

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0201.10
(137 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
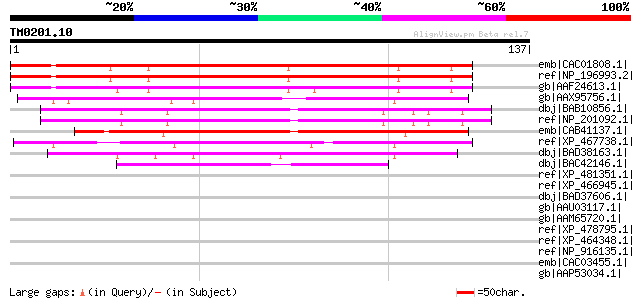
Score E
Sequences producing significant alignments: (bits) Value
emb|CAC01808.1| putative protein [Arabidopsis thaliana] gi|11357... 126 1e-28
ref|NP_196993.2| NHL repeat-containing protein [Arabidopsis thal... 126 1e-28
gb|AAF24613.1| hypothetical protein [Arabidopsis thaliana] gi|26... 120 9e-27
gb|AAX95756.1| expressed protein [Lycopersicon esculentum] 100 1e-20
dbj|BAB10856.1| unnamed protein product [Arabidopsis thaliana] g... 99 2e-20
ref|NP_201092.1| F-box family protein-related [Arabidopsis thali... 99 2e-20
emb|CAB41137.1| hypothetical protein [Arabidopsis thaliana] gi|4... 96 2e-19
ref|XP_467738.1| NHL repeat-containing protein-like [Oryza sativ... 94 7e-19
dbj|BAD38163.1| NHL repeat-containing protein-like [Oryza sativa... 86 2e-16
dbj|BAC42146.1| unknown protein [Arabidopsis thaliana] gi|288276... 51 7e-06
ref|XP_481351.1| unknown protein [Oryza sativa (japonica cultiva... 42 0.003
ref|XP_466945.1| unknown protein [Oryza sativa (japonica cultiva... 39 0.035
dbj|BAD37606.1| unknown protein [Oryza sativa (japonica cultivar... 39 0.035
gb|AAU03117.1| unknown protein [Oryza sativa (japonica cultivar-... 38 0.060
gb|AAM65720.1| unknown [Arabidopsis thaliana] gi|10176808|dbj|BA... 36 0.18
ref|XP_478795.1| unknown protein [Oryza sativa (japonica cultiva... 35 0.30
ref|XP_464348.1| unknown protein [Oryza sativa (japonica cultiva... 35 0.39
ref|NP_916135.1| P0046E05.20 [Oryza sativa (japonica cultivar-gr... 35 0.39
emb|CAC03455.1| putative protein [Arabidopsis thaliana] gi|21593... 34 0.67
gb|AAP53034.1| hypothetical protein [Oryza sativa (japonica cult... 33 1.1
>emb|CAC01808.1| putative protein [Arabidopsis thaliana] gi|11357718|pir||T51434
hypothetical protein F2G14_10 - Arabidopsis thaliana
Length = 733
Score = 126 bits (317), Expect = 1e-28
Identities = 68/140 (48%), Positives = 87/140 (61%), Gaps = 19/140 (13%)
Query: 1 MQEVLFAKQACICYLIPCSSSSETS----LSWWERVRSP---ENKEWWWAHGWNKVREWS 53
M E +FAK+ C C+++PC SS+ S WW+R+R+ E E WW GW K+REWS
Sbjct: 579 MHEAMFAKRGC-CFILPCLGSSQPSGPNGSVWWQRIRTVDKLEPDERWWVSGWMKMREWS 637
Query: 54 EIIVGPKWKTFIRRFNRNN----NRAGAASYDKKGSFHYDSLSYALNFDDGE-----EDV 104
EI+ GPKWKTFIRRF RN+ G + + SF YDS SY+LNFDDG+ ED
Sbjct: 638 EIVAGPKWKTFIRRFGRNHCCNGGIDGGCNRPEHVSFRYDSWSYSLNFDDGKQTGHFEDE 697
Query: 105 YSYGGFSTRFA--SVPASTK 122
