 865
Score = 363 bits (933), Expect = 6e-99
Identities = 205/562 (36%), Positives = 326/562 (57%), Gaps = 56/562 (9%)
Query: 12 LVCQRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLE 71
LV +R+PL+FL G+KF+E D ++R +KG P +F+ L SLY K I+E+L++ E
Sbjct: 291 LVPRRLPLNFLSGEKFKECLDRFLRMNFSKGCPPVFNTLRSLYRDKEKVAIVEELVVGYE 350
Query: 72 HSIRTSGQYPGSME--KEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDL 129
S+++ + + + +EPP+TL+W + LAQHYD+ GQ +A+ I+ AIE TPT+I+L
Sbjct: 351 TSLKSCRLFNPNDDGKEEPPTTLLWVQYYLAQHYDKIGQPSIALEYINTAIESTPTLIEL 410
Query: 130 YSVKSRILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTK 189
+ VK++I KHAG+ AA + DEA+ +D ADR++NS+C K +L+A+ + AE+ FT+
Sbjct: 411 FLVKAKIYKHAGNIKEAARWMDEAQALDTADRFINSKCAKYVLKANLIKEAEEMCSKFTR 470
Query: 190 EGDQH-NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCL 248
EG NL++MQCMW++ +++ G ALKK +E+H+ +I +DQFDFH+YC+
Sbjct: 471 EGTSAVENLNEMQCMWFQTECAQAYKAMNKFGEALKKCHEIERHFIEITDDQFDFHTYCM 530
Query: 249 RKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSP-PKSTAEEDEEMSKLLPA 307
RK+TLR+Y+++LK +D L H ++ KAA AI Y+KLHD+P E + + + +
Sbjct: 531 RKITLRSYVDLLKLEDVLRQHPFYFKAARIAIEIYLKLHDNPLTDENKEHEADTANMSDK 590
Query: 308 QKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPH----------GE 357
+ KK+R KQR+A+ +A+ E+KN E K + K D D E
Sbjct: 591 ELKKLRNKQRRAQKKAQIEEEKKNAE-----KEKQQRNQKKKKDDDDEEIGGPKEELIPE 645
Query: 358 KLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEH 417
KL +V+ PL EAIK+L L+ + +ETHL +FE+Y RK+K LL Q+VK+ +D+ H
Sbjct: 646 KLAKVETPLEEAIKFLTPLKNLVKNKIETHLFAFEIYFRKEKFLLMLQSVKRAFAIDSGH 705
Query: 418 PDSHRCLVCQLHE-------------------KTLFEA------NNSFLEKHKDSLMHRA 452
P H C++ H LF A N +FL+++ DSL HR
Sbjct: 706 PWLHECMIRLFHSVCESKDLPETVRTVLKQEMNRLFGATNPKNFNETFLKRNSDSLPHRL 765
Query: 453 AFAETLYILDPNRKSEAVKLIEESTNNIVPRNGALGPIREWKLKDCIAVHKLL--GTVLL 510
+ A+ +Y LD + + A++L ++ RN L+ C+ V + L G++
Sbjct: 766 SAAKMVYYLDSSSQKRAIELATTLDGSLTNRN----------LQTCMEVLEALCDGSLGD 815
Query: 511 DQDAALRWKVRCAEYFPYSRYF 532
++AA ++ C + FPY+ F
Sbjct: 816 CKEAAEAYRASCHKLFPYALAF 837
>gb|AAQ91276.1| transcriptional coactivator tubedown-100 [Danio rerio]
gi|42627867|ref|NP_976066.1| NMDA receptor-regulated
gene 1b [Danio rerio]
Length = 863
Score = 363 bits (931), Expect = 1e-98
Identities = 198/507 (39%), Positives = 304/507 (59%), Gaps = 41/507 (8%)
Query: 12 LVCQRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLE 71
LV +R+PL FL GD FRE D Y+R +KG P +F+ L SLY K I+E+L++ E
Sbjct: 291 LVPRRLPLSFLSGDTFRECLDRYLRMNFSKGCPPVFTTLRSLYQDKEKVSIIEELVVGFE 350
Query: 72 HSIRTSGQYPGSME--KEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDL 129
S+R+ + + +EPP+TL+W + LAQHYD+ GQ+ +A+ I+ AIE TPT+I+L
Sbjct: 351 TSLRSCRMFSPDDDGKEEPPTTLLWVQYFLAQHYDQIGQHSMALENINAAIESTPTLIEL 410
