

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0225.5
(295 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
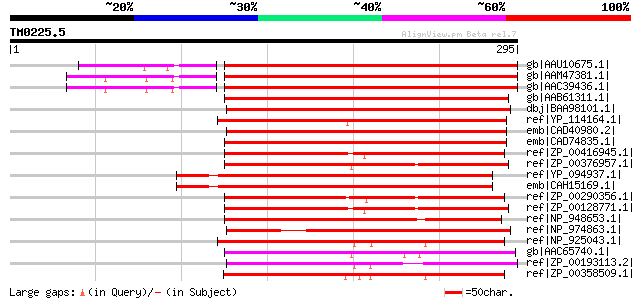
Sequences producing significant alignments: (bits) Value
gb|AAU10675.1| putative DegP protease [Oryza sativa (japonica cu... 316 4e-85
gb|AAM47381.1| At3g27925/K16N12.18 [Arabidopsis thaliana] gi|145... 307 2e-82
gb|AAC39436.1| DegP protease precursor [Arabidopsis thaliana] 305 8e-82
gb|AAB61311.1| htrA-like protein [Haematococcus pluvialis] 217 4e-55
dbj|BAA98101.1| unnamed protein product [Arabidopsis thaliana] g... 162 8e-39
ref|YP_114164.1| serine protease, putative [Methylococcus capsul... 159 7e-38
emb|CAD40980.2| OSJNBa0072F16.5 [Oryza sativa (japonica cultivar... 159 7e-38
emb|CAD74835.1| protease Do-like (S2 serine-type protease) [Rhod... 155 2e-36
ref|ZP_00416945.1| Peptidase S1, chymotrypsin:PDZ/DHR/GLGF domai... 154 2e-36
ref|ZP_00376957.1| serine protease [Erythrobacter litoralis HTCC... 148 2e-34
ref|YP_094937.1| DegP protease (Do-like, S2-serine-like) [Legion... 142 9e-33
emb|CAH15169.1| hypothetical protein [Legionella pneumophila str... 142 9e-33
ref|ZP_00290356.1| COG0265: Trypsin-like serine proteases, typic... 142 1e-32
ref|ZP_00128771.1| COG0265: Trypsin-like serine proteases, typic... 139 7e-32
ref|NP_948653.1| putative DegP protease precursor [Rhodopseudomo... 139 1e-31
ref|NP_974863.1| DegP protease, putative [Arabidopsis thaliana] 136 6e-31
ref|NP_925043.1| probable serine protease [Gloeobacter violaceus... 131 3e-29
gb|AAC65740.1| periplasmic serine protease DO (htrA-1) [Treponem... 129 1e-28
ref|ZP_00193113.2| COG0265: Trypsin-like serine proteases, typic... 122 1e-26
ref|ZP_00358509.1| COG0265: Trypsin-like serine proteases, typic... 122 2e-26
>gb|AAU10675.1| putative DegP protease [Oryza sativa (japonica cultivar-group)]
gi|51038126|gb|AAT93929.1| putative DegP protease [Oryza
sativa (japonica cultivar-group)]
Length = 437
Score = 316 bits (810), Expect = 4e-85
Identities = 157/170 (92%), Positives = 166/170 (97%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLDSSGNLIG+NTAIYSPSGASSGVGFSIP+DTV GIVDQL+KF
Sbjct: 268 VIQTDAAINPGNSGGPLLDSSGNLIGVNTAIYSPSGASSGVGFSIPVDTVGGIVDQLIKF 327
Query: 186 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
GKVTRPILGIKFAPDQSVEQLG+SGVLVLDAP NGPAGKAGLQSTKRDSYGRLILGDIIT
Sbjct: 328 GKVTRPILGIKFAPDQSVEQLGLSGVLVLDAPPNGPAGKAGLQSTKRDSYGRLILGDIIT 387
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDES 295
SVNGTKVT+GSDLYRILDQCKVG+KVTVEVLRGD KEKIPVILEPKPDES
Sbjct: 388 SVNGTKVTNGSDLYRILDQCKVGEKVTVEVLRGDQKEKIPVILEPKPDES 437
