

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
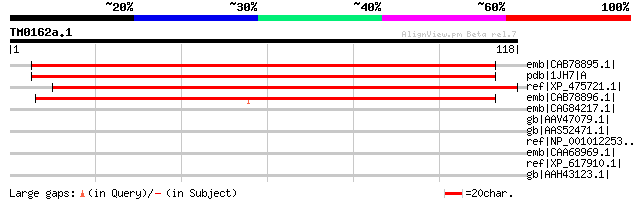
Query= TM0162a.1
(118 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB78895.1| putative protein [Arabidopsis thaliana] gi|28326... 131 3e-30
pdb|1JH7|A Chain A, Semi-Reduced Inhibitor-Bound Cyclic Nucleoti... 131 3e-30
ref|XP_475721.1| unknown protein [Oryza sativa (japonica cultiva... 125 3e-28
emb|CAB78896.1| hypothetical protein [Arabidopsis thaliana] gi|2... 105 2e-22
emb|CAG84217.1| unnamed protein product [Yarrowia lipolytica CLI... 44 0.001
gb|AAV47079.1| probable cysteine desulfurase [Haloarcula marismo... 34 0.72
gb|AAS52471.1| AEL214Cp [Ashbya gossypii ATCC 10895] gi|45190393... 33 1.6
ref|NP_001012253.1| XK-related protein 7b [Danio rerio] gi|46948... 31 6.1
emb|CAA68969.1| unnamed protein product [Saccharomyces cerevisia... 31 8.0
ref|XP_617910.1| PREDICTED: similar to Kelch repeat and BTB doma... 31 8.0
gb|AAH43123.1| Ank2 protein [Mus musculus] 31 8.0
>emb|CAB78895.1| putative protein [Arabidopsis thaliana] gi|2832621|emb|CAA16750.1|
putative protein [Arabidopsis thaliana]
gi|2065013|emb|CAA72363.1| cyclic phosphodiesterase
[Arabidopsis thaliana] gi|20148603|gb|AAM10192.1|
protein kinase-like protein [Arabidopsis thaliana]
gi|17065492|gb|AAL32900.1| protein kinase - like protein
[Arabidopsis thaliana] gi|15234068|ref|NP_193628.1|
cyclic phosphodiesterase [Arabidopsis thaliana]
gi|7484882|pir||T05030 cyclic phosphodiesterase (EC
3.1.4.-) - Arabidopsis thaliana
gi|51315738|sp|O04147|CPD_ARATH Cyclic phosphodiesterase
(CPDase)
Length = 181
Score = 131 bits (330), Expect = 3e-30
Identities = 61/108 (56%), Positives = 75/108 (68%)
Query: 6 IETQKKEVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKL 65
+E KK+VYSVWA+P + PR KLM LRSEF GP F PH+TV + LTAD+A
Sbjct: 1 MEEVKKDVYSVWALPDEESEPRFKKLMEALRSEFTGPRFVPHVTVAVSAYLTADEAKKMF 60
Query: 66 RSACEELKAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHF 113
SAC+ LKA+ TVDRV+ GTFF+QCV+LLL +P V+E HC NHF
Sbjct: 61 ESACDGLKAYTATVDRVSTGTFFFQCVFLLLQTTPEVMEAGEHCKNHF 108
>pdb|1JH7|A Chain A, Semi-Reduced Inhibitor-Bound Cyclic Nucleotide
Phosphodiesterase From Arabidopsis Thaliana
gi|18655430|pdb|1JH6|B Chain B, Semi-Reduced Cyclic
Nucleotide Phosphodiesterase From Arabidopsis Thaliana
gi|18655429|pdb|1JH6|A Chain A, Semi-Reduced Cyclic
Nucleotide Phosphodiesterase From Arabidopsis Thaliana
gi|11513546|pdb|1FSI|C Chain C, Crystal Structure Of
Cyclic Nucleotide Phosphodiesterase Of Appr>p From
Arabidopsis Thaliana gi|11513545|pdb|1FSI|B Chain B,
Crystal Structure Of Cyclic Nucleotide Phosphodiesterase
Of Appr>p From Arabidopsis Thaliana
gi|11513544|pdb|1FSI|A Chain A, Crystal Structure Of
Cyclic Nucleotide Phosphodiesterase Of Appr>p From
Arabidopsis Thaliana
Length = 189
Score = 131 bits (330), Expect = 3e-30
Identities = 61/108 (56%), Positives = 75/108 (68%)
Query: 6 IETQKKEVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKL 65
+E KK+VYSVWA+P + PR KLM LRSEF GP F PH+TV + LTAD+A
Sbjct: 1 MEEVKKDVYSVWALPDEESEPRFKKLMEALRSEFTGPRFVPHVTVAVSAYLTADEAKKMF 60
Query: 66 RSACEELKAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHF 113