Query: 130 YSVKSRILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTK 189
+ +K++I KHAG+ AA + DEA+ +D ADR++NS+C K +L+A V AE+ FT+
Sbjct: 411 FLIKAKIYKHAGNIREAARWMDEAQALDTADRFINSKCAKYLLKAGLVKEAEEMCAKFTR 470
Query: 190 EG-DQHNNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCL 248
EG NL++MQCMW++ ++ G ALKK +E+H+ +I +DQFDFH+YC+
Sbjct: 471 EGASAVENLNEMQCMWFQTECALAYKSMKKCGEALKKCHEIERHFVEITDDQFDFHTYCM 530
Query: 249 RKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSPPKSTAEE-DEEMSKLLPA 307
RKMTLR+Y+++LK +D L H ++ KA+ AI+ Y+ LHD+P ++E + L
Sbjct: 531 RKMTLRSYVDLLKLEDVLRMHPFYFKASHTAIQIYLNLHDNPLTDDSKELQADTGNLTDK 590
Query: 308 QKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHG-------EKLL 360
+ KK+R KQR+A+ +A+ E+KN E K+ K+ + D + G EKL
Sbjct: 591 ELKKLRNKQRRAQKKAQLEEEKKNAEKEKQ--LKNQKKKKEDDDEEIGGPKEELIPEKLA 648
Query: 361 QVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEHPDS 420
+V++PL EA+K+L L+ + +ETHLL+FE+Y RK+K LL Q+VK+ +D +HP
Sbjct: 649 RVENPLDEAVKFLTPLKNLVKNKIETHLLAFEIYFRKEKFLLMLQSVKRAYAIDPDHPWL 708
Query: 421 HRCLV-------------------------CQLHEKTLFEANNSFLEKHKDSLMHRAAFA 455
H+CLV E N +FL KH +S+ HR A A
Sbjct: 709 HQCLVRFFKGVSESKDLADSVSMVLKQEISRLFGESNAKSFNQAFLTKHSNSIPHRVAAA 768
Query: 456 ETLYILDPNRKSEAVKL---IEESTNN 479
+ +Y LD + + +AV+L ++ES +N
Sbjct: 769 KMMYYLDSSTQKKAVELAIALDESLDN 795
>gb|AAF73953.2| acetyltransferase Tubedown-1 [Mus musculus]
Length = 594
Score = 362 bits (929), Expect = 2e-98
Identities = 205/565 (36%), Positives = 330/565 (58%), Gaps = 61/565 (10%)
Query: 12 LVCQRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLE 71
LV +++PL+FL G+KF+E D ++R +KG P +F+ L SLY K I+E+L++ E
Sbjct: 19 LVPRKLPLNFLSGEKFKECLDRFLRMNFSKGCPPVFNTLRSLYRDKEKVAIVEELVVGYE 78
Query: 72 HSIRTSGQYPGSME--KEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDL 129
S+++ + + + +EPP+TL+W + LAQHYD+ GQ +A+ I+ AIE TPT+I+L
Sbjct: 79 TSLKSCRLFNPNDDGKEEPPTTLLWVQYYLAQHYDKIGQPSIALEYINTAIESTPTLIEL 138
Query: 130 YSVKSRILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTK 189
+ VK++I KHAG+ AA + DEA+ +D ADR++NS+C K ML+A+ + AE+ FT+
Sbjct: 139 FLVKAKIYKHAGNIKEAARWMDEAQALDTADRFINSKCAKYMLKANLIKEAEEMCSKFTR 198
Query: 190 EGDQH-NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCL 248
EG NL++MQCMW++ +++ G ALKK +E+H+ +I +DQFDFH+YC+
Sbjct: 199 EGTSAVENLNEMQCMWFQTECAQAYKAMNKFGEALKKCHEIERHFIEITDDQFDFHTYCM 258
Query: 249 RKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSP-PKSTAEEDEEMSKLLPA 307
RK+TLR+Y+++LK +D L H ++ KAA AI Y+KLHD+P E + + + +
Sbjct: 259 RKITLRSYVDLLKLEDVLRQHPFYFKAARIAIEIYLKLHDNPLTDENKEHEADTANMSDK 318
Query: 308 QKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHG-------EKLL 360
+ KK+R KQR+A+ +A+ E+KN E ++ K+ D + G EKL
Sbjct: 319 ELKKLRNKQRRAQKKAQIEEEKKNAEKEKP--QRNPKKKKDDDDEEIGGPKEELIPEKLA 376
