

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.11
(231 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
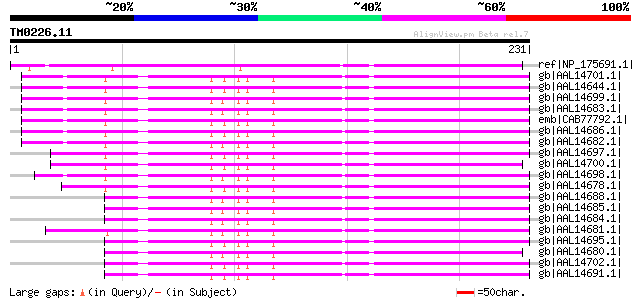
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_175691.1| 2-oxoglutarate-dependent dioxygenase, putative ... 231 9e-60
gb|AAL14701.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis... 180 2e-44
gb|AAL14644.1| AOP1.2 [Arabidopsis thaliana] 180 2e-44
gb|AAL14699.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis... 177 2e-43
gb|AAL14683.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis... 177 2e-43
emb|CAB77792.1| putative oxidoreductase [Arabidopsis thaliana] g... 176 6e-43
gb|AAL14686.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis... 176 6e-43
gb|AAL14682.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis... 173 3e-42
gb|AAL14697.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis... 171 1e-41
gb|AAL14700.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis... 170 3e-41
gb|AAL14698.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis... 170 3e-41
gb|AAL14678.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis... 169 4e-41
gb|AAL14688.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis... 168 1e-40
gb|AAL14685.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis... 168 1e-40
gb|AAL14684.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis... 168 1e-40
gb|AAL14681.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis... 168 1e-40
gb|AAL14695.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis... 167 2e-40
gb|AAL14680.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis... 167 2e-40
gb|AAL14702.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis... 167 3e-40
gb|AAL14691.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis... 167 3e-40
>ref|NP_175691.1| 2-oxoglutarate-dependent dioxygenase, putative [Arabidopsis
thaliana] gi|12324633|gb|AAG52269.1| putative
oxidoreductase; 38288-39393 [Arabidopsis thaliana]
gi|25285705|pir||D96569 probable oxidoreductase,
38288-39393 [imported] - Arabidopsis thaliana
Length = 317
Score = 231 bits (590), Expect = 9e-60
Identities = 131/295 (44%), Positives = 167/295 (56%), Gaps = 71/295 (24%)
Query: 1 MGSETTL---KLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEYMQL------------- 44
MGSET L +LPVIDF+N NL P W+ ++ V KAL +Y
Sbjct: 1 MGSETPLLPLRLPVIDFSN-KNLKPGEPEWDLTRADVQKALQDYGYFEASFDRIPFELRK 59
Query: 45 -------------------NVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKIL 85
NVSKKP+ GY GQ+P +PL++SMGIDD+++ EKV++ T+ L
Sbjct: 60 SVFGALEELFDLPLQTKLRNVSKKPFHGYVGQYPMVPLYESMGIDDSDIAEKVDAFTEKL 119
Query: 86 WPNGNPSFRYASQPIT--------------------------------YFLKTMKYKGPQ 113
WP GN SF Q + Y L+ MKYKGP
Sbjct: 120 WPQGNISFSTTIQSFSKKLSELDITIRRMIMESFGLDKYIDEHLHSTNYLLRVMKYKGPD 179
Query: 114 TSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHA 173
T +TKVGL+ HTD I+TIL+Q+ V+GLEV TKD + WI KP+ +S F VM+ D +A
Sbjct: 180 TEETKVGLNAHTDKNIVTILYQNHVEGLEVQTKD-KNWIKVKPTQDS--FTVMIGDSLYA 236
Query: 174 WSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFD 228
NGRLHSP+HRVMMTG E RYS LF +PK G I+ +P+ELVDEEHP L+KPFD
Sbjct: 237 LLNGRLHSPYHRVMMTGTETRYSLGLFSIPKAGHIVSSPDELVDEEHPRLFKPFD 291
>gb|AAL14701.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 300
Score = 180 bits (457), Expect = 2e-44
Identities = 122/296 (41%), Positives = 156/296 (52%), Gaps = 78/296 (26%)
Query: 6 TLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------------ 41
+ +LPVIDF++ N G+S W+ VK+ V KAL +Y
Sbjct: 1 SFQLPVIDFSDQNLKPGSS-KWDEVKADVLKALEDYGCFEAFFDKLSVELNRSVFEAMED 59
Query: 42 --------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPS 92
Q NVS KP+ GY L++S+GIDDANV EKV T+ LWP+ GN S
Sbjct: 60 LFELPIPTKQRNVSSKPFHGYLCH----NLYESLGIDDANVLEKVNDFTQQLWPDHGNKS 115
Query: 93 FR-----YASQPI--------------------------TYFL-KTMKYKGPQTSD---- 116
++ Q + TY+L + MKY P D
Sbjct: 116 ISETIHLFSEQLVELDLMVRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDE 175
Query: 117 -TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 175
TK+GL +HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A
Sbjct: 176 ETKLGLRSHTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALL 232