SAC+ LKA+ TVDRV+ GTFF+QCV+LLL +P V+E HC NHF
Sbjct: 61 ESACDGLKAYTATVDRVSTGTFFFQCVFLLLQTTPEVMEAGEHCKNHF 108
>ref|XP_475721.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|46575962|gb|AAT01323.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 185
Score = 125 bits (313), Expect = 3e-28
Identities = 60/108 (55%), Positives = 73/108 (67%)
Query: 11 KEVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSACE 70
KEVYSVWA+PP+ VR R+ +M LR+ GGP FEPH TVVGAI L A+ LR+A
Sbjct: 9 KEVYSVWALPPEPVRARLRGVMAGLRAAHGGPAFEPHATVVGAIRLRRSAAVEALRAAAA 68
Query: 71 ELKAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHFGYKSS 118
++ + V VA G FFYQC+YLLL P+P VVE S HC HFGY+ S
Sbjct: 69 GVRPYTARVVGVARGDFFYQCIYLLLEPTPEVVEASDHCCGHFGYERS 116
>emb|CAB78896.1| hypothetical protein [Arabidopsis thaliana]
gi|2832622|emb|CAA16751.1| hypothetical protein
[Arabidopsis thaliana] gi|22136634|gb|AAM91636.1|
unknown protein [Arabidopsis thaliana]
gi|15234071|ref|NP_193629.1| cyclic phosphodiesterase,
putative [Arabidopsis thaliana] gi|7484883|pir||T05031
cyclic phosphodiesterase homolog F13C5.110 - Arabidopsis
thaliana
Length = 196
Score = 105 bits (263), Expect = 2e-22
Identities = 48/109 (44%), Positives = 72/109 (66%), Gaps = 2/109 (1%)
Query: 7 ETQKKEVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAI--TLTADDALNK 64
+ ++K +Y++WA+P D V R+ +LM LRSEFGGP F+PH+T+VG LTA +A
Sbjct: 18 DEEEKAMYAIWAVPEDDVEDRLQRLMEGLRSEFGGPAFDPHLTLVGPFPYKLTASEAKRM 77
Query: 65 LRSACEELKAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHF 113
+SACE K + TVD+V+AGT ++QC+Y+ L + V+ + H HF
Sbjct: 78 FKSACEGFKVYPATVDQVSAGTSYFQCLYVSLRHTVEVMNAAGHFMAHF 126
>emb|CAG84217.1| unnamed protein product [Yarrowia lipolytica CLIB99]
gi|50545483|ref|XP_500279.1| hypothetical protein
[Yarrowia lipolytica]
Length = 219
Score = 43.5 bits (101), Expect = 0.001
Identities = 30/109 (27%), Positives = 57/109 (51%), Gaps = 8/109 (7%)
Query: 15 SVWAIPPDH--VRPRIAKLMTDLRSEFG-GPHFEPHITVVGAITLTA----DDALNKLRS 67
S+W PP + + ++A + L+ F +FEPHIT+ I++ D L++ +
Sbjct: 7 SLWLQPPRNSPIWQQLATTIQGLKPIFSDSENFEPHITLTSNISVNTQGQVDFVLDRAVA 66
Query: 68 ACEEL-KAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHFGY 115
A + + + FQ+ + V G+ F++ VYL + P+P ++ + C F Y
Sbjct: 67 AAKCVPQGFQIHLSSVKYGSRFFKKVYLQVEPTPELLSLARICREDFVY 115
>gb|AAV47079.1| probable cysteine desulfurase [Haloarcula marismortui ATCC 43049]
gi|55378935|ref|YP_136785.1| probable cysteine
desulfurase [Haloarcula marismortui ATCC 43049]
Length = 367
Score = 34.3 bits (77), Expect = 0.72
Identities = 26/99 (26%), Positives = 42/99 (42%), Gaps = 19/99 (19%)
Query: 13 VYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSACEEL 72
+ +V AI D ++PR+ +L L+ G P G +T TADD
Sbjct: 276 IETVEAIGFDTIQPRVERLTDRLKDGLGDRLLSPRDYESGLVTFTADDP----------- 324
Query: 73 KAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSN 111
+ TV+R+AA + + + P+P + S H N
Sbjct: 325 ---EATVERLAA-----EGIIIRSLPNPDALRASVHAFN 355
>gb|AAS52471.1| AEL214Cp [Ashbya gossypii ATCC 10895]
gi|45190393|ref|NP_984647.1| AEL214Cp [Eremothecium
gossypii]
Length = 216
Score = 33.1 bits (74), Expect = 1.6