Query: 361 QVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEHPDS 420
+V+ PL EAIK+L L+ + +ETHL +FE+Y RK+K LL Q+VK+ +D+ HP
Sbjct: 377 KVETPLEEAIKFLTPLKNLVKNKIETHLFAFEIYFRKEKFLLMLQSVKRAFAIDSSHPWL 436
Query: 421 HRCLVCQLHE-------------------KTLFEA------NNSFLEKHKDSLMHRAAFA 455
H C++ H LF A N +FL+++ DSL HR + A
Sbjct: 437 HECMIRLFHSVCESKDLPETVRTVLKQEMNRLFGATNPKNFNETFLKRNSDSLPHRLSAA 496
Query: 456 ETLYILDPNRKSEAVKLIEESTNNIVPR-------NGALG-PIREWKLKDCIAVHKLLGT 507
+ +Y LD + + A++L +T + +P +G++G P + L+DC
Sbjct: 497 KMVYYLDSSSQKRAIEL--ATTLDGIPHQQKPSDLHGSVGKPCCDGSLRDC--------- 545
Query: 508 VLLDQDAALRWKVRCAEYFPYSRYF 532
++AA ++ C + FPY+ F
Sbjct: 546 ----KEAAEAYRASCHKLFPYALAF 566
>gb|AAF91333.1| putative N-terminal acetyltransferase [Xenopus laevis]
Length = 846
Score = 361 bits (927), Expect = 3e-98
Identities = 206/557 (36%), Positives = 329/557 (58%), Gaps = 47/557 (8%)
Query: 12 LVCQRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLE 71
LV +R+PL+FL G KFRE D Y+R +KG P +F+ L SLY+ K +I+E L++ E
Sbjct: 273 LVSRRLPLNFLSGLKFRECLDKYLRMNFSKGCPPVFNTLRSLYSDKEKVEIIEDLVVGYE 332
Query: 72 HSIRTSGQYPGSME--KEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDL 129
S+++ + + + +EPP+TL+W + LAQHYD+ GQ +A+ I+ AI+ TPT+I+L
Sbjct: 333 TSLKSCRLFNMNDDGKEEPPTTLLWVQYYLAQHYDKIGQQSIALEYINAAIDSTPTLIEL 392
Query: 130 YSVKSRILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTK 189
+ VK++I KH+G+ A + DEA+ +D ADR++NS+C K ML+A+ + AE+ FT+
Sbjct: 393 FLVKAKIYKHSGNIKEAVRWMDEAQALDTADRFINSKCAKYMLKANLIKEAEEMCSKFTR 452
Query: 190 EGDQH-NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCL 248
EG NL++MQCMW++ +++ G ALKK +E+H+ +I +DQFDFH+YC+
Sbjct: 453 EGTSAVENLNEMQCMWFQTECAQAYKSMNKYGEALKKCHEIERHFVEITDDQFDFHTYCM 512
Query: 249 RKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSP-PKSTAEEDEEMSKLLPA 307
RK+TLR+Y ++LK +D L H ++ KAA AI Y+KLHD+P E + + + +
Sbjct: 513 RKITLRSYADLLKLEDVLRQHPFYFKAARIAIEIYLKLHDNPLTDENKEHEADTANMSDK 572
Query: 308 QKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHG-------EKLL 360
+ KK+R KQR+A+ +A+ E+KN E ++ K+ + D + G EKL
Sbjct: 573 ELKKLRNKQRRAQKKAQLEEEKKNAEKEKQ--QRNQKKKKEDDDEEIGGPKEELIPEKLA 630
Query: 361 QVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEHPDS 420
+V++PL EAIK+L L+ + +ETHL +FE+Y RK K LL Q+VK+ +D HP
Sbjct: 631 KVENPLEEAIKFLTPLKNLVKNKIETHLYAFEIYFRKDKFLLMLQSVKRAYAIDPNHPWL 690
Query: 421 HRCLV---CQLHE-KTLFEA---------------------NNSFLEKHKDSLMHRAAFA 455
H+CL+ C + E K L E+ NNSFL+++ +S+ HR A A
Sbjct: 691 HQCLIRFFCAVSESKELNESVRTVLKQEMCRLFGETSPANFNNSFLKENINSIPHRLAAA 750
Query: 456 ETLYILDPNRKSEAVKLIEESTNNIVPRNGALGPIREWKLKDCIAVHKLLGTVLLDQDAA 515
+ +Y LD + + +V+L ++ NG+L + + L L D++AA
Sbjct: 751 KMMYYLDHSSQKRSVELGTSLDESLC--NGSLQTCTD-------VLEALRDGSLGDKEAA 801
Query: 516 LRWKVRCAEYFPYSRYF 532
