

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.14
(896 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
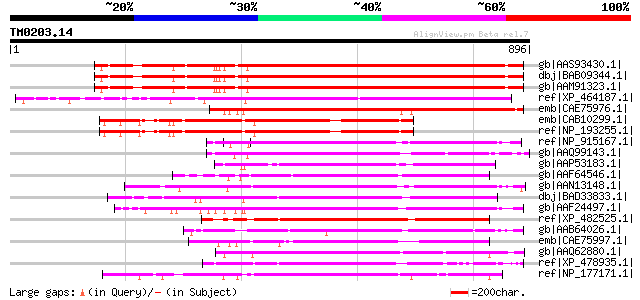
Score E
Sequences producing significant alignments: (bits) Value
gb|AAS93430.1| formin homology 2 domain-containing protein 5 [Ar... 650 0.0
dbj|BAB09344.1| unnamed protein product [Arabidopsis thaliana] g... 648 0.0
gb|AAM91323.1| unknown protein [Arabidopsis thaliana] gi|1459602... 645 0.0
ref|XP_464187.1| putative formin I2I isoform [Oryza sativa (japo... 558 e-157
emb|CAE75976.1| B1160F02.7 [Oryza sativa (japonica cultivar-grou... 525 e-147
emb|CAB10299.1| p140mDia like protein [Arabidopsis thaliana] gi|... 466 e-129
ref|NP_193255.1| formin homology 2 domain-containing protein / F... 466 e-129
ref|NP_915167.1| putative formin-like protein AHF1 [Oryza sativa... 413 e-113
gb|AAQ99143.1| formin-like protein AtFH6 [Arabidopsis thaliana] ... 395 e-108
gb|AAP53183.1| putative formin-like protein [Oryza sativa (japon... 394 e-108
gb|AAF64546.1| unknown protein [Arabidopsis thaliana] gi|1522999... 384 e-105
gb|AAN13148.1| putative formin protein AHF1 [Arabidopsis thalian... 383 e-104
dbj|BAD33833.1| diaphanous protein-like [Oryza sativa (japonica ... 373 e-101
gb|AAF24497.1| FH protein NFH2 [Nicotiana tabacum] 363 1e-98
ref|XP_482525.1| putative formin homology(FH)protein [Oryza sati... 363 1e-98
gb|AAB64026.1| unknown protein [Arabidopsis thaliana] gi|2540890... 356 2e-96
emb|CAE75997.1| B1358B12.6 [Oryza sativa (japonica cultivar-grou... 355 3e-96
gb|AAQ62880.1| At1g59910 [Arabidopsis thaliana] gi|5080823|gb|AA... 342 3e-92
ref|XP_478935.1| putative formin homology(FH) domain-containing ... 331 6e-89
ref|NP_177171.1| formin homology 2 domain-containing protein / F... 326 2e-87
>gb|AAS93430.1| formin homology 2 domain-containing protein 5 [Arabidopsis
thaliana] gi|47716755|gb|AAT37554.1| formin homology 2
domain-containing protein 5 [Arabidopsis thaliana]
Length = 900
Score = 650 bits (1676), Expect = 0.0
Identities = 399/811 (49%), Positives = 497/811 (61%), Gaps = 95/811 (11%)
Query: 147 GLESGHHGRNL-----ISDPPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTP 201
G +GH+ + L + D P L+ P G+ PSP P+ P P P SP P
Sbjct: 114 GKLNGHNPKYLELLSSMLDIPRRNLATKP-GSSPSPSPSRPPKRSRGPPRPPTRPKSPPP 172
Query: 202 TPENSSPSSTPSPTAMVPPTSPPRQHPPPSPEKLQHVTYAPDPSPPDKGDRKQKAIILAV 261
+ PS +P P PP ++ T AP SP K + +K II+AV
Sbjct: 173 RKASFPPSRSPPP-------------PPAKKNASKNSTSAP-VSPAKKKEDHEKTIIIAV 218
Query: 262 TVSGIIIMIGLALCYRESRR-----SDRADKDEGPLLILTSSDYSGGKQKVVRLGNANNE 316
V+ + + AL + R S DE PLL L+SSDYS G + G+ +
Sbjct: 219 VVTAVSTFLLAALFFLCCSRVCGNGSGGRKNDERPLLSLSSSDYSVG-SSINYGGSVKGD 277
Query: 317 GSGISNGKKPSNVRNWSMKAG---DNNTSVVEATSSEGV--------------------- 352
G + SN S G D + S+ E S EG+
Sbjct: 278 KQGHQSFNIYSNQGKMSSFDGSNSDTSDSLEERLSHEGLRNNSITNHGLPPLKPPPGRTA 337
Query: 353 ---------GQV----PEPPN---------GKPPPPPP--------GPPPPPPPRKPP-A 381
G+V PEPP PPPP P GPP PPPP PP +
Sbjct: 338 SVLSGKSFSGKVEPLPPEPPKFLKVSSKKASAPPPPVPAPPMPSSAGPPRPPPPAPPPGS 397
Query: 382 PAPRPPPPPRAGQPPPAPPKPVKVGNQG----ANQGGTSDGDAPKPKLKPFFWDKVAAKP 437
P+PPPPP G P PP P+ +G + + D DAPK KLKPFFWDKV A P
Sbjct: 398 GGPKPPPPP--GPKGPRPPPPMSLGPKAPRPPSGPADALDDDAPKTKLKPFFWDKVQANP 455
Query: 438 DQQMVWHEISAGSFVFNEEKMESLFGCANQNRNERKKDSPSVDTSV-QYIQIIDPKKAQN 496
+ MVW++I +GSF FNEE +ESLFG A ++N+ K S ++ Q++QI++PKK QN
Sbjct: 456 EHSMVWNDIRSGSFQFNEEMIESLFGYAAADKNKNDKKGSSGQAALPQFVQILEPKKGQN 515
Query: 497 LSILLRALNVTTDEVIDALREGNEIPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAE 556
LSILLRALN TT+EV DALREGNE+PVE IQTLLKMAPT EEELKLRL+ GE++QLG AE
Sbjct: 516 LSILLRALNATTEEVCDALREGNELPVEFIQTLLKMAPTPEEELKLRLYCGEIAQLGSAE 575
Query: 557 RFLKVLVDIPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVL 616
RFLK +VDIPFAFKRLE LLFM L EE + +KESF TLEVA +LR SRLFLKLLEAVL
Sbjct: 576 RFLKAVVDIPFAFKRLEALLFMCTLHEEMAFVKESFQTLEVACKELRGSRLFLKLLEAVL 635
Query: 617 KTGNRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGIRAVRTERE 676
KTGNRMNDGT+RGGAQAF+LDTLLKL+DVKGTDGKTTLLHFVVQEIIR+EG+RA RT RE
Sbjct: 636 KTGNRMNDGTFRGGAQAFKLDTLLKLADVKGTDGKTTLLHFVVQEIIRTEGVRAARTIRE 695
Query: 677 SRSVVSSVGTDQDVDESPEASEEHYRSLGLQVVSSLSNELEDVKRAALIDGDALSSAVSK 736
S+S S D V+E+ E SEE+YR+LGL+ VS LS+ELE VK++A ID D L+ V K
Sbjct: 696 SQSFSSVKTEDLLVEETSEESEENYRNLGLEKVSGLSSELEHVKKSANIDADGLTGTVLK 755
Query: 737 LGHSLAKTQEFMNTDLKSLDEESEFQRCTERFVEKAREEVTWLQEEEKRIIAQVKSTADY 796
+GH+L+K ++F+N+++KS EES F+ E F++ A + + EEEKRI+A VKST DY
Sbjct: 756 MGHALSKARDFVNSEMKSSGEESGFREALEDFIQNAEGSIMSILEEEKRIMALVKSTGDY 815
Query: 797 FHGNAGRDEGLRLFLIVRDFLIILDKVCKEVRDKTLRAVKSSQRKEALSVPSSPDTRHQQ 856
FHG AG+DEGLRLF+IVRDFLIILDK CKEVR+ R V+ + RK+ + +S +T Q
Sbjct: 816 FHGKAGKDEGLRLFVIVRDFLIILDKSCKEVREARGRPVRMA-RKQGSTASASSETPRQ- 873
Query: 857 SPPAPS-DLHRRLFPAIAGRRIVEDSSSDDD 886
PS D ++LFPAI RR V+ SSSD D
Sbjct: 874 ---TPSLDPRQKLFPAITERR-VDQSSSDSD 900
>dbj|BAB09344.1| unnamed protein product [Arabidopsis thaliana]
gi|15239699|ref|NP_200276.1| formin homology 2
domain-containing protein / FH2 domain-containing
protein [Arabidopsis thaliana]
gi|30696498|ref|NP_851191.1| formin homology 2
domain-containing protein / FH2 domain-containing
protein [Arabidopsis thaliana]
Length = 900
Score = 648 bits (1671), Expect = 0.0
Identities = 398/811 (49%), Positives = 496/811 (61%), Gaps = 95/811 (11%)
Query: 147 GLESGHHGRNL-----ISDPPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTP 201
G +GH+ + L + D P L+ P G+ PSP P+ P P P SP P
Sbjct: 114 GKLNGHNPKYLELLSSMLDIPRRNLATKP-GSSPSPSPSRPPKRSRGPPRPPTRPKSPPP 172
Query: 202 TPENSSPSSTPSPTAMVPPTSPPRQHPPPSPEKLQHVTYAPDPSPPDKGDRKQKAIILAV 261
+ PS +P P PP ++ T AP SP K + +K II+AV
Sbjct: 173 RKSSFPPSRSPPP-------------PPAKKNASKNSTSAP-VSPAKKKEDHEKTIIIAV 218
Query: 262 TVSGIIIMIGLALCYRESRR-----SDRADKDEGPLLILTSSDYSGGKQKVVRLGNANNE 316
V+ + + AL + R S DE PLL L+SSDYS G + G+ +
Sbjct: 219 VVTAVSTFLLAALFFLCCSRVCGNGSGGRKNDERPLLSLSSSDYSVG-SSINYGGSVKGD 277
Query: 317 GSGISNGKKPSNVRNWSMKAG---DNNTSVVEATSSEGV--------------------- 352
G + SN S G D + S+ E S EG+
Sbjct: 278 KQGHQSFNIYSNQGKMSSFDGSNSDTSDSLEERLSHEGLRNNSITNHGLPPLKPPPGRTA 337
Query: 353 ---------GQV----PEPPN---------GKPPPPPP--------GPPPPPPPRKPP-A 381
G+V PEPP PPPP P GPP PPPP PP +
Sbjct: 338 SVLSGKSFSGKVEPLPPEPPKFLKVSSKKASAPPPPVPAPQMPSSAGPPRPPPPAPPPGS 397
Query: 382 PAPRPPPPPRAGQPPPAPPKPVKVGNQG----ANQGGTSDGDAPKPKLKPFFWDKVAAKP 437
P+PPPPP G P PP P+ +G + + D DAPK KLKPFFWDKV A P
Sbjct: 398 GGPKPPPPP--GPKGPRPPPPMSLGPKAPRPPSGPADALDDDAPKTKLKPFFWDKVQANP 455
Query: 438 DQQMVWHEISAGSFVFNEEKMESLFGCANQNRNERKKDSPSVDTSV-QYIQIIDPKKAQN 496
+ MVW++I +GSF FNEE +ESLFG A ++N+ K S ++ Q++QI++PKK QN
Sbjct: 456 EHSMVWNDIRSGSFQFNEEMIESLFGYAAADKNKNDKKGSSGQAALPQFVQILEPKKGQN 515
Query: 497 LSILLRALNVTTDEVIDALREGNEIPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAE 556
LSILLRALN TT+EV DALREGNE+PVE IQTLLKMAPT EEELKLRL+ GE++QLG AE
Sbjct: 516 LSILLRALNATTEEVCDALREGNELPVEFIQTLLKMAPTPEEELKLRLYCGEIAQLGSAE 575
Query: 557 RFLKVLVDIPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVL 616
RFLK +VDIPFAFKRLE LLFM L EE + +KESF LEVA +LR SRLFLKLLEAVL
Sbjct: 576 RFLKAVVDIPFAFKRLEALLFMCTLHEEMAFVKESFQKLEVACKELRGSRLFLKLLEAVL 635
Query: 617 KTGNRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGIRAVRTERE 676
KTGNRMNDGT+RGGAQAF+LDTLLKL+DVKGTDGKTTLLHFVVQEIIR+EG+RA RT RE
Sbjct: 636 KTGNRMNDGTFRGGAQAFKLDTLLKLADVKGTDGKTTLLHFVVQEIIRTEGVRAARTIRE 695
Query: 677 SRSVVSSVGTDQDVDESPEASEEHYRSLGLQVVSSLSNELEDVKRAALIDGDALSSAVSK 736
S+S S D V+E+ E SEE+YR+LGL+ VS LS+ELE VK++A ID D L+ V K
Sbjct: 696 SQSFSSVKTEDLLVEETSEESEENYRNLGLEKVSGLSSELEHVKKSANIDADGLTGTVLK 755
Query: 737 LGHSLAKTQEFMNTDLKSLDEESEFQRCTERFVEKAREEVTWLQEEEKRIIAQVKSTADY 796
+GH+L+K ++F+N+++KS EES F+ E F++ A + + EEEKRI+A VKST DY
Sbjct: 756 MGHALSKARDFVNSEMKSSGEESGFREALEDFIQNAEGSIMSILEEEKRIMALVKSTGDY 815
Query: 797 FHGNAGRDEGLRLFLIVRDFLIILDKVCKEVRDKTLRAVKSSQRKEALSVPSSPDTRHQQ 856
FHG AG+DEGLRLF+IVRDFLIILDK CKEVR+ R V+ + RK+ + +S +T Q
Sbjct: 816 FHGKAGKDEGLRLFVIVRDFLIILDKSCKEVREARGRPVRMA-RKQGSTASASSETPRQ- 873
Query: 857 SPPAPS-DLHRRLFPAIAGRRIVEDSSSDDD 886
PS D ++LFPAI RR V+ SSSD D
Sbjct: 874 ---TPSLDPRQKLFPAITERR-VDQSSSDSD 900
>gb|AAM91323.1| unknown protein [Arabidopsis thaliana] gi|14596027|gb|AAK68741.1|
Unknown protein [Arabidopsis thaliana]
Length = 900
Score = 645 bits (1664), Expect = 0.0
Identities = 397/811 (48%), Positives = 495/811 (60%), Gaps = 95/811 (11%)
Query: 147 GLESGHHGRNL-----ISDPPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTP 201
