

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
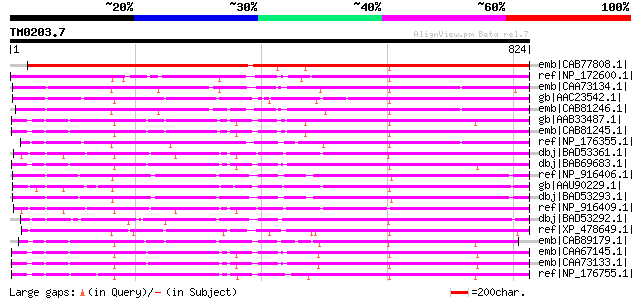
Query= TM0203.7
(824 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB77808.1| putative receptor kinase [Arabidopsis thaliana] ... 1023 0.0
ref|NP_172600.1| S-locus lectin protein kinase family protein [A... 641 0.0
emb|CAA73134.1| serine/threonine kinase [Brassica oleracea] gi|7... 625 e-177
gb|AAC23542.1| receptor protein kinase [Ipomoea trifida] 624 e-177
emb|CAB81246.1| serine/threonine kinase-like protein [Arabidopsi... 615 e-174
gb|AAB33487.1| ARK3 product/receptor-like serine/threonine prote... 614 e-174
emb|CAB81245.1| receptor-like serine/threonine protein kinase AR... 612 e-173
ref|NP_176355.1| S-locus lectin protein kinase family protein [A... 604 e-171
dbj|BAD53361.1| putative receptor-like protein kinase ARK1 [Oryz... 603 e-171
dbj|BAB69683.1| receptor kinase 5 [Brassica rapa] 601 e-170
ref|NP_916406.1| putative serine/threonine kinase [Oryza sativa ... 600 e-170
gb|AAU90229.1| putative receptor-like protein kinase [Oryza sati... 600 e-170
dbj|BAD53293.1| putative serine/threonine kinase [Oryza sativa (... 600 e-170
ref|NP_916409.1| putative receptor kinase [Oryza sativa (japonic... 599 e-169
dbj|BAD53292.1| putative serine/threonine kinase [Oryza sativa (... 598 e-169
ref|XP_478649.1| putative S-receptor kinase KIK1 precursor [Oryz... 598 e-169
emb|CAB89179.1| S-locus receptor kinase [Brassica napus var. nap... 596 e-169
emb|CAA67145.1| receptor-like kinase [Brassica oleracea var. ace... 596 e-169
emb|CAA73133.1| serine /threonine kinase [Brassica oleracea] 596 e-168
ref|NP_176755.1| S-receptor protein kinase, putative [Arabidopsi... 595 e-168
>emb|CAB77808.1| putative receptor kinase [Arabidopsis thaliana]
gi|15236268|ref|NP_192232.1| S-locus lectin protein
kinase family protein [Arabidopsis thaliana]
gi|4262151|gb|AAD14451.1| putative receptor kinase
[Arabidopsis thaliana] gi|25287707|pir||A85041 probable
receptor kinase [imported] - Arabidopsis thaliana
Length = 852
Score = 1023 bits (2645), Expect = 0.0
Identities = 506/823 (61%), Positives = 637/823 (76%), Gaps = 33/823 (4%)
Query: 29 LQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGIRYHKLAPQTVVWVANRDNPLPDSGG 88
+ D +TL+SAG FELGFFTPNGSS+ RRY+GI ++ L P TVVWVANR++P+ D
Sbjct: 36 INDSHGETLVSAGQRFELGFFTPNGSSDERRYLGIWFYNLHPLTVVWVANRESPVLDRSC 95
Query: 89 AFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMTVKLLDSGNLI-VSDDKVKKILWQSF 147
F+I++DGNL V+D G+ +W T ++ +S S VKL+D+GNL+ +SD ++WQSF
Sbjct: 96 IFTISKDGNLEVIDSKGRVYWDTGVKPSSVSAERMVKLMDNGNLVLISDGNEANVVWQSF 155
Query: 148 ANPTDTFLPGMKMDDSITLTSWRSHDDPAPGNFSFEQDQGEN-QFVIWKRSMKYWKSSVS 206
NPTDTFLPGM+MD+++TL+SWRS +DP+ GNF+F+ DQ E+ QF+IWKRSM+YWKS +S
Sbjct: 156 QNPTDTFLPGMRMDENMTLSSWRSFNDPSHGNFTFQMDQEEDKQFIIWKRSMRYWKSGIS 215
Query: 207 GKFVGTGEMSSAISYLLSNFTLRISPNN-TIPFLTSSLYSNTRLVMTYWGQLQYLKMDSM 265
GKF+G+ EM AISY LSNFT ++ +N ++P L +SLY+NTR M+ GQ QY ++D
Sbjct: 216 GKFIGSDEMPYAISYFLSNFTETVTVHNASVPPLFTSLYTNTRFTMSSSGQAQYFRLDGE 275
Query: 266 KMWLMVWVEPRDRCSVFNACGNFGSCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRK 325
+ W +W EPRD CSV+NACGNFGSCNSK + MCKCLPGFRPN +E W GDFSGGCSR+
Sbjct: 276 RFWAQIWAEPRDECSVYNACGNFGSCNSKNEEMCKCLPGFRPNFLEKWVKGDFSGGCSRE 335
Query: 326 TNVCSEDAK--SDTFLSLRMMKVGNPDAQFNAKNEEECKLECLNNCQCYAYSYEEYEKAR 383
+ +C +D D FL+L +++VG+PD+QF+A NE+EC+ ECLNNCQC AYSYEE
Sbjct: 336 SRICGKDGVVVGDMFLNLSVVEVGSPDSQFDAHNEKECRAECLNNCQCQAYSYEEV---- 391
Query: 384 QGDSVDPNAICWIWSQDLNNLEEEYEGGCDLHVRVAFSDI-----EGDSYQTERNLSSPT 438
D + N CWIW +DLNNL+E Y G ++ +RVA DI G E
Sbjct: 392 --DILQSNTKCWIWLEDLNNLKEGYLGSRNVFIRVAVPDIGSHVERGRGRYGEAKTPVVL 449
Query: 439 IILVTFTAVVFLIILSSTC--IYLRRRRQAKI-----RGIKLYGSERYVRDLIESGRLQE 491
II+VTFT+ L++LSST ++L+RR+ K RG+ L SER++++LIESGR ++
Sbjct: 450 IIVVTFTSAAILVVLSSTASYVFLQRRKVNKELGSIPRGVHLCDSERHIKELIESGRFKQ 509
Query: 492 DDAKAIDIPHFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQG 551
DD++ ID+P F LE+IL AT+NF+ ANKLGQGGFGPVYKG FPG QEIAVKRLS CSGQG
Sbjct: 510 DDSQGIDVPSFELETILYATSNFSNANKLGQGGFGPVYKGMFPGDQEIAVKRLSRCSGQG 569
Query: 552 LEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WD 603
LEEFKNEVVLIA+LQHRNLVRLLGYCV G+EK+L+YEYMP++SLD FIFD W
Sbjct: 570 LEEFKNEVVLIAKLQHRNLVRLLGYCVAGEEKLLLYEYMPHKSLDFFIFDRKLCQRLDWK 629
Query: 604 MRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVG 663
MR IILGIARGLLYLH+DSRLRIIHRDLK SNILLDEE NPKISDFGLARIFGG +T
Sbjct: 630 MRCNIILGIARGLLYLHQDSRLRIIHRDLKTSNILLDEEMNPKISDFGLARIFGGSETSA 689
Query: 664 NTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYA 723
NT+RVVGTYGYMSPEYAL+G FS KSDVFSFGVVV+E ISGKRNTGF++PE LSLLG+A
Sbjct: 690 NTNRVVGTYGYMSPEYALEGLFSFKSDVFSFGVVVIETISGKRNTGFHEPEKSLSLLGHA 749
Query: 724 WHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLG-SESN 782
W LWK R ++ +DQ L ++C+ E +KC+NVGLLC+QEDP +RPTMSNV+FMLG SE+
Sbjct: 750 WDLWKAERGIELLDQALQESCETEGFLKCLNVGLLCVQEDPNDRPTMSNVVFMLGSSEAA 809
Query: 783 TLPSPREPAFVIRRCP-SSRASTSSKMETFSRNDLTVTLENGR 824
TLP+P++PAFV+RRCP SS+AS+S+K ET S N+LT+TLE+GR
Sbjct: 810 TLPTPKQPAFVLRRCPSSSKASSSTKPETCSENELTITLEDGR 852
>ref|NP_172600.1| S-locus lectin protein kinase family protein [Arabidopsis thaliana]
Length = 840
Score = 641 bits (1654), Expect = 0.0
Identities = 365/866 (42%), Positives = 511/866 (58%), Gaps = 80/866 (9%)
Query: 2 VPIFLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYV 61
V + L C+ + + LC +D IT ++ ++D +TL+ G F GFFTP S+ RYV
Sbjct: 12 VLLLLACTCLLSRRLCFGEDRITFSSPIKDSESETLLCKSGIFRFGFFTPVNSTTRLRYV 71
Query: 62 GIRYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKN 121
GI Y K+ QTVVWVAN+D+P+ D+ G SI +DGNL V D + W TN+ +
Sbjct: 72 GIWYEKIPIQTVVWVANKDSPINDTSGVISIYQDGNLAVTDGRNRLVWSTNVSVPVAPNA 131
Query: 122 MTVKLLDSGNLIVSDDKVK-KILWQSFANPTDTFLPGMKMDD------SITLTSWRSHDD 174
V+L+DSGNL++ D++ +ILW+SF +P D+F+P M + ++ LTSW SHDD
Sbjct: 132 TWVQLMDSGNLMLQDNRNNGEILWESFKHPYDSFMPRMTLGTDGRTGGNLKLTSWTSHDD 191
Query: 175 PAPGN-------FSFEQDQGENQFVIWKRSMKYWKSSV-SGK-FVGTGEMSSAISYLLSN 225
P+ GN F+F + +IWK ++ W+S +G+ F+G M S + L
Sbjct: 192 PSTGNYTAGIAPFTFPE------LLIWKNNVPTWRSGPWNGQVFIGLPNMDSLL--FLDG 243
Query: 226 FTLRISPNNTIPFLTSSLYSNTRLVMTYWGQLQ---YLK--MDSMKMWLMVWVEPRDRCS 280
F L TI S Y+N + + + Y K SM+ W + P C
Sbjct: 244 FNLNSDNQGTI----SMSYANDSFMYHFNLDPEGIIYQKDWSTSMRTWRIGVKFPYTDCD 299
Query: 281 VFNACGNFGSCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSE--------- 331
+ CG FGSC++ + CKC+ GF P + W+ G++S GC RK + E
Sbjct: 300 AYGRCGRFGSCHAGENPPCKCVKGFVPKNNTEWNGGNWSNGCMRKAPLQCERQRNVSNGG 359
Query: 332 -DAKSDTFLSLRMMKVGNPDAQFNAKNEEECKLECLNNCQCYAYSYEEYEKARQGDSVDP 390
K+D FL L+ MKV A+ + +E+ C CL+NC C AY+Y D
Sbjct: 360 GGGKADGFLKLQKMKVPI-SAERSEASEQVCPKVCLDNCSCTAYAY------------DR 406
Query: 391 NAICWIWSQDLNNLEEEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFL 450
C +WS DL +++ G DL +RVA S+++ T NL+ ++ V+ +
Sbjct: 407 GIGCMLWSGDLVDMQSFLGSGIDLFIRVAHSELK-----THSNLA-----VMIAAPVIGV 456
Query: 451 IILSSTCIYLR----RRRQAKIRGIKLYGSERYVRDLIESGRLQEDDAKAIDIPHFHLES 506
+++++ C+ L ++R AK R +L + L + K ++P F +
Sbjct: 457 MLIAAVCVLLACRKYKKRPAKDRSAELMFKR--MEALTSDNESASNQIKLKELPLFEFQV 514
Query: 507 ILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQ 566
+ +T++F++ NKLGQGGFGPVYKGK P GQEIAVKRLS SGQGLEE NEVV+I++LQ
Sbjct: 515 LATSTDSFSLRNKLGQGGFGPVYKGKLPEGQEIAVKRLSRKSGQGLEELMNEVVVISKLQ 574
Query: 567 HRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLY 618
HRNLV+LLG C+EG+E+MLVYEYMP +SLDA++FD W RF I+ GI RGLLY
Sbjct: 575 HRNLVKLLGCCIEGEERMLVYEYMPKKSLDAYLFDPMKQKILDWKTRFNIMEGICRGLLY 634
Query: 619 LHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPE 678
LH DSRL+IIHRDLKASNILLDE NPKISDFGLARIF + NT RVVGTYGYMSPE
Sbjct: 635 LHRDSRLKIIHRDLKASNILLDENLNPKISDFGLARIFRANEDEANTRRVVGTYGYMSPE 694
Query: 679 YALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQ 738
YA++G FS KSDVFS GV+ LEIISG+RN+ ++ E+ L+LL YAW LW G D
Sbjct: 695 YAMEGFFSEKSDVFSLGVIFLEIISGRRNSSSHKEENNLNLLAYAWKLWNDGEAASLADP 754
Query: 739 TLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCP 798
+ C +E KCV++GLLC+QE +RP +SNV++ML +E+ +L P++PAF++RR
Sbjct: 755 AVFDKCFEKEIEKCVHIGLLCVQEVANDRPNVSNVIWMLTTENMSLADPKQPAFIVRRGA 814
Query: 799 SSRASTSSKMETFSRNDLTVTLENGR 824
S S+ + S ND+++T GR
Sbjct: 815 SEAESSDQSSQKVSINDVSLTAVTGR 840
>emb|CAA73134.1| serine/threonine kinase [Brassica oleracea] gi|7434416|pir||T14450
serine/threonine kinase (EC 2.7.1.-) BRLK - wild cabbage
Length = 850
