

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.5
(852 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
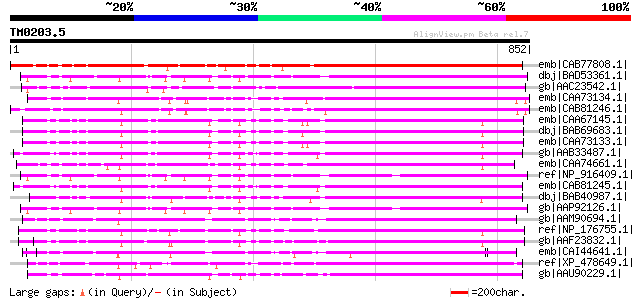
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB77808.1| putative receptor kinase [Arabidopsis thaliana] ... 758 0.0
dbj|BAD53361.1| putative receptor-like protein kinase ARK1 [Oryz... 583 e-164
gb|AAC23542.1| receptor protein kinase [Ipomoea trifida] 579 e-163
emb|CAA73134.1| serine/threonine kinase [Brassica oleracea] gi|7... 578 e-163
emb|CAB81246.1| serine/threonine kinase-like protein [Arabidopsi... 570 e-161
emb|CAA67145.1| receptor-like kinase [Brassica oleracea var. ace... 567 e-160
dbj|BAB69683.1| receptor kinase 5 [Brassica rapa] 565 e-159
emb|CAA73133.1| serine /threonine kinase [Brassica oleracea] 564 e-159
gb|AAB33487.1| ARK3 product/receptor-like serine/threonine prote... 561 e-158
emb|CAA74661.1| SFR1 [Brassica oleracea] gi|7434410|pir||T14519 ... 561 e-158
ref|NP_916409.1| putative receptor kinase [Oryza sativa (japonic... 560 e-158
emb|CAB81245.1| receptor-like serine/threonine protein kinase AR... 559 e-157
dbj|BAB40987.1| SRKb [Arabidopsis lyrata] 559 e-157
gb|AAP92126.1| receptor-like protein kinase ARK1 [Oryza sativa (... 557 e-157
gb|AAM90694.1| S-locus receptor-like kinase RLK14 [Oryza sativa] 557 e-157
ref|NP_176755.1| S-receptor protein kinase, putative [Arabidopsi... 556 e-157
gb|AAF23832.1| F1E22.15 [Arabidopsis thaliana] 556 e-157
emb|CAI44641.1| OSJNBb0015D13.18 [Oryza sativa (japonica cultiva... 555 e-156
ref|XP_478649.1| putative S-receptor kinase KIK1 precursor [Oryz... 553 e-156
gb|AAU90229.1| putative receptor-like protein kinase [Oryza sati... 549 e-154
>emb|CAB77808.1| putative receptor kinase [Arabidopsis thaliana]
gi|15236268|ref|NP_192232.1| S-locus lectin protein
kinase family protein [Arabidopsis thaliana]
gi|4262151|gb|AAD14451.1| putative receptor kinase
[Arabidopsis thaliana] gi|25287707|pir||A85041 probable
receptor kinase [imported] - Arabidopsis thaliana
Length = 852
Score = 758 bits (1956), Expect = 0.0
Identities = 423/869 (48%), Positives = 565/869 (64%), Gaps = 65/869 (7%)
Query: 1 MFFTPFTNMITIFLFHMHCWLLCF-SQLCFAGDTLNVGQEITGNGTVLVSAAKKFELGFF 59
M + F M + + + C++ S+ F G TL +G LVSA ++FELGFF
Sbjct: 1 MILSVFFYMFLLHIRRLDCFVAVQDSKTLFKGSTLINDS----HGETLVSAGQRFELGFF 56
Query: 60 SPDLNVTGGKGRYLGIWYYREEGSGLSP--VVWVANRDNPVADDSIGVFRIADDGNLVVL 117
+P N + + RYLGIW+Y L P VVWVANR++PV D S +F I+ DGNL V+
Sbjct: 57 TP--NGSSDERRYLGIWFYN-----LHPLTVVWVANRESPVLDRSC-IFTISKDGNLEVI 108
Query: 118 DTSGIRYWSASNLTNSSSATNRSVKLMDSGNLVLL-DEHVGMKLWESFEHPTDTFLPGMK 176
D+ G YW + + SS + R VKLMD+GNLVL+ D + +W+SF++PTDTFLPGM+
Sbjct: 109 DSKGRVYWD-TGVKPSSVSAERMVKLMDNGNLVLISDGNEANVVWQSFQNPTDTFLPGMR 167
Query: 177 MDKTLELTCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPD 236
MD+ + L+ W+S +DP GNFTF+MD++ + +F I + YW+S G+ D
Sbjct: 168 MDENMTLSSWRSFNDPSHGNFTFQMDQEEDKQFIIWKRSMRYWKS-----GISGKFIGSD 222
Query: 237 DISNDVYNLLTNFKE---LKNKTV-----SSYDNTRLLLNSTGVIKVLYRVNFQSDIVW- 287
++ + L+NF E + N +V S Y NTR ++S+G + + W
Sbjct: 223 EMPYAISYFLSNFTETVTVHNASVPPLFTSLYTNTRFTMSSSGQAQYF---RLDGERFWA 279
Query: 288 --WYQPRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSPLNDYTVGGDTSSLLCTRKS 345
W +PR C YN CGNF SCN N+++C CLPGF R + V GD S C+R+S
Sbjct: 280 QIWAEPRDECSVYNACGNFGSCNSKNEEMCKCLPGF--RPNFLEKWVKGDFSGG-CSRES 336
Query: 346 TSCGANT----NTFLNLTMMKIGSPDIKVSAQDENECKFRCISMCSQTQCQACSYVPIPV 401
CG + + FLNL+++++GSPD + A +E EC+ C++ C QCQA SY + +
Sbjct: 337 RICGKDGVVVGDMFLNLSVVEVGSPDSQFDAHNEKECRAECLNNC---QCQAYSYEEVDI 393
Query: 402 QQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPT-----RKGNPKSTLS 456
Q + CWIW ++L LKE YLG R +F+RVA DI R G K+ +
Sbjct: 394 LQSN---TKCWIWLEDLNNLKEGYLGS--RNVFIRVAVPDIGSHVERGRGRYGEAKTPVV 448
Query: 457 LILGIALPGVVILACICILA---YVCRRKIALKLKQESESILRQRGRFYDSERHVKDLID 513
LI+ + IL + A ++ RRK+ +L + DSERH+K+LI+
Sbjct: 449 LIIVVTFTSAAILVVLSSTASYVFLQRRKVNKELGSIPRGV-----HLCDSERHIKELIE 503
Query: 514 KEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLS 573
++ D++GI+VP F+ E+IL AT FS+ANKLG+GG+GPVYKG G +EIAVKRLS
Sbjct: 504 SGRFKQDDSQGIDVPSFELETILYATSNFSNANKLGQGGFGPVYKGMFPGDQEIAVKRLS 563
Query: 574 SVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKS 633
S QG++EFKNEVVLIAKLQHRNLVRL GYC+ G+EK+L+YEYMP+KSLD F+FD
Sbjct: 564 RCSGQGLEEFKNEVVLIAKLQHRNLVRLLGYCVAGEEKLLLYEYMPHKSLDFFIFDRKLC 623
Query: 634 ALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFG 693
LDW+MR +I+LGIARGLLYLHQDSRLR+IHRDLKTSNILLD EM PKISDFGLARIFG
Sbjct: 624 QRLDWKMRCNIILGIARGLLYLHQDSRLRIIHRDLKTSNILLDEEMNPKISDFGLARIFG 683
Query: 694 GKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTL 753
G ET ANT RVVGTYGYMSPEYAL+G FS KSD+FSFGVV++E ISGK+NTGF++ + +L
Sbjct: 684 GSETSANTNRVVGTYGYMSPEYALEGLFSFKSDVFSFGVVVIETISGKRNTGFHEPEKSL 743
Query: 754 SLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIML 813
SLLG+AW LW + ++L+D +L E+ F++C +VGLLCVQ++P+DRP MSNVV ML
Sbjct: 744 SLLGHAWDLWKAERGIELLDQALQESCETEGFLKCLNVGLLCVQEDPNDRPTMSNVVFML 803
Query: 814 -DSETATLPTPKQPTFFTRKDLSSTASSS 841
SE ATLPTPKQP F R+ SS+ +SS
Sbjct: 804 GSSEAATLPTPKQPAFVLRRCPSSSKASS 832
>dbj|BAD53361.1| putative receptor-like protein kinase ARK1 [Oryza sativa (japonica
cultivar-group)]
Length = 846
Score = 583 bits (1502), Expect = e-164
Identities = 363/873 (41%), Positives = 496/873 (56%), Gaps = 86/873 (9%)
Query: 19 CWLLCFSQL------CFAGDTLNVGQEITGNGTVLVSAAKKFELGFFSPDLNVTGGKGRY 72
C+LL F + C A DT+ G+ + N T++ F LGFF+P G Y
Sbjct: 9 CYLLLFVVVVVLTGSCRARDTVVPGRPLAANETLVSGGDANFVLGFFTPP----GANSTY 64
Query: 73 LGIWYYREEGSGLSPVVWVANRDNP----VADDSIGVFRIADDGNLVVLDTSGIRYWSAS 128
+G+WY + + VVWVANR++P VAD+ ++ G L ++ + WS +
Sbjct: 65 VGVWYNKVS---VRTVVWVANREDPLPGDVADNPDATLSVSPTGTLAIVAGNSTVVWSVT 121
Query: 129 NLTNSSSATNRSVKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKMD------KTLE 182
+S T R +MDSGNLV+ D G W+ F++PTDT LP M++ +
Sbjct: 122 PAAKLASPTAR---IMDSGNLVIADGAGGGVAWQGFDYPTDTLLPEMRLGVDYVKGRNRT 178
Query: 183 LTCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPDDI--SN 240
LT WKS SDP G MD + + I N + W+S DGV PD + S
Sbjct: 179 LTAWKSPSDPSPGPVVMAMDTSGDPQVFIWNGAEKVWRSGPW-DGVQFT-GVPDTVTYSG 236
Query: 241 DVYNLLTNFKELK------NKTVSSYDNTRLLLNSTGVIKVLYRVNFQSDI----VWWYQ 290
++ + N KE+ N ++ S RL LNSTG +L R + ++WY
Sbjct: 237 FTFSFINNAKEVTYSFQVHNVSIIS----RLGLNSTGSYGLLQRSTWVEAAGTWNLYWYA 292
Query: 291 PRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSP----LNDYTVGGDTSSLLCTRKST 346
P+ C + CG C+ +N +C+CL GF +SP L D G C R +
Sbjct: 293 PKDQCDEVSPCGANGVCDTNNLPVCSCLRGFTPKSPEAWALRDGRAG-------CVRSTP 345
Query: 347 -SCGANTNTFLNLTMMKIGSPDIKVSAQDEN----ECKFRCISMCSQTQCQACSYVPIPV 401
C T+ F+ + K+ PD + S D +C+ C+ CS C A + +
Sbjct: 346 LDCQNGTDGFVAVEHAKV--PDTERSVVDLGLSLEQCRKACLMNCS---CTAYASANVSG 400
Query: 402 QQRGLNLSP-CWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILG 460
RG C +WT LT L+ G D LFVR+A +D+ ++ + +++++
Sbjct: 401 GGRGHGAGTGCVMWTTGLTDLRVYPEFGQD--LFVRLAAADLGLTSKSNKARVIIAIVVS 458
Query: 461 IALPGVVILACIC-ILAYVCRRKIALKLKQESESI-LRQRGRFYDSERHVKDLIDKEGLE 518
I+ V L+ + L + ++K A K S R GR Y+ H D
Sbjct: 459 IS--SVTFLSVLAGFLVWTRKKKRARKTGSSKWSGGSRSTGRRYEGSSHHDD-------- 508
Query: 519 EKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQ 578
+E+P FD +I ATD FS NKLG GG+GPVYKGKL+ G+EIAVK LS S Q
Sbjct: 509 -----DLELPIFDLGTIAAATDGFSINNKLGEGGFGPVYKGKLEDGQEIAVKTLSKTSVQ 563
Query: 579 GIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDW 638
G+ EFKNEV+LIAKLQHRNLVRL G+ I G E+IL+YEYM NKSLD F+F+ + S LLDW
Sbjct: 564 GLDEFKNEVMLIAKLQHRNLVRLLGFSISGQERILVYEYMANKSLDYFLFEKSNSVLLDW 623
Query: 639 QMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETE 698
Q R+ I+ GI RGLLYLHQDSR R+IHRDLK SN+LLD EM PKISDFG+AR+FG +ETE
Sbjct: 624 QARYRIIEGITRGLLYLHQDSRYRIIHRDLKASNVLLDKEMTPKISDFGMARMFGSEETE 683
Query: 699 ANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGY 758
NT++VVGTYGYMSPEYA+DG FS KSD+FSFGV+LLEIISG++N G Y Y L+LLG+
Sbjct: 684 INTRKVVGTYGYMSPEYAMDGVFSVKSDVFSFGVLLLEIISGRRNRGVYSYSNHLNLLGH 743
Query: 759 AWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIML-DSET 817
AW LW E K L+L D ++ ++++++ ++C VGLLCVQ+ PDDRP MS V++ML ++
Sbjct: 744 AWSLWNEGKSLELADETMNGSFDSDEVLKCIRVGLLCVQENPDDRPLMSQVLLMLATTDA 803
Query: 818 ATLPTPKQPTFFTRKDLSSTASSSLQFDSSIVE 850
TLPTPKQP F R+ L T +SS + D SI +
Sbjct: 804 TTLPTPKQPGFAARRILMETDTSSSKPDCSIFD 836
>gb|AAC23542.1| receptor protein kinase [Ipomoea trifida]
Length = 853
Score = 579 bits (1492), Expect = e-163
Identities = 372/864 (43%), Positives = 500/864 (57%), Gaps = 73/864 (8%)
Query: 20 WLLCFSQL-------CFAGDTLNVGQEITGNGTVLVSAAKKFELGFFSPDLNVTGGKGRY 72
W SQ+ A D++ Q + GN T LVS+ FELGFF+P+ G Y
Sbjct: 13 WFFLISQIFIGNLAVALAVDSITPTQPLAGNRT-LVSSDGLFELGFFTPN----GSDQSY 67
Query: 73 LGIWYYREEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTN 132
+GIWY E VVWV NRD + S G+ +I +DGN+ ++D G WS TN
Sbjct: 68 VGIWYKEIEPK---TVVWVGNRDG-ASRGSAGILKIGEDGNIHLVDGGGNFIWSP---TN 120
Query: 133 SSSATNRSVKLMDSGNLVLL---DEHVGMKLWESFEHPTDTFLPGMKM---DKT---LEL 183
S+A N +L+DSGN VL DE+ LW+SF++PTDT LPGMK+ KT +
Sbjct: 121 QSAARNTVAQLLDSGNFVLRREDDENPENYLWQSFDYPTDTLLPGMKLGWDSKTGLNRYI 180
Query: 184 TCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSEEQG----DGVMNPESNPDDIS 239
+ WKSL+DPG G +FK+D + N+ ++ ++S GV PE P
Sbjct: 181 SAWKSLNDPGEGPISFKLDINGLPEIFLRNRDKIVYRSGPWNGVRFSGV--PEMKPTATI 238
Query: 240 NDVYNLLTNFK----ELKNKTVSSYDNTRLLLNSTGVIKVLYRVNFQSDIVW---WYQPR 292
+ + N + EL NKT+ S RLL+ G ++ + + +W WY P+
Sbjct: 239 TFSFVMTKNERYYSFELHNKTLYS----RLLVTRNGNLERYAWI--PTSKIWSKFWYAPK 292
Query: 293 TTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSPLN-DYTVGGDTSSLLCTRKSTSCGAN 351
C +Y CG F C+ + +C CL GF +SP D G D C R
Sbjct: 293 DQCDSYKECGTFGFCDTNMSPVCQCLVGFRPKSPQAWDLRDGSDG----CVRYH-ELECR 347
Query: 352 TNTFLNLTMMKIGSPDIKVSAQDENECKFRCISMCSQTQCQACSYVPIPVQQRGLNLSPC 411
+ FL + MK+ PD S D C+ MC + C +Y + G S C
