

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0197.5
(317 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
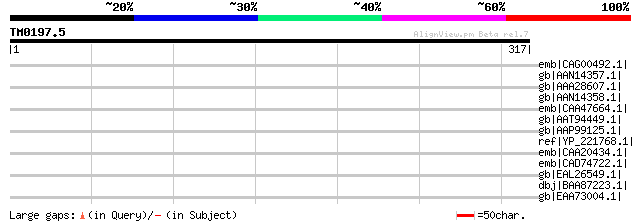
Score E
Sequences producing significant alignments: (bits) Value
emb|CAG00492.1| unnamed protein product [Tetraodon nigroviridis] 35 2.2
gb|AAN14357.1| CG5460-PD, isoform D [Drosophila melanogaster] gi... 35 2.9
gb|AAA28607.1| hairless protein 35 2.9
gb|AAN14358.1| CG5460-PC, isoform C [Drosophila melanogaster] gi... 35 2.9
emb|CAA47664.1| hairless [Drosophila melanogaster] gi|462278|sp|... 35 2.9
gb|AAT94449.1| RE41617p [Drosophila melanogaster] 35 2.9
gb|AAP99125.1| ABC-type uncharacterized transport system permeas... 35 3.8
ref|YP_221768.1| hypothetical protein BruAb1_1059 [Brucella abor... 34 4.9
emb|CAA20434.1| SPCC962.02c [Schizosaccharomyces pombe] gi|31834... 34 6.4
emb|CAD74722.1| probable response-regulator [Rhodopirellula balt... 34 6.4
gb|EAL26549.1| GA15693-PA [Drosophila pseudoobscura] 34 6.4
dbj|BAA87223.1| Hypothetical nuclear protein [Schizosaccharomyce... 34 6.4
gb|EAA73004.1| hypothetical protein FG08043.1 [Gibberella zeae P... 33 8.4
>emb|CAG00492.1| unnamed protein product [Tetraodon nigroviridis]
Length = 477
Score = 35.4 bits (80), Expect = 2.2
Identities = 42/221 (19%), Positives = 88/221 (39%), Gaps = 29/221 (13%)
Query: 65 EMEGLVVFSNGVSTARKP--ETVVGE-SSSERDQYRPESDWNRSSVSRKANIAREVSSPV 121
E EG V + A K + +VG ++++ D+ R + + ++ S++ + E S P
Sbjct: 140 EAEGEGVAAEEARRADKAIEQLIVGHLTAAQGDEVRERDEEDETTRSQERGWSTEESGPE 199
Query: 122 AVEQVAGSKPQAENANHEEKPLDLGRMSHLVCGESKGATEEAMSVTAEARHVVKNADVGL 181
+V + G E + K DLG + + G+ + +S+ + R +N D+GL
Sbjct: 200 SVSEAEGK----EKDQSQGKGTDLGALL-----KETGSKGDELSLQRKIRGYFQNIDLGL 250
Query: 182 MRKEVTVLMVG----------------WVHNWLVGSLRMGTRWVVQINNVWLR-SEQVCG 224
E+ + G W NW+ G ++ E+V
Sbjct: 251 QDNEILPPLKGYKAYNTQLARAGKKPHWQENWMGKQPAKGGNFMDDFEGEGEELEEEVED 310
Query: 225 DTQKAAKEDSWGASDASLQQVMVGEHEVKDFSPDQARVSQV 265
+ Q + + + A Q+V+ + E + ++ R++ +
Sbjct: 311 EEQSLTRMEEEARAQAEKQEVLRQQEEAERAREEEQRLADI 351
>gb|AAN14357.1| CG5460-PD, isoform D [Drosophila melanogaster]
gi|7300641|gb|AAF55790.1| CG5460-PA, isoform A
[Drosophila melanogaster] gi|24648465|ref|NP_732534.1|
CG5460-PD, isoform D [Drosophila melanogaster]
gi|24648463|ref|NP_732533.1| CG5460-PA, isoform A
[Drosophila melanogaster]
Length = 1077
Score = 35.0 bits (79), Expect = 2.9
Identities = 23/97 (23%), Positives = 49/97 (49%), Gaps = 3/97 (3%)
Query: 64 LEMEGLVVFSNGVSTAR--KPETVVGESSSERDQYRPESDWNRSSVSRKANIAREVSSPV 121
+E+E + V + + KP+T+ GE +ER + P+ + S S++A+ ++V P
