

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0178.7
(1813 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
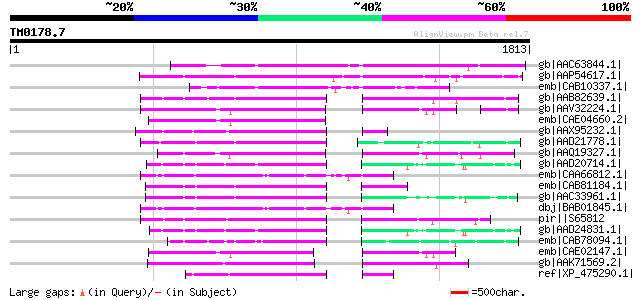
Score E
Sequences producing significant alignments: (bits) Value
gb|AAC63844.1| putative non-LTR retroelement reverse transcripta... 513 e-143
gb|AAP54617.1| putative non-LTR retroelement reverse transcripta... 487 e-135
emb|CAB10337.1| reverse transcriptase like protein [Arabidopsis ... 446 e-123
gb|AAB82639.1| putative non-LTR retroelement reverse transcripta... 345 1e-92
gb|AAV32224.1| hypothetical protein [Oryza sativa (japonica cult... 336 5e-90
emb|CAE04660.2| OSJNBa0061G20.16 [Oryza sativa (japonica cultiva... 333 2e-89
gb|AAX95232.1| retrotransposon protein, putative, unclassified [... 331 2e-88
gb|AAD21778.1| putative non-LTR retroelement reverse transcripta... 330 3e-88
gb|AAQ19327.1| bZIP-like protein [Oryza sativa (japonica cultiva... 314 2e-83
gb|AAD20714.1| putative non-LTR retroelement reverse transcripta... 312 8e-83
emb|CAA66812.1| non-ltr retrotransposon reverse transcriptase-li... 305 7e-81
emb|CAB81184.1| putative protein [Arabidopsis thaliana] gi|45393... 305 9e-81
gb|AAC33961.1| contains similarity to reverse trancriptase (Pfam... 305 9e-81
dbj|BAB01845.1| non-LTR retroelement reverse transcriptase-like ... 305 1e-80
pir||S65812 RNA-directed DNA polymerase (EC 2.7.7.49) (clone DW1... 303 3e-80
gb|AAD24831.1| putative non-LTR retroelement reverse transcripta... 302 6e-80
emb|CAB78094.1| RNA-directed DNA polymerase-like protein [Arabid... 293 3e-77
emb|CAE02147.1| OSJNBa0081G05.2 [Oryza sativa (japonica cultivar... 288 9e-76
gb|AAK71569.2| putative reverse transcriptase [Oryza sativa (jap... 278 1e-72
ref|XP_475290.1| hypothetical protein [Oryza sativa (japonica cu... 276 4e-72
>gb|AAC63844.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25408124|pir||C84716 hypothetical protein
At2g31080 [imported] - Arabidopsis thaliana
Length = 1231
Score = 513 bits (1322), Expect = e-143
Identities = 387/1279 (30%), Positives = 595/1279 (46%), Gaps = 102/1279 (7%)
Query: 561 IYASPIPVKREELWSHLLHIRSNLVLPWMIVGDMNEILHPNEVRGG*F-LPTRASRFAAI 619
+YA+P +R LW L + + L P +I GD N IL +E GG L + F
Sbjct: 6 VYAAPSVSRRSGLWGELKDVVNGLEGPLLIGGDFNTILWVDERMGGNGRLSPDSLAFGDW 65
Query: 620 LEGCGMVDLGMSGGRYTWFRKQNNRLIMSKRLDRAMGDGAWRIAFPEASVEALHRIHSDH 679
+ ++DLG G ++TW R + +++KRLDR R+ + EA V L + SDH
Sbjct: 66 INELSLIDLGFKGNKFTWRRGRQESTVVAKRLDRVFVCAHARLKWQEAVVSHLPFMASDH 125
Query: 680 CPLLINCSGPEIVHHDRPFRFLASWVEHPQYKEVVQSAWTGETETVAGKLNKVRTKSCKF 739
PL + E +Q K+R K+
Sbjct: 126 APLYVQL-------------------------EPLQQ-------------RKLR----KW 143
Query: 740 NKEVYGSIFQRKRWVEGRLRGVQRELNYRVTFDMTRLETELQAEYRFLLRQEELLWFQKA 799
N+EV+G I RK + ++ VQ L ++ D+ E L E +L QEE LWFQK+
Sbjct: 144 NREVFGDIHVRKEKLVADIKEVQDLLGVVLSDDLLAKEEVLLKEMDLVLEQEETLWFQKS 203
Query: 800 RENRVCFDDRNTAYFHAHTIIRRKRNKIHRLKLNDGSWCDDRQLLAEEIQNYFQSLFAAD 859
RE + DRNT +FH TIIRR+RN+I LK +D W D+ L Y++ L++ +
Sbjct: 204 REKYIELGDRNTTFFHTSTIIRRRRNRIESLKGDDDRWVTDKVELEAMALTYYKRLYSLE 263
Query: 860 PDNTDVNMQHVMMPR-----LTEAERHKAEAEVIKEEVFQALLRMRSYSALGPDGFHPFF 914
DV+ M+P ++EAE+ K EV A+ M + A GPDG+ P F
Sbjct: 264 ----DVSEVRNMLPTGGFASISEAEKAALLQAFTKAEVVSAVKSMGRFKAPGPDGYQPVF 319
Query: 915 FKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKEFRPISLCNVVYK 974
+++ WE VG + + V + FETG++ S + L+VLI KV P +++FRP+SLCNV++K
Sbjct: 320 YQQCWETVGPSVTRFVLEFFETGVLPASTNDALLVLIAKVAKPERIQQFRPVSLCNVLFK 379
Query: 975 LITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEIIHHMGASMAKKGTLAFKID 1034
+ITK++V R++ +++++IGP Q SF+PG +DN L QE +H M +KG + K+D
Sbjct: 380 IITKMMVTRLKNVISKLIGPAQASFIPGRLSIDNIVLVQEAVHSMRRKKGRKGWMLLKLD 439
Query: 1035 LEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGSRLPSFKPGRGLRKG 1094
LEKAYD V W FL++TL G IM V+ ++S+LWNG R SF P RGLR+G
Sbjct: 440 LEKAYDRVRWDFLQETLEAAGLSEGWTSRIMAGVTDPSMSVLWNGERTDSFVPARGLRQG 499
Query: 1095 DPLSPYLFVL-----CSLHGAIV----Y*YS*VGGNRSLEAN*NFPKGTTYFASLFCR*R 1145
DPLSPYLFVL C L A V + V S ++ F FA +
Sbjct: 500 DPLSPYLFVLCLERLCHLIEASVGKREWKPIAVSCGGSKLSHVCFADDLILFAEASVA-Q 558
Query: 1146 ALVLSSII--FSSAIGGGDS*TILCLF---GVKDKYEQI*SDCFQGSTKSCERGDPEYCP 1200
++ ++ F A G S +F V + EQ+ S+ S C + +Y
Sbjct: 559 IRIIRRVLERFCEASGQKVSLEKSKIFFSHNVSREMEQLISE---ESGIGCTKELGKYLG 615
Query: 1201 YPFCAKPWQVSWLEEGFQEVDSITYWIIFKKKLSSWKTNLLNMVGQVCLAKSVISAIPTY 1260
P K E + V + +L+ WK L++ G++ L K+V+S+IP +
Sbjct: 616 MPILQKRMNKETFGEVLERVSA---------RLAGWKGRSLSLAGRITLTKAVLSSIPVH 666
Query: 1261 TMQVFWLPRSINHKINQEMRRFIWLKNSGQRGWHRVNWKTLTQPKEHGGLAIKDMGDFNT 1320
M LP S +++ R F+W ++ H ++W+ + +PK GG+ ++ D N
Sbjct: 667 VMSAILLPVSTLDTLDRYSRTFLWGSTMEKKKQHLLSWRKICKPKAEGGIGLRSARDMNK 726
Query: 1321 SLLGKVVWLLANNSSKLWVQVLAHKYLHG--ESIFSVHRRANASPIWQGI-LKARDQLQE 1377
+L+ KV W L + LW +V+ KY G + + + S W+ + + R+ + +
Sbjct: 727 ALVAKVGWRLLQDKESLWARVVRKKYKVGGVQDTSWLKPQPRWSSTWRSVAVGLREVVVK 786
Query: 1378 GFQFRMGDGSS-SVWYHDWTGKGKLAE-SLDFVHISD-TKLCLHELVDNNQWNLNRCMTV 1434
G + GDG + W W + L E D + + K+ + + WNL
Sbjct: 787 GVGWVPGDGCTIRFWLDRWLLQEPLVELGTDMIPEGERIKVAADYWLPGSGWNLEILGLY 846
Query: 1435 FPLHIRDWFARVEPRVVANGIDRWSWKDNSDGISSVHDGYQWLREKKMAGVVHGDSWLWI 1494
P ++ V +V D SWK DG +V Y L+ G + I
Sbjct: 847 LPETVKRRLLSVVVQVFLGNGDEISWKGTQDGAFTVRSAYSLLQGDVGDRPNMGSFFNRI 906
Query: 1495 *KLHAPEKL*VFIWLVLHDALPTNANRYRCKLAVSPGCTRCSSPEEDTIHCLRDCPHSQE 1554
KL PE++ VFIWLV + + TN R R L+ + C+ C+ EE +H LRDCP +
Sbjct: 907 WKLITPERVRVFIWLVSQNVIMTNVERVRRHLSENAICSVCNGAEETILHVLRDCPAMEP 966
Query: 1555 LWGK-LGAWGWSTFRLPDLKA*VKTQLQGSNNI---RFLAGLWGVWKWR-------NNVV 1603
+W + L F L + T + I F G+W WKWR +
Sbjct: 967 IWRRLLPLRRHHEFFSQSLLEWLFTNMDPVKGIWPTLFGMGIWWAWKWRCCDVFGERKIC 1026
Query: 1604 LDPLPWTLDGAWQKLRHDHDELLQFSSGDLLDEHHFLINLWKPPPPEFVKLNTDGSFQDT 1663
D L + D A +++R H + + E + W+ P +VK+ TDG+ +
Sbjct: 1027 RDRLKFIKDMA-EEVRRVHVGAVGNRPNGVRVER---MIRWQVPSDGWVKITTDGASRGN 1082
Query: 1664 ASSMGGGGLIRDAKGAWLGGFMSHTSIGNPFLAEVLALRDGLCLAWVQGHRRIIREVDCA 1723
GG IR+ +G WLGGF + LAE+ GL +AW +G RR+ ++DC
Sbjct: 1083 HGLAAAGGAIRNGQGEWLGGFALNIGSCAAPLAELWGAYYGLLIAWDKGFRRVELDLDCK 1142
Query: 1724 GLSDILRFEDRCRLHDHAGVLQEIRELMERPWRCSVHWTNRDSNASADFLARRGGFLSTI 1783
+ L H + +++ + R W V R++N AD LA F +
Sbjct: 1143 LVVGFLS-TGVSNAHPLSFLVRLCQGFFTRDWLVRVSHVYREANRLADGLANY-AFTLPL 1200
Query: 1784 GVVELMVAPLELEHLLLKD 1802
G+ P + LLL D
Sbjct: 1201 GLHCFDACPEGVRLLLLAD 1219
>gb|AAP54617.1| putative non-LTR retroelement reverse transcriptase [Oryza sativa
(japonica cultivar-group)] gi|37536056|ref|NP_922330.1|
putative non-LTR retroelement reverse transcriptase
[Oryza sativa (japonica cultivar-group)]
gi|10140689|gb|AAG13524.1| putative non-LTR retroelement
reverse transcriptase [Oryza sativa (japonica
cultivar-group)]
Length = 1382
Score = 487 bits (1253), Expect = e-135
Identities = 389/1398 (27%), Positives = 644/1398 (45%), Gaps = 89/1398 (6%)
Query: 453 IEDLKIISW--NVRGALNEHGQLFLKDLIRCKKPDIMFLLETRCQFSRASNFWNSTRFSP 510
+E K+++W N RG + L+ L++ +P ++FL ET+ + +A N S FS
Sbjct: 1 MEPNKLVTWGRNCRGLGSAATVGELRWLVKSLRPSLVFLSETKMRDKQARNLMWSLGFSG 60
Query: 511 LFISEANGFSGGIWVLVNTATNLSSRLLYSHQQAVTFEVWRENLSWVCSAIYASPIPVKR 570
F G SGG+ + TA +S R SH V E W S +Y P R
Sbjct: 61 SFAVSCEGLSGGLALFWTTAYTVSLRGFNSHFIDVLVST-EELPPWRISFVYGEPKRELR 119
Query: 571 EELWSHLLHIRSNLVLPWMIVGDMNEILHPNEVRGG*FLPTRAS----RFAAILEGCGMV 626
W+ L + PW+ GD NE+L +E G + R+ F + L+ CG++
Sbjct: 120 HFFWNLLRRLHDQWRGPWLCCGDFNEVLCLDEHLG---MRERSEPHMQHFRSCLDDCGLI 176
Query: 627 DLGMSGGRYTWFRKQNNRLIMSKRLDRAMGDGAWRIAFPEASVEALHRIHSDHCPLLINC 686
DLG G ++TW KQ+ RLDRA+ +G + F + VE + SDH + I+
Sbjct: 177 DLGFVGPKFTWSNKQDANSNSKVRLDRAVANGEFSRYFEDCLVENVITTSSDHYAISIDL 236
Query: 687 S----GPEIVHHDRPFRFLASWVEHPQYKEVVQSAWTGETETVAGK------LNKVRTKS 736
S G + + FRF A+W+ Y+EVV+++W + G L +V
Sbjct: 237 SRRNHGQRRIPIQQGFRFEAAWLRAEDYREVVENSWRISSAGCVGLRGVWSVLQQVAVSL 296
Query: 737 CKFNKEVYGSIFQRKRWVEGRLRGVQRELNYRVTFDMTRLETELQAEYRFLLRQEELLWF 796
++K +GS+ ++ +E +L+ +++ V +L + E L +EE++
Sbjct: 297 KDWSKASFGSVRRKILKMERKLKSLRQSPVNDVVIQEEKLIEQQLCE---LFEKEEIMAR 353
Query: 797 QKARENRVCFDDRNTAYFHAHTIIRRKRNKIHRLKLNDGSWCDDRQLLAEEIQNYFQSLF 856
Q++R + + DRNTA+FHA RR+ N+I L +DGS C ++ + + ++++LF
Sbjct: 354 QRSRVDWLREGDRNTAFFHARASARRRTNRIKELVRDDGSRCISQEGIKRMAEVFYENLF 413
Query: 857 AADPDNTDVNMQHVMMPRLTEAERHKAEAEVIKEEVFQALLRMRSYSALGPDGFHPFFFK 916
+++P ++ + + ++ + + + EE+ AL +M S A GPDGF F++
Sbjct: 414 SSEPCDSMEEVLDAIPNKVGDFINGELGKQYTNEEIKTALFQMGSTKAPGPDGFPALFYQ 473
Query: 917 KYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKEFRPISLCNVVYKLI 976
+W ++ + + VR I + L ++++VLIPKV++ S + +FRPISLCNV+YK+
Sbjct: 474 THWGILEEHICNAVRGFLLGEEIPEGLCDSVVVLIPKVNNASHLSKFRPISLCNVLYKIA 533
Query: 977 TKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEIIHHMGASMAKKGTLAFKIDLE 1036
+KV+ NR++P L I+ Q++F+PG I D+A +A E +H + K A KID+
Sbjct: 534 SKVLANRLKPFLPDIVSEFQSAFVPGRLITDSALVAYECLHTIRKQHNKNPFFALKIDMM 593
Query: 1037 KAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGSRLPSFKPGRGLRKGDP 1096
KAYD V WA+L L + GF + +MRCVS ++ NG P RG+R+GDP
Sbjct: 594 KAYDRVEWAYLSGCLSKLGFSQDWINTVMRCVSSVRYAVKINGELTKPVVPSRGIRQGDP 653
Query: 1097 LSPYLFVLCSLHGAIVY*YS*VGGNRSLEAN*N---------FPKGTTYFASLFCR*RAL 1147
+SPYLF+LC+ + + V G N F + +FA R
Sbjct: 654 ISPYLFLLCTEGLSCLLHKKEVAGELQGIKNGRHGPPISHLLFADDSIFFAKADSR-NVQ 712
Query: 1148 VLSSIIFSSAIGGGDS*TILCLFGVKDKYEQI*SDCFQGSTKSCERGDPEYCPYPFCAKP 1207
L + + S G + L + + D + S KSC + D E + P
Sbjct: 713 ALKNTLRSYCSASGQK---INLHKSSIFFGKRCPDAVKISVKSCLQVDNEVLQDSYLGMP 769
Query: 1208 WQVSWLEEGFQEVDSITYWIIFKKKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWL 1267
++ F + W K+++ W L+ G + K+V AIP Y M F +
Sbjct: 770 TEIGLATTNFFKFLPERIW----KRVNGWTDRPLSRAGMETMLKAVAQAIPNYVMSCFRI 825
Query: 1268 PRSINHKINQEMRRFIWLKNSGQRGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVV 1327
P SI K+ + W G++ H +W L+ PK GG+ ++ FN ++LG+
Sbjct: 826 PVSICEKMKTCIADHWWGFEDGKKKMHWKSWSWLSTPKFLGGMGFREFTTFNQAMLGRQC 885
Query: 1328 WLLANNSSKLWVQVLAHKYLHGESIFSVHRRANASPIWQGILKARDQLQEGFQFRMGDGS 1387
W L + L +VL +Y S + + + S W+ +L R+ L +G ++ +GDG
Sbjct: 886 WRLLTDPDSLCSRVLKGRYFPNSSFWEAAQPKSPSFTWRSLLFGRELLAKGVRWGVGDGK 945
Query: 1388 S-SVWYHDWTG--KGKLAESLDFVHISDTKLCLHELVDNNQWNLNRCMTVFPLHIRDWFA 1444
