

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.8
(124 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
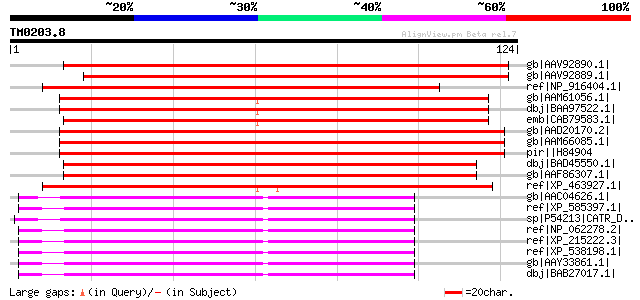
Score E
Sequences producing significant alignments: (bits) Value
gb|AAV92890.1| Avr9/Cf-9 rapidly elicited protein 20 [Nicotiana ... 149 1e-35
gb|AAV92889.1| Avr9/Cf-9 rapidly elicited protein 19 [Nicotiana ... 145 3e-34
ref|NP_916404.1| B1100D10.32 [Oryza sativa (japonica cultivar-gr... 136 1e-31
gb|AAM61056.1| EF-hand Ca2+-binding protein CCD1 [Arabidopsis th... 136 1e-31
dbj|BAA97522.1| unnamed protein product [Arabidopsis thaliana] g... 136 1e-31
emb|CAB79583.1| putative protein [Arabidopsis thaliana] gi|32692... 133 1e-30
gb|AAD20170.2| putative caltractin [Arabidopsis thaliana] gi|201... 130 7e-30
gb|AAM66085.1| putative caltractin [Arabidopsis thaliana] 130 7e-30
pir||H84904 probable caltractin [imported] - Arabidopsis thaliana 130 7e-30
dbj|BAD45550.1| putative EF-hand Ca2+-binding protein CCD1 [Oryz... 125 2e-28
gb|AAF86307.1| EF-hand Ca2+-binding protein CCD1 [Triticum aesti... 125 2e-28
ref|XP_463927.1| putative EF-hand Ca2+-binding protein CCD1 [Ory... 118 3e-26
gb|AAC04626.1| centrin [Marsilea vestita] 60 2e-08
ref|XP_585397.1| PREDICTED: similar to caltractin, partial [Bos ... 58 6e-08
sp|P54213|CATR_DUNSA Caltractin (Centrin) gi|1293682|gb|AAB67855... 58 6e-08
ref|NP_062278.2| centrin 2 [Mus musculus] gi|7619722|emb|CAB8816... 57 8e-08
ref|XP_215222.3| PREDICTED: centrin 2 [Rattus norvegicus] 57 8e-08
ref|XP_538198.1| PREDICTED: similar to centrin [Canis familiaris] 57 8e-08
gb|AAY33861.1| centrin 2 [Sus scrofa] 57 8e-08
dbj|BAB27017.1| unnamed protein product [Mus musculus] 57 8e-08
>gb|AAV92890.1| Avr9/Cf-9 rapidly elicited protein 20 [Nicotiana tabacum]
Length = 114
Score = 149 bits (377), Expect = 1e-35
Identities = 72/109 (66%), Positives = 88/109 (80%)
Query: 14 FEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVS 73
F D LP+MA KLGG+GLI ELC GF +LMDKD+ VIT ESLK N+A+LGLQD+ +D L S
Sbjct: 6 FNDFLPLMAEKLGGDGLIGELCKGFQLLMDKDKRVITFESLKKNSALLGLQDLSDDGLKS 65
Query: 74 MIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHELDNNY 122
M++EGD DGDGAL QMEFCVLMFRLSPELME+S L+EAL + + ++
Sbjct: 66 MLKEGDFDGDGALNQMEFCVLMFRLSPELMEQSQFLLDEALNQDSEFSF 114
>gb|AAV92889.1| Avr9/Cf-9 rapidly elicited protein 19 [Nicotiana tabacum]
Length = 104
Score = 145 bits (365), Expect = 3e-34
Identities = 69/104 (66%), Positives = 86/104 (82%)
Query: 19 PVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREG 78
P+MA KLGG+GLI ELC GF +LMDKD+ VIT ESLK N+A+LGLQD+ +D+L SM++EG
Sbjct: 1 PLMAEKLGGDGLIGELCKGFQLLMDKDKRVITFESLKKNSALLGLQDLSDDDLKSMLKEG 60
Query: 79 DLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHELDNNY 122
D DGDGAL QMEFCVLMFRLSPELME+S L+EAL + + ++
Sbjct: 61 DFDGDGALNQMEFCVLMFRLSPELMEQSQFLLDEALNQDSEFSF 104