++V C + +PY+ F
Sbjct: 802 ECYRVSCHKLYPYALAF 818
>ref|NP_080108.1| NMDA receptor regulated 1-like [Mus musculus]
gi|30851380|gb|AAH52445.1| NMDA receptor regulated
1-like [Mus musculus] gi|57012971|sp|Q9DBB4|NARGL_MOUSE
NMDA receptor regulated 1-like protein (NARG1-like
protein) gi|12836718|dbj|BAB23782.1| unnamed protein
product [Mus musculus]
Length = 864
Score = 358 bits (918), Expect = 3e-97
Identities = 196/504 (38%), Positives = 295/504 (57%), Gaps = 34/504 (6%)
Query: 15 QRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLEHSI 74
+R+PL F G KFRE D ++RP +KG P LF+ L SLY K I+++L+ + E S+
Sbjct: 294 RRLPLSFAPGKKFRELMDKFLRPNFSKGCPPLFTTLKSLYCDTEKVSIIQELVTNYEASL 353
Query: 75 RTSGQYPG--SMEKEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDLYSV 132
+ +G + + EKEPP+TL+W + LAQHYD+ GQY LA+ I+ I TPT+I+L+ +
Sbjct: 354 KMNGYFSPYENGEKEPPTTLIWVQYFLAQHYDKLGQYFLALEYINAVIASTPTLIELFYM 413
Query: 133 KSRILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTKEGD 192
K++I KH G+ AA + DEA+ +D ADR++NS+C K ML+A+ + AE+ FT+EG
Sbjct: 414 KAKIYKHMGNLKEAAQWMDEAQSLDTADRFINSKCAKYMLRANMIKEAEEMCSRFTREGT 473
Query: 193 QH-NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCLRKM 251
NL++MQCMW+E ++ R G G ALKK VE+H+ +I +DQFDFH+YC+RKM
Sbjct: 474 SAMENLNEMQCMWFETECISAYQRLGRYGDALKKCHEVERHFLEITDDQFDFHTYCMRKM 533
Query: 252 TLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSP-PKSTAEEDEEMSKLLPAQKK 310
TLR Y+ +L+ +D L H+++ KAA AI Y+KLHD+P + ++D + L + K
Sbjct: 534 TLRAYVGLLRLEDALRRHTFYFKAARSAIEIYLKLHDNPLTNDSKQQDIDSENLSAKEMK 593
Query: 311 KMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHGEKLL-----QVDDP 365
KM KQR+A+ +AK E K+ E ++ KR + H E+L+ +VD+P
Sbjct: 594 KMLSKQRRAQKKAKVEEERKHTERERQQKNQKKKREEEEEVTSGHKEELIPEKLERVDNP 653
Query: 366 LSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEHPDSHRCLV 425
L EAIK+L L+ + +S++THLL+FE+Y RK K LL Q+VK+ +++ +P H CL+
Sbjct: 654 LEEAIKFLTPLKTLAAESIDTHLLAFEIYFRKGKFLLMLQSVKRAFAIESNNPWLHECLI 713
Query: 426 -----CQLH--------------------EKTLFEANNSFLEKHKDSLMHRAAFAETLYI 460
H K L N FL + SL H A A+ +Y
Sbjct: 714 KFSKSVSNHSNLPDIVSKVLAQEMKKIFVNKDLHSFNEDFLRHNATSLQHLLAGAKMMYF 773
Query: 461 LDPNRKSEAVKLIEESTNNIVPRN 484
LD +R+ +A+ I +N
Sbjct: 774 LDKSRQEKAIATATRLDETIKNKN 797
>ref|XP_394637.2| PREDICTED: similar to ENSANGP00000006226 [Apis mellifera]
Length = 856
Score = 357 bits (917), Expect = 4e-97
Identities = 206/560 (36%), Positives = 319/560 (56%), Gaps = 53/560 (9%)
Query: 12 LVCQRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLE 71
L +R+ L++ D+F+ D Y+R L KGVP LF +L SLY K D ++ L+L+ +
Sbjct: 289 LAPRRLQLNYAVEDEFKTLVDRYLRRGLHKGVPPLFVNLRSLYTDQQKVDTIQSLVLEYK 348
Query: 72 HSIRTSGQYPGSME---KEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVID 128
+++ G + + +EP S L+WT + LAQHYD G E A+ +ID AIEHTPT+I+