Sbjct: 391 VEIENVAVADTTTNEIKIEKPDTIKGEDDAERLEKEPKKAVSDDSESKEASPGQQV-EPQ 449
Query: 122 AVEQVAGSKPQAENANHEEKPLDLGRMSHLVCGESKG 158
++ + + + EE +L R+++ V G+ G
Sbjct: 450 PKDETVDVEMKMNTSEDEEPMTELPRITNAVNGDLNG 486
>gb|AAA28607.1| hairless protein
Length = 1059
Score = 35.0 bits (79), Expect = 2.9
Identities = 23/97 (23%), Positives = 49/97 (49%), Gaps = 3/97 (3%)
Query: 64 LEMEGLVVFSNGVSTAR--KPETVVGESSSERDQYRPESDWNRSSVSRKANIAREVSSPV 121
+E+E + V + + KP+T+ GE +ER + P+ + S S++A+ ++V P
Sbjct: 373 VEIENVAVADTTTNEIKIEKPDTIKGEDDAERLEKEPKKAVSDDSESKEASPGQQV-EPQ 431
Query: 122 AVEQVAGSKPQAENANHEEKPLDLGRMSHLVCGESKG 158
++ + + + EE +L R+++ V G+ G
Sbjct: 432 PKDETVDVEMKMNTSEDEEPMTELPRITNAVNGDLNG 468
>gb|AAN14358.1| CG5460-PC, isoform C [Drosophila melanogaster]
gi|7300642|gb|AAF55791.1| CG5460-PB, isoform B
[Drosophila melanogaster] gi|24648467|ref|NP_732535.1|
CG5460-PB, isoform B [Drosophila melanogaster]
gi|24648469|ref|NP_524418.2| CG5460-PC, isoform C
[Drosophila melanogaster]
Length = 1059
Score = 35.0 bits (79), Expect = 2.9
Identities = 23/97 (23%), Positives = 49/97 (49%), Gaps = 3/97 (3%)
Query: 64 LEMEGLVVFSNGVSTAR--KPETVVGESSSERDQYRPESDWNRSSVSRKANIAREVSSPV 121
+E+E + V + + KP+T+ GE +ER + P+ + S S++A+ ++V P
Sbjct: 373 VEIENVAVADTTTNEIKIEKPDTIKGEDDAERLEKEPKKAVSDDSESKEASPGQQV-EPQ 431
Query: 122 AVEQVAGSKPQAENANHEEKPLDLGRMSHLVCGESKG 158
++ + + + EE +L R+++ V G+ G
Sbjct: 432 PKDETVDVEMKMNTSEDEEPMTELPRITNAVNGDLNG 468
>emb|CAA47664.1| hairless [Drosophila melanogaster] gi|462278|sp|Q02308|HLES_DROME
Hairless protein
Length = 1077
Score = 35.0 bits (79), Expect = 2.9
Identities = 23/97 (23%), Positives = 49/97 (49%), Gaps = 3/97 (3%)
Query: 64 LEMEGLVVFSNGVSTAR--KPETVVGESSSERDQYRPESDWNRSSVSRKANIAREVSSPV 121
+E+E + V + + KP+T+ GE +ER + P+ + S S++A+ ++V P
Sbjct: 391 VEIENVAVADTTTNEIKIEKPDTIKGEDDAERLEKEPKKAVSDDSESKEASPGQQV-EPQ 449
Query: 122 AVEQVAGSKPQAENANHEEKPLDLGRMSHLVCGESKG 158
++ + + + EE +L R+++ V G+ G
Sbjct: 450 PKDETVDVEMKMNTSEDEEPMTELPRITNAVNGDLNG 486
>gb|AAT94449.1| RE41617p [Drosophila melanogaster]
Length = 1077
Score = 35.0 bits (79), Expect = 2.9
Identities = 23/97 (23%), Positives = 49/97 (49%), Gaps = 3/97 (3%)
Query: 64 LEMEGLVVFSNGVSTAR--KPETVVGESSSERDQYRPESDWNRSSVSRKANIAREVSSPV 121
+E+E + V + + KP+T+ GE +ER + P+ + S S++A+ ++V P
Sbjct: 391 VEIENVAVADTTTNEIKIEKPDTIKGEDDAERLEKEPKKAVSDDSESKEASPGQQV-EPQ 449
Query: 122 AVEQVAGSKPQAENANHEEKPLDLGRMSHLVCGESKG 158
++ + + + EE +L R+++ V G+ G
Sbjct: 450 PKDETVDVEMKMNTSEDEEPMTELPRITNAVNGDLNG 486
>gb|AAP99125.1| ABC-type uncharacterized transport system permease and ATPase
component [Prochlorococcus marinus subsp. marinus str.
