

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0225.8
(423 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
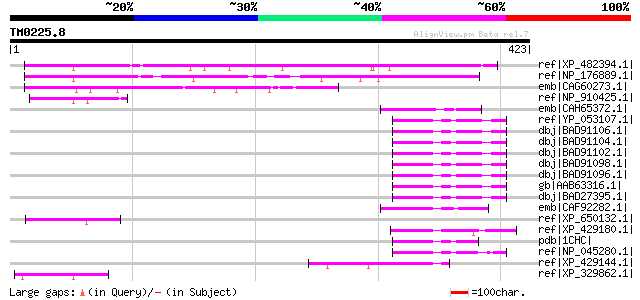
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_482394.1| unknown protein [Oryza sativa (japonica cultiva... 212 2e-53
ref|NP_176889.1| zinc finger (C3HC4-type RING finger) family pro... 205 2e-51
emb|CAG60273.1| unnamed protein product [Candida glabrata CBS138... 68 6e-10
ref|NP_910425.1| unknown protein [Oryza sativa (japonica cultiva... 57 8e-07
emb|CAH65372.1| hypothetical protein [Gallus gallus] gi|61098412... 55 5e-06
ref|YP_053107.1| transcriptional activator [Equid herpesvirus 1]... 54 7e-06
dbj|BAD91106.1| transcriptional activator [Equid herpesvirus 1] 54 7e-06
dbj|BAD91104.1| transcriptional activator [Equid herpesvirus 1] 54 7e-06
dbj|BAD91102.1| transcriptional activator [Equid herpesvirus 1] 54 7e-06
dbj|BAD91098.1| transcriptional activator [Equid herpesvirus 1] 54 7e-06
dbj|BAD91096.1| transcriptional activator [Equid herpesvirus 1] 54 7e-06
gb|AAB63316.1| contains a deletion of 399 base pairs as compared... 54 7e-06
dbj|BAD27395.1| transactivator protein [Equine herpesvirus 1] 54 7e-06
emb|CAF92282.1| unnamed protein product [Tetraodon nigroviridis] 53 2e-05
ref|XP_650132.1| hypothetical protein 244.t00016 [Entamoeba hist... 53 2e-05
ref|XP_429180.1| PREDICTED: hypothetical protein XP_429180 [Gall... 52 3e-05
pdb|1CHC| Equine Herpes Virus-1 (C3hc4, Or Ring Domain) (Nmr, ... 52 3e-05
ref|NP_045280.1| 63 [Equid herpesvirus 4] gi|2606010|gb|AAC59582... 52 3e-05
ref|XP_429144.1| PREDICTED: similar to p53-binding protein-3 [Ga... 52 3e-05
ref|XP_329862.1| hypothetical protein [Neurospora crassa] gi|289... 52 3e-05
>ref|XP_482394.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|37806204|dbj|BAC99707.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 531
Score = 212 bits (540), Expect = 2e-53
Identities = 151/461 (32%), Positives = 215/461 (45%), Gaps = 80/461 (17%)
Query: 13 IVATVTGYHGLERFNLIKLINYAGGSYIGSMSKSITHL---KFEGKKYDIARRLSIPVVN 69
+VATV+GYHG ER L++LI G SY+G+MS+SITHL + EGKKYDIARRL VV+
Sbjct: 10 VVATVSGYHGDERHRLVRLIAETGASYVGAMSRSITHLVCWRLEGKKYDIARRLRTRVVS 69
Query: 70 HRWIEDCIREGKRLPIDSYTLQSGQEVGPLLLEVPASFGPNSWIKNKVVSDGLCDIESER 129
HRW EDC++EG+RLP Y L+SG+E GP + E+P P S K + C E
Sbjct: 70 HRWFEDCLKEGRRLPEKPYMLESGEEAGP-VPELPTF--PRSRSKRNASMEDRCLKELPD 126
Query: 130 QNTVISFGAPSYVLEDS---CLMKKYEETSL-----------------HSSRRLKKVKRN 169
S+ V+ DS C +++ ++SL H R K++K
Sbjct: 127 DFCNTSYATDVLVVADSGSDCNHQRWSDSSLLKENFVGDRDNSKIGATHVKERRKRLKHA 186
Query: 170 ISCDNEVS---------TAAGPSRKEKKFVKKVVDEVVSDPVILDLSPDDRLCRMDRDRL 220
+ NE + A R E + ++ L DD R+ L
Sbjct: 187 QNSTNEDALDAEDNISRLMARQGRYESSYTSSRSASNQKGDLLKLLHNDDASMMRKRNSL 246
Query: 221 H------------TEAAATSSLSGGVDVNDLQENSSGPDARLCNQNRAMNGSSDGIEQIT 268