Score = 65.1 bits (157), Expect = 2e-09
Identities = 46/103 (44%), Positives = 55/103 (52%), Gaps = 25/103 (24%)
Query: 41 SILILCTSIALSFTNADSAYAFVVTPPRKLQSDELAT----------VVYITNLAVKMRL 90
S+++ S AL +A SA AFVV PRKLQ+DELAT VVYITNLAV+
Sbjct: 83 SLIVALASAALILGDAGSASAFVVATPRKLQADELATVRLFQENTPSVVYITNLAVRQDA 142
Query: 91 -------------RWMCWRFLKEGNIVTNYHVIPGASDLSLDL 120
W K G+IVTN+HVI GASDL + L
Sbjct: 143 FTLDVLEVPQGSGSGFVWD--KSGHIVTNFHVIRGASDLRVTL 183
>gb|AAM47381.1| At3g27925/K16N12.18 [Arabidopsis thaliana]
gi|14517500|gb|AAK62640.1| K16N12.18/K16N12.18
[Arabidopsis thaliana] gi|51338737|sp|O22609|DEGP1_ARATH
Protease Do-like 1, chloroplast precursor
gi|22331378|ref|NP_189431.2| DegP protease, putative
[Arabidopsis thaliana]
Length = 439
Score = 307 bits (787), Expect = 2e-82
Identities = 153/170 (90%), Positives = 163/170 (95%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLDSSG LIGINTAIYSPSGASSGVGFSIP+DTV GIVDQLV+F
Sbjct: 270 VIQTDAAINPGNSGGPLLDSSGTLIGINTAIYSPSGASSGVGFSIPVDTVGGIVDQLVRF 329
Query: 186 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAP +GPAGKAGLQSTKRD YGRL+LGDIIT
Sbjct: 330 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPPSGPAGKAGLQSTKRDGYGRLVLGDIIT 389
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDES 295
SVNGTKV++GSDLYRILDQCKVGD+VTVEVLRGDHKEKI V LEPKPDES
Sbjct: 390 SVNGTKVSNGSDLYRILDQCKVGDEVTVEVLRGDHKEKISVTLEPKPDES 439
Score = 73.2 bits (178), Expect = 9e-12
Identities = 51/116 (43%), Positives = 64/116 (54%), Gaps = 31/116 (26%)
Query: 34 TPKTCFNSILILCTSIALSFT------NADSAYAFVVTPPRKLQSDELATV--------- 78
TP + +LCTS+ALSF+ +SA AFVV+ P+KLQ+DELATV
Sbjct: 72 TPFSAVKPFFLLCTSVALSFSLFAASPAVESASAFVVSTPKKLQTDELATVRLFQENTPS 131
Query: 79 -VYITNLAVKMRLRWM-------------CWRFLKEGNIVTNYHVIPGASDLSLDL 120
VYITNLAV+ + W K+G+IVTNYHVI GASDL + L
Sbjct: 132 VVYITNLAVRQDAFTLDVLEVPQGSGSGFVWD--KQGHIVTNYHVIRGASDLRVTL 185
>gb|AAC39436.1| DegP protease precursor [Arabidopsis thaliana]
Length = 437
Score = 305 bits (782), Expect = 8e-82
Identities = 152/170 (89%), Positives = 163/170 (95%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLDSSG LIGINTAIYSPSGASSGVGFSIP+DTV GIVDQLV+F
Sbjct: 268 VIQTDAAINPGNSGGPLLDSSGTLIGINTAIYSPSGASSGVGFSIPVDTVGGIVDQLVRF 327
Query: 186 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
GKVTRPILGIKFAPDQSVEQLGVSGVL+LDAP +GPAGKAGLQSTKRD YGRLILGDIIT
Sbjct: 328 GKVTRPILGIKFAPDQSVEQLGVSGVLLLDAPPSGPAGKAGLQSTKRDGYGRLILGDIIT 387
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDES 295
SVNGTKV++GSDLYRILDQCKVGD+VTV+VLRGDHKEKI V LEPKPDES
Sbjct: 388 SVNGTKVSNGSDLYRILDQCKVGDEVTVQVLRGDHKEKISVTLEPKPDES 437
Score = 73.2 bits (178), Expect = 9e-12
