

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0219.2
(365 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
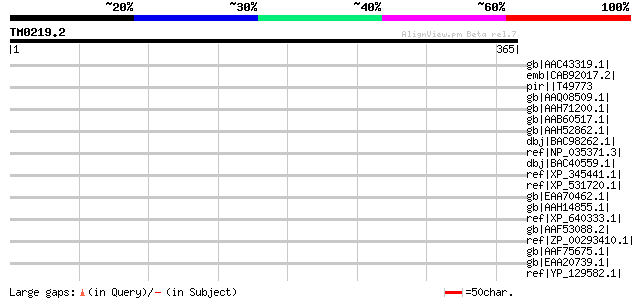
Score E
Sequences producing significant alignments: (bits) Value
gb|AAC43319.1| M protein precursor; MSzW60 gi|1096249|prf||21113... 45 0.003
emb|CAB92017.2| related to actin-interacting protein AIP3 [Neuro... 45 0.005
pir||T49773 related to actin-interacting protein AIP3 [imported]... 45 0.005
gb|AAQ08509.1| Szp protein [Streptococcus equi subsp. zooepidemi... 44 0.006
gb|AAH71200.1| Rangap1 protein [Mus musculus] gi|42490971|gb|AAH... 44 0.010
gb|AAB60517.1| RNA1 homolog 44 0.010
gb|AAH52862.1| Rangap1 protein [Mus musculus] 44 0.010
dbj|BAC98262.1| mKIAA1835 protein [Mus musculus] 44 0.010
ref|NP_035371.3| RAN GTPase activating protein 1 [Mus musculus] ... 44 0.010
dbj|BAC40559.1| unnamed protein product [Mus musculus] 44 0.010
ref|XP_345441.1| PREDICTED: similar to ABTAP [Rattus norvegicus]... 43 0.013
ref|XP_531720.1| PREDICTED: similar to Ran GTPase-activating pro... 43 0.013
gb|EAA70462.1| hypothetical protein FG00869.1 [Gibberella zeae P... 43 0.017
gb|AAH14855.1| Rangap1 protein [Mus musculus] 42 0.022
ref|XP_640333.1| hypothetical protein DDB0205159 [Dictyostelium ... 42 0.022
gb|AAF53088.2| CG6392-PA [Drosophila melanogaster] gi|24583666|r... 42 0.029
ref|ZP_00293410.1| COG1196: Chromosome segregation ATPases [Ther... 42 0.029
gb|AAF75675.1| M-like protein Szp3 precursor [Streptococcus equi... 42 0.029
gb|EAA20739.1| hypothetical protein [Plasmodium yoelii yoelii] 42 0.029
ref|YP_129582.1| Hypothetical methyl-accepting chemotaxis protei... 42 0.038
>gb|AAC43319.1| M protein precursor; MSzW60 gi|1096249|prf||2111310A M-like protein
Length = 376
Score = 45.4 bits (106), Expect = 0.003
Identities = 44/187 (23%), Positives = 81/187 (42%), Gaps = 19/187 (10%)
Query: 163 SAASIVAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTD 222
+A + +A + Q + +L EK+ A + EQ K A ++N A E+R +L+
Sbjct: 94 TAEATIASLEQKMNDL--ESKIQEKQKAFEQVVLEQKKEQANPVDRNGDAEEDRLDELSG 151
Query: 223 DL----AASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNASILG------RASL 272
++ AA ++ LQKTK +T + ++ + Q K + N + G +A
Sbjct: 152 EIRREKAALEVELQKTKEALDTAKRAYAGIEERKQVAAVKLDAANKAFAGVEEKHDQAMA 211
Query: 273 LFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQVVGDDDISLDLL------PQFDDES 326
FA+ F A KE + S + K+I V + +++ P+ + E+
Sbjct: 212 KFAEAFAAYKEAVKAELKAAGASDF-YTKKIDSANTVAGVNTLREMILDSIAKPEVEPEA 270
Query: 327 EPEEDGE 333
+PE E
Sbjct: 271 KPEPKPE 277
>emb|CAB92017.2| related to actin-interacting protein AIP3 [Neurospora crassa]
Length = 1127
Score = 44.7 bits (104), Expect = 0.005
Identities = 28/91 (30%), Positives = 45/91 (48%), Gaps = 8/91 (8%)
Query: 282 KEQISVVEPGFDLSRIGWLKEIKDGQVVGDDDISLD-------LLPQFDDESEPEEDGED 334