>ref|NP_916404.1| B1100D10.32 [Oryza sativa (japonica cultivar-group)]
gi|20804873|dbj|BAB92555.1| putative EF-hand
Ca2+-binding protein CCD1 [Oryza sativa (japonica
cultivar-group)]
Length = 111
Score = 136 bits (343), Expect = 1e-31
Identities = 66/97 (68%), Positives = 79/97 (81%)
Query: 9 GKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKE 68
G G++FED LP MA KLG EGLI+ELC GF++LMD G IT SLK NAA+LGL ++++
Sbjct: 7 GTGVQFEDFLPSMARKLGVEGLIEELCKGFELLMDPGAGKITFRSLKRNAAMLGLGELRD 66
Query: 69 DELVSMIREGDLDGDGALTQMEFCVLMFRLSPELMEE 105
DEL M+REGDLDGDGAL QMEFCVLM RLSPELM++
Sbjct: 67 DELSEMMREGDLDGDGALDQMEFCVLMVRLSPELMQD 103
>gb|AAM61056.1| EF-hand Ca2+-binding protein CCD1 [Arabidopsis thaliana]
Length = 127
Score = 136 bits (342), Expect = 1e-31
Identities = 67/106 (63%), Positives = 83/106 (78%), Gaps = 1/106 (0%)
Query: 13 EFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAA-VLGLQDMKEDEL 71
+F+D P MA KLGGEGLI+E+C GF++LMDKD+GVIT ESL+ NA+ VLGL D+ +D++
Sbjct: 18 QFQDFFPTMAGKLGGEGLIEEICKGFELLMDKDKGVITFESLRRNASTVLGLGDLTDDDV 77
Query: 72 VSMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHE 117
MI EGD D DGAL QMEFCVLMFRLSPELME S + E ++ E
Sbjct: 78 RYMINEGDFDRDGALNQMEFCVLMFRLSPELMEASRCVVTEVIEEE 123
>dbj|BAA97522.1| unnamed protein product [Arabidopsis thaliana]
gi|15239651|ref|NP_200260.1| calcium-binding EF-hand
protein, putative [Arabidopsis thaliana]
gi|38304366|gb|AAR16086.1| KIC-related protein 2
[Arabidopsis thaliana]
Length = 127
Score = 136 bits (342), Expect = 1e-31
Identities = 67/106 (63%), Positives = 83/106 (78%), Gaps = 1/106 (0%)
Query: 13 EFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAA-VLGLQDMKEDEL 71
+F+D P MA KLGGEGLI+E+C GF++LMDKD+GVIT ESL+ NA+ VLGL D+ +D++
Sbjct: 18 QFQDFFPTMAGKLGGEGLIEEICKGFELLMDKDKGVITFESLRRNASTVLGLGDLTDDDV 77
Query: 72 VSMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHE 117
MI EGD D DGAL QMEFCVLMFRLSPELME S + E ++ E
Sbjct: 78 RYMINEGDFDRDGALNQMEFCVLMFRLSPELMEASRCVVTEVIEEE 123
>emb|CAB79583.1| putative protein [Arabidopsis thaliana] gi|3269289|emb|CAA19722.1|
putative protein [Arabidopsis thaliana]
gi|21537328|gb|AAM61669.1| EF-hand Ca2+-binding protein
CCD1 [Arabidopsis thaliana] gi|20466123|gb|AAM19983.1|
AT4g27280/M4I22_90 [Arabidopsis thaliana]
gi|15237043|ref|NP_194458.1| calcium-binding EF hand
family protein [Arabidopsis thaliana]
gi|16648777|gb|AAL25579.1| AT4g27280/M4I22_90
[Arabidopsis thaliana] gi|7486809|pir||T05752
hypothetical protein M4I22.90 - Arabidopsis thaliana
gi|38304364|gb|AAR16085.1| KIC-related protein
[Arabidopsis thaliana]
Length = 130
Score = 133 bits (334), Expect = 1e-30
Identities = 66/105 (62%), Positives = 82/105 (77%), Gaps = 1/105 (0%)
Query: 14 FEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAA-VLGLQDMKEDELV 72
F D LP MA LGGEGLI ELCNGF++LMD+++GVIT ESL+ NAA VLGL D+ ++++
Sbjct: 19 FHDFLPTMAGNLGGEGLIGELCNGFELLMDREKGVITFESLRRNAAAVLGLGDLTDEDVR 78