Identities = 51/116 (43%), Positives = 64/116 (54%), Gaps = 31/116 (26%)
Query: 34 TPKTCFNSILILCTSIALSFT------NADSAYAFVVTPPRKLQSDELATV--------- 78
TP + +LCTS+ALSF+ +SA AFVV+ P+KLQ+DELATV
Sbjct: 70 TPFSAVKPFFLLCTSVALSFSLFAASPAVESASAFVVSTPKKLQTDELATVRLFQENTPS 129
Query: 79 -VYITNLAVKMRLRWM-------------CWRFLKEGNIVTNYHVIPGASDLSLDL 120
VYITNLAV+ + W K+G+IVTNYHVI GASDL + L
Sbjct: 130 VVYITNLAVRQDAFTLDVLEVPQGSGSGFVWD--KQGHIVTNYHVIRGASDLRVTL 183
>gb|AAB61311.1| htrA-like protein [Haematococcus pluvialis]
Length = 398
Score = 217 bits (552), Expect = 4e-55
Identities = 105/165 (63%), Positives = 131/165 (78%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLDSSG +IGINTAIYSPSG +SGVGF+IP DTV V Q+++F
Sbjct: 231 VIQTDAAINPGNSGGPLLDSSGCVIGINTAIYSPSGTNSGVGFAIPADTVRSSVTQILEF 290
Query: 186 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
GKV RP+LGI FAPDQ+VE LGV G++VL+A GPA KAG+ T RD YGRL+LGDII
Sbjct: 291 GKVVRPMLGIAFAPDQAVEALGVKGIMVLNAREGGPAWKAGIVGTSRDEYGRLVLGDIIR 350
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEP 290
+VNGT + S +DLYR+LD+ +VG+ + +EVLRG E + V L P
Sbjct: 351 TVNGTVIRSSTDLYRVLDKAQVGETLDIEVLRGSSTEHVNVTLAP 395
>dbj|BAA98101.1| unnamed protein product [Arabidopsis thaliana]
gi|19699228|gb|AAL90980.1| AT5g39830/K13H13_10
[Arabidopsis thaliana] gi|18421917|ref|NP_568575.1| DegP
protease, putative [Arabidopsis thaliana]
gi|15912207|gb|AAL08237.1| AT5g39830/K13H13_10
[Arabidopsis thaliana] gi|18203244|sp|Q9LU10|DEGP8_ARATH
Protease Do-like 8, chloroplast precursor
Length = 448
Score = 162 bits (411), Expect = 8e-39
Identities = 86/166 (51%), Positives = 114/166 (67%), Gaps = 1/166 (0%)
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFG 186
+ +DAAINPGNSGGPLLDS GNLIGINTAI++ +G S+GVGF+IP TV IV QL++F
Sbjct: 281 IQTDAAINPGNSGGPLLDSKGNLIGINTAIFTQTGTSAGVGFAIPSSTVLKIVPQLIQFS 340
Query: 187 KVTRPILGIKFAPDQSVEQLGV-SGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
KV R + I+ APD QL V +G LVL P A KAGL T R G ++LGDII
Sbjct: 341 KVLRAGINIELAPDPVANQLNVRNGALVLQVPGKSLAEKAGLHPTSRGFAGNIVLGDIIV 400
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPK 291
+V+ V + ++L +ILD+ VGDKVT+++ RG+ ++ + LE K
Sbjct: 401 AVDDKPVKNKAELMKILDEYSVGDKVTLKIKRGNEDLELKISLEEK 446
>ref|YP_114164.1| serine protease, putative [Methylococcus capsulatus str. Bath]
gi|53757716|gb|AAU92007.1| putative serine protease
[Methylococcus capsulatus str. Bath]
Length = 374
Score = 159 bits (403), Expect = 7e-38
Identities = 90/171 (52%), Positives = 115/171 (66%), Gaps = 3/171 (1%)
Query: 122 TLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQ 181
T+ L+ +DAAINPGNSGGPLLDS+G L+GINTAIYSPSGA SGVGF++P+DTVN +V Q