E+ SL+ D + + D R + R + D I
Sbjct: 247 MKKETRTKLAGYLIESCENGSLTDSFDEPQMSDTPPTEDRRKIRKTRLRQSTLDSIYDYG 306
Query: 269 DSNQIEDPQTVEQTSIDLCSSDGKF---------------------ID-------GDQVD 300
++++ + ++ +Q + +L S F ID GD
Sbjct: 307 EASEHDPEKSEDQENFELGESSRSFQSSDSSRQEPAFCTEKTNQGSIDIAADDDKGDDEK 366
Query: 301 NVAGSPIS----NEWSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPL 356
P S E SCVIC+ DFSST G+LPC H FC +CIQ+WA S GK+STCPL
Sbjct: 367 ATLEEPTSCQGQAELSCVICWTDFSSTRGILPCGHRFCYSCIQEWADSLSSRGKVSTCPL 426
Query: 357 CKASFAWYKIVEHAATADKELNSQTVPCDNLASVTFISMDQ 397
CK SFAW ++ A T+D+++ SQT+PC + ++ TFI D+
Sbjct: 427 CKTSFAWISKIDEAGTSDQKIYSQTIPC-STSTDTFIFDDR 466
>ref|NP_176889.1| zinc finger (C3HC4-type RING finger) family protein / BRCT
domain-containing protein [Arabidopsis thaliana]
gi|4204282|gb|AAD10663.1| Hypothetical protein
[Arabidopsis thaliana] gi|25404671|pir||G96695
hypothetical protein F5A8.9 [imported] - Arabidopsis
thaliana
Length = 453
Score = 205 bits (521), Expect = 2e-51
Identities = 139/390 (35%), Positives = 211/390 (53%), Gaps = 42/390 (10%)
Query: 13 IVATVTGYHGLERFNLIKLINYAGGSYIGSMSKSITHL---KFEGKKYDIARRLSIPVVN 69
+VATV+GYHG +RF LIKLI+++G SY+G+MS+SITHL KFEGKKYD+A++ VVN
Sbjct: 4 VVATVSGYHGSDRFKLIKLISHSGASYVGAMSRSITHLVCWKFEGKKYDLAKKFGTVVVN 63
Query: 70 HRWIEDCIREGKRLPIDSYTLQSGQEVGPLLLEVPASFGPNSWIKNKVVSDGLCDIESER 129
HRW+E+C++EG+R+ Y SG+EVGPL++E+PA + + KV + SE
Sbjct: 64 HRWVEECVKEGRRVSETPYMFDSGEEVGPLMIELPA-VSEEAKVTKKV------NKASET 116
Query: 130 QNTVISFGAPSYVLEDSCL---MKKYEETSLHSSRRLKKVKRNISCDNEVSTAAGPSRKE 186
+ S G + S L M+K E + HS R K +I + E S A SRK
Sbjct: 117 FDKYFSNGGENRSGSTSELATWMEKNVEANRHSVRLRTKRPSSILENKENSGVAESSRKG 176
Query: 187 KKFVKKVVDEVVSDPVILDLSPDDRLCRMDRDRLHTEAAATSSLSGGVDVNDLQENSSGP 246
KK V K S ++DL D+ + D H + + + N+ Q++
Sbjct: 177 KKRVVKQR----SYRNLIDLESDE-----ESDNNHHDNSDENQ-------NETQDHREPA 220
Query: 247 DARLCN------QNRAMNGSSDGIEQITDSNQIEDPQTVEQTSI----DLCSSDGKFIDG 296
D + + A+ D D ++IE+ + +++ S + K D
Sbjct: 221 DENVRGFVFEQGETSALRHPGDLATPNWDVDEIEESENWSHSAVFKRPRSFSPEIKPQDD 280
Query: 297 DQV---DNVAGSPISNEWSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKIST 353
D+ + A + SC+IC+ +FSS+ G+LPC H FC +CIQKWA + +S K +T
Sbjct: 281 DESTREETEATEKAPAQVSCIICWTEFSSSRGILPCGHRFCYSCIQKWADRLVSERKKTT 340
Query: 354 CPLCKASFAWYKIVEHAATADKELNSQTVP 383
CPLCK++F +E A ++D+++ SQTVP
Sbjct: 341 CPLCKSNFITITKIEDADSSDQKIYSQTVP 370
>emb|CAG60273.1| unnamed protein product [Candida glabrata CBS138]
gi|50289809|ref|XP_447336.1| unnamed protein product
[Candida glabrata]
Length = 1099
Score = 67.8 bits (164), Expect = 6e-10
Identities = 71/291 (24%), Positives = 128/291 (43%), Gaps = 40/291 (13%)
Query: 13 IVATVTGYHGLERFNLIKLINYAGGSYIGSMSKSITHLKFE---GKKYDIARRLS----- 64
+ T T Y G +R L +LI GGS +SK THL GKKY++A +