Query: 73 SMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHE 117
MI+EGD D DGAL QMEFCVLMFRLSP+LME S + E ++ E
Sbjct: 79 CMIKEGDFDCDGALNQMEFCVLMFRLSPDLMEASRCLVTEVIEEE 123
>gb|AAD20170.2| putative caltractin [Arabidopsis thaliana]
gi|20147271|gb|AAM10349.1| At2g46600/F13A10.13
[Arabidopsis thaliana] gi|15294244|gb|AAK95299.1|
At2g46600/F13A10.13 [Arabidopsis thaliana]
gi|18407118|ref|NP_566082.1| calcium-binding protein,
putative [Arabidopsis thaliana]
gi|38325077|gb|AAR17001.1| KCBP interacting Ca2+-binding
protein [Arabidopsis thaliana]
Length = 135
Score = 130 bits (327), Expect = 7e-30
Identities = 60/109 (55%), Positives = 82/109 (75%)
Query: 13 EFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELV 72
++ED+LPVMA K+ E + ELC GF +L D +R +IT ESL+ N+ +LG++ M +++
Sbjct: 21 KYEDMLPVMAEKMDVEEFVSELCKGFSLLADPERHLITAESLRRNSGILGIEGMSKEDAQ 80
Query: 73 SMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHELDNN 121
M+REGDLDGDGAL Q EFCVLM RLSPE+ME++ WLE+AL EL N+
Sbjct: 81 GMVREGDLDGDGALNQTEFCVLMVRLSPEMMEDAETWLEKALTQELCNH 129
>gb|AAM66085.1| putative caltractin [Arabidopsis thaliana]
Length = 118
Score = 130 bits (327), Expect = 7e-30
Identities = 60/109 (55%), Positives = 82/109 (75%)
Query: 13 EFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELV 72
++ED+LPVMA K+ E + ELC GF +L D +R +IT ESL+ N+ +LG++ M +++
Sbjct: 4 KYEDMLPVMAEKMDVEEFVSELCKGFSLLADPERHLITAESLRRNSGILGIEGMSKEDAQ 63
Query: 73 SMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHELDNN 121
M+REGDLDGDGAL Q EFCVLM RLSPE+ME++ WLE+AL EL N+
Sbjct: 64 GMVREGDLDGDGALNQTEFCVLMVRLSPEMMEDAETWLEKALTQELCNH 112
>pir||H84904 probable caltractin [imported] - Arabidopsis thaliana
Length = 156
Score = 130 bits (327), Expect = 7e-30
Identities = 60/109 (55%), Positives = 82/109 (75%)
Query: 13 EFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELV 72
++ED+LPVMA K+ E + ELC GF +L D +R +IT ESL+ N+ +LG++ M +++
Sbjct: 42 KYEDMLPVMAEKMDVEEFVSELCKGFSLLADPERHLITAESLRRNSGILGIEGMSKEDAQ 101
Query: 73 SMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHELDNN 121
M+REGDLDGDGAL Q EFCVLM RLSPE+ME++ WLE+AL EL N+
Sbjct: 102 GMVREGDLDGDGALNQTEFCVLMVRLSPEMMEDAETWLEKALTQELCNH 150
>dbj|BAD45550.1| putative EF-hand Ca2+-binding protein CCD1 [Oryza sativa (japonica
cultivar-group)]
Length = 125
Score = 125 bits (314), Expect = 2e-28
Identities = 60/101 (59%), Positives = 79/101 (77%)
Query: 14 FEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVS 73
FED LPVMA +LG EGL++EL +GF +LMD G+IT +SL+ NA +LGL M +D+L
Sbjct: 16 FEDYLPVMAERLGEEGLMQELASGFRLLMDPASGLITFDSLRRNAPLLGLGGMSDDDLRG 75
Query: 74 MIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEAL 114
M+ EGD DGDGAL++MEFCVLM RLSP+LM+E WL++A+
Sbjct: 76 MLAEGDFDGDGALSEMEFCVLMVRLSPDLMDEPRRWLDDAV 116
>gb|AAF86307.1| EF-hand Ca2+-binding protein CCD1 [Triticum aestivum]
Length = 129