Query: 176 NGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
NGRLHSP+HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 233 NGRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 288
>gb|AAL14644.1| AOP1.2 [Arabidopsis thaliana]
Length = 321
Score = 180 bits (457), Expect = 2e-44
Identities = 122/296 (41%), Positives = 156/296 (52%), Gaps = 78/296 (26%)
Query: 6 TLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------------ 41
+ +LPVIDF++ N G+S W+ VK+ V KAL +Y
Sbjct: 11 SFQLPVIDFSDQNLKPGSS-KWDEVKADVLKALEDYGCFEAFFDKLSVELNRSVFEAMED 69
Query: 42 --------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPS 92
Q NVS KP+ GY L++S+GIDDANV EKV T+ LWP+ GN S
Sbjct: 70 LFELPIPTKQRNVSSKPFHGYLCH----NLYESLGIDDANVLEKVNDFTQQLWPDHGNKS 125
Query: 93 FR-----YASQPI--------------------------TYFL-KTMKYKGPQTSD---- 116
++ Q + TY+L + MKY P D
Sbjct: 126 ISETIHLFSEQLVELDLMVRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDE 185
Query: 117 -TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 175
TK+GL +HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A
Sbjct: 186 ETKLGLRSHTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALL 242
Query: 176 NGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
NGRLHSP+HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 243 NGRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 298
>gb|AAL14699.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 300
Score = 177 bits (449), Expect = 2e-43
Identities = 120/295 (40%), Positives = 155/295 (51%), Gaps = 77/295 (26%)
Query: 6 TLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------------ 41
+ +LPVIDF++ N G+S W+ V + V KAL +Y
Sbjct: 3 SFQLPVIDFSDQNLKPGSS-KWDEVTADVLKALEDYGCFEASFDKLSVELNRSVFEAMED 61
Query: 42 --------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPS 92
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S
Sbjct: 62 LFELPIPTKQRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKS 117
Query: 93 F-----------------------------RYASQPI--TYFL-KTMKYKGPQTSD---- 116
+Y + + TY+L + MKY P D
Sbjct: 118 ISETIHLFSEQLVELDLMVRRMIMESFGIEKYIDEHLNSTYYLTRLMKYTSPPDDDDDEE 177
Query: 117 TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSN 176
TK+GL +HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A N
Sbjct: 178 TKLGLRSHTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLN 234
Query: 177 GRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
GRLHSP+HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 235 GRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 289
>gb|AAL14683.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 320
Score = 177 bits (449), Expect = 2e-43
Identities = 120/295 (40%), Positives = 155/295 (51%), Gaps = 77/295 (26%)
Query: 6 TLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------------ 41
+ +LPVIDF++ N G+S W+ V + V KAL +Y
Sbjct: 11 SFQLPVIDFSDQNLKPGSS-KWDEVTADVLKALEDYGCFEASFDKLSVELNRSVFEAMED 69
Query: 42 --------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPS 92
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S
Sbjct: 70 LFELPIPTKQRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKS 125
Query: 93 F-----------------------------RYASQPI--TYFL-KTMKYKGPQTSD---- 116
+Y + + TY+L + MKY P D
Sbjct: 126 ISETIHLFSEQLVELDLMVRRMIMESFGIEKYIDEHLNSTYYLTRLMKYTSPPDDDDDEE 185
Query: 117 TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSN 176
TK+GL +HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A N
Sbjct: 186 TKLGLRSHTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLN 242
Query: 177 GRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
GRLHSP+HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 243 GRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 297
>emb|CAB77792.1| putative oxidoreductase [Arabidopsis thaliana]
gi|15236243|ref|NP_192216.1| 2-oxoglutarate-dependent
dioxygenase (AOP1.2) [Arabidopsis thaliana]
gi|16118885|gb|AAL14643.1| AOP1.1 [Arabidopsis thaliana]
gi|3924597|gb|AAC79098.1| putative oxidoreductase
[Arabidopsis thaliana] gi|7487987|pir||T01386
oxidoreductase homolog T4I9.5 - Arabidopsis thaliana
Length = 322
Score = 176 bits (445), Expect = 6e-43
Identities = 120/297 (40%), Positives = 155/297 (51%), Gaps = 79/297 (26%)