Sbjct: 201 TIEHLIQTDAAINPGNSGGPLLDSAGRLVGINTAIYSPSGAFSGVGFAVPVDTVNRVVPQ 260
Query: 182 LVKFGKVTRPILGI---KFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRL 238
L+ G+ RP LGI + ++V++LGV+GVLVL A AGL+ GRL
Sbjct: 261 LIGRGQYIRPALGIAVDEGLNQRAVQRLGVTGVLVLKVNPGSAAEAAGLKGATLLPDGRL 320
Query: 239 ILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILE 289
I GDII +V G V S S L +LD ++G KV + V RGD + I V L+
Sbjct: 321 IPGDIIVAVEGRPVDSVSKLSALLDDYQIGQKVRLSVRRGDTEMDIAVQLQ 371
>emb|CAD40980.2| OSJNBa0072F16.5 [Oryza sativa (japonica cultivar-group)]
gi|50924798|ref|XP_472748.1| OSJNBa0072F16.5 [Oryza
sativa (japonica cultivar-group)]
Length = 420
Score = 159 bits (403), Expect = 7e-38
Identities = 83/164 (50%), Positives = 111/164 (67%), Gaps = 1/164 (0%)
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFG 186
+ +DAAINPGNSGGPLLDS G++IGINTAI++ +G S+GVGF+IP TV I QL++FG
Sbjct: 253 IQTDAAINPGNSGGPLLDSKGHMIGINTAIFTQTGTSAGVGFAIPSSTVLKIAPQLIQFG 312
Query: 187 KVTRPILGIKFAPDQSVEQLGV-SGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
KV R L ++FAPD QL V +G L+L P A KAGL T R G ++LGD+I
Sbjct: 313 KVRRAGLNVEFAPDPIAYQLNVRTGSLILQVPGGSAAAKAGLVPTSRGFAGNIVLGDVIV 372
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILE 289
+V+G + SDL R+LD VGDKV++ + RG ++ + LE
Sbjct: 373 AVDGKPIKGKSDLSRVLDDYGVGDKVSLTIQRGAETLEVTLPLE 416
>emb|CAD74835.1| protease Do-like (S2 serine-type protease) [Rhodopirellula baltica
SH 1] gi|32474295|ref|NP_867289.1| protease Do-like (S2
serine-type protease) [Rhodopirellula baltica SH 1]
Length = 399
Score = 155 bits (391), Expect = 2e-36
Identities = 80/165 (48%), Positives = 114/165 (68%), Gaps = 1/165 (0%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLD SG LIG+NTAIYSPSGA +G+GF+IP+DTV +V +L++
Sbjct: 232 VIQTDAAINPGNSGGPLLDRSGQLIGVNTAIYSPSGAYAGIGFAIPVDTVRWVVPELIEH 291
Query: 186 GKVTRPILGIKFAPDQSVEQLGV-SGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDII 244
G++ RP + I A D ++ + GVL+LD P G A +AGL+ T+R +G ++LGDII
Sbjct: 292 GRIIRPGIAITVASDSMSKRFKLPPGVLILDMPERGNAERAGLRPTRRTRFGDIVLGDII 351
Query: 245 TSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILE 289
+V+ V S +DL I + + GD V + V+R + +PV LE
Sbjct: 352 VAVDEMPVASTADLTLIFENYESGDVVDLTVIRQGTELVLPVELE 396
>ref|ZP_00416945.1| Peptidase S1, chymotrypsin:PDZ/DHR/GLGF domain [Azotobacter
vinelandii AvOP] gi|67087118|gb|EAM06585.1| Peptidase
S1, chymotrypsin:PDZ/DHR/GLGF domain [Azotobacter
vinelandii AvOP]
Length = 365
Score = 154 bits (390), Expect = 2e-36
Identities = 82/168 (48%), Positives = 110/168 (64%), Gaps = 7/168 (4%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