Score = 625 bits (1612), Expect = e-177
Identities = 373/871 (42%), Positives = 519/871 (58%), Gaps = 82/871 (9%)
Query: 5 FLLCSFIFTFNLCSAKDTITINNNLQDGG-EDTLISAGGYFELGFFTPNGSSNGRRYVGI 63
F L F+F + +A+DTI L+DG L+S FELGFF+P GSS GR Y+GI
Sbjct: 11 FPLFIFLFLYESSTAQDTIRRGGFLRDGSTHKPLVSPQKTFELGFFSP-GSSPGR-YLGI 68
Query: 64 RYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMT 123
Y + + VVWVANR+NP+ D G +I+ DGNL +L+ + W +N+ T++ N
Sbjct: 69 WYGNIEDKAVVWVANRENPISDRSGVLTISNDGNLVLLNGQNITVWSSNITSTNNDNNRV 128
Query: 124 VKLLDSGNLIVSDDKVKKILWQSFANPTDTFLPGMKM------DDSITLTSWRSHDDPAP 177
+LD+GN + + ++++W+SF +PTDTFLP M++ D++ SWRS +DP+P
Sbjct: 129 GSILDTGNFELIEVSSERVIWESFNHPTDTFLPHMRVRVNPQTGDNLAFVSWRSENDPSP 188
Query: 178 GNFSFEQD-QGENQFVIWKRS-MKYWKSSV--SGKFVGTGEMSSAISYLLSNFTLRISPN 233
GNFS D G + V+W R+ + W+S S F G M+ +YL F L P+
Sbjct: 189 GNFSLGVDPSGAPEIVLWGRNNTRRWRSGQWNSAIFTGIPNMALLTNYLYG-FKLSSPPD 247
Query: 234 NT----IPFLTSSLYSNTRLVMTYWGQLQYLKM-DSMKMWLMVWVEPRDRCSVFNACGNF 288
T ++ S R + + G + L+ ++ K W P C +N CG+F
Sbjct: 248 ETGSVYFTYVPSDPSVLLRFKVLHNGTEEELRWNETSKRWTKFQAAPESECDKYNRCGSF 307
Query: 289 GSCNSKYDS-MCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSE----DAKSDTFLSLRM 343
G C+ + D+ +C C+ G+ P S+ NWS G C R+T + E + D FL+L+
Sbjct: 308 GICDMRGDNGICSCVKGYEPVSLGNWSRG-----CRRRTPLRCERNVSNVGEDEFLTLKS 362
Query: 344 MKVGNPDA-QFNAKNEEECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAI-CWIWSQDL 401
+K+ + + + + + E+CK CL NC C A+++ N I C IW+QDL
Sbjct: 363 VKLPDFETPEHSLADPEDCKDRCLKNCSCTAFTFV-------------NGIGCMIWNQDL 409
Query: 402 NNLEEEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLR 461
+L++ GG LHVR+A S+I G+S +T+ I+V +V +++L + L
Sbjct: 410 VDLQQFEAGGSSLHVRLADSEI-GESKKTK--------IVVIVAVLVGVLLLGIFALLLW 460
Query: 462 RRRQAKIRGIKLYGSERYVRDLIESGRLQED-------------DAKAI---DIPHFHLE 505
R ++ K G + ++ +D + KA+ ++P F L+
Sbjct: 461 RFKRKKDVSGTYCGHDADTSVVVVDMTKAKDTTTAFTGSVDIMIEGKAVNTSELPVFCLK 520
Query: 506 SILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARL 565
I+ ATN+F+ N+LG+GGFGPVYKG GQEIAVKRLS SGQG++EFKNE++LIA+L
Sbjct: 521 VIVKATNDFSRENELGRGGFGPVYKGVLEDGQEIAVKRLSGKSGQGVDEFKNEIILIAKL 580
Query: 566 QHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLL 617
QHRNLVRLLG C EG+EKMLVYEYMPN+SLD FIFD W +RF II GIARGLL
Sbjct: 581 QHRNLVRLLGCCFEGEEKMLVYEYMPNKSLDFFIFDEMKQELVDWKLRFAIIEGIARGLL 640
Query: 618 YLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSP 677
YLH DSRLRIIHRDLK SN+LLD E NPKISDFG+ARIFGG NT RVVGTYGYMSP
Sbjct: 641 YLHRDSRLRIIHRDLKVSNVLLDGEMNPKISDFGMARIFGGNQNEANTVRVVGTYGYMSP 700
Query: 678 EYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFID 737
EYA++G FSVKSDV+SFGV++LEIISGKRNT EH SL+GYAW L+ GR+ + +D
Sbjct: 701 EYAMEGLFSVKSDVYSFGVLLLEIISGKRNTSLRASEHG-SLIGYAWFLYTHGRSEELVD 759
Query: 738 QTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRC 797
+ TC E ++C++V +LC+Q+ ERP M+ VL ML S++ TLP PR+P F
Sbjct: 760 PKIRATCNKREALRCIHVAMLCVQDSAAERPNMAAVLLMLESDTATLPVPRQPTFTTSTR 819
Query: 798 PSSR----ASTSSKMETFSRNDLTVTLENGR 824
+S A SS+ S N++T T+ GR
Sbjct: 820 RNSMDVNFALDSSQQYIVSSNEITSTVVLGR 850
>gb|AAC23542.1| receptor protein kinase [Ipomoea trifida]
Length = 853
Score = 624 bits (1608), Expect = e-177
Identities = 373/864 (43%), Positives = 502/864 (57%), Gaps = 68/864 (7%)
Query: 5 FLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGIR 64
F L S IF NL A +I G TL+S+ G FELGFFTPNGS YVGI
Sbjct: 14 FFLISQIFIGNLAVALAVDSITPTQPLAGNRTLVSSDGLFELGFFTPNGSDQS--YVGIW 71
Query: 65 YHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMTV 124
Y ++ P+TVVWV NRD S G I EDGN+ ++D G W + S+++N
Sbjct: 72 YKEIEPKTVVWVGNRDGASRGSAGILKIGEDGNIHLVDGGGNFIWSPTNQ--SAARNTVA 129
Query: 125 KLLDSGNLIV---SDDKVKKILWQSFANPTDTFLPGMKMD-DSIT-----LTSWRSHDDP 175
+LLDSGN ++ D+ + LWQSF PTDT LPGMK+ DS T +++W+S +DP
Sbjct: 130 QLLDSGNFVLRREDDENPENYLWQSFDYPTDTLLPGMKLGWDSKTGLNRYISAWKSLNDP 189
Query: 176 APGNFSFEQD-QGENQFVIWKRSMKYWKSSVSG--KFVGTGEMSSAISYLLSNFTLRISP 232
G SF+ D G + + R ++S +F G EM + S +
Sbjct: 190 GEGPISFKLDINGLPEIFLRNRDKIVYRSGPWNGVRFSGVPEMKPTATITFSFVMTKNER 249
Query: 233 NNTIPFLTSSLYSNTRLVMTYWGQLQ-YLKMDSMKMWLMVWVEPRDRCSVFNACGNFGSC 291
+ +LYS RL++T G L+ Y + + K+W W P+D+C + CG FG C
Sbjct: 250 YYSFELHNKTLYS--RLLVTRNGNLERYAWIPTSKIWSKFWYAPKDQCDSYKECGTFGFC 307
Query: 292 NSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSLRMMKVGNPDA 351
++ +C+CL GFRP S + W D S GC R + + + D FL++ MK+ + +
Sbjct: 308 DTNMSPVCQCLVGFRPKSPQAWDLRDGSDGCVRYHEL---ECRKDGFLTMNFMKLPDTSS 364
Query: 352 QF--NAKNEEECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLEEEYE 409
F N +EC C NNC C AY+ G C IW+ +L L+
Sbjct: 365 SFVDTTMNLDECMKMCKNNCSCTAYTNSNISNGGSG--------CVIWTTEL--LDAAVR 414
Query: 410 GG-----CDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRR 464
GG C LH R A +G + II V +++ + + +++ +RR
Sbjct: 415 GGRRWPSC-LHPRSASDVAQGGDSGDASGRTKRIIIACGIAVGVGILLFALSALFILKRR 473
Query: 465 QAKI---RGIKLYGSERYVRDLIE-----------SGRLQEDDAKAIDIPHFHLESILDA 510
Q+K + +L G +DL+ SG D+ ++P F +I+ A
Sbjct: 474 QSKRALGKNTELRGFRDRSQDLLMNAAVIPSKREYSGETMTDE---FELPLFDFSTIVVA 530
Query: 511 TNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNL 570
T+NFA NKLGQGGFG VYKG G +EIAVKRLS SGQG+EEFKNE+ LIARLQHRNL
Sbjct: 531 TDNFADVNKLGQGGFGCVYKGMVEG-EEIAVKRLSKNSGQGVEEFKNELRLIARLQHRNL 589
Query: 571 VRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHED 622
VRLLG CV+ +EK+L+YEYM N+SLD+ +F+ W RF II GIARGLLYLH+D
Sbjct: 590 VRLLGCCVDMEEKILIYEYMENKSLDSTLFNKQRSSLLNWQTRFNIICGIARGLLYLHQD 649
Query: 623 SRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDT-VGNTDRVVGTYGYMSPEYAL 681
SR RIIHRDLKASNILLD+E NPKISDFG+ARIFGG +T NT RVVGTYGYMSPEYA+
Sbjct: 650 SRFRIIHRDLKASNILLDKEMNPKISDFGMARIFGGDETDANNTKRVVGTYGYMSPEYAM 709
Query: 682 DGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLS 741
DG FSVKSDVFSFGV+VLEI++GK+N GFY ++ +LLG+AW LW+ R + +D +
Sbjct: 710 DGLFSVKSDVFSFGVLVLEIVTGKKNRGFYNQNNQQNLLGHAWRLWRERRGSELLDSAIG 769
Query: 742 QTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSR 801
++ E M+C+ VGLLC+QE +RP M+ V+ MLGSES TLP P+ P F + P+
Sbjct: 770 ESYSLCEVMRCIQVGLLCVQEQAEDRPNMATVVLMLGSESATLPQPKHPGFCLGSRPADM 829
Query: 802 -ASTSSKMETFSRNDLTVTLENGR 824
+STS+ E+ + N +TVT+ +GR
Sbjct: 830 DSSTSNCDESCTVNQVTVTMLDGR 853
>emb|CAB81246.1| serine/threonine kinase-like protein [Arabidopsis thaliana]
gi|3402758|emb|CAA20204.1| serine/threonine kinase-like
protein [Arabidopsis thaliana]
gi|15234429|ref|NP_193870.1| S-locus lectin protein
kinase family protein [Arabidopsis thaliana]
gi|7428021|pir||T05181 S-receptor kinase (EC 2.7.1.-)
T6K22.120 precursor - Arabidopsis thaliana
Length = 849
Score = 615 bits (1586), Expect = e-174
Identities = 368/866 (42%), Positives = 508/866 (58%), Gaps = 83/866 (9%)
Query: 10 FIFTFNLCSAKDTITINNNLQDG-GEDTLISAGGYFELGFFTPNGSSNGRRYVGIRYHKL 68
+ F + A +TI +L+DG L+S FELGFF+P S++ R++GI Y +
Sbjct: 16 YFFLYESSMAANTIRRGESLRDGINHKPLVSPQKTFELGFFSPGSSTH--RFLGIWYGNI 73
Query: 69 APQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLER-TSSSKNMTVKLL 127
+ VVWVANR P+ D G I+ DGNL +LD + W +N+E T+++ N V +
Sbjct: 74 EDKAVVWVANRATPISDQSGVLMISNDGNLVLLDGKNITVWSSNIESSTTNNNNRVVSIH 133
Query: 128 DSGNLIVSDDKVKKILWQSFANPTDTFLPGMKM------DDSITLTSWRSHDDPAPGNFS 181
D+GN ++S+ + +W+SF +PTDTFLP M++ D+ SWRS DP+PGN+S
Sbjct: 134 DTGNFVLSETDTDRPIWESFNHPTDTFLPQMRVRVNPQTGDNHAFVSWRSETDPSPGNYS 193
Query: 182 FEQD-QGENQFVIWK-RSMKYWKSSV--SGKFVGTGEMSSAISYLLSNFTLRISPNNT-- 235
D G + V+W+ + W+S S F G MS +YL F L P+ T
Sbjct: 194 LGVDPSGAPEIVLWEGNKTRKWRSGQWNSAIFTGIPNMSLLTNYLYG-FKLSSPPDETGS 252
Query: 236 --IPFLTSSLYSNTRLVMTYWGQLQYLKM-DSMKMWLMVWVEPRDRCSVFNACGNFGSCN 292
++ S R + Y G + L+ +++K W EP C +N CG FG C+
Sbjct: 253 VYFTYVPSDPSVLLRFKVLYNGTEEELRWNETLKKWTKFQSEPDSECDQYNRCGKFGICD 312
Query: 293 SK-YDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSE---DAKSDTFLSLRMMKVGN 348
K + +C C+ G+ SV NWS G C R+T + E D FL+L+ +K+
Sbjct: 313 MKGSNGICSCIHGYEQVSVGNWSRG-----CRRRTPLKCERNISVGEDEFLTLKSVKL-- 365
Query: 349 PDAQF---NAKNEEECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLE 405
PD + N + E+C+ CL NC C AYS + C IW+QDL +L+
Sbjct: 366 PDFEIPEHNLVDPEDCRERCLRNCSCNAYS------------LVGGIGCMIWNQDLVDLQ 413
Query: 406 EEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRR-RR 464
+ GG LH+R+A S++ G++ +T+ I V +V +I++ + L R +R
Sbjct: 414 QFEAGGSSLHIRLADSEV-GENRKTK--------IAVIVAVLVGVILIGIFALLLWRFKR 464
Query: 465 QAKIRGI---KLYGSERYVRDLIESGRLQEDDAKAIDI------------PHFHLESILD 509
