

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0154.6
(227 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
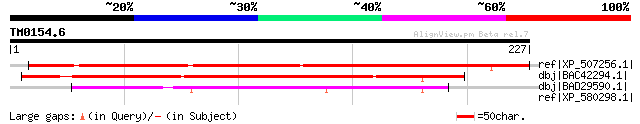
ref|XP_507256.1| PREDICTED P0493A04.32 gene product [Oryza sativ... 234 1e-60
dbj|BAC42294.1| unknown protein [Arabidopsis thaliana] gi|306782... 141 1e-32
dbj|BAD29590.1| ferredoxin-like [Oryza sativa (japonica cultivar... 124 2e-27
ref|XP_580298.1| PREDICTED: similar to KIAA1582 protein, partial... 35 1.2
>ref|XP_507256.1| PREDICTED P0493A04.32 gene product [Oryza sativa (japonica
cultivar-group)] gi|50946509|ref|XP_482782.1| unknown
protein [Oryza sativa (japonica cultivar-group)]
gi|42408414|dbj|BAD09597.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 226
Score = 234 bits (597), Expect = 1e-60
Identities = 119/220 (54%), Positives = 153/220 (69%), Gaps = 6/220 (2%)
Query: 9 ILSFLHSASTGSAATNTQIKVTNNPADKLVAAINENRTAYKVSELYDNAGLACIALQYIK 68
+L+ + +A SAA+ K NNPAD+LVA IN NRTA K S L DN GL CIALQYIK
Sbjct: 12 LLAAVLAALLLSAASAADSK--NNPADQLVALINSNRTASKASTLDDNQGLGCIALQYIK 69
Query: 69 AYQGDCGAVGGPDAKKPPESQFAEVFAPNCGVKASSLAPITGRFLGCQTKYVHAPEAFSE 128
AY+G C VG ++KKPPE+ FAE FAPNCGV+A++L ITGR L CQ+ Y +AF+
Sbjct: 70 AYEGQCNQVG--ESKKPPETSFAETFAPNCGVQAATLTKITGRLLACQSNYATPDQAFN- 126
Query: 129 VLIQNQRSLEILHSKNHTQVGAAVTGTDGGSPYFWCVLFSGGKPNSTFAFEGGVAKLTKP 188
L+ + +S+++LHSKNHT+VGAAV+GT GG PYFWCVLFS GKP ++F +GGV K +P
Sbjct: 127 FLVNDAKSIQVLHSKNHTEVGAAVSGTSGGGPYFWCVLFSSGKPTTSFKVDGGVPKSVRP 186
Query: 189 GCFSGANDECSGAHDWSPLSV-MWLFAASVLIALGFAFPL 227
GCFSG ND+C GA+ + W A++L + F L
Sbjct: 187 GCFSGNNDDCMGANAAVSIGAGTWRLVAALLFSAACVFAL 226
>dbj|BAC42294.1| unknown protein [Arabidopsis thaliana] gi|30678256|ref|NP_171720.2|
ferredoxin-related [Arabidopsis thaliana]
Length = 226
Score = 141 bits (356), Expect = 1e-32
Identities = 75/195 (38%), Positives = 119/195 (60%), Gaps = 8/195 (4%)
Query: 6 LVVILSFLHSASTGSAATNTQIKVTNNPADKLVAAINENRTAYKVSELYDNAGLACIALQ 65
L +IL FL +S +++ K+ N A ++V+ +N+NRTA K+ +L ++ GL C+ALQ
Sbjct: 8 LELILLFLSLSSVLASS-----KLHGNSAHEMVSILNQNRTARKLGKLNESPGLGCMALQ 62
Query: 66 YIKAYQGDCGAVGGPDAKKPPESQFAEVFAPNCGVKASSLAPITGRFLGCQTKYVHAPEA 125
Y++ +G+C V + + PE F +VFAPNCGV+ + ITG LGC +KY A
Sbjct: 63 YVELCEGNCN-VNNTLSCEHPEDDFTQVFAPNCGVELPTFGTITGHILGCSSKYAAPEVA 121
Query: 126 FSEVLIQNQRSLEILHSKNHTQVGAAVTGTDGGSPYFWCVLFSGGKPNSTFAFE-GGVAK 184