G +GH+ + L + D P L+ P G+ PSP P+ P P P SP P
Sbjct: 114 GKLNGHNPKYLELLSSMLDIPRRNLATKP-GSSPSPSPSRPPKRSRGPPRPPTRPKSPPP 172
Query: 202 TPENSSPSSTPSPTAMVPPTSPPRQHPPPSPEKLQHVTYAPDPSPPDKGDRKQKAIILAV 261
+ PS +P P PP ++ T AP SP K + +K II+AV
Sbjct: 173 RKSSFPPSRSPPP-------------PPAKKNASKNSTSAP-VSPAKKKEDHEKTIIIAV 218
Query: 262 TVSGIIIMIGLALCYRESRR-----SDRADKDEGPLLILTSSDYSGGKQKVVRLGNANNE 316
V+ + + AL + R S DE PLL L+SSDYS G + G+ +
Sbjct: 219 VVTAVSTFLLAALFFLCCSRVCGNGSGGRKNDERPLLSLSSSDYSVG-SSINYGGSVKGD 277
Query: 317 GSGISNGKKPSNVRNWSMKAG---DNNTSVVEATSSEGV--------------------- 352
G + SN S G D + S+ E S EG+
Sbjct: 278 KQGHQSFNIYSNQGKMSSFDGSNSDTSDSLEERLSHEGLRNNSITNHGLPPLKPPPGRTA 337
Query: 353 ---------GQV----PEPPN---------GKPPPPPP--------GPPPPPPPRKPP-A 381
G+V PEPP PPPP P GPP PPPP PP +
Sbjct: 338 SVLSGKSFSGKVEPLPPEPPKFLKVSSKKASAPPPPVPAPQMPSSAGPPRPPPPAPPPGS 397
Query: 382 PAPRPPPPPRAGQPPPAPPKPVKVGNQG----ANQGGTSDGDAPKPKLKPFFWDKVAAKP 437
P+PPPPP G P PP P+ +G + + D DAPK KLKPFFWDKV A P
Sbjct: 398 GGPKPPPPP--GPKGPRPPPPMSLGPKAPRPPSGPADALDDDAPKTKLKPFFWDKVQANP 455
Query: 438 DQQMVWHEISAGSFVFNEEKMESLFGCANQNRNERKKDSPSVDTSV-QYIQIIDPKKAQN 496
+ MVW++I +GSF FNEE +ESLFG A ++N+ K S ++ Q++QI++PKK QN
Sbjct: 456 EHSMVWNDIRSGSFQFNEEMIESLFGYAAADKNKNDKKGSSGQAALPQFVQILEPKKGQN 515
Query: 497 LSILLRALNVTTDEVIDALREGNEIPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAE 556
LSILLRALN TT+EV DALREGNE+PVE IQTLLKMAPT EEELKLRL+ GE++QLG AE
Sbjct: 516 LSILLRALNATTEEVCDALREGNELPVEFIQTLLKMAPTPEEELKLRLYCGEIAQLGSAE 575
Query: 557 RFLKVLVDIPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVL 616
RFLK +VDIPFAFKRLE LLFM L EE + +KESF LEVA +LR SRLFLKLLEAVL
Sbjct: 576 RFLKAVVDIPFAFKRLEALLFMCTLHEEMAFVKESFQKLEVACKELRGSRLFLKLLEAVL 635
Query: 617 KTGNRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGIRAVRTERE 676
KTGNRMNDGT+RGGAQAF+LDTLLKL+DVKGTDGKTTLLHFVVQEIIR+EG+RA RT RE
Sbjct: 636 KTGNRMNDGTFRGGAQAFKLDTLLKLADVKGTDGKTTLLHFVVQEIIRTEGVRAARTIRE 695
Query: 677 SRSVVSSVGTDQDVDESPEASEEHYRSLGLQVVSSLSNELEDVKRAALIDGDALSSAVSK 736
S+S S D V+E+ E SEE+YR+LGL+ VS LS+ELE VK++A ID D L+ V K
Sbjct: 696 SQSFSSVKTEDLLVEETSEESEENYRNLGLEKVSGLSSELEHVKKSANIDADGLTGTVLK 755
Query: 737 LGHSLAKTQEFMNTDLKSLDEESEFQRCTERFVEKAREEVTWLQEEEKRIIAQVKSTADY 796
+GH+L+K ++F+N+++KS E S F+ E F++ A + + EEEKRI+A VKST DY
Sbjct: 756 MGHALSKARDFVNSEMKSSGEGSGFREALEDFIQNAEGSIMSILEEEKRIMALVKSTGDY 815
Query: 797 FHGNAGRDEGLRLFLIVRDFLIILDKVCKEVRDKTLRAVKSSQRKEALSVPSSPDTRHQQ 856
FHG AG+DEGLRLF+IVRDFLIILDK CKEVR+ R V+ + RK+ + +S +T Q
Sbjct: 816 FHGKAGKDEGLRLFVIVRDFLIILDKSCKEVREARGRPVRMA-RKQGSTASASSETPRQ- 873
Query: 857 SPPAPS-DLHRRLFPAIAGRRIVEDSSSDDD 886
PS D ++LFPAI RR V+ SSSD D
Sbjct: 874 ---TPSLDPRQKLFPAITERR-VDQSSSDSD 900
>ref|XP_464187.1| putative formin I2I isoform [Oryza sativa (japonica
cultivar-group)] gi|50251274|dbj|BAD28054.1| putative
formin I2I isoform [Oryza sativa (japonica
cultivar-group)] gi|49389244|dbj|BAD25206.1| putative
formin I2I isoform [Oryza sativa (japonica
cultivar-group)]
Length = 881
Score = 558 bits (1438), Expect = e-157
Identities = 358/902 (39%), Positives = 491/902 (53%), Gaps = 80/902 (8%)
Query: 10 VIYVILLCALAQ--------GGSEGKKGKDEDFSSCAGGGPSFSSSFHVHGKQVENVGKH 61
V++++LL A + GG E K GK + GP+ +SS + G V+ +
Sbjct: 6 VVFLLLLVAASALVKSSRGGGGGEEKLGKFYGWRRHLSSGPA-ASSLVLSGDLVDKIWSV 64
Query: 62 CRKELVEKSNDDVKDFDLHLLERGSDLRCIHLLENEIPKTT-----KSLSPVMRKKVLDC 116
C +++V S +D F D H E+E+ T L P DC
Sbjct: 65 CLQDIV--SPEDTFGFGESF---AWDELSSHSTEDELKATLFMELMALLPPEKSSFTYDC 119
Query: 117 IRRSTPPAPPVSQKKTGSRYLLTKHSRSTLGLESGHHGRNLISDPPHNELSPSPAGTVPS 176
IR + G + + + L + G N P L G PS
Sbjct: 120 IRANC--------FSLGVPQIFSVALSNYLESQKSLVGSNFY---PRRRLVDKLIGDAPS 168
Query: 177 PGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHPPPSPEKLQ 236
PA + G +SP S P TPS + + P SP +P S K Q
Sbjct: 169 MAPAFAPSMSSGGEV--HSPLSVAEAP------LTPSNSLNMEPPSP--YYPSKSAHKHQ 218
Query: 237 HVTYAPDPSPPDKGDRKQKAIILAVTVSGIIIMIGLALCY-----RESRRSDRADKDEGP 291
V AP SP ++ K +++AV + + + LC+ +S+ S +D+ P
Sbjct: 219 GV--APPVSPSEEYHDYMKVVLIAVLPTAALSFLAAFLCFYCCGCNKSKVSVGEQRDDHP 276
Query: 292 LLILTSSDYSGGKQKV-VRLGNANNEGSGISNGKKPSNVRNWSMK--------AGDNNTS 342
LL L S+ G V V + + G+ +PSN K + D T
Sbjct: 277 LLHLQFSNLPGSSPDVHVPASPLHKDDHGV----RPSNAGVSMSKCFPCCFKTSSDATTP 332
Query: 343 VVEATSSEGVGQVPEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKP 402
+ ++ + P PPPPPP PPPPPPP PP P PPPP + G PPPAPPK
Sbjct: 333 TLVTGGTQENNATSDAPKLMPPPPPPPPPPPPPPPPPPPRPPPPPPPIKKGAPPPAPPKA 392
Query: 403 VKV-----------------GNQGANQGGTSDGDAPKPKLKPFFWDKVAAKPDQQMVWHE 445
++ A++ ++ +AP+ KL+PF+WDKV A PDQ M WH+
Sbjct: 393 TMARFPKLSPTESSRSEESSASELASESSETEVNAPRAKLRPFYWDKVLANPDQSMAWHD 452
Query: 446 ISAGSFVFNEEKMESLFGCANQNRNERKKDSPSV-DTSVQYIQIIDPKKAQNLSILLRAL 504
I GSF NEE +E LFG N+N K S+ D S Q++ ++D KK+ NL+++ +A+
Sbjct: 453 IKFGSFHVNEEMIEELFGYGAGNQNNVKDKEISIADPSPQHVSLLDVKKSCNLAVVFKAM 512
Query: 505 NVTTDEVIDALREGNEIPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAERFLKVLVD 564
NV +E+ DAL EGNE+P L++T+L+M PT EEE KLRL+ G+ SQLG AE+ +K L+D
Sbjct: 513 NVRAEEIHDALVEGNELPRLLLETILRMKPTDEEEQKLRLYNGDCSQLGLAEQVMKALID 572
Query: 565 IPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGNRMND 624
IPFAF+R+ LLFM L+E+ASS++ESF LE A +L K RLFLKLLEA+LKTGNR+ND
Sbjct: 573 IPFAFERIRALLFMSSLQEDASSLRESFLQLEAACGEL-KHRLFLKLLEAILKTGNRLND 631
Query: 625 GTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGIRAVRTERES-RSVVSS 683
GT+RGGA AF+LDTLLKLSDVKG DGKTTLLHFVVQEIIRSEG+R R E+ RS
Sbjct: 632 GTFRGGANAFKLDTLLKLSDVKGADGKTTLLHFVVQEIIRSEGVREARLAMENGRSPPFP 691
Query: 684 VGTDQDVDESPEASEEHYRSLGLQVVSSLSNELEDVKRAALIDGDALSSAVSKLGHSLAK 743
+D + +ES + +Y +LGL++VS LSNEL++VKR A +D DALS++V+ L H L +
Sbjct: 692 STSDDNSNESLQEDGNYYSNLGLKIVSGLSNELDNVKRVAALDADALSTSVANLRHELLR 751
Query: 744 TQEFMNTDLKSLDEESEFQRCTERFVEKAREEVTWLQEEEKRIIAQVKSTADYFHGNAGR 803
+EF+N+D+ SL+E S F R E F+E A E +L +E+KR+ VK T YFHGN +
Sbjct: 752 AKEFLNSDMASLEENSGFHRSLESFIEHAETETNFLLKEDKRLRMLVKRTIRYFHGNDEK 811
Query: 804 DEGLRLFLIVRDFLIILDKVCKEVRDKTLRAVKSSQRKEALSVPSSPDTRHQQSPPAPSD 863
D+G RLF+IVRDFL++LDK CKEV +A SQ + PSS +Q PA D
Sbjct: 812 DDGFRLFVIVRDFLVMLDKACKEVGASQKKATNKSQANGNSNNPSSQSNPQEQQFPAVLD 871
Query: 864 LH 865
H
Sbjct: 872 HH 873
>emb|CAE75976.1| B1160F02.7 [Oryza sativa (japonica cultivar-group)]
gi|50921155|ref|XP_470938.1| B1160F02.7 [Oryza sativa
(japonica cultivar-group)]
Length = 906
Score = 525 bits (1351), Expect = e-147
Identities = 285/596 (47%), Positives = 383/596 (63%), Gaps = 55/596 (9%)
Query: 346 ATSSEGVGQVPEPPN--GKPPPPPP-----------GPPPPPPPR-----KPPAPAPRPP 387
A + G G P PP P PPPP GPPPPPPP +PP P P PP
Sbjct: 312 AGAGAGTGAPPPPPAHPAAPAPPPPAPSPSAAGAGSGPPPPPPPAAPAAPRPPGPGPGPP 371
Query: 388 PPPRA------GQPPPAPPK------PVKVGNQGANQGGTSDGDAPKPKLKPFFWDKVAA 435
PPP A G PPPA P P + D K KLKPFFWDKV A
Sbjct: 372 PPPGAAGRGGGGPPPPALPGGPRARGPPPFKKSPGAAAAAAQADPNKAKLKPFFWDKVTA 431
Query: 436 KPDQQMVWHEISAGSFVFNEEKMESLFGCANQNRNERKKDSPSVDTSVQYIQIIDPKKAQ 495
P+Q MVW +I AGSF FNEE +ESLFG + + S + Q+++I+DPKKAQ
Sbjct: 432 NPNQAMVWDQIKAGSFQFNEEMIESLFGAQSTEKKSTDAKKESGKEATQFVRILDPKKAQ 491
Query: 496 NLSILLRALNVTTDEVIDALREGNEIPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPA 555
NL+I L+AL+V+ ++V A+ EG+++P +LIQTL++ +PT++EEL+LRL+ GE +QLGPA
Sbjct: 492 NLAISLKALSVSAEQVRAAVMEGHDLPPDLIQTLVRWSPTSDEELRLRLYAGEPAQLGPA 551
Query: 556 ERFLKVLVDIPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAV 615
E+F++ ++D+P+ ++RL+ LLFM L EEA+++++SFATLEVA ++LR SRLF KLLEAV
Sbjct: 552 EQFMRAIIDVPYLYQRLDALLFMAALPEEAAAVEQSFATLEVACEELRGSRLFKKLLEAV 611
Query: 616 LKTGNRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGIRAVRTER 675
LKTGNRMNDGT+RGGAQAF+LDTLLKL+DVKG DGKTTLLHFVVQEIIRSEG+RA R
Sbjct: 612 LKTGNRMNDGTFRGGAQAFKLDTLLKLADVKGVDGKTTLLHFVVQEIIRSEGVRAARAAS 671
Query: 676 --------------------ESRSVVSSVGTDQDVD----ESPEASEEHYRSLGLQVVSS 711
+S+S + S VD E + E YR LGL VVSS
Sbjct: 672 GGGGGSSISSISSSDDLILLQSQSSIGSNSGRSSVDASSLEQEQDETERYRQLGLGVVSS 731
Query: 712 LSNELEDVKRAALIDGDALSSAVSKLGHSLAKTQEFMNTDLKSLDEESEFQRCTERFVEK 771
L ++L++V++AA D DAL+ V+ LGH L K EF++T ++SL+E+S FQR FV++
Sbjct: 732 LGDDLQNVRKAASFDADALTITVASLGHRLVKANEFLSTGMRSLEEDSGFQRRLASFVQQ 791
Query: 772 AREEVTWLQEEEKRIIAQVKSTADYFHGNAGRDEGLRLFLIVRDFLIILDKVCKEVRDKT 831
++E+VT L E+EKR+ + V++T DYFHG+ G+DEGLRLF++VRDFL ILDKVC+EV+++
Sbjct: 792 SQEQVTRLLEDEKRLRSLVRATVDYFHGSTGKDEGLRLFVVVRDFLGILDKVCREVKEQA 851
Query: 832 LRAVKSSQRKEALSVPSSPDTRHQQSPPAPSDLHRRLFPAIAGRRIVEDSSSDDDD 887
K+ ++++ P S + + R A++ R SSSD DD
Sbjct: 852 AANAKAKKQQQPTPAPRSRQSSQSSFRDPRQQIQDRRAAALS-RNNSSSSSSDSDD 906
