

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0154.8
(247 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
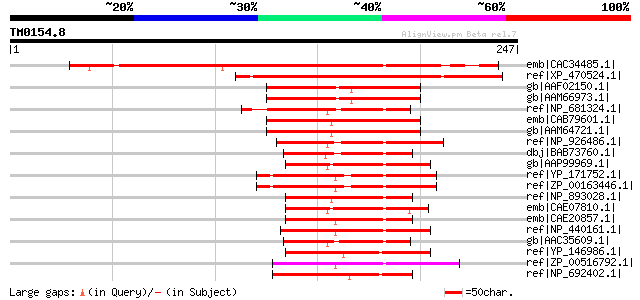
Score E
Sequences producing significant alignments: (bits) Value
emb|CAC34485.1| putative protein [Arabidopsis thaliana] gi|29294... 220 2e-56
ref|XP_470524.1| Hypothetical protein [Oryza sativa (japonica cu... 181 2e-44
gb|AAF02150.1| unknown protein [Arabidopsis thaliana] gi|3002367... 80 5e-14
gb|AAM66973.1| unknown [Arabidopsis thaliana] 80 5e-14
ref|NP_681324.1| hypothetical protein tsr0534 [Thermosynechococc... 80 5e-14
emb|CAB79601.1| putative protein [Arabidopsis thaliana] gi|44553... 79 1e-13
gb|AAM64721.1| unknown [Arabidopsis thaliana] 79 1e-13
ref|NP_926486.1| hypothetical protein gvip478 [Gloeobacter viola... 79 1e-13
dbj|BAB73760.1| asl2061 [Nostoc sp. PCC 7120] gi|17229553|ref|NP... 77 6e-13
gb|AAP99969.1| Uncharacterized YGGT family conserved membrane pr... 76 1e-12
ref|YP_171752.1| hypothetical protein YCF19 [Synechococcus elong... 75 2e-12
ref|ZP_00163446.1| COG0762: Predicted integral membrane protein ... 75 2e-12
ref|NP_893028.1| conserved hypothetical membrane protein [Prochl... 72 1e-11
emb|CAE07810.1| conserved hypothetical membrane protein [Synecho... 72 1e-11
emb|CAE20857.1| conserved hypothetical membrane protein [Prochlo... 71 3e-11
ref|NP_440161.1| hypothetical protein ssr2142 [Synechocystis sp.... 69 9e-11
gb|AAC35609.1| hypothetical chloroplast RF19 [Guillardia theta] ... 64 4e-09
ref|YP_146986.1| hypothetical protein GK1133 [Geobacillus kausto... 62 1e-08
ref|ZP_00516792.1| Protein of unknown function YGGT [Crocosphaer... 62 2e-08
ref|NP_692402.1| hypothetical protein OB1481 [Oceanobacillus ihe... 60 7e-08
>emb|CAC34485.1| putative protein [Arabidopsis thaliana] gi|29294067|gb|AAO73904.1|
expressed protein [Arabidopsis thaliana]
gi|26451921|dbj|BAC43053.1| unknown protein [Arabidopsis
thaliana] gi|28950811|gb|AAO63329.1| At5g21920
[Arabidopsis thaliana] gi|22326932|ref|NP_680180.1| YGGT
family protein [Arabidopsis thaliana]
Length = 251
Score = 220 bits (561), Expect = 2e-56
Identities = 123/215 (57%), Positives = 150/215 (69%), Gaps = 19/215 (8%)
Query: 30 TQTPFALP-----ILKPPISNLTSFLSPPELHKSFTTATDNCFRFLHSLASQNPFLNKIL 84
TQTP +L PP+++ + + H+S +AT+ FLHSLAS+NP + +
Sbjct: 33 TQTPLVRSNKPNLLLLPPVADSVKLIQ--DFHQSLISATEKFKGFLHSLASKNPLFQEAV 90
Query: 85 SLRSEFDSLCFQIRCSNY-RSRRWLSSHNFAAVLPGDSVAGLVVGNGIQNFLNIYNTLLV 143
L SEF LC +IR N R R +S+H FAAVLPGDSVAGLVV NG+ NFLNIYNT+LV