Sbjct: 348 KDGFLTMNFMKL--PDTSSSFVDTTMNLDECMKMC-KNNCSCTAYTNSNISNGG---SGC 401
Query: 412 WIWTQNLTTLKEEYLGGDDRK---LFVRVAKSDIEEPTRKGNPKSTLSLIL---GIALPG 465
IWT T L + + G R L R A SD+ + G+ I+ GIA+ G
Sbjct: 402 VIWT---TELLDAAVRGGRRWPSCLHPRSA-SDVAQGGDSGDASGRTKRIIIACGIAV-G 456
Query: 466 VVILACICILAYVCRRKIALKLKQESESILRQRGRFYDSERHVKDLIDK-EGLEEKDNEG 524
V IL ++ +R+ + + ++ + R R D + + K E E +
Sbjct: 457 VGILLFALSALFILKRRQSKRALGKNTELRGFRDRSQDLLMNAAVIPSKREYSGETMTDE 516
Query: 525 IEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFK 584
E+P FDF +I+VATD F+D NKLG+GG+G VYKG ++G EIAVKRLS S QG++EFK
Sbjct: 517 FELPLFDFSTIVVATDNFADVNKLGQGGFGCVYKGMVEG-EEIAVKRLSKNSGQGVEEFK 575
Query: 585 NEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDI 644
NE+ LIA+LQHRNLVRL G C+ +EKILIYEYM NKSLD+ +F+ +S+LL+WQ RF+I
Sbjct: 576 NELRLIARLQHRNLVRLLGCCVDMEEKILIYEYMENKSLDSTLFNKQRSSLLNWQTRFNI 635
Query: 645 LLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEA-NTQR 703
+ GIARGLLYLHQDSR R+IHRDLK SNILLD EM PKISDFG+ARIFGG ET+A NT+R
Sbjct: 636 ICGIARGLLYLHQDSRFRIIHRDLKASNILLDKEMNPKISDFGMARIFGGDETDANNTKR 695
Query: 704 VVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLW 763
VVGTYGYMSPEYA+DG FS KSD+FSFGV++LEI++GKKN GFY +LLG+AW+LW
Sbjct: 696 VVGTYGYMSPEYAMDGLFSVKSDVFSFGVLVLEIVTGKKNRGFYNQNNQQNLLGHAWRLW 755
Query: 764 TENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTP 823
E + +L+D ++GE+Y+ + +RC VGLLCVQ++ +DRPNM+ VV+ML SE+ATLP P
Sbjct: 756 RERRGSELLDSAIGESYSLCEVMRCIQVGLLCVQEQAEDRPNMATVVLMLGSESATLPQP 815
Query: 824 KQPTFFTRKDLSSTASSSLQFDSS 847
K P F + SS+ D S
Sbjct: 816 KHPGFCLGSRPADMDSSTSNCDES 839
>emb|CAA73134.1| serine/threonine kinase [Brassica oleracea] gi|7434416|pir||T14450
serine/threonine kinase (EC 2.7.1.-) BRLK - wild cabbage
Length = 850
Score = 578 bits (1490), Expect = e-163
Identities = 364/869 (41%), Positives = 503/869 (56%), Gaps = 89/869 (10%)
Query: 30 AGDTLNVGQEITGNGT--VLVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYREEGSGLSP 87
A DT+ G + T LVS K FELGFFSP GRYLGIWY E
Sbjct: 25 AQDTIRRGGFLRDGSTHKPLVSPQKTFELGFFSPG----SSPGRYLGIWYGNIEDKA--- 77
Query: 88 VVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNSSSATNRSVKLMDSG 147
VVWVANR+NP++D S GV I++DGNLV+L+ I WS SN+T++++ NR ++D+G
Sbjct: 78 VVWVANRENPISDRS-GVLTISNDGNLVLLNGQNITVWS-SNITSTNNDNNRVGSILDTG 135
Query: 148 NLVLLDEHVGMKLWESFEHPTDTFLPGMKM------DKTLELTCWKSLSDPGRGNFTFKM 201
N L++ +WESF HPTDTFLP M++ L W+S +DP GNF+ +
Sbjct: 136 NFELIEVSSERVIWESFNHPTDTFLPHMRVRVNPQTGDNLAFVSWRSENDPSPGNFSLGV 195
Query: 202 DKKWENRFAILNQGQLY-WQSEEQGDGVMNPESNPDDISNDVYNLLTNFKELKNKTVSSY 260
D + + W+S + + N ++N +Y ++T S Y
Sbjct: 196 DPSGAPEIVLWGRNNTRRWRSGQWNSAIFTGIPNMALLTNYLYGF--KLSSPPDETGSVY 253
Query: 261 -----DNTRLLLNSTGVIKVLYRVNFQSDIVW------WYQ----PRTTCLTYNVCGNFS 305
+ +LL KVL+ + ++ W W + P + C YN CG+F
Sbjct: 254 FTYVPSDPSVLLR----FKVLHN-GTEEELRWNETSKRWTKFQAAPESECDKYNRCGSFG 308
Query: 306 SCNDDNDK-LCTCLPGFGRRSPLNDYTVG-GDTSSLLCTRKSTSCGANTNTFLNLTMMKI 363
C+ D +C+C+ G+ S L +++ G + L C R ++ G + FL L +K+
Sbjct: 309 ICDMRGDNGICSCVKGYEPVS-LGNWSRGCRRRTPLRCERNVSNVGEDE--FLTLKSVKL 365
Query: 364 GSPDIKV---SAQDENECKFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIWTQNLTT 420
PD + S D +CK RC+ CS C A ++V N C IW Q+L
Sbjct: 366 --PDFETPEHSLADPEDCKDRCLKNCS---CTAFTFV---------NGIGCMIWNQDLVD 411
Query: 421 LKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGVVILACICILAYVCR 480
L++ GG L VR+A S+I G K T +++ L GV++L +L + +
Sbjct: 412 LQQFEAGGSS--LHVRLADSEI------GESKKTKIVVIVAVLVGVLLLGIFALLLWRFK 463
Query: 481 RKIALK-----LKQESESILRQRGRFYDSERHVKDLIDKEGLEEKDNEGIEVPYFDFESI 535
RK + ++ ++ + D+ +D +E K E+P F + I
Sbjct: 464 RKKDVSGTYCGHDADTSVVVVDMTKAKDTTTAFTGSVDIM-IEGKAVNTSELPVFCLKVI 522
Query: 536 LVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQH 595
+ AT+ FS N+LGRGG+GPVYKG L+ G+EIAVKRLS S QG+ EFKNE++LIAKLQH
Sbjct: 523 VKATNDFSRENELGRGGFGPVYKGVLEDGQEIAVKRLSGKSGQGVDEFKNEIILIAKLQH 582
Query: 596 RNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYL 655
RNLVRL G C +G+EK+L+YEYMPNKSLD F+FD K L+DW++RF I+ GIARGLLYL
Sbjct: 583 RNLVRLLGCCFEGEEKMLVYEYMPNKSLDFFIFDEMKQELVDWKLRFAIIEGIARGLLYL 642
Query: 656 HQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEY 715
H+DSRLR+IHRDLK SN+LLDGEM PKISDFG+ARIFGG + EANT RVVGTYGYMSPEY
Sbjct: 643 HRDSRLRIIHRDLKVSNVLLDGEMNPKISDFGMARIFGGNQNEANTVRVVGTYGYMSPEY 702
Query: 716 ALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLS 775
A++G FS KSD++SFGV+LLEIISGK+NT + SL+GYAW L+T + +L+D
Sbjct: 703 AMEGLFSVKSDVYSFGVLLLEIISGKRNTSLRASEHG-SLIGYAWFLYTHGRSEELVDPK 761
Query: 776 LGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPTFFT----- 830
+ N + +RC HV +LCVQD +RPNM+ V++ML+S+TATLP P+QPTF T
Sbjct: 762 IRATCNKREALRCIHVAMLCVQDSAAERPNMAAVLLMLESDTATLPVPRQPTFTTSTRRN 821
Query: 831 RKDLSSTASSSLQF-------DSSIVEGR 852
D++ SS Q+ S++V GR
Sbjct: 822 SMDVNFALDSSQQYIVSSNEITSTVVLGR 850
>emb|CAB81246.1| serine/threonine kinase-like protein [Arabidopsis thaliana]
gi|3402758|emb|CAA20204.1| serine/threonine kinase-like
protein [Arabidopsis thaliana]
gi|15234429|ref|NP_193870.1| S-locus lectin protein
kinase family protein [Arabidopsis thaliana]
gi|7428021|pir||T05181 S-receptor kinase (EC 2.7.1.-)
T6K22.120 precursor - Arabidopsis thaliana
Length = 849
Score = 570 bits (1468), Expect = e-161
Identities = 359/902 (39%), Positives = 517/902 (56%), Gaps = 106/902 (11%)
Query: 2 FFTPFTNMITIFLFHMHCWLLCFSQLCFAGDTLNVGQEITG--NGTVLVSAAKKFELGFF 59
FF + +++FL+ + A +T+ G+ + N LVS K FELGFF
Sbjct: 3 FFRKTSLYLSLFLYFF------LYESSMAANTIRRGESLRDGINHKPLVSPQKTFELGFF 56
Query: 60 SPDLNVTGGKGRYLGIWYYREEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDT 119
SP + R+LGIWY E VVWVANR P++D S GV I++DGNLV+LD
Sbjct: 57 SPGSSTH----RFLGIWYGNIEDKA---VVWVANRATPISDQS-GVLMISNDGNLVLLDG 108
Query: 120 SGIRYWSASNLTNSSSATNRSVKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKMDK 179
I WS++ +++++ NR V + D+GN VL + +WESF HPTDTFLP M++
Sbjct: 109 KNITVWSSNIESSTTNNNNRVVSIHDTGNFVLSETDTDRPIWESFNHPTDTFLPQMRVRV 168
Query: 180 TLE------LTCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQLY-WQSEEQGDGVMNPE 232
+ W+S +DP GN++ +D + + W+S + +
Sbjct: 169 NPQTGDNHAFVSWRSETDPSPGNYSLGVDPSGAPEIVLWEGNKTRKWRSGQWNSAIFTGI 228
Query: 233 SNPDDISNDVYNLLTNFKELKNKTVSSY-----DNTRLLLNSTGVIKVLYRVNFQSDIVW 287
N ++N +Y ++T S Y + +LL KVLY + ++ W
Sbjct: 229 PNMSLLTNYLYGF--KLSSPPDETGSVYFTYVPSDPSVLLR----FKVLYN-GTEEELRW 281
Query: 288 ------WY----QPRTTCLTYNVCGNFSSCN-DDNDKLCTCLPGFGRRSPLNDYTVGGDT 336
W +P + C YN CG F C+ ++ +C+C+ G+ + S + +++ G
Sbjct: 282 NETLKKWTKFQSEPDSECDQYNRCGKFGICDMKGSNGICSCIHGYEQVS-VGNWSRGCRR 340
Query: 337 SSLLCTRKSTSCGANTNTFLNLTMMKIGSPDIKVSAQ---DENECKFRCISMCSQTQCQA 393
+ L ++ S G + FL L +K+ PD ++ D +C+ RC+ CS C A
Sbjct: 341 RTPLKCERNISVGEDE--FLTLKSVKL--PDFEIPEHNLVDPEDCRERCLRNCS---CNA 393
Query: 394 CSYVPIPVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKS 453
S V G+ C IW Q+L L++ GG L +R+A S++ E N K+
Sbjct: 394 YSLVG------GIG---CMIWNQDLVDLQQFEAGGSS--LHIRLADSEVGE-----NRKT 437
Query: 454 TLSLILGIALPGVVILACICILAYVCRRKIALKLKQESESILRQRGRFYDSERHVKDLID 513
+++I+ + L GV+++ +L + +RK K S + G+ D+ V DL
Sbjct: 438 KIAVIVAV-LVGVILIGIFALLLWRFKRK-----KDVSGAYC---GKNTDTSVVVADLTK 488
Query: 514 KEG------------LEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKL 561
+ +E K E+P F +I +AT+ F N+LGRGG+GPVYKG L
Sbjct: 489 SKETTSAFSGSVDIMIEGKAVNTSELPVFSLNAIAIATNDFCKENELGRGGFGPVYKGVL 548
Query: 562 QGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNK 621
+ GREIAVKRLS S QG+ EFKNE++LIAKLQHRNLVRL G C +G+EK+L+YEYMPNK
Sbjct: 549 EDGREIAVKRLSGKSGQGVDEFKNEIILIAKLQHRNLVRLLGCCFEGEEKMLVYEYMPNK 608
Query: 622 SLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQP 681
SLD F+FD TK AL+DW++RF I+ GIARGLLYLH+DSRLR+IHRDLK SN+LLD EM P
Sbjct: 609 SLDFFLFDETKQALIDWKLRFSIIEGIARGLLYLHRDSRLRIIHRDLKVSNVLLDAEMNP 668
Query: 682 KISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGK 741
KISDFG+ARIFGG + EANT RVVGTYGYMSPEYA++G FS KSD++SFGV+LLEI+SGK
Sbjct: 669 KISDFGMARIFGGNQNEANTVRVVGTYGYMSPEYAMEGLFSVKSDVYSFGVLLLEIVSGK 728
Query: 742 KNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPD 801
+NT + SL+GYAW L+T + +L+D + + + +RC HV +LCVQD
Sbjct: 729 RNTSLRSSEHG-SLIGYAWYLYTHGRSEELVDPKIRVTCSKREALRCIHVAMLCVQDSAA 787
Query: 802 DRPNMSNVVIMLDSETATLPTPKQPTFFTRK----DLSSTASSSLQF-------DSSIVE 850
+RPNM++V++ML+S+TATL P+QPTF + + D++ SS Q+ S++V
Sbjct: 788 ERPNMASVLLMLESDTATLAAPRQPTFTSTRRNSIDVNFALDSSQQYIVSSNEITSTVVL 847
Query: 851 GR 852
GR
Sbjct: 848 GR 849
>emb|CAA67145.1| receptor-like kinase [Brassica oleracea var. acephala]
gi|7434413|pir||T14470 receptor-like kinase (EC 2.7.1.-)
SFR2 - wild cabbage
Length = 847
Score = 567 bits (1462), Expect = e-160
Identities = 361/863 (41%), Positives = 498/863 (56%), Gaps = 88/863 (10%)
Query: 21 LLCFSQLCFAGDTLNVGQEIT-GNGTVLVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYR 79
LL F F+ +TL+ + +T + + S FELGFF P + YLGIWY
Sbjct: 14 LLLFPAFSFSANTLSATESLTISSNKTISSPGNIFELGFFKP----SSSSRWYLGIWY-- 67
Query: 80 EEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNSSSATNR 139
+ VWVANRD+P++ S G +I+D NLVV+D S WS +NLT +
Sbjct: 68 -KAISKRTYVWVANRDHPLST-STGTLKISDS-NLVVVDGSDTAVWS-TNLTGGGDVRSP 123
Query: 140 SV-KLMDSGNLVLLDEHVG---MKLWESFEHPTDTFLPGMKMDKTLE------LTCWKSL 189
V +L+D+GN VL D + + LW+SF+ PTDT LP MK+ L+ L WKS
Sbjct: 124 VVAELLDNGNFVLRDSNNNDPDIVLWQSFDFPTDTLLPEMKLGWDLKTGFNWFLRSWKSP 183
Query: 190 SDPGRGNFTFKMDKK-WENRFAILNQGQLYWQSEEQG---DGVMNPESNPDDISNDVYNL 245
DP G+++FK+ + + F Q+Y G GV PE P D +N
Sbjct: 184 DDPSSGDYSFKLKTRGFPEAFLWNKASQVYRSGPWNGIRFSGV--PEMQPFDYIE--FNF 239
Query: 246 LTNFKELKNKTVSSYDN--TRLLLNSTGVIKVLYRVN-FQSDIVWWYQPRTTCLTYNVCG 302
T+ +E+ + DN +RL L+STG ++ + Q+ +WY P+ C Y CG
Sbjct: 240 TTSNQEVTYSFHITKDNMYSRLSLSSTGSLQRFTWIEAIQNWNQFWYAPKDQCDDYKECG 299
Query: 303 NFSSCNDDNDKLCTCLPGFGRRSP----LNDYTVGGDTSSLLCTRKSTSCGANTNTFLNL 358
+ C+ + +C C+ GF R+P L D + G C RK+ + F+ L