Sbjct: 376 LAVTYTNYFGEQRTYLQQLIELLGGSATMELSKQNTHLICNLPFGKKYEVAMKWKESGSK 435
Query: 65 IPVVNHRWIEDCIREGKRLPID-----------SYTLQSGQEVGPLLLEVPASFGPNSWI 113
I + +HRW+E+C GK++P+D +Y+L GQ++ P L +S
Sbjct: 436 IVICSHRWLEECYISGKKVPVDEAYTRLERDDNTYSLTLGQQLSPKLDSENSSDEETDIE 495
Query: 114 KNKVVSDGLCDIESERQNTVISFGAPSYVLEDSCLMKKYEETSLHSSRRLKK-----VKR 168
+++V + R + + S+ LE S KKY +++ + + + + +
Sbjct: 496 ESQVFALEYLKTSQSRAHDLTGLDDISH-LESSIRKKKYGKSNANENASISRTPSASISN 554
Query: 169 NISCDNEVSTAAGPS-------RKEKKFVKKVVDEVVSDPVILDLSPDD----RLCRMDR 217
+ S D+E A + K K K+++ DP L L D LC +D
Sbjct: 555 DSSRDDEAYETAPTNFLTSTELNKSAKLDKQLLITSTVDPSNLRLKDTDYAGSALC-LDE 613
Query: 218 DRLHTEAAATSSLSGGVDVNDLQENSSGPDARLCNQNRAMNGSSDGIEQIT 268
D+ H + L+ ++ + +N + + + + N+ G+SD E+++
Sbjct: 614 DK-HESNKTDAKLN--IEGHVALKNGAKRTSDVSSTNKVSAGNSDKNEKLS 661
>ref|NP_910425.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|22831177|dbj|BAC16036.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 1335
Score = 57.4 bits (137), Expect = 8e-07
Identities = 34/90 (37%), Positives = 52/90 (57%), Gaps = 11/90 (12%)
Query: 17 VTGYHGLERFNLIKLINYAGGSYIGSMSK-SITHL---KFEGKKYDIARR------LSIP 66
+TGY R +++K+ + G + S THL KFEG+KY +A+R ++
Sbjct: 119 LTGYQKNWRDDIMKMASLMGAEFSKSFDALKDTHLICYKFEGEKYKVAKRENTAKRANVS 178
Query: 67 VVNHRWIEDCIREGKRLPIDSYTLQSGQEV 96
+VNH+W+EDC+ K LP D YT +SG E+
Sbjct: 179 LVNHQWLEDCLMAWKILPADDYT-KSGWEI 207
>emb|CAH65372.1| hypothetical protein [Gallus gallus]
gi|61098412|ref|NP_001012953.1| similar to zinc finger
protein [Gallus gallus]
Length = 244
Score = 54.7 bits (130), Expect = 5e-06
Identities = 29/82 (35%), Positives = 42/82 (50%), Gaps = 6/82 (7%)
Query: 303 AGSPISNEWSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFA 362
A + ++SC IC E F +G+ C HTFC C+Q + CPLC+ F
Sbjct: 30 AAEALEAQFSCPICLEVFHRAVGIAGCGHTFCGECLQPCLQVPSPL-----CPLCRMPFD 84
Query: 363 WYKIVEHAATADKELNSQTVPC 384
K VE A++ +K+L+S PC
Sbjct: 85 -PKKVEKASSVEKQLSSYKAPC 105
>ref|YP_053107.1| transcriptional activator [Equid herpesvirus 1]
gi|61287189|dbj|BAD91100.1| transcriptional activator
[Equid herpesvirus 1] gi|49617047|gb|AAT67320.1|
transcriptional activator [Equine herpesvirus 1]
gi|124137|sp|P28990|ICP0_EHV1B Trans-acting
transcriptional protein ICP0 gi|42795190|gb|AAS45947.1|
transcriptional regulator [Equine herpesvirus 1]
gi|60389885|sp|P84445|ICP0_EHV1V Trans-acting
transcriptional protein ICP0 (Infected cell protein 0)
Length = 532
Score = 54.3 bits (129), Expect = 7e-06
Identities = 33/93 (35%), Positives = 39/93 (41%), Gaps = 16/93 (17%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC ED S+ LPC H FC CI +W Q TCPLCK + V H
Sbjct: 8 CPICLEDPSNYSMALPCLHAFCYVCITRWIRQN------PTCPLCKVP---VESVVHTIE 58
Query: 373 ADKELNSQTVPCDNLASVTFISMDQELPDNSFE 405
+D E V D D E ++SFE
Sbjct: 59 SDSEFKETKVSVD-------FDYDSEEDEDSFE 84