Identities = 27/94 (28%), Positives = 44/94 (46%), Gaps = 14/94 (14%)
Query: 15 SVWAIPPDH--VRPRIAKLMTDLRSEF-GGPHFEPHITVVGAITL-TADDALNKLRSACE 70
++W PP + + L+ L++ F FEPH+T+ + +ADDA L +AC
Sbjct: 4 ALWYCPPANSPCYDTVNSLILSLQTLFPDAVLFEPHVTITSQLRCDSADDAAAVLAAACA 63
Query: 71 ELKAFQ----------VTVDRVAAGTFFYQCVYL 94
L+A + V D VA G ++ V+L
Sbjct: 64 ALRACRPQLDARGSPVVRFDAVAVGRRYFDKVHL 97
>ref|NP_001012253.1| XK-related protein 7b [Danio rerio] gi|46948433|gb|AAT07128.1|
XK-related protein 7b [Danio rerio]
Length = 404
Score = 31.2 bits (69), Expect = 6.1
Identities = 15/33 (45%), Positives = 17/33 (51%)
Query: 77 VTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHC 109
V V A GTFF Y LLHP P++ T C
Sbjct: 238 VVVCSYALGTFFMFVYYCLLHPDGPILGTQWSC 270
>emb|CAA68969.1| unnamed protein product [Saccharomyces cerevisiae]
gi|6321686|ref|NP_011763.1| Cyclic nucleotide
phosphodiesterase, hydrolyzes ADP-ribose 1'',
2''-cyclic phosphate to ADP-ribose 1''-phosphate; no
detectable phenotype is conferred by null mutation or
by overexpression; Cpd1p [Saccharomyces cerevisiae]
gi|1323448|emb|CAA97276.1| unnamed protein product
[Saccharomyces cerevisiae]
gi|1723762|sp|P53314|YG59_YEAST Hypothetical 26.7 kDa
protein in TDS4-SOL4 intergenic region
gi|45270652|gb|AAS56707.1| YGR247W [Saccharomyces
cerevisiae]
Length = 239
Score = 30.8 bits (68), Expect = 8.0
Identities = 11/28 (39%), Positives = 18/28 (64%)
Query: 42 PHFEPHITVVGAITLTADDALNKLRSAC 69
P FEPH+TV + + D +NK+ ++C
Sbjct: 34 PVFEPHVTVTSHLVCNSKDDVNKILTSC 61
>ref|XP_617910.1| PREDICTED: similar to Kelch repeat and BTB domain containing
protein 9, partial [Bos taurus]
Length = 309
Score = 30.8 bits (68), Expect = 8.0
Identities = 24/78 (30%), Positives = 31/78 (38%), Gaps = 8/78 (10%)
Query: 27 RIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSACEELKAFQVTVDRVAAGT 86
R K TDL+ G FE H V+ + +L D + + AF VD AG
Sbjct: 86 RRVKAFTDLKIVVEGREFEVHQNVLASCSLYFKDLIQRF--------AFLPAVDGTLAGI 137
Query: 87 FFYQCVYLLLHPSPPVVE 104
F C L L P V+
Sbjct: 138 SFPICCVLSLRAELPGVQ 155
>gb|AAH43123.1| Ank2 protein [Mus musculus]
Length = 1038
Score = 30.8 bits (68), Expect = 8.0
Identities = 25/86 (29%), Positives = 38/86 (44%), Gaps = 5/86 (5%)
Query: 6 IETQKKEVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAIT-----LTADD 60
+ET+ ++ ++WA D +A +D S G + IT A + DD
Sbjct: 29 LETESEQKSALWAAQSDAPPLAVAPTASDAASVTGEQASKVIITKTDADADSWSEIREDD 88
Query: 61 ALNKLRSACEELKAFQVTVDRVAAGT 86
A + R EE K F + VDR + GT
Sbjct: 89 AAFEARVKEEEQKIFGLMVDRQSQGT 114
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.134 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 204,269,570
Number of Sequences: 2540612
Number of extensions: 6955002
Number of successful extensions: 15426
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 15417
Number of HSP's gapped (non-prelim): 11
length of query: 118
length of database: 863,360,394
effective HSP length: 94
effective length of query: 24
effective length of database: 624,542,866
effective search space: 14989028784
effective search space used: 14989028784
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0162a.1