Score = 125 bits (314), Expect = 2e-28
Identities = 61/101 (60%), Positives = 78/101 (76%)
Query: 14 FEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVS 73
FED LPVMA +LG EGL++EL GF +LMD G+IT +SL+ NA +LGL M +D+L
Sbjct: 19 FEDYLPVMAERLGEEGLMEELAAGFRLLMDPASGLITFDSLRRNAPLLGLGGMSDDDLRG 78
Query: 74 MIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEAL 114
M+ EGD DGDGAL+QMEFCVLM RLSP+LM+E WL++A+
Sbjct: 79 MLAEGDFDGDGALSQMEFCVLMVRLSPDLMDEPRRWLDDAV 119
>ref|XP_463927.1| putative EF-hand Ca2+-binding protein CCD1 [Oryza sativa (japonica
cultivar-group)] gi|41053013|dbj|BAD07944.1| putative
EF-hand Ca2+-binding protein CCD1 [Oryza sativa
(japonica cultivar-group)]
Length = 147
Score = 118 bits (296), Expect = 3e-26
Identities = 65/115 (56%), Positives = 82/115 (70%), Gaps = 5/115 (4%)
Query: 9 GKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAA-VLGLQ--- 64
G G E+EDL+PVMA +LG EGL+ EL GF +L D RG IT ESL+ +AA VLGL
Sbjct: 23 GGGEEYEDLMPVMAGRLGAEGLLSELRAGFRLLADPARGAITAESLRRSAASVLGLGGGG 82
Query: 65 -DMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHEL 118
+M +E +M+REGD DGDGAL++ EFCVLM RLSP +M ++ WLEEA+ EL
Sbjct: 83 GEMTVEEAAAMVREGDQDGDGALSEAEFCVLMVRLSPGIMGDAEGWLEEAIADEL 137
>gb|AAC04626.1| centrin [Marsilea vestita]
Length = 170
Score = 59.7 bits (143), Expect = 2e-08
Identities = 35/97 (36%), Positives = 55/97 (56%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G+GT+ +FED L +M K+G +E+ F + D + G I+ ++LK A LG
Sbjct: 78 GSGTI-----DFEDFLQMMTTKMGERDSKEEIMKAFRLFDDDETGKISFKNLKRVAKELG 132
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
++M ++EL MI E D DGDG + + EF +M + S
Sbjct: 133 -ENMTDEELQEMIDEADRDGDGEINEEEFYRIMKKTS 168
Score = 33.5 bits (75), Expect = 1.2
Identities = 20/64 (31%), Positives = 30/64 (46%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I + LK LG + KE E+ MI + D DG G + +F
Sbjct: 29 QEIREAFDLFDTDGSGTIDAKELKVAMRALGFEPKKE-EIKKMIADIDKDGSGTIDFEDF 87
Query: 92 CVLM 95
+M
Sbjct: 88 LQMM 91
>ref|XP_585397.1| PREDICTED: similar to caltractin, partial [Bos taurus]
Length = 171
Score = 57.8 bits (138), Expect = 6e-08
Identities = 34/97 (35%), Positives = 53/97 (54%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
GTG +N F D L VM K+ + +E+ F + D + G I+ ++LK A LG
Sbjct: 79 GTGKMN-----FSDFLTVMTQKMSEKDTKEEILKAFKLFDDDETGKISFKNLKRVAKELG 133
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
+++ ++EL MI E D DGDG + + EF +M + S
Sbjct: 134 -ENLSDEELQEMIDEADRDGDGEVNEQEFLRIMKKTS 169
Score = 33.9 bits (76), Expect = 0.92
Identities = 20/64 (31%), Positives = 31/64 (48%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I ++ LK LG + KE E+ MI E D +G G + +F
Sbjct: 30 QEIREAFDLFDADGTGTIDVKELKVAMRALGFEPKKE-EIKKMISEIDKEGTGKMNFSDF 88
Query: 92 CVLM 95
+M
Sbjct: 89 LTVM 92
>sp|P54213|CATR_DUNSA Caltractin (Centrin) gi|1293682|gb|AAB67855.1| caltractin-like
protein [Dunaliella salina]
Length = 169