+ ++ +W + +L +L T CL D W+ + ++FP+ I
Sbjct: 946 TIKIFSDNWIPGFRPQLVTTLSPFPTDATVSCLMN-EDARCWDGDLIRSLFPVDIAKEIL 1004
Query: 1445 RVEPRVVANGIDRWSWKDNSDGISSVHDGYQWLREKKMAG-------------VVHGDSW 1491
++ P D SW + G+ SV Y R + + W
Sbjct: 1005 QI-PISRHGDADFASWPHDKLGLYSVRSAYNLARSEAFFADQSNSGRGMASRLLESQKDW 1063
Query: 1492 LWI*KLHAPEKL*VFIWLVLHDALPTNANRYRCKLAVSPGCTRCSSPEEDTI-HCLRDCP 1550
+ K++AP K+ + +W H+ L T R + + GC C+ +DT+ H CP
Sbjct: 1064 KGLWKINAPGKMKITLWRAAHECLATGFQLRRRHIPSTDGCVFCN--RDDTVEHVFLFCP 1121
Query: 1551 HSQELWG--------KLGAWGWSTFR---LPDLKA*VKTQLQGSNNIRFLAGL--WGVWK 1597
+ ++W KLG G+ST R LK +GS++ L + W +W+
Sbjct: 1122 FAAQIWEEIKGKCAVKLGRNGFSTMRQWIFDFLK-------RGSSHANTLLAVTFWHIWE 1174
Query: 1598 WRNNVVLDPLPWTLDGAWQKLRHDHDELLQFSSGDLLDE---HHFLINLWKPPPPEFVKL 1654
RNN + K+ D +L+ ++ + + + I W+PPP +
Sbjct: 1175 ARNNTKNNNGTVHPQRVVIKILSYVDMILKHNTKTVDGQRGGNTQAIPRWQPPPASVWMI 1234
Query: 1655 NTDGSFQDTASSMGGGGLIRDAKGAWLGGFMSHTS-IGNPFLAEVLALRDGLCLAWVQGH 1713
N+D + ++ +MG G LIRD G L S + P LAE LA+R L LA +G
Sbjct: 1235 NSDAAIFSSSRTMGVGALIRDNTGKCLVACSEMISDVVLPELAEALAIRRALGLAKEEGL 1294
Query: 1714 RRIIREVDCAGLSDILRFEDRCRLHDHAG-VLQEIRELMERPWRCSVHWTNRDSNASADF 1772
I+ DC L+ I R + R G V+++I++L CS NR SN +A
Sbjct: 1295 EHIVMASDC--LTVIRRIQTSGRDRSGVGCVIEDIKKLASTFVLCSFMHVNRLSNLAAHS 1352
Query: 1773 LARRGGFLSTIGVVELMV 1790
LAR LST V ++
Sbjct: 1353 LARNAE-LSTCTVYRSVI 1369
>emb|CAB10337.1| reverse transcriptase like protein [Arabidopsis thaliana]
gi|7268307|emb|CAB78601.1| reverse transcriptase like
protein [Arabidopsis thaliana] gi|7485171|pir||G71420
hypothetical protein - Arabidopsis thaliana
Length = 929
Score = 446 bits (1147), Expect = e-123
Identities = 296/937 (31%), Positives = 466/937 (49%), Gaps = 97/937 (10%)
Query: 628 LGMSGGRYTWFRKQNNRLIMSKRLDRAMGDGAWRIAFPEASVEALHRIHSDHCPLLINCS 687
+G G R+TW R ++KRLDR + R+ + EA + C
Sbjct: 1 MGFKGNRFTWRRGLVESTFVAKRLDRVLFCAHARLKWQEALL----------------CP 44
Query: 688 GPEIVHHDRPFRFLASWVEHPQYKEVVQSAW-TGETETVAGKLNKVRTKSCKFNKEVYGS 746
+ RPFRF A+W+ H +KE++ ++W TG + VA LN++R + K+NKEV+G+
Sbjct: 45 AQNVDARRRPFRFEAAWLSHEGFKELLTASWDTGLSTPVA--LNRLRWQLKKWNKEVFGN 102
Query: 747 IFQRKRWVEGRLRGVQRELNYRVTFDMTRLETELQAEYRFLLRQEELLWFQKARENRVCF 806
I RK V L+ VQ L T D+ E L E+ LL QEE LWFQK+RE +
Sbjct: 103 IHVRKEKVVSDLKAVQDLLEVVQTDDLLMKEDTLLKEFDVLLHQEETLWFQKSREKLLAL 162
Query: 807 DDRNTAYFHAHTIIRRKRNKIHRLKLNDGSWCDDRQLLAEEIQNYFQSLFAADPDNTDVN 866
DRNT +FH T+IRR+RN+I LK ++ W +++ L + +Y++ L++ + DV+
Sbjct: 163 GDRNTTFFHTSTVIRRRRNRIEMLKDSEDRWVTEKEALEKLAMDYYRKLYSLE----DVS 218
Query: 867 MQHVMMP-----RLTEAERHKAEAEVIKEEVFQALLRMRSYSALGPDGFHPFFFKKYWEL 921
+ +P RLT E++ ++EV A+ M + A GPDG+ P F+++ WE
Sbjct: 219 VVRGTLPTEGFPRLTREEKNNLNRPFTRDEVVVAVRSMGRFKAPGPDGYQPVFYQQCWET 278
Query: 922 VGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKEFRPISLCNVVYKLITKVVV 981
VG+ + K V + FE+G++ S + L+VL+ KV P + +FRP+SLCNV++K+ITK++V
Sbjct: 279 VGESVSKFVMEFFESGVLPKSTNDVLLVLLAKVAKPERITQFRPVSLCNVLFKIITKMMV 338
Query: 982 NRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEIIHHMGASMAKKGTLAFKIDLEKAYDS 1041
R++ +++++IGP Q SF+PG DN + QE +H M +KG + K+DLEKAYD
Sbjct: 339 IRLKNVISKLIGPAQASFIPGRLSFDNIVVVQEAVHSMRRKKGRKGWMLLKLDLEKAYDR 398
Query: 1042 VSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGSRLPSFKPGRGLRKGDPLSPYL 1101
+ W FL +TL G ++ IM CV+G +S+LWNG + SF P RGLR+GDP+SPYL
Sbjct: 399 IRWDFLAETLEAAGLSEGWIKRIMECVAGPEMSLLWNGEKTDSFTPERGLRQGDPISPYL 458
Query: 1102 FVLC------SLHGAIVY*YS*VGGNRSLEAN*NFPKGT---------TYFASLFCR*RA 1146
FVLC + A+ G +S+ + PK + + + + +
Sbjct: 459 FVLCIERLCHQIETAVGR-----GDWKSISISQGGPKVSHVCFADDLILFAEASVAQKVS 513
Query: 1147 LVLSSIIFSSAIGGGDS*TILCLFGVKDKYEQI*SDCFQGSTKSCERGDPEYCPYPFCAK 1206
L S I FS+ + +D I ++ GST+ E G +Y P K
Sbjct: 514 LEKSKIFFSNNVS-------------RDLEGLITAETGIGSTR--ELG--KYLGMPVLQK 556
Query: 1207 PWQVSWLEEGFQEVDSITYWIIFKKKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFW 1266
E + V S +LS WK+ L++ G++ L K+V+ +IP +TM
Sbjct: 557 RINKDTFGEVLERVSS---------RLSGWKSRSLSLAGRITLTKAVLMSIPIHTMSSIL 607
Query: 1267 LPRSINHKINQEMRRFIWLKNSGQRGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKV 1326
LP S+ ++++ R F+W +R H ++WK + +PK GGL ++ D N +LL KV
Sbjct: 608 LPASLLEQLDKVSRNFLWGSTVEKRKQHLLSWKKVCRPKAAGGLGLRASKDMNRALLAKV 667
Query: 1327 VWLLANNSSKLWVQVLAHKYLHGESIFSVH------RRANASPIWQGILKARDQLQEGFQ 1380
W L N+ LW +VL KY + VH +A S W+ I G
Sbjct: 668 GWRLLNDKVSLWARVLRRKY----KVTDVHDSSWLVPKATWSSTWRSI---------GVG 714
Query: 1381 FRMGDGSSSVWYHDWTGKGKLAESLDFVHISDTKLCLHELVDNNQWNLNRCMTVFPLHIR 1440
R G + + + S + ++ + + W++ + P +
Sbjct: 715 LREGVAKGWILHEPLCTRATCLLSPEELNARVEEF----WTEGVGWDMVKLGQCLPRSVT 770
Query: 1441 DWFARVEPRVVANGIDRWSWKDNSDGISSVHDGYQWLREKKMAGVVHGDSWLWI*KLHAP 1500
D V + V DR SW+ SDG +V Y L +++ + + I + AP
Sbjct: 771 DRLHAVVIKGVLGLRDRISWQGTSDGDFTVGSAYVLLTQEEESKPCMESFFKRIWGVIAP 830
Query: 1501 EKL*VFIWLVLHDALPTNANRYRCKLAVSPGCTRCSS 1537
E++ VF+WLV + TN R R + C R S
Sbjct: 831 ERVRVFLWLVGQQVIMTNVERVRRHIGDIEVCQRLPS 867
>gb|AAB82639.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25408936|pir||A84888 hypothetical protein
At2g45230 [imported] - Arabidopsis thaliana
Length = 1374
Score = 345 bits (884), Expect = 1e-92
Identities = 209/661 (31%), Positives = 354/661 (52%), Gaps = 23/661 (3%)
Query: 456 LKIISWNVRGALNEHGQLFLKDLIRCKKPDIMFLLETRCQFSRASNFWNSTRFSPLFISE 515
++I+SWN +G N L+++ P+++FL ET+ + + N F L E
Sbjct: 1 MRILSWNCQGVGNTPTVRHLREIRGLYFPEVIFLCETKKRRNYLENVVGHLGFFDLHTVE 60
Query: 516 ANGFSGGIWVLVNTATNLSSRLLYSHQQAVTFEVWRENLSWVCSAIYASPIPVKREELWS 575
G SGG+ ++ + + ++L S ++ + + ++ + + IY P+ +R ELW
Sbjct: 61 PIGKSGGLALMWKDSVQI--KVLQSDKRLIDALLIWQDKEFYLTCIYGEPVQAERGELWE 118
Query: 576 HLLHIRSNLVLPWMIVGDMNEILHPNEVRGG*FLPTRAS----RFAAILEGCGMVDLGMS 631
L + + PWM+ GD NE++ P+E GG P R F +L CG+ ++ S
Sbjct: 119 RLTRLGLSRSGPWMLTGDFNELVDPSEKIGG---PARKESSCLEFRQMLNSCGLWEVNHS 175
Query: 632 GGRYTWFRKQNNRLIMSKRLDRAMGDGAWRIAFPEASVEALHRIHSDHCPLLINCSGPEI 691
G +++W+ +N+ L+ RLDR + + AW FP+A L +I SDH PL+ N G
Sbjct: 176 GYQFSWYGNRNDELVQC-RLDRTVANQAWMELFPQAKATYLQKICSDHSPLINNLVGDNW 234
Query: 692 VHHDRPFRFLASWVEHPQYKEVVQSAWTGE-TETVAGKLNKV---RTKSCKFNKEVYGSI 747
F++ WV+ +K+++ + W+ + T+T A + K+ R + K+ + S
Sbjct: 235 -RKWAGFKYDKRWVQREGFKDLLCNFWSQQSTKTNALMMEKIASCRREISKWKRVSKPSS 293
Query: 748 FQRKRWVEGRLRGVQRELNYRVTFDMTRLETELQAEYRFLLRQEELLWFQKARENRVCFD 807
R + ++ +L +++ + ++ RL+ EL EY EE W +K+R +
Sbjct: 294 AVRIQELQFKLDAATKQIPFDRR-ELARLKKELSQEYN----NEEQFWQEKSRIMWMRNG 348
Query: 808 DRNTAYFHAHTIIRRKRNKIHRLKLNDG-SWCDDRQLLAEEIQNYFQSLFAADPDNTDVN 866
DRNT YFHA T RR +N+I +L +G W D L + YF+ LFA++ V
Sbjct: 349 DRNTKYFHAATKNRRAQNRIQKLIDEEGREWTSDEDL-GRVAEAYFKKLFASEDVGYTVE 407
Query: 867 MQHVMMPRLTEAERHKAEAEVIKEEVFQALLRMRSYSALGPDGFHPFFFKKYWELVGDDL 926
+ P +++ + A + KEEV +A + + GPDG + F ++++WE +GD +
Sbjct: 408 ELENLTPLVSDQMNNNLLAPITKEEVQRATFSINPHKCPGPDGMNGFLYQQFWETMGDQI 467
Query: 927 WKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKEFRPISLCNVVYKLITKVVVNRIRP 986
++V+ F +G I++ + +T I LIPK+ + +FRPISLCNV+YK+I K++ NR++
Sbjct: 468 TEMVQAFFRSGSIEEGMNKTNICLIPKILKAEKMTDFRPISLCNVIYKVIGKLMANRLKK 527
Query: 987 LLNRIIGPMQNSFLPGMGIMDNAFLAQEIIHHMGA-SMAKKGTLAFKIDLEKAYDSVSWA 1045
+L +I Q +F+ G I DN +A E++H + + + + +A K D+ KAYD V W
Sbjct: 528 ILPSLISETQAAFVKGRLISDNILIAHELLHALSSNNKCSEEFIAIKTDISKAYDRVEWP 587
Query: 1046 FLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGSRLPSFKPGRGLRKGDPLSPYLFVLC 1105
FLE + GF + ++LIM CV +L NG+ P RGLR+GDPLSPYLFV+C
Sbjct: 588 FLEKAMRGLGFADHWIRLIMECVKSVRYQVLINGTPHGEIIPSRGLRQGDPLSPYLFVIC 647
Query: 1106 S 1106
+
Sbjct: 648 T 648
Score = 179 bits (453), Expect = 1e-42
Identities = 155/576 (26%), Positives = 252/576 (42%), Gaps = 35/576 (6%)
Query: 1231 KKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSGQ 1290
KK+ W++N L+ G+ L K+V A+PTYTM F +P++I +I M F W
Sbjct: 774 KKVLGWQSNFLSPGGKEILLKAVAMALPTYTMSCFKIPKTICQQIESVMAEFWWKNKKEG 833
Query: 1291 RGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLAHKYLHGE 1350
RG H W L++PK GGL K++ FN +LLGK +W + L +V +Y
Sbjct: 834 RGLHWKAWCHLSRPKAVGGLGFKEIEAFNIALLGKQLWRMITEKDSLMAKVFKSRYFSKS 893
Query: 1351 SIFSVHRRANASPIWQGILKARDQLQEGFQFRMGDGSS-SVWYHDWTG--KGKLAESLDF 1407
+ + S W+ I +A+ +++G + +G+G + +VW W G K A+++
Sbjct: 894 DPLNAPLGSRPSFAWKSIYEAQVLIKQGIRAVIGNGETINVWTDPWIGAKPAKAAQAVKR 953
Query: 1408 VHISDTKLC--LHE-----LVDNNQWNLNRCMTVFPLHIRDWFARVEPRVVANGIDRWSW 1460
H+ +H L D WN N +FP + ++ + P DR++W
Sbjct: 954 SHLVSQYAANSIHVVKDLLLPDGRDWNWNLVSLLFPDNTQENILALRPGGKETR-DRFTW 1012
Query: 1461 KDNSDGISSVHDGYQWLRE--------KKMAGVVHGDSWLWI*KLHAPEKL*VFIWLVLH 1512
+ + G SV GY + E +++ + I KL P K+ F+W ++
Sbjct: 1013 EYSRSGHYSVKSGYWVMTEIINQRNNPQEVLQPSLDPIFQQIWKLDVPPKIHHFLWRCVN 1072
Query: 1513 DALPTNANRYRCKLAVSPGCTRCSSPEEDTIHCLRDCPHSQELWG-------KLGAWGWS 1565
+ L +N LA C RC S E H L CP ++ W G W S
Sbjct: 1073 NCLSVASNLAYRHLAREKSCVRCPSHGETVNHLLFKCPFARLTWAISPLPAPPGGEWAES 1132
Query: 1566 TFR-LPDLKA*VKTQLQGSNNIRFLAG-LWGVWKWRNNVVLDPLPWTLDGAWQKLRHDHD 1623
FR + + + K+Q + S++ + LW +WK RN++V +T K D D
Sbjct: 1133 LFRNMHHVLSVHKSQPEESDHHALIPWILWRLWKNRNDLVFKGREFTAPQVILKATEDMD 1192
Query: 1624 --ELLQFSSGDLLDEHHFLINLWKPPPPEFVKLNTDGSFQDTASSMGGGGLIRDAKG--A 1679
+ + W+PP +VK NTDG++ + G G ++R+ G
Sbjct: 1193 AWNNRKEPQPQVTSSTRDRCVKWQPPSHGWVKCNTDGAWSKDLGNCGVGWVLRNHTGRLL 1252
Query: 1680 WLGGFMSHTSIGNPFLAEVLALRDGLCLAWVQGHRRIIREVDCAGLSDILRFEDRCRLHD 1739
WL G + S + EV ALR + +RR+I E D L ++ ++ +
Sbjct: 1253 WL-GLRALPSQQSVLETEVEALRWAVLSLSRFNYRRVIFESDSQYLVSLI--QNEMDIPS 1309
Query: 1740 HAGVLQEIRELMERPWRCSVHWTNRDSNASADFLAR 1775
A +Q+IR L+ +T R+ N AD AR
Sbjct: 1310 LAPRIQDIRNLLRHFEEVKFQFTRREGNNVADRTAR 1345
>gb|AAV32224.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|45267888|gb|AAS55787.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 1936
Score = 336 bits (861), Expect = 5e-90
Identities = 208/659 (31%), Positives = 334/659 (50%), Gaps = 20/659 (3%)
Query: 456 LKIISWNVRGALNEHGQLFLKDLIRCKKPDIMFLLETRCQFSRASNFWNSTRFSPLFISE 515
+ ++WN RG N L+ LI+ ++FL ETR + S F
Sbjct: 636 MSCLAWNCRGLGNTATVQDLRALIQKAGSQLVFLCETRQSVEKMSRLRRKLAFRGFVGVS 695
Query: 516 ANGFSGGIWVLVNTATNLSSRLLYSHQQAVTFEVWRENLSWVCSAIYASPIPVKREELWS 575
+ G SGG+ + + + ++ + + + + W + +Y P R +WS