Query: 6 TLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------------ 41
+ +LPVIDF++ N G+S W+ V + V KAL +Y
Sbjct: 11 SFQLPVIDFSDQNLKPGSS-KWDEVTADVLKALEDYGCFEASFDKLSVELNRSVFEAMED 69
Query: 42 --------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPS 92
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S
Sbjct: 70 LFELPIPTKQRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKS 125
Query: 93 FR-----YASQPI--------------------------TYFL-KTMKYKGPQTSD---- 116
++ Q + TY+L + MKY P D
Sbjct: 126 ISETIHLFSEQLVELDLMVRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDD 185
Query: 117 --TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAW 174
TK+GL +HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A
Sbjct: 186 EETKLGLRSHTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCAL 242
Query: 175 SNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
NGRLHSP+HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 243 LNGRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 299
>gb|AAL14686.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 309
Score = 176 bits (445), Expect = 6e-43
Identities = 120/297 (40%), Positives = 155/297 (51%), Gaps = 79/297 (26%)
Query: 6 TLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------------ 41
+ +LPVIDF++ N G+S W+ V + V KAL +Y
Sbjct: 5 SFQLPVIDFSDQNLKPGSS-KWDEVTADVLKALEDYGCFEASFDKLSVELNRSVFEAMED 63
Query: 42 --------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPS 92
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S
Sbjct: 64 LFELPIPTKQRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKS 119
Query: 93 FR-----YASQPI--------------------------TYFL-KTMKYKGPQTSD---- 116
++ Q + TY+L + MKY P D
Sbjct: 120 ISETIHLFSEQLVELDLMVRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDD 179
Query: 117 --TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAW 174
TK+GL +HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A
Sbjct: 180 EETKLGLRSHTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCAL 236
Query: 175 SNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
NGRLHSP+HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 237 LNGRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 293
>gb|AAL14682.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 301
Score = 173 bits (439), Expect = 3e-42
Identities = 119/295 (40%), Positives = 154/295 (51%), Gaps = 77/295 (26%)
Query: 6 TLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------------ 41
+ +LPVIDF++ N G+S W+ V + V KAL +Y
Sbjct: 4 SFQLPVIDFSDQNLKPGSS-KWDEVTADVLKALEDYGCFEASFDKLSVELNRSVFEAMED 62
Query: 42 --------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPS 92
Q NVS K + GY L++S+GI+DANV EKV T+ LWP+ GN S
Sbjct: 63 LFELPIPTKQRNVSSKLFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKS 118
Query: 93 F-----------------------------RYASQPI--TYFL-KTMKYKGPQTSD---- 116
+Y + + TY+L + MKY P D
Sbjct: 119 ISETIHLFSEQLVELDLMVRRMIMESFGIEKYIDEHLNSTYYLTRLMKYTSPPDDDDDEE 178
Query: 117 TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSN 176
TK+GL +HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A N
Sbjct: 179 TKLGLRSHTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLN 235
Query: 177 GRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
GRLHSP+HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 236 GRLHSPYHRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 290
>gb|AAL14697.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
gi|16118996|gb|AAL14696.1| 2-oxoglutarate-dependent
dioxygenase [Arabidopsis thaliana]
Length = 278
Score = 171 bits (433), Expect = 1e-41
Identities = 116/283 (40%), Positives = 145/283 (50%), Gaps = 77/283 (27%)
Query: 19 NLGGTSPNWEAVKSKVHKALVEY--------------------------------MQLNV 46
NL S W+ VK+ V KAL +Y Q NV
Sbjct: 2 NLKPGSSKWDEVKADVLKALEDYGCFEAFFDKLSVELNRSVFEAMEDLFELPIPTKQRNV 61
Query: 47 SKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPSFR-----YASQPI 100
S KP+ GY L++S+GIDDANV EKV T+ LWP+ GN S ++ Q +
Sbjct: 62 SSKPFHGYLCH----NLYESLGIDDANVLEKVNDFTQQLWPDHGNKSISETIHLFSEQLV 117
Query: 101 --------------------------TYFL-KTMKYKGPQTSD-----TKVGLDTHTDTA 128
TY+L + MKY P D TK+GL +HTD
Sbjct: 118 ELDLMVRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDEETKLGLRSHTDKN 177
Query: 129 ILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMM 188
I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A NGRLHSP+HRV+M
Sbjct: 178 IITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLNGRLHSPYHRVIM 234
Query: 189 TGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
TG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 235 TGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 277
>gb|AAL14700.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 276
Score = 170 bits (430), Expect = 3e-41
Identities = 115/280 (41%), Positives = 144/280 (51%), Gaps = 77/280 (27%)
Query: 19 NLGGTSPNWEAVKSKVHKALVEY--------------------------------MQLNV 46
NL S W+ VK+ V KAL +Y Q NV
Sbjct: 2 NLKPGSSKWDEVKADVLKALEDYGCFEAFFDKLSVELNRSVFEAMEDLFELPIPTKQRNV 61
Query: 47 SKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPSFR-----YASQPI 100
S KP+ GY L++S+GIDDANV EKV T+ LWP+ GN S ++ Q +
Sbjct: 62 SSKPFHGYLCH----NLYESLGIDDANVLEKVNDFTQQLWPDHGNKSISETIHLFSEQLV 117
Query: 101 --------------------------TYFL-KTMKYKGPQTSD-----TKVGLDTHTDTA 128
TY+L + MKY P D TK+GL +HTD
Sbjct: 118 ELDLMVRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDEETKLGLRSHTDKN 177
Query: 129 ILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMM 188
I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A NGRLHSP+HRV+M
Sbjct: 178 IITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLNGRLHSPYHRVIM 234
Query: 189 TGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFD 228
TG + RYS LF +PK G II +PEELVD+EHP ++KPF+
Sbjct: 235 TGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFE 274
>gb|AAL14698.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 301
Score = 170 bits (430), Expect = 3e-41
Identities = 117/292 (40%), Positives = 150/292 (51%), Gaps = 80/292 (27%)
Query: 12 IDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------------------ 41
IDF++ G+S W+ VK+ V KAL Y
Sbjct: 1 IDFSDQTLKPGSS-KWDEVKADVRKALENYGCFEASFDKLSVELNRSVFEAMEDLFELPI 59
Query: 42 --MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPSF----- 93
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S
Sbjct: 60 PTKQRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKSISETIH 115
Query: 94 RYASQPI--------------------------TYFL-KTMKYKGPQTSD-------TKV 119
R++ + + TY+L + MKY P D TK+
Sbjct: 116 RFSEKLVELDLMVRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDDDEETKL 175
Query: 120 GLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRL 179
GL +HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A NGRL
Sbjct: 176 GLRSHTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLNGRL 232
Query: 180 HSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
HSP+HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 233 HSPYHRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 284
>gb|AAL14678.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 272
Score = 169 bits (429), Expect = 4e-41
Identities = 114/278 (41%), Positives = 143/278 (51%), Gaps = 77/278 (27%)
Query: 24 SPNWEAVKSKVHKALVEY--------------------------------MQLNVSKKPY 51
S W+ VK+ V KAL +Y Q NVS KP+
Sbjct: 1 SSKWDEVKADVLKALEDYGCFEAFFDKLSVELNRSVFEAMEDLFELPIPTKQRNVSSKPF 60
Query: 52 FGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPSFR-----YASQPI----- 100
GY L++S+GIDDANV EKV T+ LWP+ GN S ++ Q +
Sbjct: 61 HGYLCH----NLYESLGIDDANVLEKVNDFTQQLWPDHGNKSISETIHLFSEQLVELDLM 116
Query: 101 ---------------------TYFL-KTMKYKGPQTSD-----TKVGLDTHTDTAILTIL 133
TY+L + MKY P D TK+GL +HTD I+TIL
Sbjct: 117 VRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDEETKLGLRSHTDKNIITIL 176
Query: 134 FQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEA 193
Q QVDGLEV TKD KWI KPS +S +VM+ D A NGRLHSP+HRV+MTG +
Sbjct: 177 HQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLNGRLHSPYHRVIMTGKKT 233
Query: 194 RYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 234 RYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 271
>gb|AAL14688.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 271
Score = 168 bits (426), Expect = 1e-40
Identities = 104/226 (46%), Positives = 131/226 (57%), Gaps = 44/226 (19%)
Query: 43 QLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPSF-------- 93
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S