Sbjct: 349 EALKAHGHFSDQEKDNPREPASALLWTYYYLAQHYDHLGLTEKALNEIDAAIEHTPTLIE 408
Query: 129 LYSVKSRILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFT 188
L+ K RI KHAG+ A + DEA+ +D ADRY+NS+C K ML+A+ + AE+T FT
Sbjct: 409 LFVTKGRIYKHAGNVQEAYKWLDEAQGLDTADRYINSKCAKYMLRANLIKEAEETCSKFT 468
Query: 189 KEGD-QHNNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYC 247
+EG NL++MQCMW + + ++ R G G ALKK V++H+++I EDQFDFH+YC
Sbjct: 469 REGVLAMENLNEMQCMWIQTEAANAYKRLGKYGEALKKCHEVDRHFSEIIEDQFDFHTYC 528
Query: 248 LRKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSP-PKSTAEEDEEMSKLLP 306
+RKMTLR+Y+ +L+ +D L +H ++ KAA A+ Y++LHD P P T ++ + L P
Sbjct: 529 MRKMTLRSYVGLLRLEDVLRAHPFYFKAAKCAMEVYLRLHDEPLPDPTQAQEIDTENLAP 588
Query: 307 AQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPH--------GEK 358
++ KK+R KQRK + +K E+ + A + + + DPD EK
Sbjct: 589 SELKKLRNKQRK---QRRKAELERQQAAQAQEKREQHNKSRQQTDPDLEQPTLDELIPEK 645
Query: 359 LLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEHP 418
L +++DPL +AIK+L+ LQ + + +ETHL++FE+Y RK + LL +++K+ LD +P
Sbjct: 646 LERIEDPLEQAIKFLQPLQDLASNRIETHLMAFEIYIRKGRTLLMLRSIKRAHGLDPNNP 705
Query: 419 DSHRCLVCQL------------------------HEKTLFEANNSFLEKHKDSLMHRAAF 454
D H CLV + T + N FL+K+++SL H
Sbjct: 706 DLHTCLVRFMLYINRSPLEGPVGEVVKRQTSGIYSASTATQLNAEFLKKNRNSLPHLLQG 765
Query: 455 AETLYILDPNRKSEAVKLIEESTNNIVPRNGALGPIREWKLKDCIAVHKLL--GTVLLDQ 512
A LY+LDP+ +++A+ LI TN + + L++C V + L G
Sbjct: 766 ARMLYVLDPSAQTKALSLI---TN--------IDGLESVTLQNCTKVLESLCNGDFGHCD 814
Query: 513 DAALRWKVRCAEYFPYSRYF 532
+ V+C +YFPY+ F
Sbjct: 815 STIADYMVKCHKYFPYATAF 834
>pir||JC7720 acetyltransferase (EC 2.3.1.-) 1, Xat-1 - African clawed frog
Length = 846
Score = 357 bits (916), Expect = 6e-97
Identities = 205/557 (36%), Positives = 327/557 (57%), Gaps = 47/557 (8%)
Query: 12 LVCQRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLE 71
LV +R+PL+FL G KFRE D Y+R +KG P +F+ L SLY+ K +I E L++ E
Sbjct: 273 LVSRRLPLNFLSGLKFRECLDKYLRMNFSKGCPPVFNTLRSLYSDKEKVEITEDLVVGYE 332
Query: 72 HSIRTSGQYPGSME--KEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDL 129
S+++ + + + +EPP+TL+W + LAQHYD+ GQ +A+ I+ AI+ TPT+I+L
Sbjct: 333 TSLKSCRLFNMNDDGKEEPPTTLLWVQYYLAQHYDKIGQQSIALEYINAAIDSTPTLIEL 392
Query: 130 YSVKSRILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTK 189
+ VK++I KH+G+ A + DEA+ +D ADR++NS+C K ML+A+ + AE+ FT+
Sbjct: 393 FLVKAKIYKHSGNIKEAVRWMDEAQALDTADRFINSKCAKYMLKANLIKEAEEMCSKFTR 452
Query: 190 EGDQH-NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCL 248
EG NL++MQCMW++ +++ G ALKK +E+H+ +I +DQFDFH+YC+
Sbjct: 453 EGTSAVENLNEMQCMWFQTECAQAYKSMNKYGEALKKCHEIERHFVEITDDQFDFHTYCM 512
Query: 249 RKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSP-PKSTAEEDEEMSKLLPA 307
RK+TLR+Y ++LK +D L H ++ KAA AI Y+KLHD+P E + + + +