+ Y +S RFA S+P STK
Sbjct: 698 FPYRDYSMRFAAPSLPVSTK 717
>ref|NP_196993.2| NHL repeat-containing protein [Arabidopsis thaliana]
Length = 754
Score = 126 bits (317), Expect = 1e-28
Identities = 68/140 (48%), Positives = 87/140 (61%), Gaps = 19/140 (13%)
Query: 1 MQEVLFAKQACICYLIPCSSSSETS----LSWWERVRSP---ENKEWWWAHGWNKVREWS 53
M E +FAK+ C C+++PC SS+ S WW+R+R+ E E WW GW K+REWS
Sbjct: 600 MHEAMFAKRGC-CFILPCLGSSQPSGPNGSVWWQRIRTVDKLEPDERWWVSGWMKMREWS 658
Query: 54 EIIVGPKWKTFIRRFNRNN----NRAGAASYDKKGSFHYDSLSYALNFDDGE-----EDV 104
EI+ GPKWKTFIRRF RN+ G + + SF YDS SY+LNFDDG+ ED
Sbjct: 659 EIVAGPKWKTFIRRFGRNHCCNGGIDGGCNRPEHVSFRYDSWSYSLNFDDGKQTGHFEDE 718
Query: 105 YSYGGFSTRFA--SVPASTK 122
+ Y +S RFA S+P STK
Sbjct: 719 FPYRDYSMRFAAPSLPVSTK 738
>gb|AAF24613.1| hypothetical protein [Arabidopsis thaliana]
gi|26452871|dbj|BAC43514.1| unknown protein [Arabidopsis
thaliana] gi|28973291|gb|AAO63970.1| unknown protein
[Arabidopsis thaliana] gi|15232117|ref|NP_186792.1|
expressed protein [Arabidopsis thaliana]
Length = 180
Score = 120 bits (300), Expect = 9e-27
Identities = 66/153 (43%), Positives = 88/153 (57%), Gaps = 32/153 (20%)
Query: 1 MQEVLFAKQACICYLIPCSSSSETSLS----WWERVRSP---ENKEWWWAHGWNKVREWS 53
M E LFAK+ C C+L+PC +SS+ S WW+R+ + E E WW GW ++REWS
Sbjct: 17 MHEALFAKRGC-CFLMPCLASSQPSTRGGSVWWQRITTVDKLEPDERWWIRGWRRMREWS 75
Query: 54 EIIVGPKWKTFIRRFNRNN------NRAGAAS-----------YDKKGSFHYDSLSYALN 96
E++ GP+WKT+IRRF R+N R G +S +G F YD LSY+LN
Sbjct: 76 ELVAGPRWKTYIRRFGRSNCCGGGGGRVGNSSGGCGGGAMPNRSSDQGKFRYDQLSYSLN 135
Query: 97 FDDGE-----EDVYSYGGFSTRFA--SVPASTK 122
FDDG +D + Y +S RFA S+P STK
Sbjct: 136 FDDGNQTGHFDDEFPYRDYSMRFAAPSLPVSTK 168
>gb|AAX95756.1| expressed protein [Lycopersicon esculentum]
Length = 156
Score = 100 bits (248), Expect = 1e-20
Identities = 59/133 (44%), Positives = 77/133 (57%), Gaps = 20/133 (15%)
Query: 3 EVLFAKQA-CICY---LIPCSSSSETSLSWWERVRSPENKEWW-WAHGWN---KVREWSE 54
E LF +++ C C+ P + WW R+RS KE WA G N K+REWSE
Sbjct: 20 ETLFQRRSRCFCFPSFRSPNQLTKYVKFDWWHRLRSGAVKEGSVWAKGINALKKLREWSE 79
Query: 55 IIVGPKWKTFIRRFNRNNNRAGAASYDKKGSFHYDSLSYALNFDDG------EEDVYSYG 108
I+ GP+WKTFIRRF+RN S ++ F YD LSY+LNFD+G EE+
Sbjct: 80 IVAGPRWKTFIRRFHRNK------SGNRNAKFQYDPLSYSLNFDEGRGNNGEEEEELVLR 133
Query: 109 GFSTRFASVPAST 121
FSTR+AS+PAS+
Sbjct: 134 NFSTRYASIPASS 146
>dbj|BAB10856.1| unnamed protein product [Arabidopsis thaliana]
gi|26450527|dbj|BAC42376.1| unknown protein [Arabidopsis
thaliana]