KE S VE G L + G ++E++ + DD I + L+ + DD+ E EED E+
Sbjct: 984 KELASFVEEG-KLKKSGGVEEVERARKAKDDQIRREVWERMNGLIAENDDDGEDEEDDEE 1042
Query: 335 GNEQHKNEDHEKEDPQAGTSQGNNANNENLA 365
E+ + E+ E+E+ + G Q E A
Sbjct: 1043 EEEEEEEEEGEEEEEEEGEEQEEGEEGEEKA 1073
>pir||T49773 related to actin-interacting protein AIP3 [imported] - Neurospora
crassa
Length = 972
Score = 44.7 bits (104), Expect = 0.005
Identities = 28/91 (30%), Positives = 45/91 (48%), Gaps = 8/91 (8%)
Query: 282 KEQISVVEPGFDLSRIGWLKEIKDGQVVGDDDISLD-------LLPQFDDESEPEEDGED 334
KE S VE G L + G ++E++ + DD I + L+ + DD+ E EED E+
Sbjct: 829 KELASFVEEG-KLKKSGGVEEVERARKAKDDQIRREVWERMNGLIAENDDDGEDEEDDEE 887
Query: 335 GNEQHKNEDHEKEDPQAGTSQGNNANNENLA 365
E+ + E+ E+E+ + G Q E A
Sbjct: 888 EEEEEEEEEGEEEEEEEGEEQEEGEEGEEKA 918
>gb|AAQ08509.1| Szp protein [Streptococcus equi subsp. zooepidemicus]
Length = 375
Score = 44.3 bits (103), Expect = 0.006
Identities = 46/187 (24%), Positives = 82/187 (43%), Gaps = 19/187 (10%)
Query: 163 SAASIVAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTD 222
+A + +A + Q + +L EK+ A + EQ K A ++N A E+R +L+
Sbjct: 93 TAEATIAELEQKMNDL--ESKIQEKQKAFEQVVLEQKKEQANPVDRNGDAEEDRLDELSG 150
Query: 223 DL----AASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNASILG------RASL 272
++ AA ++ LQKTK +T + ++ + Q K + N + G +A
Sbjct: 151 EIRREKAALEVELQKTKEALDTAKRAYAGIEERKQVAAAKLDAANKAFAGVEEKHAQAMA 210
Query: 273 LFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQVVGD----DDISLDLL--PQFDDES 326
FA+ F A KE + S + K+I V ++ LD + P+ + E+
Sbjct: 211 KFAEAFAAYKEAVKAELKAAGASDF-YTKKIDSANTVDGVKTLREMILDSIAKPEVEPEA 269
Query: 327 EPEEDGE 333
+PE E
Sbjct: 270 KPEPKPE 276
>gb|AAH71200.1| Rangap1 protein [Mus musculus] gi|42490971|gb|AAH66213.1| Rangap1
protein [Mus musculus]
Length = 589
Score = 43.5 bits (101), Expect = 0.010
Identities = 29/107 (27%), Positives = 53/107 (49%), Gaps = 14/107 (13%)
Query: 249 QAKCQKLEKKYERLNASILGRASLLFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQV 308
+A K E + LN + LG EQ+ V F+++++ L + D +
Sbjct: 315 EAVADKAELEKLDLNGNALGEEGC----------EQLQEVMDSFNMAKV--LASLSDDE- 361
Query: 309 VGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQ 355
G+D+ + + D+E E EED ED +E+ + ++ E+E PQ G+ +
Sbjct: 362 -GEDEDEEEEGEEDDEEEEDEEDEEDDDEEEEEQEEEEEPPQRGSGE 407
>gb|AAB60517.1| RNA1 homolog
Length = 589
Score = 43.5 bits (101), Expect = 0.010
Identities = 29/107 (27%), Positives = 53/107 (49%), Gaps = 14/107 (13%)
Query: 249 QAKCQKLEKKYERLNASILGRASLLFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQV 308
+A K E + LN + LG EQ+ V F+++++ L + D +
Sbjct: 315 EAVADKAELEKLDLNGNALGEEGC----------EQLQEVMDSFNMAKV--LASLSDDE- 361
Query: 309 VGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQ 355
G+D+ + + D+E E EED ED +E+ + ++ E+E PQ G+ +
Sbjct: 362 -GEDEDEEEEGEEDDEEEEDEEDEEDDDEEEEEQEEEEEPPQRGSGE 407