Sbjct: 696 SEGKSGGLALYWDESVSVDVKDINKRYIDAYVRLSPDEPQWHITFVYGEPRVENRHRMWS 755
Query: 576 HLLHIRSNLVLPWMIVGDMNEILHPNE-VRGG*FLPTRASRFAAILEGCGMVDLGMSGGR 634
L IR + LPWM++GD NE L E T+ F L C + DLG G
Sbjct: 756 LLRTIRQSSALPWMVIGDFNETLWQFEHFSKNPRCETQMQNFRDALYDCDLQDLGFKGVP 815
Query: 635 YTWFRKQNNRLIMSKRLDRAMGDGAWRIAFPEASVEALHRIHSDHCPLLINCSGPEIVH- 693
+T+ +++ + RLDRA+ D WR FPEA V L SDH P+L+ +
Sbjct: 816 HTYDNRRDGWRNVKVRLDRAVADDKWRDLFPEAQVSHLVSPCSDHSPILLEFIVKDTTRP 875
Query: 694 HDRPFRFLASWVEHPQYKEVVQSAWTGETETVAGKLNKVRTKSCKFNKEVYGSIFQRKRW 753
+ + W P+ +V++ AW AG V+T N + + + W
Sbjct: 876 RQKCLHYEIVWEREPESVQVIEEAWIN-----AG----VKTDLGDINIALGRVMSALRSW 926
Query: 754 VEGRLRGVQRELNYR-------VTFDMTRLETELQAEY-RFLLRQEELLWFQKARENRVC 805
+ +++ V +EL + + R ++ +L +EE+LW Q++R N +
Sbjct: 927 SKTKVKNVGKELEKARKKLEDLIASNAARSSIRQATDHMNEMLYREEMLWLQRSRVNWLK 986
Query: 806 FDDRNTAYFHAHTIIRRKRNKIHRLKLNDGSWCDDRQLLAEEIQNYFQSLFAADPDNTDV 865
DRNT +FH+ + R K+NKI +L+ +G+ +L YFQ ++ ADP
Sbjct: 987 EGDRNTRFFHSRAVWRAKKNKISKLRDENGAIHSTTSVLETMATEYFQGVYKADPSLNPE 1046
Query: 866 NMQHVMMPRLTEAERHKAEAEVIKEEVFQALLRMRSYSALGPDGFHPFFFKKYWELVGDD 925
++ + ++T+A K E +EE+ QA+ ++ + PDGF F+++ W + D
Sbjct: 1047 SVTRLFQEKVTDAMNEKLCQEFKEEEIAQAIFQIGPLKSPRPDGFPARFYQRNWGTLKSD 1106
Query: 926 LWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKEFRPISLCNVVYKLITKVVVNRIR 985
+ VR+ F++G++ + +T IVLIPK D P +K++RPISLCNVVYK+++K +VNR+R
Sbjct: 1107 IILAVRNFFQSGLMPKGVNDTAIVLIPKKDQPIDLKDYRPISLCNVVYKVVSKCLVNRLR 1166
Query: 986 PLLNRIIGPMQNSFLPGMGIMDNAFLAQEIIHHMGAS-MAKKGTLAFKIDLEKAYDSVSW 1044
P+L+ ++ Q++F+ G I DNA LA E H + + A A+K+DL KAYD V W
Sbjct: 1167 PILDDLVSKEQSAFIQGRMITDNALLAFECFHSIQKNKKANSAACAYKLDLSKAYDRVDW 1226
Query: 1045 AFLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGSRLPSFKPGRGLRKGDPLSPYLFV 1103
FLE L + GF +R V IM CV+ S+ +NG+ L SF P RGLR+G+PLSP+LF+
Sbjct: 1227 RFLELALNKLGFAHRWVSWIMLCVTTVRYSVKFNGTLLRSFAPTRGLRQGEPLSPFLFL 1285
Score = 137 bits (346), Expect = 3e-30
Identities = 95/344 (27%), Positives = 151/344 (43%), Gaps = 23/344 (6%)
Query: 1231 KKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSGQ 1290
K++ W N L+ G+ L K+VI AIP Y M +F P S+ ++ + R F W ++G+
Sbjct: 1414 KRVIQWGENFLSSGGKEILIKAVIQAIPVYVMGLFKFPDSVYDELTKMTRNFWWGADNGR 1473
Query: 1291 RGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLAHKYLHGE 1350
R H W +LT+ K +GGL +D FN +LL + W L + L QVL KY
Sbjct: 1474 RRTHWRAWDSLTKAKINGGLGFRDYKLFNQALLTRQAWRLIEFPNSLCAQVLKAKYFPHG 1533
Query: 1351 SIFSVHRRANASPIWQGILKARDQLQEGFQFRMGDGSS-SVWYHDWTGKGKLAESLDFVH 1409
S+ ANASP W GI D L++G +R+G+G+S +W W + +
Sbjct: 1534 SLTDTTFSANASPTWHGIEYGLDLLKKGIIWRIGNGNSVRIWRDPWIPRDLSRRPVSSKA 1593
Query: 1410 ISDTKLCLHELVDNNQWNLNRCMTVFPLHIRDWFARVEPRVVAN-------GIDRWSWKD 1462
K + ++ W+ + I +F +++ ++ D +W
Sbjct: 1594 NCRLKWVSDLIAEDGTWDSAK--------INQYFLKIDADIIQKICISARLEEDFIAWHP 1645
Query: 1463 NSDGISSVHDGYQWLRE-------KKMAGVVHGDSWLWI*KLHAPEKL*VFIWLVLHDAL 1515
+ G SV Y+ + + SW I K + P+K+ +F W V ++L
Sbjct: 1646 DKTGRFSVRSAYKLALQLADMNNCSSSSSSRLNKSWELIWKCNVPQKVRIFAWRVASNSL 1705
Query: 1516 PTNANRYRCKLAVSPGCTRCSSPEEDTIHCLRDCPHSQELWGKL 1559
T N+ + L C C +ED H L C H+ LW L
Sbjct: 1706 ATMENKKKRNLERFDVCGICDREKEDAGHALCRCVHANSLWVNL 1749
Score = 50.1 bits (118), Expect = 7e-04
Identities = 40/134 (29%), Positives = 60/134 (43%), Gaps = 2/134 (1%)
Query: 1644 WKPPPPEFVKLNTDGSFQDTASSMGGGGLIRDAKG-AWLGGFMSHTSIGNPFLAEVLALR 1702
W+ P ++KLN DGSF + G G ++R+ G S S P AE+ A
Sbjct: 1777 WERPRNGWMKLNVDGSFDINSEKGGIGMILRNCLGNVIFSSCRSLDSCSGPLEAELHACV 1836
Query: 1703 DGLCLAWVQGHRRIIREVDCAGLSDILRFEDRCRLHDHAGVLQEIRELMERPWRCSVHWT 1762
+GL LA I E DC+ + +L D+ R A + QE + LM + ++
Sbjct: 1837 EGLHLALHWTLLPIQVETDCSSVIQLLNHPDKDR-SVLANIAQEAKSLMAGDRQIAISKV 1895
Query: 1763 NRDSNASADFLARR 1776
R N + FLA +
Sbjct: 1896 QRSQNVISHFLANK 1909
>emb|CAE04660.2| OSJNBa0061G20.16 [Oryza sativa (japonica cultivar-group)]
Length = 1285
Score = 333 bits (855), Expect = 2e-89
Identities = 203/629 (32%), Positives = 323/629 (51%), Gaps = 20/629 (3%)
Query: 486 IMFLLETRCQFSRASNFWNSTRFSPLFISEANGFSGGIWVLVNTATNLSSRLLYSHQQAV 545
++FL ETR + S F + G SGG+ + + + ++ + +
Sbjct: 578 LVFLCETRQSVEKMSRLRRKLAFRGFVGVSSEGMSGGLALYWDESVSVDVKDINKRYIDA 637
Query: 546 TFEVWRENLSWVCSAIYASPIPVKREELWSHLLHIRSNLVLPWMIVGDMNEILHPNE-VR 604
+ + W + +Y P R +WS L IR + LPWM++GD NE L E
Sbjct: 638 YVRLSSDEPQWHITFVYGEPRVENRHRMWSLLRTIRQSSALPWMVIGDFNETLWQFEHFS 697
Query: 605 GG*FLPTRASRFAAILEGCGMVDLGMSGGRYTWFRKQNNRLIMSKRLDRAMGDGAWRIAF 664
T+ F L C + DLG G +T+ +++ + RLDRA+ D WR F
Sbjct: 698 KNPRCETQMQNFRDALYDCDLQDLGFKGVPHTYDNRRDGWRNVKVRLDRAVADDKWRDLF 757
Query: 665 PEASVEALHRIHSDHCPLLINCSGPEIVH-HDRPFRFLASWVEHPQYKEVVQSAWTGETE 723
PEA V L SDH P+L+ + + + W P+ +V++ AW
Sbjct: 758 PEAQVSHLVSPCSDHSPILLEFIVKDTTRPRQKCLHYEIVWEREPESVQVIEEAWIN--- 814
Query: 724 TVAGKLNKVRTKSCKFNKEVYGSIFQRKRWVEGRLRGVQRELNYR-------VTFDMTRL 776
AG V+T N + + + W + +++ V REL + + R
Sbjct: 815 --AG----VKTDLGDINIALGRVMSALRSWGKTKVKNVGRELEKARKKLEDLIASNAARS 868
Query: 777 ETELQAEY-RFLLRQEELLWFQKARENRVCFDDRNTAYFHAHTIIRRKRNKIHRLKLNDG 835
++ +L +EE+LW Q++R N + DRNT +FH+ + R K+NKI +L+ +G
Sbjct: 869 SIRQATDHMNEMLYREEMLWLQRSRVNWLKEGDRNTRFFHSRAVWRAKKNKISKLRDENG 928
Query: 836 SWCDDRQLLAEEIQNYFQSLFAADPDNTDVNMQHVMMPRLTEAERHKAEAEVIKEEVFQA 895
+ +L YFQ ++ ADP ++ + ++T+A K E +EE+ QA
Sbjct: 929 AIHSMTSVLETMATEYFQEVYKADPSLNPESVTRLFQEKVTDAMNEKLCQEFKEEEIAQA 988
Query: 896 LLRMRSYSALGPDGFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVD 955
+ ++ + GPDGF F+++ W + D+ VR+ F++G++ + + +T IVLIPK D
Sbjct: 989 IFQIGPLKSPGPDGFPARFYQRNWGTLKSDIILAVRNFFQSGLMPEGVNDTAIVLIPKKD 1048
Query: 956 SPSTVKEFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEI 1015
P +K++RPISLCNVVYK+++K +VNR+RP+L+ ++ Q++F+ G I DNA LA E
Sbjct: 1049 QPIDLKDYRPISLCNVVYKVVSKCLVNRLRPILDDLVSKEQSAFIQGRMITDNALLAFEC 1108
Query: 1016 IHHMGAS-MAKKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNLS 1074
H + + A A+K+DL KAYD V W FLE L + GF +R V IM CV+ S
Sbjct: 1109 FHSIQKNKKANSAACAYKLDLSKAYDRVDWRFLELALNKLGFAHRWVSWIMSCVTTVRYS 1168
Query: 1075 MLWNGSRLPSFKPGRGLRKGDPLSPYLFV 1103
+ +NG+ L SF P RGLR+GDPLSP+LF+
Sbjct: 1169 VKFNGTLLRSFAPTRGLRQGDPLSPFLFL 1197
>gb|AAX95232.1| retrotransposon protein, putative, unclassified [Oryza sativa
(japonica cultivar-group)] gi|62733010|gb|AAX95129.1|
retrotransposon protein, putative, unclassified [Oryza
sativa (japonica cultivar-group)]
Length = 1324
Score = 331 bits (848), Expect = 2e-88
Identities = 228/686 (33%), Positives = 337/686 (48%), Gaps = 29/686 (4%)
Query: 439 GVLPLSQPISFMIGIEDLKIISWNVRGALNEHGQLFLKDLIRCKKPDIMFLLETRCQFSR 498
GV +SQ F I +KII+WN RG N L +L + PDI+FL T+ ++
Sbjct: 431 GVKQISQASCFCINKYPMKIIAWNCRGLGNGPTVRGLLNLQKEDDPDILFLSNTKMDTNK 490
Query: 499 ASNFWNSTRFSPLFISEANGFSGGIWVLVNTATNLSSRLLYSHQQAVTFEVWREN-LSWV 557
F L + + G SGG+ V NL RL + + +V ++ W
Sbjct: 491 LQCFRWKLGMPNLVVKDCQGKSGGLAVFWKKEINL--RLRTVSRLFMDVDVMEDDGFWWR 548
Query: 558 CSAIYASPIPVKREELWSHLLHIRSNLVLPWMIVGDMNEILHPNEV-RGG*FLPTRASRF 616
+ +Y P K++ W L + + W+ VGD NE+L E RG + RF
Sbjct: 549 FTGVYGEPRSDKKDLTWKALRTLNATKGELWLCVGDFNEVLFSWEKERGHAKAQSCMDRF 608
Query: 617 AAILEGCGMVDLGMSGGRYTWFRKQNNRLI----MSKRLDRAMGDGAWRIAFPEASVEAL 672
L+ C + +LG +G +TW +NN + + +RLDRA+ D W FP V
Sbjct: 609 RQTLDCCSLSNLGFTGDPFTW---RNNWCVRDGYIRERLDRAVADSDWCCRFPSFRVRKG 665
Query: 673 HRIHSDHCPLLINCSGPEIVHHDRP---FRFLASWVEHPQYKEVVQSAWTGETETVAGK- 728
HSDH P+++ S I + R FRF A W +V++AW GK
Sbjct: 666 DPRHSDHHPVIVTTSNEVIWNGGRSKPGFRFEAGWAREEHCAPIVENAWKLSVGPRGGKV 725
Query: 729 ---LNKVRTKSCKFNKEVYGSIFQRKRWVEGRLRGVQRELNYRVTFDMTRLETELQAEYR 785
+ +V ++K G + +R + + L E + R + E +YR
Sbjct: 726 MDAIREVAANLWDWSKNFLGDLEKRIKKAKQAL-----EAHRRSPISSSTASREAVLKYR 780
Query: 786 F--LLRQEELLWFQKARENRVCFDDRNTAYFHAHTIIRRKRNKIHRLKLNDGSWCDDRQL 843
L Q EL W Q+A ++ + DRNT +FH R+KRNKI +L+ +DG D
Sbjct: 781 LDKLEEQRELYWRQRANQHWLEKGDRNTKFFHECASERKKRNKIKKLRRDDGEVITDEAG 840
Query: 844 LAEEIQNYFQSLFAAD-PDNTDVNMQHVMMPRLTEAERHKAEAEVIKEEVFQALLRMRSY 902
I +++ LF A P N D +Q+V R+T + V EE+ +AL +
Sbjct: 841 SLSLISEFYKQLFTAGVPLNLDELLQNVPK-RVTSTMNDELMKGVTTEEIKKALDSIGDL 899
Query: 903 SAL-GPDGFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVK 961
A G DG F+K++WE VGDD+ V+D G + DS +T++VLIPKV +P +K
Sbjct: 900 KAPPGSDGMPALFYKQFWECVGDDIVHEVKDFLGGGEMPDSWNDTVVVLIPKVPNPERIK 959
Query: 962 EFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEIIHHMGA 1021
+ RPISLCNVVYK+ +KV+ NR++PLL+ II P+Q +F+P I DN LA E+ H +
Sbjct: 960 DLRPISLCNVVYKIASKVLANRLKPLLSEIISPIQIAFVPQRMITDNILLAYELTHFLKT 1019
Query: 1022 S-MAKKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGS 1080
G A K + KAYD V W FLE +++ GF V ++M+CV+ + NG
Sbjct: 1020 KRRGSVGFAAVKQGMSKAYDRVEWGFLEKMMLKLGFDRNWVFIVMKCVTSVTYRIKVNGE 1079
Query: 1081 RLPSFKPGRGLRKGDPLSPYLFVLCS 1106
P GLR+GDP+SPYLF +C+
Sbjct: 1080 FTEQINPTGGLRQGDPISPYLFFICA 1105
Score = 57.8 bits (138), Expect = 3e-06
Identities = 30/94 (31%), Positives = 46/94 (48%), Gaps = 5/94 (5%)
Query: 1231 KKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSGQ 1290
KK+ WK LL+ G+ L K+V AIP + M F L + + +I + RF W + +
Sbjct: 1231 KKIQGWKEKLLSRAGKDVLIKAVAQAIPAFAMSCFNLTKGLCDEITSLICRFFWAQQERE 1290
Query: 1291 RGWHRVNWKTLTQPKEHGGLAIK-----DMGDFN 1319
H + W+ L KE GG + D+ +FN
Sbjct: 1291 NKMHWIAWEHLCSRKEKGGFFFRRTTAGDLPEFN 1324
>gb|AAD21778.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25410938|pir||G84429 hypothetical protein
At2g01840 [imported] - Arabidopsis thaliana
Length = 1715
Score = 330 bits (846), Expect = 3e-88
Identities = 207/659 (31%), Positives = 339/659 (51%), Gaps = 19/659 (2%)
Query: 456 LKIISWNVRGALNEHGQLFLKDLIRCKKPDIMFLLETRCQFSRASNFWNSTRFSPLFISE 515
+++ WN +G L+++ R D++FL+ET+ Q + + F + I
Sbjct: 363 MRVGFWNCQGLGQPLTVRRLEEVQRVYFLDMLFLIETKQQDNYTRDLGVKMGFEDMCIIS 422
Query: 516 ANGFSGGIWVLVNTATNLSSRLLYSHQQAVTFEVWRENLSWVCSAIYASPIPVKREELWS 575
G SGG+ +V +LS +++ + V V +N ++ S IY PIP +R LW
Sbjct: 423 PRGLSGGL--VVYWKKHLSIQVISHDVRLVDLYVEYKNFNFYLSCIYGHPIPSERHHLWE 480
Query: 576 HLLHIRSNLVLPWMIVGDMNEILHPNEVRGG*FLPTRA-SRFAAILEGCGMVDLGMSGGR 634
L + ++ PWM+ GD NEIL+ NE +GG + F ++ C M DL G
Sbjct: 481 KLQRVSAHRSGPWMMCGDFNEILNLNEKKGGRRRSIGSLQNFTNMINCCNMKDLKSKGNP 540
Query: 635 YTWFRKQNNRLIMSKRLDRAMGDGAWRIAFPEASVEALHRIHSDHCPLLINCSGPEIVHH 694