Sbjct: 513 RKITLRSYADLLKLEDVLRQHPFYFKAARIAIEIYLKLHDNPLTDENKEHEADTANMSDK 572
Query: 308 QKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHG-------EKLL 360
+ KK+R KQR+A+ +A+ E+KN E ++ K+ + D + G EKL
Sbjct: 573 ELKKLRNKQRRAQKKAQLEEEKKNAEKEKQ--QRNQKKKKEDDDEEIGGPKEELIPEKLA 630
Query: 361 QVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEHPDS 420
+V++PL EAIK+L L+ + +ETHL +F +Y RK K LL Q+VK+ +D HP
Sbjct: 631 KVENPLEEAIKFLTPLKNLVKNKIETHLYAFFIYFRKDKFLLMLQSVKRAYAIDPNHPWL 690
Query: 421 HRCLV---CQLHE-KTLFEA---------------------NNSFLEKHKDSLMHRAAFA 455
H+CL+ C + E K L E+ NNSFL+++ +S+ HR A A
Sbjct: 691 HQCLIRFFCAVSESKELNESVRTVLKQEMCRLFGETSPANFNNSFLKENINSIPHRLAAA 750
Query: 456 ETLYILDPNRKSEAVKLIEESTNNIVPRNGALGPIREWKLKDCIAVHKLLGTVLLDQDAA 515
+ +Y LD + + +V+L ++ NG+L + + L L D++AA
Sbjct: 751 KMMYYLDHSSQKRSVELGTSLDESLC--NGSLQTCTD-------VLEALRDGSLGDKEAA 801
Query: 516 LRWKVRCAEYFPYSRYF 532
++V C + +PY+ F
Sbjct: 802 ECYRVSCHKLYPYALAF 818
>ref|NP_573384.1| CG12202-PA [Drosophila melanogaster] gi|7293585|gb|AAF48957.1|
CG12202-PA [Drosophila melanogaster]
gi|47271170|gb|AAT27255.1| SD09860p [Drosophila
melanogaster]
Length = 890
Score = 356 bits (914), Expect = 1e-96
Identities = 208/563 (36%), Positives = 320/563 (55%), Gaps = 57/563 (10%)
Query: 15 QRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLEHSI 74
+R+PL+ GD+FR D Y+R L KG+P LF ++ +L+ P +A ++E+L L ++
Sbjct: 294 RRLPLNIANGDEFRVVTDEYLRRGLRKGIPPLFVNVRTLHQIPERAAVIEELALQYFENL 353
Query: 75 RTSGQYP-----GSMEKEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDL 129
SG + + EP S L+WT LAQHYD + A+ I+ AI+HTPT+I+L
Sbjct: 354 TRSGHFSREDADAGIPVEPASALVWTALFLAQHYDYMRDTDRALEYINVAIDHTPTLIEL 413
Query: 130 YSVKSRILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTK 189
K RI KHAGD A + +EA+ MD ADRY+NS+C K ML+A+ V AE+ FT+
Sbjct: 414 LITKGRIFKHAGDPVEAYVWLEEAQSMDTADRYINSKCAKYMLRANMVQEAEEICAKFTR 473
Query: 190 EG-DQHNNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCL 248
EG +NL++MQCMW++ ++ R G G +LKK VE+H+A+I EDQFDFH+YC+
Sbjct: 474 EGVSAMDNLNEMQCMWFQTECALAYQRMGRWGESLKKCHEVERHFAEIVEDQFDFHTYCM 533
Query: 249 RKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSPPKS-TAEEDEEMSKLLPA 307
RKMTLR Y+ +L+ +D L H ++ KAA AI YI+L+D P KS T E+ ++ L P+
Sbjct: 534 RKMTLRAYVGLLRLEDVLRQHPFYFKAAKCAIEVYIRLYDKPLKSETTIEEIDIENLPPS 593
Query: 308 QKKKMRQKQRKAEARAK-KGAEEKNEELSASGVSKSGKRHVKPVDPDPH------GEKLL 360
+ KK+R KQRKA+ +A+ + A+ ++ KS ++ + DPD EKL
Sbjct: 594 ELKKLRSKQRKAKKKAELESAQAAQAQVKREQHQKSKQQANQETDPDAPQLDELVAEKLE 653
Query: 361 QVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEHPDS 420
+ DDPL +AI++LK LQ+ + + +ETHLL+FELY RK K+LL Q +++ +DA HP
Sbjct: 654 RTDDPLDKAIEFLKPLQQLAKERIETHLLAFELYYRKNKLLLMLQCIRRARAVDASHPVI 713