>dbj|BAD91106.1| transcriptional activator [Equid herpesvirus 1]
Length = 531
Score = 54.3 bits (129), Expect = 7e-06
Identities = 33/93 (35%), Positives = 39/93 (41%), Gaps = 16/93 (17%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC ED S+ LPC H FC CI +W Q TCPLCK + V H
Sbjct: 8 CPICLEDPSNYSMALPCLHAFCYVCITRWIRQN------PTCPLCKVP---VESVVHTIE 58
Query: 373 ADKELNSQTVPCDNLASVTFISMDQELPDNSFE 405
+D E V D D E ++SFE
Sbjct: 59 SDSEFKETKVSVD-------FDYDSEEDEDSFE 84
>dbj|BAD91104.1| transcriptional activator [Equid herpesvirus 1]
Length = 531
Score = 54.3 bits (129), Expect = 7e-06
Identities = 33/93 (35%), Positives = 39/93 (41%), Gaps = 16/93 (17%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC ED S+ LPC H FC CI +W Q TCPLCK + V H
Sbjct: 8 CPICLEDPSNYSMALPCLHAFCYVCITRWIRQN------PTCPLCKVP---VESVVHTIE 58
Query: 373 ADKELNSQTVPCDNLASVTFISMDQELPDNSFE 405
+D E V D D E ++SFE
Sbjct: 59 SDSEFKETKVSVD-------FDYDSEEDEDSFE 84
>dbj|BAD91102.1| transcriptional activator [Equid herpesvirus 1]
Length = 531
Score = 54.3 bits (129), Expect = 7e-06
Identities = 33/93 (35%), Positives = 39/93 (41%), Gaps = 16/93 (17%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC ED S+ LPC H FC CI +W Q TCPLCK + V H
Sbjct: 8 CPICLEDPSNYSMALPCLHAFCYVCITRWIRQN------PTCPLCKVP---VESVVHTIE 58
Query: 373 ADKELNSQTVPCDNLASVTFISMDQELPDNSFE 405
+D E V D D E ++SFE
Sbjct: 59 SDSEFKETKVSVD-------FDYDSEEDEDSFE 84
>dbj|BAD91098.1| transcriptional activator [Equid herpesvirus 1]
Length = 532
Score = 54.3 bits (129), Expect = 7e-06
Identities = 33/93 (35%), Positives = 39/93 (41%), Gaps = 16/93 (17%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC ED S+ LPC H FC CI +W Q TCPLCK + V H
Sbjct: 8 CPICLEDPSNYSMALPCLHAFCYVCITRWIRQN------PTCPLCKVP---VESVVHTIE 58
Query: 373 ADKELNSQTVPCDNLASVTFISMDQELPDNSFE 405
+D E V D D E ++SFE
Sbjct: 59 SDSEFKETKVSVD-------FDYDSEEDEDSFE 84
>dbj|BAD91096.1| transcriptional activator [Equid herpesvirus 1]
Length = 532
Score = 54.3 bits (129), Expect = 7e-06
Identities = 33/93 (35%), Positives = 39/93 (41%), Gaps = 16/93 (17%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC ED S+ LPC H FC CI +W Q TCPLCK + V H
Sbjct: 8 CPICLEDPSNYSMALPCLHAFCYVCITRWIRQN------PTCPLCKVP---VESVVHTIE 58
Query: 373 ADKELNSQTVPCDNLASVTFISMDQELPDNSFE 405
+D E V D D E ++SFE
Sbjct: 59 SDSEFKETKVSVD-------FDYDSEEDEDSFE 84
>gb|AAB63316.1| contains a deletion of 399 base pairs as compared to ICPO protein
of the Ab4p strain of Equine herpesvirus 1, encoded by
Genbank Accession Number M86664; transcriptional protein
Length = 419
Score = 54.3 bits (129), Expect = 7e-06
Identities = 33/93 (35%), Positives = 39/93 (41%), Gaps = 16/93 (17%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC ED S+ LPC H FC CI +W Q TCPLCK + V H
Sbjct: 8 CPICLEDPSNYSMALPCLHAFCYVCITRWIRQN------PTCPLCKVP---VESVVHTIE 58