L+ +DAAINPGNSGGPLLDS+G LIGINTAIYSPSGAS+G+GF++P+DTVN +V QL+
Sbjct: 196 LIQTDAAINPGNSGGPLLDSAGRLIGINTAIYSPSGASAGIGFAVPVDTVNRVVPQLIDT 255
Query: 186 GKVTRPILGIKFAPDQSVEQ-----LGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLIL 240
GK +P LGI+ D V Q G+ GV VL A AGL+ G ++
Sbjct: 256 GKYVQPTLGIQV--DSGVNQRLGELSGIEGVFVLGVKPGSAAEAAGLEGAALTRDGGIVP 313
Query: 241 GDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVIL 288
GDI+T+V+G V S L ILD + GD+V + V RG+ + ++ ++L
Sbjct: 314 GDIVTAVDGKAVDSVERLLAILDDYRAGDRVRLSVKRGERQREVELVL 361
>ref|ZP_00376957.1| serine protease [Erythrobacter litoralis HTCC2594]
gi|60735610|gb|EAL73871.1| serine protease
[Erythrobacter litoralis HTCC2594]
Length = 332
Score = 148 bits (373), Expect = 2e-34
Identities = 80/168 (47%), Positives = 110/168 (64%), Gaps = 4/168 (2%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
L+ +DAAINPGNSGGPLLDS+G LIGINTAIYSPSGAS+G+GF++P+DTV +V QL+
Sbjct: 164 LIQTDAAINPGNSGGPLLDSAGRLIGINTAIYSPSGASAGIGFAVPVDTVMRVVPQLISE 223
Query: 186 GKVTRPILGIKF---APDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGD 242
G+ TRP LG++ D+ G+ GV VL A +AGL + +R G + GD
Sbjct: 224 GRYTRPSLGLESDDDINDRLKRASGIEGVFVLRVDPGSSADRAGLVAAQRTRRG-VAPGD 282
Query: 243 IITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEP 290
I+T++NG V+ DL LD +VG V + ++RG + + + LEP
Sbjct: 283 IVTALNGKPVSRVGDLLARLDDFRVGQSVVLTLMRGGAERTVRLELEP 330
>ref|YP_094937.1| DegP protease (Do-like, S2-serine-like) [Legionella pneumophila
subsp. pneumophila str. Philadelphia 1]
gi|53750709|emb|CAH12116.1| hypothetical protein
[Legionella pneumophila str. Paris]
gi|54296924|ref|YP_123293.1| hypothetical protein
lpp0965 [Legionella pneumophila str. Paris]
gi|52628249|gb|AAU26990.1| DegP protease (Do-like,
S2-serine-like) [Legionella pneumophila subsp.
pneumophila str. Philadelphia 1]
Length = 363
Score = 142 bits (359), Expect = 9e-33
Identities = 73/185 (39%), Positives = 112/185 (60%), Gaps = 6/185 (3%)
Query: 98 LKEGNIVTNYHVIPGASDLSLDLTTLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIY 157
L +G I +PG + T+ ++ +D INPGNSGGPLL+S+G LIG+NT IY
Sbjct: 171 LSKGVISALGRKVPGIGGV-----TIYDMIQTDTPINPGNSGGPLLNSAGQLIGMNTMIY 225
Query: 158 SPSGASSGVGFSIPLDTVNGIVDQLVKFGKVTRPILGIKFAPDQSVEQLGV-SGVLVLDA 216
S SG+S+G+GF++P + + I Q++ G+V +GI+ E+LGV G+L+ D
Sbjct: 226 SRSGSSAGIGFAVPAEDIQKIASQIINHGRVVLSGIGIQRVEPHLAERLGVKKGILIADV 285
Query: 217 PANGPAGKAGLQSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVL 276
PA K L+ T R+ +GR++LGD+I VN V + LY +L + KVG+++TV ++
Sbjct: 286 VPGTPADKLKLRGTHRNQWGRIVLGDVIVGVNAHPVPNYDALYNLLTEIKVGEQITVSII 345
Query: 277 RGDHK 281
R K
