

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0209.4
(342 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
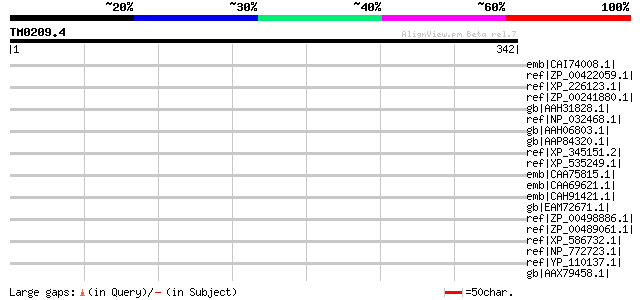
Score E
Sequences producing significant alignments: (bits) Value
emb|CAI74008.1| hypothetical protein, conserved [Theileria annul... 38 0.50
ref|ZP_00422059.1| hypothetical protein Bcep1808DRAFT_6073 [Burk... 35 2.5
ref|XP_226123.1| PREDICTED: similar to cyclin D binding myb-like... 35 3.2
ref|ZP_00241880.1| COG0665: Glycine/D-amino acid oxidases (deami... 35 3.2
gb|AAH31828.1| Kinesin heavy chain member 2 [Homo sapiens] 35 4.2
ref|NP_032468.1| kinesin family member 2A [Mus musculus] gi|2204... 35 4.2
gb|AAH06803.1| Kif2a protein [Mus musculus] 35 4.2
gb|AAP84320.1| kinesin heavy chain member 2 [Homo sapiens] 35 4.2
ref|XP_345151.2| PREDICTED: kinesin heavy chain member 2 [Rattus... 35 4.2
ref|XP_535249.1| PREDICTED: similar to Kinesin heavy chain membe... 35 4.2
emb|CAA75815.1| kinesin-like protein 2beta [Mus musculus] 35 4.2
emb|CAA69621.1| kinesin-2 [Homo sapiens] gi|3024057|sp|O00139|KI... 35 4.2
emb|CAH91421.1| hypothetical protein [Pongo pygmaeus] 35 4.2
gb|EAM72671.1| ATP-binding region, ATPase-like:Histidine kinase,... 34 5.5
ref|ZP_00498886.1| COG0542: ATPases with chaperone activity, ATP... 34 5.5
ref|ZP_00489061.1| COG0542: ATPases with chaperone activity, ATP... 34 5.5
ref|XP_586732.1| PREDICTED: similar to Kinesin-like protein KIF2... 34 5.5
ref|NP_772723.1| putative hydrolase [Bradyrhizobium japonicum US... 34 5.5
ref|YP_110137.1| putative chaperone-related protein [Burkholderi... 34 7.2
gb|AAX79458.1| hypothetical protein, conserved [Trypanosoma brucei] 34 7.2
>emb|CAI74008.1| hypothetical protein, conserved [Theileria annulata]
Length = 710
Score = 37.7 bits (86), Expect = 0.50
Identities = 28/88 (31%), Positives = 42/88 (46%), Gaps = 12/88 (13%)
Query: 186 CVDIASHRVDVSARTILVSTLGILAKVPSLGGRIRPLGVGGALPIWRSGLASSSFATAIR 245
C+DIA H VDV V T L K PS+ I P+ S LA S+ + AIR
Sbjct: 553 CMDIAQHAVDVGHPP--VETCWYLVKPPSMHCAIEPV----------SNLAISASSVAIR 600
Query: 246 AIMASSVCGVGGVEPLEPVAVDLVSTIL 273
+ +S++ + G + P+ + T+L
Sbjct: 601 DVFSSTILALSGPYMMIPMGIYASWTLL 628
>ref|ZP_00422059.1| hypothetical protein Bcep1808DRAFT_6073 [Burkholderia vietnamiensis
G4] gi|67534825|gb|EAM31566.1| hypothetical protein
Bcep1808DRAFT_6073 [Burkholderia vietnamiensis G4]