Y+W K+ N I S LDR + W+ +FP E L SDH P++I+ + E+
Sbjct: 541 YSWVGKRQNETIESC-LDRVFINSDWQASFPAFETEFLPIAGSDHAPVIIDIA-EEVCTK 598
Query: 695 DRPFRFLASWVEHPQYKEVVQSAWT-GETETVAG---KLNKVRTKSCKFNKEVYGSIFQR 750
FR+ + + + VQ W G +++ G KL+ R + K+ + + ++
Sbjct: 599 RGQFRYDRRHFQFEDFVDSVQRGWNRGRSDSHGGYYEKLHCCRQELAKWKRRTKTNTAEK 658
Query: 751 KRWVEGRLRGVQRE--LNYRVTFDMTRLETELQAEYRFLLRQEELLWFQKARENRVCFDD 808
++ R+ +R+ L ++ + RL +L YR EEL W K+R + D
Sbjct: 659 IETLKYRVDAAERDHTLPHQT---ILRLRQDLNQAYR----DEELYWHLKSRNRWMLLGD 711
Query: 809 RNTAYFHAHTIIRRKRNKIHRLKLNDGSWCDDRQLLAEEIQNYFQSLFAADPDNTDVNMQ 868
RNT +F+A T +R+ RN+I + G + + +NYF LF + +
Sbjct: 712 RNTMFFYASTKLRKSRNRIKAITDAQGIENFRDDTIGKVAENYFADLFTTTQTSDWEEII 771
Query: 869 HVMMPRLTEAERHKAEAEVIKEEVFQALLRMRSYSALGPDGFHPFFFKKYWELVGDDLWK 928
+ P++TE H+ V +EV A+ + + A G DGF F+ W+L+G+D+
Sbjct: 772 SGIAPKVTEQMNHELLQSVTDQEVRDAVFAIGADRAPGFDGFTAAFYHHLWDLIGNDVCL 831
Query: 929 LVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKEFRPISLCNVVYKLITKVVVNRIRPLL 988
+VR FE+ ++D+ + +T I LIPK+ P + ++RPISLC YK+I+K+++ R++ L
Sbjct: 832 MVRHFFESDVMDNQINQTQICLIPKIIDPKHMSDYRPISLCTASYKIISKILIKRLKQCL 891
Query: 989 NRIIGPMQNSFLPGMGIMDNAFLAQEIIHHMGASM-AKKGTLAFKIDLEKAYDSVSWAFL 1047
+I Q +F+PG I DN +A E++H + + + G +A K D+ KAYD V W FL
Sbjct: 892 GDVISDSQAAFVPGQNISDNVLVAHELLHSLKSRRECQSGYVAVKTDISKAYDRVEWNFL 951
Query: 1048 EDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGSRLPSFKPGRGLRKGDPLSPYLFVLCS 1106
E ++Q GF R V+ IM CV+ + +L NGS P RG+R+GDPLSPYLF+ C+
Sbjct: 952 EKVMIQLGFAPRWVKWIMTCVTSVSYEVLINGSPYGKIFPSRGIRQGDPLSPYLFLFCA 1010
Score = 146 bits (369), Expect = 6e-33
Identities = 146/610 (23%), Positives = 244/610 (39%), Gaps = 77/610 (12%)
Query: 1216 GFQEVDSITYWII-FKKKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHK 1274
G ++V+ Y + K++ W N L+ G+ + K++ A+P Y+M F LP I ++
Sbjct: 1120 GRRKVELFEYIVTKVKERTEGWAYNYLSPAGKEIVIKAIAMALPVYSMNCFLLPTLICNE 1179
Query: 1275 INQEMRRFIWLKNSGQRGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNS 1334
IN + F W K + G L KD+ FN +LL K W + N
Sbjct: 1180 INSLITAFWWGKEN------------------EGDLGFKDLHQFNRALLAKQAWRILTNP 1221
Query: 1335 SKLWVQVLAHKYLHGESIFSVHRRANASPIWQGILKARDQLQEGFQFRMGDGSSS-VWYH 1393
L ++ Y + ++ +AS W I + + LQ+G + R+GDG ++ +W
Sbjct: 1222 QSLLARLYKGLYYPNTTYLRANKGGHASYGWNSIQEGKLLLQQGLRVRLGDGQTTKIWED 1281
Query: 1394 DWTGKGKLAESLDFVHISDTKLCLHELVDNNQW---------NLNRCMTVFPLHIRDWFA 1444
W + + D K+ + +W N L++ ++ A
Sbjct: 1282 PWLPTLPPRPARGPILDEDMKVADLWRENKREWDPVIFEGVLNPEDQQLAKSLYLSNYAA 1341
Query: 1445 RVEPRVVANGIDRWSWKDNSDGISSVHDGYQW------LREKKMAGVVHGDSWLW--I*K 1496
R D + W + +V GY W L E+++ + GD L I +
Sbjct: 1342 R----------DSYKWAYTRNTQYTVRSGY-WVATHVNLTEEEIINPLEGDVPLKQEIWR 1390
Query: 1497 LHAPEKL*VFIWLVLHDALPTNANRYRCKLAVSPGCTRCSSPEEDTIHCLRDCPHSQELW 1556
L K+ FIW L AL T + P C RC + +E H + C ++Q +W
Sbjct: 1391 LKITPKIKHFIWRCLSGALSTTTQLRNRNIPADPTCQRCCNADETINHIIFTCSYAQVVW 1450
Query: 1557 GKLGAWGWSTFRLPD-LKA*VKTQLQG--SNNIRFLAGL------WGVWKWRNNVV---L 1604
G + D L+ ++ LQG + N+ L GL W +WK RN + L
Sbjct: 1451 RSANFSGSNRLCFTDNLEENIRLILQGKKNQNLPILNGLMPFWIMWRLWKSRNEYLFQQL 1510
Query: 1605 DPLPWTLDGAWQKLRHDHDELLQFSSGDLLDEHHFL---------INLWKPPPPEFVKLN 1655
D PW + QK + E ++ D H+ W PP F+K N
Sbjct: 1511 DRFPWKVA---QKAEQEATEWVETMVNDTAISHNTAQSNDRPLSRSKQWSSPPEGFLKCN 1567
Query: 1656 TDGSFQDTASSMGGGGLIRDAKGAWL-GGFMSHTSIGNPFLAEVLALRDGLCLAWVQGHR 1714
D + G ++RD G L G + AE L L + W++G+
Sbjct: 1568 FDSGYVQGRDYTSTGWILRDCNGRVLHSGCAKLQQSYSALQAEALGFLHALQMVWIRGYC 1627
Query: 1715 RIIREVDCAGLSDIL-RFEDRCRLHDHAGVLQEIRELMERPWRCSVHWTNRDSNASADFL 1773
+ E D L++++ + ED H +L +IR M + S+ + NR+ N +AD L
Sbjct: 1628 YVWFEGDNLELTNLINKTEDH---HLLETLLYDIRFWMTKLPFSSIGYVNRERNLAADKL 1684
Query: 1774 ARRGGFLSTI 1783
+ +S++
Sbjct: 1685 TKYANSMSSL 1694
>gb|AAQ19327.1| bZIP-like protein [Oryza sativa (japonica cultivar-group)]
Length = 2367
Score = 314 bits (804), Expect = 2e-83
Identities = 190/604 (31%), Positives = 308/604 (50%), Gaps = 30/604 (4%)
Query: 516 ANGFSGGIWVLVNTATNLSSRLLYSHQQAVTFEVWRENLSWVCSAIYASPIPVKREELWS 575
+ G SGG+ + + + ++ + + ++ E W + +Y P R +WS
Sbjct: 790 SEGMSGGLALYWDESVSVDVKDINKRYIDAYVQLSPEEPQWHVTFVYGEPRVENRHRMWS 849
Query: 576 HLLHIRSNLVLPWMIVGDMNEIL----HPNEVRGG*FLPTRASRFAAILEGCGMVDLGMS 631
L I + LPW ++GD NE + H + G + F +L+ C + DLG
Sbjct: 850 LLRTIHQSSSLPWAVIGDFNETMWQFEHFSRTPRG---EPQMQDFRDVLQDCELHDLGFK 906
Query: 632 GGRYTWFRKQNNRLIMSKRLDRAMGDGAWRIAFPEASVEALHRIHSDHCPLLINC--SGP 689
G +T+ K+ + RLDR + D WR + A V L SDHCP+L+N P
Sbjct: 907 GVPHTYDNKREGWRNVKVRLDRVVADDKWRDIYSTAQVVHLVSPCSDHCPILLNLVVKDP 966
Query: 690 EIVHHDRPFRFLASWVEHPQYKEVVQSAWTGETETVAGKLNKVRTKSCKFNKEVYGSIFQ 749
+ + + W P+ +V++ AW VAG+ + NK + +
Sbjct: 967 HQLRQ-KCLHYEIVWEREPEATQVIEEAWV-----VAGE----KADLGDINKALAKVMTA 1016
Query: 750 RKRWVEGRLRGVQRELN---------YRVTFDMTRLETELQAEYRFLLRQEELLWFQKAR 800
+ W +++ V REL D T + LL +EE+LW Q++R
Sbjct: 1017 LRSWSRAKVKNVGRELEKARKKLAELIESNADRTVIRNATD-HMNELLYREEMLWLQRSR 1075
Query: 801 ENRVCFDDRNTAYFHAHTIIRRKRNKIHRLKLNDGSWCDDRQLLAEEIQNYFQSLFAADP 860
N + +DRNT +FH+ + R K+NKI +L+ + + L YFQ ++ ADP
Sbjct: 1076 VNWLKDEDRNTKFFHSRAVWRAKKNKISKLRDANETVHSSTMKLESMATEYFQDVYTADP 1135
Query: 861 DNTDVNMQHVMMPRLTEAERHKAEAEVIKEEVFQALLRMRSYSALGPDGFHPFFFKKYWE 920
+ + ++ ++T+ K + ++E+ QA+ ++ + GPDGF F+++ W
Sbjct: 1136 NLNPETVTRLIQEKVTDIMNEKLCEDFTEDEISQAIFQIGPLKSPGPDGFPARFYQRNWG 1195
Query: 921 LVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKEFRPISLCNVVYKLITKVV 980
+ D+ VR F+TG++ + + +T IVLIPK + P +++FRPISLCNVVYK+++K +
Sbjct: 1196 TIKADIIGAVRRFFQTGLMPEGVNDTAIVLIPKKEQPVDLRDFRPISLCNVVYKVVSKCL 1255
Query: 981 VNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEIIHHMGAS-MAKKGTLAFKIDLEKAY 1039
VNR+RP+L+ ++ Q++F+ G I DNA LA E H M + A A+K+DL KAY
Sbjct: 1256 VNRLRPILDDLVSVEQSAFVQGRMITDNALLAFECFHAMQKNKKANHAACAYKLDLSKAY 1315
Query: 1040 DSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGSRLPSFKPGRGLRKGDPLSP 1099
D V W FLE + + GF R V IM+CV+ + +NG+ L SF P RGLR+GDPL P
Sbjct: 1316 DRVDWRFLEMAMNKLGFARRWVNWIMKCVTSVRYMVKFNGTLLQSFAPTRGLRQGDPLLP 1375
Query: 1100 YLFV 1103
+LF+
Sbjct: 1376 FLFL 1379
Score = 174 bits (440), Expect = 3e-41
Identities = 151/581 (25%), Positives = 246/581 (41%), Gaps = 61/581 (10%)
Query: 1231 KKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSGQ 1290
K++ W N L+ G+ L K+VI AIP Y M +F LP S+ + + + F W +GQ
Sbjct: 1508 KRVIQWGENHLSTGGKEVLIKAVIQAIPVYVMGIFKLPESVIDDLTKLTKNFWWDSMNGQ 1567
Query: 1291 RGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLAHKYLHGE 1350
R H W +LT+PK GGL +D FN +LL + W L L +VL KY
Sbjct: 1568 RKTHWKAWDSLTKPKSLGGLGFRDYRLFNQALLARQAWRLITYPDSLCARVLKAKYFPHG 1627
Query: 1351 SIFSVHRRANASPIWQGILKARDQLQEGFQFRMGDGSS-SVWYHDWTGKGKLAESLDFVH 1409
S+ +N+SP W+ I D L++G +R+G+G+S +W W + +
Sbjct: 1628 SLIDTSFGSNSSPAWRSIEYGLDLLKKGIIWRVGNGNSIRIWRDSWLPRDHSRRPITGKA 1687
Query: 1410 ISDTKLCLHELVDNNQWNLNRCMTVFPLHIRDWFARVEPRVVAN-------GIDRWSWKD 1462
K + ++ W++ + I +F ++ V+ N D +W
Sbjct: 1688 NCRLKWVSDLITEDGSWDVPK--------IHQYFHNLDAEVILNICISSRSEEDFIAWHP 1739
Query: 1463 NSDGISSVHDGYQW------LREKKMAGVVH-GDSWLWI*KLHAPEKL*VFIWLVLHDAL 1515
+ +G+ SV Y+ + E +G + +W I K P+K+ +F W V + L
Sbjct: 1740 DKNGMFSVRSAYRLAAQLVNIEESSSSGTNNINKAWEMIWKCKVPQKVKIFAWRVASNCL 1799
Query: 1516 PTNANRYRCKLAVSPGCTRCSSPEEDTIHCLRDCPHSQELWGKLGAWGWSTFRLPDLKA* 1575
T N+ + KL S C C ED H L C + +LW + G + D+KA
Sbjct: 1800 ATMVNKKKRKLEQSDMCQICDRENEDDAHALCRCIQASQLWSCMHKSGSVSV---DIKAS 1856
Query: 1576 V---------KTQLQGSNNIRFLAGLWGVWKWRNNVVLDPLPWTLDGAWQKLRHDHDELL 1626
V ++ FL LW W RN ++ + + + ++ D L
Sbjct: 1857 VLGRFWLFDCLEKIPEYEQAMFLMTLWRNWYVRNELIHGKSAPPTETSQRFIQSYVDLLF 1916
Query: 1627 QF---SSGDLLDEHHFLINL-----------------WKPPPPEFVKLNTDGSFQDTASS 1666
Q DL+ H + + W+ P ++KLN DGSF ++
Sbjct: 1917 QIRQAPQADLVKGKHVVRTVPLKGGPKYRVLNNHQPCWERPKDGWMKLNVDGSFDASSGK 1976
Query: 1667 MGGGGLIRDAKGAWLGGFMSHTSI---GNPFLAEVLALRDGLCLAWVQGHRRIIREVDCA 1723
G G ++R++ G + F S + NP +E+ A +GL LA I E DCA
Sbjct: 1977 GGLGMILRNSAGDII--FTSCKPLERCNNPLESELRACVEGLKLAIHWTLLPIQVETDCA 2034
Query: 1724 GLSDILRFEDRCRLHDHAGVLQEIRELMERPWRCSVHWTNR 1764
+ +L+ R A ++ E R L++ P R + T +
Sbjct: 2035 SVVQLLQGIGR-DFSVLANIIHEARHLLQTPSRTNTKVTTQ 2074
>gb|AAD20714.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25412331|pir||G84649 hypothetical protein
At2g25550 [imported] - Arabidopsis thaliana
Length = 1750
Score = 312 bits (799), Expect = 8e-83
Identities = 202/644 (31%), Positives = 323/644 (49%), Gaps = 34/644 (5%)
Query: 478 LIRCKKPDIMFLLETRCQFSRASNFWNSTRFSPLFISEANGFSGGIWVLVNTATNLSSRL 537
L R DI+FL+ET Q + F + NG SGG+ ++ N+S L
Sbjct: 404 LFRMYNYDILFLVETLNQCDKVCKLAYDLGFPNVITQPPNGRSGGLALMWKN--NVSLSL 461
Query: 538 LYSHQQAVTFEVWRENLSWVCSAIYASPIPVKREELWSHLLHIRSNLVLPWMIVGDMNEI 597
+ ++ + V N S+ S +Y P +R +LW L HI N W++VGD NEI
Sbjct: 462 ISQDERLIDSHVTFNNKSFYLSCVYGHPTQSERHQLWQTLEHISDNRNAEWLLVGDFNEI 521
Query: 598 LHPNEVRGG*FLPTRAS----RFAAILEGCGMVDLGMSGGRYTWFRKQNNRLIMSKRLDR 653
L E GG P R F ++ C + D+ G R++W +++ + LDR
Sbjct: 522 LSNAEKIGG---PMREEWTFRNFRNMVSHCDIEDMRSKGDRFSWVGERHTHTVKCC-LDR 577
Query: 654 AMGDGAWRIAFPEASVEALHRIHSDHCPLLINCSGPEIVHHDRPFRFLASWVEHPQYKEV 713
+ AW FP A +E L SDH P+L++ + + FRF ++ P +K +
Sbjct: 578 VFINSAWTATFPYAEIEFLDFTGSDHKPVLVHFN-ESFPRRSKLFRFDNRLIDIPTFKRI 636
Query: 714 VQSAWTGETETVAGKLNKVRTKSCK--FNKEVYGSIFQRKRWVEGRLRGVQRELNYRVTF 771
VQ++W + + + + R SC+ + + S E R++ +Q LN +
Sbjct: 637 VQTSWRTNRNSRSTPITE-RISSCRQAMARLKHASNLNS----EQRIKKLQSSLNRAM-- 689
Query: 772 DMTR-----LETELQAEYRFLLRQEELLWFQKARENRVCFDDRNTAYFHAHTIIRRKRNK 826
+ TR L +LQ EE+ W QK+R + D+NT YFHA T R +N+
Sbjct: 690 ESTRRVDRQLIPQLQESLAKAFSDEEIYWKQKSRNQWMKEGDQNTGYFHACTKTRYSQNR 749
Query: 827 IHRLKLNDGSWCDDRQLLAEEIQNYFQSLFAADPDNTDVNMQHVMMPRLTEAERHKAEAE 886
++ + + G + + Q++F ++F+ + + + + + +
Sbjct: 750 VNTIMDDQGRMFTGDKEIGNHAQDFFTNIFSTN----GIKVSPIDFADFKSTVTNTVNLD 805
Query: 887 VIKE----EVFQALLRMRSYSALGPDGFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDS 942
+ KE E++ A+ ++ A GPDG F+K W++VG D+ V+ FET + S
Sbjct: 806 LTKEFSDTEIYDAICQIGDDKAPGPDGLTARFYKNCWDIVGYDVILEVKKFFETSFMKPS 865
Query: 943 LLETLIVLIPKVDSPSTVKEFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPG 1002
+ T I +IPK+ +P+T+ ++RPI+LCNV+YK+I+K +VNR++ LN I+ Q +F+PG