Length = 167
Score = 99.0 bits (245), Expect = 2e-20
Identities = 60/137 (43%), Positives = 76/137 (54%), Gaps = 20/137 (14%)
Query: 9 QACICYLIPCSSSSETSLSW--WERVRSPENKEW---------WWAHGWNKVREWSEIIV 57
Q C C+ S S T++ + W R+R+ ++ WW K+REWSEI+
Sbjct: 18 QRCCCFPSFRRSRSSTAVGYSSWGRIRTVDDSNHSGDHGDEPRWWIRASLKIREWSEIVA 77
Query: 58 GPKWKTFIRRFNRNNNRAGAASYDKKGSFHYDSLSYALNF-DDGEEDVY-SYGG---FST 112
GP+WKTFIRRFNR+ R +D F YD LSY+LNF DD EED Y GG FST
Sbjct: 78 GPRWKTFIRRFNRDPRR--GRDWDASEKFQYDPLSYSLNFDDDDEEDEYVGLGGLRSFST 135
Query: 113 RFASVP--ASTKPTFTP 127
RFASVP + P +P
Sbjct: 136 RFASVPVYSGKAPAISP 152
>ref|NP_201092.1| F-box family protein-related [Arabidopsis thaliana]
Length = 377
Score = 99.0 bits (245), Expect = 2e-20
Identities = 60/137 (43%), Positives = 76/137 (54%), Gaps = 20/137 (14%)
Query: 9 QACICYLIPCSSSSETSLSW--WERVRSPENKEW---------WWAHGWNKVREWSEIIV 57
Q C C+ S S T++ + W R+R+ ++ WW K+REWSEI+
Sbjct: 228 QRCCCFPSFRRSRSSTAVGYSSWGRIRTVDDSNHSGDHGDEPRWWIRASLKIREWSEIVA 287
Query: 58 GPKWKTFIRRFNRNNNRAGAASYDKKGSFHYDSLSYALNF-DDGEEDVY-SYGG---FST 112
GP+WKTFIRRFNR+ R +D F YD LSY+LNF DD EED Y GG FST
Sbjct: 288 GPRWKTFIRRFNRDPRR--GRDWDASEKFQYDPLSYSLNFDDDDEEDEYVGLGGLRSFST 345
Query: 113 RFASVP--ASTKPTFTP 127
RFASVP + P +P
Sbjct: 346 RFASVPVYSGKAPAISP 362
>emb|CAB41137.1| hypothetical protein [Arabidopsis thaliana]
gi|48310531|gb|AAT41834.1| At3g48020 [Arabidopsis
thaliana] gi|15228321|ref|NP_190385.1| expressed protein
[Arabidopsis thaliana] gi|51970374|dbj|BAD43879.1|
unknown protein [Arabidopsis thaliana]
gi|7487117|pir||T06681 hypothetical protein T17F15.110 -
Arabidopsis thaliana
Length = 135
Score = 95.9 bits (237), Expect = 2e-19
Identities = 48/111 (43%), Positives = 68/111 (61%), Gaps = 10/111 (9%)
Query: 18 CSSSSETSLSWWERVRSPENKE-WWWAHGWNKVREWSEIIVGPKWKTFIRRFNRNNNRAG 76
C S++ S SWW+R+ ++E WW + K+REWSEI+ GP+WKTFIRRFNR+ R
Sbjct: 15 CCSTTVKS-SWWQRIHRNNHQEPRWWVRAFLKIREWSEIVAGPRWKTFIRRFNRDPRR-- 71
Query: 77 AASYDKKGSFHYDSLSYALNFDDGEED------VYSYGGFSTRFASVPAST 121
+D F YD +SY L+F+D ++D V FS R+ASVP ++
Sbjct: 72 GQDWDDSDKFRYDPVSYTLSFEDEDKDDDDEAGVGGVRSFSMRYASVPVAS 122
>ref|XP_467738.1| NHL repeat-containing protein-like [Oryza sativa (japonica
cultivar-group)] gi|46390621|dbj|BAD16104.1| NHL
repeat-containing protein-like [Oryza sativa (japonica
cultivar-group)] gi|46390293|dbj|BAD15743.1| NHL
repeat-containing protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 171
Score = 94.0 bits (232), Expect = 7e-19