+ + G K + V DL +S + ++DI P F L +I
Sbjct: 465 KKDVSGAYCGKNTDTSVVVADLTKSKETTSAFSGSVDIMIEGKAVNTSELPVFSLNAIAI 524
Query: 510 ATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRN 569
ATN+F N+LG+GGFGPVYKG G+EIAVKRLS SGQG++EFKNE++LIA+LQHRN
Sbjct: 525 ATNDFCKENELGRGGFGPVYKGVLEDGREIAVKRLSGKSGQGVDEFKNEIILIAKLQHRN 584
Query: 570 LVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHE 621
LVRLLG C EG+EKMLVYEYMPN+SLD F+FD W +RF II GIARGLLYLH
Sbjct: 585 LVRLLGCCFEGEEKMLVYEYMPNKSLDFFLFDETKQALIDWKLRFSIIEGIARGLLYLHR 644
Query: 622 DSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYAL 681
DSRLRIIHRDLK SN+LLD E NPKISDFG+ARIFGG NT RVVGTYGYMSPEYA+
Sbjct: 645 DSRLRIIHRDLKVSNVLLDAEMNPKISDFGMARIFGGNQNEANTVRVVGTYGYMSPEYAM 704
Query: 682 DGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLS 741
+G FSVKSDV+SFGV++LEI+SGKRNT EH SL+GYAW+L+ GR+ + +D +
Sbjct: 705 EGLFSVKSDVYSFGVLLLEIVSGKRNTSLRSSEHG-SLIGYAWYLYTHGRSEELVDPKIR 763
Query: 742 QTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPS-- 799
TC E ++C++V +LC+Q+ ERP M++VL ML S++ TL +PR+P F R S
Sbjct: 764 VTCSKREALRCIHVAMLCVQDSAAERPNMASVLLMLESDTATLAAPRQPTFTSTRRNSID 823
Query: 800 -SRASTSSKMETFSRNDLTVTLENGR 824
+ A SS+ S N++T T+ GR
Sbjct: 824 VNFALDSSQQYIVSSNEITSTVVLGR 849
>gb|AAB33487.1| ARK3 product/receptor-like serine/threonine protein kinase ARK3
[Arabidopsis thaliana, Columbia, Peptide, 851 aa]
Length = 851
Score = 614 bits (1583), Expect = e-174
Identities = 366/864 (42%), Positives = 517/864 (59%), Gaps = 74/864 (8%)
Query: 4 IFLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGI 63
+ L ++ + N SA +++TI++N +T++S G FELGFF P S R Y+GI
Sbjct: 19 LILFPAYSISANTLSASESLTISSN------NTIVSPGNVFELGFFKPGLDS--RWYLGI 70
Query: 64 RYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMT 123
Y ++ +T VWVANRD PL S G I+ D NL VLD++ W TNL +
Sbjct: 71 WYKAISKRTYVWVANRDTPLSSSIGTLKIS-DSNLVVLDQSDTPVWSTNLTGGDVRSPLV 129
Query: 124 VKLLDSGNLIVSDDKVKK---ILWQSFANPTDTFLPGMKMD-DSIT-----LTSWRSHDD 174
+LLD+GN ++ D K +LWQSF PTDT LP MK+ D+ T + SW+S DD
Sbjct: 130 AELLDNGNFVLRDSKNSAPDGVLWQSFDFPTDTLLPEMKLGWDAKTGFNRFIRSWKSPDD 189
Query: 175 PAPGNFSFE-QDQGENQFVIWKRSMKYWKSSVSG--KFVGTGEMSSAISYLLSNFTL-RI 230
P+ G+FSF+ + +G + +W R + ++S +F G EM Y++ NFT +
Sbjct: 190 PSSGDFSFKLETEGFPEIFLWNRESRMYRSGPWNGIRFSGVPEMQP-FEYMVFNFTTSKE 248
Query: 231 SPNNTIPFLTSSLYSNTRLVMTYWGQLQ-YLKMDSMKMWLMVWVEPRDRCSVFNACGNFG 289
+ S +YS RL ++ G LQ + +++ + W W P+D+C + CG +G
Sbjct: 249 EVTYSFRITKSDVYS--RLSISSSGLLQRFTWIETAQNWNQFWYAPKDQCDEYKECGVYG 306
Query: 290 SCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSLRMMKVGNP 349
C+S +C C+ GF+P + + W D S GC RKT + D F+ L+ MK+ P
Sbjct: 307 YCDSNTSPVCNCIKGFKPRNPQVWGLRDGSDGCVRKTLLSC--GGGDGFVRLKKMKL--P 362
Query: 350 DAQFNAKNE----EECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLE 405
D + + +EC+ +CL +C C A++ + + G C W+ +L ++
Sbjct: 363 DTTTASVDRGIGVKECEQKCLRDCNCTAFANTDIRGSGSG--------CVTWTGELFDIR 414
Query: 406 EEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRRQ 465
+GG DL+VR+A +D+E +RN S+ I+ + V L++LS +L +R+Q
Sbjct: 415 NYAKGGQDLYVRLAATDLED-----KRNRSAK--IIGSSIGVSVLLLLSFIIFFLWKRKQ 467
Query: 466 AK-------IRGIKLYGSERYVRDLIESGRL---QEDDAKAIDIPHFHLESILDATNNFA 515
+ I +L + + +++ S R +E++ +++P E + ATNNF+
Sbjct: 468 KRSILIETPIVDHQLRSRDLLMNEVVISSRRHISRENNTDDLELPLMEFEEVAMATNNFS 527
Query: 516 IANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLG 575
ANKLGQGGFG VYKGK GQE+AVKRLS S QG +EFKNEV LIARLQH NLVRLL
Sbjct: 528 NANKLGQGGFGIVYKGKLLDGQEMAVKRLSKTSVQGTDEFKNEVKLIARLQHINLVRLLA 587
Query: 576 YCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRI 627
CV+ EKML+YEY+ N SLD+ +FD W MRF II GIARGLLYLH+DSR RI
Sbjct: 588 CCVDAGEKMLIYEYLENLSLDSHLFDKSRNSKLNWQMRFDIINGIARGLLYLHQDSRFRI 647
Query: 628 IHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSV 687
IHRDLKASNILLD+ PKISDFG+ARIFG +T NT +VVGTYGYMSPEYA+DG FS+
Sbjct: 648 IHRDLKASNILLDKYMTPKISDFGMARIFGRDETEANTRKVVGTYGYMSPEYAMDGIFSM 707
Query: 688 KSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFID----QTLSQT 743
KSDVFSFGV++LEIIS KRN GFY + +L+LLG W WK G+ L+ ID +LS T
Sbjct: 708 KSDVFSFGVLLLEIISSKRNKGFYNSDRDLNLLGCVWRNWKEGKGLEIIDPIITDSLSST 767
Query: 744 CKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRAS 803
+ E ++C+ +GLLC+QE +RPTMS V+ MLGSES T+P P+ P + + R S
Sbjct: 768 FRQHEILRCIQIGLLCVQERAEDRPTMSLVILMLGSESTTIPQPKAPGYCLERSLLDTDS 827
Query: 804 TSSKM---ETFSRNDLTVTLENGR 824
+SSK E+++ N +TV++ + R
Sbjct: 828 SSSKQRDDESWTVNQITVSVLDAR 851
>emb|CAB81245.1| receptor-like serine/threonine protein kinase ARK3 [Arabidopsis
thaliana] gi|3402757|emb|CAA20203.1| receptor-like
serine/threonine protein kinase ARK3 [Arabidopsis
thaliana] gi|29824117|gb|AAP04019.1| putative receptor
serine/threonine protein kinase ARK3 [Arabidopsis
thaliana] gi|26452798|dbj|BAC43479.1| putative
receptor-like serine/threonine protein kinase ARK3
[Arabidopsis thaliana] gi|15234427|ref|NP_193869.1|
S-locus protein kinase, putative (ARK3) [Arabidopsis
thaliana] gi|7428017|pir||T05180 S-receptor kinase (EC
2.7.1.-) ARK3 precursor - Arabidopsis thaliana
Length = 850
Score = 612 bits (1579), Expect = e-173
Identities = 365/863 (42%), Positives = 516/863 (59%), Gaps = 73/863 (8%)
Query: 4 IFLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGI 63
+ L ++ + N SA +++TI++N +T++S G FELGFF P S R Y+GI
Sbjct: 19 LILFPAYSISANTLSASESLTISSN------NTIVSPGNVFELGFFKPGLDS--RWYLGI 70
Query: 64 RYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMT 123
Y ++ +T VWVANRD PL S G I+ D NL VLD++ W TNL +
Sbjct: 71 WYKAISKRTYVWVANRDTPLSSSIGTLKIS-DSNLVVLDQSDTPVWSTNLTGGDVRSPLV 129
Query: 124 VKLLDSGNLIVSDDKVKK---ILWQSFANPTDTFLPGMKMD-DSIT-----LTSWRSHDD 174
+LLD+GN ++ D K +LWQSF PTDT LP MK+ D+ T + SW+S DD
Sbjct: 130 AELLDNGNFVLRDSKNSAPDGVLWQSFDFPTDTLLPEMKLGWDAKTGFNRFIRSWKSPDD 189
Query: 175 PAPGNFSFE-QDQGENQFVIWKRSMKYWKSSVSG--KFVGTGEMSSAISYLLSNFTL-RI 230
P+ G+FSF+ + +G + +W R + ++S +F G EM Y++ NFT +
Sbjct: 190 PSSGDFSFKLETEGFPEIFLWNRESRMYRSGPWNGIRFSGVPEMQP-FEYMVFNFTTSKE 248
Query: 231 SPNNTIPFLTSSLYSNTRLVMTYWGQLQ-YLKMDSMKMWLMVWVEPRDRCSVFNACGNFG 289
+ S +YS RL ++ G LQ + +++ + W W P+D+C + CG +G
Sbjct: 249 EVTYSFRITKSDVYS--RLSISSSGLLQRFTWIETAQNWNQFWYAPKDQCDEYKECGVYG 306
Query: 290 SCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSLRMMKVGNP 349
C+S +C C+ GF+P + + W D S GC RKT + D F+ L+ MK+ P
Sbjct: 307 YCDSNTSPVCNCIKGFKPRNPQVWGLRDGSDGCVRKTLLSC--GGGDGFVRLKKMKL--P 362
Query: 350 DAQFNAKNE----EECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLE 405
D + + +EC+ +CL +C C A++ + + G C W+ +L ++
Sbjct: 363 DTTTASVDRGIGVKECEQKCLRDCNCTAFANTDIRGSGSG--------CVTWTGELFDIR 414
Query: 406 EEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRRQ 465
+GG DL+VR+A +D+E +RN S+ I+ + V L++LS +L +R+Q
Sbjct: 415 NYAKGGQDLYVRLAATDLED-----KRNRSAK--IIGSSIGVSVLLLLSFIIFFLWKRKQ 467
Query: 466 AK-------IRGIKLYGSERYVRDLIESGRL---QEDDAKAIDIPHFHLESILDATNNFA 515
+ I +L + + +++ S R +E++ +++P E + ATNNF+
Sbjct: 468 KRSILIETPIVDHQLRSRDLLMNEVVISSRRHISRENNTDDLELPLMEFEEVAMATNNFS 527
Query: 516 IANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLG 575
ANKLGQGGFG VYKGK GQE+AVKRLS S QG +EFKNEV LIARLQH NLVRLL
Sbjct: 528 NANKLGQGGFGIVYKGKLLDGQEMAVKRLSKTSVQGTDEFKNEVKLIARLQHINLVRLLA 587
Query: 576 YCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRI 627
CV+ EKML+YEY+ N SLD+ +FD W MRF II GIARGLLYLH+DSR RI
Sbjct: 588 CCVDAGEKMLIYEYLENLSLDSHLFDKSRNSKLNWQMRFDIINGIARGLLYLHQDSRFRI 647
Query: 628 IHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSV 687
IHRDLKASNILLD+ PKISDFG+ARIFG +T NT +VVGTYGYMSPEYA+DG FS+
Sbjct: 648 IHRDLKASNILLDKYMTPKISDFGMARIFGRDETEANTRKVVGTYGYMSPEYAMDGIFSM 707
Query: 688 KSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTL---SQTC 744
KSDVFSFGV++LEIIS KRN GFY + +L+LLG W WK G+ L+ ID + S T
Sbjct: 708 KSDVFSFGVLLLEIISSKRNKGFYNSDRDLNLLGCVWRNWKEGKGLEIIDPIITDSSSTF 767
Query: 745 KAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRAST 804
+ E ++C+ +GLLC+QE +RPTMS V+ MLGSES T+P P+ P + + R S+
Sbjct: 768 RQHEILRCIQIGLLCVQERAEDRPTMSLVILMLGSESTTIPQPKAPGYCLERSLLDTDSS 827
Query: 805 SSKM---ETFSRNDLTVTLENGR 824
SSK E+++ N +TV++ + R
Sbjct: 828 SSKQRDDESWTVNQITVSVLDAR 850
>ref|NP_176355.1| S-locus lectin protein kinase family protein [Arabidopsis thaliana]
gi|25287702|pir||E96641 hypothetical protein T25B24.4
[imported] - Arabidopsis thaliana
gi|4585876|gb|AAD25549.1| Putative serine/threonine
kinase [Arabidopsis thaliana]
Length = 842