Sbjct: 866 INHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPG 925
Query: 1003 MGIMDNAFLAQEIIHHMGA-SMAKKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLV 1061
I DN +A E++H + K +A K D+ KAYD V W FLE T+ FGF N+ +
Sbjct: 926 RIINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWI 985
Query: 1062 QLIMRCVSGSNLSMLWNGSRLPSFKPGRGLRKGDPLSPYLFVLC 1105
IM V + S+L NGS P RG+R+GDPLSPYLF+LC
Sbjct: 986 GWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILC 1029
Score = 158 bits (399), Expect = 2e-36
Identities = 152/609 (24%), Positives = 234/609 (37%), Gaps = 99/609 (16%)
Query: 1230 KKKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSG 1289
KK+ S+W L+ G+ + KSV A+P Y M F LP+ I +I + F W K S
Sbjct: 1155 KKRTSTWSARFLSPAGKEIMLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASN 1214
Query: 1290 QRGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLAHKYLHG 1349
QRG V WK L K+ GGL +D+ FN +LL K W L + L+ +V+ +Y
Sbjct: 1215 QRGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKD 1274
Query: 1350 ESIFSVHRRANASPIWQGILKARDQLQEGFQFRMGDG-------------------SSSV 1390
SI R S W +L L++G + +GDG ++
Sbjct: 1275 VSILDAKVRKQQSYGWASLLDGIALLKKGTRHLIGDGQNIRIGLDNIVDSHPPRPLNTEE 1334
Query: 1391 WYHDWTGKGKLAESLDFVHISDTKLCLHELVDNNQWNLNRCMTVFPLHIRDWFARVEPRV 1450
Y + T + D+K + + VD + +H R + A+
Sbjct: 1335 TYKEMTINNLFERKGSYYFWDDSK--ISQFVDQSDHGF--------IH-RIYLAK----- 1378
Query: 1451 VANGIDRWSWKDNSDGISSVHDGYQWLREKKMAGV-----VHG--DSWLWI*KLHAPEKL 1503
+ D+ W N+ G +V GY L + HG D I L KL
Sbjct: 1379 -SKKPDKIIWNYNTTGEYTVRSGYWLLTHDPSTNIPAINPPHGSIDLKTRIWNLPIMPKL 1437
Query: 1504 *VFIWLVLHDALPTNANRYRCKLAVSPGCTRCSSPEEDTIHCLRDCPHSQELWGKLGAWG 1563
F+W L AL T + + P C RC E H L CP + W
Sbjct: 1438 KHFLWRALSQALATTERLTTRGMRIDPSCPRCHRENESINHALFTCPFATMAW------- 1490
Query: 1564 WSTFRLPDLKA*VKTQLQG-------SNNIRFLAG--------------LWGVWKWRNNV 1602
RL D + ++ QL SN + F+ +W +WK RNNV
Sbjct: 1491 ----RLSD-SSLIRNQLMSNDFEENISNILNFVQDTTMSDFHKLLPVWLIWRIWKARNNV 1545
Query: 1603 VLDPL---PWTLDGAWQKLRHDHDELLQFSSGDLLDEHHFLINL--WKPPPPEFVKLNTD 1657
V + P + + HD Q N W+ PP +VK N D
Sbjct: 1546 VFNKFRESPSKTVLSAKAETHDWLNATQSHKKTPSPTRQIAENKIEWRNPPATYVKCNFD 1605
Query: 1658 GSFQDTASSMGGGGLIRDAKG---AWLGGFMSHTSIGNPFLAEVLALRDGLCLAWVQGHR 1714
F GG +IR+ G +W ++HTS NP AE AL L W++G+
Sbjct: 1606 AGFDVQKLEATGGWIIRNHYGTPISWGSMKLAHTS--NPLEAETKALLAALQQTWIRGYT 1663
Query: 1715 RIIREVDCAGLSDILRFEDRCRLHDHAGVLQEIRELMERPWRCSVHWTNRDSNASADFLA 1774
++ E DC L +++ + H+ + + ++ W N+ ++ F+
Sbjct: 1664 QVFMEGDCQTLINLIN-----GISFHSSLANHLEDIS--------FWANKFASIQFGFIR 1710
Query: 1775 RRGGFLSTI 1783
++G L+ +
Sbjct: 1711 KKGNKLAHV 1719
>emb|CAA66812.1| non-ltr retrotransposon reverse transcriptase-like protein
[Arabidopsis thaliana]
Length = 893
Score = 305 bits (782), Expect = 7e-81
Identities = 257/926 (27%), Positives = 444/926 (47%), Gaps = 82/926 (8%)
Query: 457 KIISWNVRGA-LNEHGQLFLKDLIRCKKPDIMFLLETRCQFSRASNFWNSTRFSPLFISE 515
K+ WNVRG ++ H + F K + KP L+ET + + F ++ F+ E
Sbjct: 4 KLFCWNVRGFNISSHRRGFKKWFL-LNKPLFGGLIETHVKQPKEKKFISNLLPGWSFV-E 61
Query: 516 ANGFS--GGIWVLVNTATNLSSRLLYSHQQAVTFEVWR-ENLSW-VCSAIYASPIPVKRE 571
FS G IWVL + + + ++ Q +T E+ ++ SW V S +YAS R+
Sbjct: 62 NYEFSVLGKIWVLWDPSVKVV--VIGRSLQMITCELLLPDSPSWFVVSIVYASNEEGTRK 119
Query: 572 ELWSHLLHIRSNLVL---PWMIVGDMNEILHPNEVRGG*FLPTRASRFAAILEGCGMVDL 628
ELW+ L+ + + V+ W+++GD N+IL+P + + F + L + DL
Sbjct: 120 ELWNELVQLALSPVVVGRSWIVLGDFNQILNPESAINA-NIGRKIRAFRSCLLDSDLYDL 178
Query: 629 GMSGGRYTWFRKQNNRLIMSKRLDRAMGDGAWRIAFPEASVEALHRIHSDH--CPLLINC 686
G YTW+ K ++R + +K++DR + + W FP A SDH C ++++
Sbjct: 179 VYKGSSYTWWNKCSSRPL-AKKIDRILVNDHWNTLFPSAYANFGEPDFSDHSSCEVVLD- 236
Query: 687 SGPEIVHHDRPFRFLASWVEHPQYKEVVQSAWTGETET------VAGKLNKVRTKSCKFN 740
P ++ RPFRF ++ +P + ++++ W + V+ KL ++ C F+
Sbjct: 237 --PAVLKAKRPFRFFNYFLHNPDFLQLIRENWYSCNVSGSAMYRVSKKLKHLKLPICCFS 294
Query: 741 KEVYGSIFQRKRWVEGRLRGVQRELNYRVTFDMTRLETELQAEYRFLLRQEELLWFQKAR 800
+E Y I +R + QR + LE E +++ L + EE + QK+
Sbjct: 295 RENYSDIEKRVSEAHAIVLHRQRITLTNPSVVHATLELEATRKWQILAKAEESFFCQKSS 354
Query: 801 ENRVCFDDRNTAYFHAHTIIRRKRNKIHRLKLNDGSWCDDRQLLAEEIQ----NYFQSLF 856
+ + D NTAYFH +R+ N I+ L + G + +Q + E I+ N+F+SL
Sbjct: 355 ISWLYEGDNNTAYFHKMADMRKSINTINFLIDDFGERIETQQGIKEGIKEHSCNFFESLL 414
Query: 857 AA-DPDNT----DVNMQ---HVMMPRLTEAERHKAEAEVIKEEVFQALLRMRSYSALGPD 908
+ +N+ D+N+ + ++ + ER ++ ++ +E F +L R + A GPD
Sbjct: 415 CGVEGENSLAQSDMNLLLSFRCSVDQINDLERSFSDLDI--QEAFFSLPRNK---ASGPD 469
Query: 909 GFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKEFRPISL 968
G+ FFK W +VG ++ + V++ F +G + T +VLIPK+ + S + +FRPIS
Sbjct: 470 GYSSEFFKGVWFVVGPEVTEAVQEFFRSGQLLKQWNATTLVLIPKITNSSKMTDFRPISC 529
Query: 969 CNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEIIH-HMGASMAKKG 1027
N +YK+I K++ +R++ LLN +I P Q++FLPG + +N LA EI+H + +++ +G
Sbjct: 530 LNTLYKVIAKLLTSRLKKLLNEVISPSQSAFLPGRLLSENVLLATEIVHGYNTKNISSRG 589
Query: 1028 TLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGSRLPSFKP 1087
L K+DL KA+DSV W F+ P + V I +C+S S++ NGS FK
Sbjct: 590 ML--KVDLRKAFDSVRWDFIISAFRALAVPEKFVCWINQCISTPYFSVMVNGSSSGFFKS 647
Query: 1088 GRGLRKGDPLSPYLFVLCSLHGAIVY*YS*VGGNRSLEAN*NFPKGTTYFASLFCR*RAL 1147
+GLR+GDPLSPYLFVL V+ + L+A F G + + L
Sbjct: 648 NKGLRQGDPLSPYLFVL----AMEVF-------SSLLKA--RFDAG---YIQYHPKTADL 691
Query: 1148 VLSSIIFSSAI----GGGDS*TILCLFGVKDKYEQI*S-----------DCFQGSTKSCE 1192
+S ++F+ + GG S L G+ + + S + + T E
Sbjct: 692 SISHLMFADDVMVFFDGGSS----SLHGISEALDDFASWSGLHVNKDKTNLYLAGTDEVE 747
Query: 1193 RGDPEYCPYPFCAKPWQVSWLEEGFQEVDSITYWIIFKKKLSSWKTNLLNMVGQVCLAKS 1252
+ +P P + L +++ Y ++ K+ SW L+ G+V L S
Sbjct: 748 ALAISHYGFPISTLPIRYLGLPLMSRKLKISEYELV--KRFRSWAVKSLSFAGRVQLITS 805
Query: 1253 VISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSGQRGWHRVNWKTLTQPKEHGGLAI 1312
VI+ + + M F L KI RF+W + ++ W + PK GG+A+
Sbjct: 806 VITGLVNFWMSTFVLLLGCVKKIESLCSRFLWSGSIDASKGAKIAWSGVCLPKNEGGVAL 865
Query: 1313 KDMGDFNTSLLGKVVWLLANNSSKLW 1338
+ +N + + +W L ++ LW
Sbjct: 866 RRFTPWNKTFYLRFIWPLFADNDVLW 891
>emb|CAB81184.1| putative protein [Arabidopsis thaliana] gi|4539357|emb|CAB40051.1|
putative protein [Arabidopsis thaliana]
gi|7486147|pir||T04278 hypothetical protein F25I24.40 -
Arabidopsis thaliana
Length = 1294
Score = 305 bits (781), Expect = 9e-81
Identities = 195/641 (30%), Positives = 317/641 (49%), Gaps = 20/641 (3%)
Query: 475 LKDLIRCKKPDIMFLLETRCQFSRASNFWNSTRFSPLFISEANGFSGGIWVLVNTATNLS 534
L +L + K D++FL+ET + SN + F + G SGG+ +L + LS
Sbjct: 381 LSNLCKVFKFDVLFLIETLNKCEVISNLASVLGFPNVITQPPQGHSGGLALLWKDSVRLS 440
Query: 535 SRLLYSHQQAVTFEVWRENLSWVCSAIYASPIPVKREELWSHLLHIRSNLVLPWMIVGDM 594
+ LY + + + N+++ S +Y P +R LW+H ++ PW+++GD
Sbjct: 441 N--LYQDDRHIDVHISINNINFYLSRVYGHPCQSERHSLWTHFENLSKTRNDPWILIGDF 498
Query: 595 NEILHPNEVRGG*FLPTRAS----RFAAILEGCGMVDLGMSGGRYTWFRKQNNRLIMSKR 650
NEIL NE GG P R F ++ C + D+ G R++W ++++ +
Sbjct: 499 NEILSNNEKIGG---PQRDEWTFRGFRNMVSTCDLKDIRSIGDRFSWVGERHSHTVKCC- 554
Query: 651 LDRAMGDGAWRIAFPEASVEALHRIHSDHCPLLINCSGPEIVHHDRPFRFLASWVEHPQY 710
LDRA + FP A +E L SDH PL ++ E RPFRF +E P +
Sbjct: 555 LDRAFINSEGAFLFPFAELEFLEFTGSDHKPLFLSLEKTE-TRKMRPFRFDKRLLEVPHF 613
Query: 711 KEVVQSAWT----GETETVAGKLNKVRTKSCKFNKEVYGSIFQRKRWVEGRLRGVQRELN 766
K V++ W G+ + + ++ R K + + R ++ L +N
Sbjct: 614 KTYVKAGWNKAINGQRKHLPDQVRTCRQAMAKLKHKSNLNSRIRINQLQAALDKAMSSVN 673
Query: 767 YRVTFDMTRLETELQAEYRFLLRQEELLWFQKARENRVCFDDRNTAYFHAHTIIRRKRNK 826
++ ++ EL YR EE W QK+R + DRNT +FHA T R N+
Sbjct: 674 RTERRTISHIQRELTVAYR----DEERYWQQKSRNQWMKEGDRNTEFFHACTKTRFSVNR 729
Query: 827 IHRLKLNDGSWCDDRQLLAEEIQNYFQSLFAADPDNTDVNMQHVMMPRLTEAERHKAEAE 886
+ +K +G + + Q +F ++ ++ + P +TE +
Sbjct: 730 LVTIKDEEGMIYRGDKEIGVHAQEFFTKVYESNGRPVSIIDFAGFKPIVTEQINDDLTKD 789
Query: 887 VIKEEVFQALLRMRSYSALGPDGFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLET 946
+ E++ A+ + A GPDG F+K WE+VG D+ K V+ F T + S+ T
Sbjct: 790 LSDLEIYNAICHIGDDKAPGPDGLTARFYKSCWEIVGPDVIKEVKIFFRTSYMKQSINHT 849
Query: 947 LIVLIPKVDSPSTVKEFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIM 1006
I +IPK+ +P T+ ++RPI+LCNV+YK+I+K +V R++ L+ I+ Q +F+PG +
Sbjct: 850 NICMIPKITNPETLSDYRPIALCNVLYKIISKCLVERLKGHLDAIVSDSQAAFIPGRLVN 909
Query: 1007 DNAFLAQEIIHHMGA-SMAKKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIM 1065
DN +A E++H + + +A K D+ KAYD V W FLE T+ FGF ++ IM
Sbjct: 910 DNVMIAHEMMHSLKTRKRVSQSYMAVKTDVSKAYDRVEWNFLETTMRLFGFSETWIKWIM 969
Query: 1066 RCVSGSNLSMLWNGSRLPSFKPGRGLRKGDPLSPYLFVLCS 1106
V N S+L NG + +P RG+R+GDPLSPYLF+LC+
Sbjct: 970 GAVKSVNYSVLVNGIPHGTIQPQRGIRQGDPLSPYLFILCA 1010
Score = 107 bits (268), Expect = 3e-21
Identities = 54/159 (33%), Positives = 86/159 (53%)
Query: 1230 KKKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSG 1289
KK+ SSW L+ G+ + KSV ++P Y M F LP +I +I + F W KN+
Sbjct: 1135 KKRTSSWSAKYLSPAGKEIMLKSVAMSMPVYAMSCFKLPLNIVSEIEALLMNFWWEKNAK 1194
Query: 1290 QRGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLAHKYLHG 1349
+R + WK L K+ GGL +D+ FN +LL K VW + NN + L+ +++ +Y
Sbjct: 1195 KREIPWIAWKRLQYSKKEGGLGFRDLAKFNDALLAKQVWRMINNPNSLFARIMKARYFRE 1254
Query: 1350 ESIFSVHRRANASPIWQGILKARDQLQEGFQFRMGDGSS 1388
+SI R+ S W +L D +++G +F +GDG +
Sbjct: 1255 DSILDAKRQRYQSYGWTSMLAGLDVIKKGSRFIVGDGKT 1293
>gb|AAC33961.1| contains similarity to reverse trancriptase (Pfam: rvt.hmm, score:
42.57) [Arabidopsis thaliana] gi|7486711|pir||T01893
hypothetical protein F8M12.22 - Arabidopsis thaliana
Length = 1662
Score = 305 bits (781), Expect = 9e-81
Identities = 195/641 (30%), Positives = 317/641 (49%), Gaps = 20/641 (3%)
Query: 475 LKDLIRCKKPDIMFLLETRCQFSRASNFWNSTRFSPLFISEANGFSGGIWVLVNTATNLS 534
L +L + K D++FL+ET + SN + F + G SGG+ +L + LS
Sbjct: 401 LSNLCKVFKFDVLFLIETLNKCEVISNLASVLGFPNVITQPPQGHSGGLALLWKDSVRLS 460
Query: 535 SRLLYSHQQAVTFEVWRENLSWVCSAIYASPIPVKREELWSHLLHIRSNLVLPWMIVGDM 594
+ LY + + + N+++ S +Y P +R LW+H ++ PW+++GD
Sbjct: 461 N--LYQDDRHIDVHISINNINFYLSRVYGHPCQSERHSLWTHFENLSKTRNDPWILIGDF 518
Query: 595 NEILHPNEVRGG*FLPTRAS----RFAAILEGCGMVDLGMSGGRYTWFRKQNNRLIMSKR 650
NEIL NE GG P R F ++ C + D+ G R++W ++++ +
Sbjct: 519 NEILSNNEKIGG---PQRDEWTFRGFRNMVSTCDLKDIRSIGDRFSWVGERHSHTVKCC- 574
Query: 651 LDRAMGDGAWRIAFPEASVEALHRIHSDHCPLLINCSGPEIVHHDRPFRFLASWVEHPQY 710
LDRA + FP A +E L SDH PL ++ E RPFRF +E P +