Identities = 57/136 (41%), Positives = 73/136 (52%), Gaps = 23/136 (16%)
Query: 2 QEVLFAKQA--CICYLIPCSSSSETSLSWWERVRSPENKEWWW--AHGWNKVREWSEIIV 57
+ FA++ C C+ P S+SS +RV E + WW KVREWSE++
Sbjct: 18 EAAFFARRGRRCCCFPWPSSASSH------QRVGGAEEESWWQRAVDAVLKVREWSELVA 71
Query: 58 GPKWKTFIRRFNRNNNRAGAA------SYDKKGSFHYDSLSYALNFDDG-----EEDVYS 106
GP+WKTFIRRF R G +Y +K +YD+LSYALNFD+G E D
Sbjct: 72 GPRWKTFIRRFGRGGGGGGGGGGPRPHNYGRK--LNYDALSYALNFDEGHGASPEGDYTG 129
Query: 107 YGGFSTRFASVPASTK 122
Y FS RFA+ PAS K
Sbjct: 130 YRDFSARFAAPPASAK 145
>dbj|BAD38163.1| NHL repeat-containing protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 211
Score = 85.9 bits (211), Expect = 2e-16
Identities = 52/132 (39%), Positives = 66/132 (49%), Gaps = 24/132 (18%)
Query: 11 CICYLIPCSSSSETSLS---------WWERVRSPEN-KEWWWAHGWN---KVREWSEIIV 57
C C+ P +SSS + WW RV + WW G + KVREWSE++
Sbjct: 36 CCCFWGPWASSSYSRAGGPAAAAEEEWWHRVGGGGGERRRWWRRGVDALMKVREWSELVA 95
Query: 58 GPKWKTFIRRFNR----NNNRAGAASYDKKGSFHYDSLSYALNFDDG-------EEDVYS 106
GP+WKTFIRRF R +++ G +YD LSYALNFD+G E D
Sbjct: 96 GPRWKTFIRRFRRSPRHHHHGGGGGGGGGGRKLNYDPLSYALNFDEGHGGACSPEGDYAG 155
Query: 107 YGGFSTRFASVP 118
Y FSTRF + P
Sbjct: 156 YRDFSTRFVAPP 167
>dbj|BAC42146.1| unknown protein [Arabidopsis thaliana] gi|28827680|gb|AAO50684.1|
unknown protein [Arabidopsis thaliana]
gi|15238747|ref|NP_197906.1| expressed protein
[Arabidopsis thaliana]
Length = 131
Score = 50.8 bits (120), Expect = 7e-06
Identities = 27/72 (37%), Positives = 36/72 (49%), Gaps = 5/72 (6%)
Query: 29 WERVRSPENKEWWWAHGWNKVREWSEIIVGPKWKTFIRRFNRNNNRAGAASYDKKGSFHY 88
W E + W + ++E SE I GPKWK FIR F+ +G + F Y
Sbjct: 46 WSGCLQEERRGNWGSEKLKGLKEISEKIAGPKWKNFIRSFS-----SGRKKMRRDVDFTY 100
Query: 89 DSLSYALNFDDG 100
D +Y+LNFDDG
Sbjct: 101 DLKNYSLNFDDG 112
>ref|XP_481351.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|38175512|dbj|BAD01207.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|28564623|dbj|BAC57790.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 132
Score = 42.0 bits (97), Expect = 0.003
Identities = 24/49 (48%), Positives = 27/49 (54%), Gaps = 4/49 (8%)
Query: 84 GSFHYDSLSYALNFD---DGEEDVYSYGGFSTRF-ASVPASTKPTFTPV 128
G F YD +SYALNF+ DGE + Y FS R AS P PT PV
Sbjct: 80 GEFRYDPVSYALNFEEDGDGEAQPFKYMAFSARLPASPPPPPPPTALPV 128
>ref|XP_466945.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|49388702|dbj|BAD25883.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|49387977|dbj|BAD25085.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 81