Sbjct: 300 TYGYCDSNTYPVCNCMRGFEPRNPQAWGLRDGSDG-------CVRKTALSCNGGDGFVRL 352
Query: 359 TMMKIGSPDIKVSAQDEN----ECKFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIW 414
MK+ PD ++ D EC+ +C S C+ T ++ ++ G S C +W
Sbjct: 353 KKMKL--PDTAATSVDRGIGIKECEEKCKSDCNCT-----AFANTDIRGGG---SGCVVW 402
Query: 415 TQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGVVILACICI 474
T ++ + GG D L+VR+A +D+E+ T + + I+G + GV +L +C
Sbjct: 403 TGDILDTRNYAKGGQD--LYVRLAATDLEDTTNRN------AKIIGSCI-GVSVLLLLCF 453
Query: 475 LAYVC-----RRKIALKL----KQESESILRQRGRFYDSERHVKDLIDKEGLEEKDNEGI 525
+ Y +R IA++ + S+ +L + RH+ E + +
Sbjct: 454 IFYRFWKRKQKRSIAIETSFVDQVRSQDLLMNEVVIPPNRRHIS--------RENKTDDL 505
Query: 526 EVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKN 585
E+P DFE++ +ATD FS+ANKLG+GG+G VYKG+L G+EIAVKRLS +S QG EFKN
Sbjct: 506 ELPLMDFEAVAIATDNFSNANKLGQGGFGIVYKGRLLDGQEIAVKRLSKMSVQGTDEFKN 565
Query: 586 EVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDIL 645
EV LIA+LQH NLVRL G C+ EK+LIYEY+ N SLD+ +FD T+S L+WQ RFDI
Sbjct: 566 EVKLIARLQHINLVRLLGCCVDEGEKMLIYEYLENLSLDSHLFDKTRSCKLNWQKRFDIT 625
Query: 646 LGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVV 705
GIARGLLYLHQDSR R+IHRDLK SN+LLD +M PKISDFG+ARIFG ETEANT++VV
Sbjct: 626 NGIARGLLYLHQDSRFRIIHRDLKASNVLLDKDMTPKISDFGMARIFGRDETEANTRKVV 685
Query: 706 GTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTE 765
GTYGYMSPEYA+DG FSTKSD+FSFGV+LLEIISGK+N GFY L+LLG W+ W +
Sbjct: 686 GTYGYMSPEYAMDGIFSTKSDVFSFGVLLLEIISGKRNKGFYNSDHDLNLLGCVWRNWKK 745
Query: 766 NKLLDLMDL----SLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLP 821
K LD++D S Y + +RC +GLLCVQ+ +DRP MS+VV+ML SETA +P
Sbjct: 746 GKGLDIVDPIILDSSPSTYRPLEILRCIKIGLLCVQERANDRPTMSSVVMMLGSETAAIP 805
Query: 822 TPKQPTFFT-RKDLSSTASSSLQ 843
P+QP + R L + +SSS Q
Sbjct: 806 QPEQPGYCVGRSPLDTDSSSSNQ 828
>dbj|BAB69683.1| receptor kinase 5 [Brassica rapa]
Length = 838
Score = 565 bits (1457), Expect = e-159
Identities = 360/859 (41%), Positives = 492/859 (56%), Gaps = 84/859 (9%)
Query: 21 LLCFSQLCFAGDTLNVGQEIT-GNGTVLVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYR 79
LL F F+ +TL+ + +T + + S FELGFF P + YLGIWY
Sbjct: 9 LLLFPAFSFSANTLSATESLTISSNKTISSPGNIFELGFFKP----SSSSRWYLGIWY-- 62
Query: 80 EEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNSSSATNR 139
+ VWVANRD+P++ S G +I+D NLVV+D S WS +NLT +
Sbjct: 63 -KAISKRTYVWVANRDHPLST-STGTLKISDS-NLVVVDGSDTAVWS-TNLTGGGDVRSP 118
Query: 140 SV-KLMDSGNLVLLDEHVGMK---LWESFEHPTDTFLPGMKMDKTLE------LTCWKSL 189
V +L+D+GNLVL D + LW+SF+ PTDT LP MK+ L+ L WKS
Sbjct: 119 VVAELLDNGNLVLRDSNNNDPDGVLWQSFDFPTDTLLPEMKLGWDLKTGFNRFLRSWKSP 178
Query: 190 SDPGRGNFTFKMDKK-WENRFAILNQGQLYWQSEEQG---DGVMNPESNPDDISNDVYNL 245
DP G+++FK++ + + F Q+Y G GV PE P D +N
Sbjct: 179 DDPSSGDYSFKLETRGFPEAFLWNKASQVYRSGPWNGIRFSGV--PEMQPFDYIE--FNF 234
Query: 246 LTNFKELKNKTVSSYDN--TRLLLNSTGVIKVLYRVN-FQSDIVWWYQPRTTCLTYNVCG 302
T+ +E+ + DN +RL L+STG ++ + Q+ +WY P+ C Y CG
Sbjct: 235 TTSNQEVTYSFHITKDNMYSRLSLSSTGSLQRFTWIEAIQNWNQFWYAPKDQCDEYKECG 294
Query: 303 NFSSCNDDNDKLCTCLPGFGRRSP----LNDYTVGGDTSSLLCTRKSTSCGANTNTFLNL 358
F C+ + +C C+ GF R+P L D + G C RK+ + F+ L
Sbjct: 295 TFGYCDSNTYPVCNCMRGFEPRNPQAWALRDGSDG-------CVRKTALSCNGGDGFVRL 347
Query: 359 TMMKIGSPDIKVSAQDEN----ECKFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIW 414
MK+ PD ++ D EC+ +C S C+ T ++ ++ G S C +W
Sbjct: 348 KKMKL--PDTAATSVDRGIGIKECEEKCKSDCNCT-----AFANTDIRGGG---SGCVVW 397
Query: 415 TQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGVVILACICI 474
T ++ + GG D L+VR+A +D+E+ T + + I+G + GV +L +C
Sbjct: 398 TGDILDTRNYAKGGQD--LYVRLAATDLEDTTNRN------AKIIGSCI-GVSVLLLLCF 448
Query: 475 LAYVC-----RRKIALKLKQESESILRQRGRFYDSERHVKDLIDKEGLEEKDNEGIEVPY 529
+ Y +R IA++ L S RH+ E + E+P
Sbjct: 449 IFYRFWKRKQKRSIAIETSFVRSQDLLMNEVVIPSRRHIS--------RENKTDDFELPL 500
Query: 530 FDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVL 589
DFE++ +ATD F++ANKLG+GG+G VYKG+L G+EIAVKRLS +S QG EFKNEV L
Sbjct: 501 MDFEAVAIATDNFTNANKLGQGGFGIVYKGRLLDGQEIAVKRLSKMSVQGTDEFKNEVKL 560
Query: 590 IAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIA 649
IA+LQH NLVRL G C+ EK+LIYEY+ N SLD+ +FD T+S L+WQ RFDI GIA
Sbjct: 561 IARLQHINLVRLLGCCVDEGEKMLIYEYLENLSLDSHLFDKTRSCKLNWQKRFDITNGIA 620
Query: 650 RGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYG 709
RGLLYLHQDSR R+IHRDLK SN+LLD +M PKISDFG+ARIFG ETEANT++VVGTYG
Sbjct: 621 RGLLYLHQDSRFRIIHRDLKASNVLLDKDMTPKISDFGMARIFGRDETEANTRKVVGTYG 680
Query: 710 YMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLL 769
YMSPEYA+DG FSTKSD+FSFGV+LLEIISGK+N GFY L+LLG W+ W + K L
Sbjct: 681 YMSPEYAMDGIFSTKSDVFSFGVLLLEIISGKRNKGFYNSDHDLNLLGCVWRNWKKGKGL 740
Query: 770 DLMDL----SLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQ 825
D++D S Y + +RC +GLLCVQ+ +DRP MS+VV+ML SET +P P+
Sbjct: 741 DIVDPIILDSSPSTYRPLEILRCIKIGLLCVQERANDRPTMSSVVMMLGSETTAIPQPEP 800
Query: 826 PTFFT-RKDLSSTASSSLQ 843
P + R L + +SSS Q
Sbjct: 801 PGYCVGRSPLDTDSSSSNQ 819
>emb|CAA73133.1| serine /threonine kinase [Brassica oleracea]
Length = 847
Score = 564 bits (1453), Expect = e-159
Identities = 360/863 (41%), Positives = 497/863 (56%), Gaps = 88/863 (10%)
Query: 21 LLCFSQLCFAGDTLNVGQEIT-GNGTVLVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYR 79
LL F F+ +TL+ + +T + + S FELGFF P + YLGIWY
Sbjct: 14 LLLFPAFSFSSNTLSATESLTISSNKTISSPGNIFELGFFKP----SSSSRWYLGIWY-- 67
Query: 80 EEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNSSSATNR 139
+ VWVANRD+P++ S G +I+D NLVV+D S WS +NLT +
Sbjct: 68 -KAISKRTYVWVANRDHPLST-STGTLKISDS-NLVVVDGSDTAVWS-TNLTGGGDVRSP 123
Query: 140 SV-KLMDSGNLVLLDEHVG---MKLWESFEHPTDTFLPGMKMDKTLE------LTCWKSL 189
V +L+D+GN VL D + + LW+SF+ PTDT LP MK+ L+ L WKS
Sbjct: 124 VVAELLDNGNFVLRDSNNNDPDIVLWQSFDFPTDTLLPEMKLGWDLKTGFNWFLRSWKSP 183
Query: 190 SDPGRGNFTFKMDKK-WENRFAILNQGQLYWQSEEQG---DGVMNPESNPDDISNDVYNL 245
DP G+++FK+ + + F Q+Y G GV PE P D +N
Sbjct: 184 DDPSSGDYSFKLKTRGFPEAFLWNKASQVYRSGPWNGIRFSGV--PEMQPFDYIE--FNF 239
Query: 246 LTNFKELKNKTVSSYDN--TRLLLNSTGVIKVLYRVN-FQSDIVWWYQPRTTCLTYNVCG 302
T+ +E+ + DN +RL L+STG ++ + Q+ +WY P+ C Y CG
Sbjct: 240 TTSNQEVTYSFHITKDNMYSRLSLSSTGSLQRFTWIEAIQNWNQFWYAPKDQCDDYKECG 299
Query: 303 NFSSCNDDNDKLCTCLPGFGRRSP----LNDYTVGGDTSSLLCTRKSTSCGANTNTFLNL 358
+ C+ + +C C+ GF R+P L D + G C RK+ + F+ L
Sbjct: 300 TYGYCDSNTYPVCNCMRGFEPRNPQAWGLRDGSDG-------CVRKTALSCNGGDGFVRL 352
Query: 359 TMMKIGSPDIKVSAQDEN----ECKFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIW 414
MK+ PD ++ D EC+ +C S C+ T ++ ++ G S C +W
Sbjct: 353 KKMKL--PDTAATSVDRGIGIKECEEKCKSDCNCT-----AFANTDIRGGG---SGCVVW 402
Query: 415 TQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGVVILACICI 474
T ++ + GG D L+VR+A +D+E+ T + + I+G + GV +L +C
Sbjct: 403 TGDILDTRNYAKGGQD--LYVRLAATDLEDTTNRN------AKIIGSCI-GVSVLLLLCF 453
Query: 475 LAYVC-----RRKIALKL----KQESESILRQRGRFYDSERHVKDLIDKEGLEEKDNEGI 525
+ Y +R IA++ + S+ +L + RH+ E + +
Sbjct: 454 IFYRFWKRKQKRSIAIETSFVDQVRSQDLLMNEVVIPPNRRHIS--------RENKTDDL 505
Query: 526 EVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKN 585
E+P DFE++ +ATD FS+ANKLG+GG+G VYKG+L G+EIAVKRLS +S QG EFKN
Sbjct: 506 ELPLMDFEAVAIATDNFSNANKLGQGGFGIVYKGRLLDGQEIAVKRLSKMSVQGTDEFKN 565
Query: 586 EVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDIL 645
EV LIA+LQH NLVRL G C+ EK+LIYEY+ N SLD+ +FD T+S L+WQ RF I
Sbjct: 566 EVKLIARLQHINLVRLLGCCVDEGEKMLIYEYLENLSLDSHLFDKTRSCKLNWQKRFVIT 625
Query: 646 LGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVV 705
GIARGLLYLHQDSR R+IHRDLK SN+LLD +M PKISDFG+ARIFG ETEANT++VV
Sbjct: 626 NGIARGLLYLHQDSRFRIIHRDLKASNVLLDKDMTPKISDFGMARIFGRDETEANTRKVV 685
Query: 706 GTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTE 765
GTYGYMSPEYA+DG FSTKSD+FSFGV+LLEIISGK+N GFY L+LLG W+ W +
Sbjct: 686 GTYGYMSPEYAMDGIFSTKSDVFSFGVLLLEIISGKRNKGFYNSDHDLNLLGCVWRNWKK 745
Query: 766 NKLLDLMDL----SLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLP 821
K LD++D S Y + +RC +GLLCVQ+ +DRP MS+VV+ML SETA +P
Sbjct: 746 GKGLDIVDPIILDSSPSTYRPLEILRCIKIGLLCVQERANDRPTMSSVVMMLGSETAAIP 805
Query: 822 TPKQPTFFT-RKDLSSTASSSLQ 843
P+QP + R L + +SSS Q
Sbjct: 806 QPEQPGYCVGRSPLDTDSSSSNQ 828
>gb|AAB33487.1| ARK3 product/receptor-like serine/threonine protein kinase ARK3
[Arabidopsis thaliana, Columbia, Peptide, 851 aa]
Length = 851
Score = 561 bits (1447), Expect = e-158
Identities = 358/874 (40%), Positives = 500/874 (56%), Gaps = 89/874 (10%)
Query: 6 FTNMITIFLFHMHCWLLCFSQLCFAGDTLNVGQEIT-GNGTVLVSAAKKFELGFFSPDLN 64
F + T F F + L+ F + +TL+ + +T + +VS FELGFF P L+
Sbjct: 7 FYHSYTFFFFFL---LILFPAYSISANTLSASESLTISSNNTIVSPGNVFELGFFKPGLD 63
Query: 65 VTGGKGRYLGIWYYREEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRY 124
YLGIWY + VWVANRD P++ SIG +I+D NLVVLD S
Sbjct: 64 SRW----YLGIWY---KAISKRTYVWVANRDTPLSS-SIGTLKISDS-NLVVLDQSDTPV 114
Query: 125 WSASNLTNSSSATNRSVKLMDSGNLVLLDEHVGMK---LWESFEHPTDTFLPGMKMDKTL 181
WS +NLT + +L+D+GN VL D LW+SF+ PTDT LP MK+
Sbjct: 115 WS-TNLTGGDVRSPLVAELLDNGNFVLRDSKNSAPDGVLWQSFDFPTDTLLPEMKLGWDA 173
Query: 182 E------LTCWKSLSDPGRGNFTFKMDKKWENRFAILN-QGQLYWQSEEQG---DGVMNP 231
+ + WKS DP G+F+FK++ + + N + ++Y G GV P
Sbjct: 174 KTGFNRFIRSWKSPDDPSSGDFSFKLETEGFPEIFLWNRESRMYRSGPWNGIRFSGV--P 231
Query: 232 ESNPDDISNDVYNLLTNFKELKN--KTVSSYDNTRLLLNSTGVI-KVLYRVNFQSDIVWW 288
E P + V+N T+ +E+ + S +RL ++S+G++ + + Q+ +W
Sbjct: 232 EMQPFEYM--VFNFTTSKEEVTYSFRITKSDVYSRLSISSSGLLQRFTWIETAQNWNQFW 289
Query: 289 YQPRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSP----LNDYTVGGDTSSLLCTRK 344
Y P+ C Y CG + C+ + +C C+ GF R+P L D + G C RK
Sbjct: 290 YAPKDQCDEYKECGVYGYCDSNTSPVCNCIKGFKPRNPQVWGLRDGSDG-------CVRK 342
Query: 345 ST-SCGANTNTFLNLTMMKIGSPDIKVSAQDEN----ECKFRCISMCSQTQCQACSYVPI 399
+ SCG F+ L MK+ PD ++ D EC+ +C+ C+ C A + I
Sbjct: 343 TLLSCGGGDG-FVRLKKMKL--PDTTTASVDRGIGVKECEQKCLRDCN---CTAFANTDI 396
Query: 400 PVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLIL 459
RG S C WT L ++ GG D L+VR+A +D+E+ + + I+