Sbjct: 575 LDRAFINSEGAFLFPFAELEFLEFTGSDHKPLFLSLEKTE-TRKMRPFRFDKRLLEVPHF 633
Query: 711 KEVVQSAWT----GETETVAGKLNKVRTKSCKFNKEVYGSIFQRKRWVEGRLRGVQRELN 766
K V++ W G+ + + ++ R K + + R ++ L +N
Sbjct: 634 KTYVKAGWNKAINGQRKHLPDQVRTCRQAMAKLKHKSNLNSRIRINQLQAALDKAMSSVN 693
Query: 767 YRVTFDMTRLETELQAEYRFLLRQEELLWFQKARENRVCFDDRNTAYFHAHTIIRRKRNK 826
++ ++ EL YR EE W QK+R + DRNT +FHA T R N+
Sbjct: 694 RTERRTISHIQRELTVAYR----DEERYWQQKSRNQWMKEGDRNTEFFHACTKTRFSVNR 749
Query: 827 IHRLKLNDGSWCDDRQLLAEEIQNYFQSLFAADPDNTDVNMQHVMMPRLTEAERHKAEAE 886
+ +K +G + + Q +F ++ ++ + P +TE +
Sbjct: 750 LVTIKDEEGMIYRGDKEIGVHAQEFFTKVYESNGRPVSIIDFAGFKPIVTEQINDDLTKD 809
Query: 887 VIKEEVFQALLRMRSYSALGPDGFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLET 946
+ E++ A+ + A GPDG F+K WE+VG D+ K V+ F T + S+ T
Sbjct: 810 LSDLEIYNAICHIGDDKAPGPDGLTARFYKSCWEIVGPDVIKEVKIFFRTSYMKQSINHT 869
Query: 947 LIVLIPKVDSPSTVKEFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIM 1006
I +IPK+ +P T+ ++RPI+LCNV+YK+I+K +V R++ L+ I+ Q +F+PG +
Sbjct: 870 NICMIPKITNPETLSDYRPIALCNVLYKIISKCLVERLKGHLDAIVSDSQAAFIPGRLVN 929
Query: 1007 DNAFLAQEIIHHMGA-SMAKKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIM 1065
DN +A E++H + + +A K D+ KAYD V W FLE T+ FGF ++ IM
Sbjct: 930 DNVMIAHEMMHSLKTRKRVSQSYMAVKTDVSKAYDRVEWNFLETTMRLFGFSETWIKWIM 989
Query: 1066 RCVSGSNLSMLWNGSRLPSFKPGRGLRKGDPLSPYLFVLCS 1106
V N S+L NG + +P RG+R+GDPLSPYLF+LC+
Sbjct: 990 GAVKSVNYSVLVNGIPHGTIQPQRGIRQGDPLSPYLFILCA 1030
Score = 149 bits (376), Expect = 9e-34
Identities = 144/571 (25%), Positives = 226/571 (39%), Gaps = 119/571 (20%)
Query: 1230 KKKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSG 1289
KK+ SSW L+ G+ + KSV ++P Y M F LP +I +I + F W KN+
Sbjct: 1155 KKRTSSWSAKYLSPAGKEIMLKSVAMSMPVYAMSCFKLPLNIVSEIEALLMNFWWEKNAK 1214
Query: 1290 QRGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLAHKYLHG 1349
+R + WK L K+ GGL +D+ FN +LL K VW + NN + L+ +++ +Y
Sbjct: 1215 KREIPWIAWKRLQYSKKEGGLGFRDLAKFNDALLAKQVWRMINNPNSLFARIMKARYFRE 1274
Query: 1350 ESIFSVHRRANASPIWQGILKARDQLQEGFQFRMGDGSSSVWYHDWTGKGKLAESLDFVH 1409
+SI R+ S W +L D +++G +F +GDG + Y W H
Sbjct: 1275 DSILDAKRQRYQSYGWTSMLAGLDVIKKGSRFIVGDGKTGS-YRYWN-----------AH 1322
Query: 1410 ISDTKLCLHELV--DNNQWNLNRCMTVFPLHIRDWFARVEPRVVANGIDRWSWKDNSDGI 1467
+ + +LV D++++ +N ++ R+V D+ W +S
Sbjct: 1323 L------ISQLVSPDDHRFVMNHHLS---------------RIVHQ--DKLVWNYSSS-- 1357
Query: 1468 SSVHDGYQWLREKKMAGVVHGDSWLWI*KLHAPEKL*VFIWLVLHDALPTNANRYRCKLA 1527
GD LW KL K+ +W + ALPT + +
Sbjct: 1358 --------------------GDYTLW--KLPIIPKIKYMLWRTISKALPTRSRLLTRGMD 1395
Query: 1528 VSPGCTRCSSPEEDTIHCLRDCPHSQELWGKLGAWGWSTFRLPDLKA*VKTQLQGSNNIR 1587
+ P C RC + EE H L CP++ +WG L + W LP T+ NI
Sbjct: 1396 IDPHCPRCPTEEETINHVLFTCPYAASIWG-LSNFPW----LPGHTFSQDTE----ENIS 1446
Query: 1588 FLAG------------------LWGVWKWRNNVVLDPLPWTLDGAWQKLRHDHDELLQF- 1628
FL +W +WK RNN+V + + + + +E LQ
Sbjct: 1447 FLINSFSNNTLNTEQRLAPFWLIWRLWKARNNLVFNKFSESCSRVVTQTEAEVNEWLQSV 1506
Query: 1629 ----SSGDLLDEHHFLINLWKPPPPEFVKLNTDGSFQDTASSMGGGGLIRDAKGAWLGGF 1684
+ L H WK P VK N D F + GG +I + K
Sbjct: 1507 NRREDASVLTRSSHRANVRWKKPVFPLVKCNFDAGFTGNNTQSTGGWIIPETK------- 1559
Query: 1685 MSHTSIGNPFLAEVLALRDGLCLAWVQGHRRIIREVDCAGLSDILRFE-DRCRLHDHAGV 1743
AL + AWV+G++ + E DC L + RC + +
Sbjct: 1560 ---------------ALLIAMQQAWVRGYKCVQFEGDCEILIKAINGAISRCEI---TSL 1601
Query: 1744 LQEIRELMERPWRCSVHWTNRDSNASADFLA 1774
L+++ R +TNR N +A LA
Sbjct: 1602 LRDVDFWASRFSTVVFTFTNRLCNNTAHLLA 1632
>dbj|BAB01845.1| non-LTR retroelement reverse transcriptase-like protein [Arabidopsis
thaliana]
Length = 893
Score = 305 bits (780), Expect = 1e-80
Identities = 256/926 (27%), Positives = 444/926 (47%), Gaps = 82/926 (8%)
Query: 457 KIISWNVRGA-LNEHGQLFLKDLIRCKKPDIMFLLETRCQFSRASNFWNSTRFSPLFISE 515
K+ WNVRG ++ H + F K + KP L+ET + + F ++ F+ E
Sbjct: 4 KLFCWNVRGFNISSHRRGFKKWFL-LNKPLFGGLIETHVKQPKEKKFISNLLPGWSFV-E 61
Query: 516 ANGFS--GGIWVLVNTATNLSSRLLYSHQQAVTFEVWR-ENLSW-VCSAIYASPIPVKRE 571
FS G IWVL + + + ++ Q +T E+ ++ SW V S +YAS R+
Sbjct: 62 NYEFSVLGKIWVLWDPSVKVV--VIGRSLQMITCELLLPDSPSWFVVSIVYASNEEGTRK 119
Query: 572 ELWSHLLHIRSNLVL---PWMIVGDMNEILHPNEVRGG*FLPTRASRFAAILEGCGMVDL 628
ELW+ L+ + + V+ W+++GD N+IL+P + + F + L + DL
Sbjct: 120 ELWNELVQLALSPVVVGRSWIVLGDFNQILNPESAINA-NIGRKIRAFRSCLLDSDLYDL 178
Query: 629 GMSGGRYTWFRKQNNRLIMSKRLDRAMGDGAWRIAFPEASVEALHRIHSDH--CPLLINC 686
G YTW+ K ++R + +K++DR + + W FP A SDH C ++++
Sbjct: 179 VYKGSSYTWWNKCSSRPL-AKKIDRILVNDHWNTLFPSAYANFGEPDFSDHSSCEVVLD- 236
Query: 687 SGPEIVHHDRPFRFLASWVEHPQYKEVVQSAWTGETET------VAGKLNKVRTKSCKFN 740
P ++ RPFRF ++ +P + ++++ W + V+ KL ++ C F+
Sbjct: 237 --PAVLKAKRPFRFFNYFLHNPDFLQLIRENWYSCNVSGSAMYRVSKKLKHLKLPICCFS 294
Query: 741 KEVYGSIFQRKRWVEGRLRGVQRELNYRVTFDMTRLETELQAEYRFLLRQEELLWFQKAR 800
+E Y I +R + QR + LE E +++ L + EE + QK+
Sbjct: 295 RENYSDIEKRVSEAHAIVLHRQRITLTNPSVVHATLELEATRKWQILAKAEESFFCQKSS 354
Query: 801 ENRVCFDDRNTAYFHAHTIIRRKRNKIHRLKLNDGSWCDDRQLLAEEIQ----NYFQSLF 856
+ + D NTAYFH +R+ N I+ L + G + +Q + E I+ N+F+SL
Sbjct: 355 ISWLYEGDNNTAYFHKMADMRKSINTINFLIDDFGERIETQQGIKEGIKEHSCNFFESLL 414
Query: 857 AA-DPDNT----DVNMQ---HVMMPRLTEAERHKAEAEVIKEEVFQALLRMRSYSALGPD 908
+ +N+ D+N+ + ++ + ER ++ ++ +E F +L R + A GPD
Sbjct: 415 CGVEGENSLAQSDMNLLLSFRCSVDQINDLERSFSDLDI--QEAFFSLPRNK---ASGPD 469
Query: 909 GFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKEFRPISL 968
G+ FFK W +VG ++ + V++ F +G + T +VLIPK+ + S + +FRPIS
Sbjct: 470 GYSSEFFKGVWFVVGPEVTEAVQEFFRSGQLLKQWNATTLVLIPKITNSSKMTDFRPISC 529
Query: 969 CNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEIIH-HMGASMAKKG 1027
N +YK+I K++ +R++ LLN +I P Q++FLPG + +N LA EI+H + +++ +G
Sbjct: 530 LNTLYKVIAKLLTSRLKKLLNEVISPSQSAFLPGRLLSENVLLATEIVHGYNTKNISSRG 589
Query: 1028 TLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGSRLPSFKP 1087
L K+DL KA+DSV W F+ P + V I +C+S S++ NGS FK
Sbjct: 590 ML--KVDLRKAFDSVRWDFIISAFRALAVPEKFVCWINQCISTPYFSVMVNGSSSGFFKS 647
Query: 1088 GRGLRKGDPLSPYLFVLCSLHGAIVY*YS*VGGNRSLEAN*NFPKGTTYFASLFCR*RAL 1147
+GLR+GDPLSPYLFVL V+ + L+A F G ++ + L
Sbjct: 648 NKGLRQGDPLSPYLFVL----AMEVF-------SSLLKA--RFDAGYIHYHP---KTADL 691
Query: 1148 VLSSIIFSSAI----GGGDS*TILCLFGVKDKYEQI*S-----------DCFQGSTKSCE 1192
+S ++F+ + GG S L G+ + + S + + T E
Sbjct: 692 SISHLMFADDVMVFFDGGSS----SLHGISEALDDFASWSGLHVNKDKTNLYLAGTDEVE 747
Query: 1193 RGDPEYCPYPFCAKPWQVSWLEEGFQEVDSITYWIIFKKKLSSWKTNLLNMVGQVCLAKS 1252
+ +P P + L +++ Y ++ K+ SW L+ G+V L S
Sbjct: 748 ALAISHYGFPISTLPIRYLGLPLMSRKLKISEYELV--KRFRSWAVKSLSFAGRVQLITS 805
Query: 1253 VISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSGQRGWHRVNWKTLTQPKEHGGLAI 1312
VI+ + + M F L KI RF+W + ++ W + PK GG+ +
Sbjct: 806 VITGLVNFWMSTFVLLLGCVKKIESLCSRFLWSGSIDASKGAKIAWSGVCLPKNEGGVGL 865
Query: 1313 KDMGDFNTSLLGKVVWLLANNSSKLW 1338
+ +N + + +W L ++ LW
Sbjct: 866 RRFTPWNKTFYLRFIWPLFADNDVLW 891
>pir||S65812 RNA-directed DNA polymerase (EC 2.7.7.49) (clone DW15) - Arabidopsis
thaliana retrotransposon Ta11-1 gi|976278|gb|AAA75254.1|
reverse transcriptase
Length = 1333
Score = 303 bits (777), Expect = 3e-80
Identities = 204/663 (30%), Positives = 325/663 (48%), Gaps = 25/663 (3%)
Query: 456 LKIISWNVRGALNEHGQLFLKDLIRCKK---PDIMFLLETRCQFSRASNFWNSTRFSPLF 512
+ ++SWN +G L L + L+ + P+++FL+ET+ + + + +F
Sbjct: 1 MSLVSWNCQG-LGWSQDLTIPRLMEMRLSHFPEVLFLMETKNCSNVVVDLQEWLGYERVF 59
Query: 513 ISEANGFSGGIWVLVNTATNLSSRLLYSHQQAVTFEVWRENLSWVCSAIYASPIPVKREE 572
G SGG+ + ++ + Y+ + + F++ + + S +Y +P +
Sbjct: 60 TVNPIGLSGGLALFWKKGVDIVIK--YADKNLIDFQIQFGSHEFYVSCVYGNPAFSDKHL 117
Query: 573 LWSHLLHIRSNLVLPWMIVGDMNEILHPNEVRGG*FLPTRASR----FAAILEGCGMVDL 628
+W + I N PW ++GD N ILH E RGG P R F +L+ C M++L
Sbjct: 118 VWEKITRIGINRKEPWCMLGDFNPILHNGEKRGG---PRRGDSSFLPFTDMLDSCDMLEL 174
Query: 629 GMSGGRYTWFRKQNNRLIMSKRLDRAMGDGAWRIAFPEASVEALHRIHSDHCPLLINCSG 688
G +TW K N I S RLDR G+ W FP ++ E L + SDH P+L+ +
Sbjct: 175 PSIGNPFTWGGKTNEMWIQS-RLDRCFGNKNWFRFFPISNQEFLDKRGSDHRPVLVRLTK 233
Query: 689 PEIVHHDRPFRFLASWVEHPQYKEVVQSAWTG----ETETVAGKLNKVRTKSCKFNKEVY 744
+ + FRF P KE + AW G E V KL R+ ++ KE
Sbjct: 234 TKEEYRGN-FRFDKRLFNQPNVKETIVQAWNGSQRNENLLVLDKLKHCRSALSRWKKE-- 290
Query: 745 GSIFQRKRWVEGRLRGVQRELNYRVTFDMTRLETELQAEYRFLLRQEELLWFQKARENRV 804
+I R + R EL F L L+ + EE+ W QK+R +
Sbjct: 291 NNINSSTRITQAR---AALELEQSSGFPRADLVFSLKNDLCKANHDEEVFWSQKSRAKWM 347
Query: 805 CFDDRNTAYFHAHTIIRRKRNKIHRLKLNDGSWCDDRQLLAEEIQNYFQSLFAADPDNTD 864
D+NT++FHA R + I +L +G + D + YF LF + ++
Sbjct: 348 HSGDKNTSFFHASVKDNRGKQHIDQLCDVNGLFHKDEMNKGAIAEAYFSDLFKSTDPSSF 407
Query: 865 VNMQHVMMPRLTEAERHKAEAEVIKEEVFQALLRMRSYSALGPDGFHPFFFKKYWELVGD 924
V++ PR+TE+ + A V K E+ +A+ +RS SA G DGF FFF+KYW ++
Sbjct: 408 VDLFEDYQPRVTESMNNTLIAAVSKNEIREAVFAIRSSSAPGVDGFTGFFFQKYWSIICL 467
Query: 925 DLWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVKEFRPISLCNVVYKLITKVVVNRI 984
+ K +++ F G S T + L+PK P + + RPISLC+V+YK+I+K++V R+
Sbjct: 468 QVTKEIQNFFLLGYFPKSWNFTHLCLLPKKKKPDKMTDLRPISLCSVLYKIISKIMVRRL 527
Query: 985 RPLLNRIIGPMQNSFLPGMGIMDNAFLAQEIIHHMGA-SMAKKGTLAFKIDLEKAYDSVS 1043
+P L ++ P Q++F+ I DN +A E++H + KG +A K ++ KA+D V
Sbjct: 528 QPFLPDLVSPNQSAFVAERLIFDNILIAHEVVHGLRTHKSVSKGFIAIKSNMSKAFDRVE 587
Query: 1044 WAFLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGSRLPSFKPGRGLRKGDPLSPYLFV 1103
W ++ L GF + V IM +S + S+L N + P RGLR+GDPLSP+LFV
Sbjct: 588 WNYVRALLDALGFHQKWVGWIMFMISSVSYSVLINDKAFGNIVPSRGLRQGDPLSPFLFV 647
Query: 1104 LCS 1106
LCS
Sbjct: 648 LCS 650
Score = 135 bits (339), Expect = 2e-29
Identities = 124/480 (25%), Positives = 204/480 (41%), Gaps = 38/480 (7%)
Query: 1230 KKKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSG 1289
K +LS W L+M G+ L K+ A+ Y M F L ++ + M F W
Sbjct: 775 KTRLSGWFARTLSMGGKETLLKAFALALLFYAMSCFKLTKTTCVNMTSAMSDFWWNALEH 834
Query: 1290 QRGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLAHKYLHG 1349
+R H V+ + + KE+GGL +D+ FN +LL K W L + L+ + +Y
Sbjct: 835 KRKTHWVSCEKMCLSKENGGLGFRDIESFNQALLAKQAWRLLQFPNSLFARFFKSRYYDE 894
Query: 1350 ESIFSVHRRANASPIWQGILKARDQLQEGFQFRMGDGSS-SVWYHDWTGKGKLAESLDFV 1408
E +A S W+ IL RD L +GF+ ++G+GSS SVW W L
Sbjct: 895 EDFLDAELKATPSYAWRSILHGRDLLIKGFRKKVGNGSSTSVWMDPWIYDNDPRLPLQKH 954
Query: 1409 HISDTKLCLHELVDNNQWNLNRCMTVFPLHIRDWFARVEPRVVANGI----DRWSWKDNS 1464