Length = 168
Score = 35.4 bits (80), Expect = 2.5
Identities = 34/108 (31%), Positives = 45/108 (41%), Gaps = 13/108 (12%)
Query: 23 PHGREPQKKLIVGDLPCPKMRNDNQSCDTTPTLT----QSESRTINSLFTSMT-SLLTVR 77
P REP++ + G N SCDT PTLT S + L TS++ S LT+
Sbjct: 6 PRAREPRRAGLSGISRVGSRPNRRSSCDTAPTLTVRFHSVASGCVGELSTSISRSPLTLE 65
Query: 78 VTDCSRH--------TDRGGLIDSPGGHALIASLMCCDGRLDRSVDSA 117
V S RG ID P + S+ CDG + + SA
Sbjct: 66 VHSVSSDRGKGHSFPNRRGRSIDLPSLTVGMHSVSSCDGWVLSTAKSA 113
>ref|XP_226123.1| PREDICTED: similar to cyclin D binding myb-like transcription
factor 1 [Rattus norvegicus]
Length = 409
Score = 35.0 bits (79), Expect = 3.2
Identities = 36/140 (25%), Positives = 57/140 (40%), Gaps = 12/140 (8%)
Query: 149 DAMADRFSRKTKLAWIESQRQDLQVVGIKDRDISQPACVDIASHRVDVSARTILVSTLGI 208
D +A+ +S W+ S+R ++ +D+S PA D + VD T+ TL
Sbjct: 84 DLLAEGWSSVRSPQWLRSKRWAIKRQIANHKDVSFPASTDSPAASVDSETITLNSGTLQT 143
Query: 209 LAKVPSLGGRIRPLGVGGALPIWRSG-------LASSSFATAIRAIMASSVCGVGGVEPL 261
+PS ++P G G + S L +S T A AS + V L
Sbjct: 144 FEILPSF--HLQPTGTPGTYLLQTSSSQGLPLTLTTSPTVTLAAAAPASPEQII--VHAL 199
Query: 262 EPV-AVDLVSTILLQCHTDR 280
P ++ + +QCHT R
Sbjct: 200 SPEHLLNTSDNVTVQCHTPR 219
>ref|ZP_00241880.1| COG0665: Glycine/D-amino acid oxidases (deaminating) [Rubrivivax
gelatinosus PM1]
Length = 444
Score = 35.0 bits (79), Expect = 3.2
Identities = 14/39 (35%), Positives = 24/39 (60%)
Query: 298 CCDGRLDRSVDSAVNRGRVSHRSGQSIISVLALEYFRMD 336
C DG +R+V V GR SH S +++++ +EY R++
Sbjct: 96 CTDGAFERNVQQLVALGRYSHESLKALVAQTGIEYHRLE 134
>gb|AAH31828.1| Kinesin heavy chain member 2 [Homo sapiens]
Length = 679
Score = 34.7 bits (78), Expect = 4.2
Identities = 19/50 (38%), Positives = 31/50 (62%), Gaps = 5/50 (10%)
Query: 153 DRFSRKTKLAWIESQRQDLQVVGIKDRDISQPACVDIASHRVDV--SART 200
D +RKTKL +E +Q +QVVG+++R++ CV+ +D+ S RT
Sbjct: 349 DLLNRKTKLRVLEDGKQQVQVVGLQEREVK---CVEDVLKLIDIGNSCRT 395
>ref|NP_032468.1| kinesin family member 2A [Mus musculus] gi|220468|dbj|BAA02165.1|
KIF2 protein [Mus musculus] gi|346875|pir||A44259
kinesin-related protein KIF2 - mouse
gi|125402|sp|P28740|KIF2_MOUSE Kinesin-like protein KIF2
Length = 716
Score = 34.7 bits (78), Expect = 4.2
Identities = 19/50 (38%), Positives = 31/50 (62%), Gaps = 5/50 (10%)
Query: 153 DRFSRKTKLAWIESQRQDLQVVGIKDRDISQPACVDIASHRVDV--SART 200
D +RKTKL +E +Q +QVVG+++R++ CV+ +D+ S RT
Sbjct: 348 DLLNRKTKLRVLEDGKQQVQVVGLQEREVK---CVEDVLKLIDIGNSCRT 394