Sbjct: 346 RNGKK 350
>emb|CAH15169.1| hypothetical protein [Legionella pneumophila str. Lens]
gi|54293879|ref|YP_126294.1| hypothetical protein
lpl0935 [Legionella pneumophila str. Lens]
Length = 363
Score = 142 bits (359), Expect = 9e-33
Identities = 73/185 (39%), Positives = 112/185 (60%), Gaps = 6/185 (3%)
Query: 98 LKEGNIVTNYHVIPGASDLSLDLTTLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIY 157
L +G I +PG + T+ ++ +D INPGNSGGPLL+S+G LIG+NT IY
Sbjct: 171 LSKGVISALGRKVPGIGGV-----TIYDMIQTDTPINPGNSGGPLLNSAGQLIGMNTMIY 225
Query: 158 SPSGASSGVGFSIPLDTVNGIVDQLVKFGKVTRPILGIKFAPDQSVEQLGV-SGVLVLDA 216
S SG+S+G+GF++P + + I Q++ G+V +GI+ E+LGV G+L+ D
Sbjct: 226 SRSGSSAGIGFAVPAEDIQKIASQIINHGRVVLSGIGIQRVEPHLAERLGVKKGILIADV 285
Query: 217 PANGPAGKAGLQSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVL 276
PA K L+ T R+ +GR++LGD+I VN V + LY +L + KVG+++TV ++
Sbjct: 286 VPGTPADKLKLRGTHRNQWGRIVLGDVIVGVNAHPVPNYDALYNLLTEIKVGEQITVSII 345
Query: 277 RGDHK 281
R K
Sbjct: 346 RNGKK 350
>ref|ZP_00290356.1| COG0265: Trypsin-like serine proteases, typically periplasmic,
contain C-terminal PDZ domain [Magnetococcus sp. MC-1]
Length = 368
Score = 142 bits (358), Expect = 1e-32
Identities = 76/167 (45%), Positives = 113/167 (67%), Gaps = 6/167 (3%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
L+ +DAAINPGNSGGPLLDS+G LIGINTAIYSPSGA +G+GF++P+D VN +V QL+
Sbjct: 204 LIQTDAAINPGNSGGPLLDSAGRLIGINTAIYSPSGAYAGIGFAVPVDEVNRVVPQLIAQ 263
Query: 186 GKVTRPILGIKFAPDQSVEQL----GVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILG 241
G+ RP LGI+ A D+S Q+ ++GVLVL + A +AG+Q+++ D G ++LG
Sbjct: 264 GRYQRPSLGIQ-ASDRSSAQILSRFEITGVLVLGVASGSAAQRAGIQASRLDERG-IVLG 321
Query: 242 DIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVIL 288
D+I ++ + L + L + +VGD V + + R +++ V+L
Sbjct: 322 DVIVAIADQPTENIDQLQKALAKYRVGDTVKITLWRQGENQQLEVVL 368
>ref|ZP_00128771.1| COG0265: Trypsin-like serine proteases, typically periplasmic,
contain C-terminal PDZ domain [Desulfovibrio
desulfuricans G20]
Length = 383
Score = 139 bits (351), Expect = 7e-32
Identities = 82/170 (48%), Positives = 105/170 (61%), Gaps = 8/170 (4%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
L+ +DAAINPGNSGGPLLDS+G LIGINTAIYSPSGAS+G+GF++P+DTV +V QL+K
Sbjct: 215 LIQTDAAINPGNSGGPLLDSAGRLIGINTAIYSPSGASAGIGFAVPVDTVMRVVPQLIKT 274
Query: 186 GKVTRPILGIKFAPDQSVEQ-----LGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLIL 240
GK RP LGI+ D+ + Q G GV VL A KAGL + G ++
Sbjct: 275 GKYIRPALGIQV--DEQLNQRLLALTGSKGVFVLRVTPGSAAHKAGLAGVEVTPQG-IVP 331
Query: 241 GDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEP 290
GD I V+G + + L LD KVGD V + V R ++ V L+P