+ L +H+L++ +RC L + A +E V N + D W W +
Sbjct: 955 FSVNLDLRVHDLINVE----DRCRRRDRLEELFYPADIEIIVKRNPVVSMDDFWVWLHSK 1010
Query: 1465 DGISSVHDGY---------QWLREKKMAGVVHG-DSWLWI*KLHAPEKL*VFIWLVLHDA 1514
G SV GY + +RE ++ +G +W L +P K+ +F+W +L A
Sbjct: 1011 SGEYSVKSGYWLAFQTNKPELIREARVQPSTNGLKEKIWS-TLTSP-KIKLFLWRILSSA 1068
Query: 1515 LPTNANRYRCKLAVSPGCTRCSSPEEDTIHCLRDCPHSQELWGKLGA-WGWSTFRLPDLK 1573
LP R + + P C C E H L C ++++W G F+ +
Sbjct: 1069 LPVAYQIIRRGMPIDPRCQVCGEEGESINHVLFTCSLARQVWALSGVPTSQFGFQNSSIF 1128
Query: 1574 A*VK--TQLQGSNNI------RFLAGLWGVWKWRNNVVLDPLPWTLDGAWQKLRHDHDE- 1624
A ++ +L+G I + LW +WK R+ + + ++ + +K+R D E
Sbjct: 1129 ANIQYLLELKGKGLIPEQIKKSWPWVLWRLWKNRDKLFFEGTIFSPLKSIEKIRDDVQEW 1188
Query: 1625 ------LLQFSSGDLLDEHHFLINLWKPPPPEFVKLNTDGSFQDTASSMGGGGLIRDAKG 1678
+ +G+ + + W+PPP +VK N G + GG ++RD G
Sbjct: 1189 FLAQALVASVDAGETVCSAP-CPSSWEPPPLGWVKCNISGVWSGKKRVCGGAWVLRDDHG 1247
>gb|AAD24831.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25408166|pir||G84721 hypothetical protein
At2g31520 [imported] - Arabidopsis thaliana
Length = 1524
Score = 302 bits (774), Expect = 6e-80
Identities = 197/632 (31%), Positives = 315/632 (49%), Gaps = 34/632 (5%)
Query: 490 LETRCQFSRASNFWNSTRFSPLFISEANGFSGGIWVLVNTATNLSSRLLYSHQQAVTFEV 549
LET Q + F + NG SGG+ ++ N+S L+ ++ + V
Sbjct: 190 LETLNQCDKVCKLAYDLGFPNVITQPPNGRSGGLALMWKN--NVSLSLISQDERLIDSHV 247
Query: 550 WRENLSWVCSAIYASPIPVKREELWSHLLHIRSNLVLPWMIVGDMNEILHPNEVRGG*FL 609
N S+ S +Y P +R +LW L HI N W++VGD NEIL E GG
Sbjct: 248 TFNNKSFYLSCVYGHPTQSERHQLWQTLEHISDNRNAEWLLVGDFNEILSNAEKIGG--- 304
Query: 610 PTRAS----RFAAILEGCGMVDLGMSGGRYTWFRKQNNRLIMSKRLDRAMGDGAWRIAFP 665
P R F ++ C + D+ G R++W +++ + LDR + AW FP
Sbjct: 305 PMREEWTFRNFRNMVSHCDIEDMRSKGDRFSWVGERHTHTVKCC-LDRVFINSAWTATFP 363
Query: 666 EASVEALHRIHSDHCPLLINCSGPEIVHHDRPFRFLASWVEHPQYKEVVQSAWTGETETV 725
A E L SDH P+L++ + + FRF ++ P +K +VQ++W +
Sbjct: 364 YAETEFLDFTGSDHKPVLVHFN-ESFPRRSKLFRFDNRLIDIPTFKRIVQTSWRTNRNSR 422
Query: 726 AGKLNKVRTKSCK--FNKEVYGSIFQRKRWVEGRLRGVQRELNYRVTFDMTR-----LET 778
+ + + R SC+ + + S E R++ +Q LN + + TR L
Sbjct: 423 STPITE-RISSCRQAMARLKHASNLNS----EQRIKKLQSSLNRAM--ESTRRVDRQLIP 475
Query: 779 ELQAEYRFLLRQEELLWFQKARENRVCFDDRNTAYFHAHTIIRRKRNKIHRLKLNDGSWC 838
+LQ EE+ W QK+R + D+NT YFHA T R +N+++ + + G
Sbjct: 476 QLQESLAKAFSDEEIYWKQKSRNQWMKEGDQNTGYFHACTKTRYSQNRVNTIMDDQGRMF 535
Query: 839 DDRQLLAEEIQNYFQSLFAADPDNTDVNMQHVMMPRLTEAERHKAEAEVIKE----EVFQ 894
+ + Q++F ++F+ + + + + + ++ KE E++
Sbjct: 536 TGDKEIGNHAQDFFTNIFSTN----GIKVSPIDFADFKSTVTNTVNLDLTKEFSDTEIYD 591
Query: 895 ALLRMRSYSALGPDGFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKV 954
A+ ++ A GPDG F+K W++VG D+ V+ FET + S+ T I +IPK+
Sbjct: 592 AICQIGDDKAPGPDGLTARFYKNCWDIVGYDVILEVKKFFETSFMKPSINHTNICMIPKI 651
Query: 955 DSPSTVKEFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQE 1014
+P+T+ ++RPI+LCNV+YK+I+K +VNR++ LN I+ Q +F+PG I DN +A E
Sbjct: 652 TNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPGRIINDNVMIAHE 711
Query: 1015 IIHHMGA-SMAKKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNL 1073
++H + K +A K D+ KAYD V W FLE T+ FGF N+ + IM V +
Sbjct: 712 VMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWIGWIMAAVKSVHY 771
Query: 1074 SMLWNGSRLPSFKPGRGLRKGDPLSPYLFVLC 1105
S+L NGS P RG+R+GDPLSPYLF+LC
Sbjct: 772 SVLINGSPHGYITPTRGIRQGDPLSPYLFILC 803
Score = 155 bits (393), Expect = 9e-36
Identities = 153/609 (25%), Positives = 234/609 (38%), Gaps = 99/609 (16%)
Query: 1230 KKKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSG 1289
KK+ S+W L+ G+ + KSV A+P Y M F LP+ I +I + F W K S
Sbjct: 929 KKRTSTWSARFLSPAGKEIMLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASN 988
Query: 1290 QRGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLAHKYLHG 1349
QRG V WK L K+ GGL +D+ FN +LL K W L + L+ +V+ +Y
Sbjct: 989 QRGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKD 1048
Query: 1350 ESIFSVHRRANASPIWQGILKARDQLQEGFQFRMGDG-------------------SSSV 1390
SI R S W +L L++G + +GDG ++
Sbjct: 1049 VSILDAKVRKQQSYGWASLLDGIALLKKGTRHLIGDGQNIRIGLDNIVDSHPPRPLNTEE 1108
Query: 1391 WYHDWTGKGKLAESLDFVHISDTKLCLHELVDNNQWNLNRCMTVFPLHIRDWFARVEPRV 1450
Y + T + D+K + + VD + +H R + A+
Sbjct: 1109 TYKEMTINNLFERKGSYYFWDDSK--ISQFVDQSDHGF--------IH-RIYLAK----- 1152
Query: 1451 VANGIDRWSWKDNSDGISSVHDGYQWLREKKMAGV-----VHG--DSWLWI*KLHAPEKL 1503
+ D+ W N+ G +V GY L + HG D I L KL
Sbjct: 1153 -SKKPDKIIWNYNTTGEYTVRSGYWLLTHDPSTNIPAINPPHGSIDLKTRIWNLPIMPKL 1211
Query: 1504 *VFIWLVLHDALPTNANRYRCKLAVSPGCTRCSSPEEDTIHCLRDCPHSQELWGKLGAWG 1563
F+W L AL T + + P C RC E H L CP + W W
Sbjct: 1212 KHFLWRALSQALATTERLTTRGMRIDPICPRCHRENESINHALFTCPFATMAW-----W- 1265
Query: 1564 WSTFRLPDLKA*VKTQLQG-------SNNIRFLAG--------------LWGVWKWRNNV 1602
L D + ++ QL SN + F+ +W +WK RNNV
Sbjct: 1266 -----LSD-SSLIRNQLMSNDFEENISNILNFVQDTTMSDFHKLLPVWLIWRIWKARNNV 1319
Query: 1603 VLDPL---PWTLDGAWQKLRHDHDELLQFSSGDLLDEHHFLINL--WKPPPPEFVKLNTD 1657
V + P + + HD Q N W+ PP +VK N D
Sbjct: 1320 VFNKFRESPSKTVLSAKAETHDWLNATQSHKKTPSPTRQIAENKIEWRNPPATYVKCNFD 1379
Query: 1658 GSFQDTASSMGGGGLIRDAKG---AWLGGFMSHTSIGNPFLAEVLALRDGLCLAWVQGHR 1714
F GG +IR+ G +W ++HTS NP AE AL L W++G+
Sbjct: 1380 AGFDVQKLEATGGWIIRNHYGTPISWGSMKLAHTS--NPLEAETKALLAALQQTWIRGYT 1437
Query: 1715 RIIREVDCAGLSDILRFEDRCRLHDHAGVLQEIRELMERPWRCSVHWTNRDSNASADFLA 1774
++ E DC L +++ + H+ + + ++ W N+ ++ F+
Sbjct: 1438 QVFMEGDCQTLINLIN-----GISFHSSLANHLEDIS--------FWANKFASIQFGFIR 1484
Query: 1775 RRGGFLSTI 1783
R+G L+ +
Sbjct: 1485 RKGNKLAHV 1493
>emb|CAB78094.1| RNA-directed DNA polymerase-like protein [Arabidopsis thaliana]
gi|4538901|emb|CAB39638.1| RNA-directed DNA
polymerase-like protein [Arabidopsis thaliana]
gi|7485606|pir||T04018 hypothetical protein F17A8.60 -
Arabidopsis thaliana
Length = 1274
Score = 293 bits (751), Expect = 3e-77
Identities = 186/575 (32%), Positives = 302/575 (52%), Gaps = 41/575 (7%)
Query: 550 WRENLSWVCSAIYASPIPV-KREELWSHLLHIRSNLVLPWMIVGDMNEILHPNEVRGG*F 608
W+EN+ + A+P + R W + + + W++ GD N+IL +E +GG
Sbjct: 38 WKENVE--VEILEAAPNFIDNRSVFWDKISSLGAQRSSAWLLTGDFNDILDNSEKQGG-- 93
Query: 609 LPTRAS----RFAAILEGCGMVDLGMSGGRYTWFRKQNNRLIMSKRLDRAMGDGAWRIAF 664
P R F + + G+ D+ +G +W + + I S RLDRA+G+ +W F
Sbjct: 94 -PLRWEGFFLAFRSFVSQNGLWDINHTGNSLSWRGTRYSHFIKS-RLDRALGNCSWSELF 151
Query: 665 PEASVEALHRIHSDHCPLLINCSGPEIVHHDRPFRFLASWVEHPQYKEVVQSAWT-GETE 723
P + E L SDH PL+ P + +PFRF E + + +V+ W +
Sbjct: 152 PMSKCEYLRFEGSDHRPLVTYFGAPPL-KRSKPFRFDRRLREKEEIRALVKEVWELARQD 210
Query: 724 TVAGKLNKVRTKSCKFNKEVYGSIFQRKRWVEGRLRGVQRELNYRVTFDMT------RLE 777
+V K+++ R K+ KE + + ++ Q+ L ++ D+ +
Sbjct: 211 SVLYKISRCRQSIIKWTKEQNSNSAKA-------IKKAQQALESALSADIPDPSLIGSIT 263
Query: 778 TELQAEYRFLLRQEELLWFQKARENRVCFDDRNTAYFHAHTIIRRKRNKIHRLKLNDGSW 837
EL+A YR QEEL W Q +R + DRN YFHA T RR N + ++ G
Sbjct: 264 QELEAAYR----QEELFWKQWSRVQWLNSGDRNKGYFHATTRTRRMLNNLSVIEDGSGQE 319
Query: 838 CDDRQLLAEEIQNYFQSLFAADPDNTDVNM-QHVMMPRLTEAERHKAEAEVIKE----EV 892
+ + +A I +YFQ++F +N+D+ + Q + P ++ E+IK E+
Sbjct: 320 FHEEEQIASTISSYFQNIFTTS-NNSDLQVVQEALSPIISS----HCNEELIKISSLLEI 374
Query: 893 FQALLRMRSYSALGPDGFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIP 952
+AL + + A GPDGF FF YW+++ D+ + +R F + L ET + LIP
Sbjct: 375 KEALFSISADKAPGPDGFSASFFHAYWDIIEADVSRDIRSFFVDSCLSPRLNETHVTLIP 434
Query: 953 KVDSPSTVKEFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLA 1012
K+ +P V ++RPI+LCNV YK++ K++ R++P L+ +I Q++F+PG I DN +
Sbjct: 435 KISAPRKVSDYRPIALCNVQYKIVAKILTRRLQPWLSELISLHQSAFVPGRAIADNVLIT 494
Query: 1013 QEIIHHMGASMAKK-GTLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGS 1071
EI+H + S AKK ++A K D+ KAYD + W FL++ L++ GF ++ ++ +M+CV
Sbjct: 495 HEILHFLRVSGAKKYCSMAIKTDMSKAYDRIKWNFLQEVLMRLGFHDKWIRWVMQCVCTV 554
Query: 1072 NLSMLWNGSRLPSFKPGRGLRKGDPLSPYLFVLCS 1106
+ S L NGS S P RGLR+GDPLSPYLF+LC+
Sbjct: 555 SYSFLINGSPQGSVVPSRGLRQGDPLSPYLFILCT 589
Score = 135 bits (339), Expect = 2e-29
Identities = 140/570 (24%), Positives = 228/570 (39%), Gaps = 47/570 (8%)
Query: 1230 KKKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSG 1289
+++ SW L+ G+ L K+V+S++P+Y M F LP S+ +I + RF W
Sbjct: 714 RQRSHSWSIRFLSSAGKQILLKAVLSSMPSYAMMCFKLPASLCKQIQSVLTRFWWDSKPD 773
Query: 1290 QRGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLAHKYLHG 1349
+R V+W LT P GGL ++ + K+ W + L +VL KY +
Sbjct: 774 KRKMAWVSWDKLTLPINEGGLGFRE-------IEAKLSWRILKEPHSLLSRVLLGKYCNT 826
Query: 1350 ESIFSVHRRAN-ASPIWQGILKARDQLQEGFQFRMGDGSS-SVWYHDWTGKGKLAESLDF 1407
S + AS W+GIL RD L++G + +G G S +VW W +
Sbjct: 827 SSFMDCSASPSFASHGWRGILAGRDLLRKGLGWSIGQGDSINVWTEAWLSPSSPQTPIGP 886
Query: 1408 VHISDTKLCLHELV--DNNQWNLNRCMTVFPLHIRDWFARVEPRVVANGI---DRWSWKD 1462
++ L +H+L+ D WN+ H+ + ++ ++ N + D W
Sbjct: 887 PTETNKDLSVHDLICHDVKSWNVE----AIRKHLPQYEDQIR-KITINALPLQDSLVWLP 941
Query: 1463 NSDGISSVHDGYQWLREKKMAGVVHGDSW---LWI*KLHAPEKL*VFIWLVLHDALPTNA 1519
G + GY + +W +W K+H K+ F+W + ALP
Sbjct: 942 VKSGEYTTKTGYALAKLNSFPASQLDFNWQKNIW--KIHTSPKVKHFLWKAMKGALPVGE 999
Query: 1520 NRYRCKLAVSPGCTRCSSPEEDTIHCLRDCPHSQELW--------GKLGAWGWSTFRLPD 1571
R + C RC E ++H + CP+++++W L D
Sbjct: 1000 ALSRRNIEAEVTCKRCGQ-TESSLHLMLLCPYAKKVWELAPVLFNPSEATHSSVALLLVD 1058
Query: 1572 LKA*VKTQLQGSNNIRFLAG-LWGVWKWRNNVVLDPLPWTLDGAWQKLRHDHDELLQFSS 1630
K V G + LW +WK RN ++ D + +G K D ++
Sbjct: 1059 AKRMVALPPTGLGSAPLYPWLLWHLWKARNRLIFDNHSCSEEGLVLKAILDARAWME--- 1115
Query: 1631 GDLLDEHHFLINLWKPPPPEFVKLNTDGSFQDTASSMGG----GGLIRDAKGAWLGGFMS 1686
LL H I+ + P P L F D A + G G ++D + S
Sbjct: 1116 AQLLIHHPSPISDYPSPTP---NLKVTSCFVDAAWTTSGYCGMGWFLQDPYKVKIKENQS 1172
Query: 1687 HTS-IGNPFLAEVLALRDGLCLAWVQGHRRIIREVDCAGLSDILRFEDRCRLHDHAGVLQ 1745
+S +G+ +AE LA+ L A G R++ DC L +L + + G+L
Sbjct: 1173 SSSFVGSALMAETLAVHLALVDALSTGVRQLNVFSDCKELISLL--NSGKSIVELRGLLH 1230
Query: 1746 EIRELMERPWRCSVHWTNRDSNASADFLAR 1775
+IREL + R SN AD LA+
Sbjct: 1231 DIRELSVSFTHLCFFFIPRLSNVVADSLAK 1260
>emb|CAE02147.1| OSJNBa0081G05.2 [Oryza sativa (japonica cultivar-group)]
gi|50923489|ref|XP_472105.1| OSJNBa0081G05.2 [Oryza
sativa (japonica cultivar-group)]
Length = 1711
Score = 288 bits (738), Expect = 9e-76
Identities = 179/585 (30%), Positives = 292/585 (49%), Gaps = 20/585 (3%)
Query: 486 IMFLLETRCQFSRASNFWNSTRFSPLFISEANGFSGGIWVLVNTATNLSSRLLYSHQQAV 545