FS++L ++ +L +L +++HT+VG + G+ +FWC+LFS G NS+F E G
Sbjct: 122 FSDILFRDSSALSVLRNRSHTEVGVGMARLHKGT-FFWCLLFSDGVKNSSFVLEDNGRGI 180
Query: 185 LTKPGCFSGANDECS 199
+ GC+SG+ CS
Sbjct: 181 KQRTGCYSGSAFPCS 195
>dbj|BAD29590.1| ferredoxin-like [Oryza sativa (japonica cultivar-group)]
gi|50251426|dbj|BAD28464.1| ferredoxin-like [Oryza
sativa (japonica cultivar-group)]
Length = 230
Score = 124 bits (311), Expect = 2e-27
Identities = 74/175 (42%), Positives = 93/175 (52%), Gaps = 14/175 (8%)
Query: 28 KVTNNPADKLVAAINENRTAYKVSELYDNAGLACIALQYIKAYQGDCGAVG------GPD 81
K+ NPA+ LVA +N NRTA K+ L +AGL C+ALQYI DC +G
Sbjct: 24 KIHGNPANDLVALVNANRTATKLPHLRTSAGLGCMALQYI----SDCIGIGIGCAGDNTV 79
Query: 82 AKKPPESQFAEVFAPNCGVKASSLAPITGRFLGCQTKYVHAPEAFSEVLIQNQRSL---E 138
A +PPE+ EV+A NCGV+ ++ ITGR LGC + A A VL + S
Sbjct: 80 ACQPPEAHITEVYAANCGVELPTVDVITGRLLGCHRQRSDAEAALEAVLSGSGNSTAARA 139
Query: 139 ILHSKNHTQVGAAVTGTDGGSPYFWCVLFSGGKPNSTFAFE-GGVAKLTKPGCFS 192
++ K HTQVGA P+FWC+LFS G NSTF E G GCFS
Sbjct: 140 VIRGKEHTQVGAGFDRAHRRGPFFWCLLFSSGSANSTFLLEAAGKGVHQSHGCFS 194
>ref|XP_580298.1| PREDICTED: similar to KIAA1582 protein, partial [Bos taurus]
Length = 1746
Score = 35.4 bits (80), Expect = 1.2
Identities = 37/134 (27%), Positives = 53/134 (38%), Gaps = 15/134 (11%)
Query: 73 DCGA---VGGPDAKKPPESQFAEVFAPNCGVKASSLAP--ITGRFLGCQTKYVHAPEAFS 127
D GA + G + + E A NCG + SS+A G F G QTK + +
Sbjct: 252 DAGAWPSITGAETESASECTTDTDSASNCGSENSSMATGSAQGSFTG-QTKKTNGNNGSN 310
Query: 128 EVLIQNQRSLEILHSKNHTQVGAAVTGTDGGSPYFWCVLFSGGKPNSTFAFEGGVAKLTK 187
L+QN + L GA ++GG+ W V S G + + GG K+
Sbjct: 311 GTLVQNPSAQSAL--------GAGGANSNGGAARVWGVAASAGSGLAHCSLGGGDGKMDN 362
Query: 188 PGCFSGANDECSGA 201
G + C GA
Sbjct: 363 M-IGDGRSQNCWGA 375
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.134 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 402,279,737
Number of Sequences: 2540612
Number of extensions: 17103988
Number of successful extensions: 33597
Number of sequences better than 10.0: 4
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 33588
Number of HSP's gapped (non-prelim): 4
length of query: 227
length of database: 863,360,394
effective HSP length: 124
effective length of query: 103
effective length of database: 548,324,506
effective search space: 56477424118
effective search space used: 56477424118
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0154.6