Score = 48.1 bits (113), Expect = 0.001
Identities = 34/104 (32%), Positives = 38/104 (35%), Gaps = 9/104 (8%)
Query: 152 HHGRNLISDPPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPEN--SSPS 209
HHG + PPH P P G G P P H PA P +P+P+ S P
Sbjct: 297 HHGHH--PPPPH----PLPPGAGAGAGTGAPPPPPAHPAAPAPPPPAPSPSAAGAGSGPP 350
Query: 210 STPSPTAMVPPTSP-PRQHPPPSPEKLQHVTYAPDPSPPDKGDR 252
P P A P P P PPP P P P G R
Sbjct: 351 PPPPPAAPAAPRPPGPGPGPPPPPGAAGRGGGGPPPPALPGGPR 394
Score = 37.0 bits (84), Expect = 2.9
Identities = 26/85 (30%), Positives = 31/85 (35%), Gaps = 11/85 (12%)
Query: 168 PSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSS-----TPSPTAMVPPTS 222
P PAG P P L P+H + P P P P + + P P P
Sbjct: 277 PPPAGPPPPAPPPLP---PSHHHHHGHHPPPPHPLPPGAGAGAGTGAPPPPPAHPAAPAP 333
Query: 223 PPRQHPPPSPEKLQHVTYAPDPSPP 247
PP P PSP + P P PP
Sbjct: 334 PP---PAPSPSAAGAGSGPPPPPPP 355
>emb|CAB10299.1| p140mDia like protein [Arabidopsis thaliana]
gi|7268266|emb|CAB78562.1| p140mDia like protein
[Arabidopsis thaliana] gi|7485017|pir||A71416
hypothetical protein - Arabidopsis thaliana
Length = 645
Score = 466 bits (1198), Expect = e-129
Identities = 284/579 (49%), Positives = 359/579 (61%), Gaps = 82/579 (14%)
Query: 155 RNLISDP-------PHNELSPSPAGTVPSPGPALE-SPTPNHGP----TPANSPSSP-TP 201
R LIS P P+ P+P+ P PGP+ P PN PA+SP+ P
Sbjct: 98 RKLISYPKKFSVSAPNLAFGPAPS-FAPGPGPSFAPGPAPNPRSYDWLAPASSPNEPPAE 156
Query: 202 TPENSSPSSTPSPTAMVPPTS----PPRQHPPPSPEKLQHVTYAPDPSPPDKGDRKQKAI 257
TP+ SSPS + ++V P+ PPR PPP EK K D K +
Sbjct: 157 TPDESSPSPSEETPSVVAPSQSVPGPPR--PPPQREK--------------KDDILMK-L 199
Query: 258 ILAVTVSGIIIMIGLAL----CYRESRR----SDRADKDEGPLLILTSSDYSGGKQKVVR 309
I+AV + ++ + +AL C++ + S +DEGPLL L++ G +
Sbjct: 200 IIAVASTAVLTFVFVALMFLCCFKRNCNNAVGSRDGPRDEGPLLRLST----GSTENSPT 255
Query: 310 LGNANNEGSGISNGKKPSNVRNWSMKAGDNNTSVVEATSSEGVGQVPEPPNGKPPPPPPG 369
+ + + + +++ KK S + S+K + S E++S+ G+ + PP PPPPP
Sbjct: 256 VASTSRKMFSVASSKKRSFLSRVSLKRNGHEFSTAESSSAAGLPPLKLPPGRSAPPPPPA 315
Query: 370 PPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKPVKVGNQGANQGGTSDGDA--------- 420
PPP P PP P P+PPPPP+ +PPPAPPK G QG TS GDA
Sbjct: 316 AAPPPQPPPPPPPKPQPPPPPKIARPPPAPPK----GAAPKRQGNTSSGDASDVDSETGA 371
Query: 421 PKPKLKPFFWDKVAAKPDQQMVWHEISAGSFVFNEEKMESLFGCANQNRN---ERKKDSP 477
PK KLKPFFWDK+A PDQ+MVWHEISAGSF FNEE MESLFG + N+N ++ DS
Sbjct: 372 PKTKLKPFFWDKMA-NPDQKMVWHEISAGSFQFNEEAMESLFGYNDGNKNKNGQKSTDSS 430
Query: 478 SVDTSVQYIQIIDPKKAQNLSILLRALNVTTDEVIDALREGNEIPVELIQTLLKMAPTTE 537
++ +QYIQIID +KAQNLSILLRALNVTT+EV+DA++EGNE+PVEL+QTLLKMAPT+E
Sbjct: 431 LRESPLQYIQIIDTRKAQNLSILLRALNVTTEEVVDAIKEGNELPVELLQTLLKMAPTSE 490
Query: 538 EELKLRLFTGELSQLGPAERFLKVLVDIPFAFKRLETLLFMFILREEASSIKESFATLEV 597
EELKLRL++G+L LGP R E+LLFM L+EE S +KE+ TLEV
Sbjct: 491 EELKLRLYSGDLHLLGP----------------RSESLLFMISLQEEVSGLKEALGTLEV 534
Query: 598 ASDKLRKSRLFLKLLEAVLKTGNRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLHF 657
A KLR SRLFLKLLEAVLKTGNRMN GT+RG AQAF+LDTLLKLSDVKGTDGKTTLLHF
Sbjct: 535 ACKKLRNSRLFLKLLEAVLKTGNRMNVGTFRGDAQAFKLDTLLKLSDVKGTDGKTTLLHF 594
Query: 658 VVQEIIRSEGIRAVRTERESRSVVSSVGTDQDVDESPEA 696
VV EIIRSEG+RA+R +SRS S D + D SP++
Sbjct: 595 VVLEIIRSEGVRALRL--QSRSFSSVKTDDSNADSSPQS 631
>ref|NP_193255.1| formin homology 2 domain-containing protein / FH2 domain-containing
protein [Arabidopsis thaliana]
Length = 600
Score = 466 bits (1198), Expect = e-129
Identities = 284/579 (49%), Positives = 359/579 (61%), Gaps = 82/579 (14%)
Query: 155 RNLISDP-------PHNELSPSPAGTVPSPGPALE-SPTPNHGP----TPANSPSSP-TP 201
R LIS P P+ P+P+ P PGP+ P PN PA+SP+ P
Sbjct: 53 RKLISYPKKFSVSAPNLAFGPAPS-FAPGPGPSFAPGPAPNPRSYDWLAPASSPNEPPAE 111
Query: 202 TPENSSPSSTPSPTAMVPPTS----PPRQHPPPSPEKLQHVTYAPDPSPPDKGDRKQKAI 257
TP+ SSPS + ++V P+ PPR PPP EK K D K +
Sbjct: 112 TPDESSPSPSEETPSVVAPSQSVPGPPR--PPPQREK--------------KDDILMK-L 154
Query: 258 ILAVTVSGIIIMIGLAL----CYRESRR----SDRADKDEGPLLILTSSDYSGGKQKVVR 309
I+AV + ++ + +AL C++ + S +DEGPLL L++ G +
Sbjct: 155 IIAVASTAVLTFVFVALMFLCCFKRNCNNAVGSRDGPRDEGPLLRLST----GSTENSPT 210
Query: 310 LGNANNEGSGISNGKKPSNVRNWSMKAGDNNTSVVEATSSEGVGQVPEPPNGKPPPPPPG 369
+ + + + +++ KK S + S+K + S E++S+ G+ + PP PPPPP
Sbjct: 211 VASTSRKMFSVASSKKRSFLSRVSLKRNGHEFSTAESSSAAGLPPLKLPPGRSAPPPPPA 270
Query: 370 PPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKPVKVGNQGANQGGTSDGDA--------- 420
PPP P PP P P+PPPPP+ +PPPAPPK G QG TS GDA
Sbjct: 271 AAPPPQPPPPPPPKPQPPPPPKIARPPPAPPK----GAAPKRQGNTSSGDASDVDSETGA 326
Query: 421 PKPKLKPFFWDKVAAKPDQQMVWHEISAGSFVFNEEKMESLFGCANQNRN---ERKKDSP 477
PK KLKPFFWDK+A PDQ+MVWHEISAGSF FNEE MESLFG + N+N ++ DS
Sbjct: 327 PKTKLKPFFWDKMA-NPDQKMVWHEISAGSFQFNEEAMESLFGYNDGNKNKNGQKSTDSS 385
Query: 478 SVDTSVQYIQIIDPKKAQNLSILLRALNVTTDEVIDALREGNEIPVELIQTLLKMAPTTE 537
++ +QYIQIID +KAQNLSILLRALNVTT+EV+DA++EGNE+PVEL+QTLLKMAPT+E
Sbjct: 386 LRESPLQYIQIIDTRKAQNLSILLRALNVTTEEVVDAIKEGNELPVELLQTLLKMAPTSE 445
Query: 538 EELKLRLFTGELSQLGPAERFLKVLVDIPFAFKRLETLLFMFILREEASSIKESFATLEV 597
EELKLRL++G+L LGP R E+LLFM L+EE S +KE+ TLEV
Sbjct: 446 EELKLRLYSGDLHLLGP----------------RSESLLFMISLQEEVSGLKEALGTLEV 489
Query: 598 ASDKLRKSRLFLKLLEAVLKTGNRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLHF 657
A KLR SRLFLKLLEAVLKTGNRMN GT+RG AQAF+LDTLLKLSDVKGTDGKTTLLHF
Sbjct: 490 ACKKLRNSRLFLKLLEAVLKTGNRMNVGTFRGDAQAFKLDTLLKLSDVKGTDGKTTLLHF 549
Query: 658 VVQEIIRSEGIRAVRTERESRSVVSSVGTDQDVDESPEA 696
VV EIIRSEG+RA+R +SRS S D + D SP++
Sbjct: 550 VVLEIIRSEGVRALRL--QSRSFSSVKTDDSNADSSPQS 586
>ref|NP_915167.1| putative formin-like protein AHF1 [Oryza sativa (japonica
cultivar-group)] gi|22093600|dbj|BAC06896.1| putative FH
protein NFH2 [Oryza sativa (japonica cultivar-group)]
gi|19386691|dbj|BAB86073.1| putative FH protein NFH2
[Oryza sativa (japonica cultivar-group)]
Length = 960
Score = 413 bits (1062), Expect = e-113
Identities = 260/590 (44%), Positives = 349/590 (59%), Gaps = 72/590 (12%)
Query: 340 NTSVVEATSSEGVGQVPEPPNGKPPPPPP---GPPPPPPPRKPP---------------- 380
NTS + +T++ +P P +PPPPP GPPPPPPP PP
Sbjct: 390 NTSALRSTTTTDT-TIPRNPFVQPPPPPTHTHGPPPPPPPPPPPPVGYWESRVRKPGTGT 448
Query: 381 -----APAPRPPPPP---RAGQPPPAPPKPVKVGNQGA------NQGGTSDGDAPKPKLK 426
+PA PPP ++G P A P + A G S+ P+PKLK
Sbjct: 449 SKETRSPALSPPPQAASFKSGLPTDAFPGRLADNADHAAAAAAGGGGDKSEETTPRPKLK 508
Query: 427 PFFWDKVAAKPDQQMVWHEISAGSFVFNEEKMESLFGCANQNRNERKKDS---PSVDTSV 483
P WDKV A D+ MVW ++ + SF NEE +E+LF C N K+ + P + T
Sbjct: 509 PLHWDKVRASSDRVMVWDQLKSSSFQVNEEMIETLFICNPANSAPPKEPATRRPVLPTPK 568
Query: 484 QYIQIIDPKKAQNLSILLRALNVTTDEVIDALREGN--EIPVELIQTLLKMAPTTEEELK 541
+++DPKK+QN++ILLRALNV+ ++V DAL EGN EL++TLLKMAPT EEE+K
Sbjct: 569 TDNKVLDPKKSQNIAILLRALNVSKEQVCDALCEGNTENFGAELLETLLKMAPTKEEEIK 628
Query: 542 LRLFTGELS--QLGPAERFLKVLVDIPFAFKRLETLLFMFILREEASSIKESFATLEVAS 599
LR F E S +LGPAE+FLK ++DIPFAFKR++ +L++ E + +K+SF TLE A
Sbjct: 629 LREFKEETSPIKLGPAEKFLKAVLDIPFAFKRVDAMLYIANFESEVNYLKKSFETLETAC 688
Query: 600 DKLRKSRLFLKLLEAVLKTGNRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLHFVV 659
D+LR SRLFLKLLEAVLKTGNRMN GT RG A AF+LDTLLKL DVKGTDGKTTLLHFVV
Sbjct: 689 DELRNSRLFLKLLEAVLKTGNRMNVGTNRGDAHAFKLDTLLKLVDVKGTDGKTTLLHFVV 748
Query: 660 QEIIRSEGIRAVRTERESRSVVSSVGTDQDVDESPEASEEHYRSLGLQVVSSLSNELEDV 719
QEIIR+EG S S+ T + +P E + LGLQVV+ L NEL +V
Sbjct: 749 QEIIRTEG---------SHLSASNQSTPR-TQANPLRDELECKKLGLQVVAGLGNELSNV 798
Query: 720 KRAALIDGDALSSAVSKLGHSLAKTQEF--MNTDLKSLDEESEFQRCTERFVEKAREEVT 777
K+AA +D D LSS VSKL + K E +N ++KS ++ F ++F+++A +++
Sbjct: 799 KKAAAMDSDVLSSYVSKLAGGIEKITEVLRLNEEVKSREDAWRFHDSMQKFLKRADDDII 858
Query: 778 WLQEEEKRIIAQVKSTADYFHGNAGRDEG--LRLFLIVRDFLIILDKVCKEVRDKTLRAV 835
+Q +E ++ VK +YFHG++ ++E R+F++VRDFL +LD+VCKEV R +
Sbjct: 859 RVQAQESVALSLVKEITEYFHGDSAKEEAHPFRIFMVVRDFLSVLDQVCKEVGRINDRTI 918
Query: 836 KSSQRKEALSVPSSPDTRHQQSPPAPSDLHRRLFPAIAGRR--IVEDSSS 883
SS R VP +P + +LFP I R I +D SS
Sbjct: 919 ASSVRH--FPVPVNP-------------MMPQLFPRIHALRAGISDDESS 953
Score = 49.3 bits (116), Expect = 6e-04
Identities = 22/48 (45%), Positives = 26/48 (53%)
Query: 369 GPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKPVKVGNQGANQGGTS 416
G PP PPP P AP PPPPPR P P+PP + N A + T+
Sbjct: 352 GQPPAPPPPPPFAPTLPPPPPPRRKPPSPSPPSSPLIENTSALRSTTT 399
Score = 48.5 bits (114), Expect = 0.001
Identities = 29/63 (46%), Positives = 32/63 (50%), Gaps = 4/63 (6%)
Query: 359 PNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKPVKVGNQGANQGGTSDG 418
P+ PP P PP P PP P P P PPPP AGQ P P V + N GA GG +
Sbjct: 42 PDQSSSPPTPAPPGPAPPFFPALPVP-PPPPATAGQEQPTYPALV-LPNTGA--GGAAAT 97