++FL ETR + S F + G SGG+ + + + ++ + +
Sbjct: 578 LVFLCETRQSVEKMSRLRRKLAFRGFVGVSSEGMSGGLALYWDESVSVDVKDINKRYIDA 637
Query: 546 TFEVWRENLSWVCSAIYASPIPVKREELWSHLLHIRSNLVLPWMIVGDMNEILHPNE-VR 604
+ + W + +Y P R +WS L IR + LPWM++GD NE L E
Sbjct: 638 YVRLSSDEPQWHITFVYGEPRVENRHRMWSLLRTIRQSSALPWMVIGDFNETLWQFEHFS 697
Query: 605 GG*FLPTRASRFAAILEGCGMVDLGMSGGRYTWFRKQNNRLIMSKRLDRAMGDGAWRIAF 664
T+ F L C + DLG G +T+ +++ + RLDRA+ D WR F
Sbjct: 698 KNPRCETQMQNFRDALYDCDLQDLGFKGVPHTYDNRRDGWRNVKVRLDRAVADDKWRDLF 757
Query: 665 PEASVEALHRIHSDHCPLLINCSGPEIVH-HDRPFRFLASWVEHPQYKEVVQSAWTGETE 723
PEA V L SDH P+L+ + + + W P+ +V++ AW
Sbjct: 758 PEAQVSHLVSPCSDHSPILLEFIVKDTTRPRQKCLHYEIVWEREPESVQVIEEAWIN--- 814
Query: 724 TVAGKLNKVRTKSCKFNKEVYGSIFQRKRWVEGRLRGVQRELNYR-------VTFDMTRL 776
AG V+T N + + + W + +++ V REL + + R
Sbjct: 815 --AG----VKTDLGDINIALGRVMSALRSWGKTKVKNVGRELEKARKKLEDLIASNAARS 868
Query: 777 ETELQAEY-RFLLRQEELLWFQKARENRVCFDDRNTAYFHAHTIIRRKRNKIHRLKLNDG 835
++ +L +EE+LW Q++R N + DRNT +FH+ + R K+NKI +L+ +G
Sbjct: 869 SIRQATDHMNEMLYREEMLWLQRSRVNWLKEGDRNTRFFHSRAVWRAKKNKISKLRDENG 928
Query: 836 SWCDDRQLLAEEIQNYFQSLFAADPDNTDVNMQHVMMPRLTEAERHKAEAEVIKEEVFQA 895
+ +L YFQ ++ ADP ++ + ++T+A K E +EE+ QA
Sbjct: 929 AIHSMTSVLETMATEYFQEVYKADPSLNPESVTRLFQEKVTDAMNEKLCQEFKEEEIAQA 988
Query: 896 LLRMRSYSALGPDGFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVD 955
+ ++ + GPDGF F+++ W + D+ VR+ F++G++ + + +T IVLIPK D
Sbjct: 989 IFQIGPLKSPGPDGFPARFYQRNWGTLKSDIILAVRNFFQSGLMPEGVNDTAIVLIPKKD 1048
Query: 956 SPSTVKEFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEI 1015
P +K++RPISLCNVVYK+++K +VNR+RP+L+ ++ Q++F+ G I DNA LA E
Sbjct: 1049 QPIDLKDYRPISLCNVVYKVVSKCLVNRLRPILDDLVSKEQSAFIQGRMITDNALLAFEC 1108
Query: 1016 IHHMGAS-MAKKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNR 1059
H + + A A+K+DL KAYD V W FLE L + GF +R
Sbjct: 1109 FHSIQKNKKANSAACAYKLDLSKAYDRVDWRFLELALNKLGFAHR 1153
Score = 138 bits (348), Expect = 2e-30
Identities = 95/341 (27%), Positives = 149/341 (42%), Gaps = 23/341 (6%)
Query: 1231 KKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSGQ 1290
K++ W N L+ G+ L K+VI AIP Y M +F P S+ + + R F W+ ++G+
Sbjct: 1258 KRVILWGENFLSSGGKEILIKAVIQAIPVYVMGLFKFPESVCDDLTKMTRNFWWVADNGR 1317
Query: 1291 RGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLAHKYLHGE 1350
R H W +LT+ K +GGL +D FN +LL W L + L QVL KY
Sbjct: 1318 RRTHWRAWDSLTKAKINGGLGFRDYKLFNQALLAPQAWRLIEFPNSLCAQVLKAKYFPHG 1377
Query: 1351 SIFSVHRRANASPIWQGILKARDQLQEGFQFRMGDGSS-SVWYHDWTGKGKLAESLDFVH 1409
S+ ANASP W GI D L++G +R+G+G+S +W W K +
Sbjct: 1378 SLTDTTFSANASPTWHGIEYGLDLLKKGIIWRIGNGNSVRIWRDPWIPKDLSRRPVSSKA 1437
Query: 1410 ISDTKLCLHELVDNNQWNLNRCMTVFPLHIRDWFARVEPRVVAN-------GIDRWSWKD 1462
K + ++ W+ + I +F +++ ++ D +W
Sbjct: 1438 NCRLKWVSDLIAEDGTWDSAK--------INQYFLKIDADIIQKICISARLEEDFIAWHP 1489
Query: 1463 NSDGISSVHDGYQWLRE-------KKMAGVVHGDSWLWI*KLHAPEKL*VFIWLVLHDAL 1515
+ G SV Y+ + + SW I K + P+K+ +F W V ++L
Sbjct: 1490 DKTGRFSVRSAYKLALQLADMNNCSSSSSSRLNKSWELIWKCNVPQKVRIFAWRVASNSL 1549
Query: 1516 PTNANRYRCKLAVSPGCTRCSSPEEDTIHCLRDCPHSQELW 1556
T N+ + L C C +ED H L C H+ LW
Sbjct: 1550 ATMENKKKRNLERFDVCGICDREKEDAGHALCCCVHANSLW 1590
>gb|AAK71569.2| putative reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|50918647|ref|XP_469720.1| putative
reverse transcriptase [Oryza sativa (japonica
cultivar-group)]
Length = 1833
Score = 278 bits (711), Expect = 1e-72
Identities = 186/591 (31%), Positives = 290/591 (48%), Gaps = 17/591 (2%)
Query: 483 KPDIMFLLETRCQFSRASNFWNSTRFSPLFISEANGFSGGIWVLVNTATNLSSRLLYS-H 541
+P I+FL ETR N R F G GG+ + + + N+ + S H
Sbjct: 11 RPKIVFLAETRQDQYMVQNRIGLRR---CFTVNGVGKGGGLALFWDESMNVVLKSYNSRH 67
Query: 542 QQAVTFEVWRENLSWVCSAIYASPIPVKREELWSHLLHIRSNLVLPWMIVGDMNEILHPN 601
+ E+ + +W + +Y P R +WS L IRSN PW+++GD NE +
Sbjct: 68 IDVLITEL--DCCTWRATFVYGEPRAQDRHLMWSLLRRIRSNSGDPWLMIGDFNEAMWQT 125
Query: 602 EVRGG*FLPTRASR-FAAILEGCGMVDLGMSGGRYTWFRKQNNRLIMSKRLDRAMGDGAW 660
E + R F +L C + D+G G +T+ Q + RLDR + AW
Sbjct: 126 EHKSHRKRSESQMRDFREVLSECDLHDIGFQGAPWTFCNMQREGRNVKVRLDRGVASPAW 185
Query: 661 RIAFPEASVEALHRIHSDHCPLLINCSGPEIVHHDRPFRFLASWVEHPQYKEVVQSAWT- 719
FP+A + L SDH PLL+ + + R+ W EV+Q AWT
Sbjct: 186 SSRFPQAVITHLTTPSSDHAPLLLEREETTLARPMKIMRYEEVWERESSLPEVIQEAWTM 245
Query: 720 GETETVAGKLNK----VRTKSCKFNKEVYGSIFQRKRWVEGRLRGVQRELNYRVTFDMTR 775
G + G +N TK ++K+ G++ RK+ + LR EL D
Sbjct: 246 GADASTLGDINDKMKVTMTKLVSWSKDKIGNV--RKKIKD--LREKLGELRNIGLLDTDN 301
Query: 776 LETELQAEYRFLLRQEELLWFQKARENRVCFDDRNTAYFHAHTIIRRKRNKIHRLKLNDG 835
++ E +L +EE+ W Q++R + D NT YFH R K+NKI +LK NDG
Sbjct: 302 EVHSVKKELEEMLHREEIWWKQRSRITWLKEGDLNTRYFHLKASWRAKKNKIKKLKKNDG 361
Query: 836 SWCDDRQLLAEEIQNYFQSLFAADPDNTDVNMQHVMMPRLTEAERHKAEAEVIKEEVFQA 895
S +++ + E +++FQ L+ D + VN+ ++ +++E EE+ A
Sbjct: 362 STTMNKKEMKEINRSFFQQLYTKDDNLNPVNLLNMFHEKISEQMNADLIKPFTNEEISDA 421
Query: 896 LLRMRSYSALGPDGFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVD 955
L ++ A GPDGF F ++ W L+ ++ VR+ FE ++ + + +T+IV+IPK +
Sbjct: 422 LFQIGPLKAPGPDGFPARFLQRNWGLLKGEVIAAVRNFFEDEVMQEGVNDTVIVMIPKKN 481
Query: 956 SPSTVKEFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEI 1015
+K+FRPISLCNVVYK++ K +VNR+RP+L II Q++F+PG I DNA +A E
Sbjct: 482 LAEDMKDFRPISLCNVVYKVVAKCLVNRMRPMLQEIISEAQSAFIPGRLITDNALIAFEC 541
Query: 1016 IHHM-GASMAKKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIM 1065
H + + A K+DL KAYD V W FL + + GF ++ + IM
Sbjct: 542 FHSIKNCKRENQNFCALKLDLSKAYDRVDWEFLNGVMERLGFCDKWRRWIM 592
Score = 137 bits (344), Expect = 4e-30
Identities = 99/391 (25%), Positives = 170/391 (43%), Gaps = 19/391 (4%)
Query: 1231 KKLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSGQ 1290
K++S W ++ + L KSV A+PTY M VF + + +R F W ++G+
Sbjct: 686 KRMSDWNEKYMSSGAKEVLIKSVAQALPTYIMGVFKMTDKFCDDYTRMVRNFWWGHDNGE 745
Query: 1291 RGWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQVLAHKYLHGE 1350
R H + W+ + PK HGGL +DM FN +LL + W L + L +V KY
Sbjct: 746 RKVHWIAWEKILYPKSHGGLGFRDMKCFNQALLARQAWRLLTSPDSLCARVFKAKYYPNG 805
Query: 1351 SIFSVHRRANASPIWQGILKARDQLQEGFQFRMGDGSS-SVWYHDWTGKGKLAESLDFVH 1409
++ +S +W+GI D L++G +R+G+G + +W H W G+ L+
Sbjct: 806 NLVDTAFPTVSSSVWKGIQHGLDLLKQGTIWRVGNGHNIKIWRHKWVPHGEHINVLE-KK 864
Query: 1410 ISDTKLCLHELVDNNQWNLNRCMTVFPLHIRDWFARVEPRVVANGIDRW-SWKDNSDGIS 1468
+ + ++EL++ N + L D ++ R+ + +D + +W G+
Sbjct: 865 GRNRLIYVNELINEENRCWNETLVRHVLKEEDANEVLKIRLPNHQMDDFPAWHHEKSGLF 924
Query: 1469 SVHDGYQWLREKKMAGVVHGDS----------WLWI*KLHAPEKL*VFIWLVLHDALPTN 1518
+V Y+ GVV S W + K+ +FIW + D LPT
Sbjct: 925 TVKSAYKLAWNLSGKGVVQSSSSTATSGERKIWSRVWNAKVQAKVKIFIWKLAQDKLPTW 984
Query: 1519 ANRYRCKLAVSPGCTRCSSPEEDTIHCLRDCPHSQELWGKLGA-W---GWSTFRL--PDL 1572
N+ R K+ ++ C C + E++ H +C ++ L L A W G F PD
Sbjct: 985 ENKRRRKIEMNGTCPVCGTKGENSYHATVECTKARALREALRAVWHLPGEDKFLWTGPDW 1044
Query: 1573 KA*VKTQLQGSNNIRFLAGLWGVWKWRNNVV 1603
+ + + LW W RN+++
Sbjct: 1045 LLILLDGVNEEQRTHIMYMLWRAWYLRNDLI 1075
>ref|XP_475290.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|49328177|gb|AAT58873.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 1264
Score = 276 bits (707), Expect = 4e-72
Identities = 172/506 (33%), Positives = 263/506 (50%), Gaps = 20/506 (3%)
Query: 614 SRFAAILEGCGMVDLGMSGGRYTWFRKQNNRLIMS----KRLDRAMGDGAWRIAFPEASV 669
++F LE C + DLG G +TW +N+ + S +RLDRA+ +GAWR FP V
Sbjct: 498 AKFRQALEDCQLHDLGFVGDAFTW---RNHHHLASNYIKERLDRAVANGAWRARFPLVRV 554
Query: 670 EALHRIHSDHCPLLINCSGPEIVHHDRPF----RFLASWVEHPQYKEVVQSAWTGETETV 725
HSDH +++ E +P +F A W+E + + V+ AW E
Sbjct: 555 INGDPRHSDHRSVIVETGATEKQQWGQPLEIMQKFEARWLEEEECQARVEEAWENALEGG 614
Query: 726 AGKLNKVRTKSCK----FNKEVYGSIFQRKRWVEGRLRGVQRELNYRVTFDMTRLETELQ 781
+L +++++ K +++ V G + +R + + L +RE ++ E L+
Sbjct: 615 QTRLMEIQSRVLKELWAWDRTVLGELKKRVKNLRKELEKCRRE---PISNRQVNREHLLR 671
Query: 782 AEYRFLLRQEELLWFQKARENRVCFDDRNTAYFHAHTIIRRKRNKIHRLKLNDGSWCDDR 841
+ LL Q+ + W Q+A + DRNT +FHA ++KRN I +L+ G
Sbjct: 672 YKLERLLDQQHIYWKQRAHSTWLTKGDRNTKFFHAQASEKKKRNTIQKLQDGHGGLVAGN 731
Query: 842 QLLAEEIQNYFQSLFAADPDNTDVNMQHVMMPRLTEAERHKAEAEVIKEEVFQALLRMRS 901
QL + I N +Q LF ++ + + + R+T R A +EEV+ AL M
Sbjct: 732 QLKSF-ISNQYQQLFRSNGCSQMDAVLQCVQARVTPEMREGLAAPYQREEVWVALKDMGD 790
Query: 902 YSALGPDGFHPFFFKKYWELVGDDLWKLVRDAFETGMIDDSLLETLIVLIPKVDSPSTVK 961
A G DG F+KK+ L GD + V G + +T++VLIPK P T+K
Sbjct: 791 LKAPGADGIPAIFYKKFLSLAGDKVKDEVLAVLNGGDMPQGWNDTVVVLIPKTKQPDTLK 850
Query: 962 EFRPISLCNVVYKLITKVVVNRIRPLLNRIIGPMQNSFLPGMGIMDNAFLAQEIIHHMG- 1020
+ RPISLCNV YKLI+KV+VNR++ +L II P Q++F+P I N LA E+ H++
Sbjct: 851 DLRPISLCNVAYKLISKVIVNRLKVVLLEIISPSQSAFVPRRLITYNVLLAYELTHYLNQ 910
Query: 1021 ASMAKKGTLAFKIDLEKAYDSVSWAFLEDTLVQFGFPNRLVQLIMRCVSGSNLSMLWNGS 1080
K G A K+D+ KAYD V W FL +++ GF ++ V L+M+CV+ + NG
Sbjct: 911 RKKGKNGVAAIKLDMSKAYDRVEWDFLRHMMLRLGFHDQWVNLVMKCVTSVTYRIKINGE 970
Query: 1081 RLPSFKPGRGLRKGDPLSPYLFVLCS 1106
P RGLR+GDPLSPYLF++C+
Sbjct: 971 HSDQIYPQRGLRQGDPLSPYLFIICA 996
Score = 74.3 bits (181), Expect = 4e-11
Identities = 36/110 (32%), Positives = 60/110 (53%)
Query: 1232 KLSSWKTNLLNMVGQVCLAKSVISAIPTYTMQVFWLPRSINHKINQEMRRFIWLKNSGQR 1291
++ W+ LL+ G+ L K+V AIPTY M F L + + +IN + ++ W +N +
Sbjct: 1123 RIQGWQEKLLSKAGKEILVKAVAQAIPTYAMSCFDLTKGLCDEINSMISKWWWSQNDKEN 1182
Query: 1292 GWHRVNWKTLTQPKEHGGLAIKDMGDFNTSLLGKVVWLLANNSSKLWVQV 1341
H ++W+ +T PK+ GG +D+ FN ++L + W L N L QV
Sbjct: 1183 KIHWLSWEKMTLPKKLGGPGFRDLHLFNMAMLARQAWRLLLNVDSLCGQV 1232
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.333 0.144 0.485
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,159,001,482
Number of Sequences: 2540612
Number of extensions: 134557497
Number of successful extensions: 306109
Number of sequences better than 10.0: 1215
Number of HSP's better than 10.0 without gapping: 766
Number of HSP's successfully gapped in prelim test: 449
Number of HSP's that attempted gapping in prelim test: 302450
Number of HSP's gapped (non-prelim): 2258
length of query: 1813
length of database: 863,360,394
effective HSP length: 143
effective length of query: 1670
effective length of database: 500,052,878
effective search space: 835088306260
effective search space used: 835088306260
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 83 (36.6 bits)
Lotus: description of TM0178.7