Sbjct: 397 ----RGSG-SGCVTWTGELFDIRNYAKGGQD--LYVRLAATDLEDKRNRS------AKII 443
Query: 460 GIALPGVVILACICILAYVCRRKIALKLKQESESI---LRQRGRFYD-----SERHVKDL 511
G ++ V+L I+ ++ +RK + E+ + LR R + S RH+
Sbjct: 444 GSSIGVSVLLLLSFIIFFLWKRKQKRSILIETPIVDHQLRSRDLLMNEVVISSRRHIS-- 501
Query: 512 IDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKR 571
E + + +E+P +FE + +AT+ FS+ANKLG+GG+G VYKGKL G+E+AVKR
Sbjct: 502 ------RENNTDDLELPLMEFEEVAMATNNFSNANKLGQGGFGIVYKGKLLDGQEMAVKR 555
Query: 572 LSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPT 631
LS S QG EFKNEV LIA+LQH NLVRL C+ EK+LIYEY+ N SLD+ +FD +
Sbjct: 556 LSKTSVQGTDEFKNEVKLIARLQHINLVRLLACCVDAGEKMLIYEYLENLSLDSHLFDKS 615
Query: 632 KSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARI 691
+++ L+WQMRFDI+ GIARGLLYLHQDSR R+IHRDLK SNILLD M PKISDFG+ARI
Sbjct: 616 RNSKLNWQMRFDIINGIARGLLYLHQDSRFRIIHRDLKASNILLDKYMTPKISDFGMARI 675
Query: 692 FGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKG 751
FG ETEANT++VVGTYGYMSPEYA+DG FS KSD+FSFGV+LLEIIS K+N GFY
Sbjct: 676 FGRDETEANTRKVVGTYGYMSPEYAMDGIFSMKSDVFSFGVLLLEIISSKRNKGFYNSDR 735
Query: 752 TLSLLGYAWKLWTENKLLDLMDL----SLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMS 807
L+LLG W+ W E K L+++D SL + ++ +RC +GLLCVQ+ +DRP MS
Sbjct: 736 DLNLLGCVWRNWKEGKGLEIIDPIITDSLSSTFRQHEILRCIQIGLLCVQERAEDRPTMS 795
Query: 808 NVVIMLDSETATLPTPKQPTFFTRKDLSSTASSS 841
V++ML SE+ T+P PK P + + L T SSS
Sbjct: 796 LVILMLGSESTTIPQPKAPGYCLERSLLDTDSSS 829
>emb|CAA74661.1| SFR1 [Brassica oleracea] gi|7434410|pir||T14519 probable S-receptor
kinase (EC 2.7.1.-) SFR1 - wild cabbage
Length = 849
Score = 561 bits (1445), Expect = e-158
Identities = 353/848 (41%), Positives = 493/848 (57%), Gaps = 72/848 (8%)
Query: 11 TIFLFHMHCWLLCFSQLCFAGDTLNVGQEIT--GNGTVLVSAAKKFELGFFSPDLNVTGG 68
T+ L + L+ F + +T + + +T N T+L S ++ FELGFF+P
Sbjct: 12 TVVLMFIFLVLILFHAFPVSANTFSATESLTISSNKTIL-SRSEIFELGFFNPP----SS 66
Query: 69 KGRYLGIWYYREEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSAS 128
YLGIWY + VWVANRDNP+ + G I+D NLV+ D S WS +
Sbjct: 67 SRWYLGIWYKKVS---TRTYVWVANRDNPLLSSN-GTLNISDS-NLVIFDQSDTPVWS-T 120
Query: 129 NLTNSSSATNRSVKLMDSGNLVLLDEHVGMK------LWESFEHPTDTFLPGMKMD---- 178
NLT + +L+D+GN VL H+ LW+SF+ PTDT LP M++
Sbjct: 121 NLTEGEVRSPVVAELLDNGNFVL--RHLNNNNDPDGYLWQSFDFPTDTLLPEMRLGWDHK 178
Query: 179 --KTLELTCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVM---NPES 233
+ L WK+ DP G+F K+ K F + ++ + ++S +G+ +PE+
Sbjct: 179 TGRDRFLRSWKTPDDPSSGDFFTKLKTKGFPEFYVCSKDSIIYRSGPW-NGIRFSSSPET 237
Query: 234 NPDDISNDVYNLLTNFKELKNKTVSSYDNT--RLLLNSTGVIKVLYRVNF-QSDIVWWYQ 290
P D VYN +E+ + + N R+ L+S G+++ L + QS WY
Sbjct: 238 KPLDYI--VYNFTATNEEVSYSYLITKTNIYERVRLSSAGLLERLTWIETAQSWKQLWYS 295
Query: 291 PRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSPLNDYTVGGDTSSLLCTRKSTSCGA 350
P+ C Y CG++ C+ + +C C+ GFG + +T+ D++ C RK+
Sbjct: 296 PKDLCDNYKECGSYGYCDSNTSPICNCIKGFGPGNQ-QPWTLRDDSAG--CVRKTRLSCD 352
Query: 351 NTNTFLNLTMMKIGSPDIKVSAQDEN----ECKFRCISMCSQTQCQACSYVPIPVQQRGL 406
+ F+ L MK+ PD + D EC+ RC+ C+ T ++ ++ G
Sbjct: 353 GRDGFVRLKKMKL--PDTTATTVDRGIGLKECEERCLKDCNCT-----AFANTDIRNGG- 404
Query: 407 NLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEP-TRKGNPKSTLSLILGIALPG 465
S C IWT + +K GG D LFVR+A +D+E+ T+K N +ILG+++
Sbjct: 405 --SGCVIWTGEIFDIKNFAKGGQD--LFVRLAAADLEDKRTKKRN------IILGLSIGV 454
Query: 466 VVILACICILAYVCRRKIALKLKQESESILRQRGRFYDSERHVKDLIDKEGLE-EKDNEG 524
++L I+ +RK +++S +I + DS + + K L + E
Sbjct: 455 SILLLLSFIIFRFWKRK-----QKQSVAIPKPIVTSQDSLMNEVVISSKRHLSGDMKTED 509
Query: 525 IEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFK 584
+E+P DFE+I AT FS NKLG+GG+G VYKG+L G+EIAVKRLS +S QG EFK
Sbjct: 510 LELPLMDFEAIATATHNFSSTNKLGQGGFGIVYKGRLLDGKEIAVKRLSKMSLQGTDEFK 569
Query: 585 NEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDI 644
NEV LIA+LQH NLVRL G C+ EK+LIYEY+ N SLD+ +FD ++ + L+WQ+RFDI
Sbjct: 570 NEVRLIARLQHINLVRLLGCCVDKGEKMLIYEYLENLSLDSHLFDKSRRSNLNWQLRFDI 629
Query: 645 LLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRV 704
GIARGLLYLHQDSR R+IHRDLK SNILLD M PKISDFG+ARIF ETEANT++V
Sbjct: 630 ANGIARGLLYLHQDSRFRIIHRDLKVSNILLDKNMIPKISDFGMARIFRRDETEANTRKV 689
Query: 705 VGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWT 764
VGTYGYMSPEYA++G FS KSD+FSFGV+LLEIISGK++TGFY G LSLLG W+ W
Sbjct: 690 VGTYGYMSPEYAMNGIFSVKSDVFSFGVLLLEIISGKRSTGFYNSSGDLSLLGCVWRNWK 749
Query: 765 ENKLLDLMDL----SLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATL 820
E K LD++D SL + ++ +RC H+GLLCVQ+ +DRP MS+V++ML SET TL
Sbjct: 750 ERKGLDIIDPIIIDSLSSTFKTHEILRCIHIGLLCVQERAEDRPAMSSVMVMLGSETTTL 809
Query: 821 PTPKQPTF 828
P PKQP F
Sbjct: 810 PEPKQPAF 817
>ref|NP_916409.1| putative receptor kinase [Oryza sativa (japonica cultivar-group)]
Length = 835
Score = 560 bits (1444), Expect = e-158
Identities = 357/873 (40%), Positives = 488/873 (55%), Gaps = 97/873 (11%)
Query: 19 CWLLCFSQL------CFAGDTLNVGQEITGNGTVLVSAAKKFELGFFSPDLNVTGGKGRY 72
C+LL F + C A DT+ G+ + N T++ F LGFF+P G Y
Sbjct: 9 CYLLLFVVVVVLTGSCRARDTVVPGRPLAANETLVSGGDANFVLGFFTPP----GANSTY 64
Query: 73 LGIWYYREEGSGLSPVVWVANRDNP----VADDSIGVFRIADDGNLVVLDTSGIRYWSAS 128
+G+WY + + VVWVANR++P VAD+ ++ G L ++ + WS +
Sbjct: 65 VGVWYNKVS---VRTVVWVANREDPLPGDVADNPDATLSVSPTGTLAIVAGNSTVVWSVT 121
Query: 129 NLTNSSSATNRSVKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKMD------KTLE 182
+S T R +MDSGNLV+ D G W+ F++PTDT LP M++ +
Sbjct: 122 PAAKLASPTAR---IMDSGNLVIADGAGGGVAWQGFDYPTDTLLPEMRLGVDYVKGRNRT 178
Query: 183 LTCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPDDI--SN 240
LT WKS SDP G MD + + I N + W+S DGV PD + S
Sbjct: 179 LTAWKSPSDPSPGPVVMAMDTSGDPQVFIWNGAEKVWRSGPW-DGVQFT-GVPDTVTYSG 236
Query: 241 DVYNLLTNFKELK------NKTVSSYDNTRLLLNSTGVIKVLYRVNFQSDI----VWWYQ 290
++ + N KE+ N ++ S RL LNSTG +L R + ++WY
Sbjct: 237 FTFSFINNAKEVTYSFQVHNVSIIS----RLGLNSTGSYGLLQRSTWVEAAGTWNLYWYA 292
Query: 291 PRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSP----LNDYTVGGDTSSLLCTRKST 346
P+ C + CG C+ +N +C+CL GF +SP L D G C R +
Sbjct: 293 PKDQCDEVSPCGANGVCDTNNLPVCSCLRGFTPKSPEAWALRDGRAG-------CVRSTP 345
Query: 347 -SCGANTNTFLNLTMMKIGSPDIKVSAQDEN----ECKFRCISMCSQTQCQACSYVPIPV 401
C T+ F+ + K+ PD + S D +C+ C+ CS C A + +
Sbjct: 346 LDCQNGTDGFVAVEHAKV--PDTERSVVDLGLSLEQCRKACLMNCS---CTAYASANVSG 400
Query: 402 QQRGLNLSP-CWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILG 460
RG C +WT LT L+ G D LFVR+A +D+ ++ + +++++
Sbjct: 401 GGRGHGAGTGCVMWTTGLTDLRVYPEFGQD--LFVRLAAADLGLTSKSNKARVIIAIVVS 458
Query: 461 IALPGVVILACIC-ILAYVCRRKIALKLKQESESI-LRQRGRFYDSERHVKDLIDKEGLE 518
I+ V L+ + L + ++K A K S R GR Y+ H D
Sbjct: 459 IS--SVTFLSVLAGFLVWTRKKKRARKTGSSKWSGGSRSTGRRYEGSSHHDD-------- 508
Query: 519 EKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQ 578
+E+P FD +I ATD FS NKLG GG+GPVYKGKL+ G+EIAVK LS S Q
Sbjct: 509 -----DLELPIFDLGTIAAATDGFSINNKLGEGGFGPVYKGKLEDGQEIAVKTLSKTSVQ 563
Query: 579 GIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDW 638
G+ EFKNEV+LIAKLQHRNLVRL G+ I G E+IL+YEYM NKSLD F+F
Sbjct: 564 GLDEFKNEVMLIAKLQHRNLVRLLGFSISGQERILVYEYMANKSLDYFLF---------- 613
Query: 639 QMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETE 698
R+ I+ GI RGLLYLHQDSR R+IHRDLK SN+LLD EM PKISDFG+AR+FG +ETE
Sbjct: 614 -ARYRIIEGITRGLLYLHQDSRYRIIHRDLKASNVLLDKEMTPKISDFGMARMFGSEETE 672
Query: 699 ANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGY 758
NT++VVGTYGYMSPEYA+DG FS KSD+FSFGV+LLEIISG++N G Y Y L+LLG+
Sbjct: 673 INTRKVVGTYGYMSPEYAMDGVFSVKSDVFSFGVLLLEIISGRRNRGVYSYSNHLNLLGH 732
Query: 759 AWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIML-DSET 817
AW LW E K L+L D ++ ++++++ ++C VGLLCVQ+ PDDRP MS V++ML ++
Sbjct: 733 AWSLWNEGKSLELADETMNGSFDSDEVLKCIRVGLLCVQENPDDRPLMSQVLLMLATTDA 792
Query: 818 ATLPTPKQPTFFTRKDLSSTASSSLQFDSSIVE 850
TLPTPKQP F R+ L T +SS + D SI +
Sbjct: 793 TTLPTPKQPGFAARRILMETDTSSSKPDCSIFD 825
>emb|CAB81245.1| receptor-like serine/threonine protein kinase ARK3 [Arabidopsis
thaliana] gi|3402757|emb|CAA20203.1| receptor-like
serine/threonine protein kinase ARK3 [Arabidopsis
thaliana] gi|29824117|gb|AAP04019.1| putative receptor
serine/threonine protein kinase ARK3 [Arabidopsis
thaliana] gi|26452798|dbj|BAC43479.1| putative
receptor-like serine/threonine protein kinase ARK3
[Arabidopsis thaliana] gi|15234427|ref|NP_193869.1|
S-locus protein kinase, putative (ARK3) [Arabidopsis
thaliana] gi|7428017|pir||T05180 S-receptor kinase (EC
2.7.1.-) ARK3 precursor - Arabidopsis thaliana
Length = 850
Score = 559 bits (1441), Expect = e-157
Identities = 356/873 (40%), Positives = 501/873 (56%), Gaps = 88/873 (10%)
Query: 6 FTNMITIFLFHMHCWLLCFSQLCFAGDTLNVGQEIT-GNGTVLVSAAKKFELGFFSPDLN 64
F + T F F + L+ F + +TL+ + +T + +VS FELGFF P L+
Sbjct: 7 FYHSYTFFFFFL---LILFPAYSISANTLSASESLTISSNNTIVSPGNVFELGFFKPGLD 63
Query: 65 VTGGKGRYLGIWYYREEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRY 124
YLGIWY + VWVANRD P++ SIG +I+D NLVVLD S
Sbjct: 64 SRW----YLGIWY---KAISKRTYVWVANRDTPLSS-SIGTLKISDS-NLVVLDQSDTPV 114
Query: 125 WSASNLTNSSSATNRSVKLMDSGNLVLLDEHVGMK---LWESFEHPTDTFLPGMKMDKTL 181
WS +NLT + +L+D+GN VL D LW+SF+ PTDT LP MK+
Sbjct: 115 WS-TNLTGGDVRSPLVAELLDNGNFVLRDSKNSAPDGVLWQSFDFPTDTLLPEMKLGWDA 173
Query: 182 E------LTCWKSLSDPGRGNFTFKMDKKWENRFAILN-QGQLYWQSEEQG---DGVMNP 231
+ + WKS DP G+F+FK++ + + N + ++Y G GV P
Sbjct: 174 KTGFNRFIRSWKSPDDPSSGDFSFKLETEGFPEIFLWNRESRMYRSGPWNGIRFSGV--P 231
Query: 232 ESNPDDISNDVYNLLTNFKELKN--KTVSSYDNTRLLLNSTGVI-KVLYRVNFQSDIVWW 288
E P + V+N T+ +E+ + S +RL ++S+G++ + + Q+ +W
Sbjct: 232 EMQPFEYM--VFNFTTSKEEVTYSFRITKSDVYSRLSISSSGLLQRFTWIETAQNWNQFW 289
Query: 289 YQPRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSP----LNDYTVGGDTSSLLCTRK 344
Y P+ C Y CG + C+ + +C C+ GF R+P L D + G C RK
Sbjct: 290 YAPKDQCDEYKECGVYGYCDSNTSPVCNCIKGFKPRNPQVWGLRDGSDG-------CVRK 342
Query: 345 ST-SCGANTNTFLNLTMMKIGSPDIKVSAQDEN----ECKFRCISMCSQTQCQACSYVPI 399