Sbjct: 91 RLSSEFHILCDEIRLRNTTRVRFAMSNHGFAAVLPGDSVAGLVVANGLINFLNIYNTILV 150
Query: 144 VRLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAFLVLNAFTSASA 203
VRLVLTWFP++PPAIV PLST+CDPYLN+FRG IPPLGG LDLSPILAFLVLNAFTS++
Sbjct: 151 VRLVLTWFPSAPPAIVNPLSTLCDPYLNIFRGFIPPLGG-LDLSPILAFLVLNAFTSSAM 209
Query: 204 ALPAELPVTQQSEQGVAATLQSTDITSSQKKWMKR 238
ALP ELP S G + SS+ KW++R
Sbjct: 210 ALPCELP----SADGAVSP------ASSETKWVRR 234
>ref|XP_470524.1| Hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|27436744|gb|AAO13463.1| Hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 157
Score = 181 bits (458), Expect = 2e-44
Identities = 91/130 (70%), Positives = 110/130 (84%), Gaps = 2/130 (1%)
Query: 111 HNFAAVLPGDSVAGLVVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTICDPYL 170
H FAAVL GDSVAG+VV +GI NFL++YNT+LVVRLVLTWFPN+PPAIV PLSTICDPYL
Sbjct: 19 HCFAAVL-GDSVAGVVVSSGINNFLSLYNTVLVVRLVLTWFPNTPPAIVAPLSTICDPYL 77
Query: 171 NLFRGLIPPLGGTLDLSPILAFLVLNAFTSASAALPAELPVTQQSEQGVAATLQSTDITS 230
N+FRG+IPPLGGTLDLSPILAFLVLNA +S +AALPAELP AT S+ +T+
Sbjct: 78 NIFRGIIPPLGGTLDLSPILAFLVLNALSSTAAALPAELP-DPAPPTSRGATSSSSVLTA 136
Query: 231 SQKKWMKRLQ 240
+++KWM+R++
Sbjct: 137 NRRKWMRRIR 146
>gb|AAF02150.1| unknown protein [Arabidopsis thaliana] gi|30023676|gb|AAP13371.1|
At3g07430 [Arabidopsis thaliana]
gi|20466762|gb|AAM20698.1| unknown protein [Arabidopsis
thaliana] gi|18397948|ref|NP_566307.1| YGGT family
protein [Arabidopsis thaliana]
Length = 232
Score = 80.1 bits (196), Expect = 5e-14
Identities = 43/78 (55%), Positives = 56/78 (71%), Gaps = 4/78 (5%)
Query: 126 VVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTI---CDPYLNLFRGLIPPLGG 182
VV GI+ +L+IY+ +L+VR++L+WFPN P PLS I CDPYLNLFR +IPP+
Sbjct: 149 VVAVGIKKWLDIYSGVLMVRVLLSWFPNIPWERQ-PLSAIRDLCDPYLNLFRNIIPPIFD 207
Query: 183 TLDLSPILAFLVLNAFTS 200
TLD+SP+LAF VL S
Sbjct: 208 TLDVSPLLAFAVLGTLGS 225
>gb|AAM66973.1| unknown [Arabidopsis thaliana]
Length = 234
Score = 80.1 bits (196), Expect = 5e-14
Identities = 43/78 (55%), Positives = 56/78 (71%), Gaps = 4/78 (5%)
Query: 126 VVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTI---CDPYLNLFRGLIPPLGG 182
VV GI+ +L+IY+ +L+VR++L+WFPN P PLS I CDPYLNLFR +IPP+
Sbjct: 151 VVAVGIKKWLDIYSGVLMVRVLLSWFPNIPWERQ-PLSAIRDLCDPYLNLFRNIIPPIFD 209
Query: 183 TLDLSPILAFLVLNAFTS 200
TLD+SP+LAF VL S
Sbjct: 210 TLDVSPLLAFAVLGTLGS 227
>ref|NP_681324.1| hypothetical protein tsr0534 [Thermosynechococcus elongatus BP-1]
gi|22294255|dbj|BAC08086.1| ycf19 [Thermosynechococcus
elongatus BP-1]