Sbjct: 52 QRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKSISETIHLFS 107
Query: 94 ---------------------RYASQPI--TYFL-KTMKYKGPQTSD----TKVGLDTHT 125
+Y + + TY+L + MKY P D TK+GL +HT
Sbjct: 108 EQLVELDLMVRRMIMESFGIEKYIDEHLNSTYYLTRLMKYTSPPDDDDDEETKLGLRSHT 167
Query: 126 DTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHR 185
D I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A NGRLHSP+HR
Sbjct: 168 DKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLNGRLHSPYHR 224
Query: 186 VMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
V+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 225 VIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 270
>gb|AAL14685.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 285
Score = 168 bits (426), Expect = 1e-40
Identities = 104/226 (46%), Positives = 131/226 (57%), Gaps = 44/226 (19%)
Query: 43 QLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPSF-------- 93
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S
Sbjct: 56 QRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKSISETIHLFS 111
Query: 94 ---------------------RYASQPI--TYFL-KTMKYKGPQTSD----TKVGLDTHT 125
+Y + + TY+L + MKY P D TK+GL +HT
Sbjct: 112 EQLVELDLMVRRMIMESFGIEKYIDEHLNSTYYLTRLMKYTSPPDDDDDEETKLGLRSHT 171
Query: 126 DTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHR 185
D I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A NGRLHSP+HR
Sbjct: 172 DKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLNGRLHSPYHR 228
Query: 186 VMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
V+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 229 VIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 274
>gb|AAL14684.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 276
Score = 168 bits (426), Expect = 1e-40
Identities = 104/226 (46%), Positives = 131/226 (57%), Gaps = 44/226 (19%)
Query: 43 QLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPSF-------- 93
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S
Sbjct: 57 QRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKSISETIHLFS 112
Query: 94 ---------------------RYASQPI--TYFL-KTMKYKGPQTSD----TKVGLDTHT 125
+Y + + TY+L + MKY P D TK+GL +HT
Sbjct: 113 EQLVELDLMVRRMIMESFGIEKYIDEHLNSTYYLTRLMKYTSPPDDDDDEETKLGLRSHT 172
Query: 126 DTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHR 185
D I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A NGRLHSP+HR
Sbjct: 173 DKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLNGRLHSPYHR 229
Query: 186 VMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
V+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 230 VIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 275
>gb|AAL14681.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 274
Score = 168 bits (426), Expect = 1e-40
Identities = 109/266 (40%), Positives = 147/266 (54%), Gaps = 58/266 (21%)
Query: 17 LNNLGGTSPNWEAVKSKVHKALVEYM-----------QLNVSKKPYFGYAGQFPNIPLFK 65
L N G +++ + ++++++ E M Q NVS KP+ GY L++
Sbjct: 15 LENYGCFEASFDKLSVELNRSVFEAMEDLFELPIPTKQRNVSSKPFHGYLCH----NLYE 70
Query: 66 SMGIDDANVYEKVESLTKILWPN-GNPSF-----RYASQPI------------------- 100
S+GI+DANV EKV T+ LWP+ GN S R++ + +
Sbjct: 71 SLGINDANVLEKVNDFTQQLWPDHGNKSISETIHRFSEKLVELDLMVRRMIMESFGIENY 130
Query: 101 -------TYFL-KTMKYKGPQTSD-------TKVGLDTHTDTAILTILFQSQVDGLEVLT 145
TY+L + MKY P D TK+GL +HTD I+TIL Q QVDGLEV T
Sbjct: 131 IDEHLNSTYYLTRLMKYTSPPDDDDDDDDEETKLGLRSHTDKNIITILHQYQVDGLEVKT 190
Query: 146 KDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKG 205
KD KWI KPS +S +VM+ D A NGRLHSP+HRV+MTG + RYS LF +PK
Sbjct: 191 KDD-KWIKVKPSQDS--VLVMVGDSLCALLNGRLHSPYHRVIMTGKKTRYSTGLFSIPKT 247
Query: 206 GCIIKAPEELVDEEHPLLYKPFDRED 231
G II +PEELVD+EHP ++KPF+ D
Sbjct: 248 GVIIDSPEELVDKEHPRIFKPFEYTD 273
>gb|AAL14695.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 272
Score = 167 bits (423), Expect = 2e-40
Identities = 104/227 (45%), Positives = 131/227 (56%), Gaps = 45/227 (19%)
Query: 43 QLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPSFR-----YA 96
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S ++
Sbjct: 52 QRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKSISETIHLFS 107
Query: 97 SQPI--------------------------TYFL-KTMKYKGPQTSD-----TKVGLDTH 124