Score = 57.8 bits (138), Expect = 6e-08
Identities = 34/98 (34%), Positives = 57/98 (57%), Gaps = 6/98 (6%)
Query: 2 AGTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVL 61
AG+GT+ +FE+ L +M +K+G +E+ F + D + G ITL++LK A L
Sbjct: 76 AGSGTI-----DFEEFLQMMTSKMGERDSREEIIKAFKLFDDDNTGFITLKNLKRVAKEL 130
Query: 62 GLQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
G +++ ++EL M E D +GDG + + EF +M + S
Sbjct: 131 G-ENLTDEELQEMTDEADRNGDGQIDEDEFYRIMKKTS 167
Score = 31.6 bits (70), Expect = 4.6
Identities = 20/64 (31%), Positives = 29/64 (45%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I + LK LG + KE E+ MI + D G G + EF
Sbjct: 28 QEIREAFDLFDTDGSGTIDAKELKVAMRALGFEPKKE-EIKKMIADIDKAGSGTIDFEEF 86
Query: 92 CVLM 95
+M
Sbjct: 87 LQMM 90
>ref|NP_062278.2| centrin 2 [Mus musculus] gi|7619722|emb|CAB88169.1| Caltractin [Mus
musculus] gi|15488829|gb|AAH13545.1| Centrin 2 [Mus
musculus] gi|23396523|sp|Q9R1K9|CETN2_MOUSE Centrin-2
(Caltractin isoform 1) gi|5669593|gb|AAD46391.1| centrin
[Mus musculus] gi|12835124|dbj|BAB23161.1| unnamed
protein product [Mus musculus]
Length = 172
Score = 57.4 bits (137), Expect = 8e-08
Identities = 34/97 (35%), Positives = 53/97 (54%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
GTG +N F D L VM K+ + +E+ F + D + G I+ ++LK A LG
Sbjct: 80 GTGKMN-----FSDFLTVMTQKMSEKDTKEEILKAFKLFDDDETGKISFKNLKRVAKELG 134
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
+++ ++EL MI E D DGDG + + EF +M + S
Sbjct: 135 -ENLTDEELQEMIDEADRDGDGEVNEQEFLRIMKKTS 170
Score = 34.3 bits (77), Expect = 0.71
Identities = 20/64 (31%), Positives = 31/64 (48%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I ++ LK LG + KE E+ MI E D +G G + +F
Sbjct: 31 QEIREAFDLFDADGTGTIDIKELKVAMRALGFEPKKE-EIKKMISEIDKEGTGKMNFSDF 89
Query: 92 CVLM 95
+M
Sbjct: 90 LTVM 93
>ref|XP_215222.3| PREDICTED: centrin 2 [Rattus norvegicus]
Length = 335
Score = 57.4 bits (137), Expect = 8e-08
Identities = 34/97 (35%), Positives = 53/97 (54%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
GTG +N F D L VM K+ + +E+ F + D + G I+ ++LK A LG
Sbjct: 243 GTGKMN-----FSDFLTVMTQKMSEKDTKEEILKAFKLFDDDETGKISFKNLKRVAKELG 297
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
+++ ++EL MI E D DGDG + + EF +M + S
Sbjct: 298 -ENLTDEELQEMIDEADRDGDGEVNEQEFLRIMKKTS 333
Score = 34.3 bits (77), Expect = 0.71
Identities = 20/64 (31%), Positives = 31/64 (48%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I ++ LK LG + KE E+ MI E D +G G + +F
Sbjct: 194 QEIREAFDLFDADGTGTIDMKELKVAMRALGFEPKKE-EIKKMISEIDKEGTGKMNFSDF 252
Query: 92 CVLM 95
+M
Sbjct: 253 LTVM 256
>ref|XP_538198.1| PREDICTED: similar to centrin [Canis familiaris]
Length = 250
Score = 57.4 bits (137), Expect = 8e-08
Identities = 34/97 (35%), Positives = 53/97 (54%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
GTG +N F D L VM K+ + +E+ F + D + G I+ ++LK A LG