>gb|AAH52862.1| Rangap1 protein [Mus musculus]
Length = 589
Score = 43.5 bits (101), Expect = 0.010
Identities = 29/107 (27%), Positives = 53/107 (49%), Gaps = 14/107 (13%)
Query: 249 QAKCQKLEKKYERLNASILGRASLLFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQV 308
+A K E + LN + LG EQ+ V F+++++ L + D +
Sbjct: 315 EAVADKAELEKLDLNGNALGEEGC----------EQLQEVMDSFNMAKV--LASLSDDE- 361
Query: 309 VGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQ 355
G+D+ + + D+E E EED ED +E+ + ++ E+E PQ G+ +
Sbjct: 362 -GEDEDEEEEGEEDDEEEEDEEDEEDDDEEEEEQEEEEEPPQRGSGE 407
>dbj|BAC98262.1| mKIAA1835 protein [Mus musculus]
Length = 646
Score = 43.5 bits (101), Expect = 0.010
Identities = 29/107 (27%), Positives = 53/107 (49%), Gaps = 14/107 (13%)
Query: 249 QAKCQKLEKKYERLNASILGRASLLFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQV 308
+A K E + LN + LG EQ+ V F+++++ L + D +
Sbjct: 372 EAVADKAELEKLDLNGNALGEEGC----------EQLQEVMDSFNMAKV--LASLSDDE- 418
Query: 309 VGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQ 355
G+D+ + + D+E E EED ED +E+ + ++ E+E PQ G+ +
Sbjct: 419 -GEDEDEEEEGEEDDEEEEDEEDEEDDDEEEEEQEEEEEPPQRGSGE 464
>ref|NP_035371.3| RAN GTPase activating protein 1 [Mus musculus]
gi|1172923|sp|P46061|RGP1_MOUSE Ran GTPase-activating
protein 1 gi|472852|gb|AAA17681.1| homolog of yeast
RNA1, RNA production and processing protein, Swiss Prot
Accession Number P11745
Length = 589
Score = 43.5 bits (101), Expect = 0.010
Identities = 29/107 (27%), Positives = 53/107 (49%), Gaps = 14/107 (13%)
Query: 249 QAKCQKLEKKYERLNASILGRASLLFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQV 308
+A K E + LN + LG EQ+ V F+++++ L + D +
Sbjct: 315 EAVADKAELEKLDLNGNALGEEGC----------EQLQEVMDSFNMAKV--LASLSDDE- 361
Query: 309 VGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQ 355
G+D+ + + D+E E EED ED +E+ + ++ E+E PQ G+ +
Sbjct: 362 -GEDEDEEEEGEEDDEEEEDEEDEEDDDEEEEEQEEEEEPPQRGSGE 407
>dbj|BAC40559.1| unnamed protein product [Mus musculus]
Length = 589
Score = 43.5 bits (101), Expect = 0.010
Identities = 29/107 (27%), Positives = 53/107 (49%), Gaps = 14/107 (13%)
Query: 249 QAKCQKLEKKYERLNASILGRASLLFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQV 308
+A K E + LN + LG EQ+ V F+++++ L + D +
Sbjct: 315 EAVADKAELEKLDLNGNALGEEGC----------EQLQEVMDSFNMAKV--LASLSDDE- 361
Query: 309 VGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQ 355
G+D+ + + D+E E EED ED +E+ + ++ E+E PQ G+ +
Sbjct: 362 -GEDEDEEEEGEEDDEEEEDEEDEEDDDEEEEEQEEEEEPPQRGSGE 407
>ref|XP_345441.1| PREDICTED: similar to ABTAP [Rattus norvegicus]
gi|42491219|dbj|BAD10946.1| ABTAP [Rattus rattus]
Length = 842
Score = 43.1 bits (100), Expect = 0.013
Identities = 40/169 (23%), Positives = 72/169 (41%), Gaps = 27/169 (15%)
Query: 193 KTAYEQAKADAETANKNLKAAEERCAKLTDDLAASDLLLQKTKSLKETINDKHTAVQAKC 252