+ SCG F+ L MK+ PD ++ D EC+ +C+ C+ C A + I
Sbjct: 343 TLLSCGGGDG-FVRLKKMKL--PDTTTASVDRGIGVKECEQKCLRDCN---CTAFANTDI 396
Query: 400 PVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLIL 459
RG S C WT L ++ GG D L+VR+A +D+E+ + + I+
Sbjct: 397 ----RGSG-SGCVTWTGELFDIRNYAKGGQD--LYVRLAATDLEDKRNRS------AKII 443
Query: 460 GIALPGVVILACICILAYVCRRKIALKLKQESESI---LRQRGRFYD-----SERHVKDL 511
G ++ V+L I+ ++ +RK + E+ + LR R + S RH+
Sbjct: 444 GSSIGVSVLLLLSFIIFFLWKRKQKRSILIETPIVDHQLRSRDLLMNEVVISSRRHIS-- 501
Query: 512 IDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKR 571
E + + +E+P +FE + +AT+ FS+ANKLG+GG+G VYKGKL G+E+AVKR
Sbjct: 502 ------RENNTDDLELPLMEFEEVAMATNNFSNANKLGQGGFGIVYKGKLLDGQEMAVKR 555
Query: 572 LSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPT 631
LS S QG EFKNEV LIA+LQH NLVRL C+ EK+LIYEY+ N SLD+ +FD +
Sbjct: 556 LSKTSVQGTDEFKNEVKLIARLQHINLVRLLACCVDAGEKMLIYEYLENLSLDSHLFDKS 615
Query: 632 KSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARI 691
+++ L+WQMRFDI+ GIARGLLYLHQDSR R+IHRDLK SNILLD M PKISDFG+ARI
Sbjct: 616 RNSKLNWQMRFDIINGIARGLLYLHQDSRFRIIHRDLKASNILLDKYMTPKISDFGMARI 675
Query: 692 FGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKG 751
FG ETEANT++VVGTYGYMSPEYA+DG FS KSD+FSFGV+LLEIIS K+N GFY
Sbjct: 676 FGRDETEANTRKVVGTYGYMSPEYAMDGIFSMKSDVFSFGVLLLEIISSKRNKGFYNSDR 735
Query: 752 TLSLLGYAWKLWTENKLLDLMDLSLGEA---YNANQFIRCTHVGLLCVQDEPDDRPNMSN 808
L+LLG W+ W E K L+++D + ++ + ++ +RC +GLLCVQ+ +DRP MS
Sbjct: 736 DLNLLGCVWRNWKEGKGLEIIDPIITDSSSTFRQHEILRCIQIGLLCVQERAEDRPTMSL 795
Query: 809 VVIMLDSETATLPTPKQPTFFTRKDLSSTASSS 841
V++ML SE+ T+P PK P + + L T SSS
Sbjct: 796 VILMLGSESTTIPQPKAPGYCLERSLLDTDSSS 828
>dbj|BAB40987.1| SRKb [Arabidopsis lyrata]
Length = 853
Score = 559 bits (1441), Expect = e-157
Identities = 356/849 (41%), Positives = 500/849 (57%), Gaps = 87/849 (10%)
Query: 33 TLNVGQEIT-GNGTVLVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYREEGSGLSPVVWV 91
TL+ + +T + +VS + FELGFF+P G YLGIW+ + + VWV
Sbjct: 31 TLSSTESLTISSKQTIVSPGEVFELGFFNPAATSRDGDRWYLGIWF---KTNLERTYVWV 87
Query: 92 ANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNSSSATNRSV--KLMDSGNL 149
ANRDNP+ + S G +I+D NLV+LD WS TN + V +L+ +GNL
Sbjct: 88 ANRDNPLYN-STGTLKISDT-NLVLLDQFDTLVWS----TNLTGVLRSPVVAELLSNGNL 141
Query: 150 VLLDEHVGMK---LWESFEHPTDTFLPGMKMDKTLE------LTCWKSLSDPGRGNFTFK 200
VL D K LW+SF++PTDT LP MKM ++ L WKS DP G+F++K
Sbjct: 142 VLKDSKTNDKDGILWQSFDYPTDTLLPQMKMGWDVKKGLNRFLRSWKSQYDPSSGDFSYK 201
Query: 201 MDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPDDISNDVYNLLTNFKELKNKTVSSY 260
++ + F +L + ++S DG+ S ++ Y +++NF E + + ++
Sbjct: 202 LETRGFPEFFLLWRNSRVFRSGPW-DGLRF--SGIPEMQQWEY-MVSNFTENREEVAYTF 257
Query: 261 DNT------RLLLNSTGVIKVLYRVNFQSDIVW---WYQPRTTCLTYNVCGNFSSCNDDN 311
T R ++STG +K ++ + W W +P C Y CG +S C+ +
Sbjct: 258 QITNHNIYSRFTMSSTGALKRFRWISSSEE--WNQLWNKPNDHCDMYKRCGPYSYCDMNT 315
Query: 312 DKLCTCLPGFGRRSPLNDYTVGGDTSSLLCTRKST-SCGANTNTFLNLTMMKIGSPDIKV 370
+C C+ GF R+ L+++T+ S+ C RK+ +CG + FL L MK+ PD
Sbjct: 316 SPICNCIGGFKPRN-LHEWTLRN--GSIGCVRKTRLNCGGDG--FLCLRKMKL--PDSSA 368
Query: 371 SAQDEN----ECKFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIWTQNLTTLKEEYL 426
+ D ECK RC++ C+ T +Y +Q GL C IW + L ++
Sbjct: 369 AIVDRTIDLGECKKRCLNDCNCT-----AYASTDIQNGGLG---CVIWIEELLDIRNYAS 420
Query: 427 GGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGVVILACICILAYVCRRKIALK 486
GG D L+VR+A DI G+ ++ I+G+A+ VIL I+ V RRK L
Sbjct: 421 GGQD--LYVRLADVDI------GDERNIRGKIIGLAVGASVILFLSSIMFCVWRRKQKLL 472
Query: 487 LKQES--------ESILRQRGRFYDSERHVKDLIDKEGLEEKDNEGIEVPYFDFESILVA 538
E+ + +L R S RH+ E+ E +E+P +FE++++A
Sbjct: 473 RATEAPIVYPTINQGLLMNRLEI-SSGRHLS--------EDNQTEDLELPLVEFEAVVMA 523
Query: 539 TDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNL 598
T+ FS++NKLG GG+G VYKG+L G+EIAVKRLS+ S QGI EF+NEV LI+KLQH NL
Sbjct: 524 TENFSNSNKLGEGGFGVVYKGRLLDGQEIAVKRLSTTSIQGICEFRNEVKLISKLQHINL 583
Query: 599 VRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQD 658
VRL+G C+ +EK+LIYEY+ N SLD+ +F+ + S L+WQMRFDI GIARGLLYLHQD
Sbjct: 584 VRLFGCCVDENEKMLIYEYLENLSLDSHLFNKSLSCKLNWQMRFDITNGIARGLLYLHQD 643
Query: 659 SRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALD 718
SR R+IHRDLK SN+LLD +M PKISDFG+ARIFG ETEANT++VVGTYGYMSPEYA+D
Sbjct: 644 SRFRIIHRDLKASNVLLDKDMTPKISDFGMARIFGRDETEANTRKVVGTYGYMSPEYAMD 703
Query: 719 GQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMD----- 773
G FS KSD+FSFGV++LEI+SGKKN GFY +LLGYAW+ W E K L+++D
Sbjct: 704 GIFSVKSDVFSFGVLVLEIVSGKKNRGFYNSNQDNNLLGYAWRNWKEGKGLEILDPFIVD 763
Query: 774 -LSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPTFFTRK 832
S A+ ++ +RC +GLLCVQ+ +DRP MS+VV+ML SET T+P PK P + +
Sbjct: 764 SSSSPSAFRPHEVLRCIQIGLLCVQERAEDRPVMSSVVVMLRSETETIPQPKPPGYCVGR 823
Query: 833 DLSSTASSS 841
T SS+
Sbjct: 824 SPFETDSST 832
>gb|AAP92126.1| receptor-like protein kinase ARK1 [Oryza sativa (japonica
cultivar-group)]
Length = 835
Score = 557 bits (1435), Expect = e-157
Identities = 356/873 (40%), Positives = 487/873 (55%), Gaps = 97/873 (11%)
Query: 19 CWLLCFSQL------CFAGDTLNVGQEITGNGTVLVSAAKKFELGFFSPDLNVTGGKGRY 72
C+LL F + C A DT+ G+ + N T++ F LGFF+ G Y
Sbjct: 9 CYLLLFVVVVVLTGSCRARDTVVPGRPLAANETLVSGGDANFVLGFFTRP----GANSTY 64
Query: 73 LGIWYYREEGSGLSPVVWVANRDNP----VADDSIGVFRIADDGNLVVLDTSGIRYWSAS 128
+G+WY + + VVWVANR++P VAD+ ++ G L ++ + WS +
Sbjct: 65 VGVWYNKVS---VRTVVWVANREDPLPGDVADNPDATLSVSPTGTLAIVAGNSTVVWSVT 121
Query: 129 NLTNSSSATNRSVKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKMD------KTLE 182
+S T R +MDSGNLV+ D G W+ F++PTDT LP M++ +
Sbjct: 122 PAAKLASPTAR---IMDSGNLVIADGAGGGVAWQGFDYPTDTLLPEMRLGVDYVKGRNRT 178
Query: 183 LTCWKSLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPDDI--SN 240
LT WKS SDP G MD + + I N + W+S DGV PD + S
Sbjct: 179 LTAWKSPSDPSPGPVVMAMDTSGDPQVFIWNGAEKVWRSGPW-DGVQFT-GVPDTVTYSG 236
Query: 241 DVYNLLTNFKELK------NKTVSSYDNTRLLLNSTGVIKVLYRVNFQSDI----VWWYQ 290
++ + N KE+ N ++ S RL LNSTG +L R + ++WY
Sbjct: 237 FTFSFINNAKEVTYSFQVHNVSIIS----RLGLNSTGSYGLLQRSTWVEAAGTWNLYWYA 292
Query: 291 PRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFGRRSP----LNDYTVGGDTSSLLCTRKST 346
P+ C + CG C+ +N +C+CL GF +SP L D G C R +
Sbjct: 293 PKDQCDEVSPCGANGVCDTNNLPVCSCLRGFTPKSPEAWALRDGRAG-------CVRSTP 345
Query: 347 -SCGANTNTFLNLTMMKIGSPDIKVSAQDEN----ECKFRCISMCSQTQCQACSYVPIPV 401
C T+ F+ + K+ PD + S D +C+ C+ CS C A + +
Sbjct: 346 LDCQNGTDGFVAVEHAKV--PDTERSVVDLGLSLEQCRKACLMNCS---CTAYASANVSG 400
Query: 402 QQRGLNLSP-CWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILG 460
RG C +WT LT L+ G D LFVR+A +D+ ++ + +++++
Sbjct: 401 GGRGHGAGTGCVMWTTGLTDLRVYPEFGQD--LFVRLAAADLGLTSKSNKARVIIAIVVS 458
Query: 461 IALPGVVILACIC-ILAYVCRRKIALKLKQESESI-LRQRGRFYDSERHVKDLIDKEGLE 518
I+ V L+ + L + ++K A K S R GR Y+ H D
Sbjct: 459 IS--SVTFLSVLAGFLVWTRKKKRARKTGSSKWSGGSRSTGRRYEGSSHHDD-------- 508
Query: 519 EKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQ 578
+E+P FD +I ATD FS NKLG GG+GPVYKGKL+ G+EIAVK LS S Q
Sbjct: 509 -----DLELPIFDLGTIAAATDGFSINNKLGEGGFGPVYKGKLEDGQEIAVKTLSKTSVQ 563
Query: 579 GIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDW 638
G+ EFKNEV+LIAKLQHRNLVRL G+ I G E+IL+YEYM NKSLD F+F
Sbjct: 564 GLDEFKNEVMLIAKLQHRNLVRLLGFSISGQERILVYEYMANKSLDYFLF---------- 613
Query: 639 QMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETE 698
R+ I+ GI RGLLYLHQDSR R+IHRDLK SN+LLD EM PKISDFG+AR+FG +ETE
Sbjct: 614 -ARYRIIEGITRGLLYLHQDSRYRIIHRDLKASNVLLDKEMTPKISDFGMARMFGSEETE 672
Query: 699 ANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGY 758
NT++VVGTYGYMSPEYA+DG FS KSD+FSFGV+LLEIISG++N G Y Y L+LLG+
Sbjct: 673 INTRKVVGTYGYMSPEYAMDGVFSVKSDVFSFGVLLLEIISGRRNRGVYSYSNHLNLLGH 732
Query: 759 AWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIML-DSET 817
AW LW E K L+L D ++ ++++++ ++C VGLLCVQ+ PDDRP MS V++ML ++
Sbjct: 733 AWSLWNEGKSLELADETMNGSFDSDEVLKCIRVGLLCVQENPDDRPLMSQVLLMLATTDA 792
Query: 818 ATLPTPKQPTFFTRKDLSSTASSSLQFDSSIVE 850
TLPTPKQP F R+ L T +SS + D SI +
Sbjct: 793 TTLPTPKQPGFAARRILMETDTSSSKPDCSIFD 825
>gb|AAM90694.1| S-locus receptor-like kinase RLK14 [Oryza sativa]
Length = 813
Score = 557 bits (1435), Expect = e-157
Identities = 337/824 (40%), Positives = 465/824 (55%), Gaps = 59/824 (7%)
Query: 21 LLCFSQLCFAGDTLNVGQEITGNGTVLVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYRE 80
L+ LC + D L + + G +L+S F LGFFSP Y+GIWY++
Sbjct: 11 LVFLISLCKSDDQLTPAKPLHP-GDMLISDGGVFALGFFSP---TKSNATLYVGIWYHKI 66
Query: 81 EGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASN--LTNSSSATN 138
VVWVANRDNP+ S + I++ +LV+ ++ G W A N T S AT
Sbjct: 67 PNR---TVVWVANRDNPITAPSSAMLFISNSSDLVLSESGGHTLWEARNNITTGGSGAT- 122
Query: 139 RSVKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKM------DKTLELTCWKSLSDP 192
V L++SGNLVL + + LW+SF+H TDT LPGMK+ + WK DP
Sbjct: 123 --VVLLNSGNLVLRSPNHTI-LWQSFDHLTDTILPGMKLLLKYNGQVAQRIVSWKGPDDP 179
Query: 193 GRGNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPDDISNDVYNLLTNFKEL 252
GNF+ D + + + N YW+S +++ + S ++ E+
Sbjct: 180 STGNFSLSGDPNSDFQVLVWNGTSPYWRSGAWNGALVSATFQSNTSSVTYQTIINKGNEI 239
Query: 253 KNKTVSSYDNT--RLLLNSTGVIKVL-YRVNFQSDIVWWYQPRTTCLTYNVCGNFSSCND 309
S D+ RL+L+ TG IK+L + N + V + P TC Y CG F C+
Sbjct: 240 YMMYSVSDDSPSMRLMLDYTGTIKMLIWNSNLFAWSVLFSNPSYTCERYASCGPFGYCDA 299
Query: 310 -DNDKLCTCLPGFGRRSPLNDYTVGGDTSSLLCTRKSTSCGANTNTFLNLTMMKIGSPDI 368
+ C CL GF G S C RK + ++FL L MK +
Sbjct: 300 AEAFPTCKCLDGF---------KPDGLNISRGCVRKEQMKCSYGDSFLTLPGMKTPDKFL 350
Query: 369 KVSAQDENECKFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIWTQNLTTLKEEYLGG 428
+ + +EC C CS C A +Y + + S C +W L L + GG
Sbjct: 351 YIRNRSLDECMEECRHNCS---CTAYAYANLSTASMMGDTSRCLVWMGELLDLAKVTGGG 407
Query: 429 DDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGVVILACICILAYVCRRKIALKLK 488
++ L++R + PT + ++L + + ++IL CIC L ++C+ + + K
Sbjct: 408 EN--LYLR-----LPSPTAVKKETDVVKIVLPV-VASLLILTCIC-LVWICKSRGKQRSK 458