Sbjct: 332 GDRIIGVDGKATDNVAKLLARLDDRKVGDVVVLSVERAGKTREVRVELQP 381
>ref|NP_948653.1| putative DegP protease precursor [Rhodopseudomonas palustris
CGA009] gi|39650232|emb|CAE28755.1| putative DegP
protease precursor [Rhodopseudomonas palustris CGA009]
Length = 399
Score = 139 bits (349), Expect = 1e-31
Identities = 75/161 (46%), Positives = 101/161 (62%), Gaps = 4/161 (2%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLDS+G LIGINTAI S SGAS+G+GF+IP+D VN +V L+
Sbjct: 232 VIQTDAAINPGNSGGPLLDSAGRLIGINTAIISGSGASAGIGFAIPVDAVNRVVTALITN 291
Query: 186 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
G V P +GI A + QLG+ GV++L + PA +AGL+ D Y R D+IT
Sbjct: 292 GSVPVPGIGIVAARETETAQLGIDGVVILRTLPDSPAAQAGLEGATDDGYVR----DVIT 347
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPV 286
NG+ + S SDL L++ +G V + V R + V
Sbjct: 348 GANGSDIHSMSDLAAALEEAGIGRDVKLTVERDGRARTVTV 388
>ref|NP_974863.1| DegP protease, putative [Arabidopsis thaliana]
Length = 434
Score = 136 bits (343), Expect = 6e-31
Identities = 78/166 (46%), Positives = 103/166 (61%), Gaps = 15/166 (9%)
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFG 186
+ +DAAINPGNSGGPLLDS GNLIGINTAI++ TV IV QL++F
Sbjct: 281 IQTDAAINPGNSGGPLLDSKGNLIGINTAIFT--------------QTVLKIVPQLIQFS 326
Query: 187 KVTRPILGIKFAPDQSVEQLGV-SGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
KV R + I+ APD QL V +G LVL P A KAGL T R G ++LGDII
Sbjct: 327 KVLRAGINIELAPDPVANQLNVRNGALVLQVPGKSLAEKAGLHPTSRGFAGNIVLGDIIV 386
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPK 291
+V+ V + ++L +ILD+ VGDKVT+++ RG+ ++ + LE K
Sbjct: 387 AVDDKPVKNKAELMKILDEYSVGDKVTLKIKRGNEDLELKISLEEK 432
>ref|NP_925043.1| probable serine protease [Gloeobacter violaceus PCC 7421]
gi|35212664|dbj|BAC90038.1| gll2097 [Gloeobacter
violaceus PCC 7421]
Length = 400
Score = 131 bits (329), Expect = 3e-29
Identities = 74/176 (42%), Positives = 111/176 (63%), Gaps = 9/176 (5%)
Query: 122 TLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQ 181
TL L+ +DAAINPGNSGGPLLDS G LIG+NTAI+S SG+S+G+GF++P+DTV ++ +
Sbjct: 215 TLRNLIQTDAAINPGNSGGPLLDSQGRLIGVNTAIFSTSGSSAGIGFAVPVDTVRQVLPE 274
Query: 182 LVKFGKVTRPILGIKFAP--DQSVEQLGVS---GVLVLDAPANGPAGKAGLQSTKRDSY- 235
L+ G V R LG++ P VE L +S G LV G A +AGL++ + ++
Sbjct: 275 LISRGTVRRASLGVQVLPLSPMVVETLKLSVKEGALVAAVVPGGAAARAGLRAGRLETID 334
Query: 236 GRLIL---GDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVIL 288
G L L D+I +++ + DL + + K GDKVT+ ++R + + ++PV L
Sbjct: 335 GNLQLPVGADVIVAIDRVAIKDAQDLINQIQKHKPGDKVTLTIVRNNAEVQVPVTL 390
>gb|AAC65740.1| periplasmic serine protease DO (htrA-1) [Treponema pallidum subsp.