Length = 96
Score = 80.1 bits (196), Expect = 5e-14
Identities = 45/86 (52%), Positives = 60/86 (69%), Gaps = 14/86 (16%)
Query: 114 AAVLPGDSVAGLVVGNGIQNFLNIYNTLLVVRLVLTWFPN----SPPAIVGPLSTICDPY 169
AAVLP ++ GI NFL IY +L++R++L+WFPN +PP + LS + DPY
Sbjct: 4 AAVLP-------ILLMGIANFLQIYLIILLIRVLLSWFPNINWYNPPFSI--LSQLTDPY 54
Query: 170 LNLFRGLIPPLGGTLDLSPILAFLVL 195
LN+FRGLIPP+GG LD SPI+AF +L
Sbjct: 55 LNIFRGLIPPIGG-LDFSPIIAFFLL 79
>emb|CAB79601.1| putative protein [Arabidopsis thaliana] gi|4455358|emb|CAB36768.1|
putative protein [Arabidopsis thaliana]
gi|15234345|ref|NP_194528.1| YGGT family protein
[Arabidopsis thaliana] gi|7486950|pir||T02900
hypothetical protein T13J8.100 - Arabidopsis thaliana
Length = 218
Score = 79.0 bits (193), Expect = 1e-13
Identities = 38/77 (49%), Positives = 54/77 (69%), Gaps = 2/77 (2%)
Query: 126 VVGNGIQNFLNIYNTLLVVRLVLTWFPNSP--PAIVGPLSTICDPYLNLFRGLIPPLGGT 183
VV G+ +L+IY+ +L+VR++L+WFPN P + + +CDPYLNLFR +IPP+ T
Sbjct: 135 VVAAGLSKWLDIYSGVLMVRVLLSWFPNIPWDRQPLSAIRDLCDPYLNLFRNIIPPVFDT 194
Query: 184 LDLSPILAFLVLNAFTS 200
LD+SP+LAF VL S
Sbjct: 195 LDVSPLLAFAVLGTLGS 211
>gb|AAM64721.1| unknown [Arabidopsis thaliana]
Length = 218
Score = 79.0 bits (193), Expect = 1e-13
Identities = 38/77 (49%), Positives = 54/77 (69%), Gaps = 2/77 (2%)
Query: 126 VVGNGIQNFLNIYNTLLVVRLVLTWFPNSP--PAIVGPLSTICDPYLNLFRGLIPPLGGT 183
VV G+ +L+IY+ +L+VR++L+WFPN P + + +CDPYLNLFR +IPP+ T
Sbjct: 135 VVAAGLSKWLDIYSGVLMVRVLLSWFPNIPWDGQPLSAIRDLCDPYLNLFRNIIPPVFDT 194
Query: 184 LDLSPILAFLVLNAFTS 200
LD+SP+LAF VL S
Sbjct: 195 LDVSPLLAFAVLGTLGS 211
>ref|NP_926486.1| hypothetical protein gvip478 [Gloeobacter violaceus PCC 7421]
gi|35214112|dbj|BAC91481.1| ycf19 [Gloeobacter violaceus
PCC 7421]
Length = 95
Score = 78.6 bits (192), Expect = 1e-13
Identities = 46/86 (53%), Positives = 56/86 (64%), Gaps = 9/86 (10%)
Query: 131 IQNFLNIYNTLLVVRLVLTWFPN-----SPPAIVGPLSTICDPYLNLFRGLIPPLGGTLD 185
+ NF+ IY LL VR++L+WFPN +P AI LS + DPYLNLFR +IPPLGG +D
Sbjct: 14 LSNFIQIYVVLLFVRVLLSWFPNIDWSSNPWAI---LSQLTDPYLNLFRSIIPPLGG-ID 69
Query: 186 LSPILAFLVLNAFTSASAALPAELPV 211
LSPILAFL L + A LPV
Sbjct: 70 LSPILAFLALQVVGGLLVSGLASLPV 95
>dbj|BAB73760.1| asl2061 [Nostoc sp. PCC 7120] gi|17229553|ref|NP_486101.1|
hypothetical protein asl2061 [Nostoc sp. PCC 7120]
gi|25305475|pir||AG2063 hypothetical protein asl2061
[imported] - Nostoc sp. (strain PCC 7120)
gi|53763393|ref|ZP_00351282.1| COG0762: Predicted
integral membrane protein [Anabaena variabilis ATCC
29413]
Length = 92