Query: 489 QESESILRQRGRFYDSERHVKDLIDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKL 548
+ I+ Q Y S + E E ++ P+ FE +++AT+ FS N L
Sbjct: 459 EIQNKIMVQ----YLSASN-----------ELGAEDVDFPFIGFEEVVIATNNFSSYNML 503
Query: 549 GRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKG 608
G+GG+G VYKG L+GG+E+AVKRLS S QGI+EF+NEVVLIA+LQHRNLV+L G CI
Sbjct: 504 GKGGFGKVYKGILEGGKEVAVKRLSKGSGQGIEEFRNEVVLIARLQHRNLVKLVGCCIHE 563
Query: 609 DEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDL 668
DEK+LIYEY+PNKSLDAF+FD T+ +LDW RF I+ G+ARGLLYLHQDSRL +IHRDL
Sbjct: 564 DEKLLIYEYLPNKSLDAFLFDATRKTVLDWPNRFKIIKGVARGLLYLHQDSRLTIIHRDL 623
Query: 669 KTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIF 728
K NILLD EM PKISDFG+ARIFGG + +ANT RVVGTYGYMSPEYA++G FS KSDI+
Sbjct: 624 KAGNILLDAEMSPKISDFGMARIFGGNQQQANTTRVVGTYGYMSPEYAMEGIFSVKSDIY 683
Query: 729 SFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRC 788
SFG++LLEIISG + + + G +L+ Y+W LW + DL+D S+ E+ ++ +RC
Sbjct: 684 SFGILLLEIISGFRISSPHLIMGFPNLIAYSWSLWKDGNARDLVDSSVVESCPLHEVLRC 743
Query: 789 THVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPTFFTRK 832
H+ LLC+QD PDDRP MS+VV ML++ TA LP PKQP FF K
Sbjct: 744 IHIALLCIQDHPDDRPLMSSVVFMLENNTAPLPQPKQPIFFVHK 787
>ref|NP_176755.1| S-receptor protein kinase, putative [Arabidopsis thaliana]
gi|2129703|pir||S70769 S-receptor kinase (EC 2.7.1.-)
Ark1 precursor - Arabidopsis thaliana
gi|166692|gb|AAA32786.1| receptor kinase
gi|445123|prf||1908429A receptor kinase
Length = 843
Score = 556 bits (1434), Expect = e-157
Identities = 345/859 (40%), Positives = 499/859 (57%), Gaps = 70/859 (8%)
Query: 15 FHMHCWLLCFSQLCFAGDTLNVGQEIT-GNGTVLVSAAKKFELGFFSPDLNVTGGKGRYL 73
F + L+ F + +TL+ + +T + ++S ++ FELGFF+P YL
Sbjct: 11 FFIFLILILFLAFSVSPNTLSATESLTISSNKTIISPSQIFELGFFNP----ASSSRWYL 66
Query: 74 GIWYYREEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNS 133
GIWY + + VWVANRDNP++ + G +I+ + NLV+ D S WS +N+T
Sbjct: 67 GIWY---KIIPIRTYVWVANRDNPLSSSN-GTLKISGN-NLVIFDQSDRPVWS-TNITGG 120
Query: 134 SSATNRSVKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKMDKTLE------LTCWK 187
+ + +L+D+GN +L D + + LW+SF+ PTDT L MK+ + L WK
Sbjct: 121 DVRSPVAAELLDNGNFLLRDSNNRL-LWQSFDFPTDTLLAEMKLGWDQKTGFNRILRSWK 179
Query: 188 SLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPDDISNDVYNLLT 247
+ DP G F+ K++ F I ++ + ++S M S P I D ++
Sbjct: 180 TTDDPSSGEFSTKLETSEFPEFYICSKESILYRSGPWNG--MRFSSVPGTIQVDY--MVY 235
Query: 248 NFKELKNKTVSSYD------NTRLLLNSTGVIKVLYRVNFQSDIVW---WYQPRTTCLTY 298
NF K + SY +RL LNS G+++ L F++ W WY P+ C Y
Sbjct: 236 NFTASKEEVTYSYRINKTNLYSRLYLNSAGLLQRL--TWFETTQSWKQLWYSPKDLCDNY 293
Query: 299 NVCGNFSSCNDDNDKLCTCLPGFGRRSPLNDYTVGGDTSSLLCTRKSTSCGANTNTFLNL 358
VCGNF C+ ++ C C+ GF P+N+ S C RK+ + F L
Sbjct: 294 KVCGNFGYCDSNSLPNCYCIKGF---KPVNEQAWDLRDGSAGCMRKTRLSCDGRDGFTRL 350
Query: 359 TMMKIGSPDIKVSAQDEN----ECKFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIW 414
MK+ PD + D CK RC+ C+ T ++ ++ G S C IW
Sbjct: 351 KRMKL--PDTTATIVDREIGLKVCKERCLEDCNCT-----AFANADIRNGG---SGCVIW 400
Query: 415 TQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGVVILACICI 474
T+ + ++ GG D L+VR+A +++E+ K I+G ++ ++L +
Sbjct: 401 TREILDMRNYAKGGQD--LYVRLAAAELEDKRIKNEK------IIGSSIGVSILLLLSFV 452
Query: 475 LAYVCRRKIALKLKQESESILRQRGRFYDSERHVKDLI-DKEGL--EEKDNEGIEVPYFD 531
+ + +RK + ++ ++ + R + + + D++ + G +EK +E +E+P +
Sbjct: 453 IFHFWKRKQKRSITIQTPNVDQVRSQ----DSLINDVVVSRRGYTSKEKKSEYLELPLLE 508
Query: 532 FESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIA 591
E++ AT+ FS+ NKLG+GG+G VYKG+L G+EIAVKRLS +SSQG EF NEV LIA
Sbjct: 509 LEALATATNNFSNDNKLGQGGFGIVYKGRLLDGKEIAVKRLSKMSSQGTDEFMNEVRLIA 568
Query: 592 KLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARG 651
KLQH NLVRL G C+ EK+LIYEY+ N SLD+ +FD T+S+ L+WQ RFDI+ GIARG
Sbjct: 569 KLQHINLVRLLGCCVDKGEKMLIYEYLENLSLDSHLFDQTRSSNLNWQKRFDIINGIARG 628
Query: 652 LLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYM 711
LLYLHQDSR R+IHRDLK SN+LLD M PKISDFG+ARIFG +ETEANT+RVVGTYGYM
Sbjct: 629 LLYLHQDSRCRIIHRDLKASNVLLDKNMTPKISDFGMARIFGREETEANTRRVVGTYGYM 688
Query: 712 SPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDL 771
SPEYA+DG FS KSD+FSFGV+LLEIISGK+N GFY L+LLG+ W+ W E L++
Sbjct: 689 SPEYAMDGIFSMKSDVFSFGVLLLEIISGKRNKGFYNSNRDLNLLGFVWRHWKEGNELEI 748
Query: 772 MDL----SLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPT 827
+D SL + ++ +RC +GLLCVQ+ +DRP MS+V++ML SET +P PK+P
Sbjct: 749 VDPINIDSLSSKFPTHEILRCIQIGLLCVQERAEDRPVMSSVMVMLGSETTAIPQPKRPG 808
Query: 828 F-FTRKDLSSTASSSLQFD 845
F R L + +SSS Q D
Sbjct: 809 FCIGRSPLEADSSSSTQRD 827
>gb|AAF23832.1| F1E22.15 [Arabidopsis thaliana]
Length = 1662
Score = 556 bits (1434), Expect = e-157
Identities = 345/859 (40%), Positives = 499/859 (57%), Gaps = 70/859 (8%)
Query: 15 FHMHCWLLCFSQLCFAGDTLNVGQEIT-GNGTVLVSAAKKFELGFFSPDLNVTGGKGRYL 73
F + L+ F + +TL+ + +T + ++S ++ FELGFF+P YL
Sbjct: 11 FFIFLILILFLAFSVSPNTLSATESLTISSNKTIISPSQIFELGFFNP----ASSSRWYL 66
Query: 74 GIWYYREEGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNS 133
GIWY + + VWVANRDNP++ + G +I+ + NLV+ D S WS +N+T
Sbjct: 67 GIWY---KIIPIRTYVWVANRDNPLSSSN-GTLKISGN-NLVIFDQSDRPVWS-TNITGG 120
Query: 134 SSATNRSVKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKMDKTLE------LTCWK 187
+ + +L+D+GN +L D + + LW+SF+ PTDT L MK+ + L WK
Sbjct: 121 DVRSPVAAELLDNGNFLLRDSNNRL-LWQSFDFPTDTLLAEMKLGWDQKTGFNRILRSWK 179
Query: 188 SLSDPGRGNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPDDISNDVYNLLT 247
+ DP G F+ K++ F I ++ + ++S M S P I D ++
Sbjct: 180 TTDDPSSGEFSTKLETSEFPEFYICSKESILYRSGPWNG--MRFSSVPGTIQVDY--MVY 235
Query: 248 NFKELKNKTVSSYD------NTRLLLNSTGVIKVLYRVNFQSDIVW---WYQPRTTCLTY 298
NF K + SY +RL LNS G+++ L F++ W WY P+ C Y
Sbjct: 236 NFTASKEEVTYSYRINKTNLYSRLYLNSAGLLQRL--TWFETTQSWKQLWYSPKDLCDNY 293
Query: 299 NVCGNFSSCNDDNDKLCTCLPGFGRRSPLNDYTVGGDTSSLLCTRKSTSCGANTNTFLNL 358
VCGNF C+ ++ C C+ GF P+N+ S C RK+ + F L
Sbjct: 294 KVCGNFGYCDSNSLPNCYCIKGF---KPVNEQAWDLRDGSAGCMRKTRLSCDGRDGFTRL 350
Query: 359 TMMKIGSPDIKVSAQDEN----ECKFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIW 414
MK+ PD + D CK RC+ C+ T ++ ++ G S C IW
Sbjct: 351 KRMKL--PDTTATIVDREIGLKVCKERCLEDCNCT-----AFANADIRNGG---SGCVIW 400
Query: 415 TQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGVVILACICI 474
T+ + ++ GG D L+VR+A +++E+ K I+G ++ ++L +
Sbjct: 401 TREILDMRNYAKGGQD--LYVRLAAAELEDKRIKNEK------IIGSSIGVSILLLLSFV 452
Query: 475 LAYVCRRKIALKLKQESESILRQRGRFYDSERHVKDLI-DKEGL--EEKDNEGIEVPYFD 531
+ + +RK + ++ ++ + R + + + D++ + G +EK +E +E+P +
Sbjct: 453 IFHFWKRKQKRSITIQTPNVDQVRSQ----DSLINDVVVSRRGYTSKEKKSEYLELPLLE 508
Query: 532 FESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIA 591
E++ AT+ FS+ NKLG+GG+G VYKG+L G+EIAVKRLS +SSQG EF NEV LIA
Sbjct: 509 LEALATATNNFSNDNKLGQGGFGIVYKGRLLDGKEIAVKRLSKMSSQGTDEFMNEVRLIA 568
Query: 592 KLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARG 651
KLQH NLVRL G C+ EK+LIYEY+ N SLD+ +FD T+S+ L+WQ RFDI+ GIARG
Sbjct: 569 KLQHINLVRLLGCCVDKGEKMLIYEYLENLSLDSHLFDQTRSSNLNWQKRFDIINGIARG 628
Query: 652 LLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYM 711
LLYLHQDSR R+IHRDLK SN+LLD M PKISDFG+ARIFG +ETEANT+RVVGTYGYM
Sbjct: 629 LLYLHQDSRCRIIHRDLKASNVLLDKNMTPKISDFGMARIFGREETEANTRRVVGTYGYM 688
Query: 712 SPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDL 771
SPEYA+DG FS KSD+FSFGV+LLEIISGK+N GFY L+LLG+ W+ W E L++
Sbjct: 689 SPEYAMDGIFSMKSDVFSFGVLLLEIISGKRNKGFYNSNRDLNLLGFVWRHWKEGNELEI 748
Query: 772 MDL----SLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPT 827
+D SL + ++ +RC +GLLCVQ+ +DRP MS+V++ML SET +P PK+P
Sbjct: 749 VDPINIDSLSSKFPTHEILRCIQIGLLCVQERAEDRPVMSSVMVMLGSETTAIPQPKRPG 808
Query: 828 F-FTRKDLSSTASSSLQFD 845
F R L + +SSS Q D
Sbjct: 809 FCIGRSPLEADSSSSTQRD 827
Score = 545 bits (1404), Expect = e-153
Identities = 343/832 (41%), Positives = 489/832 (58%), Gaps = 64/832 (7%)
Query: 40 ITGNGTVLVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYREEGSGLSPVVWVANRDNPVA 99
I+ N T+ +S ++ FELGFF+PD YLGIWY + + VWVANRDNP++
Sbjct: 853 ISSNKTI-ISPSQIFELGFFNPD----SSSRWYLGIWY---KIIPIRTYVWVANRDNPLS 904
Query: 100 DDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNSSSATNRSVKLMDSGNLVLLDEHVGMK 159
+ G +I+D+ NLV+ D S WS +N+T + + +L+D GN VL D
Sbjct: 905 SSN-GTLKISDN-NLVIFDQSDRPVWS-TNITGGDVRSPVAAELLDYGNFVLRDSKNNKP 961
Query: 160 ---LWESFEHPTDTFLPGMKMDKTLE-------LTCWKSLSDPGRGNFTFKMDKKWENRF 209
LW+SF+ PTDT L MKM + L WK+ DP G+F+ K+ F
Sbjct: 962 SGFLWQSFDFPTDTLLSDMKMGWDNKSGGFNRILRSWKTTDDPSSGDFSTKLRTSGFPEF 1021
Query: 210 AILNQGQLYWQSEEQGDGVMNPESNPDDISNDVYNLLTNFKELKNKTVSSYDNTR----- 264
I N+ + ++S G + N S+ + Y + +F E + V SY +
Sbjct: 1022 YIYNKESITYRS---GPWLGNRFSSVPGMKPVDY-IDNSFTENNQQVVYSYRVNKTNIYS 1077
Query: 265 -LLLNSTGVIKVL-YRVNFQSDIVWWYQPRTTCLTYNVCGNFSSCNDDNDKLCTCLPGFG 322
L L+STG+++ L + QS WY P+ C Y CGN+ C+ + +C C+ GF
Sbjct: 1078 ILSLSSTGLLQRLTWMEAAQSWKQLWYSPKDLCDNYKECGNYGYCDANTSPICNCIKGF- 1136
Query: 323 RRSPLNDYTVGGDTSSLLCTRKSTSCGANTNTFLNLTMMKIGSPDIKVSAQDEN----EC 378
P+N+ D S+ C RK+ + F+ L M++ PD ++ D+ EC
Sbjct: 1137 --EPMNEQAALRD-DSVGCVRKTKLSCDGRDGFVRLKKMRL--PDTTETSVDKGIGLKEC 1191
Query: 379 KFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVA 438
+ RC+ C+ T ++ ++ G S C IW+ L ++ GG D L+VRVA
Sbjct: 1192 EERCLKGCNCT-----AFANTDIRNGG---SGCVIWSGGLFDIRNYAKGGQD--LYVRVA 1241
Query: 439 KSDIEEPTRKGNPKSTLSLILGIALPGVVILACICILAYVCRRKIALKLKQESESILRQR 498
D+E+ K K + +G+++ +++L+ I + ++K ++ ++ ++R +
Sbjct: 1242 AGDLEDKRIKS--KKIIGSSIGVSI--LLLLSFIIFHFWKRKQKRSITIQTPIVDLVRSQ 1297
Query: 499 GRFYDSERHVKDLIDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYK 558
+ VK E K + +E+P +++++ +AT+ FS NKLG+GG+G VYK
Sbjct: 1298 DSLMNEL--VKASRSYTSKENK-TDYLELPLMEWKALAMATNNFSTDNKLGQGGFGIVYK 1354
Query: 559 GKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYM 618
G L G+EIAVKRLS +SSQG EF NEV LIAKLQH NLVRL G C+ EK+LIYEY+
Sbjct: 1355 GMLLDGKEIAVKRLSKMSSQGTDEFMNEVRLIAKLQHINLVRLLGCCVDKGEKMLIYEYL 1414