Q + TY+L + MKY P D TK+GL +H
Sbjct: 108 EQLVELDLMVRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDEETKLGLRSH 167
Query: 125 TDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFH 184
TD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A NGRLHSP+H
Sbjct: 168 TDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLNGRLHSPYH 224
Query: 185 RVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
RV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 225 RVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 271
>gb|AAL14680.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 275
Score = 167 bits (423), Expect = 2e-40
Identities = 103/223 (46%), Positives = 130/223 (58%), Gaps = 44/223 (19%)
Query: 43 QLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPSF-------- 93
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S
Sbjct: 58 QRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKSISETIHLFS 113
Query: 94 ---------------------RYASQPI--TYFL-KTMKYKGPQTSD----TKVGLDTHT 125
+Y + + TY+L + MKY P D TK+GL +HT
Sbjct: 114 EQLVELDLMVRRMIMESFGIEKYIDEHLNSTYYLTRLMKYTSPPDDDDDEETKLGLRSHT 173
Query: 126 DTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHR 185
D I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A NGRLHSP+HR
Sbjct: 174 DKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLNGRLHSPYHR 230
Query: 186 VMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFD 228
V+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+
Sbjct: 231 VIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFE 273
>gb|AAL14702.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 279
Score = 167 bits (422), Expect = 3e-40
Identities = 104/228 (45%), Positives = 131/228 (56%), Gaps = 46/228 (20%)
Query: 43 QLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPSFR-----YA 96
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S ++
Sbjct: 58 QRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKSISETIHLFS 113
Query: 97 SQPI--------------------------TYFL-KTMKYKGPQTSD------TKVGLDT 123
Q + TY+L + MKY P D TK+GL +
Sbjct: 114 EQLVELDLMVRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDDEETKLGLRS 173
Query: 124 HTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPF 183
HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A NGRLHSP+
Sbjct: 174 HTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLNGRLHSPY 230
Query: 184 HRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 231 HRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 278
>gb|AAL14691.1| 2-oxoglutarate-dependent dioxygenase [Arabidopsis thaliana]
Length = 273
Score = 167 bits (422), Expect = 3e-40
Identities = 104/228 (45%), Positives = 131/228 (56%), Gaps = 46/228 (20%)
Query: 43 QLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN-GNPSFR-----YA 96
Q NVS KP+ GY L++S+GI+DANV EKV T+ LWP+ GN S ++
Sbjct: 52 QRNVSSKPFHGYLCH----NLYESLGINDANVLEKVNDFTQQLWPDHGNKSISETIHLFS 107
Query: 97 SQPI--------------------------TYFL-KTMKYKGPQTSD------TKVGLDT 123
Q + TY+L + MKY P D TK+GL +
Sbjct: 108 EQLVELDLMVRRMIMESFGIENYIDEHLNSTYYLTRLMKYTSPPDDDDDDDEETKLGLRS 167
Query: 124 HTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPF 183
HTD I+TIL Q QVDGLEV TKD KWI KPS +S +VM+ D A NGRLHSP+
Sbjct: 168 HTDKNIITILHQYQVDGLEVKTKDD-KWIKVKPSQDS--VLVMVGDSLCALLNGRLHSPY 224
Query: 184 HRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
HRV+MTG + RYS LF +PK G II +PEELVD+EHP ++KPF+ D
Sbjct: 225 HRVIMTGKKTRYSTGLFSIPKTGVIIDSPEELVDKEHPRIFKPFEYTD 272
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 426,023,079
Number of Sequences: 2540612
Number of extensions: 17599613
Number of successful extensions: 34532
Number of sequences better than 10.0: 1014
Number of HSP's better than 10.0 without gapping: 540
Number of HSP's successfully gapped in prelim test: 474
Number of HSP's that attempted gapping in prelim test: 32914
Number of HSP's gapped (non-prelim): 1145
length of query: 231
length of database: 863,360,394
effective HSP length: 124
effective length of query: 107
effective length of database: 548,324,506
effective search space: 58670722142
effective search space used: 58670722142
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0226.11


