

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0199.9
(56 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
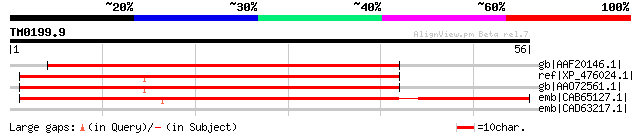
gb|AAF20146.1| vacuolar ATP synthase subunit C [Arabidopsis thal... 71 7e-12
ref|XP_476024.1| putative vacuolar ATP synthase subunit C [Oryza... 60 2e-08
gb|AAO72561.1| putative vacuolar ATP synthase subunit C [Oryza s... 60 2e-08
emb|CAB65127.1| vacuolar H+-ATPase subunit C [Hordeum vulgare su... 51 7e-06
emb|CAD63217.1| ATPase of the PilT family [Lactobacillus plantar... 32 5.9
>gb|AAF20146.1| vacuolar ATP synthase subunit C [Arabidopsis thaliana]
gi|12248023|gb|AAG50103.1| putative vacuolar ATP
synthase subunit C [Arabidopsis thaliana]
gi|20259972|gb|AAM13333.1| vacuolar ATP sythase subunit
C [Arabidopsis thaliana] gi|8698731|gb|AAF78489.1|
Identical to vacuolar ATP sythase subunit C (DET3) from
Arabidopsis thaliana gb|AF208261. ESTs gb|AA067533,
gb|Z37481, gb|AA721838, gb|Z37180, gb|T21206 come from
this gene gi|18391442|ref|NP_563916.1| vacuolar ATP
synthase subunit C (VATC) / V-ATPase C subunit /
vacuolar proton pump C subunit (DET3) [Arabidopsis
thaliana] gi|16649005|gb|AAL24354.1| Identical to
vacuolar ATP sythase subunit C (DET3) [Arabidopsis
thaliana] gi|25529228|pir||T52300 H+-exporting ATPase
(EC 3.6.3.6) chain C, vacuolar [validated] -
Arabidopsis thaliana gi|12585488|sp|Q9SDS7|VATC_ARATH
Vacuolar ATP synthase subunit C (V-ATPase C subunit)
(Vacuolar proton pump C subunit)
Length = 375
Score = 71.2 bits (173), Expect = 7e-12
Identities = 30/38 (78%), Positives = 35/38 (91%)
Query: 5 WWVVSLPVHNSASELWNRLQQNVSKHSFDTPLYRFNIP 42
+WVVSLPV +SAS LWNRLQ+ +SKHSFDTP+YRFNIP
Sbjct: 5 YWVVSLPVKDSASSLWNRLQEQISKHSFDTPVYRFNIP 42
>ref|XP_476024.1| putative vacuolar ATP synthase subunit C [Oryza sativa (japonica
cultivar-group)] gi|48475236|gb|AAT44305.1| putative
vacuolar ATP synthase subunit C [Oryza sativa (japonica
cultivar-group)]
Length = 377
Score = 60.1 bits (144), Expect = 2e-08
Identities = 26/44 (59%), Positives = 35/44 (79%), Gaps = 3/44 (6%)
Query: 2 AVVWWVVSLPVH---NSASELWNRLQQNVSKHSFDTPLYRFNIP 42
A +W+VSLPV ++A+ LW RLQ ++S+HSFDTPLYRFN+P
Sbjct: 2 ATRYWIVSLPVQTPGSTANSLWARLQDSISRHSFDTPLYRFNVP 45
>gb|AAO72561.1| putative vacuolar ATP synthase subunit C [Oryza sativa (japonica
cultivar-group)]
Length = 408
Score = 60.1 bits (144), Expect = 2e-08
Identities = 26/44 (59%), Positives = 35/44 (79%), Gaps = 3/44 (6%)
Query: 2 AVVWWVVSLPVH---NSASELWNRLQQNVSKHSFDTPLYRFNIP 42
A +W+VSLPV ++A+ LW RLQ ++S+HSFDTPLYRFN+P
Sbjct: 42 ATRYWIVSLPVQTPGSTANSLWARLQDSISRHSFDTPLYRFNVP 85
>emb|CAB65127.1| vacuolar H+-ATPase subunit C [Hordeum vulgare subsp. vulgare]
gi|12585487|sp|Q9SCB9|VATC_HORVU Vacuolar ATP synthase
subunit C (V-ATPase C subunit) (Vacuolar proton pump C
subunit)
Length = 354
Score = 51.2 bits (121), Expect = 7e-06
Identities = 25/60 (41%), Positives = 38/60 (62%), Gaps = 7/60 (11%)
Query: 2 AVVWWVVSLPVHNS-----ASELWNRLQQNVSKHSFDTPLYRFNIPISGSEP*TLSAMIS 56
A +W+ +LPV + + LW RLQ+ +S+HSFDTPLYRF +P P TL ++++
Sbjct: 2 ATRYWIAALPVADDNVAAGKTALWARLQEAISRHSFDTPLYRFTVP--DLRPGTLDSLLA 59
>emb|CAD63217.1| ATPase of the PilT family [Lactobacillus plantarum WCFS1]
gi|28377484|ref|NP_784376.1| ATPase of the PilT family
[Lactobacillus plantarum WCFS1]
Length = 391
Score = 31.6 bits (70), Expect = 5.9
Identities = 15/37 (40%), Positives = 27/37 (72%)
Query: 1 GAVVWWVVSLPVHNSASELWNRLQQNVSKHSFDTPLY 37
GA++ WV+SL + N+A L +R+++N+SK + T L+
Sbjct: 47 GALILWVLSLFLANAAVRLVDRVEKNLSKQNPMTLLF 83
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.327 0.135 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 90,668,424
Number of Sequences: 2540612
Number of extensions: 2419930
Number of successful extensions: 7420
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 7413
Number of HSP's gapped (non-prelim): 5
length of query: 56
length of database: 863,360,394
effective HSP length: 32
effective length of query: 24
effective length of database: 782,060,810
effective search space: 18769459440
effective search space used: 18769459440
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0199.9