K +++ K D+E + K+ + + + K D +DL ++ + K D T+ K
Sbjct: 142 KNSHKSPKIDSEVSPKDSEESLQSRKKKRD---TTDLSVEASPKGKLRTKDPSTSAMVK- 197
Query: 253 QKLEKKYERLNASILGRASLLFAQGFLAAKEQISVV-------EPGFDLSRIGWLKEIKD 305
++++ G + Q + AK+ + E S +G +E +
Sbjct: 198 ----------SSTVSGSKAKREKQAVIMAKDNAGRMLHEEAPEEDSDSASELGRDEE-SE 246
Query: 306 GQVVGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTS 354
G++ DD S D DDE+E EE+ EDG E+ + E+ E E S
Sbjct: 247 GEITSDDRASAD-----DDENEDEEEEEDGEEEEEEEEEEDESDDESDS 290
>ref|XP_531720.1| PREDICTED: similar to Ran GTPase-activating protein 1 [Canis
familiaris]
Length = 991
Score = 43.1 bits (100), Expect = 0.013
Identities = 31/116 (26%), Positives = 58/116 (49%), Gaps = 16/116 (13%)
Query: 249 QAKCQKLEKKYERLNASILGRASLLFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQV 308
+A K E + LN ++LG EQ+ V GF+++R+ L + D +
Sbjct: 695 EAVADKAELEKLDLNGNMLGEEGC----------EQLQEVLDGFNMARV--LASLSDDE- 741
Query: 309 VGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQGNNANNENL 364
G+DD + + ++E E EE+ ED E+ + E+ E+E+PQ G + ++ + +
Sbjct: 742 -GEDDEEEEEEEEEEEEGEEEEEEED--EEEEEEEEEEEEPQQGQGEESSTPSRKI 794
>gb|EAA70462.1| hypothetical protein FG00869.1 [Gibberella zeae PH-1]
gi|46107974|ref|XP_381045.1| hypothetical protein
FG00869.1 [Gibberella zeae PH-1]
Length = 1116
Score = 42.7 bits (99), Expect = 0.017
Identities = 72/316 (22%), Positives = 119/316 (36%), Gaps = 59/316 (18%)
Query: 58 AGVKPLHQSTLDPKGRPTEKKKGHDNVP--------PH-----QPDSGALINRPSTPFIQ 104
AG K L T GRP + G+ +P PH + D +P P ++
Sbjct: 167 AGSKVLRLRTFGKDGRP---QLGNSRLPTSFANNDHPHPKNGDEDDDSDSQKKPPVPTVE 223
Query: 105 AGPSSAIGGEALPPLLNLSDPRFNGLDFMNRTFDNRIHKDVSGQGPPNIASVAIHHA-LS 163
A P G A + SD T + R V+ P I + LS
Sbjct: 224 AAPVLD-GNNA-----STSDVTATVPSQRRNTVNRRHSPSVASMDQPQIKLPLVDGPELS 277
Query: 164 AASIVAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTDD 223
+ KE+ T +Y K+ A+ + E+ K + ET + LK EE+ A+L +
Sbjct: 278 LDELNRRFESTRKEVDETLAQYAKEEAELQQQEEELKQEKETKRQALKEKEEQTAQLKAN 337
Query: 224 L--------AASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNASILGRASLLFA 275
+ AA +K + LK+ N K + V+ KLE ER+
Sbjct: 338 VRTTMEQMRAAEKDRAKKEQQLKDKEN-KKSKVRDTATKLENDIERMKKE---------R 387
Query: 276 QGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQVVGDDDISL-----DLLPQFDDESEPEE 330
+GF A K++++ K +DGQ + D + L +L + ++ + +
Sbjct: 388 EGFEAQKKELAA-------------KRDRDGQDLDDSNAELQENCAELEAELKEKGKQLQ 434
Query: 331 DGEDGNEQHKNEDHEK 346
D + E+ D E+
Sbjct: 435 DLKATREELPGADDEQ 450
>gb|AAH14855.1| Rangap1 protein [Mus musculus]
Length = 589
Score = 42.4 bits (98), Expect = 0.022
Identities = 28/107 (26%), Positives = 53/107 (49%), Gaps = 14/107 (13%)
Query: 249 QAKCQKLEKKYERLNASILGRASLLFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQV 308
+A K E + LN + LG EQ+ V F+++++ L + D +
Sbjct: 315 EAVADKAELEKLDLNGNALGEEGC----------EQLQEVMDSFNMAKV--LASLSDDE- 361