>gb|AAH06803.1| Kif2a protein [Mus musculus]
Length = 678
Score = 34.7 bits (78), Expect = 4.2
Identities = 19/50 (38%), Positives = 31/50 (62%), Gaps = 5/50 (10%)
Query: 153 DRFSRKTKLAWIESQRQDLQVVGIKDRDISQPACVDIASHRVDV--SART 200
D +RKTKL +E +Q +QVVG+++R++ CV+ +D+ S RT
Sbjct: 348 DLLNRKTKLRVLEDGKQQVQVVGLQEREVK---CVEDVLKLIDIGNSCRT 394
>gb|AAP84320.1| kinesin heavy chain member 2 [Homo sapiens]
Length = 509
Score = 34.7 bits (78), Expect = 4.2
Identities = 19/50 (38%), Positives = 31/50 (62%), Gaps = 5/50 (10%)
Query: 153 DRFSRKTKLAWIESQRQDLQVVGIKDRDISQPACVDIASHRVDV--SART 200
D +RKTKL +E +Q +QVVG+++R++ CV+ +D+ S RT
Sbjct: 179 DLLNRKTKLRVLEDGKQQVQVVGLQEREVK---CVEDVLKLIDIGNSCRT 225
>ref|XP_345151.2| PREDICTED: kinesin heavy chain member 2 [Rattus norvegicus]
Length = 705
Score = 34.7 bits (78), Expect = 4.2
Identities = 19/50 (38%), Positives = 31/50 (62%), Gaps = 5/50 (10%)
Query: 153 DRFSRKTKLAWIESQRQDLQVVGIKDRDISQPACVDIASHRVDV--SART 200
D +RKTKL +E +Q +QVVG+++R++ CV+ +D+ S RT
Sbjct: 375 DLLNRKTKLRVLEDGKQQVQVVGLQEREVK---CVEDVLKLIDIGNSCRT 421
>ref|XP_535249.1| PREDICTED: similar to Kinesin heavy chain member 2 [Canis
familiaris]
Length = 679
Score = 34.7 bits (78), Expect = 4.2
Identities = 19/50 (38%), Positives = 31/50 (62%), Gaps = 5/50 (10%)
Query: 153 DRFSRKTKLAWIESQRQDLQVVGIKDRDISQPACVDIASHRVDV--SART 200
D +RKTKL +E +Q +QVVG+++R++ CV+ +D+ S RT
Sbjct: 349 DLLNRKTKLRVLEDGKQQVQVVGLQEREVK---CVEDVLKLIDIGNSCRT 395
>emb|CAA75815.1| kinesin-like protein 2beta [Mus musculus]
Length = 659
Score = 34.7 bits (78), Expect = 4.2
Identities = 19/50 (38%), Positives = 31/50 (62%), Gaps = 5/50 (10%)
Query: 153 DRFSRKTKLAWIESQRQDLQVVGIKDRDISQPACVDIASHRVDV--SART 200
D +RKTKL +E +Q +QVVG+++R++ CV+ +D+ S RT
Sbjct: 329 DLLNRKTKLRVLEDGKQQVQVVGLQEREVK---CVEDVLKLIDIGNSCRT 375
>emb|CAA69621.1| kinesin-2 [Homo sapiens] gi|3024057|sp|O00139|KIF2_HUMAN
Kinesin-like protein KIF2 (Kinesin-2) (HK2)
gi|4758644|ref|NP_004511.1| kinesin heavy chain member 2
[Homo sapiens]
Length = 679
Score = 34.7 bits (78), Expect = 4.2
Identities = 19/50 (38%), Positives = 31/50 (62%), Gaps = 5/50 (10%)
Query: 153 DRFSRKTKLAWIESQRQDLQVVGIKDRDISQPACVDIASHRVDV--SART 200
D +RKTKL +E +Q +QVVG+++R++ CV+ +D+ S RT
Sbjct: 349 DLLNRKTKLRVLEDGKQQVQVVGLQEREVK---CVEDVLKLIDIGNSCRT 395
>emb|CAH91421.1| hypothetical protein [Pongo pygmaeus]