Score = 38.5 bits (88), Expect = 0.035
Identities = 20/37 (54%), Positives = 22/37 (59%), Gaps = 1/37 (2%)
Query: 65 IRRFNRNNNR-AGAASYDKKGSFHYDSLSYALNFDDG 100
IRR R G + SFHYD+LSYALNFDDG
Sbjct: 36 IRRMQSERGRWRGGRRDHARFSFHYDALSYALNFDDG 72
>dbj|BAD37606.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|51535534|dbj|BAD37453.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 102
Score = 38.5 bits (88), Expect = 0.035
Identities = 20/46 (43%), Positives = 24/46 (51%), Gaps = 9/46 (19%)
Query: 84 GSFHYDSLSYALNFDDGEED---------VYSYGGFSTRFASVPAS 120
G F YD LSYALNFDDG+ D + Y FS+R P +
Sbjct: 45 GEFGYDPLSYALNFDDGDGDDDAADDAAAAFRYKNFSSRLPPSPVA 90
>gb|AAU03117.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|46575947|gb|AAT01308.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 119
Score = 37.7 bits (86), Expect = 0.060
Identities = 20/53 (37%), Positives = 26/53 (48%), Gaps = 7/53 (13%)
Query: 61 WKTFIRRFNRNN-------NRAGAASYDKKGSFHYDSLSYALNFDDGEEDVYS 106
W +RR R + +R G A +FHYD+ SYA NFDDG Y+
Sbjct: 54 WSRLLRRLVRESRSFCSLGSRHGGAMAAATTTFHYDAASYAKNFDDGRRAHYA 106
>gb|AAM65720.1| unknown [Arabidopsis thaliana] gi|10176808|dbj|BAB10016.1| unnamed
protein product [Arabidopsis thaliana]
gi|15238394|ref|NP_198359.1| expressed protein
[Arabidopsis thaliana]
Length = 159
Score = 36.2 bits (82), Expect = 0.18
Identities = 19/40 (47%), Positives = 22/40 (54%), Gaps = 4/40 (10%)
Query: 86 FHYDSLSYALNFDDGEE----DVYSYGGFSTRFASVPAST 121
FHYD SYALNFD G+E D + FS R P S+
Sbjct: 105 FHYDPSSYALNFDKGDEDDNIDRFPLRNFSARLPHSPPSS 144
>ref|XP_478795.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|34393550|dbj|BAC83148.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 119
Score = 35.4 bits (80), Expect = 0.30
Identities = 17/42 (40%), Positives = 23/42 (54%), Gaps = 2/42 (4%)
Query: 60 KWKTFIRRF--NRNNNRAGAASYDKKGSFHYDSLSYALNFDD 99
+WK + R+ G + +K +F YDS SYALNFDD
Sbjct: 72 RWKAVAQEIMARRSGGGGGGSGRRRKTAFSYDSKSYALNFDD 113
>ref|XP_464348.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|49387525|dbj|BAD25058.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 103
Score = 35.0 bits (79), Expect = 0.39
Identities = 17/29 (58%), Positives = 19/29 (64%), Gaps = 4/29 (13%)
Query: 84 GSFHYDSLSYALNFDDG----EEDVYSYG 108
G F YD LSYALNFD+G E+D Y G
Sbjct: 52 GEFRYDPLSYALNFDEGAADDEDDDYEAG 80
>ref|NP_916135.1| P0046E05.20 [Oryza sativa (japonica cultivar-group)]