Query: 619 PNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGE 678
N SLD+ +FD T+S+ L+WQ RFDI+ GIARGLLYLHQDSR R+IHRDLK SN+LLD
Sbjct: 1415 ENLSLDSHLFDQTRSSNLNWQKRFDIINGIARGLLYLHQDSRCRIIHRDLKASNVLLDKN 1474
Query: 679 MQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEII 738
M PKISDFG+ARIFG +ETEANT+RVVGTYGYMSPEYA+DG FS KSD+FSFGV+LLEII
Sbjct: 1475 MTPKISDFGMARIFGREETEANTRRVVGTYGYMSPEYAMDGIFSMKSDVFSFGVLLLEII 1534
Query: 739 SGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDL----SLGEAYNANQFIRCTHVGLL 794
SGK+N GFY L+LLG+ W+ W E K L+++D +L + ++ +RC +GLL
Sbjct: 1535 SGKRNKGFYNSNRDLNLLGFVWRHWKEGKELEIVDPINIDALSSEFPTHEILRCIQIGLL 1594
Query: 795 CVQDEPDDRPNMSNVVIMLDSETATLPTPKQPTFFT-RKDLSSTASSSLQFD 845
CVQ+ +DRP MS+V++ML SET +P PK+P F R L +SSS Q D
Sbjct: 1595 CVQERAEDRPVMSSVMVMLGSETTAIPQPKRPGFCVGRSSLEVDSSSSTQRD 1646
>emb|CAI44641.1| OSJNBb0015D13.18 [Oryza sativa (japonica cultivar-group)]
Length = 3307
Score = 555 bits (1431), Expect = e-156
Identities = 332/801 (41%), Positives = 456/801 (56%), Gaps = 58/801 (7%)
Query: 44 GTVLVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYREEGSGLSPVVWVANRDNPVADDSI 103
G +L+S F LGFFSP Y+GIWY++ VVWVANRDNP+ S
Sbjct: 2527 GDMLISDGGVFALGFFSP---TNSNATLYVGIWYHKIPNR---TVVWVANRDNPITAPSS 2580
Query: 104 GVFRIADDGNLVVLDTSGIRYWSASN--LTNSSSATNRSVKLMDSGNLVLLDEHVGMKLW 161
+ I++ +LV+ ++ G W A N T S AT V L++SGNLVL + + LW
Sbjct: 2581 AMLFISNSSDLVLSESGGHTLWEARNNITTGGSGAT---VVLLNSGNLVLRSPNHTI-LW 2636
Query: 162 ESFEHPTDTFLPGMKM------DKTLELTCWKSLSDPGRGNFTFKMDKKWENRFAILNQG 215
+SF+H TDT LPGMK+ + WK DP GNF+ D + + + N
Sbjct: 2637 QSFDHLTDTILPGMKLLLKYNGQVAQRIVSWKGPDDPSTGNFSLSGDPNSDFQVLVWNGT 2696
Query: 216 QLYWQSEEQGDGVMNPESNPDDISNDVYNLLTNFKELKNKTVSSYDNT--RLLLNSTGVI 273
YW+S +++ + S ++ E+ S D+ RL+L+ TG I
Sbjct: 2697 SPYWRSGAWNGALVSAMFQSNTSSVTYQTIINKGNEIYMMYSVSDDSPSMRLMLDYTGTI 2756
Query: 274 KVL-YRVNFQSDIVWWYQPRTTCLTYNVCGNFSSCND-DNDKLCTCLPGFGRRSPLNDYT 331
K+L + N + V + P TC Y CG F C+ + C CL GF
Sbjct: 2757 KMLIWNSNLFAWSVLFSNPSYTCERYASCGPFGYCDAAEAFPTCKCLDGF---------K 2807
Query: 332 VGGDTSSLLCTRKSTSCGANTNTFLNLTMMKIGSPDIKVSAQDENECKFRCISMCSQTQC 391
G S C RK + ++FL L MK + + + +EC C CS C
Sbjct: 2808 PDGLNISRGCVRKEQMKCSYGDSFLTLPGMKTPDKFLYIRNRSLDECMEECRHNCS---C 2864
Query: 392 QACSYVPIPVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNP 451
A +Y + + S C +W L L + GG++ L++R + PT
Sbjct: 2865 TAYAYANLSTASMMGDTSRCLVWMGELLDLAKVTGGGEN--LYLR-----LPSPTAVKKE 2917
Query: 452 KSTLSLILGIALPGVVILACICILAYVCRRKIALKLKQESESILRQRGRFYDSERHVKDL 511
+ ++L + + ++IL CIC L ++C+ + + K+ I+ Q Y S +
Sbjct: 2918 TDVVKIVLPV-VASLLILTCIC-LVWICKSRGKQRSKEIQNKIMVQ----YLSASN---- 2967
Query: 512 IDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKR 571
E E ++ P+ FE +++AT+ FS N LG+GG+G VYKG L+GG+E+AVKR
Sbjct: 2968 -------ELGAEDVDFPFIGFEEVVIATNNFSSYNMLGKGGFGKVYKGILEGGKEVAVKR 3020
Query: 572 LSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPT 631
LS S QGI+EF+NEVVLIA+LQHRNLV+L G CI DEK+LIYEY+PNKSLDAF+FD T
Sbjct: 3021 LSKGSGQGIEEFRNEVVLIARLQHRNLVKLVGCCIHEDEKLLIYEYLPNKSLDAFLFDAT 3080
Query: 632 KSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARI 691
+ +LDW RF I+ G+ARGLLYLHQDSRL +IHRDLK NILLD EM PKISDFG+ARI
Sbjct: 3081 RKTVLDWPNRFKIIKGVARGLLYLHQDSRLTIIHRDLKAGNILLDAEMSPKISDFGMARI 3140
Query: 692 FGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKG 751
FGG + +ANT RVVGTYGYMSPEYA++G FS KSDI+SFG++LLEIISG + + + G
Sbjct: 3141 FGGNQQQANTTRVVGTYGYMSPEYAMEGIFSVKSDIYSFGILLLEIISGFRISSPHLIMG 3200
Query: 752 TLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVI 811
+L+ Y+W LW + DL+D S+ E+ ++ +RC H+ LLC+QD PDDRP MS+VV
Sbjct: 3201 FPNLIAYSWSLWKDGNARDLVDSSVVESCPLHEVLRCIHIALLCIQDHPDDRPLMSSVVF 3260
Query: 812 MLDSETATLPTPKQPTFFTRK 832
ML++ TA LP PKQP FF K
Sbjct: 3261 MLENNTAPLPQPKQPIFFVHK 3281
Score = 515 bits (1326), Expect = e-144
Identities = 325/836 (38%), Positives = 451/836 (53%), Gaps = 103/836 (12%)
Query: 21 LLCFSQLCFAGDTLNVGQEITGNGTVLVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYRE 80
LL C D L + G VL+S + F LGFFSP + +LGIWY+
Sbjct: 1601 LLFLISSCKGDDQLTQANRLISPGDVLISKGRVFALGFFSP---TASNQSFFLGIWYHNI 1657
Query: 81 EGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNSSSATNRS 140
S + VWVANRDNP+ S I++ NLV+ D+ W+ +N+T ++
Sbjct: 1658 SESERT-YVWVANRDNPITTPSFATLAISNSSNLVLSDSGNHTLWT-TNVT-ATGGDGAY 1714
Query: 141 VKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGM------KMDKTLELTCWKSLSDPGR 194
L+DSGNLVL + G +W+SF+HPTDT L GM K + WK DP
Sbjct: 1715 AALLDSGNLVLRLPN-GTTIWQSFDHPTDTLLMGMRFLVSYKAQVAMRCIAWKGPDDPST 1773
Query: 195 GNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPDDISNDVYNLLTNFKELKN 254
G+F+ D + + N + Y + G P + + V++ T+ +
Sbjct: 1774 GDFSISGDPSSNLQIFLWNGTRPYIRFIGFG---------PSSMWSSVFSFSTSL--IYE 1822
Query: 255 KTVSSYDN-------------TRLLLNSTGVIKVLYRVNFQSD---IVWWYQPRTTCLTY 298
+VS+ D RL L+ TG +K L + S +V P C Y
Sbjct: 1823 TSVSTDDEFYIIYTTSDGSPYKRLQLDYTGTLKFLAWNDSASSWTVVVQRPSPTIVCDPY 1882
Query: 299 NVCGNFSSCNDDND-KLCTCLPGFGRRSPLNDYTVGGDTSSLLCTRKST-SCGANTNTFL 356
CG F C+ C CL GF G ++SS C RK C + F+
Sbjct: 1883 ASCGPFGYCDATAAIPRCQCLDGFEPD--------GSNSSSRGCRRKQQLRCRGRDDRFV 1934
Query: 357 NLTMMKIGSPDIKVSAQDENECKFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIWTQ 416
+ MK+ + V + +EC C CS C A +Y + G + + C +W+
Sbjct: 1935 TMAGMKVPDKFLHVRNRSFDECAAECSRNCS---CTAYAYANLT----GADQARCLLWSG 1987
Query: 417 NLTTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGVVILACICILA 476
L +G L++R+A S + + KS + I+ + ++IL CIC LA
Sbjct: 1988 ELADTGRANIG---ENLYLRLADSTVNKK------KSDIPKIVLPVITSLLILMCIC-LA 2037
Query: 477 YVCRRKIALKLKQESESILRQRGRFYDSERHVKDLIDKEGLEEKDNEGIEVPYFDFESIL 536
++C+ + + K+ ++++ R +H+KD E +N+ +E+P+ E I+
Sbjct: 2038 WICKSRGIHRSKE-----IQKKHRL----QHLKDS------SELENDNLELPFICLEDIV 2082
Query: 537 VATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHR 596
AT+ FSD N LG+GG+G VYKG L+GG+EIAVKRLS S QG++EF+NEVVLIAKLQHR
Sbjct: 2083 TATNNFSDHNMLGKGGFGKVYKGVLEGGKEIAVKRLSKGSQQGVEEFRNEVVLIAKLQHR 2142
Query: 597 NLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLH 656
NLVRL YCI DEK+LIYEY+PNKSLD F+FD + ++LDW RF I+ GIARGLLYLH
Sbjct: 2143 NLVRLISYCIHEDEKLLIYEYLPNKSLDTFLFDAKRKSVLDWTTRFMIIKGIARGLLYLH 2202
Query: 657 QDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYA 716
QDSRL +IHRDLK SNILLD M PKISDFG+ARIF G + + NT RVVGTYGYMSPEYA
Sbjct: 2203 QDSRLTIIHRDLKASNILLDTNMSPKISDFGMARIFEGNKQQENTTRVVGTYGYMSPEYA 2262
Query: 717 LDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSL 776
L+G FS KSD +SFGV+LLE+ AW LW + +DL+D S+
Sbjct: 2263 LEGSFSVKSDTYSFGVLLLEL---------------------AWSLWKDGNAMDLVDSSI 2301
Query: 777 GEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETATLPTPKQPTFFTRK 832
E+ ++ +RC + L CVQD+P RP MS++V ML++ETA LPTPK+ + T +
Sbjct: 2302 RESCLLHEVLRCIQIALSCVQDDPTARPLMSSIVFMLENETAALPTPKESAYLTAR 2357
Score = 465 bits (1197), Expect = e-129
Identities = 302/784 (38%), Positives = 419/784 (52%), Gaps = 74/784 (9%)
Query: 21 LLCFSQLCFAGDTLNVGQEITGNGTVLVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYRE 80
LL LC D L +G+ I + +L+S F LGFF P Y+G+W++
Sbjct: 9 LLLSIPLCKTDDQLTLGKPIFPS-EMLISKGGIFALGFFPP---ANFSNSLYVGVWFHNI 64
Query: 81 EGSGLSPVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNSSSATNRS 140
VVWVANRDNP+ S I + +V+ D+ G W+A S S
Sbjct: 65 PQR---TVVWVANRDNPITTPSSATLAITNSSGMVLSDSQGDILWTAK-----ISVIGAS 116
Query: 141 VKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGM------KMDKTLELTCWKSLSDPGR 194
L+D+GN VL + G +W+SF+HPTDT L GM K + LT W+S DP
Sbjct: 117 AVLLDTGNFVLRLAN-GTDIWQSFDHPTDTILAGMMFLMSYKSEIIGRLTAWRSHDDPST 175
Query: 195 GNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPDDISNDVYNLLTNF--KEL 252
G+F+F +D + + N + Y ++ + ++ P + S +Y L + K
Sbjct: 176 GDFSFSLDPSSDLQGMTWNGTKPYCRNGVRTSVTVSGAQYPSNSSLFMYQTLIDSGNKLY 235
Query: 253 KNKTVS-SYDNTRLLLNSTGVIKVLYRVNFQSDIVWWYQPRT--TCLTYNVCGNFSSCND 309
+ TVS S TRL L+STG + L N S + +Q +C Y CG F C+
Sbjct: 236 YSYTVSDSSIYTRLTLDSTGTMMFLSWDNSSSSWMLIFQRPAAGSCEVYGSCGPFGYCDF 295
Query: 310 DND-KLCTCLPGFGRRSPLNDYTVGGDTSSLLCTRKST-SCGANTNTFLNLTMMKIGSPD 367
C CL GF P S C RK CG + F++L MK+
Sbjct: 296 TGAVPACRCLDGFEPVDP--------SISQSGCRRKEELRCGEGGHRFVSLPDMKVPDKF 347
Query: 368 IKVSAQDENECKFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIWTQNLTTLKEEYLG 427
+++ + ++C C S CS C+A +Y + + S C +WT L +++
Sbjct: 348 LQIRNRSFDQCAAECSSNCS---CKAYAYANLSSGGTMADPSRCLVWTGELVDSEKKASL 404
Query: 428 GDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGVVILACICILAYVCRRKIALKL 487
G++ L++R+A+ + G L +++ I + +++L CI +L ++C+ +
Sbjct: 405 GEN--LYLRLAEPPV------GKKNRLLKIVVPITVC-MLLLTCI-VLTWICKHRGKQNK 454
Query: 488 KQESESILRQRGRFYDSERHVKDLIDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANK 547
+ + +L G E E ++ P+ F I+ ATD F ++N
Sbjct: 455 EIQKRLMLEYPGTS----------------NELGGENVKFPFISFGDIVAATDNFCESNL 498
Query: 548 LGRGGYGPVYK-----------GKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHR 596
LGRGG+G VYK G L+GG E+AVKRL+ S QGI+EF+NEVVLIAKLQHR
Sbjct: 499 LGRGGFGKVYKRFPIYIDDNMKGILEGGTEVAVKRLNEGSGQGIEEFRNEVVLIAKLQHR 558
Query: 597 NLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLH 656
NLVRL G CI DEK+LIYEY+PNKSLDAF+FD T+ +LDW RF I+ GIA+GLLYLH
Sbjct: 559 NLVRLLGCCIHEDEKLLIYEYLPNKSLDAFLFDATRKYVLDWPTRFKIIKGIAKGLLYLH 618
Query: 657 QDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYA 716
QDSRL +IHRDLK SNILLD EM PKISDFG+ARIF G + +ANT RVVGTYGYMSPEY
Sbjct: 619 QDSRLTIIHRDLKASNILLDTEMNPKISDFGIARIFHGNQQQANTTRVVGTYGYMSPEYV 678
Query: 717 LDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSL 776
L G FS KSD +SFGV+LLEI+SG K + SL YAW+LW + +L+D
Sbjct: 679 LGGAFSVKSDTYSFGVLLLEIVSGLKISSSKLTPNFFSLTAYAWRLWKDGNATELLDKFF 738
Query: 777 GEAY 780
++Y
Sbjct: 739 VDSY 742
Score = 300 bits (769), Expect = 1e-79
Identities = 237/769 (30%), Positives = 346/769 (44%), Gaps = 143/769 (18%)
Query: 28 CFAGDTLNVGQE-ITGNGTVLVSAAKKFELGFFSPDLNVTGGKGRYLGIWYYREEGSGLS 86