Query: 309 VGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQ 355
G+D+ + + D+E E EED ED +++ + ++ E+E PQ G+ +
Sbjct: 362 -GEDEDEEEEGEEDDEEEEDEEDEEDDDDEEEEQEEEEEPPQRGSGE 407
>ref|XP_640333.1| hypothetical protein DDB0205159 [Dictyostelium discoideum]
gi|60468350|gb|EAL66357.1| hypothetical protein
DDB0205159 [Dictyostelium discoideum]
Length = 356
Score = 42.4 bits (98), Expect = 0.022
Identities = 35/146 (23%), Positives = 64/146 (42%), Gaps = 9/146 (6%)
Query: 227 SDLLLQKTKSLKET----INDKHTAVQAKCQKLEKKYERLNASILGRASLLFAQGFLAAK 282
S+L+L+ K L E I +K + + K+ K+ + LN S+ + FL
Sbjct: 47 SELILKSLKELTENEIKNIQNKKVIKKEQFFKVRKEKKSLNKLKNRLKSIEKKKQFLLKD 106
Query: 283 EQISVVEPGFDLSRIGWL-----KEIKDGQVVGDDDISLDLLPQFDDESEPEEDGEDGNE 337
E IS + ++ R L K+ D Q++ D + L+ + ++ E E+D + N
Sbjct: 107 ESISKEDKDKEILRQNQLLNDFQKQKNDQQLILDSILKLNNVNNNNNNKENEKDNNNNNN 166
Query: 338 QHKNEDHEKEDPQAGTSQGNNANNEN 363
+ N ++ + + NN NN N
Sbjct: 167 NNNNNNNNNNNNNNNNNNNNNNNNNN 192
>gb|AAF53088.2| CG6392-PA [Drosophila melanogaster] gi|24583666|ref|NP_524993.2|
CG6392-PA [Drosophila melanogaster]
Length = 2013
Score = 42.0 bits (97), Expect = 0.029
Identities = 26/101 (25%), Positives = 50/101 (48%), Gaps = 3/101 (2%)
Query: 177 ELIATKNRYEKKAADYKTAYE---QAKADAETANKNLKAAEERCAKLTDDLAASDLLLQK 233
E+++ +++ E + + A Q D E +K L ++ +L + A D Q
Sbjct: 704 EIVSVRDKLESVESAFNLASSEIIQKATDCERLSKELSTSQNAFGQLQERYDALDQQWQA 763
Query: 234 TKSLKETINDKHTAVQAKCQKLEKKYERLNASILGRASLLF 274
++ T+++KH VQ K QKL+++YE+L + +S F
Sbjct: 764 QQAGITTLHEKHEHVQEKYQKLQEEYEQLESRARSASSAEF 804
>ref|ZP_00293410.1| COG1196: Chromosome segregation ATPases [Thermobifida fusca]
Length = 1183
Score = 42.0 bits (97), Expect = 0.029
Identities = 39/156 (25%), Positives = 69/156 (44%), Gaps = 11/156 (7%)
Query: 175 VKELIATKNRYEKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTDDLAASDLLLQKT 234
++ L ++AA + + A+ADAE+ + +E + ++LAA++ L +
Sbjct: 411 IERLTQAAAEARERAAQAQEEFADARADAESLDLGDSELDEAYEQAREELAAAEARLAEL 470
Query: 235 KSLKETINDKHTAVQAKCQKLEKKYERLNASILGRASLLFAQ----GFLAAKEQISVVEP 290
+ + + A++A+ L+ ER + G A+LL A G + A + V
Sbjct: 471 RDAERNAERERAALEARRDALQMGLERRD----GAAALLTADGDALGLIGALTSLIDVAA 526
Query: 291 GFDLSRIGWLKEIKDGQVVGDDDI---SLDLLPQFD 323
G + + L D VVG D +LDLL Q D
Sbjct: 527 GDESAIAAALGAAADAVVVGTHDAARSALDLLKQRD 562
Score = 34.7 bits (78), Expect = 4.7
Identities = 26/96 (27%), Positives = 49/96 (50%), Gaps = 3/96 (3%)
Query: 189 AADYKTAYEQAKADAETANKNLKAAEERCAKLTDDLAASDLLLQKTKSLKETINDKHTAV 248
AA + E A ETA++ L AAEE C ++T+DL + L ++ + + I +
Sbjct: 651 AATGPSQLEVRAAIDETASQ-LAAAEEVCKRITEDLGDAQLERERAAAAVDAITARRREA 709
Query: 249 QAKCQKLEKKYERLNASILGRASLLFAQGFLAAKEQ 284
+ ++ ++ RL A + R++ A+ + AA E+
Sbjct: 710 DRRRSEIAQRVGRLGAQV--RSATQEAERYTAAVEK 743
>gb|AAF75675.1| M-like protein Szp3 precursor [Streptococcus equi subsp.