CCMP1375] gi|33239531|ref|NP_874473.1| ABC-type
uncharacterized transport system permease and ATPase
component [Prochlorococcus marinus subsp. marinus str.
CCMP1375]
Length = 662
Score = 34.7 bits (78), Expect = 3.8
Identities = 20/72 (27%), Positives = 39/72 (53%), Gaps = 2/72 (2%)
Query: 43 RFMLAELTIEAMIDGWLFIVLLEMEGLVVFSNGVSTARKPETVVGESSSERDQYRPESD- 101
RF+ A M++G LF ++ ++E L F+ G+S ++ V + S ++D + D
Sbjct: 385 RFIQASFAF-GMVEGSLFFIVNQIEELAKFTAGISRLEGFQSKVEKVSRQKDSSQENIDS 443
Query: 102 WNRSSVSRKANI 113
WN S + + A++
Sbjct: 444 WNNSIIIKNADL 455
>ref|YP_221768.1| hypothetical protein BruAb1_1059 [Brucella abortus biovar 1 str.
9-941] gi|62196107|gb|AAX74407.1| conserved hypothetical
protein [Brucella abortus biovar 1 str. 9-941]
gi|23347876|gb|AAN29974.1| conserved hypothetical
protein [Brucella suis 1330] gi|17982886|gb|AAL52113.1|
metal-dependent hydrolase [Brucella melitensis 16M]
gi|17987215|ref|NP_539849.1| ALANYL-TRNA SYNTHETASE
[Brucella melitensis 16M] gi|25529279|pir||AF3368
alanine-tRNA ligase (EC 6.1.1.7) [imported] - Brucella
melitensis (strain 16M) gi|23501932|ref|NP_698059.1|
hypothetical protein BR1054 [Brucella suis 1330]
Length = 247
Score = 34.3 bits (77), Expect = 4.9
Identities = 23/99 (23%), Positives = 44/99 (44%), Gaps = 8/99 (8%)
Query: 86 VGESSSERDQYRPESDWNRSSVSRKANIAREVSSPVAVEQVAG----SKPQAENANHEEK 141
VGE S D P+ + + SV+ K + ++PV++ ++ + P + +
Sbjct: 126 VGEDESRVDFDLPDQSYTKESVTEKLMELVQANAPVSIHWISDEEFLANPDIVKSKNVRP 185
Query: 142 PLDLGRMSHLVCGESKGATEEAMSVTAEARHVVKNADVG 180
P+ LGR+ + GE+ + T HV + +VG
Sbjct: 186 PVGLGRIRLVAIGENGSVDSQPCGGT----HVSETQEVG 220
>emb|CAA20434.1| SPCC962.02c [Schizosaccharomyces pombe]
gi|3183409|sp|O14064|BIR1_SCHPO Bir1 protein (Chromosome
segregation protein cut17) gi|5738948|dbj|BAA83415.1|
Cut17 [Schizosaccharomyces pombe]
Length = 997
Score = 33.9 bits (76), Expect = 6.4
Identities = 25/67 (37%), Positives = 32/67 (47%), Gaps = 9/67 (13%)
Query: 76 VSTARKPETVVGESSSE--RDQYRPESDWNRSSVSRKANIAREVSSPVAVEQVAGSKPQA 133
V KP+T + E E R ES R S+ R R+VSSPV+ E ++
Sbjct: 642 VDFIEKPKTEISEVLPEEKRKAICDESQTVRVSIDRGVTKTRDVSSPVSDE-------KS 694
Query: 134 ENANHEE 140
EN NHEE
Sbjct: 695 ENVNHEE 701
>emb|CAD74722.1| probable response-regulator [Rhodopirellula baltica SH 1]
gi|32474183|ref|NP_867177.1| probable response-regulator
[Rhodopirellula baltica SH 1]
Length = 257
Score = 33.9 bits (76), Expect = 6.4
Identities = 50/172 (29%), Positives = 74/172 (42%), Gaps = 19/172 (11%)