Score = 604 bits (1558), Expect = e-171
Identities = 351/847 (41%), Positives = 493/847 (57%), Gaps = 69/847 (8%)
Query: 17 CSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGIRYHKLAPQTVVWV 76
CS ++ T N+ +++G D+LIS FELGFFTP S+ RYVGI Y + PQTVVWV
Sbjct: 26 CSTSNSFTRNHTIREG--DSLISEDESFELGFFTPKNST--LRYVGIWYKNIEPQTVVWV 81
Query: 77 ANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMTVKLLDSGNLIV-S 135
ANR+ PL D GA I +DGNL +++ ++ W TN+E S N L +G+L++ S
Sbjct: 82 ANREKPLLDHKGALKIADDGNLVIVNGQNETIWSTNVE--PESNNTVAVLFKTGDLVLCS 139
Query: 136 DDKVKKILWQSFANPTDTFLPGMK------MDDSITLTSWRSHDDPAPGNFSFEQDQ-GE 188
D +K W+SF NPTDTFLPGM+ + ++ W+S DP+PG +S D G
Sbjct: 140 DSDRRKWYWESFNNPTDTFLPGMRVRVNPSLGENRAFIPWKSESDPSPGKYSMGIDPVGA 199
Query: 189 NQFVIWKRSMKYWKSSV--SGKFVGTGEMSSAISYLLSNFTLRISPNNTIPFLTSSLYSN 246
+ VIW+ + W+S S F G +M +Y+ F L P+ + + S+
Sbjct: 200 LEIVIWEGEKRKWRSGPWNSAIFTGIPDMLRFTNYIYG-FKLSSPPDRDGSVYFTYVASD 258
Query: 247 TRLVMTYW-----GQLQYLKMDSMKMWLMVWVEPRDRCSVFNACGNFGSCNS--KYDS-M 298
+ + +W + Q+ ++ W ++ +P C +N CGN+ C+ ++DS
Sbjct: 259 SSDFLRFWIRPDGVEEQFRWNKDIRNWNLLQWKPSTECEKYNRCGNYSVCDDSKEFDSGK 318
Query: 299 CKCLPGFRPNSVENWSAGDFSGGCSRKTNV-CSED---AKSDTFLSLRMMKVGNPDAQFN 354
C C+ GF P + W+ DFSGGC R+ + C++ + D F L+ +KV + +
Sbjct: 319 CSCIDGFEPVHQDQWNNRDFSGGCQRRVPLNCNQSLVAGQEDGFTVLKGIKVPDFGSVVL 378
Query: 355 AKNEEECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLEEEYEGGCDL 414
N E CK C +C C AY+ + C IW++DL ++E GG +
Sbjct: 379 HNNSETCKDVCARDCSCKAYA------------LVVGIGCMIWTRDLIDMEHFERGGNSI 426
Query: 415 HVRVAFSDIEGDSYQTERNLSSPTIILVTFTAV-VFLIILSSTCIYLRRRRQAKIRGIKL 473
++R+A S + G + T+ ++ F+ + FL+ L CI++ + + ++
Sbjct: 427 NIRLAGSKLGGGK-------ENSTLWIIVFSVIGAFLLGL---CIWILWKFKKSLKAFLW 476
Query: 474 YGSERYVRDLIESGRLQEDDAKAI--------DIPHFHLESILDATNNFAIANKLGQGGF 525
+ V D+IE+ K + D+P F +S+ AT +FA NKLGQGGF
Sbjct: 477 KKKDITVSDIIENRDYSSSPIKVLVGDQVDTPDLPIFSFDSVASATGDFAEENKLGQGGF 536
Query: 526 GPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKML 585
G VYKG F G+EIAVKRLS S QGLEEFKNE++LIA+LQHRNLVRLLG C+E +EKML
Sbjct: 537 GTVYKGNFSEGREIAVKRLSGKSKQGLEEFKNEILLIAKLQHRNLVRLLGCCIEDNEKML 596
Query: 586 VYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNI 637
+YEYMPN+SLD F+FD W R+++I GIARGLLYLH DSRL+IIHRDLKASNI
Sbjct: 597 LYEYMPNKSLDRFLFDESKQGSLDWRKRWEVIGGIARGLLYLHRDSRLKIIHRDLKASNI 656
Query: 638 LLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVV 697
LLD E NPKISDFG+ARIF + NT RVVGTYGYM+PEYA++G FS KSDV+SFGV+
Sbjct: 657 LLDTEMNPKISDFGMARIFNYRQDHANTIRVVGTYGYMAPEYAMEGIFSEKSDVYSFGVL 716
Query: 698 VLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGL 757
+LEI+SG++N F +H SL+GYAWHLW G+T + ID + T E M+C++VG+
Sbjct: 717 ILEIVSGRKNVSFRGTDHG-SLIGYAWHLWSQGKTKEMIDPIVKDTRDVTEAMRCIHVGM 775
Query: 758 LCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSSKMETFSRNDLT 817
LC Q+ I RP M +VL ML S+++ LP PR+P F + S ND+T
Sbjct: 776 LCTQDSVIHRPNMGSVLLMLESQTSQLPPPRQPTFHSFLNSGDIELNFDGHDVASVNDVT 835
Query: 818 VTLENGR 824
T GR
Sbjct: 836 FTTIVGR 842
>dbj|BAD53361.1| putative receptor-like protein kinase ARK1 [Oryza sativa (japonica
cultivar-group)]
Length = 846
Score = 603 bits (1556), Expect = e-171
Identities = 354/859 (41%), Positives = 505/859 (58%), Gaps = 60/859 (6%)
Query: 7 LCSFIFTFNL-------CSAKDTITINNNLQDGGEDTLISAG-GYFELGFFTPNGSSNGR 58
LC ++ F + C A+DT+ L +TL+S G F LGFFTP G+++
Sbjct: 7 LCCYLLLFVVVVVLTGSCRARDTVVPGRPL--AANETLVSGGDANFVLGFFTPPGANS-- 62
Query: 59 RYVGIRYHKLAPQTVVWVANRDNPLP-----DSGGAFSITEDGNLRVLDRTGKSFWGTNL 113
YVG+ Y+K++ +TVVWVANR++PLP + S++ G L ++ W
Sbjct: 63 TYVGVWYNKVSVRTVVWVANREDPLPGDVADNPDATLSVSPTGTLAIVAGNSTVVWSVTP 122
Query: 114 ERTSSSKNMTVKLLDSGNLIVSDDKVKKILWQSFANPTDTFLPGMKMDDSI------TLT 167
+S T +++DSGNL+++D + WQ F PTDT LP M++ TLT
Sbjct: 123 AAKLASP--TARIMDSGNLVIADGAGGGVAWQGFDYPTDTLLPEMRLGVDYVKGRNRTLT 180
Query: 168 SWRSHDDPAPGNFSFEQD-QGENQFVIWKRSMKYWKSSVSGKFVGTGEMSSAISYLLSNF 226
+W+S DP+PG D G+ Q IW + K W+S TG + ++Y S F
Sbjct: 181 AWKSPSDPSPGPVVMAMDTSGDPQVFIWNGAEKVWRSGPWDGVQFTG-VPDTVTY--SGF 237
Query: 227 TLRISPNN---TIPFLTSSLYSNTRLVMTYWGQLQYLK----MDSMKMWLMVWVEPRDRC 279
T N T F ++ +RL + G L+ +++ W + W P+D+C
Sbjct: 238 TFSFINNAKEVTYSFQVHNVSIISRLGLNSTGSYGLLQRSTWVEAAGTWNLYWYAPKDQC 297
Query: 280 SVFNACGNFGSCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFL 339
+ CG G C++ +C CL GF P S E W+ D GC R T + ++ +D F+
Sbjct: 298 DEVSPCGANGVCDTNNLPVCSCLRGFTPKSPEAWALRDGRAGCVRSTPLDCQNG-TDGFV 356
Query: 340 SLRMMKVGNPDAQFNAKNE----EECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICW 395
++ KV PD + + + E+C+ CL NC C AY+ +G C
Sbjct: 357 AVEHAKV--PDTERSVVDLGLSLEQCRKACLMNCSCTAYASANVSGGGRGHGAGTG--CV 412
Query: 396 IWSQDLNNLEEEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSS 455
+W+ L +L E G DL VR+A +D+ S + + I+V+ ++V FL +L+
Sbjct: 413 MWTTGLTDLRVYPEFGQDLFVRLAAADLGLTSKSNKARVI--IAIVVSISSVTFLSVLAG 470
Query: 456 TCIYLRRRRQAKIRGI-KLYGSERYVRDLIESGRLQEDDAKAIDIPHFHLESILDATNNF 514
++ R++++A+ G K G R E +DD +++P F L +I AT+ F
Sbjct: 471 FLVWTRKKKRARKTGSSKWSGGSRSTGRRYEGSSHHDDD---LELPIFDLGTIAAATDGF 527
Query: 515 AIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLL 574
+I NKLG+GGFGPVYKGK GQEIAVK LS S QGL+EFKNEV+LIA+LQHRNLVRLL
Sbjct: 528 SINNKLGEGGFGPVYKGKLEDGQEIAVKTLSKTSVQGLDEFKNEVMLIAKLQHRNLVRLL 587
Query: 575 GYCVEGDEKMLVYEYMPNRSLDAFIF--------DWDMRFKIILGIARGLLYLHEDSRLR 626
G+ + G E++LVYEYM N+SLD F+F DW R++II GI RGLLYLH+DSR R
Sbjct: 588 GFSISGQERILVYEYMANKSLDYFLFEKSNSVLLDWQARYRIIEGITRGLLYLHQDSRYR 647
Query: 627 IIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFS 686
IIHRDLKASN+LLD+E PKISDFG+AR+FG ++T NT +VVGTYGYMSPEYA+DG FS
Sbjct: 648 IIHRDLKASNVLLDKEMTPKISDFGMARMFGSEETEINTRKVVGTYGYMSPEYAMDGVFS 707
Query: 687 VKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKA 746
VKSDVFSFGV++LEIISG+RN G Y + L+LLG+AW LW G++L+ D+T++ + +
Sbjct: 708 VKSDVFSFGVLLLEIISGRRNRGVYSYSNHLNLLGHAWSLWNEGKSLELADETMNGSFDS 767
Query: 747 EECMKCVNVGLLCLQEDPIERPTMSNVLFMLG-SESNTLPSPREPAFVIRRCPSSRASTS 805
+E +KC+ VGLLC+QE+P +RP MS VL ML +++ TLP+P++P F RR ++S
Sbjct: 768 DEVLKCIRVGLLCVQENPDDRPLMSQVLLMLATTDATTLPTPKQPGFAARRILMETDTSS 827
Query: 806 SKMETFSRNDLTVTLENGR 824
SK + + TVT+ GR
Sbjct: 828 SKPDCSIFDSATVTILEGR 846
>dbj|BAB69683.1| receptor kinase 5 [Brassica rapa]
Length = 838
Score = 601 bits (1549), Expect = e-170
Identities = 357/860 (41%), Positives = 508/860 (58%), Gaps = 69/860 (8%)
Query: 4 IFLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGI 63
+ L +F F+ N SA +++TI++N T+ S G FELGFF P+ SS R Y+GI
Sbjct: 9 LLLFPAFSFSANTLSATESLTISSN------KTISSPGNIFELGFFKPSSSS--RWYLGI 60
Query: 64 RYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMT 123
Y ++ +T VWVANRD+PL S G I+ D NL V+D + + W TNL ++
Sbjct: 61 WYKAISKRTYVWVANRDHPLSTSTGTLKIS-DSNLVVVDGSDTAVWSTNLTGGGDVRSPV 119
Query: 124 V-KLLDSGNLIVSDDKVKK---ILWQSFANPTDTFLPGMKMDDSIT------LTSWRSHD 173
V +LLD+GNL++ D +LWQSF PTDT LP MK+ + L SW+S D
Sbjct: 120 VAELLDNGNLVLRDSNNNDPDGVLWQSFDFPTDTLLPEMKLGWDLKTGFNRFLRSWKSPD 179
Query: 174 DPAPGNFSFE-QDQGENQFVIWKRSMKYWKSSVSG--KFVGTGEMSSAISYLLSNFTLRI 230
DP+ G++SF+ + +G + +W ++ + ++S +F G EM Y+ NFT
Sbjct: 180 DPSSGDYSFKLETRGFPEAFLWNKASQVYRSGPWNGIRFSGVPEMQP-FDYIEFNFTTS- 237
Query: 231 SPNNTIPFLTSSLYSNTRLVMTYWGQLQ-YLKMDSMKMWLMVWVEPRDRCSVFNACGNFG 289
+ T F + +RL ++ G LQ + +++++ W W P+D+C + CG FG
Sbjct: 238 NQEVTYSFHITKDNMYSRLSLSSTGSLQRFTWIEAIQNWNQFWYAPKDQCDEYKECGTFG 297
Query: 290 SCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSLRMMKVGNP 349
C+S +C C+ GF P + + W+ D S GC RKT + D F+ L+ MK+ P
Sbjct: 298 YCDSNTYPVCNCMRGFEPRNPQAWALRDGSDGCVRKTALSCNGG--DGFVRLKKMKL--P 353
Query: 350 DAQFNAKNE----EECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLE 405
D + + +EC+ +C ++C C A++ + G C +W+ D+ +
Sbjct: 354 DTAATSVDRGIGIKECEEKCKSDCNCTAFANTDIRGGGSG--------CVVWTGDILDTR 405
Query: 406 EEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRRQ 465
+GG DL+VR+A +D+E T RN I+ + V L++L +R+Q
Sbjct: 406 NYAKGGQDLYVRLAATDLEDT---TNRNAK----IIGSCIGVSVLLLLCFIFYRFWKRKQ 458
Query: 466 AKIRGIK---LYGSERYVRDLIESGRL---QEDDAKAIDIPHFHLESILDATNNFAIANK 519
+ I+ + + + +++ R +E+ ++P E++ AT+NF ANK
Sbjct: 459 KRSIAIETSFVRSQDLLMNEVVIPSRRHISRENKTDDFELPLMDFEAVAIATDNFTNANK 518
Query: 520 LGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVE 579
LGQGGFG VYKG+ GQEIAVKRLS S QG +EFKNEV LIARLQH NLVRLLG CV+
Sbjct: 519 LGQGGFGIVYKGRLLDGQEIAVKRLSKMSVQGTDEFKNEVKLIARLQHINLVRLLGCCVD 578
Query: 580 GDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRIIHRD 631