zooepidemicus]
Length = 224
Score = 42.0 bits (97), Expect = 0.029
Identities = 34/131 (25%), Positives = 60/131 (44%), Gaps = 12/131 (9%)
Query: 163 SAASIVAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTD 222
+A + +A + Q + +L EK+ A + EQ K A ++N A E+R +L+
Sbjct: 95 TAEATIASLEQKMNDL--ESKIQEKQKAFEQVVLEQKKEQANPVDRNGDAEEDRLDELSG 152
Query: 223 DL----AASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNASILG------RASL 272
++ AA ++ LQKTK +T + ++ + Q K + N + G +A
Sbjct: 153 EIRREKAALEVELQKTKEALDTAKRAYAGIEERKQVAATKLDAANKAFAGVEEKHAQAMA 212
Query: 273 LFAQGFLAAKE 283
F + F A KE
Sbjct: 213 AFEEAFAAYKE 223
>gb|EAA20739.1| hypothetical protein [Plasmodium yoelii yoelii]
Length = 483
Score = 42.0 bits (97), Expect = 0.029
Identities = 36/127 (28%), Positives = 59/127 (46%), Gaps = 3/127 (2%)
Query: 163 SAASIVAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKN-LKAAEERCAKLT 221
S + A + + ELI +N EK + + T + K +T NKN +K E KL
Sbjct: 78 SKDELYADLIKFETELIKEQNENEKYSLEILTLTNKNKILTDTINKNNIKIKENEIEKLK 137
Query: 222 DDLAASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNASILGRASLLFAQGFLAA 281
+ L K K L T +++ K + +EK ++LN L + +LLF L+
Sbjct: 138 LKELKDEYL--KEKLLLTTKVEEYDKELKKYKIIEKYLDKLNKEDLNKFNLLFGLNILSL 195
Query: 282 KEQISVV 288
+EQ +V+
Sbjct: 196 EEQQNVI 202
>ref|YP_129582.1| Hypothetical methyl-accepting chemotaxis protein [Photobacterium
profundum SS9] gi|46912991|emb|CAG19780.1| Hypothetical
methyl-accepting chemotaxis protein [Photobacterium
profundum SS9]
Length = 646
Score = 41.6 bits (96), Expect = 0.038
Identities = 27/90 (30%), Positives = 45/90 (50%), Gaps = 4/90 (4%)
Query: 157 AIHHALSAA----SIVAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKNLKA 212
A+ H+L A + + +A V EL+AT N+ +D E+ KADA+ + K +
Sbjct: 388 AVSHSLENAQQQQAESSSVATAVNELLATSNQIAANISDAALTAEKVKADAQESLKINQQ 447
Query: 213 AEERCAKLTDDLAASDLLLQKTKSLKETIN 242
AE L D++ S +L++K +IN
Sbjct: 448 AESSIKGLVSDISQSQVLIEKLAQESVSIN 477
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.308 0.127 0.350
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 638,776,565
Number of Sequences: 2540612
Number of extensions: 28721798
Number of successful extensions: 206326
Number of sequences better than 10.0: 1032
Number of HSP's better than 10.0 without gapping: 357
Number of HSP's successfully gapped in prelim test: 709
Number of HSP's that attempted gapping in prelim test: 198892
Number of HSP's gapped (non-prelim): 4952
length of query: 365
length of database: 863,360,394
effective HSP length: 129
effective length of query: 236
effective length of database: 535,621,446
effective search space: 126406661256
effective search space used: 126406661256
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0219.2