Query: 23 RVKRLGAIKVLLEFEIGEDMRFMLAELTIEAMIDGWLFIVL--LEMEGLV-VFSNGVSTA 79
R++R G + V +E IG+ + + L+ EA L +V L +E +V F NG T
Sbjct: 56 RLERPGCLVVDVEL-IGDGFQRVQKALS-EASCSAPLILVAGELPIETVVHAFENGAWTV 113
Query: 80 --RKPETVVGESSSERDQYRPESDWNRSSVS-----RKAN------IAREVSSPVAVEQV 126
+ + S S RD R DW+R V RK N R+ V +
Sbjct: 114 VLKSADNAQKFSGSLRDHIRQAIDWDRFQVGMEKAHRKRNRILDGLTERQRKVLNCVMEG 173
Query: 127 AGSKPQAENANHEEKPLDLGRMSHLVCGESKGATEEAMSVTAEARHVVKNAD 178
+K A N N ++ ++ R SHL+ + G T E +V E R V K D
Sbjct: 174 MPTKAIAANYNVSKRLIEFER-SHLLSAFNVGGTAELTAVVGEHRIVEKLLD 224
>gb|EAL26549.1| GA15693-PA [Drosophila pseudoobscura]
Length = 956
Score = 33.9 bits (76), Expect = 6.4
Identities = 20/61 (32%), Positives = 32/61 (51%), Gaps = 12/61 (19%)
Query: 247 VGEHEVKD--------FSPDQARVSQVVDARNTNIQDGFRSGIAVDDDVSQVCVTPQEGS 298
VGE +V+D F+P A S + RN N+ + +R G +DVS +C+ PQ+
Sbjct: 892 VGECQVQDTVTPVKPRFAPMSASTSTPLSNRNINLVEEYRGG----EDVSPICMRPQKAP 947
Query: 299 K 299
+
Sbjct: 948 R 948
>dbj|BAA87223.1| Hypothetical nuclear protein [Schizosaccharomyces pombe]
Length = 234
Score = 33.9 bits (76), Expect = 6.4
Identities = 25/67 (37%), Positives = 32/67 (47%), Gaps = 9/67 (13%)
Query: 76 VSTARKPETVVGESSSE--RDQYRPESDWNRSSVSRKANIAREVSSPVAVEQVAGSKPQA 133
V KP+T + E E R ES R S+ R R+VSSPV+ E ++
Sbjct: 66 VDFIEKPKTEISEVLPEEKRKAICDESQTVRVSIDRGVTKTRDVSSPVSDE-------KS 118
Query: 134 ENANHEE 140
EN NHEE
Sbjct: 119 ENVNHEE 125
>gb|EAA73004.1| hypothetical protein FG08043.1 [Gibberella zeae PH-1]
gi|46127331|ref|XP_388219.1| hypothetical protein
FG08043.1 [Gibberella zeae PH-1]
Length = 545
Score = 33.5 bits (75), Expect = 8.4
Identities = 22/50 (44%), Positives = 26/50 (52%), Gaps = 1/50 (2%)
Query: 229 AAKEDSWGASDASLQQVMVGEHEVKDFSPDQARVSQVVDARNTNIQDGFR 278
AA+ DS G S A + GE D QAR+SQV +A T QDG R
Sbjct: 114 AAQADS-GFSLAPTPALRTGEERPPDIGSLQARLSQVAEAERTTWQDGAR 162
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.131 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 492,001,046
Number of Sequences: 2540612
Number of extensions: 18696396
Number of successful extensions: 46113
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 46111
Number of HSP's gapped (non-prelim): 13
length of query: 317
length of database: 863,360,394
effective HSP length: 128
effective length of query: 189
effective length of database: 538,162,058
effective search space: 101712628962
effective search space used: 101712628962
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0197.5