Query: 421 HRCLVC----------------------------QLHEKTLFEANNSFLEKHKDSLMHRA 452
H C++ + KT + N+ F+ KH S++H
Sbjct: 714 HSCIIRFVKSLTSAAKEQPFNEHVQQVLEKATKELIGSKTPQQLNDEFIAKHNASILHLY 773
Query: 453 AFAETLYILDPNRKSEAVKLIEESTNNIVPRNGALGPIREWKLKDCIAVHKLL--GTVLL 510
A +LY LD ++K+ A+KL+ + + +L++ ++ L G V
Sbjct: 774 EGARSLYELDNSKKAAAIKLVTSFN------------LAKLRLEEATKIYTALRDGDVFG 821
Query: 511 DQDA-ALRWKVRCAEYFPYSRYF 532
+ +A A ++ C + F Y+R F
Sbjct: 822 ECEAEAASYQQACHQRFQYARIF 844
>gb|AAH45491.2| NMDA receptor-regulated gene 1 [Danio rerio]
Length = 867
Score = 354 bits (909), Expect = 4e-96
Identities = 203/561 (36%), Positives = 323/561 (57%), Gaps = 53/561 (9%)
Query: 12 LVCQRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLE 71
LV +R+PL FL G+KFRE D Y+R +KG P +F+ L SLY+H K I+E+L++ +
Sbjct: 291 LVPRRLPLSFLTGEKFRECLDRYLRMNFSKGCPPVFTTLKSLYHHKDKVAIIEELVVGYD 350
Query: 72 HSIRTSGQYPGSMEK--EPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDL 129
S++T + + + EPP+TL+W + LAQH++ G++ +A+ I+ AIE TPT+I+L
Sbjct: 351 KSLKTCKMFNQNDDGKIEPPTTLLWVQYFLAQHFNHIGKHTVALEYINTAIESTPTLIEL 410
Query: 130 YSVKSRILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTK 189
+ +K++I KHAG+ AA + DEA+ +D ADR++NS+C K +L+A + AE+ FT+
Sbjct: 411 FLIKAKIYKHAGNIREAARWMDEAQALDTADRFINSKCAKYLLKAGLIKEAEEMCSKFTR 470
Query: 190 EGDQH-NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCL 248
EG NL++MQCMW++ ++ G ALKK +E+H+ +I +DQFDFH+YC+
Sbjct: 471 EGTSAVENLNEMQCMWFQTECALAYKSLNKYGEALKKCHEIERHFVEITDDQFDFHTYCM 530
Query: 249 RKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSPPKSTAEEDE-EMSKLLPA 307
RKMTLR+Y+++L +D L H ++ KAA AI Y+ LHD+P +E + + + L
Sbjct: 531 RKMTLRSYVDLLNLEDVLRMHPFYFKAARTAIEIYLSLHDNPLSDDNKESQADNANLTDK 590
Query: 308 QKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHG--------EKL 359
+ KK+R KQR+A+ +A+ E+KN E ++ K K D + G EKL
Sbjct: 591 ELKKLRNKQRRAQKKAQLEEEKKNAEKEKQLKNQKKK---KEEDEEEIGGPKEELIPEKL 647
Query: 360 LQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEHPD 419
++++PL EA+K+L L+ + +ETHLL+FE+Y RK+K LL Q+VK+ LD +HP
Sbjct: 648 AKIENPLEEAVKFLTPLKNLVKNKIETHLLAFEIYLRKEKYLLMLQSVKRAYSLDPDHPW 707
Query: 420 SHRCLVCQLH----EKTLFEA---------------------NNSFLEKHKDSLMHRAAF 454
H+CLV K + EA N +FL KH +S+ HR A
Sbjct: 708 LHQCLVRFFKGVSDSKDMPEAVQTVLKQEITKLFGESDPKTFNKNFLSKHANSIPHRVAA 767
Query: 455 AETLYILDPNRKSEAVKLIEESTNNIVPRNGALGPIREWKLKDCIAVHKLL--GTVLLDQ 512
A+ + LD + + +A +L ++ R+ L+ C V + L G++ Q
Sbjct: 768 AKMMVFLDSSSEGKAAELATSLNESLTDRS----------LQTCAEVLQALRDGSLGSQQ 817
Query: 513 DAALR-WKVRCAEYFPYSRYF 532
+ ++V C +PY+ F
Sbjct: 818 EKFTESYRVSCHGLYPYAVAF 838
>gb|AAH57466.1| NMDA receptor-regulated gene 1 [Danio rerio]
gi|41055289|ref|NP_956940.1| NMDA receptor-regulated
gene 1 [Danio rerio]
Length = 867
Score = 354 bits (908), Expect = 5e-96