EKML+YEY+ N SLD+ +FD W RF I GIARGLLYLH+DSR RIIHRD
Sbjct: 579 EGEKMLIYEYLENLSLDSHLFDKTRSCKLNWQKRFDITNGIARGLLYLHQDSRFRIIHRD 638
Query: 632 LKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDV 691
LKASN+LLD++ PKISDFG+ARIFG +T NT +VVGTYGYMSPEYA+DG FS KSDV
Sbjct: 639 LKASNVLLDKDMTPKISDFGMARIFGRDETEANTRKVVGTYGYMSPEYAMDGIFSTKSDV 698
Query: 692 FSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTL----SQTCKAE 747
FSFGV++LEIISGKRN GFY +H+L+LLG W WK G+ LD +D + T +
Sbjct: 699 FSFGVLLLEIISGKRNKGFYNSDHDLNLLGCVWRNWKKGKGLDIVDPIILDSSPSTYRPL 758
Query: 748 ECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSSK 807
E ++C+ +GLLC+QE +RPTMS+V+ MLGSE+ +P P P + + R P S+SS
Sbjct: 759 EILRCIKIGLLCVQERANDRPTMSSVVMMLGSETTAIPQPEPPGYCVGRSPLDTDSSSSN 818
Query: 808 M---ETFSRNDLTVTLENGR 824
E++S N +TV++ + R
Sbjct: 819 QRNDESWSVNQMTVSVIDPR 838
>ref|NP_916406.1| putative serine/threonine kinase [Oryza sativa (japonica
cultivar-group)]
Length = 807
Score = 600 bits (1546), Expect = e-170
Identities = 357/840 (42%), Positives = 492/840 (58%), Gaps = 69/840 (8%)
Query: 5 FLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGIR 64
FL+ + T L S T ++ N Q T++SA F LGFF+P S+ RYVGI
Sbjct: 17 FLILLVLSTCCLSSTITTDSLLPNKQISDGQTIVSANETFTLGFFSPGTSTY--RYVGIW 74
Query: 65 YHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMTV 124
Y + +TVVWVANR+NP+ D+ G GNL +LD G SF + S +K+
Sbjct: 75 YSNVPNRTVVWVANRNNPVLDTSGILMFDTSGNLVILDGRGSSF---TVAYGSGAKDTEA 131
Query: 125 KLLDSGNLIV-SDDKVKKILWQSFANPTDTFLPGMKMD----DSITLTSWRSHDDPAPGN 179
+LDSGNL++ S ++ WQSF PTDT+L GM + + LTSWRS DDPA G+
Sbjct: 132 TILDSGNLVLRSVSNRSRLRWQSFDYPTDTWLQGMNLGFVGAQNQLLTSWRSSDDPAIGD 191
Query: 180 FSFEQDQGEN-QFVIWKRSMKYWKSSV-SGKFVGTGEMSSAISYLLSNFTLRISPNNTIP 237
+SF D E F IW+R YWKS + +G+ E S +SN T+
Sbjct: 192 YSFGMDPNEKGDFFIWERGNVYWKSGLWNGQSYNFTESESMSFLYVSN-----DARTTLS 246
Query: 238 FLTSSLYSNTRLVMTYWGQLQYL-KMDS-MKMWLMVWVEPRDRCSVFNACGNFGSC--NS 293
+ + R V+ + GQL+ L +MD + WL++ P C ++ CG FG C N
Sbjct: 247 YSSIPASGMVRYVLDHSGQLKLLERMDFVLHQWLVLGSWPEGSCKAYSPCGAFGICAGNQ 306
Query: 294 KYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSLRMMKV-GNPDAQ 352
+ + CKC GF P WS+GD GC R+TN+ D F + M + GN
Sbjct: 307 DWQNRCKCPKGFNPGDGVGWSSGDTRRGCIRQTNM---HCVGDKFFQMPDMGLPGNATTI 363
Query: 353 FNAKNEEECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLEEEYEGGC 412
+ +++C+ CL NC C AY+ + + C +W ++ NL E G
Sbjct: 364 SSITGQKQCESTCLTNCSCTAYAVLQDK-------------CSLWYGNIMNLREGESGDA 410
Query: 413 --DLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRRQAKIRG 470
++R+A S++E + +I T ++V FLI S +++ R++ +K +G
Sbjct: 411 VGTFYLRLAASELESRG-------TPVVLIAATVSSVAFLIFASLIFLWMWRQK-SKAKG 462
Query: 471 IKLYGSERYVRDLIESGRLQEDDAKAIDIPHFHLESILDATNNFAIANKLGQGGFGPVYK 530
+ D + +L E + F I DAT F++ NKLG+GGFGPVYK
Sbjct: 463 V----------DTDSAIKLWESEETGSHFTSFCFSEIADATCKFSLENKLGEGGFGPVYK 512
Query: 531 GKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYM 590
G P GQEIAVKRL++ SGQGL EFKNE++LIA+LQHRNLVRLLG C++G+EK+L+YEYM
Sbjct: 513 GNLPEGQEIAVKRLAAHSGQGLLEFKNEIMLIAKLQHRNLVRLLGCCIQGEEKILIYEYM 572
Query: 591 PNRSLDAFIFDWDM----RFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPK 646
PN+SLD F+F + II GIA+GLLYLH+ SR RIIHRDLKASNILLD + NPK
Sbjct: 573 PNKSLDFFLFAGQVIQCGLEGIIEGIAQGLLYLHKHSRFRIIHRDLKASNILLDIDMNPK 632
Query: 647 ISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKR 706
ISDFG+ARIFG K+T NT+RVVGTYGYM+PEYA++G FSVKSDVFSFGV++LEI+SG R
Sbjct: 633 ISDFGMARIFGSKETEANTNRVVGTYGYMAPEYAMEGIFSVKSDVFSFGVLLLEIVSGIR 692
Query: 707 NTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIE 766
N GF+Q + L+LL YAW LWK GR + D ++ C + ++C++VGL+C+QE PI
Sbjct: 693 NAGFHQRGNSLNLLCYAWELWKEGRWSELADPSIYNACPEHKVLRCIHVGLMCVQESPIN 752
Query: 767 RPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSSKME--TFSRNDLTVTLENGR 824
RPTM+ ++ L +ES TLP P++PAFV S+ T + + T S N +T++ GR
Sbjct: 753 RPTMTEIISALDNESTTLPEPKQPAFV-----SAGIWTEAGVHGGTHSINGMTISDTQGR 807
>gb|AAU90229.1| putative receptor-like protein kinase [Oryza sativa (japonica
cultivar-group)]
Length = 837
Score = 600 bits (1546), Expect = e-170
Identities = 360/856 (42%), Positives = 501/856 (58%), Gaps = 62/856 (7%)
Query: 5 FLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAG----GYFELGFFTPNGSSNGRRY 60
F+L + A+D+I L G DTL+SAG G F LGFFTP GS++ Y
Sbjct: 8 FVLLLMLLAPATSRARDSIAPGEPL--AGHDTLVSAGAGDGGGFALGFFTPPGSND--TY 63
Query: 61 VGIRYHKLAPQTVVWVANRDNPLP-----DSGGAFSITEDGNLRVLDRTGKSFWGTNLER 115
VG+ Y +++P+TVVWVANR +P+P ++G S++ L V D W
Sbjct: 64 VGVWYARVSPRTVVWVANRADPVPGPVDGNAGATLSVSRACELAVADANSTVVWSVTPAT 123
Query: 116 TSSSKNMTVKLLDSGNLIVSDDKVKKILWQSFANPTDTFLPGMKMD------DSITLTSW 169
T T ++ D GNL+V+D++ ++ WQ F +PTDT LPGM++ +++TLT+W
Sbjct: 124 TGPC---TARIRDDGNLVVTDER-GRVAWQGFDHPTDTLLPGMRIGVDFAAGNNMTLTAW 179
Query: 170 RSHDDPAPGNFSFEQD-QGENQFVIWKRSMKYWKSSVSGKFVGTGEMSSAISYLLSNFTL 228
+S DP+P + D G+ + +W K W+S TG + I+Y +F+
Sbjct: 180 KSPSDPSPSSVVVAMDTSGDPEVFLWNGPNKVWRSGPWDGMQFTG-VPDTITYKNFSFSF 238
Query: 229 RISPNN-TIPFLTSSLYSNTRLVMTYWGQ---LQYLKMDSMKMWLMVWVEPRDRCSVFNA 284
S T F +RLV+ G ++ +++ W + W P+D+C +
Sbjct: 239 VNSAREVTYSFQVPDASIMSRLVLNSSGGGLVQRWTWVEAAGAWNLYWYAPKDQCDAVSP 298
Query: 285 CGNFGSCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSLRMM 344
CG G C++ +C CL GF P S W+ D GC+R+T + + +D F +R
Sbjct: 299 CGANGVCDTNSLPVCSCLRGFAPRSPAAWALRDGRDGCARETPLGCANG-TDGFAVVRHA 357
Query: 345 KVGNPDA---QFNAKNEEECKLECLNNCQCYAYSYEEYEK--ARQGDSVDPNAICWIWSQ 399
K + A ++A + C+ CL NC C AY+ R+G C +W+
Sbjct: 358 KAPDTTAATVDYDA-GLQLCRRRCLGNCSCTAYANANLSAPPGRRG--------CVMWTG 408
Query: 400 DLNNLEEEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIY 459
+L +L G DL+VR+A +D++ S ++ + II V + IIL+ T +Y
Sbjct: 409 ELEDLRVYPAFGQDLYVRLAAADLDSTSKSKKK---THIIIAVVVSICALAIILALTGMY 465
Query: 460 LRRRRQAKIR--GIKLYGSERYVRDLIESGRLQEDDAKAIDIPHFHLESILDATNNFAIA 517
+ R ++ K R G + + R+L G DD +D+P F LE+I ATN F+
Sbjct: 466 IWRTKKTKARRQGPSNWSGGLHSRELHSEGNSHGDD---LDLPLFDLETIASATNGFSAD 522
Query: 518 NKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYC 577
NKLG+GGFGPVYKG GQEIAVK LS S QGL+EF+NEV+LIA+LQHRNLV+L+GY
Sbjct: 523 NKLGEGGFGPVYKGTLEDGQEIAVKTLSKTSVQGLDEFRNEVMLIAKLQHRNLVQLIGYS 582
Query: 578 VEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRIIH 629
V G EKML+YE+M N+SLD F+FD W R+ II GIARGLLYLH+DSR RIIH
Sbjct: 583 VCGQEKMLLYEFMENKSLDCFLFDKSKSKLLDWQTRYHIIEGIARGLLYLHQDSRYRIIH 642
Query: 630 RDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKS 689
RDLK SNILLD+E PKISDFG+AR+FG DT NT RVVGTYGYM+PEYA+DG FSVKS
Sbjct: 643 RDLKTSNILLDKEMTPKISDFGMARMFGSDDTEINTVRVVGTYGYMAPEYAMDGVFSVKS 702
Query: 690 DVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEEC 749
DVFSFGV+VLEIISGKRN G Y L+LL AW W G +LD +D+TL+ + EE
Sbjct: 703 DVFSFGVIVLEIISGKRNRGVYSYSSHLNLLARAWSSWSEGNSLDLVDKTLNGSFNQEEV 762
Query: 750 MKCVNVGLLCLQEDPIERPTMSNVLFMLGS-ESNTLPSPREPAFVIRRCPSSRASTSSKM 808
+KC+ VGLLC+QE+P +RP MS VL ML S ++ +LP PR+P FV RR ++ ++SS+
Sbjct: 763 LKCLKVGLLCVQENPDDRPLMSQVLLMLASADATSLPDPRKPGFVARRA-ATEDTSSSRP 821
Query: 809 ETFSRNDLTVTLENGR 824
+ + +T+T+ GR
Sbjct: 822 DCSFVDSMTITMIEGR 837
>dbj|BAD53293.1| putative serine/threonine kinase [Oryza sativa (japonica
cultivar-group)] gi|53792448|dbj|BAD53356.1| putative
serine/threonine kinase [Oryza sativa (japonica
cultivar-group)]
Length = 809
Score = 600 bits (1546), Expect = e-170
Identities = 357/840 (42%), Positives = 492/840 (58%), Gaps = 69/840 (8%)
Query: 5 FLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGIR 64
FL+ + T L S T ++ N Q T++SA F LGFF+P S+ RYVGI
Sbjct: 19 FLILLVLSTCCLSSTITTDSLLPNKQISDGQTIVSANETFTLGFFSPGTSTY--RYVGIW 76
Query: 65 YHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMTV 124
Y + +TVVWVANR+NP+ D+ G GNL +LD G SF + S +K+
Sbjct: 77 YSNVPNRTVVWVANRNNPVLDTSGILMFDTSGNLVILDGRGSSF---TVAYGSGAKDTEA 133
Query: 125 KLLDSGNLIV-SDDKVKKILWQSFANPTDTFLPGMKMD----DSITLTSWRSHDDPAPGN 179
+LDSGNL++ S ++ WQSF PTDT+L GM + + LTSWRS DDPA G+
Sbjct: 134 TILDSGNLVLRSVSNRSRLRWQSFDYPTDTWLQGMNLGFVGAQNQLLTSWRSSDDPAIGD 193
Query: 180 FSFEQDQGEN-QFVIWKRSMKYWKSSV-SGKFVGTGEMSSAISYLLSNFTLRISPNNTIP 237
+SF D E F IW+R YWKS + +G+ E S +SN T+
Sbjct: 194 YSFGMDPNEKGDFFIWERGNVYWKSGLWNGQSYNFTESESMSFLYVSN-----DARTTLS 248
Query: 238 FLTSSLYSNTRLVMTYWGQLQYL-KMDS-MKMWLMVWVEPRDRCSVFNACGNFGSC--NS 293
+ + R V+ + GQL+ L +MD + WL++ P C ++ CG FG C N
Sbjct: 249 YSSIPASGMVRYVLDHSGQLKLLERMDFVLHQWLVLGSWPEGSCKAYSPCGAFGICAGNQ 308