Length = 744
Score = 34.7 bits (78), Expect = 4.2
Identities = 19/50 (38%), Positives = 31/50 (62%), Gaps = 5/50 (10%)
Query: 153 DRFSRKTKLAWIESQRQDLQVVGIKDRDISQPACVDIASHRVDV--SART 200
D +RKTKL +E +Q +QVVG+++R++ CV+ +D+ S RT
Sbjct: 376 DLLNRKTKLRVLEDGKQQVQVVGLQEREVK---CVEDVLKLIDIGNSCRT 422
>gb|EAM72671.1| ATP-binding region, ATPase-like:Histidine kinase, HAMP
region:Histidine kinase A, N-terminal [Kineococcus
radiotolerans SRS30216]
Length = 509
Score = 34.3 bits (77), Expect = 5.5
Identities = 26/77 (33%), Positives = 35/77 (44%), Gaps = 12/77 (15%)
Query: 52 TPTLTQSESRTINSLFTSMTSLLTVRVTDCSRHTDRGGLIDSPGGHALIASLMCCDGRLD 111
+P+ RT+ S TSLL V V GL+ GG +A + GR+D
Sbjct: 3 SPSRRPRRPRTLRSRLVLTTSLLLVAV----------GLVI--GGVTAVALHLSLTGRVD 50
Query: 112 RSVDSAVNRGRGPSGRP 128
+D AV+RGR G P
Sbjct: 51 AQLDQAVDRGRRDVGDP 67
>ref|ZP_00498886.1| COG0542: ATPases with chaperone activity, ATP-binding subunit
[Burkholderia pseudomallei S13]
gi|67756699|ref|ZP_00495580.1| COG0542: ATPases with
chaperone activity, ATP-binding subunit [Burkholderia
pseudomallei Pasteur] gi|67711665|ref|ZP_00481471.1|
COG0542: ATPases with chaperone activity, ATP-binding
subunit [Burkholderia pseudomallei 1710b]
gi|67685213|ref|ZP_00479089.1| COG0542: ATPases with
chaperone activity, ATP-binding subunit [Burkholderia
pseudomallei 1710a]
Length = 881
Score = 34.3 bits (77), Expect = 5.5
Identities = 29/93 (31%), Positives = 38/93 (40%), Gaps = 15/93 (16%)
Query: 208 ILAKVPSLGGRIRPLGVGGALPIWRSGLA--------SSSFATAIRAIMASSVCGVGGVE 259
+L + S GRIR + LP W G +S F AI +S++ GG
Sbjct: 129 VLLSISSAFGRIRVDELDAMLPAWIDGSPEANETPYDNSDFGAAIPGEASSAMQQKGGGS 188
Query: 260 PLEPVAVDLVSTILLQCHTDRGGLIDSHVGHAL 292
PLE +DL + R G ID VG L
Sbjct: 189 PLEQFCLDLTARA-------RAGQIDPVVGREL 214
>ref|ZP_00489061.1| COG0542: ATPases with chaperone activity, ATP-binding subunit
[Burkholderia pseudomallei 668]
Length = 881
Score = 34.3 bits (77), Expect = 5.5
Identities = 29/93 (31%), Positives = 38/93 (40%), Gaps = 15/93 (16%)
Query: 208 ILAKVPSLGGRIRPLGVGGALPIWRSGLA--------SSSFATAIRAIMASSVCGVGGVE 259
+L + S GRIR + LP W G +S F AI +S++ GG
Sbjct: 129 VLLSISSAFGRIRVDELDAMLPAWIDGSPEANETPYDNSDFGAAIPGEASSAMQQKGGGS 188
Query: 260 PLEPVAVDLVSTILLQCHTDRGGLIDSHVGHAL 292
PLE +DL + R G ID VG L
Sbjct: 189 PLEQFCLDLTARA-------RAGQIDPVVGREL 214
>ref|XP_586732.1| PREDICTED: similar to Kinesin-like protein KIF2, partial [Bos