Identities = 203/561 (36%), Positives = 322/561 (57%), Gaps = 53/561 (9%)
Query: 12 LVCQRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLE 71
LV +R+PL FL G+KFRE D Y+R +KG P +F+ L SLY H K I+E+L++ +
Sbjct: 291 LVPRRLPLSFLTGEKFRECLDRYLRMNFSKGCPPVFTTLKSLYRHKDKVAIIEELVVGYD 350
Query: 72 HSIRTSGQYPGSMEK--EPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDL 129
S++T + + + EPP+TL+W + LAQH++ G++ +A+ I+ AIE TPT+I+L
Sbjct: 351 KSLKTCKMFNQNDDGKIEPPTTLLWVQYFLAQHFNHIGKHTVALEYINTAIESTPTLIEL 410
Query: 130 YSVKSRILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTK 189
+ +K++I KHAG+ AA + DEA+ +D ADR++NS+C K +L+A + AE+ FT+
Sbjct: 411 FLIKAKIYKHAGNIREAARWMDEAQALDTADRFINSKCAKYLLKAGLIKEAEEMCSKFTR 470
Query: 190 EGDQH-NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCL 248
EG NL++MQCMW++ ++ G ALKK +E+H+ +I +DQFDFH+YC+
Sbjct: 471 EGTSAVENLNEMQCMWFQTECALAYKSLNKYGEALKKCHEIERHFVEITDDQFDFHTYCM 530
Query: 249 RKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSPPKSTAEEDE-EMSKLLPA 307
RKMTLR+Y+++L +D L H ++ KAA AI Y+ LHD+P +E + + + L
Sbjct: 531 RKMTLRSYVDLLNLEDVLRMHPFYFKAARTAIEIYLSLHDNPLSDDNKESQADNANLTDK 590
Query: 308 QKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHG--------EKL 359
+ KK+R KQR+A+ +A+ E+KN E ++ K K D + G EKL
Sbjct: 591 ELKKLRNKQRRAQKKAQLEEEKKNAEKEKQLKNQKKK---KEEDEEEIGGPKEELIPEKL 647
Query: 360 LQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEHPD 419
++++PL EA+K+L L+ + +ETHLL+FE+Y RK+K LL Q+VK+ LD +HP
Sbjct: 648 AKIENPLEEAVKFLTPLKNLVKNKIETHLLAFEIYLRKEKYLLMLQSVKRAYSLDPDHPW 707
Query: 420 SHRCLVCQLH----EKTLFEA---------------------NNSFLEKHKDSLMHRAAF 454
H+CLV K + EA N +FL KH +S+ HR A
Sbjct: 708 LHQCLVRFFKGVSDSKDMPEAVQTVLKQEITKLFGESDPKTFNKNFLSKHANSIPHRVAA 767
Query: 455 AETLYILDPNRKSEAVKLIEESTNNIVPRNGALGPIREWKLKDCIAVHKLL--GTVLLDQ 512
A+ + LD + + +A +L ++ R+ L+ C V + L G++ Q
Sbjct: 768 AKMMVFLDSSSEGKAAELATSLNESLTDRS----------LQTCAEVLQALRDGSLGSQQ 817
Query: 513 DAALR-WKVRCAEYFPYSRYF 532
+ ++V C +PY+ F
Sbjct: 818 EKFTESYRVSCHGLYPYAVAF 838
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.133 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 968,520,074
Number of Sequences: 2540612
Number of extensions: 40772505
Number of successful extensions: 158224
Number of sequences better than 10.0: 281
Number of HSP's better than 10.0 without gapping: 112
Number of HSP's successfully gapped in prelim test: 174
Number of HSP's that attempted gapping in prelim test: 156993
Number of HSP's gapped (non-prelim): 727
length of query: 583
length of database: 863,360,394
effective HSP length: 134
effective length of query: 449
effective length of database: 522,918,386
effective search space: 234790355314
effective search space used: 234790355314
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0226.2