Query: 419 DAP 421
AP
Sbjct: 98 AAP 100
Score = 40.0 bits (92), Expect = 0.34
Identities = 28/79 (35%), Positives = 35/79 (43%), Gaps = 7/79 (8%)
Query: 170 PAGTVPSP-GPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHP 228
PA P P P L P P P+ SP S +P EN+S + + T P +P Q P
Sbjct: 355 PAPPPPPPFAPTLPPPPPPRRKPPSPSPPS-SPLIENTSALRSTTTTDTTIPRNPFVQPP 413
Query: 229 PPSPEKLQHVTYAPDPSPP 247
PP T+ P P PP
Sbjct: 414 PPPTH-----THGPPPPPP 427
Score = 38.1 bits (87), Expect = 1.3
Identities = 31/118 (26%), Positives = 43/118 (36%), Gaps = 10/118 (8%)
Query: 133 GSRYLLTKHSRSTLGLESGHHGRNLISDPPHNELSPSPAGTVPSPGPALESPTPNHGP-- 190
GS + T H +E+ R+ P + +P+ PS A SP P P
Sbjct: 273 GSSKMSTSHRTLAAAVEAAVAARDRSKSPSPGSIVSTPS--YPSSPGATMSPAPASPPLF 330
Query: 191 -TPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHPPPSPEKLQHVTYAPDPSPP 247
+P S + +S + P PP P PPP P + P PSPP
Sbjct: 331 SSPGQSGRRSVKSRSDSVRTFGQPPAPPPPPPFAPTLPPPPPPRR-----KPPSPSPP 383
Score = 37.7 bits (86), Expect = 1.7
Identities = 29/96 (30%), Positives = 36/96 (37%), Gaps = 20/96 (20%)
Query: 161 PPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMV-- 218
PP P A T+P P P P P + PSSP ++ S+T + T +
Sbjct: 354 PPAPPPPPPFAPTLPPP------PPPRRKPPSPSPPSSPLIENTSALRSTTTTDTTIPRN 407
Query: 219 ----PPTSPPRQHPPPSPEKLQHVTYAPDPSPPDKG 250
PP P H PP P P P PP G
Sbjct: 408 PFVQPPPPPTHTHGPPPP--------PPPPPPPPVG 435
>gb|AAQ99143.1| formin-like protein AtFH6 [Arabidopsis thaliana]
gi|9757868|dbj|BAB08455.1| formin-like protein
[Arabidopsis thaliana] gi|15240762|ref|NP_201548.1|
formin homology 2 domain-containing protein / FH2
domain-containing protein [Arabidopsis thaliana]
Length = 899
Score = 395 bits (1014), Expect = e-108
Identities = 248/590 (42%), Positives = 341/590 (57%), Gaps = 75/590 (12%)
Query: 340 NTSVVEATSSEGVGQVPEPPNGKPPPP---PPGPPPPPPPRKPPAPAPRP---------- 386
N + +A + E + P PP + PPP PP PPPPPP PP P RP
Sbjct: 352 NRAAFQAITQE---KSPVPPPRRSPPPLQTPPPPPPPPPLAPPPPPQKRPRDFQMLRKVT 408
Query: 387 -----------PPPPRAGQPPPAPPKPVKVGNQ----GANQGGTSDGDAPKPKLKPFFWD 431
P +A + P K V+ N + G D D KPKLKP WD
Sbjct: 409 NSEATTNSTTSPSRKQAFKTPSPKTKAVEEVNSVSAGSLEKSGDGDTDPSKPKLKPLHWD 468
Query: 432 KVAAKPDQQMVWHEISAGSFVFNEEKMESLFGCANQNRNERKKDSPSV-DTSVQYIQIID 490
KV A D+ VW ++ + SF NE++ME LFGC + + ++ SV + +++D
Sbjct: 469 KVRASSDRATVWDQLKSSSFQLNEDRMEHLFGCNSGSSAPKEPVRRSVIPLAENENRVLD 528
Query: 491 PKKAQNLSILLRALNVTTDEVIDALREGN--EIPVELIQTLLKMAPTTEEELKLRLFTGE 548
PKK+QN++ILLRALNVT +EV +AL +GN + EL++TL+KMAPT EEE+KLR ++G+
Sbjct: 529 PKKSQNIAILLRALNVTREEVSEALTDGNPESLGAELLETLVKMAPTKEEEIKLREYSGD 588
Query: 549 LSQLGPAERFLKVLVDIPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLF 608
+S+LG AERFLK ++DIPFAFKR+E +L+ E ++ SF TLE AS +L+ SRLF
Sbjct: 589 VSKLGTAERFLKTILDIPFAFKRVEAMLYRANFDAEVKYLRNSFQTLEEASLELKASRLF 648
Query: 609 LKLLEAVLKTGNRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGI 668
LKLLEAVL TGNRMN GT RG A AF+LDTLLKL D+KG DGKTTLLHFVVQEI RSEG
Sbjct: 649 LKLLEAVLMTGNRMNVGTNRGDAIAFKLDTLLKLVDIKGVDGKTTLLHFVVQEITRSEGT 708
Query: 669 RAVRTERESRSVVSSVGTDQDVDESPEASEEHYRSLGLQVVSSLSNELEDVKRAALIDGD 728
+ E + + +R GLQVV+ LS +L +VK++A +D D
Sbjct: 709 TTTKDE-----------------TILHGNNDGFRKQGLQVVAGLSRDLVNVKKSAGMDFD 751
Query: 729 ALSSAVSKLGHSLAKTQEFMNTDLKSLDEESEFQRCTERFVEKAREEVTWLQEEEKRIIA 788
LSS V+KL L K + F+ T+ + F + F+++A EE+ ++ E++ ++
Sbjct: 752 VLSSYVTKLEMGLDKLRSFLKTE----TTQGRFFDSMKTFLKEAEEEIRKIKGGERKALS 807
Query: 789 QVKSTADYFHGNAGRDEG--LRLFLIVRDFLIILDKVCKEVRDKTLRAVKSSQRKEALSV 846
VK +YFHGNA R+E LR+F++VRDFL +LD VCKEV KT++ + +S +
Sbjct: 808 MVKEVTEYFHGNAAREEAHPLRIFMVVRDFLGVLDNVCKEV--KTMQEMSTS-----MGS 860
Query: 847 PSSPDTRHQQSPPAPSDLHRRLFPAIAGRRIVEDSSSDDDDDDHKEGSST 896
S+ R + P LHR + +D+SS D +H SST
Sbjct: 861 ASARSFRISATASLPV-LHRY-------KARQDDTSS---DSEHSSNSST 899
Score = 40.8 bits (94), Expect = 0.20
Identities = 53/232 (22%), Positives = 78/232 (32%), Gaps = 41/232 (17%)
Query: 196 PSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHPPPSPEKLQHVTYAPDPSPPDKGD---- 251
P S TP P + STPSP P P P +P++ T P P PP D
Sbjct: 37 PESSTPPPPDFQ--STPSPPLPDTPDQPFFPENPSTPQQ----TLFPPPPPPVSADVNGG 90
Query: 252 -------------RKQKAIILAVTVSGIIIMIGLALCYRESRRSDRADKDEGPLLILTSS 298
K+ AI+++V + + ++ LA + +D + ++T
Sbjct: 91 LPIPTATTQSAKPGKKVAIVISVGIVTLGMLSALAFFLYRHKAKHASDTQK----LVTGG 146
Query: 299 DYSGGKQKVVRLGNANNEGSGISNGKKPSNVRNWSMKAGDNNTSVVEATSSEGV--GQVP 356
GG ++ E SG P+ + + G + V A+ S G G V
Sbjct: 147 GDGGGSRRF-------QEDSG-----PPTTTSSTFLYMGTVEPTRVSASESNGGTNGPVN 194
Query: 357 EPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKPVKVGNQ 408
P K P P P P PP P P P G +
Sbjct: 195 SSPYRKLNSAKRSERYRPSPELQPLPPLAKPPQPSDNSPSALSPSSSSSGEE 246
Score = 37.4 bits (85), Expect = 2.2
Identities = 38/140 (27%), Positives = 52/140 (37%), Gaps = 41/140 (29%)
Query: 99 PKTTKSLSPVMRKKVLDCIRRSTPPA---PPVSQKKTGSRYLLTKHSRSTLGLESGHHGR 155
P T S SP M+ ++ I++ PP PP+ GLES
Sbjct: 294 PTTAASRSPEMKHVIIPSIKQKLPPPVQPPPLR------------------GLESDEQ-- 333
Query: 156 NLISDPPHNELSPSPAGTVPSPG-PALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSP 214
+ P+++ P + P P A ++ T P P S P P TP P
Sbjct: 334 ----ELPYSQNKPKFSQPPPPPNRAAFQAITQEKSPVPPPRRSPP--------PLQTPPP 381
Query: 215 TAMVPPTSPPRQHPPPSPEK 234
PP PP PPP P+K
Sbjct: 382 ----PPPPPPLA-PPPPPQK 396
Score = 35.8 bits (81), Expect = 6.5
Identities = 25/71 (35%), Positives = 28/71 (39%), Gaps = 4/71 (5%)
Query: 359 PNGKPPPPPP--GPPPPPPPRKPPAPA-PRPPPPPRAGQPPPAPPKPVKVGNQGANQGGT 415
P PPPP P PP P P P P P P+ PP PP PV G T
Sbjct: 37 PESSTPPPPDFQSTPSPPLPDTPDQPFFPENPSTPQQTLFPP-PPPPVSADVNGGLPIPT 95
Query: 416 SDGDAPKPKLK 426
+ + KP K
Sbjct: 96 ATTQSAKPGKK 106
>gb|AAP53183.1| putative formin-like protein [Oryza sativa (japonica
cultivar-group)] gi|37533188|ref|NP_920896.1| putative
formin-like protein [Oryza sativa (japonica
cultivar-group)] gi|22748365|gb|AAN05367.1| Putative
formin-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 849
Score = 394 bits (1012), Expect = e-108
Identities = 232/522 (44%), Positives = 312/522 (59%), Gaps = 60/522 (11%)
Query: 354 QVPEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAP--------PKPVK- 404
Q+ PP+ PP PPP PPPPP + P+PPPPP +PP P P P +
Sbjct: 319 QMASPPSNPPPAPPP---PPPPPSRFNNTTPKPPPPPPPPEPPTGPVSARRLLRPLPAEG 375
Query: 405 --------------------VGNQGANQGGTSDGDAPKPKLKPFFWDKVAAKPDQQMVWH 444
G + GD P+PKLKP WDKV D+ MVW
Sbjct: 376 PSIVIPRAPAMAVTKDNDATAATMSVRTRGEAAGDEPRPKLKPLHWDKVRTSSDRDMVWD 435
Query: 445 EISAGSFVFNEEKMESLFGCANQNRNERKKDSPSVDTSVQYIQ---IIDPKKAQNLSILL 501
+ +E+ +E LF N + D+P Q+ Q ++DPKKAQN++ILL
Sbjct: 436 RLK-----LDEDMIEVLF-MNNSTAVAPRMDNPKKVGMPQFKQEERVLDPKKAQNIAILL 489
Query: 502 RALNVTTDEVIDALREGNE--IPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAERFL 559
RALNVT +EV DAL +GN + EL++TL+KMAPT EEELKLR FTG+LS+LG AERFL
Sbjct: 490 RALNVTLEEVTDALLDGNAECLGAELLETLVKMAPTKEEELKLRDFTGDLSKLGSAERFL 549
Query: 560 KVLVDIPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTG 619
K ++DIPFAFKR++ +L+ E + +++SF TLE A D L+ SRLFLKLLEAVL+TG
Sbjct: 550 KAVLDIPFAFKRVDVMLYRANFENEVNYLRKSFQTLEAACDDLKGSRLFLKLLEAVLRTG 609
Query: 620 NRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGIRAVRTERESRS 679
NRMN GT RG A+AF+LDTLLKL+DVKG DGKTTLLHFVVQEI+RSE ++E+ +
Sbjct: 610 NRMNVGTNRGEAKAFKLDTLLKLADVKGADGKTTLLHFVVQEIVRSED---AKSEKAPEN 666
Query: 680 VVSSVGTDQDVDESPEASEEHYRSLGLQVVSSLSNELEDVKRAALIDGDALSSAVSKLGH 739
++++ A E R GL+VVS LS EL +VKRAA +D D L VSKL
Sbjct: 667 HITNI-----------AKVEQLRRQGLKVVSGLSTELGNVKRAATMDFDVLHGYVSKLEA 715
Query: 740 SLAKTQEFMNTDLKSLDEESEFQRCTERFVEKAREEVTWLQEEEKRIIAQVKSTADYFHG 799
L K + + + K + F F+++A +E+ ++ +EK + +VK +YFHG
Sbjct: 716 GLGKIKSVLQLE-KQCSQGVNFFATMREFLKEAEQEIEQVRHDEKAALGRVKEITEYFHG 774
Query: 800 NAGRDEG--LRLFLIVRDFLIILDKVCKEVRDKTLRAVKSSQ 839
NA ++E LR+F++VRDFL +LD VC+EV + V S++
Sbjct: 775 NAVKEEAHPLRIFMVVRDFLSMLDHVCREVSQQDRTFVGSAR 816
Score = 43.9 bits (102), Expect = 0.024
Identities = 23/84 (27%), Positives = 35/84 (41%), Gaps = 2/84 (2%)
Query: 164 NELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSP 223
++ P+ P P PA +P P P S +P T ++ + + PP++P
Sbjct: 270 SDFFPTTPAAAPVPAPAAAAPPP--APPAPRSRRTPPRTRFSAGSGAEMNKQMASPPSNP 327
Query: 224 PRQHPPPSPEKLQHVTYAPDPSPP 247
P PPP P + P P PP
Sbjct: 328 PPAPPPPPPPPSRFNNTTPKPPPP 351
Score = 42.4 bits (98), Expect = 0.069
Identities = 25/81 (30%), Positives = 31/81 (37%), Gaps = 21/81 (25%)
Query: 185 TPNHGPTPANSPSSPTPTPENSSPSSTP------------------SPTAMVPPTSPPRQ 226
TP P PA + ++P P P TP SP + PP PP
Sbjct: 276 TPAAAPVPAPAAAAPPPAPPAPRSRRTPPRTRFSAGSGAEMNKQMASPPSNPPPAPPP-- 333