taurus]
Length = 130
Score = 34.3 bits (77), Expect = 5.5
Identities = 16/44 (36%), Positives = 28/44 (63%), Gaps = 3/44 (6%)
Query: 153 DRFSRKTKLAWIESQRQDLQVVGIKDRDISQPACVDIASHRVDV 196
D +RKTKL +E +Q +QVVG+++R++ CV+ +D+
Sbjct: 85 DLLNRKTKLRVLEDGKQQVQVVGLQEREVK---CVEDVLKLIDI 125
>ref|NP_772723.1| putative hydrolase [Bradyrhizobium japonicum USDA 110]
gi|27354361|dbj|BAC51348.1| blr6083 [Bradyrhizobium
japonicum USDA 110]
Length = 287
Score = 34.3 bits (77), Expect = 5.5
Identities = 25/70 (35%), Positives = 34/70 (47%), Gaps = 3/70 (4%)
Query: 223 GVGGALPIWRSGLASSSFATAIRAIMASSVCGVGGVEPLEPVAVDLVSTILLQCHTDRGG 282
G+GGA WR LA +F RAI A + G GG PL V++ ++ L Q G
Sbjct: 32 GIGGAARAWRHQLA--TFGDRFRAI-AWDMPGYGGSAPLARVSIAALADALQQFIEQLGA 88
Query: 283 LIDSHVGHAL 292
VGH++
Sbjct: 89 TRPILVGHSI 98
>ref|YP_110137.1| putative chaperone-related protein [Burkholderia pseudomallei
K96243] gi|52211566|emb|CAH37561.1| putative
chaperone-related protein [Burkholderia pseudomallei
K96243]
Length = 881
Score = 33.9 bits (76), Expect = 7.2
Identities = 29/93 (31%), Positives = 37/93 (39%), Gaps = 15/93 (16%)
Query: 208 ILAKVPSLGGRIRPLGVGGALPIWRSGLAS--------SSFATAIRAIMASSVCGVGGVE 259
+L + S GRIR + LP W G S F AI +S++ GG
Sbjct: 129 VLLSISSAFGRIRVDELDAMLPAWIDGSPEANETPYDDSDFGAAIPGEASSAMQQKGGGS 188
Query: 260 PLEPVAVDLVSTILLQCHTDRGGLIDSHVGHAL 292
PLE +DL + R G ID VG L
Sbjct: 189 PLEQFCLDLTARA-------RAGQIDPVVGREL 214
>gb|AAX79458.1| hypothetical protein, conserved [Trypanosoma brucei]
Length = 654
Score = 33.9 bits (76), Expect = 7.2
Identities = 18/38 (47%), Positives = 22/38 (57%), Gaps = 2/38 (5%)
Query: 106 CDGRLDRSVDSAVNRGRGPSGRPRHPTRVGRDIELTSD 143
CDGR VD A+ S RPR R G+++ELTSD
Sbjct: 229 CDGRTMDFVDKAIRNNM--SSRPRIMNRKGKELELTSD 264
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 576,259,306
Number of Sequences: 2540612
Number of extensions: 24013155
Number of successful extensions: 51518
Number of sequences better than 10.0: 23
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 51502
Number of HSP's gapped (non-prelim): 33
length of query: 342
length of database: 863,360,394
effective HSP length: 129
effective length of query: 213
effective length of database: 535,621,446
effective search space: 114087367998
effective search space used: 114087367998
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0209.4