Query: 294 KYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSLRMMKV-GNPDAQ 352
+ + CKC GF P WS+GD GC R+TN+ D F + M + GN
Sbjct: 309 DWQNRCKCPKGFNPGDGVGWSSGDTRRGCIRQTNM---HCVGDKFFQMPDMGLPGNATTI 365
Query: 353 FNAKNEEECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLEEEYEGGC 412
+ +++C+ CL NC C AY+ + + C +W ++ NL E G
Sbjct: 366 SSITGQKQCESTCLTNCSCTAYAVLQDK-------------CSLWYGNIMNLREGESGDA 412
Query: 413 --DLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRRQAKIRG 470
++R+A S++E + +I T ++V FLI S +++ R++ +K +G
Sbjct: 413 VGTFYLRLAASELESRG-------TPVVLIAATVSSVAFLIFASLIFLWMWRQK-SKAKG 464
Query: 471 IKLYGSERYVRDLIESGRLQEDDAKAIDIPHFHLESILDATNNFAIANKLGQGGFGPVYK 530
+ D + +L E + F I DAT F++ NKLG+GGFGPVYK
Sbjct: 465 V----------DTDSAIKLWESEETGSHFTSFCFSEIADATCKFSLENKLGEGGFGPVYK 514
Query: 531 GKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYM 590
G P GQEIAVKRL++ SGQGL EFKNE++LIA+LQHRNLVRLLG C++G+EK+L+YEYM
Sbjct: 515 GNLPEGQEIAVKRLAAHSGQGLLEFKNEIMLIAKLQHRNLVRLLGCCIQGEEKILIYEYM 574
Query: 591 PNRSLDAFIFDWDM----RFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPK 646
PN+SLD F+F + II GIA+GLLYLH+ SR RIIHRDLKASNILLD + NPK
Sbjct: 575 PNKSLDFFLFAGQVIQCGLEGIIEGIAQGLLYLHKHSRFRIIHRDLKASNILLDIDMNPK 634
Query: 647 ISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKR 706
ISDFG+ARIFG K+T NT+RVVGTYGYM+PEYA++G FSVKSDVFSFGV++LEI+SG R
Sbjct: 635 ISDFGMARIFGSKETEANTNRVVGTYGYMAPEYAMEGIFSVKSDVFSFGVLLLEIVSGIR 694
Query: 707 NTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIE 766
N GF+Q + L+LL YAW LWK GR + D ++ C + ++C++VGL+C+QE PI
Sbjct: 695 NAGFHQRGNSLNLLCYAWELWKEGRWSELADPSIYNACPEHKVLRCIHVGLMCVQESPIN 754
Query: 767 RPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSSKME--TFSRNDLTVTLENGR 824
RPTM+ ++ L +ES TLP P++PAFV S+ T + + T S N +T++ GR
Sbjct: 755 RPTMTEIISALDNESTTLPEPKQPAFV-----SAGIWTEAGVHGGTHSINGMTISDTQGR 809
>ref|NP_916409.1| putative receptor kinase [Oryza sativa (japonica cultivar-group)]
Length = 835
Score = 599 bits (1544), Expect = e-169
Identities = 352/851 (41%), Positives = 503/851 (58%), Gaps = 55/851 (6%)
Query: 7 LCSFIFTFNL-------CSAKDTITINNNLQDGGEDTLISAG-GYFELGFFTPNGSSNGR 58
LC ++ F + C A+DT+ L +TL+S G F LGFFTP G+++
Sbjct: 7 LCCYLLLFVVVVVLTGSCRARDTVVPGRPL--AANETLVSGGDANFVLGFFTPPGANS-- 62
Query: 59 RYVGIRYHKLAPQTVVWVANRDNPLP-----DSGGAFSITEDGNLRVLDRTGKSFWGTNL 113
YVG+ Y+K++ +TVVWVANR++PLP + S++ G L ++ W
Sbjct: 63 TYVGVWYNKVSVRTVVWVANREDPLPGDVADNPDATLSVSPTGTLAIVAGNSTVVWSVTP 122
Query: 114 ERTSSSKNMTVKLLDSGNLIVSDDKVKKILWQSFANPTDTFLPGMKMDDSI------TLT 167
+S T +++DSGNL+++D + WQ F PTDT LP M++ TLT
Sbjct: 123 AAKLASP--TARIMDSGNLVIADGAGGGVAWQGFDYPTDTLLPEMRLGVDYVKGRNRTLT 180
Query: 168 SWRSHDDPAPGNFSFEQD-QGENQFVIWKRSMKYWKSSVSGKFVGTGEMSSAISYLLSNF 226
+W+S DP+PG D G+ Q IW + K W+S TG + ++Y S F
Sbjct: 181 AWKSPSDPSPGPVVMAMDTSGDPQVFIWNGAEKVWRSGPWDGVQFTG-VPDTVTY--SGF 237
Query: 227 TLRISPNN---TIPFLTSSLYSNTRLVMTYWGQLQYLK----MDSMKMWLMVWVEPRDRC 279
T N T F ++ +RL + G L+ +++ W + W P+D+C
Sbjct: 238 TFSFINNAKEVTYSFQVHNVSIISRLGLNSTGSYGLLQRSTWVEAAGTWNLYWYAPKDQC 297
Query: 280 SVFNACGNFGSCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFL 339
+ CG G C++ +C CL GF P S E W+ D GC R T + ++ +D F+
Sbjct: 298 DEVSPCGANGVCDTNNLPVCSCLRGFTPKSPEAWALRDGRAGCVRSTPLDCQNG-TDGFV 356
Query: 340 SLRMMKVGNPDAQFNAKNE----EECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICW 395
++ KV PD + + + E+C+ CL NC C AY+ +G C
Sbjct: 357 AVEHAKV--PDTERSVVDLGLSLEQCRKACLMNCSCTAYASANVSGGGRGHGAGTG--CV 412
Query: 396 IWSQDLNNLEEEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSS 455
+W+ L +L E G DL VR+A +D+ S + + I+V+ ++V FL +L+
Sbjct: 413 MWTTGLTDLRVYPEFGQDLFVRLAAADLGLTSKSNKARVI--IAIVVSISSVTFLSVLAG 470
Query: 456 TCIYLRRRRQAKIRGI-KLYGSERYVRDLIESGRLQEDDAKAIDIPHFHLESILDATNNF 514
++ R++++A+ G K G R E +DD +++P F L +I AT+ F
Sbjct: 471 FLVWTRKKKRARKTGSSKWSGGSRSTGRRYEGSSHHDDD---LELPIFDLGTIAAATDGF 527
Query: 515 AIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLL 574
+I NKLG+GGFGPVYKGK GQEIAVK LS S QGL+EFKNEV+LIA+LQHRNLVRLL
Sbjct: 528 SINNKLGEGGFGPVYKGKLEDGQEIAVKTLSKTSVQGLDEFKNEVMLIAKLQHRNLVRLL 587
Query: 575 GYCVEGDEKMLVYEYMPNRSLDAFIFDWDMRFKIILGIARGLLYLHEDSRLRIIHRDLKA 634
G+ + G E++LVYEYM N+SLD F+F R++II GI RGLLYLH+DSR RIIHRDLKA
Sbjct: 588 GFSISGQERILVYEYMANKSLDYFLF---ARYRIIEGITRGLLYLHQDSRYRIIHRDLKA 644
Query: 635 SNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSF 694
SN+LLD+E PKISDFG+AR+FG ++T NT +VVGTYGYMSPEYA+DG FSVKSDVFSF
Sbjct: 645 SNVLLDKEMTPKISDFGMARMFGSEETEINTRKVVGTYGYMSPEYAMDGVFSVKSDVFSF 704
Query: 695 GVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVN 754
GV++LEIISG+RN G Y + L+LLG+AW LW G++L+ D+T++ + ++E +KC+
Sbjct: 705 GVLLLEIISGRRNRGVYSYSNHLNLLGHAWSLWNEGKSLELADETMNGSFDSDEVLKCIR 764
Query: 755 VGLLCLQEDPIERPTMSNVLFMLG-SESNTLPSPREPAFVIRRCPSSRASTSSKMETFSR 813
VGLLC+QE+P +RP MS VL ML +++ TLP+P++P F RR ++SSK +
Sbjct: 765 VGLLCVQENPDDRPLMSQVLLMLATTDATTLPTPKQPGFAARRILMETDTSSSKPDCSIF 824
Query: 814 NDLTVTLENGR 824
+ TVT+ GR
Sbjct: 825 DSATVTILEGR 835
>dbj|BAD53292.1| putative serine/threonine kinase [Oryza sativa (japonica
cultivar-group)]
Length = 827
Score = 598 bits (1543), Expect = e-169
Identities = 364/842 (43%), Positives = 493/842 (58%), Gaps = 72/842 (8%)
Query: 17 CSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGIRYHKLAPQTVVWV 76
C D+I+ N L DG T++S F LGFF+P SS+ RYVGI Y +T+VWV
Sbjct: 24 CLGTDSISANETLPDG--QTIVSMKNVFVLGFFSPGASSH--RYVGIWYSNPVNRTIVWV 79
Query: 77 ANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMTVKLLDSGNLIVSD 136
ANR+ PL D+ G +GNL V+ G+S + +K+M +LDSGNL +S
Sbjct: 80 ANRNEPLLDASGVLMFDVNGNL-VIAHGGRSLI---VAYGQGTKDMKATILDSGNLALSS 135
Query: 137 -DKVKKILWQSFANPTDTFLPGMKMD---DSITLTSWRSHDDPAPGNFSFEQDQ------ 186
+ +WQSF +PTDT+LP MK+ + TL SW S DDPA G++ D
Sbjct: 136 MANPSRYIWQSFDSPTDTWLPEMKIGLRTTNQTLISWSSIDDPAMGDYKLGMDPAGLSHP 195
Query: 187 -GENQFVIWKRSMKYWKSSVSGKFVGTGEMSSAISYLLSNFTLRI--SPNNTIPFLTSSL 243
G +QF++W R +W SG + +G+M S I L T+ I NN+ +T +
Sbjct: 196 AGLSQFIVWWRGNNFW---TSGHW--SGDMFSLIPELKFFTTIPIFFKCNNSTNDITCTY 250
Query: 244 YSN-----TRLVMTYWGQLQYLKMDSM-KMWLMVWVEPRDRCSVFNACGNFGSCNSKYDS 297
+N T++V+ G L ++ DS+ K W+++W +P C V N CG FG CN D+
Sbjct: 251 SANPSDRMTKIVLNSTGSLSIMQFDSLEKSWILLWRQP-STCEVHNLCGAFGICNDN-DA 308
Query: 298 M--CKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSLRMMKVGNPDAQFNA 355
+ C C GF P + ++ G GC+R+T + SD F + +++ + +
Sbjct: 309 VPKCYCTKGFVPQDIIAYTNGYTREGCNRQTKL---QCSSDEFFEIPNVRLPDNRKKLPV 365
Query: 356 KNEEECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLEEEYE--GGCD 413
ECKL CL NC C AY+Y + + C +W DL NL++ Y+ G
Sbjct: 366 MGLSECKLACLMNCSCTAYAYLQLDG------------CSLWYGDLMNLQDGYDVHGAGT 413
Query: 414 LHVRVAFSDIEGDSYQTERNLSSPTIIL---VTFTAVVFLIILSSTCIYLRRRRQAKIRG 470
L +R+A S++E RN S +L VV L S + + RRR Q K +
Sbjct: 414 LCLRLAASEVESG-----RNSGSGHKMLWMACVIPPVVVLSFCSLSFVLWRRRSQNKGKE 468
Query: 471 IKLYGSERYVRDLIESGRLQEDDAKAIDIPHFHLESILDATNNFAIANKLGQGGFGPVYK 530
D + +L E + F I ++TNNF+ NKLG+GGFGPVYK
Sbjct: 469 NLHAHHSLMTLDTDSAVKLWESEEAGSQFVLFSFSQIANSTNNFSAQNKLGEGGFGPVYK 528
Query: 531 GKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYM 590
G P Q+IAVKRL++ SGQGL EFKNEV+LIA+LQH NLVRLLG C++G+EK+L+YEYM
Sbjct: 529 GNLPDRQDIAVKRLATNSGQGLVEFKNEVLLIAKLQHVNLVRLLGCCIQGEEKILIYEYM 588
Query: 591 PNRSLDAFIF--------DWDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEE 642
PN+SLD F+F DW R II GIA GLLYLH+ SRLRIIHRDLKASNILLD +
Sbjct: 589 PNKSLDFFLFEKSRSVVLDWRKRIHIIEGIAHGLLYLHKHSRLRIIHRDLKASNILLDID 648
Query: 643 KNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEII 702
NPKISDFGLARIFG K+T NT+RVVGTYGYM+PEYA+ G FSVKSDVFSFGV++LEI+
Sbjct: 649 MNPKISDFGLARIFGSKETQANTNRVVGTYGYMAPEYAMQGIFSVKSDVFSFGVLLLEIV 708
Query: 703 SGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQE 762
SG RN G ++ L+LLG+AW LW+ GR D +D + ++CV+VGL+C+QE
Sbjct: 709 SGMRNAGSHRRGRSLNLLGHAWELWREGRWFDLVDPSTRDAYPEHRVLRCVHVGLMCVQE 768
Query: 763 DPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSSKMETFSRNDLTVTLEN 822
+ ++RPTMS+V+ ML SES TLP PR+PAF+ P A + +FS+N +T+T
Sbjct: 769 NAVDRPTMSDVISMLTSESITLPDPRQPAFLSIVLP---AEMDAHDGSFSQNAMTITDLE 825
Query: 823 GR 824
GR
Sbjct: 826 GR 827
>ref|XP_478649.1| putative S-receptor kinase KIK1 precursor [Oryza sativa (japonica
cultivar-group)] gi|28971965|dbj|BAC65366.1| putative
S-receptor kinase KIK1 precursor [Oryza sativa (japonica
cultivar-group)] gi|50510068|dbj|BAD30706.1| putative
S-receptor kinase KIK1 precursor [Oryza sativa (japonica