Query: 227 HPPPSPEKLQHVTYAPDPSPP 247
PPP P + + T P P PP
Sbjct: 334 -PPPPPSRFNNTTPKPPPPPP 353
Score = 37.4 bits (85), Expect = 2.2
Identities = 32/134 (23%), Positives = 49/134 (35%), Gaps = 18/134 (13%)
Query: 89 RCIHLLENEIPKTTKSLSPVMRKKVLDCIRRSTPPAPPVSQKKTGSRYLLTKHSRSTLGL 148
R + L ++ TT + +PV + PPAPP + + R+
Sbjct: 263 RSLPSLTSDFFPTTPAAAPVPAPAAA-----APPPAPPAPRSRRTP-------PRTRFSA 310
Query: 149 ESGHHGRNLISDPPHNELSPSPAGTVPSPGPAL---ESPTPNHGPTPANSPSSPTPTPEN 205
SG ++ PP N P PA P P P+ +P P P P P+ P
Sbjct: 311 GSGAEMNKQMASPPSN---PPPAPPPPPPPPSRFNNTTPKPPPPPPPPEPPTGPVSARRL 367
Query: 206 SSPSSTPSPTAMVP 219
P P+ ++P
Sbjct: 368 LRPLPAEGPSIVIP 381
Score = 37.4 bits (85), Expect = 2.2
Identities = 25/78 (32%), Positives = 31/78 (39%), Gaps = 16/78 (20%)
Query: 168 PSPAGTVPSPGPALESPTPNHGPTPA----NSPSSPTPTPENSSPSSTPSPTAMVPPTSP 223
P+PA P P P P P TP ++ S + +SP S P P PP P
Sbjct: 283 PAPAAAAPPPAP----PAPRSRRTPPRTRFSAGSGAEMNKQMASPPSNPPPAPPPPPPPP 338
Query: 224 PRQH--------PPPSPE 233
R + PPP PE
Sbjct: 339 SRFNNTTPKPPPPPPPPE 356
Score = 35.4 bits (80), Expect = 8.5
Identities = 28/98 (28%), Positives = 35/98 (35%), Gaps = 24/98 (24%)
Query: 156 NLISD--PPHNELSPSPAGTVPSPGPALESPTPNHGPT-------------------PAN 194
+L SD P +P PA +P PA +P P P+N
Sbjct: 267 SLTSDFFPTTPAAAPVPAPAAAAPPPAPPAPRSRRTPPRTRFSAGSGAEMNKQMASPPSN 326
Query: 195 SPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHPPPSP 232
P +P P P P S + T PP PP PP P
Sbjct: 327 PPPAPPPPP---PPPSRFNNTTPKPPPPPPPPEPPTGP 361
Score = 35.4 bits (80), Expect = 8.5
Identities = 14/24 (58%), Positives = 14/24 (58%)
Query: 379 PPAPAPRPPPPPRAGQPPPAPPKP 402
P PA P P P A PPPAPP P
Sbjct: 274 PTTPAAAPVPAPAAAAPPPAPPAP 297
>gb|AAF64546.1| unknown protein [Arabidopsis thaliana] gi|15229995|ref|NP_187198.1|
formin homology 2 domain-containing protein / FH2
domain-containing protein [Arabidopsis thaliana]
Length = 884
Score = 384 bits (987), Expect = e-105
Identities = 239/597 (40%), Positives = 330/597 (55%), Gaps = 82/597 (13%)
Query: 282 SDRADKDEGPLLILTSSDYSGGKQKVVRLGNANNEGSGISNGKKPSNVRNWSMKAGDNNT 341
+D +D DE + S YS RL NA++ ++ G S + ++
Sbjct: 305 NDSSDDDESFHSVGGGSQYSNP-----RLSNASSASGSVNVGS--------SQRFSEHKL 351
Query: 342 SVVEATSSEGVGQVPEPPNGKPPPPPPGPPPPPP-------------------------- 375
+ E + S+ V PPPPPP PPP P
Sbjct: 352 DIPECSRSDFGISV-----SAPPPPPPPPPPLPQFSNKRIHTLSSPETANLQTLSSQLCE 406
Query: 376 -----------PRKPPAPAPRPPPPPRAGQPPP----------APPKPVKVGNQGANQGG 414
P P PRPPPPP PPP PP P+ + G
Sbjct: 407 KLCASSSKTSFPINVPNSQPRPPPPP----PPPQQLQVAGINKTPPPPLSLDFSERRPLG 462
Query: 415 TSDGDAPKPKLKPFFWDKVAAKPDQQMVWHEISAGSFVFNEEKMESLFGCANQNRNERKK 474
DG AP PKLKP WDKV A PD+ MVW ++ SF +EE +ESLFG Q+ + ++
Sbjct: 463 -KDG-APLPKLKPLHWDKVRATPDRTMVWDKLRTSSFELDEEMIESLFGYTMQSSTKNEE 520
Query: 475 DSPSVDTSVQYIQIIDPKKAQNLSILLRALNVTTDEVIDALREGNEIPVELIQTLLKMAP 534
+ +++ ++PK+ QN +ILL+ALN T D++ AL +G + ++ ++ L+KM P
Sbjct: 521 GKSKTPSPGKHL--LEPKRLQNFTILLKALNATADQICSALGKGEGLCLQQLEALVKMVP 578
Query: 535 TTEEELKLRLFTGELSQLGPAERFLKVLVDIPFAFKRLETLLFMFILREEASSIKESFAT 594
T EEELKLR + G + +LG AE+FL+ LV +PFAF+R E +L+ +E ++ SF+
Sbjct: 579 TKEEELKLRSYKGAVDELGSAEKFLRALVGVPFAFQRAEAMLYRETFEDEVVHLRNSFSM 638
Query: 595 LEVASDKLRKSRLFLKLLEAVLKTGNRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTL 654
LE A +L+ SRLFLKLLEAVLKTGNRMN GT RGGA+AF+LD LLKLSDVKGTDGKTTL
Sbjct: 639 LEEACKELKSSRLFLKLLEAVLKTGNRMNVGTIRGGAKAFKLDALLKLSDVKGTDGKTTL 698
Query: 655 LHFVVQEIIRSEGIRAVRTERESRSVVSSVGTDQ-DVDESPEASEEHYRSLGLQVVSSLS 713
LHFVVQEI RSEGIR S S++ + + + + +PE EE YR +GL +VS L+
Sbjct: 699 LHFVVQEISRSEGIRV------SDSIMGRIMNQRSNKNRTPEEKEEDYRRMGLDLVSGLN 752
Query: 714 NELEDVKRAALIDGDALSSAVSKLGHSLAKTQEFMNTDLKSLDEESEFQRCTERFVEKAR 773
EL +VK+ A ID + L ++VS L L + + LK +E F F+
Sbjct: 753 TELRNVKKTATIDLEGLVTSVSNLRDGLGQLSCLASEKLKGDEENRAFVSSMSSFLRYGE 812
Query: 774 EEVTWLQEEEKRIIAQVKSTADYFHGNAGRDE--GLRLFLIVRDFLIILDKVCKEVR 828
+ + L+E+EKRI+ +V A+YFHG+ DE LR+F+IVRDFL +LD VC+E+R
Sbjct: 813 KSLEELREDEKRIMERVGEIAEYFHGDVRGDEKNPLRIFVIVRDFLGMLDHVCRELR 869
>gb|AAN13148.1| putative formin protein AHF1 [Arabidopsis thaliana]
gi|19423899|gb|AAL87275.1| putative formin protein AHF1
[Arabidopsis thaliana] gi|9279730|dbj|BAB01320.1|
formin-like protein [Arabidopsis thaliana]
gi|6503010|gb|AAF14548.1| formin-like protein AHF1
[Arabidopsis thaliana] gi|15230845|ref|NP_189177.1|
formin homology 2 domain-containing protein / FH2
domain-containing protein [Arabidopsis thaliana]
Length = 1051
Score = 383 bits (983), Expect = e-104
Identities = 272/729 (37%), Positives = 380/729 (51%), Gaps = 80/729 (10%)
Query: 198 SPTPTPENSSPSSTPSPTAMVPPTSP-PRQHPPPSPEK-LQHVTYAPDPSPPDKGDRKQK 255
SPT T +SP + + + TS P + P +PE L+ +++ + P + +K
Sbjct: 360 SPTVTSLTTSPENNKKENSPLSSTSTSPERRPNDTPEAYLRSPSHSSASTSPYRCFQKSP 419
Query: 256 AIILAVTVSGIIIMIGLALCYRESRRSDRADKDEGP-----LLILTSSDYSGGKQKVVRL 310
++ A M L + S ++ G L L S S V
Sbjct: 420 EVLPA-------FMSNLRQGLQSQLLSSPSNSHGGQGFLKQLDALRSRSPSSSSSSVCSS 472
Query: 311 GNANNEGSGISNGKKPSNVRNWSMKAGDNNTSVVEATSSEGVGQVPEPPNGKPPPPPPGP 370
+ S +++ K S + D + S S P+ + PPPPP P
Sbjct: 473 PEKASHKSPVTSPKLSSRNSQSLSSSPDRDFSHSLDVSPRISNISPQILQSRVPPPPPPP 532
Query: 371 PPPP-------------PPRKPPAPAPRPPPPPRAGQPPPAPPKPVKVGNQGANQGGTSD 417
PP P +PP+ P P + P P++ +
Sbjct: 533 PPLPLWGRRSQVTTKADTISRPPSLTPPSHPFVIPSENLPVTSSPMETPETVCASEAAEE 592
Query: 418 GDAPKPKLKPFFWDKVAAKPDQQMVWHEISAGSFVFNEEKMESLFGCANQNRNERKKDSP 477
PKPKLK WDKV A D++MVW + + SF +EE +E+LF + N + +
Sbjct: 593 --TPKPKLKALHWDKVRASSDREMVWDHLRSSSFKLDEEMIETLFVAKSLNNKPNQSQTT 650
Query: 478 S---VDTSVQYIQIIDPKKAQNLSILLRALNVTTDEVIDALREGNE--IPVELIQTLLKM 532
+ + Q +++DPKKAQN++ILLRALNVT +EV +AL EGN + EL+++LLKM
Sbjct: 651 PRCVLPSPNQENRVLDPKKAQNIAILLRALNVTIEEVCEALLEGNADTLGTELLESLLKM 710
Query: 533 APTTEEELKLRLFTGELS-QLGPAERFLKVLVDIPFAFKRLETLLFMFILREEASSIKES 591
APT EEE KL+ + + +LG AE+FLK ++DIPFAFKR++ +L++ E +K+S
Sbjct: 711 APTKEEERKLKAYNDDSPVKLGHAEKFLKAMLDIPFAFKRVDAMLYVANFESEVEYLKKS 770
Query: 592 FATLEVASDKLRKSRLFLKLLEAVLKTGNRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGK 651
F TLE A ++LR SR+FLKLLEAVLKTGNRMN GT RG A AF+LDTLLKL DVKG DGK
Sbjct: 771 FETLEAACEELRNSRMFLKLLEAVLKTGNRMNVGTNRGDAHAFKLDTLLKLVDVKGADGK 830
Query: 652 TTLLHFVVQEIIRSEGIRAVRTERESRSVVSSVGTDQDVDESPEASEEHYRSLGLQVVSS 711
TTLLHFVVQEIIR+EG R +S T D + R LGLQVVSS
Sbjct: 831 TTLLHFVVQEIIRAEGTR-----------LSGNNTQTD--------DIKCRKLGLQVVSS 871
Query: 712 LSNELEDVKRAALIDGDALSSAVSKLGHSLAKTQEFMNTDLKSLDEESEFQRCTE---RF 768
L +EL +VK+AA +D + LSS VSKL +AK E + ++ EES QR +E F
Sbjct: 872 LCSELSNVKKAAAMDSEVLSSYVSKLSQGIAKINEAIQVQ-STITEESNSQRFSESMKTF 930
Query: 769 VEKAREEVTWLQEEEKRIIAQVKSTADYFHGNAGRDEG--LRLFLIVRDFLIILDKVCKE 826
+++A EE+ +Q +E ++ VK +YFHGN+ ++E R+FL+VRDFL ++D+VCKE
Sbjct: 931 LKRAEEEIIRVQAQESVALSLVKEITEYFHGNSAKEEAHPFRIFLVVRDFLGVVDRVCKE 990
Query: 827 VRDKTLRAVKSSQRKEALSVPSSPDTRHQQSPPAPSDLHRRLFPAIAGRRIVEDSS---- 882
V R + SS K VP +P P P P + GRR SS
Sbjct: 991 VGMINERTMVSSAHK--FPVPVNP------MMPQP-------LPGLVGRRQSSSSSSSSS 1035
Query: 883 -SDDDDDDH 890
S D+D+H
Sbjct: 1036 TSSSDEDEH 1044
Score = 39.3 bits (90), Expect = 0.59
Identities = 31/104 (29%), Positives = 50/104 (47%), Gaps = 13/104 (12%)
Query: 191 TPANSPSSPTPTPENSSPSSTPSPTAMVPPTSP-----PRQHPPPSPEKLQ----HVTYA 241
+P SP SP P P+ S+TP P++ P SP P PPPSP +++
Sbjct: 35 SPPPSPPSPPPLPKLPFSSTTP-PSSSDPNASPFFPLYPSSPPPPSPASFASFPANISSL 93
Query: 242 PDPSPPDKGDRKQKAIILAVTV--SGIIIMIGLALCY-RESRRS 282
P +K +I+A++ S ++ + +AL Y R S+R+
Sbjct: 94 IVPHATKSPPNSKKLLIVAISAVSSAALVALLIALLYWRRSKRN 137
Score = 35.8 bits (81), Expect = 6.5
Identities = 19/54 (35%), Positives = 21/54 (38%), Gaps = 8/54 (14%)
Query: 366 PPPGPPPPPPPRKPPAPAPRPPPPPRAGQPP--------PAPPKPVKVGNQGAN 411
PPP PP PPP K P + PP P P PP P + AN
Sbjct: 36 PPPSPPSPPPLPKLPFSSTTPPSSSDPNASPFFPLYPSSPPPPSPASFASFPAN 89
>dbj|BAD33833.1| diaphanous protein-like [Oryza sativa (japonica cultivar-group)]
Length = 788
Score = 373 bits (958), Expect = e-101
Identities = 262/721 (36%), Positives = 376/721 (51%), Gaps = 73/721 (10%)
Query: 170 PAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHPP 229
P P P + +P P G + + T + SS SS+ +A PT+P
Sbjct: 70 PDSAPPQLPPPVTTPAPAGGAGDGGTDAGAAATGDASSSSSS---SASPHPTAPANISYM 126
Query: 230 PSPEKLQHVTYAPDPSPPDKGDRKQKAIILAVTVSGIIIMIGLALCYRESRRSDRADKDE 289
P Y P + ++L V + ++ AL Y +RR R K E
Sbjct: 127 AMP------IYHSAPLRSFLSSHRLLTVLLPVAAV-LAAVLAAALVYLLTRRR-RCSKGE 178
Query: 290 GPLLILTSSDYSGGKQKVVRLGNANNEGSG-----ISNGKKPS---------NVRNWSMK 335
+ S G + G+ + G G S+ P +
Sbjct: 179 PHAAHTKAVLLSPGNSTALYDGDHDQHGRGSTATAASSASSPELRPMPPLPRQFQQTRTS 238