Query: 373 ADKELNSQTVPCDNLASVTFISMDQELPDNSFE 405
+D E V D D E ++SFE
Sbjct: 59 SDSEFKETKVSVD-------FDYDSEEDEDSFE 84
>dbj|BAD27395.1| transactivator protein [Equine herpesvirus 1]
Length = 532
Score = 54.3 bits (129), Expect = 7e-06
Identities = 33/93 (35%), Positives = 39/93 (41%), Gaps = 16/93 (17%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC ED S+ LPC H FC CI +W Q TCPLCK + V H
Sbjct: 8 CPICLEDPSNYSMALPCLHAFCYVCITRWIRQN------PTCPLCKVP---VESVVHTIE 58
Query: 373 ADKELNSQTVPCDNLASVTFISMDQELPDNSFE 405
+D E V D D E ++SFE
Sbjct: 59 SDSEFKETKVSVD-------FDYDSEEDEDSFE 84
>emb|CAF92282.1| unnamed protein product [Tetraodon nigroviridis]
Length = 473
Score = 53.1 bits (126), Expect = 2e-05
Identities = 29/89 (32%), Positives = 44/89 (48%), Gaps = 10/89 (11%)
Query: 303 AGSPISNEWSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFA 362
A S + C IC + F++ + C H FC CIQ+W+H + + CPLCK FA
Sbjct: 2 AAEDASPDSKCPICLDRFNNLAYLDRCLHRFCFPCIQEWSHNK------AECPLCKQPFA 55
Query: 363 WYKIVEHAATADKELNSQTV-PCDNLASV 390
+ H+ A+ + T+ P N +SV
Sbjct: 56 ---SILHSVRAEDDFKEYTLQPAPNTSSV 81
>ref|XP_650132.1| hypothetical protein 244.t00016 [Entamoeba histolytica HM-1:IMSS]
gi|56466698|gb|EAL44745.1| hypothetical protein
244.t00016 [Entamoeba histolytica HM-1:IMSS]
Length = 102
Score = 52.8 bits (125), Expect = 2e-05
Identities = 31/81 (38%), Positives = 45/81 (55%), Gaps = 4/81 (4%)
Query: 14 VATVTGYHGLERFNLIKLINYAGGSYIGSM-SKSITHLKFEGKKYDIAR---RLSIPVVN 69
+ V+GY ER L ++ GG Y+ M SKS+T L +G D A R +PV++
Sbjct: 18 IICVSGYSSDERLLLRGMVELCGGIYMEDMESKSVTFLLSKGLTSDKASHALRWGVPVLS 77
Query: 70 HRWIEDCIREGKRLPIDSYTL 90
H+W+ DCI E + L I+ Y L
Sbjct: 78 HQWLFDCIAERRLLSINQYVL 98
>ref|XP_429180.1| PREDICTED: hypothetical protein XP_429180 [Gallus gallus]
Length = 340
Score = 52.4 bits (124), Expect = 3e-05
Identities = 31/105 (29%), Positives = 46/105 (43%), Gaps = 12/105 (11%)
Query: 311 WSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHA 370
W+C IC + +PC H FCL CIQ+WA R +CPLC+A+ ++
Sbjct: 6 WTCAICRDVRQDIAYAIPCHHMFCLGCIQRWARLR------DSCPLCRAAMQTIRVPVRG 59
Query: 371 ATADKE--LNSQTVPCDNLASVTFISMDQELPDNSFERREEPRVP 413
E ++ VP ++F + PDN+ E P P
Sbjct: 60 DNQYVECIVSPPAVP----VPISFSTTTATGPDNAEEFLPPPPPP 100
>pdb|1CHC| Equine Herpes Virus-1 (C3hc4, Or Ring Domain) (Nmr, 1 Structure)
Length = 68
Score = 52.4 bits (124), Expect = 3e-05
Identities = 27/70 (38%), Positives = 32/70 (45%), Gaps = 9/70 (12%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC ED S+ LPC H FC CI +W Q TCPLCK + V H
Sbjct: 8 CPICLEDPSNYSMALPCLHAFCYVCITRWIRQN------PTCPLCKVP---VESVVHTIE 58
Query: 373 ADKELNSQTV 382
+D E Q +
Sbjct: 59 SDSEFGDQLI 68
>ref|NP_045280.1| 63 [Equid herpesvirus 4] gi|2606010|gb|AAC59582.1| 63 [Equine
herpesvirus 4] gi|11278251|pir||T42606 probable