cultivar-group)]
Length = 865
Score = 598 bits (1541), Expect = e-169
Identities = 359/867 (41%), Positives = 508/867 (58%), Gaps = 90/867 (10%)
Query: 19 AKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGIRYHKLAPQTVVWVAN 78
A DT++ +L G D L+SA G F++GFFTP G G+ Y+G+ Y QTV+WVAN
Sbjct: 28 AADTLSQGQSL--GANDMLVSANGTFKVGFFTPAGGDPGKVYLGVMYATSNVQTVMWVAN 85
Query: 79 RDNPLPDSGGAFSITEDGNLRVLDRTG-KSFWGTNLERTSSSKNMTVKLLDSGNLIVS-- 135
RD P+ + GA S T G+ +L + G + W TN SK+ T+ + D GNL++S
Sbjct: 86 RDAPVRTAAGAASATVTGSGELLVKEGDRVAWRTNASAAGRSKH-TLTIRDDGNLVISGS 144
Query: 136 DDKVKKILWQSFANPTDTFLPGMKM------DDSITLTSWRSHDDPAPGNFSFEQDQGEN 189
D + W+SF +PTDTF+PGM++ D TSWRS DPA G+F+ D
Sbjct: 145 DAAGTDVEWESFHHPTDTFVPGMEIALRQTNGDRTLYTSWRSDADPATGDFTLGLDASA- 203
Query: 190 QFVIWK----RSMKYWKSS--VSGKFVGTGEMSSAI-SYLLSNFTLRISPNNTIPF--LT 240
Q IW+ ++ YW+S SG FVG + + + L+ I+ + +I F
Sbjct: 204 QLYIWRSQGGKNSTYWRSGQWASGNFVGIPWRALYVYGFKLNGDPPPIAGDMSIAFTPFN 263
Query: 241 SSLYSNTRLVMTYWGQLQYLKMDSMKMWLMVWVEPRDRCSVFNACGNFGSCNSK-YDSMC 299
SSLY R V+ G + W +VW +P C +N CG+ C + + +C
Sbjct: 264 SSLY---RFVLRPNGVETCYMLLGSGDWELVWSQPTIPCHRYNLCGDNAECTADDNEPIC 320
Query: 300 KCLPGFRPNSVENWSAGDFSGGCSRKTNV-CSEDAKSDT-----------FLSLRMMKVG 347
C GF P S + ++ G+++ GC R + CS + + T F +R +K+
Sbjct: 321 TCFTGFEPKSPQEYNNGNWTQGCVRSVPLTCSSERNNTTAGGAGAGGGDGFTVIRGVKL- 379
Query: 348 NPDAQFNAK---NEEECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNL 404
PD + C+ CL NC C AYSY C W Q+L ++
Sbjct: 380 -PDFAVWGSLVGDANSCEKACLGNCSCGAYSYS-------------TGSCLTWGQELVDI 425
Query: 405 EEEYEGG----CDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYL 460
+ G DL+V+V S ++ S + + T+++V VV +++ S ++
Sbjct: 426 FQFQTGTEGAKYDLYVKVPSSLLDKSSGRWK------TVVVVVVVVVVVVLLASGLLMWK 479
Query: 461 RRRRQAKIRGIKLYGSE----RYVRDL-------IESGRLQEDDAKAIDIPHFHLESILD 509
RRR + GI ++ R RD +S + ++ K ++P F E++
Sbjct: 480 CRRRIKEKLGIGRKKAQLPLLRPARDAKQDFSGPAQSEHEKSEEGKNCELPLFAFETLAT 539
Query: 510 ATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRN 569
AT+NF+I+NKLG+GGFG VYKG+ PGG+EIAVKRLS SGQGLEEFKNEV+LIA+LQHRN
Sbjct: 540 ATDNFSISNKLGEGGFGHVYKGRLPGGEEIAVKRLSRSSGQGLEEFKNEVILIAKLQHRN 599
Query: 570 LVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHE 621
LVRLLG C++G+EK+LVYEYMPN+SLDAF+FD W RF+II G+ARGLLYLH
Sbjct: 600 LVRLLGCCIQGEEKILVYEYMPNKSLDAFLFDPERRGLLDWRTRFQIIEGVARGLLYLHR 659
Query: 622 DSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYAL 681
DSRLR++HRDLKASNILLD + NPKISDFG+ARIFGG NT+RVVGT GYMSPEYA+
Sbjct: 660 DSRLRVVHRDLKASNILLDRDMNPKISDFGMARIFGGDQNQVNTNRVVGTLGYMSPEYAM 719
Query: 682 DGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLS 741
+G FSV+SDV+SFG+++LEII+G++N+ F+ E L+++GYAW LW R + ID +
Sbjct: 720 EGLFSVRSDVYSFGILILEIITGQKNSSFHHMEGSLNIVGYAWQLWNGDRGQELIDPAIR 779
Query: 742 QTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSR 801
TC A+E ++CV++ LLC+Q+ +RP + V+ LGS+S+ LP+PR P F + +C SS
Sbjct: 780 GTCPAKEALRCVHMALLCVQDHAHDRPDIPYVVLTLGSDSSVLPTPRPPTFTL-QCTSSS 838
Query: 802 ASTS----SKMETFSRNDLTVTLENGR 824
+ K E++S NDLTVT+ GR
Sbjct: 839 SGRDMYYRDKEESYSANDLTVTMLQGR 865
>emb|CAB89179.1| S-locus receptor kinase [Brassica napus var. napus]
gi|322652|pir||JQ1677 S-receptor kinase (EC 2.7.1.-)
precursor - rape gi|167181|gb|AAA33008.1|
serine/threonine kinase receptor
Length = 858
Score = 596 bits (1537), Expect = e-169
Identities = 345/834 (41%), Positives = 489/834 (58%), Gaps = 72/834 (8%)
Query: 15 NLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGIRYHKLAPQTVV 74
N S+ +++TI+NN TL+S G FELGFF SS R Y+GI Y L +T V
Sbjct: 35 NTLSSTESLTISNNR------TLVSPGNVFELGFFRTTSSS--RWYLGIWYKNLPYKTYV 86
Query: 75 WVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMTVKLLDSGNLIV 134
WVANRDNPL DS G I+ + NL +LD + KS W TNL R + + +LL++GN ++
Sbjct: 87 WVANRDNPLSDSIGTLKIS-NMNLVLLDHSNKSVWSTNLTRGNERSPVVAELLENGNFVI 145
Query: 135 ---SDDKVKKILWQSFANPTDTFLPGMKMDDSIT------LTSWRSHDDPAPGNFSFEQD 185
+++ LWQSF PTDT LP MK+ LT+WR+ DDP+ G S++ D
Sbjct: 146 RYSNNNNASGFLWQSFDFPTDTLLPEMKLGYDRKKGLNRFLTAWRNSDDPSSGEISYQLD 205
Query: 186 --QGENQFVIWKRSMKYWKSSVSG--KFVGTGEMSSAISYLLSNFTLRISPNN-TIPFLT 240
+G +F + K ++ ++S +F G E +SY++ NFT T
Sbjct: 206 TQRGMPEFYLLKNGVRGYRSGPWNGVRFNGIPE-DQKLSYMVYNFTDNSEEAAYTFRMTD 264
Query: 241 SSLYSNTRLVMTYWGQLQYLKMDSMKM-WLMVWVEPRD-RCSVFNACGNFGSCNSKYDSM 298
S+YS RL+++ L L W + W P + C V+ CG++ C+ +
Sbjct: 265 KSIYS--RLIISNDEYLARLTFTPTSWEWNLFWTSPEEPECDVYKTCGSYAYCDVNTSPV 322
Query: 299 CKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSLRMMKVGNPDAQFNAKN- 357
C C+ GF+P +++ W ++GGC R+T + D F ++ MK+ ++
Sbjct: 323 CNCIQGFKPFNMQQWELRVWAGGCIRRTRL---SCNGDGFTRMKNMKLPETTMAIVDRSI 379
Query: 358 -EEECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLEEEYEGGCDLHV 416
+ECK CL++C C A++ + G C IW+ +L ++ ++ G DL+V
Sbjct: 380 GRKECKKRCLSDCNCTAFANADIRNGGSG--------CVIWTGELEDIRNYFDDGQDLYV 431
Query: 417 RVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRRQAKIRGIKLYGS 476
R+A +D+ +RN + TI L+ V+ L+I+ C++ R++++AK +
Sbjct: 432 RLAAADLV-----KKRNANGKTIALIVGVCVLLLMIMF--CLWKRKQKRAKTTATSIVNR 484
Query: 477 ERYVRDLIESGRLQ--------EDDAKAIDIPHFHLESILDATNNFAIANKLGQGGFGPV 528
+R +DL+ +G + E+ + +++P LE+++ AT NF+ NKLGQGGFG V
Sbjct: 485 QRN-QDLLMNGMILSSKRQLPIENKTEELELPLIELEAVVKATENFSNCNKLGQGGFGIV 543
Query: 529 YKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYE 588
YKG+ GQEIAVKRLS S QG EF NEV LIARLQH NLVR+LG C+E DEKMLVYE
Sbjct: 544 YKGRLLDGQEIAVKRLSKTSVQGTGEFMNEVRLIARLQHINLVRILGCCIEADEKMLVYE 603
Query: 589 YMPNRSLDAFIF--------DWDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLD 640
Y+ N SLD+++F +W RF I G+ARGLLYLH+DSR RIIHRD+K SNILLD
Sbjct: 604 YLENLSLDSYLFGNKRSSTLNWKDRFNITNGVARGLLYLHQDSRFRIIHRDMKVSNILLD 663
Query: 641 EEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLE 700
+ PKISDFG+ARIF +T NT +VVGTYGYMSPEYA+DG FS KSDVFSFGV+VLE
Sbjct: 664 KNMTPKISDFGMARIFARDETEANTRKVVGTYGYMSPEYAMDGVFSEKSDVFSFGVIVLE 723
Query: 701 IISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFID-------QTLSQTCKAEECMKCV 753
I+SGKRN GFY HE +LL Y W W GR L+ +D +L T + +E +KC+
Sbjct: 724 IVSGKRNRGFYNLNHENNLLSYVWSHWTEGRALEIVDPVIVDSLSSLPATFQPKEVLKCI 783
Query: 754 NVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSSK 807
+GLLC+QE RPTMS+V++MLGSE+ +P P P + + R P +SS+
Sbjct: 784 QIGLLCVQERAEHRPTMSSVVWMLGSEATEIPQPTPPGYSLGRSPYENNPSSSR 837
>emb|CAA67145.1| receptor-like kinase [Brassica oleracea var. acephala]
gi|7434413|pir||T14470 receptor-like kinase (EC 2.7.1.-)
SFR2 - wild cabbage
Length = 847
Score = 596 bits (1537), Expect = e-169
Identities = 356/864 (41%), Positives = 510/864 (58%), Gaps = 73/864 (8%)
Query: 4 IFLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGI 63
+ L +F F+ N SA +++TI++N T+ S G FELGFF P+ SS R Y+GI
Sbjct: 14 LLLFPAFSFSANTLSATESLTISSN------KTISSPGNIFELGFFKPSSSS--RWYLGI 65
Query: 64 RYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMT 123
Y ++ +T VWVANRD+PL S G I+ D NL V+D + + W TNL ++
Sbjct: 66 WYKAISKRTYVWVANRDHPLSTSTGTLKIS-DSNLVVVDGSDTAVWSTNLTGGGDVRSPV 124
Query: 124 V-KLLDSGNLIVSDDKVKK---ILWQSFANPTDTFLPGMKMDDSIT------LTSWRSHD 173
V +LLD+GN ++ D +LWQSF PTDT LP MK+ + L SW+S D
Sbjct: 125 VAELLDNGNFVLRDSNNNDPDIVLWQSFDFPTDTLLPEMKLGWDLKTGFNWFLRSWKSPD 184
Query: 174 DPAPGNFSFE-QDQGENQFVIWKRSMKYWKSSVSG--KFVGTGEMSSAISYLLSNFTLRI 230
DP+ G++SF+ + +G + +W ++ + ++S +F G EM Y+ NFT
Sbjct: 185 DPSSGDYSFKLKTRGFPEAFLWNKASQVYRSGPWNGIRFSGVPEMQP-FDYIEFNFTTS- 242
Query: 231 SPNNTIPFLTSSLYSNTRLVMTYWGQLQ-YLKMDSMKMWLMVWVEPRDRCSVFNACGNFG 289
+ T F + +RL ++ G LQ + +++++ W W P+D+C + CG +G
Sbjct: 243 NQEVTYSFHITKDNMYSRLSLSSTGSLQRFTWIEAIQNWNQFWYAPKDQCDDYKECGTYG 302
Query: 290 SCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSLRMMKVGNP 349
C+S +C C+ GF P + + W D S GC RKT + D F+ L+ MK+ P
Sbjct: 303 YCDSNTYPVCNCMRGFEPRNPQAWGLRDGSDGCVRKTALSCNGG--DGFVRLKKMKL--P 358
Query: 350 DAQFNAKNE----EECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLE 405
D + + +EC+ +C ++C C A++ + G C +W+ D+ +
Sbjct: 359 DTAATSVDRGIGIKECEEKCKSDCNCTAFANTDIRGGGSG--------CVVWTGDILDTR 410
Query: 406 EEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRRQ 465
+GG DL+VR+A +D+E T RN I+ + V L++L +R+Q
Sbjct: 411 NYAKGGQDLYVRLAATDLEDT---TNRNAK----IIGSCIGVSVLLLLCFIFYRFWKRKQ 463
Query: 466 AKIRGIKL-YGSERYVRDLIESGRL---------QEDDAKAIDIPHFHLESILDATNNFA 515
+ I+ + + +DL+ + + +E+ +++P E++ AT+NF+
Sbjct: 464 KRSIAIETSFVDQVRSQDLLMNEVVIPPNRRHISRENKTDDLELPLMDFEAVAIATDNFS 523