Query: 336 AGDNNTSVVEATSSEGVGQVPEPPNGKPPPPPPGPPPPPPPRKPPAP---APRPPPP-PR 391
+ ++ EA + + P+ PPPPPP PPPP PPR A AP PPPP PR
Sbjct: 239 MPSTSQTIHEAGAEDKRAPPPQSVRPPPPPPPPPPPPPMPPRTDNASTQAAPAPPPPLPR 298
Query: 392 AGQPPPAPPK-------------------PVKVGNQGANQGGTSDGDAPKPKLKPFFWDK 432
AG P+ P + + + + +D A +PKLKP WDK
Sbjct: 299 AGNGSGWLPRRYTERAAPTVIRASAGAVHPEESPARASPEEKAADA-AARPKLKPLHWDK 357
Query: 433 VA-AKPDQQMVWHEISAGSFVFNEEKMESLFGCANQNRNERKKDSPSVDTSV--QYIQII 489
V A + VW ++ A SF NEE +E+LF +N R K + + Q +++
Sbjct: 358 VRPASSGRPTVWDQLKASSFRVNEEMIETLF-VSNSTRRASKNGVKEANAACCNQENKVL 416
Query: 490 DPKKAQNLSILLRALNVTTDEVIDALREGN--EIPVELIQTLLKMAPTTEEELKLRLFTG 547
DPKK+QN++I+LRAL+ T +EV AL +G + EL++TLLKMAP+ EEE+KL+ F
Sbjct: 417 DPKKSQNIAIMLRALDATKEEVCKALLDGQAESLGTELLETLLKMAPSREEEIKLKEFRE 476
Query: 548 E-LSQLGPAERFLKVLVDIPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSR 606
+ +S+LGPAE FLK ++ IPFAFKR+E +L++ E +K SF TLE A ++LR SR
Sbjct: 477 DAVSKLGPAESFLKAVLAIPFAFKRVEAMLYIANFDSEVDYLKTSFKTLEAACEELRGSR 536
Query: 607 LFLKLLEAVLKTGNRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSE 666
LF K+L+AVLKTGNRMN GT RG A AF+LD LLKL DVKG DGKTTLLHFV++EI++SE
Sbjct: 537 LFHKILDAVLKTGNRMNTGTNRGNASAFKLDALLKLVDVKGADGKTTLLHFVIEEIVKSE 596
Query: 667 GIRAVRTERESRSVVSSVGTDQDVDESPEASEEHYRSLGLQVVSSLSNELEDVKRAALID 726
G + T + S S A + + +GL++V+SL EL +VK+AA +D
Sbjct: 597 GASILATGQTSN------------QGSAIADDFQCKKVGLRIVASLGGELGNVKKAAGMD 644
Query: 727 GDALSSAVSKLGHSLAKTQEF--MNTDLKSLDEESEFQRCTERFVEKAREEVTWLQEEEK 784
D L+S V+KL ++K E +N L S D F+ F++KA E+T +Q +E
Sbjct: 645 SDTLASCVAKLSAGVSKISEALQLNQQLGSDDHCKRFRASIGEFLQKAEAEITAVQAQES 704
Query: 785 RIIAQVKSTADYFHGNAGRDEG--LRLFLIVRDFLIILDKVCKEV-RDKTLRAVKSSQRK 841
++ V+ T ++FHG++ ++EG LR+F++VRDFL +LD VCK+V R A+ SS R
Sbjct: 705 LALSLVRETTEFFHGDSVKEEGHPLRIFMVVRDFLTVLDHVCKDVGRMNERTAIGSSLRL 764
Query: 842 E 842
E
Sbjct: 765 E 765
Score = 43.9 bits (102), Expect = 0.024
Identities = 29/76 (38%), Positives = 34/76 (44%), Gaps = 7/76 (9%)
Query: 348 SSEGVGQVPEP--PNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKPV-K 404
SS ++ EP P P PP PPPPP P P P PP P PP P P
Sbjct: 34 SSGSRRELHEPLFPLENAPALPPPPPPPPAPFFPFLPDSAPPQLP----PPVTTPAPAGG 89
Query: 405 VGNQGANQGGTSDGDA 420
G+ G + G + GDA
Sbjct: 90 AGDGGTDAGAAATGDA 105
>gb|AAF24497.1| FH protein NFH2 [Nicotiana tabacum]
Length = 835
Score = 363 bits (933), Expect = 1e-98
Identities = 288/830 (34%), Positives = 398/830 (47%), Gaps = 160/830 (19%)
Query: 181 LESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHPPPSP-----EKL 235
++SP P+ P PA P+ PTP P S TP PT R PPP P +
Sbjct: 39 IDSPPPSQPPIPA-PPAPPTPYPFQPS---TPDNNNPFFPTY--RSPPPPPPPPSPSSLV 92
Query: 236 QHVTYAPDPSPPDKGDRK---QKAIILAVT-VSGIIIMIGLALCYR-----------ESR 280
D + P+ K K II A+T V II++ +A+C +++
Sbjct: 93 SFPANISDINLPNTSKSKHVSSKLIITAITCVLAAIIVLSIAICLHAKKRRRHFNDPKTQ 152
Query: 281 RSDRAD-------KDEGPL--LILTSSDYSGGKQKVVRLGNANNEGSGISNGKKPS---- 327
RSD ++ K++G I S + + LG N GI++G P
Sbjct: 153 RSDNSNRLNHGSSKNDGNTNNSIPKLQQPSQTSSEFLYLGTIVNSHGGINSGSNPDTAPS 212
Query: 328 ---------------NVRNWSMKAGDNNT---------SVVEATSSEGVGQ--------- 354
N RN S + SV SS G G
Sbjct: 213 SRKMASPELRPLPPLNGRNLSQNYRNTRNDDDFYSTEESVGYIESSFGAGSLSRRGFAAV 272
Query: 355 -----VPEPPNGKPPPPPPG-------------PPPPPPPRKPPAPAPRPP-------PP 389
V +G G PP P++ P+ P PP
Sbjct: 273 EVNKFVGSSLSGSDSSSSSGSGSPNRSVSLSISPPVSVSPKRESCSRPKSPELIAVVTPP 332
Query: 390 PRAGQPPPAPP-------------KPV----------KVGNQGANQGGTSDGDAPKPKLK 426
P PPP PP PV V N + + KPKLK
Sbjct: 333 PPQRPPPPPPPFVHGPQVKVTANESPVLISPMEKNDQNVENHSIEKNEEKSEEILKPKLK 392
Query: 427 PFFWDKVAAKPDQQMVWHEISAGSFVFNEEKMESLFGCANQNRNERKKDSPSVDTSV-QY 485
WDKV A D +MVW ++ + SF NEE +E+LF N N V +S+ Q
Sbjct: 393 TLHWDKVRASSDCEMVWDQLKSSSFKLNEEMIETLFVVKNPTLNTSATAKHFVVSSMSQE 452
Query: 486 IQIIDPKKAQNLSILLRALNVTTDEVIDALREGN--EIPVELIQTLLKMAPTTEEELKLR 543
+++DPKK+QN++ILLR LN TT+E+ +A EGN I EL++ LLKMAP+ EEE KL+
Sbjct: 453 NRVLDPKKSQNIAILLRVLNGTTEEICEAFLEGNAENIGTELLEILLKMAPSKEEERKLK 512
Query: 544 LFTGELS-QLGPAERFLKVLVDIPFAFKRLETLLFMFILREEASSIKESFATLEVASDKL 602
+ + +LGPAE+FLK ++DIPFAFKR++ +L++ E + SF TLE A ++L
Sbjct: 513 EYKDDSPFKLGPAEKFLKAVLDIPFAFKRIDAMLYISNFDYEVDYLGNSFETLEAACEEL 572
Query: 603 RKSRLFLKLLEAVLKTGNRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEI 662
R SR+FLKLLEAVLKTGNRMN GT RG A AF+LDTLLKL DVKG DGKTTLLHFVVQEI
Sbjct: 573 RSSRMFLKLLEAVLKTGNRMNVGTNRGDAHAFKLDTLLKLVDVKGADGKTTLLHFVVQEI 632
Query: 663 IRSEGIRAVRTERESRSVVSSVGTDQDVDESPEASEEHYRSLGLQVVSSLSNELEDVKRA 722
I+SEG R G +Q+ +S + + LGLQVVS++S+EL +VK++
Sbjct: 633 IKSEGARL-------------SGGNQNHQQSTTNDDAKCKKLGLQVVSNISSELINVKKS 679
Query: 723 ALIDGDALSSAVSKLGHSLAKTQEFMNT-DLKSLDEES--EFQRCTERFVEKAREEVTWL 779
A +D + L + V KL + E + + + L+E S F RF++ A E++ L
Sbjct: 680 AAMDSEVLHNDVLKLSKGIQNIAEVVRSIEAVGLEESSIKRFSESMNRFMKVAEEKILRL 739
Query: 780 QEEEKRIIAQVKSTADYFHGNAGRDEG--LRLFLIVRDFLIILDKVCKEVRDKTLRAVKS 837
Q +E ++ VK +Y HG++ R+E R+F++V+DFL+ILD VCKEV R + S
Sbjct: 740 QAQETLAMSLVKEITEYVHGDSAREEAHPFRIFMVVKDFLMILDCVCKEVGTINERTIVS 799
Query: 838 SQRKEALSVPSSPDTRHQQSPPAPSDLHRRLFPAIAGRRIVEDSSSDDDD 887
S +K VP +P+ L P I+G R SS D++
Sbjct: 800 SAQK--FPVPVNPN----------------LQPVISGFRAKRLHSSSDEE 831
Score = 46.2 bits (108), Expect = 0.005
Identities = 23/59 (38%), Positives = 29/59 (48%), Gaps = 2/59 (3%)
Query: 355 VPEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPP--PPPRAGQPPPAPPKPVKVGNQGAN 411
+ PP +PP P P PP P P +P P P P R+ PPP PP P + + AN
Sbjct: 39 IDSPPPSQPPIPAPPAPPTPYPFQPSTPDNNNPFFPTYRSPPPPPPPPSPSSLVSFPAN 97
>ref|XP_482525.1| putative formin homology(FH)protein [Oryza sativa (japonica
cultivar-group)] gi|38175481|dbj|BAD01178.1| putative
formin homology(FH)protein [Oryza sativa (japonica
cultivar-group)] gi|37805923|dbj|BAC99340.1| putative
formin homology(FH)protein [Oryza sativa (japonica
cultivar-group)]
Length = 882
Score = 363 bits (932), Expect = 1e-98
Identities = 206/508 (40%), Positives = 309/508 (60%), Gaps = 54/508 (10%)
Query: 331 NWSMKAGDNNTSVVEATSSEGVGQVPEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPP 390
N +++ +S +E +SS G+ P P PPPPP+K PPP
Sbjct: 404 NSKVESSSKESSRIETSSSMGI---------------PKPAPPPPPQK------NPPPNL 442
Query: 391 RA---GQPPPAPPKP--VKVGNQGANQGGTSDGDAPKPKLKPFFWDKVAAKPDQQMVWHE 445
+ GQPPP PP P ++VG G+ P P+LKP WDKV A P++ MVW++
Sbjct: 443 KGQCYGQPPPPPPLPLQIQVGKDGS----------PLPRLKPLHWDKVRAAPNRSMVWND 492
Query: 446 ISAGSFVF--NEEKMESLFGCANQNRNERKKDSPSVDTSVQYIQ-IIDPKKAQNLSILLR 502
I + SF F +E+ ++SLF N KD +++ + + +I+ + QN +ILL+
Sbjct: 493 IRSSSFEFEFDEQMIKSLFA---YNLQGSMKDEEAMNKTASTTKHVIEHHRLQNTTILLK 549
Query: 503 ALNVTTDEVIDALREGNEIPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAERFLKVL 562
LN T +V +++ +GN + V+ ++ L+KM PT EEE KL + G+++ L PAE F+KVL
Sbjct: 550 TLNANTSQVCNSVIQGNGLSVQQLEALVKMKPTKEEEEKLLNYDGDINMLDPAENFVKVL 609
Query: 563 VDIPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGNRM 622
+ IP AF R+E +L+ +E + IK SFA +E A +L+ S+LFL+LLEAVLKTGNRM
Sbjct: 610 LTIPMAFPRMEVMLYKENFDDEVAHIKMSFAMIEGACTELKSSKLFLRLLEAVLKTGNRM 669
Query: 623 NDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGIRAVRTERESRSVVS 682
N GT RGGA AF+LD LLKL+D++GTDGKTTLLHFVV+E+ RS+G++A+ E+ S
Sbjct: 670 NVGTLRGGASAFKLDALLKLADIRGTDGKTTLLHFVVKEMARSKGLKALEKLNETPSSCH 729
Query: 683 SVGTDQDVDESPEASEEHYRSLGLQVVSSLSNELEDVKRAALIDGDALSSAVSKLGHSLA 742
T++ E Y S+G + VS LSNEL +VK+ A ID D L +++S L LA
Sbjct: 730 DTPTER----------EEYSSMGTEFVSELSNELGNVKKVASIDLDTLRNSISNLSCGLA 779
Query: 743 KTQEFMNTDLKSLDEESEFQRCTERFVEKAREEVTWLQEEEKRIIAQVKSTADYFHGNAG 802
+ + + DL S D+ + F +C + F+ A + L+ +E +++ V+ +Y+HG
Sbjct: 780 QLRNLVEKDLASDDKNNNFLQCMKSFLNHAENTMQGLKADEAQVLLNVRELTEYYHGEVS 839
Query: 803 RDEG--LRLFLIVRDFLIILDKVCKEVR 828
+DE L++F+IV+DFL +LDKVC+E+R
Sbjct: 840 KDESNLLQIFIIVKDFLGLLDKVCREMR 867
>gb|AAB64026.1| unknown protein [Arabidopsis thaliana] gi|25408904|pir||F84870
hypothetical protein At2g43800 [imported] - Arabidopsis
thaliana gi|15224360|ref|NP_181908.1| formin homology 2
domain-containing protein / FH2 domain-containing
protein [Arabidopsis thaliana]
Length = 894
Score = 356 bits (914), Expect = 2e-96
Identities = 247/606 (40%), Positives = 328/606 (53%), Gaps = 67/606 (11%)
Query: 301 SGGKQKVVRLG------NANNEGSGISNGKKPSNVRNWSMKAGDNNTSVVEATSSEGVGQ 354
SG KQ R+ N + GSG SN P+N S+ A TS+ + S V
Sbjct: 334 SGRKQSPTRVSDVDQIDNRSINGSG-SNSCSPTNFAP-SLNASPG-TSLKPKSISPPVSL 390
Query: 355 VPE--PPNGKPP---PPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKPVKVGNQG 409
+ NG P P P PPPPPPP+ PA P G