Score = 76.6 bits (187), Expect = 6e-13
Identities = 42/68 (61%), Positives = 50/68 (72%), Gaps = 9/68 (13%)
Query: 134 FLNIYNTLLVVRLVLTWFP-----NSPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSP 188
F+ IY+ LL++R++LTWFP N P A LS I DPYLNLFR +IPPLGG +D SP
Sbjct: 11 FVTIYSYLLIIRVLLTWFPQIDWYNQPFAA---LSQITDPYLNLFRSIIPPLGG-MDFSP 66
Query: 189 ILAFLVLN 196
ILAFLVLN
Sbjct: 67 ILAFLVLN 74
>gb|AAP99969.1| Uncharacterized YGGT family conserved membrane protein
[Prochlorococcus marinus subsp. marinus str. CCMP1375]
gi|33240375|ref|NP_875317.1| Uncharacterized YGGT family
conserved membrane protein [Prochlorococcus marinus
subsp. marinus str. CCMP1375]
Length = 101
Score = 75.9 bits (185), Expect = 1e-12
Identities = 42/74 (56%), Positives = 55/74 (73%), Gaps = 5/74 (6%)
Query: 135 LNIYNTLLVVRLVLTWFPN---SPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILA 191
L IY+ +L++R++LTWFPN S P I+ +S I DPYLNLFRG+IP +GG LD+SPILA
Sbjct: 16 LFIYSYILILRVLLTWFPNLDWSNP-ILSNISAITDPYLNLFRGIIPAIGG-LDISPILA 73
Query: 192 FLVLNAFTSASAAL 205
F+VLN S + L
Sbjct: 74 FIVLNLAESVLSNL 87
>ref|YP_171752.1| hypothetical protein YCF19 [Synechococcus elongatus PCC 6301]
gi|56686010|dbj|BAD79232.1| hypothetical protein YCF19
[Synechococcus elongatus PCC 6301]
Length = 99
Score = 74.7 bits (182), Expect = 2e-12
Identities = 45/92 (48%), Positives = 60/92 (64%), Gaps = 9/92 (9%)
Query: 121 SVAGLVVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPA----IVGPLSTICDPYLNLFRGL 176
S GL++ N + LNIY LL++R++L+WFPN + I+G L+ DPYLNLFR +
Sbjct: 4 SSLGLLL-NTLSITLNIYFVLLIIRVLLSWFPNFQSSQFMLILGQLT---DPYLNLFRRV 59
Query: 177 IPPLGGTLDLSPILAFLVLNAFTSASAALPAE 208
IPPLGG +D SPILAF +L A T A L +
Sbjct: 60 IPPLGG-MDFSPILAFFILQAVTQAVQQLAVQ 90
>ref|ZP_00163446.1| COG0762: Predicted integral membrane protein [Synechococcus
elongatus PCC 7942]
Length = 98
Score = 74.7 bits (182), Expect = 2e-12
Identities = 45/92 (48%), Positives = 60/92 (64%), Gaps = 9/92 (9%)
Query: 121 SVAGLVVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPA----IVGPLSTICDPYLNLFRGL 176
S GL++ N + LNIY LL++R++L+WFPN + I+G L+ DPYLNLFR +
Sbjct: 3 SSLGLLL-NTLSITLNIYFVLLIIRVLLSWFPNFQSSQFMLILGQLT---DPYLNLFRRV 58
Query: 177 IPPLGGTLDLSPILAFLVLNAFTSASAALPAE 208
IPPLGG +D SPILAF +L A T A L +
Sbjct: 59 IPPLGG-MDFSPILAFFILQAVTQAVQQLAVQ 89
>ref|NP_893028.1| conserved hypothetical membrane protein [Prochlorococcus marinus
subsp. pastoris str. CCMP1986]
gi|33634044|emb|CAE19369.1| conserved hypothetical
membrane protein [Prochlorococcus marinus subsp.