Sbjct: 158 GTGKMN-----FSDFLTVMTQKMSEKDTKEEILKAFKLFDDDETGKISFKNLKRVAKELG 212
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
+++ ++EL MI E D DGDG + + EF +M + S
Sbjct: 213 -ENLTDEELQEMIDEADRDGDGEVNEQEFLRIMKKTS 248
Score = 33.9 bits (76), Expect = 0.92
Identities = 20/64 (31%), Positives = 31/64 (48%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I ++ LK LG + KE E+ MI E D +G G + +F
Sbjct: 109 QEIREAFDLFDADGTGTIDVKELKVAMRALGFEPKKE-EIKKMISEIDKEGTGKMNFSDF 167
Query: 92 CVLM 95
+M
Sbjct: 168 LTVM 171
>gb|AAY33861.1| centrin 2 [Sus scrofa]
Length = 139
Score = 57.4 bits (137), Expect = 8e-08
Identities = 34/97 (35%), Positives = 53/97 (54%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
GTG +N F D L VM K+ + +E+ F + D + G I+ ++LK A LG
Sbjct: 47 GTGKMN-----FSDFLTVMTQKMSEKDTKEEILKAFKLFDDDETGKISFKNLKRVAKELG 101
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
+++ ++EL MI E D DGDG + + EF +M + S
Sbjct: 102 -ENLTDEELQEMIDEADRDGDGEVNEQEFLRIMKKTS 137
Score = 32.0 bits (71), Expect = 3.5
Identities = 19/58 (32%), Positives = 28/58 (47%), Gaps = 1/58 (1%)
Query: 38 FDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
FD+ G I ++ LK LG + KE E+ MI E D +G G + +F +M
Sbjct: 4 FDLFDADGTGTIDVKELKVAMRALGFEPKKE-EIKKMISEIDKEGTGKMNFSDFLTVM 60
>dbj|BAB27017.1| unnamed protein product [Mus musculus]
Length = 172
Score = 57.4 bits (137), Expect = 8e-08
Identities = 34/97 (35%), Positives = 53/97 (54%), Gaps = 6/97 (6%)
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
GTG +N F D L VM K+ + +E+ F + D + G I+ ++LK A LG
Sbjct: 80 GTGKMN-----FSDFLTVMTQKMSEKDTKEEILKAFKLFDDDETGKISFKNLKRVAKELG 134
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
+++ ++EL MI E D DGDG + + EF +M + S
Sbjct: 135 -ENLTDEELQEMIDEADRDGDGEVNEQEFLRIMKKTS 170
Score = 35.8 bits (81), Expect = 0.24
Identities = 20/64 (31%), Positives = 31/64 (48%), Gaps = 1/64 (1%)
Query: 32 KELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEF 91
+E+ FD+ G I ++ LK LG + KE E+ MI E D +G G + +F
Sbjct: 31 QEIPEAFDLFDADGTGTIDIKELKVAMRALGFEPKKE-EIKKMISENDKEGTGKMNFSDF 89
Query: 92 CVLM 95
+M
Sbjct: 90 LTVM 93
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.317 0.138 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 221,471,757
Number of Sequences: 2540612
Number of extensions: 8517395
Number of successful extensions: 21809
Number of sequences better than 10.0: 976
Number of HSP's better than 10.0 without gapping: 452
Number of HSP's successfully gapped in prelim test: 524
Number of HSP's that attempted gapping in prelim test: 20398
Number of HSP's gapped (non-prelim): 1838
length of query: 124
length of database: 863,360,394
effective HSP length: 100
effective length of query: 24
effective length of database: 609,299,194
effective search space: 14623180656
effective search space used: 14623180656
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0203.8