Sbjct: 391 HSQISSNNGIPKRLCPARPPPPPPPPPQVSEVPATMSHSLP------------------G 432
Query: 410 ANQGGTSDGDAPKPKLKPFFWDKVAAKPDQQMVWHEISAGSFVFNEEKMESLFGCANQNR 469
+ + KPKLK WDKV A + MVW +I + SF NEE +E+LF +
Sbjct: 433 DDSDPEKKVETMKPKLKTLHWDKVRASSSRVMVWDQIKSNSFQVNEEMIETLFKVNDPTS 492
Query: 470 NERKKDSPSVDTSVQYIQIIDPKKAQNLSILLRALNVTTDEVIDALREGNE--IPVELIQ 527
R SV Q + +DP+K+ N++ILLRALNVT DEV +AL EGN + EL++
Sbjct: 493 RTRDGVVQSVS---QENRFLDPRKSHNIAILLRALNVTADEVCEALIEGNSDTLGPELLE 549
Query: 528 TLLKMAPTTEEELKLRLF----TGELSQLGPAERFLKVLVDIPFAFKRLETLLFMFILRE 583
LLKMAPT EEE KL+ G S++GPAE+FLK L++IPFAFKR++ +L++
Sbjct: 550 CLLKMAPTKEEEDKLKELKDDDDGSPSKIGPAEKFLKALLNIPFAFKRIDAMLYIVKFES 609
Query: 584 EASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGNRMNDGTYRGGAQAFRLDTLLKLS 643
E + SF TLE A+ +L+ +R+FLKLLEAVLKTGNRMN GT RG A AF+LDTLLKL
Sbjct: 610 EIEYLNRSFDTLEAATGELKNTRMFLKLLEAVLKTGNRMNIGTNRGDAHAFKLDTLLKLV 669
Query: 644 DVKGTDGKTTLLHFVVQEIIRSEGIRAVRTERESRSVVSSVGTDQDVDESPEASEEHYRS 703
D+KG DGKTTLLHFVVQEII+ EG R T +S +G D ++S + +
Sbjct: 670 DIKGADGKTTLLHFVVQEIIKFEGARVPFTPSQSH-----IG-DNMAEQSAFQDDLELKK 723
Query: 704 LGLQVVSSLSNELEDVKRAALIDGDALSSAVSKLGHSLAKTQEFMNTDLKSLDEESEFQR 763
LGLQVVS LS++L +VK+AA +D ++L + +++ +AK +E + T+LK F
Sbjct: 724 LGLQVVSGLSSQLINVKKAAAMDSNSLINETAEIARGIAKVKEVI-TELKQETGVERFLE 782
Query: 764 CTERFVEKAREEVTWLQEEEKRIIAQVKSTADYFHGNAGRDEGLRLFLIVRDFLIILDKV 823
F+ K +E+T LQ ++ VK +YFHGN+ R+F +VRDFL ILD+V
Sbjct: 783 SMNSFLNKGEKEITELQSHGDNVMKMVKEVTEYFHGNS-ETHPFRIFAVVRDFLTILDQV 841
Query: 824 CKEVRDKTLRAVKSSQRKEALSVPSSPDTRHQQSPPAPSDLHRRLFPAIAGR--RIVEDS 881
CKEV R V S L PS +Q + P LFP + R+
Sbjct: 842 CKEVGRVNERTVYGSM---PLHSPS-----NQTATP--------LFPVVINNNSRLSPSG 885
Query: 882 SSDDDD 887
S DDDD
Sbjct: 886 SLDDDD 891
Score = 44.7 bits (104), Expect = 0.014
Identities = 22/49 (44%), Positives = 23/49 (46%), Gaps = 6/49 (12%)
Query: 363 PPPPPPGPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKPVKVGNQGAN 411
PPPPPP PP P P P P A PPPAPP P + AN
Sbjct: 87 PPPPPPSPPHPNP------FFPSSDPTSTASHPPPAPPPPASLPTFPAN 129
Score = 43.9 bits (102), Expect = 0.024
Identities = 50/193 (25%), Positives = 71/193 (35%), Gaps = 40/193 (20%)
Query: 152 HHGRNLISDP--PHNELSPSP-----AGTVPSPGP----------ALESPTPN------- 187
HH R+L+ P P +P P + PSP P +P P+
Sbjct: 24 HHSRHLLHQPFFPVVTAAPPPYQPPVSSQPPSPSPHTHHHHKKHLTTTTPPPHEKHLFSS 83
Query: 188 --HGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHP--PPSPEKLQHVTYAPD 243
+ P P SP P P +S P+ST S PP PP P P + L T+
Sbjct: 84 VANPPPPPPSPPHPNPFFPSSDPTSTASHPPPAPP--PPASLPTFPANISSLLFPTHNKQ 141
Query: 244 PSPPDKGDRKQKAIILAVTVSGIIIMIGLALCY------RESRRSDRADKDEG----PLL 293
PP G + I A +S ++ A+ R RRS AD + L
Sbjct: 142 SKPPSNGHIARLVTITASVISAAALLSLFAVFIIFIRRTRHRRRSSPADDTKSTRSDALQ 201
Query: 294 ILTSSDYSGGKQK 306
+ +S G K++
Sbjct: 202 LFNASPSDGSKKQ 214
>emb|CAE75997.1| B1358B12.6 [Oryza sativa (japonica cultivar-group)]
gi|38344969|emb|CAD40989.2| OSJNBa0072F16.14 [Oryza
sativa (japonica cultivar-group)]
gi|50924822|ref|XP_472757.1| OSJNBa0072F16.14 [Oryza
sativa (japonica cultivar-group)]
Length = 833
Score = 355 bits (912), Expect = 3e-96
Identities = 219/556 (39%), Positives = 307/556 (54%), Gaps = 81/556 (14%)
Query: 309 RLGNANNEGSGISN--GKKPSNVRNWSMKAGDNNTSVVEATSSEGVGQVPEPP----NGK 362
R A++ S +S+ S V++ ++ ++ EA S G G V E N +
Sbjct: 306 RSSRAHSPSSSVSDLTSVSTSVVKDHEVRRAVHSLMFPEAQSG-GAGHVKEDEAESGNMR 364
Query: 363 PPPPPPGPPPPPPPRK--------------PPAPAPRPPPPP--------RAGQP--PPA 398
PPPPPP PPPPPPP PP P P PPPPP +G P PPA
Sbjct: 365 PPPPPPPPPPPPPPAVTQQQDVKTSCGPAVPPPPPPTPPPPPPLLAPKQQSSGGPILPPA 424
Query: 399 PPKPVKVGNQGANQGGTSDGDAPKPKLKPFFWDKVAAKPDQQMVWHEISAGSFVFNEEKM 458
P P G AP PKLKP WDKV A P+++MVW I + SF +E+ +
Sbjct: 425 PAPPPLFRPWAPAVGKNG---APLPKLKPLHWDKVRAAPNRRMVWDRIRSSSFELDEKMI 481
Query: 459 ESLFG----CANQNRNERKKDSPSVDTSVQYIQIIDPKKAQNLSILLRALNVTTDEVIDA 514
ESLFG C+ ++ + + SPS+ V +D K+ QN +IL++A++ T +++ A
Sbjct: 482 ESLFGYNARCSTKHEEVQSR-SPSLGHHV-----LDTKRLQNFTILMKAVSATAEQIFAA 535
Query: 515 LREGNEIPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAERFLKVLVDIPFAFKRLET 574
L GN + + ++ L+KMAP +E KL + G++ L PAER LKV++ IP AF R+E
Sbjct: 536 LLHGNGLSAQQLEALIKMAPAKDEADKLSAYDGDVDGLVPAERLLKVVLTIPCAFARVEA 595
Query: 575 LLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGNRMNDGTYRGGAQAF 634
+L+ +E I++SF LE A +L S+LFLKLLEAVLKTGNRMN GT RGGA AF
Sbjct: 596 MLYRETFADEVGHIRKSFEMLEEACRELMSSKLFLKLLEAVLKTGNRMNVGTARGGAMAF 655
Query: 635 RLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGIRAVRTERESRSVVSSVGTDQDVDESP 694
+LD LLKL+DVKGTDGKTTLLHFVVQE+ RS A
Sbjct: 656 KLDALLKLADVKGTDGKTTLLHFVVQEMTRSRAAEAA----------------------- 692
Query: 695 EASEEHYRSLGLQVVSSLSNELEDVKRAALIDGDALSSAVSKLGHSLAKTQEFMNTDLKS 754
+ + L EL +V++ A +D D L+++VS L H L++ +E + +DL
Sbjct: 693 ------------DIAAGLGAELTNVRKTATVDLDVLTTSVSGLSHGLSRIKELVGSDLSG 740
Query: 755 LDEESEFQRCTERFVEKAREEVTWLQEEEKRIIAQVKSTADYFHGNAGRDEG--LRLFLI 812
+ F FV A E + L++ E+R++A V+ +Y+HG+ G+DE LR+F+I
Sbjct: 741 DERNQCFVAFMAPFVAHAGEVIRELEDGERRVLAHVREITEYYHGDVGKDEASPLRIFVI 800
Query: 813 VRDFLIILDKVCKEVR 828
VRDFL +L++VCKEVR
Sbjct: 801 VRDFLGMLERVCKEVR 816
>gb|AAQ62880.1| At1g59910 [Arabidopsis thaliana] gi|5080823|gb|AAD39332.1|
Hypothetical protein [Arabidopsis thaliana]
gi|15218954|ref|NP_176199.1| formin homology 2
domain-containing protein / FH2 domain-containing
protein [Arabidopsis thaliana] gi|25404245|pir||C96623
hypothetical protein F23H11.22 [imported] - Arabidopsis
thaliana
Length = 929
Score = 342 bits (877), Expect = 3e-92
Identities = 218/567 (38%), Positives = 303/567 (52%), Gaps = 48/567 (8%)
Query: 356 PEPPNGKPPPPPP-----GPPPPPPPRKPPAPAPRPPPPPRAGQPPPAPPKPVKVGNQ-- 408
P P N PPPPP PPPPPPP+K PA P PPPP + G PP PP K G
Sbjct: 376 PGPANQTSPPPPPPPSAAAPPPPPPPKKGPAAPPPPPPPGKKGAGPPPPPPMSKKGPPKP 435
Query: 409 -GANQGGTSDG----------DAPKPKLKPFFWDKVAAKPDQQMVWHEISAGSFVFNEEK 457
G +G T G D +PKLKP WDK+ + MVWH+I GSF F+ +
Sbjct: 436 PGNPKGPTKSGETSLAVGKTEDPTQPKLKPLHWDKMNPDASRSMVWHKIDGGSFNFDGDL 495
Query: 458 MESLFGCANQNRNERKK--DSPSVDTSVQYIQ--IIDPKKAQNLSILLRALNVTTDEVID 513
ME+LFG + +E + +V SV + Q I+DP+K+QN +I+L++L +T +E+ID
Sbjct: 496 MEALFGYVARKPSESNSVPQNQTVSNSVPHNQTYILDPRKSQNKAIVLKSLGMTKEEIID 555
Query: 514 ALREGNEIPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAERFL-KVLVDIPFAFKRL 572
L EG++ + ++ L +APT EE+ ++ F GE L A+ L +L +P AF R
Sbjct: 556 LLTEGHDAESDTLEKLAGIAPTPEEQTEIIDFDGEPMTLAYADSLLFHILKAVPSAFNRF 615
Query: 573 ETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGNRMNDGTYRGGAQ 632
+LF E + K S TLE A ++LR LF+KLLEA+LK GNRMN GT RG AQ
Sbjct: 616 NVMLFKINYGSEVAQQKGSLLTLESACNELRARGLFMKLLEAILKAGNRMNAGTARGNAQ 675
Query: 633 AFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGIRAVRTERESRSVVSSVGTDQDVDE 692
AF L L KLSDVK D KTTLLHFVV+E++RSEG RA + + S G+ ++ D
Sbjct: 676 AFNLTALRKLSDVKSVDAKTTLLHFVVEEVVRSEGKRAAMNK---NMMSSDNGSGENADM 732
Query: 693 SPEASEEHYRSLGLQVVSSLSNELEDVKRAALIDGDALSSAVSKLGHSLAKTQEFMNTDL 752
S E E + +GL ++ LS+E +VK+AA ID D+ + LG + +T+ ++
Sbjct: 733 SREEQEIEFIKMGLPIIGGLSSEFTNVKKAAGIDYDSFVATTLALGTRVKETKRLLD--- 789
Query: 753 KSLDEESEFQRCTERFVEKAREEVTWLQEEEKRIIAQVKSTADYFHGNAGRDEGL-RLFL 811
+S +E F E A EE+ + EE+ RI+ VK T +Y+ A ++ L +LF+
Sbjct: 790 QSKGKEDGCLTKLRSFFESAEEELKVITEEQLRIMELVKKTTNYYQAGALKERNLFQLFV 849
Query: 812 IVRDFLIILDKVCKEV----------RDKTLRAVKSSQRKEALSVPSSPDTRHQQSPPAP 861
I+RDFL ++D C E+ R T A SS E SV ++P + P P
Sbjct: 850 IIRDFLGMVDNACSEIARNQRKQQQQRPATTVAGASSSPAETPSVAAAPQRNAVRFPILP 909
Query: 862 SDLHRRLFPAIAGRRIVEDSSSDDDDD 888
P SSSD D +
Sbjct: 910 --------PNFMSESSRYSSSSDSDSE 928
Score = 42.7 bits (99), Expect = 0.053
Identities = 28/82 (34%), Positives = 35/82 (42%), Gaps = 11/82 (13%)
Query: 170 PAGTVPSPGPALESPTP-NHGP-TPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQH 227
P G AL + TP G T AN+P +P +SP P P+A PP PP +
Sbjct: 344 PPGQYMPGNAALSASTPLTPGQFTTANAPPAPPGPANQTSPPPPPPPSAAAPPPPPPPKK 403
Query: 228 PPPSPEKLQHVTYAPDPSPPDK 249
P +P P P PP K
Sbjct: 404 GPAAP---------PPPPPPGK 416
Score = 41.6 bits (96), Expect = 0.12
Identities = 31/95 (32%), Positives = 37/95 (38%), Gaps = 5/95 (5%)
Query: 171 AGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQHPPP 230
A T +PG + P P PAN +SP P P S+ + P P PP P PPP
Sbjct: 357 ASTPLTPGQFTTANAPPAPPGPANQ-TSPPPPPPPSAAAPPPPP----PPKKGPAAPPPP 411
Query: 231 SPEKLQHVTYAPDPSPPDKGDRKQKAIILAVTVSG 265
P + P P KG K T SG
Sbjct: 412 PPPGKKGAGPPPPPPMSKKGPPKPPGNPKGPTKSG 446
Score = 39.3 bits (90), Expect = 0.59
Identities = 21/67 (31%), Positives = 29/67 (42%), Gaps = 3/67 (4%)