Query: 516 IANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLG 575
ANKLGQGGFG VYKG+ GQEIAVKRLS S QG +EFKNEV LIARLQH NLVRLLG
Sbjct: 524 NANKLGQGGFGIVYKGRLLDGQEIAVKRLSKMSVQGTDEFKNEVKLIARLQHINLVRLLG 583
Query: 576 YCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRI 627
CV+ EKML+YEY+ N SLD+ +FD W RF I GIARGLLYLH+DSR RI
Sbjct: 584 CCVDEGEKMLIYEYLENLSLDSHLFDKTRSCKLNWQKRFDITNGIARGLLYLHQDSRFRI 643
Query: 628 IHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSV 687
IHRDLKASN+LLD++ PKISDFG+ARIFG +T NT +VVGTYGYMSPEYA+DG FS
Sbjct: 644 IHRDLKASNVLLDKDMTPKISDFGMARIFGRDETEANTRKVVGTYGYMSPEYAMDGIFST 703
Query: 688 KSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTL----SQT 743
KSDVFSFGV++LEIISGKRN GFY +H+L+LLG W WK G+ LD +D + T
Sbjct: 704 KSDVFSFGVLLLEIISGKRNKGFYNSDHDLNLLGCVWRNWKKGKGLDIVDPIILDSSPST 763
Query: 744 CKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRAS 803
+ E ++C+ +GLLC+QE +RPTMS+V+ MLGSE+ +P P +P + + R P S
Sbjct: 764 YRPLEILRCIKIGLLCVQERANDRPTMSSVVMMLGSETAAIPQPEQPGYCVGRSPLDTDS 823
Query: 804 TSSKM---ETFSRNDLTVTLENGR 824
+SS E++S N +TV++ + R
Sbjct: 824 SSSNQRHDESWSVNQMTVSVIDPR 847
>emb|CAA73133.1| serine /threonine kinase [Brassica oleracea]
Length = 847
Score = 596 bits (1536), Expect = e-168
Identities = 356/864 (41%), Positives = 510/864 (58%), Gaps = 73/864 (8%)
Query: 4 IFLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGI 63
+ L +F F+ N SA +++TI++N T+ S G FELGFF P+ SS R Y+GI
Sbjct: 14 LLLFPAFSFSSNTLSATESLTISSN------KTISSPGNIFELGFFKPSSSS--RWYLGI 65
Query: 64 RYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMT 123
Y ++ +T VWVANRD+PL S G I+ D NL V+D + + W TNL ++
Sbjct: 66 WYKAISKRTYVWVANRDHPLSTSTGTLKIS-DSNLVVVDGSDTAVWSTNLTGGGDVRSPV 124
Query: 124 V-KLLDSGNLIVSDDKVKK---ILWQSFANPTDTFLPGMKMDDSIT------LTSWRSHD 173
V +LLD+GN ++ D +LWQSF PTDT LP MK+ + L SW+S D
Sbjct: 125 VAELLDNGNFVLRDSNNNDPDIVLWQSFDFPTDTLLPEMKLGWDLKTGFNWFLRSWKSPD 184
Query: 174 DPAPGNFSFE-QDQGENQFVIWKRSMKYWKSSVSG--KFVGTGEMSSAISYLLSNFTLRI 230
DP+ G++SF+ + +G + +W ++ + ++S +F G EM Y+ NFT
Sbjct: 185 DPSSGDYSFKLKTRGFPEAFLWNKASQVYRSGPWNGIRFSGVPEMQP-FDYIEFNFTTS- 242
Query: 231 SPNNTIPFLTSSLYSNTRLVMTYWGQLQ-YLKMDSMKMWLMVWVEPRDRCSVFNACGNFG 289
+ T F + +RL ++ G LQ + +++++ W W P+D+C + CG +G
Sbjct: 243 NQEVTYSFHITKDNMYSRLSLSSTGSLQRFTWIEAIQNWNQFWYAPKDQCDDYKECGTYG 302
Query: 290 SCNSKYDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSLRMMKVGNP 349
C+S +C C+ GF P + + W D S GC RKT + D F+ L+ MK+ P
Sbjct: 303 YCDSNTYPVCNCMRGFEPRNPQAWGLRDGSDGCVRKTALSCNGG--DGFVRLKKMKL--P 358
Query: 350 DAQFNAKNE----EECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLE 405
D + + +EC+ +C ++C C A++ + G C +W+ D+ +
Sbjct: 359 DTAATSVDRGIGIKECEEKCKSDCNCTAFANTDIRGGGSG--------CVVWTGDILDTR 410
Query: 406 EEYEGGCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRRQ 465
+GG DL+VR+A +D+E T RN I+ + V L++L +R+Q
Sbjct: 411 NYAKGGQDLYVRLAATDLEDT---TNRNAK----IIGSCIGVSVLLLLCFIFYRFWKRKQ 463
Query: 466 AKIRGIKL-YGSERYVRDLIESGRL---------QEDDAKAIDIPHFHLESILDATNNFA 515
+ I+ + + +DL+ + + +E+ +++P E++ AT+NF+
Sbjct: 464 KRSIAIETSFVDQVRSQDLLMNEVVIPPNRRHISRENKTDDLELPLMDFEAVAIATDNFS 523
Query: 516 IANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLG 575
ANKLGQGGFG VYKG+ GQEIAVKRLS S QG +EFKNEV LIARLQH NLVRLLG
Sbjct: 524 NANKLGQGGFGIVYKGRLLDGQEIAVKRLSKMSVQGTDEFKNEVKLIARLQHINLVRLLG 583
Query: 576 YCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRI 627
CV+ EKML+YEY+ N SLD+ +FD W RF I GIARGLLYLH+DSR RI
Sbjct: 584 CCVDEGEKMLIYEYLENLSLDSHLFDKTRSCKLNWQKRFVITNGIARGLLYLHQDSRFRI 643
Query: 628 IHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSV 687
IHRDLKASN+LLD++ PKISDFG+ARIFG +T NT +VVGTYGYMSPEYA+DG FS
Sbjct: 644 IHRDLKASNVLLDKDMTPKISDFGMARIFGRDETEANTRKVVGTYGYMSPEYAMDGIFST 703
Query: 688 KSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTL----SQT 743
KSDVFSFGV++LEIISGKRN GFY +H+L+LLG W WK G+ LD +D + T
Sbjct: 704 KSDVFSFGVLLLEIISGKRNKGFYNSDHDLNLLGCVWRNWKKGKGLDIVDPIILDSSPST 763
Query: 744 CKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRAS 803
+ E ++C+ +GLLC+QE +RPTMS+V+ MLGSE+ +P P +P + + R P S
Sbjct: 764 YRPLEILRCIKIGLLCVQERANDRPTMSSVVMMLGSETAAIPQPEQPGYCVGRSPLDTDS 823
Query: 804 TSSKM---ETFSRNDLTVTLENGR 824
+SS E++S N +TV++ + R
Sbjct: 824 SSSNQRHDESWSVNQMTVSVIDPR 847
>ref|NP_176755.1| S-receptor protein kinase, putative [Arabidopsis thaliana]
gi|2129703|pir||S70769 S-receptor kinase (EC 2.7.1.-)
Ark1 precursor - Arabidopsis thaliana
gi|166692|gb|AAA32786.1| receptor kinase
gi|445123|prf||1908429A receptor kinase
Length = 843
Score = 595 bits (1535), Expect = e-168
Identities = 350/857 (40%), Positives = 496/857 (57%), Gaps = 66/857 (7%)
Query: 4 IFLLCSFIFTFNLCSAKDTITINNNLQDGGEDTLISAGGYFELGFFTPNGSSNGRRYVGI 63
+ L +F + N SA +++TI++N T+IS FELGFF P SS R Y+GI
Sbjct: 17 LILFLAFSVSPNTLSATESLTISSN------KTIISPSQIFELGFFNPASSS--RWYLGI 68
Query: 64 RYHKLAPQTVVWVANRDNPLPDSGGAFSITEDGNLRVLDRTGKSFWGTNLERTSSSKNMT 123
Y + +T VWVANRDNPL S G I+ + NL + D++ + W TN+ +
Sbjct: 69 WYKIIPIRTYVWVANRDNPLSSSNGTLKISGN-NLVIFDQSDRPVWSTNITGGDVRSPVA 127
Query: 124 VKLLDSGNLIVSDDKVKKILWQSFANPTDTFLPGMKMD-DSIT-----LTSWRSHDDPAP 177
+LLD+GN ++ D ++LWQSF PTDT L MK+ D T L SW++ DDP+
Sbjct: 128 AELLDNGNFLLRDSN-NRLLWQSFDFPTDTLLAEMKLGWDQKTGFNRILRSWKTTDDPSS 186
Query: 178 GNFSFEQDQGE--NQFVIWKRSMKYWKSSVSGKFVGTGEMSSAISYLLSNFTLRISPNNT 235
G FS + + E ++ K S+ Y +G + + + Y++ NFT T
Sbjct: 187 GEFSTKLETSEFPEFYICSKESILYRSGPWNGMRFSSVPGTIQVDYMVYNFTAS-KEEVT 245
Query: 236 IPFLTSSLYSNTRLVMTYWGQLQYLK-MDSMKMWLMVWVEPRDRCSVFNACGNFGSCNSK 294
+ + +RL + G LQ L ++ + W +W P+D C + CGNFG C+S
Sbjct: 246 YSYRINKTNLYSRLYLNSAGLLQRLTWFETTQSWKQLWYSPKDLCDNYKVCGNFGYCDSN 305
Query: 295 YDSMCKCLPGFRPNSVENWSAGDFSGGCSRKTNVCSEDAKSDTFLSLRMMKVGNPDAQFN 354
C C+ GF+P + + W D S GC RKT + + D F L+ MK+ PD
Sbjct: 306 SLPNCYCIKGFKPVNEQAWDLRDGSAGCMRKTRLSCDGR--DGFTRLKRMKL--PDTTAT 361
Query: 355 AKNEE----ECKLECLNNCQCYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLEEEYEG 410
+ E CK CL +C C A++ + G C IW++++ ++ +G
Sbjct: 362 IVDREIGLKVCKERCLEDCNCTAFANADIRNGGSG--------CVIWTREILDMRNYAKG 413
Query: 411 GCDLHVRVAFSDIEGDSYQTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRRQAKIRG 470
G DL+VR+A +++E + E+ + S V L++LS + +R+Q +
Sbjct: 414 GQDLYVRLAAAELEDKRIKNEKIIGSSI-------GVSILLLLSFVIFHFWKRKQKRSIT 466
Query: 471 IK------LYGSERYVRDLIESGR---LQEDDAKAIDIPHFHLESILDATNNFAIANKLG 521
I+ + + + D++ S R +E ++ +++P LE++ ATNNF+ NKLG
Sbjct: 467 IQTPNVDQVRSQDSLINDVVVSRRGYTSKEKKSEYLELPLLELEALATATNNFSNDNKLG 526
Query: 522 QGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGD 581
QGGFG VYKG+ G+EIAVKRLS S QG +EF NEV LIA+LQH NLVRLLG CV+
Sbjct: 527 QGGFGIVYKGRLLDGKEIAVKRLSKMSSQGTDEFMNEVRLIAKLQHINLVRLLGCCVDKG 586
Query: 582 EKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRIIHRDLK 633
EKML+YEY+ N SLD+ +FD W RF II GIARGLLYLH+DSR RIIHRDLK
Sbjct: 587 EKMLIYEYLENLSLDSHLFDQTRSSNLNWQKRFDIINGIARGLLYLHQDSRCRIIHRDLK 646
Query: 634 ASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFS 693
ASN+LLD+ PKISDFG+ARIFG ++T NT RVVGTYGYMSPEYA+DG FS+KSDVFS
Sbjct: 647 ASNVLLDKNMTPKISDFGMARIFGREETEANTRRVVGTYGYMSPEYAMDGIFSMKSDVFS 706
Query: 694 FGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFID----QTLSQTCKAEEC 749
FGV++LEIISGKRN GFY +L+LLG+ W WK G L+ +D +LS E
Sbjct: 707 FGVLLLEIISGKRNKGFYNSNRDLNLLGFVWRHWKEGNELEIVDPINIDSLSSKFPTHEI 766
Query: 750 MKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSS--K 807
++C+ +GLLC+QE +RP MS+V+ MLGSE+ +P P+ P F I R P S+SS +
Sbjct: 767 LRCIQIGLLCVQERAEDRPVMSSVMVMLGSETTAIPQPKRPGFCIGRSPLEADSSSSTQR 826
Query: 808 METFSRNDLTVTLENGR 824
+ + N +T+++ + R
Sbjct: 827 DDECTVNQITLSVIDAR 843
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.137 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,492,298,812
Number of Sequences: 2540612
Number of extensions: 67015725
Number of successful extensions: 193888
Number of sequences better than 10.0: 20609
Number of HSP's better than 10.0 without gapping: 10946
Number of HSP's successfully gapped in prelim test: 9664
Number of HSP's that attempted gapping in prelim test: 143584
Number of HSP's gapped (non-prelim): 25818
length of query: 824
length of database: 863,360,394
effective HSP length: 137
effective length of query: 687
effective length of database: 515,296,550
effective search space: 354008729850
effective search space used: 354008729850
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 80 (35.4 bits)
Lotus: description of TM0203.7