pastoris str. CCMP1986]
Length = 92
Score = 72.4 bits (176), Expect = 1e-11
Identities = 34/64 (53%), Positives = 50/64 (78%), Gaps = 3/64 (4%)
Query: 135 LNIYNTLLVVRLVLTWFPNSP--PAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAF 192
L+IY+ +L++R++LTWFP I+ L++I DPYLN+FRG+IPP+GG D+S +LAF
Sbjct: 13 LSIYSFILIIRILLTWFPGIDWSNGILSALTSITDPYLNIFRGIIPPIGG-FDISSLLAF 71
Query: 193 LVLN 196
L+LN
Sbjct: 72 LLLN 75
>emb|CAE07810.1| conserved hypothetical membrane protein [Synechococcus sp. WH 8102]
gi|33865829|ref|NP_897388.1| conserved hypothetical
membrane protein [Synechococcus sp. WH 8102]
Length = 99
Score = 72.0 bits (175), Expect = 1e-11
Identities = 41/76 (53%), Positives = 56/76 (72%), Gaps = 8/76 (10%)
Query: 135 LNIYNTLLVVRLVLTWFPN---SPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILA 191
+NIY +L VR++LTWFPN S P ++G +++I DPYLN+FRG+IPP+GG +DLS ILA
Sbjct: 17 VNIYLFVLFVRVLLTWFPNIDFSNP-VLGGVASITDPYLNMFRGVIPPIGG-IDLSAILA 74
Query: 192 FL---VLNAFTSASAA 204
F+ VL AS+A
Sbjct: 75 FIALRVLQGLLEASSA 90
>emb|CAE20857.1| conserved hypothetical membrane protein [Prochlorococcus marinus
str. MIT 9313] gi|33862954|ref|NP_894514.1| conserved
hypothetical membrane protein [Prochlorococcus marinus
str. MIT 9313]
Length = 99
Score = 70.9 bits (172), Expect = 3e-11
Identities = 37/64 (57%), Positives = 49/64 (75%), Gaps = 3/64 (4%)
Query: 135 LNIYNTLLVVRLVLTWFPNSPPA--IVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAF 192
L IY+ +L+VR++L+WFPN A ++ +S+I DPYLN FRGLIPPLGG LDLS ILA
Sbjct: 15 LEIYSLVLIVRVLLSWFPNLDWANPVLSTVSSITDPYLNAFRGLIPPLGG-LDLSAILAL 73
Query: 193 LVLN 196
+ L+
Sbjct: 74 VALS 77
>ref|NP_440161.1| hypothetical protein ssr2142 [Synechocystis sp. PCC 6803]
gi|1651915|dbj|BAA16841.1| ycf19 [Synechocystis sp. PCC
6803] gi|7429899|pir||S74690 conserved hypothetical
protein ssr2142 [imported] - Synechocystis sp. (strain
PCC 6803)
Length = 90
Score = 69.3 bits (168), Expect = 9e-11
Identities = 37/75 (49%), Positives = 50/75 (66%), Gaps = 3/75 (4%)
Query: 133 NFLNIYNTLLVVRLVLTWFPNSPPA--IVGPLSTICDPYLNLFRGLIPPLGGTLDLSPIL 190
+F+NIY LL VR++L+WF + A I+G LS + DPYLN+FR IPPLGG +D SPIL
Sbjct: 10 SFINIYLVLLFVRILLSWFQTAEWAGNIMGFLSPVTDPYLNIFRSFIPPLGG-IDFSPIL 68
Query: 191 AFLVLNAFTSASAAL 205
A L A +++
Sbjct: 69 AIFALQFLQQALSSV 83
>gb|AAC35609.1| hypothetical chloroplast RF19 [Guillardia theta]
gi|11467623|ref|NP_050675.1| hypothetical chloroplast
RF19 [Guillardia theta] gi|6136608|sp|O78424|YC19_GUITH
HYPOTHETICAL 10.4 KD PROTEIN YCF19
Length = 91
Score = 63.9 bits (154), Expect = 4e-09
Identities = 35/66 (53%), Positives = 45/66 (68%), Gaps = 7/66 (10%)
Query: 134 FLNIYNTLLVVRLVLTWFPN----SPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPI 189