Query: 168 PSPAG-TVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVPPTSPPRQ 226
P PA T P P P + P P P P++P P P + P P + PP+
Sbjct: 376 PGPANQTSPPPPPPPSAAAPPPPPPPKKGPAAPPPPPPPGKKGAGPPPPPPMSKKGPPK- 434
Query: 227 HPPPSPE 233
PP +P+
Sbjct: 435 -PPGNPK 440
Score = 39.3 bits (90), Expect = 0.59
Identities = 27/92 (29%), Positives = 36/92 (38%), Gaps = 8/92 (8%)
Query: 158 ISDPPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAM 217
+S PP + + A + +P + T N P P P P + S P P A
Sbjct: 341 VSLPPGQYMPGNAALSASTPLTPGQFTTANAPPAP------PGPANQTSPPPPPPPSAAA 394
Query: 218 VPPTSPPRQHP--PPSPEKLQHVTYAPDPSPP 247
PP PP++ P PP P P P PP
Sbjct: 395 PPPPPPPKKGPAAPPPPPPPGKKGAGPPPPPP 426
Score = 38.1 bits (87), Expect = 1.3
Identities = 32/101 (31%), Positives = 39/101 (37%), Gaps = 14/101 (13%)
Query: 166 LSPSPAGTVPSPGPALESPTPNHGPTPANSPSS---PTPTPENSSPSSTPSPTAMVPPTS 222
L+P T +P PA P P P PS+ P P P P++ P P PP
Sbjct: 361 LTPGQFTTANAP-PAPPGPANQTSPPPPPPPSAAAPPPPPPPKKGPAAPPPP----PPPG 415
Query: 223 PPRQHPPPSPEKLQHVTYAPDPSPP--DKGDRKQKAIILAV 261
PPP P ++ P PP KG K LAV
Sbjct: 416 KKGAGPPPPPP----MSKKGPPKPPGNPKGPTKSGETSLAV 452
Score = 35.4 bits (80), Expect = 8.5
Identities = 27/105 (25%), Positives = 43/105 (40%), Gaps = 11/105 (10%)
Query: 154 GRNLISDPPHNELSPSPAGTVPSPGPALES-----PTPNHGPTPANSPSSPTPTPENSSP 208
G+ + + P + +P P G P+ L + P P T N+P S + P + P
Sbjct: 286 GQYMAGNAPFSSSTPLPPGQYPAVNAQLSTSAPSVPLPPGQYTAVNAPFSTSTQPVSLPP 345
Query: 209 SSTPSPTAMVPPTSP--PRQ----HPPPSPEKLQHVTYAPDPSPP 247
A + ++P P Q + PP+P + T P P PP
Sbjct: 346 GQYMPGNAALSASTPLTPGQFTTANAPPAPPGPANQTSPPPPPPP 390
>ref|XP_478935.1| putative formin homology(FH) domain-containing protein [Oryza
sativa (japonica cultivar-group)]
gi|50509375|dbj|BAD30930.1| putative formin homology(FH)
domain-containing protein [Oryza sativa (japonica
cultivar-group)] gi|34393607|dbj|BAC83260.1| putative
formin homology(FH) domain-containing protein [Oryza
sativa (japonica cultivar-group)]
Length = 753
Score = 331 bits (849), Expect = 6e-89
Identities = 219/563 (38%), Positives = 315/563 (55%), Gaps = 65/563 (11%)
Query: 333 SMKAGDNNTSVVEATSSEGVGQVPEPPNGKPPPPPPGPPPPPPPRKP---PAPAPRPPPP 389
S + S +E T+ + P PP PP P PPPPPPPR+ P PA PPP
Sbjct: 243 SFSRSTSQHSTLEQTAMPPMA-APAPPQTNPPRPVRPPPPPPPPRQRLLRPLPAESPPPA 301
Query: 390 PRAGQPPPAPPKPVKVGNQGANQGGTSDGDAPKPKLKPFFWDKVAAKPDQQMVWHEI-SA 448
A P V ++G G + P P LKP WDK+ A + VW ++ ++
Sbjct: 302 ALANLELTGSPVKPAVEDRGGENSGAARPPKP-PHLKPLHWDKLRAISGRTTVWDQVKNS 360
Query: 449 GSFVFNEEKMESLFGCANQNRNERKKDSPSV---DTSVQYIQIIDPKKAQNLSILLRALN 505
+F +EE MESLF N P+ + Q +++DPK+ QN++I+L++LN
Sbjct: 361 DTFRVDEEAMESLF--LNSGGGGAGSSDPAARRGGSGKQERRLLDPKRLQNVAIMLKSLN 418
Query: 506 VTTDEVIDALREGN--EIPVELIQTLLKMAPTTEEELKLRLFTGELSQLGPAERFLKVLV 563
V DEVI AL GN ++ E +TL KMAPT EEELKL+ ++G+LS++ PAERFLK ++
Sbjct: 419 VAADEVIGALVRGNPEDLGSEFYETLAKMAPTKEEELKLKGYSGDLSKIDPAERFLKDVL 478
Query: 564 DIPFAFKRLETLLFMFILREEASSIKESFATLEVASDKLRKSRLFLKLLEAVLKTGNRMN 623
+PFAF+R++ +L+ E + +++SF TLE A ++LR S+LFLKLL+AVLKTGNRMN
Sbjct: 479 GVPFAFERVDAMLYRANFDNEVNYLRKSFGTLEAACEELRSSKLFLKLLDAVLKTGNRMN 538
Query: 624 DGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLHFVVQEIIRSEGIRAVRTERESRSVVSS 683
DGT RG A+AF+LDTLLKL+D+K TDG+TTLLHFVV+EIIRSEG
Sbjct: 539 DGTNRGEARAFKLDTLLKLADIKSTDGRTTLLHFVVKEIIRSEGF--------------- 583
Query: 684 VGTDQDVDESPEASEEHYRSLGLQVVSSLSNELEDVKRAALIDGDALSSAVSKLGHSLAK 743
+DQ S+E ++ GL++++ LS+EL +VKRAA ++ D LS + +L L K
Sbjct: 584 -DSDQSAVNPGSGSKEQFKRDGLKLLAGLSSELSNVKRAATLEMDTLSGNILRLEADLEK 642
Query: 744 TQEFMNTDLKSLDEESEFQRCTERFVEKAREEVTWLQEEEKRIIAQVKSTADYFHGNAGR 803
+ + L E Q +E F + V +L+ E A++K+ A
Sbjct: 643 VKLVL-----QLKETCSDQGASENFFQAM---VVFLRRAE----AEIKNMK-----TAEE 685
Query: 804 DEGLRLFLIVRDFLIILDKVCKEVRDKTLRAVKSSQRKEALSVPSSPDTRHQQSPPAPSD 863
LR+F++V +FL+ILD+VC++V R + S + + VP+ PP
Sbjct: 686 PHPLRIFVVVDEFLLILDRVCRDVGRTPERVMMGSGK--SFRVPAGTSL-----PP---- 734
Query: 864 LHRRLFPAIAGRRIVEDSSSDDD 886
HR RR++ SSSD+D
Sbjct: 735 -HRN-----ENRRVL--SSSDED 749
Score = 38.5 bits (88), Expect = 1.0
Identities = 52/215 (24%), Positives = 77/215 (35%), Gaps = 28/215 (13%)
Query: 219 PPTSPPRQHPPPSPEKLQHVTYAPDPSPPDKGDRKQKAIILAVTVSGIIIMIGLAL---- 274
PP S PPP P A S G R ++ V ++ ++ LA+
Sbjct: 53 PPMSGSEAVPPPPP--------AAAASATTGGGRSTTTVMNTVAIALSAGLVALAVASYS 104
Query: 275 CYRESRRSDRADKDEGPLLILTSSDYSGGKQKVVRLGNANNEGSGISNGKKPSNVRNWSM 334
C RR R ++D+G ++ + G V ++ GS + P S
Sbjct: 105 CCLLLRRRRREEEDDGD----RAAKRAVGAAAAVAARVPSDVGSSSRQHRSPPPSSTASD 160
Query: 335 KAG-DNNTSVVEATSSEGVGQVPEPPNGKPPPPPPGPPPPPPPRKPPAPAPRPPPPPRAG 393
D T++VE E + P+ +P P P P P PP P PPP
Sbjct: 161 AIYLDPLTTLVEVRQHE------KSPDLRPLPLLKQPSPDLRPL-PPLKRPESQPPP--- 210
Query: 394 QPPPAPPKPVKVGNQGANQGGTSDGDAPKPKLKPF 428
PPP+ P G + + APK + F
Sbjct: 211 -PPPSTPPLTTTGYSTDEEDQATYYTAPKTAMSSF 244
>ref|NP_177171.1| formin homology 2 domain-containing protein / FH2 domain-containing
protein [Arabidopsis thaliana] gi|25406031|pir||B96724
hypothetical protein F20P5.14 [imported] - Arabidopsis
thaliana gi|2194126|gb|AAB61101.1| EST gb|T43335 comes
from this gene. [Arabidopsis thaliana]
Length = 760
Score = 326 bits (836), Expect = 2e-87
Identities = 251/747 (33%), Positives = 369/747 (48%), Gaps = 93/747 (12%)
Query: 160 DPPHNELSPSPAGTVPSPGPALESPTPNHGPTPANSPSSPTPTPENSSPSSTPSPTAMVP 219
D P N + P ++ P L P+ N P P+N+ SS + T + T +V
Sbjct: 32 DSPQNIETFFPISSLSPVPPPLLPPSSNPSP-PSNNSSSSDKKTITKAVLITAASTLLVA 90
Query: 220 PTS---------PPRQHPPPSPEKLQHVTYAPDPSPPDKGDRKQKAIILAVT-------- 262
R+ P ++++ T P P PP A + T
Sbjct: 91 GVFFFCLQRCIIARRRRDRVGPVRVEN-TLPPYPPPP-----MTSAAVTTTTLAREGFTR 144
Query: 263 ---VSGIII-MIGLALCYRESRRSDRADKDEGPLLILTSSDYSGGKQKVVRLGNANNEGS 318
V G+I+ GL + Y +S R + SG +K + G +E
Sbjct: 145 FGGVKGLILDENGLDVLYWRKLQSQR--------------ERSGSFRKQIVTGEEEDEKE 190
Query: 319 GI--SNGKKPSNVRNWSMKAGDNNTS--VVEATSSEGVGQVPEPPNGKPPPPP------- 367
I N KK V + G ++TS V+ + QV + PPPPP
Sbjct: 191 VIYYKNKKKTEPVTEIPLLRGRSSTSHSVIHNEDHQPPPQVKQSEPTPPPPPPSIAVKQS 250
Query: 368 -PGPPPPPPPRKPPAPAPRPPPPPR--------AGQPPPAPPKPVKVGNQGANQGGTSDG 418
P P PPPP +K +P+P PPPP + A +PPPAP + GA+ G TS
Sbjct: 251 APTPSPPPPIKKGSSPSPPPPPPVKKVGALSSSASKPPPAPVR-------GASGGETSK- 302
Query: 419 DAPKPKLKPFFWDKVAAKPDQQMVWHEISAGSFVFNEEKMESLFGCANQNRNERKKDSPS 478
+ KLKP WDKV D MVW +I GSF F+ + ME+LFG + ++
Sbjct: 303 ---QVKLKPLHWDKVNPDSDHSMVWDKIDRGSFSFDGDLMEALFGYVAVGKKSPEQGDEK 359
Query: 479 VDTSVQYIQIIDPKKAQNLSILLRALNVTTDEVIDALREGNEIPVELIQTLLKMAPTTEE 538
S Q I I+DP+K+QN +I+L++L +T +E++++L EGN+ + ++ L ++APT EE
Sbjct: 360 NPKSTQ-IFILDPRKSQNTAIVLKSLGMTREELVESLIEGNDFVPDTLERLARIAPTKEE 418
Query: 539 ELKLRLFTGELSQLGPAERFL-KVLVDIPFAFKRLETLLFMFILREEASSIKESFATLEV 597
+ + F G+ ++L AE FL +L +P AF RL LF E + + TL++
Sbjct: 419 QSAILEFDGDTAKLADAETFLFHLLKSVPTAFTRLNAFLFRANYYPEMAHHSKCLQTLDL 478
Query: 598 ASDKLRKSRLFLKLLEAVLKTGNRMNDGTYRGGAQAFRLDTLLKLSDVKGTDGKTTLLHF 657
A +LR LF+KLLEA+LK GNRMN GT RG AQAF L LLKLSDVK DGKT+LL+F
Sbjct: 479 ACKELRSRGLFVKLLEAILKAGNRMNAGTARGNAQAFNLTALLKLSDVKSVDGKTSLLNF 538
Query: 658 VVQEIIRSEGIRAVRTERESRSVVSSVGTDQDVDE------SPEASEEHYRSLGLQVVSS 711
VV+E++RSEG R V R S S+ S ++ + S E E+ Y LGL VV
Sbjct: 539 VVEEVVRSEGKRCV-MNRRSHSLTRSGSSNYNGGNSSLQVMSKEEQEKEYLKLGLPVVGG 597
Query: 712 LSNELEDVKRAALIDGDALSSAVSKLGHSLAKTQEFMNTDLKSLDEESEFQRCTERFVEK 771
LS+E +VK+AA +D + + + S L AK + + + + E F + F++
Sbjct: 598 LSSEFSNVKKAACVDYETVVATCSALA-VRAKDAKTVIGECED-GEGGRFVKTMMTFLDS 655
Query: 772 AREEVTWLQEEEKRIIAQVKSTADYFHGNA---GRDEGLRLFLIVRDFLIILDKVCKEV- 827
EEV + EE++++ VK T DY+ A G++ L LF+IVRDFL ++DKVC ++
Sbjct: 656 VEEEVKIAKGEERKVMELVKRTTDYYQAGAVTKGKNP-LHLFVIVRDFLAMVDKVCLDIM 714
Query: 828 ----RDKTLRAVKSSQRKEALSVPSSP 850
R K + S ++ A+ P P
Sbjct: 715 RNMQRRKVGSPISPSSQRNAVKFPVLP 741
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.312 0.132 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,790,569,006
Number of Sequences: 2540612
Number of extensions: 101192349
Number of successful extensions: 2372225
Number of sequences better than 10.0: 23283
Number of HSP's better than 10.0 without gapping: 11662
Number of HSP's successfully gapped in prelim test: 12793
Number of HSP's that attempted gapping in prelim test: 1077291
Number of HSP's gapped (non-prelim): 387790
length of query: 896
length of database: 863,360,394
effective HSP length: 137
effective length of query: 759
effective length of database: 515,296,550
effective search space: 391110081450
effective search space used: 391110081450
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 80 (35.4 bits)
Lotus: description of TM0203.14