gi|20160582|dbj|BAB89529.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|15623854|dbj|BAB67913.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 150
Score = 35.0 bits (79), Expect = 0.39
Identities = 23/69 (33%), Positives = 34/69 (48%), Gaps = 3/69 (4%)
Query: 43 AHGWNKVREWSEIIVGPKWKTFIRRFNRNNNRAGAASYDKKGSFHYDSLSYALNFDDGEE 102
A GW R W ++ +T R +++ +GAAS + +F YD+ SYA NFDDG
Sbjct: 67 AAGWCGHRAWRRLLRRLAQET--RCICSSSSPSGAAS-SRPITFGYDAASYAKNFDDGRR 123
Query: 103 DVYSYGGFS 111
Y +
Sbjct: 124 PAAHYAALA 132
>emb|CAC03455.1| putative protein [Arabidopsis thaliana] gi|21593765|gb|AAM65732.1|
unknown [Arabidopsis thaliana]
gi|15238944|ref|NP_196668.1| expressed protein
[Arabidopsis thaliana] gi|11358370|pir||T51796
hypothetical protein T5K6_60 - Arabidopsis thaliana
Length = 152
Score = 34.3 bits (77), Expect = 0.67
Identities = 25/68 (36%), Positives = 32/68 (46%), Gaps = 9/68 (13%)
Query: 62 KTFIRRFNRNNNRA-------GAASYDKKGSFHYDSLSYALNFDDG--EEDVYSYGGFST 112
K+ I+ RNNN A S G F YD LSYALNF+D +D S+ F+
Sbjct: 73 KSRIKVTCRNNNCAYNNCVHHHHHSQSYPGDFSYDPLSYALNFEDNVRADDDGSFPNFTA 132
Query: 113 RFASVPAS 120
R P +
Sbjct: 133 RLPQSPVT 140
>gb|AAP53034.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|37532890|ref|NP_920747.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|22655752|gb|AAN04169.1| Hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 184
Score = 33.5 bits (75), Expect = 1.1
Identities = 20/66 (30%), Positives = 28/66 (42%), Gaps = 17/66 (25%)
Query: 82 KKGSFHYDSLSYALNFDDGEE--------------DVYSYGGFSTRFASVPASTKPTFTP 127
+ G F YD+ SYA NFD+G + D Y F++R +P S P +P
Sbjct: 102 RSGDFQYDARSYARNFDEGTDGEASGDEQAGLAAGDTLKYRSFASR---LPPSPTPALSP 158
Query: 128 VMVRRC 133
C
Sbjct: 159 SAAPVC 164
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.133 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 261,853,622
Number of Sequences: 2540612
Number of extensions: 10972323
Number of successful extensions: 22794
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 22
Number of HSP's successfully gapped in prelim test: 44
Number of HSP's that attempted gapping in prelim test: 22738
Number of HSP's gapped (non-prelim): 68
length of query: 137
length of database: 863,360,394
effective HSP length: 113
effective length of query: 24
effective length of database: 576,271,238
effective search space: 13830509712
effective search space used: 13830509712
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0201.10