C + D L + I G L+S F +GFFS + YLGIWY
Sbjct: 863 CQSDDRLTPAKPLIFPGGDKLISDGGVFAVGFFSLTTTNSTPSLLYLGIWY---NNIPER 919
Query: 87 PVVWVANRDNPVADDSIGVFRIADDGNLVVLDTSGIRYWSASNLTNSSSATNRSVKLMDS 146
VWVANRDNP+ + + + + LV+ D+ G +A+ +T + L ++
Sbjct: 920 TYVWVANRDNPITTHTARL-AVTNTSGLVLSDSKGT---TANTVTIGGGGA--TAVLQNT 973
Query: 147 GNLVLLDEHVGMKLWESFEHPTDTFLPGMKMDKTLELTCWKSLSDPGRGNFTFKMDK-KW 205
GN VL + K + + + W+ DP F+ D +W
Sbjct: 974 GNFVL------------------RYGRTYKNHEAVRVVAWRGRRDPSTCEFSLSGDPDQW 1015
Query: 206 ENRFAILNQGQLYWQSEEQGDGVMNPESNPDDISNDVYNLLTNFKELKNKTVSSYDN--T 263
I + W+S GV N + ++ +++ + + E ++ D T
Sbjct: 1016 GLHIVIWHGASPSWRS-----GVWNG-ATATGLTRYIWSQIVDNGEEIYAIYNAADGILT 1069
Query: 264 RLLLNSTGVIKVLYRVNFQSDIVW---WYQPRTTCLTYNVCGNFSSCNDDND-KLCTCLP 319
L+ TG V +R W + +P CL Y CG F C+ + C CL
Sbjct: 1070 HWKLDYTG--NVSFRAWNNVSSTWTSPFERPGHGCLHYGACGPFGYCDITGSFQECKCLD 1127
Query: 320 GFGRRSPLNDYTVGGDTSSLLCTRKSTSCGANTNTFLNLTMMKIGSPDIKVSAQDENECK 379
GF P + +++ SS C RK + F L MK+ + + + EC
Sbjct: 1128 GF---EPADGFSLN---SSRGCRRKEELRCGGQDHFFTLPGMKVPDKFLYIRNRTFEECA 1181
Query: 380 FRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIWTQNLTTLKEEYLGGDDRKLFVRVAK 439
C CS C A +Y + + S C +W L L E L++R+A
Sbjct: 1182 DECDRNCS---CTAYAYANLRTILTTGDPSRCLVWMGEL--LDSEKASAVGENLYLRLA- 1235
Query: 440 SDIEEPTRKGNPKSTLSLILGIALPGV----VILACICILAYVCRRKIALKLKQESESIL 495
G+P I+ I LP + ++ AC C++ C + + K+ +L
Sbjct: 1236 ---------GSPAVNNKNIVKIVLPAIACLLILTACSCVVLCKCESRGIRRNKE----VL 1282
Query: 496 RQRGRFYDSERHVKDLIDKEGLEEKDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGP 555
++ Y S H + ++ +E P +E + AT+ F + N LG+GG+G
Sbjct: 1283 KKTELGYLSAFH-----------DSWDQNLEFPDISYEDLTSATNGFHETNMLGKGGFG- 1330
Query: 556 VYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIY 615
+H+NLVRL G CI GDEK+LIY
Sbjct: 1331 --------------------------------------KHKNLVRLLGCCIHGDEKLLIY 1352
Query: 616 EYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILL 675
EY+PNKSLD F+FD +++DWQ RF+I+ G+ARGLLYLHQDSR+ +IHRDLKTSNILL
Sbjct: 1353 EYLPNKSLDKFLFDHAMKSVIDWQTRFNIIKGVARGLLYLHQDSRMMIIHRDLKTSNILL 1412
Query: 676 DGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLL 735
D EM PKISDFG+ARIFG E +A+T+RVVGTYGYM+PEYA++G FS KSD +SFGV+LL
Sbjct: 1413 DAEMNPKISDFGMARIFGNSEQQASTRRVVGTYGYMAPEYAMEGIFSVKSDTYSFGVLLL 1472
Query: 736 EIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGEAYNANQ 784
EI AW LW + +D + E+ N+
Sbjct: 1473 EI---------------------AWNLWKDGMAEAFVDKMVLESCLLNE 1500
>ref|XP_478649.1| putative S-receptor kinase KIK1 precursor [Oryza sativa (japonica
cultivar-group)] gi|28971965|dbj|BAC65366.1| putative
S-receptor kinase KIK1 precursor [Oryza sativa (japonica
cultivar-group)] gi|50510068|dbj|BAD30706.1| putative
S-receptor kinase KIK1 precursor [Oryza sativa (japonica
cultivar-group)]
Length = 865
Score = 553 bits (1426), Expect = e-156
Identities = 347/862 (40%), Positives = 499/862 (57%), Gaps = 100/862 (11%)
Query: 30 AGDTLNVGQEITGNGTVLVSAAKKFELGFFSPDLNVTGGKGR-YLGIWYYREEGSGLSPV 88
A DTL+ GQ + N +LVSA F++GFF+P G G+ YLG+ Y S + V
Sbjct: 28 AADTLSQGQSLGAND-MLVSANGTFKVGFFTP---AGGDPGKVYLGVMYAT---SNVQTV 80
Query: 89 VWVANRDNPVADDS-IGVFRIADDGNLVVLDTSGIRYWSASNLTNSSSA--TNRSVKLMD 145
+WVANRD PV + + G L+V + + + TN+S+A + ++ + D
Sbjct: 81 MWVANRDAPVRTAAGAASATVTGSGELLVKEGDRVAW-----RTNASAAGRSKHTLTIRD 135
Query: 146 SGNLVLL-DEHVGMKL-WESFEHPTDTFLPGMKM-------DKTLELTCWKSLSDPGRGN 196
GNLV+ + G + WESF HPTDTF+PGM++ D+TL T W+S +DP G+
Sbjct: 136 DGNLVISGSDAAGTDVEWESFHHPTDTFVPGMEIALRQTNGDRTL-YTSWRSDADPATGD 194
Query: 197 FTFKMDKK-----WENRFAILNQGQLYWQSEEQGDGVM-------------NPESNPDDI 238
FT +D W ++ + YW+S + G +P I
Sbjct: 195 FTLGLDASAQLYIWRSQGG---KNSTYWRSGQWASGNFVGIPWRALYVYGFKLNGDPPPI 251
Query: 239 SNDVYNLLTNFKELKNKTVSSYDNTRLLLNSTGVIKVLYRVNFQSDIVWWYQPRTTCLTY 298
+ D+ T F S Y R +L GV + + W QP C Y
Sbjct: 252 AGDMSIAFTPFNS------SLY---RFVLRPNGVETCYMLLGSGDWELVWSQPTIPCHRY 302
Query: 299 NVCGNFSSCN-DDNDKLCTCLPGFGRRSPLNDYTVGGDTSS------LLCTRK---STSC 348
N+CG+ + C DDN+ +CTC GF +SP +Y G T L C+ + +T+
Sbjct: 303 NLCGDNAECTADDNEPICTCFTGFEPKSP-QEYNNGNWTQGCVRSVPLTCSSERNNTTAG 361
Query: 349 GANTNTFLNLTMMK-IGSPDIKVSAQ---DENECKFRCISMCSQTQCQACSYVPIPVQQR 404
GA T+++ + PD V D N C+ C+ CS C A SY
Sbjct: 362 GAGAGGGDGFTVIRGVKLPDFAVWGSLVGDANSCEKACLGNCS---CGAYSY-------- 410
Query: 405 GLNLSPCWIWTQNLTTLKEEYLGGDDRK--LFVRVAKSDIEEPTRKGNPKSTLSLILGIA 462
+ C W Q L + + G + K L+V+V S +++ + G K+ + +++ +
Sbjct: 411 --STGSCLTWGQELVDIFQFQTGTEGAKYDLYVKVPSSLLDKSS--GRWKTVVVVVVVVV 466
Query: 463 LPGVVILACICILAYVCRRKIALKL---KQESESILRQRGRFYDSERHVKDLIDKEGLEE 519
VV+L +L + CRR+I KL +++++ L + R D+++ E +
Sbjct: 467 ---VVVLLASGLLMWKCRRRIKEKLGIGRKKAQLPLLRPAR--DAKQDFSGPAQSEHEKS 521
Query: 520 KDNEGIEVPYFDFESILVATDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQG 579
++ + E+P F FE++ ATD FS +NKLG GG+G VYKG+L GG EIAVKRLS S QG
Sbjct: 522 EEGKNCELPLFAFETLATATDNFSISNKLGEGGFGHVYKGRLPGGEEIAVKRLSRSSGQG 581
Query: 580 IQEFKNEVVLIAKLQHRNLVRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQ 639
++EFKNEV+LIAKLQHRNLVRL G CI+G+EKIL+YEYMPNKSLDAF+FDP + LLDW+
Sbjct: 582 LEEFKNEVILIAKLQHRNLVRLLGCCIQGEEKILVYEYMPNKSLDAFLFDPERRGLLDWR 641
Query: 640 MRFDILLGIARGLLYLHQDSRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEA 699
RF I+ G+ARGLLYLH+DSRLRV+HRDLK SNILLD +M PKISDFG+ARIFGG + +
Sbjct: 642 TRFQIIEGVARGLLYLHRDSRLRVVHRDLKASNILLDRDMNPKISDFGMARIFGGDQNQV 701
Query: 700 NTQRVVGTYGYMSPEYALDGQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYA 759
NT RVVGT GYMSPEYA++G FS +SD++SFG+++LEII+G+KN+ F+ +G+L+++GYA
Sbjct: 702 NTNRVVGTLGYMSPEYAMEGLFSVRSDVYSFGILILEIITGQKNSSFHHMEGSLNIVGYA 761
Query: 760 WKLWTENKLLDLMDLSLGEAYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETAT 819
W+LW ++ +L+D ++ A + +RC H+ LLCVQD DRP++ VV+ L S+++
Sbjct: 762 WQLWNGDRGQELIDPAIRGTCPAKEALRCVHMALLCVQDHAHDRPDIPYVVLTLGSDSSV 821
Query: 820 LPTPKQPTFFTRKDLSSTASSS 841
LPTP+ PTF L T+SSS
Sbjct: 822 LPTPRPPTF----TLQCTSSSS 839
>gb|AAU90229.1| putative receptor-like protein kinase [Oryza sativa (japonica
cultivar-group)]
Length = 837
Score = 549 bits (1414), Expect = e-154
Identities = 349/844 (41%), Positives = 481/844 (56%), Gaps = 78/844 (9%)
Query: 30 AGDTLNVGQEITGNGTVLVSAAKK---FELGFFSPDLNVTGGKGRYLGIWYYREEGSGLS 86
A D++ G+ + G+ T++ + A F LGFF+P G Y+G+WY R +S
Sbjct: 22 ARDSIAPGEPLAGHDTLVSAGAGDGGGFALGFFTPP----GSNDTYVGVWYAR-----VS 72
Query: 87 P--VVWVANRDNPV---ADDSIGV-FRIADDGNLVVLDTSGIRYWSASNLTNSSSATNRS 140
P VVWVANR +PV D + G ++ L V D + WS + T +
Sbjct: 73 PRTVVWVANRADPVPGPVDGNAGATLSVSRACELAVADANSTVVWSVTPATTGPC----T 128
Query: 141 VKLMDSGNLVLLDEHVGMKLWESFEHPTDTFLPGMKMD------KTLELTCWKSLSDPGR 194
++ D GNLV+ DE G W+ F+HPTDT LPGM++ + LT WKS SDP
Sbjct: 129 ARIRDDGNLVVTDER-GRVAWQGFDHPTDTLLPGMRIGVDFAAGNNMTLTAWKSPSDPSP 187
Query: 195 GNFTFKMDKKWENRFAILNQGQLYWQSEEQGDGVMNPESNPDDIS--NDVYNLLTNFKEL 252
+ MD + + N W+S DG M PD I+ N ++ + + +E+
Sbjct: 188 SSVVVAMDTSGDPEVFLWNGPNKVWRSGPW-DG-MQFTGVPDTITYKNFSFSFVNSAREV 245
Query: 253 KN--KTVSSYDNTRLLLNSTGVIKVLYRVNFQSDIVW---WYQPRTTCLTYNVCGNFSSC 307
+ + +RL+LNS+G V ++ W WY P+ C + CG C
Sbjct: 246 TYSFQVPDASIMSRLVLNSSGGGLVQRWTWVEAAGAWNLYWYAPKDQCDAVSPCGANGVC 305
Query: 308 NDDNDKLCTCLPGFGRRSP----LNDYTVGGDTSSLLCTRKST-SCGANTNTFLNLTMMK 362
+ ++ +C+CL GF RSP L D G C R++ C T+ F + K
Sbjct: 306 DTNSLPVCSCLRGFAPRSPAAWALRDGRDG-------CARETPLGCANGTDGFAVVRHAK 358
Query: 363 IGSPDIKVSAQDENE----CKFRCISMCSQTQCQACSYVPIPVQQRGLNLSPCWIWTQNL 418
+PD + D + C+ RC+ CS T A + + P +RG C +WT L
Sbjct: 359 --APDTTAATVDYDAGLQLCRRRCLGNCSCT-AYANANLSAPPGRRG-----CVMWTGEL 410
Query: 419 TTLKEEYLGGDDRKLFVRVAKSDIEEPTRKGNPKSTLSLILGIALPGVVILACICILAYV 478
L+ G D L+VR+A +D++ T K K+ + + + +++ + I+ + + Y+
Sbjct: 411 EDLRVYPAFGQD--LYVRLAAADLDS-TSKSKKKTHIIIAVVVSICALAIILALTGM-YI 466
Query: 479 CRRKIALKLKQESESILRQRGRFYDSERHVKDLIDKEGLEEKDNEGIEVPYFDFESILVA 538
R K K K + G + E H EG D+ +++P FD E+I A
Sbjct: 467 WRTK---KTKARRQGPSNWSGGLHSRELH------SEGNSHGDD--LDLPLFDLETIASA 515
Query: 539 TDYFSDANKLGRGGYGPVYKGKLQGGREIAVKRLSSVSSQGIQEFKNEVVLIAKLQHRNL 598
T+ FS NKLG GG+GPVYKG L+ G+EIAVK LS S QG+ EF+NEV+LIAKLQHRNL
Sbjct: 516 TNGFSADNKLGEGGFGPVYKGTLEDGQEIAVKTLSKTSVQGLDEFRNEVMLIAKLQHRNL 575
Query: 599 VRLWGYCIKGDEKILIYEYMPNKSLDAFVFDPTKSALLDWQMRFDILLGIARGLLYLHQD 658
V+L GY + G EK+L+YE+M NKSLD F+FD +KS LLDWQ R+ I+ GIARGLLYLHQD
Sbjct: 576 VQLIGYSVCGQEKMLLYEFMENKSLDCFLFDKSKSKLLDWQTRYHIIEGIARGLLYLHQD 635
Query: 659 SRLRVIHRDLKTSNILLDGEMQPKISDFGLARIFGGKETEANTQRVVGTYGYMSPEYALD 718
SR R+IHRDLKTSNILLD EM PKISDFG+AR+FG +TE NT RVVGTYGYM+PEYA+D
Sbjct: 636 SRYRIIHRDLKTSNILLDKEMTPKISDFGMARMFGSDDTEINTVRVVGTYGYMAPEYAMD 695
Query: 719 GQFSTKSDIFSFGVVLLEIISGKKNTGFYQYKGTLSLLGYAWKLWTENKLLDLMDLSLGE 778
G FS KSD+FSFGV++LEIISGK+N G Y Y L+LL AW W+E LDL+D +L
Sbjct: 696 GVFSVKSDVFSFGVIVLEIISGKRNRGVYSYSSHLNLLARAWSSWSEGNSLDLVDKTLNG 755
Query: 779 AYNANQFIRCTHVGLLCVQDEPDDRPNMSNVVIMLDSETAT-LPTPKQPTFFTRKDLSST 837
++N + ++C VGLLCVQ+ PDDRP MS V++ML S AT LP P++P F R+ +
Sbjct: 756 SFNQEEVLKCLKVGLLCVQENPDDRPLMSQVLLMLASADATSLPDPRKPGFVARRAATED 815
Query: 838 ASSS 841
SSS
Sbjct: 816 TSSS 819
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.138 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,537,106,779
Number of Sequences: 2540612
Number of extensions: 69703850
Number of successful extensions: 209323
Number of sequences better than 10.0: 20351
Number of HSP's better than 10.0 without gapping: 9325
Number of HSP's successfully gapped in prelim test: 11030
Number of HSP's that attempted gapping in prelim test: 162685
Number of HSP's gapped (non-prelim): 25849
length of query: 852
length of database: 863,360,394
effective HSP length: 137
effective length of query: 715
effective length of database: 515,296,550
effective search space: 368437033250
effective search space used: 368437033250
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 80 (35.4 bits)
Lotus: description of TM0203.5