pallidum str. Nichols] gi|15639760|ref|NP_219210.1|
periplasmic serine protease DO (htrA-1) [Treponema
pallidum subsp. pallidum str. Nichols]
gi|7448456|pir||B71284 probable periplasmic serine
proteinase DO (htrA-1) - syphilis spirochete
Length = 398
Score = 129 bits (323), Expect = 1e-28
Identities = 77/182 (42%), Positives = 109/182 (59%), Gaps = 13/182 (7%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLD+ G +IGINT IYS SG+SSGVGF++P+DT IV +L+++
Sbjct: 217 MIQTDAAINPGNSGGPLLDTQGRMIGINTVIYSTSGSSSGVGFAVPVDTAKRIVSELIRY 276
Query: 186 GKVTRPILGIKF----APDQSVEQLGV-SGVLVLDAPANGPAGKAGLQ---STKRDSYGR 237
G+V R + + A QL V G+LV PA +AGL+ + R GR
Sbjct: 277 GRVRRGKIDAELVQVNASIAHYAQLTVGKGLLVSQVKRGSPAAQAGLRGGTTAVRYGLGR 336
Query: 238 -----LILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKP 292
+ GD+IT+++ V + SD Y +L+ K D+V V VLRG + + V L +
Sbjct: 337 RAAVIYLGGDVITAIDNQPVANLSDYYSVLEDKKPDDEVRVTVLRGRRQHVVAVRLTERS 396
Query: 293 DE 294
DE
Sbjct: 397 DE 398
>ref|ZP_00193113.2| COG0265: Trypsin-like serine proteases, typically periplasmic,
contain C-terminal PDZ domain [Mesorhizobium sp. BNC1]
Length = 569
Score = 122 bits (306), Expect = 1e-26
Identities = 73/174 (41%), Positives = 100/174 (56%), Gaps = 16/174 (9%)
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFG 186
+ +DAAIN GNSGGPL + +G +IGINTAI SPSG S G+GF+IP + +V QL +FG
Sbjct: 289 IQTDAAINRGNSGGPLFNMAGEVIGINTAIISPSGGSIGIGFAIPSNLALNVVGQLREFG 348
Query: 187 KVTRPILGIKFAP--DQSVEQLGV---SGVLVLDAPANGPAGKAGLQSTKRDSYGRLILG 241
+ R LG++ P D+ E LG+ +GVLV GPA LQ+ G
Sbjct: 349 ETRRGWLGVRIQPVTDEIAESLGLDEAAGVLVSGIEKGGPADNGLLQA-----------G 397
Query: 242 DIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDES 295
DII NGTKV L R++ + VG ++ +E+LR +E + V L +ES
Sbjct: 398 DIIVGFNGTKVADDRQLRRLVAESGVGKEIDLEILRKGERETVKVTLGRLEEES 451
>ref|ZP_00358509.1| COG0265: Trypsin-like serine proteases, typically periplasmic,
contain C-terminal PDZ domain [Chloroflexus aurantiacus]
Length = 396
Score = 122 bits (305), Expect = 2e-26
Identities = 73/179 (40%), Positives = 111/179 (61%), Gaps = 15/179 (8%)
Query: 125 QLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVK 184
+++ SD AINPGNSGGPLLD SG +IG+N+AI SPSGA++G+GF+I TV +V L++
Sbjct: 213 EVIQSDVAINPGNSGGPLLDLSGRVIGVNSAILSPSGANAGIGFAISSRTVQRVVPVLIR 272
Query: 185 FGKVTRPILG---IKFAPDQS--VEQLGV-----SGVLVLDAPANGPAGKAGLQSTKR-D 233
G+ P LG I+ P ++ E+ G+ G+L+ + NGPA +AGL+ R
Sbjct: 273 EGRYPHPSLGVRVIELTPQRASLFERAGMQLPVTQGLLIAELITNGPAAQAGLRGPDRLV 332
Query: 234 SYGRLIL---GDIITSVNGTKVTSGSDLYRILD-QCKVGDKVTVEVLRGDHKEKIPVIL 288
G L L GD+I +VN +T+ DL L+ + +VG+ V V+++R ++ +PV L
Sbjct: 333 RVGNLNLPVGGDVIVAVNDRPITTSQDLLVYLETETQVGETVQVKIIRDGREQVVPVTL 391
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.317 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 516,714,861
Number of Sequences: 2540612
Number of extensions: 22032476
Number of successful extensions: 67584
Number of sequences better than 10.0: 993
Number of HSP's better than 10.0 without gapping: 776
Number of HSP's successfully gapped in prelim test: 218
Number of HSP's that attempted gapping in prelim test: 64857
Number of HSP's gapped (non-prelim): 1836
length of query: 295
length of database: 863,360,394
effective HSP length: 127
effective length of query: 168
effective length of database: 540,702,670
effective search space: 90838048560
effective search space used: 90838048560
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0225.5