transcription activator - equine herpesvirus 4 (strain
NS80567)
Length = 536
Score = 52.0 bits (123), Expect = 3e-05
Identities = 34/93 (36%), Positives = 40/93 (42%), Gaps = 16/93 (17%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC ED S+ LPC H FC CI +W Q TCPLCK V H
Sbjct: 9 CPICLEDPSNYSMALPCLHAFCYVCITRWIRQN------PTCPLCKVP---VNSVVHTIE 59
Query: 373 ADKELNSQTVPCDNLASVTFISMDQELPDNSFE 405
+D E V SV F D + ++SFE
Sbjct: 60 SDSEFKETKV------SVEF-DYDSDEDEDSFE 85
>ref|XP_429144.1| PREDICTED: similar to p53-binding protein-3 [Gallus gallus]
Length = 246
Score = 52.0 bits (123), Expect = 3e-05
Identities = 34/125 (27%), Positives = 51/125 (40%), Gaps = 16/125 (12%)
Query: 244 SGPDARLCNQNRAM---NGSSDGIEQITDSNQIEDPQTVEQTSIDLCSSDG-------KF 293
+GPD L A NG E I + + + T DLCS +
Sbjct: 10 AGPDRMLAAGEMAFTKENGDRHTAESIKAGAEQSGDKIFQITPKDLCSEESLEERGSAAM 69
Query: 294 IDGDQVDNVAGSPISNEWSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKIST 353
+ + S+++ C IC ++ S+ V PC H FC CI++WA R +
Sbjct: 70 ANRSHLQQEGPLEASDDYRCAICLDNISNMACVDPCFHRFCSGCIRRWATTR------AV 123
Query: 354 CPLCK 358
CPLC+
Sbjct: 124 CPLCR 128
>ref|XP_329862.1| hypothetical protein [Neurospora crassa] gi|28919794|gb|EAA29196.1|
hypothetical protein [Neurospora crassa]
Length = 1048
Score = 52.0 bits (123), Expect = 3e-05
Identities = 31/86 (36%), Positives = 46/86 (53%), Gaps = 10/86 (11%)
Query: 5 EENLV------RLKIVATVTGY-HGLERFNLIKLINYAGGSYIGSMSKSITHL---KFEG 54
E NLV R K++ TG+ ER ++I ++ GG Y G +++ +THL K EG
Sbjct: 128 EPNLVDGTIYPRRKLLVCSTGFMEPEERQHIIDMVEKGGGIYTGDLTRRVTHLVVYKPEG 187
Query: 55 KKYDIARRLSIPVVNHRWIEDCIREG 80
+KY A+ I V+ WI DC+ G
Sbjct: 188 RKYQAAQNWGIKNVSVEWINDCVERG 213
Score = 35.8 bits (81), Expect = 2.6
Identities = 19/79 (24%), Positives = 36/79 (45%), Gaps = 6/79 (7%)
Query: 16 TVTGYHGLERFNLIKLINYAGGSYIGSMSKSITHLK------FEGKKYDIARRLSIPVVN 69
+ G+ G++ ++ K I G Y + ++ L +K D+A R +P+V
Sbjct: 462 STAGFTGVDLNHVDKAIRQLGARYEERFTADVSVLVAPSVSVIRKQKLDMALRWKVPIVT 521
Query: 70 HRWIEDCIREGKRLPIDSY 88
W+ CI G + PI+ +
Sbjct: 522 AEWLWTCINSGFKCPIEDF 540
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.133 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 733,334,495
Number of Sequences: 2540612
Number of extensions: 30762664
Number of successful extensions: 76178
Number of sequences better than 10.0: 2381
Number of HSP's better than 10.0 without gapping: 267
Number of HSP's successfully gapped in prelim test: 2114
Number of HSP's that attempted gapping in prelim test: 75140
Number of HSP's gapped (non-prelim): 2742
length of query: 423
length of database: 863,360,394
effective HSP length: 131
effective length of query: 292
effective length of database: 530,540,222
effective search space: 154917744824
effective search space used: 154917744824
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0225.8