FL IY LL++R+ LTWFPN P LS I DPYL +FRG++PPL G +D+SPI
Sbjct: 15 FLQIYLILLLIRVSLTWFPNVNWYGQPFY--SLSRITDPYLKMFRGIVPPLIG-IDISPI 71
Query: 190 LAFLVL 195
L F++L
Sbjct: 72 LGFILL 77
>ref|YP_146986.1| hypothetical protein GK1133 [Geobacillus kaustophilus HTA426]
gi|56379510|dbj|BAD75418.1| hypothetical conserved
protein [Geobacillus kaustophilus HTA426]
Length = 90
Score = 62.4 bits (150), Expect = 1e-08
Identities = 31/72 (43%), Positives = 48/72 (66%), Gaps = 2/72 (2%)
Query: 135 LNIYNTLLVVRLVLTWFPNSPPAIVGP-LSTICDPYLNLFRGLIPPLGGTLDLSPILAFL 193
+ +Y+ L++ ++++WFPN+ G L+ IC+PYL FR +IPPL G +D+SPI+AF+
Sbjct: 12 IQVYSYALIIYILMSWFPNARETRFGQMLAAICEPYLEPFRRVIPPL-GIIDVSPIVAFI 70
Query: 194 VLNAFTSASAAL 205
VL T AL
Sbjct: 71 VLEFATRGLHAL 82
>ref|ZP_00516792.1| Protein of unknown function YGGT [Crocosphaera watsonii WH 8501]
gi|67854832|gb|EAM50108.1| Protein of unknown function
YGGT [Crocosphaera watsonii WH 8501]
Length = 103
Score = 61.6 bits (148), Expect = 2e-08
Identities = 35/93 (37%), Positives = 51/93 (54%), Gaps = 3/93 (3%)
Query: 129 NGIQNFLNIYNTLLVVRLVLTWFPNSPPA--IVGPLSTICDPYLNLFRGLIPPLGGTLDL 186
N + F IY L++VR++L+WF + I+ LS I DPYL++FR IPPLGG LD+
Sbjct: 9 NTLYWFFTIYYVLIIVRVLLSWFRGQEWSYNIISFLSPITDPYLDIFRSFIPPLGG-LDI 67
Query: 187 SPILAFLVLNAFTSASAALPAELPVTQQSEQGV 219
S ILA +L + L ++ G+
Sbjct: 68 SAILAIFLLQFLADRAKVLAVSAAISSGFHYGM 100
>ref|NP_692402.1| hypothetical protein OB1481 [Oceanobacillus iheyensis HTE831]
gi|22777164|dbj|BAC13437.1| hypothetical conserved
protein [Oceanobacillus iheyensis HTE831]
Length = 88
Score = 59.7 bits (143), Expect = 7e-08
Identities = 31/69 (44%), Positives = 45/69 (64%), Gaps = 2/69 (2%)
Query: 129 NGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLST-ICDPYLNLFRGLIPPLGGTLDLS 187
N I + IY+ L + ++++W P + + G L T IC+PYL +FR IPPL G +DLS
Sbjct: 6 NLISMGITIYSFALFIYIMMSWIPGARESSFGELLTKICEPYLEIFRRFIPPL-GMIDLS 64
Query: 188 PILAFLVLN 196
PI+A +VLN
Sbjct: 65 PIVAIIVLN 73
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.135 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 424,463,756
Number of Sequences: 2540612
Number of extensions: 17076001
Number of successful extensions: 42715
Number of sequences better than 10.0: 135
Number of HSP's better than 10.0 without gapping: 28
Number of HSP's successfully gapped in prelim test: 107
Number of HSP's that attempted gapping in prelim test: 42589
Number of HSP's gapped (non-prelim): 140
length of query: 247
length of database: 863,360,394
effective HSP length: 125
effective length of query: 122
effective length of database: 545,783,894
effective search space: 66585635068
effective search space used: 66585635068
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0154.8


