

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0154.14
(906 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
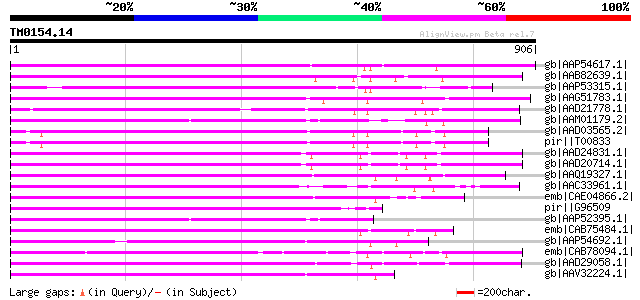
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP54617.1| putative non-LTR retroelement reverse transcripta... 560 e-157
gb|AAB82639.1| putative non-LTR retroelement reverse transcripta... 526 e-148
gb|AAP53315.1| putative reverse transcriptase [Oryza sativa (jap... 524 e-147
gb|AAG51783.1| reverse transcriptase, putative; 16838-20266 [Ara... 517 e-145
gb|AAD21778.1| putative non-LTR retroelement reverse transcripta... 507 e-142
gb|AAM01179.2| Putative retroelement [Oryza sativa (japonica cul... 506 e-141
gb|AAD03565.2| putative non-LTR retroelement reverse transcripta... 481 e-134
pir||T00833 RNA-directed DNA polymerase homolog T13L16.7 - Arabi... 481 e-134
gb|AAD24831.1| putative non-LTR retroelement reverse transcripta... 476 e-132
gb|AAD20714.1| putative non-LTR retroelement reverse transcripta... 473 e-132
gb|AAQ19327.1| bZIP-like protein [Oryza sativa (japonica cultiva... 473 e-131
gb|AAC33961.1| contains similarity to reverse trancriptase (Pfam... 470 e-130
emb|CAE04866.2| OSJNBa0086O06.14 [Oryza sativa (japonica cultiva... 469 e-130
pir||G96509 protein F27F5.21 [imported] - Arabidopsis thaliana g... 464 e-129
gb|AAP52395.1| putative retroelement [Oryza sativa (japonica cul... 463 e-128
emb|CAB75484.1| putative protein [Arabidopsis thaliana] gi|11357... 462 e-128
gb|AAP54692.1| putative reverse transcriptase [Oryza sativa (jap... 462 e-128
emb|CAB78094.1| RNA-directed DNA polymerase-like protein [Arabid... 461 e-128
gb|AAD29058.1| putative non-LTR retroelement reverse transcripta... 451 e-125
gb|AAV32224.1| hypothetical protein [Oryza sativa (japonica cult... 448 e-124
>gb|AAP54617.1| putative non-LTR retroelement reverse transcriptase [Oryza sativa
(japonica cultivar-group)] gi|37536056|ref|NP_922330.1|
putative non-LTR retroelement reverse transcriptase
[Oryza sativa (japonica cultivar-group)]
gi|10140689|gb|AAG13524.1| putative non-LTR retroelement
reverse transcriptase [Oryza sativa (japonica
cultivar-group)]
Length = 1382
Score = 560 bits (1442), Expect = e-157
Identities = 335/937 (35%), Positives = 492/937 (51%), Gaps = 34/937 (3%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+++ ALFQM TKAPG DG PALFYQ W I+ + + + L G+ + +++VL
Sbjct: 448 EIKTALFQMGSTKAPGPDGFPALFYQTHWGILEEHICNAVRGFLLGEEIPEGLCDSVVVL 507
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPKV H ++FR ISLCNV++KI +K LANRLK LPD++ E Q+AFVPGRLITD+AL
Sbjct: 508 IPKVNNASHLSKFRPISLCNVLYKIASKVLANRLKPFLPDIVSEFQSAFVPGRLITDSAL 567
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
VA+EC H ++K+ + N ALK+DM KAYDRVEW +L+ L ++GF W++ +M CVS
Sbjct: 568 VAYECLHTIRKQHNK-NPFFALKIDMMKAYDRVEWAYLSGCLSKLGFSQDWINTVMRCVS 626
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V +++ +NG +P P RG+RQGDP+SPYLF+LC E S L++K +A L GIK R
Sbjct: 627 SVRYAVKINGELTKPVVPSRGIRQGDPISPYLFLLCTEGLSCLLHKKEVAGELQGIKNGR 686
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
P ISHLLFADDS+ FA+A + ++K L SY ASGQ INL KS + + PD
Sbjct: 687 HGPPISHLLFADDSIFFAKADSRNVQALKNTLRSYCSASGQKINLHKSSIFFGKRCPDAV 746
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
++ L V D YLG+PT IG + T F F+ ER+WK++ GW +R L RAG E
Sbjct: 747 KISVKSCLQVDNEVLQDSYLGMPTEIGLATTNFFKFLPERIWKRVNGWTDRPLSRAGMET 806
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
++KAVA AIP+YVMSCF +P +C +++ I+ +WG + ++ MH SW L K G
Sbjct: 807 MLKAVAQAIPNYVMSCFRIPVSICEKMKTCIADHWWGFEDGKKKMHWKSWSWLSTPKFLG 866
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
+GFR+F FN A++ + WR+ ++P+SL +V + YFP + A + PS+ W S+
Sbjct: 867 GMGFREFTTFNQAMLGRQCWRLLTDPDSLCSRVLKGRYFPNSSFWEAAQPKSPSFTWRSL 926
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSLVSDLMLPGLLA 540
+ E+ A G W +G+GK++K +SD W+PG + + L+ + VS LM
Sbjct: 927 LFGRELLAKGVRWGVGDGKTIKIFSDNWIPGFRPQLVTT-LSPFPTDATVSCLMNEDARC 985
Query: 541 WNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHGAEAVVA 600
W+ +LI + A+ I+ IP+ D WP G YS +S Y R A
Sbjct: 986 WDGDLIRSLFPVDIAKEILQIPISRHGDADFASWPHDKLGLYSVRSAYNLARSEAFFADQ 1045
Query: 601 SSSQVPSLPR-----ADWRKLWKADA--------------LLPVRAYLHARGLVTDPTCP 641
S+S R DW+ LWK +A L L R + + C
Sbjct: 1046 SNSGRGMASRLLESQKDWKGLWKINAPGKMKITLWRAAHECLATGFQLRRRHIPSTDGCV 1105
Query: 642 RCLVALETVQHALLFCPMVQPIW--FASSLGFRLAHE--CRVHEFVSDFMQVADMSSLGA 697
C +TV+H LFCP IW +L + +++ DF++ +
Sbjct: 1106 FC-NRDDTVEHVFLFCPFAAQIWEEIKGKCAVKLGRNGFSTMRQWIFDFLKRGSSHANTL 1164
Query: 698 FLTLLYAIWMARNELCFKEKASTLEQILTHAASLTPL------PPLEGPMVPQVHRPNTW 751
+ IW ARN ++++ S + ++G W
Sbjct: 1165 LAVTFWHIWEARNNTKNNNGTVHPQRVVIKILSYVDMILKHNTKTVDGQRGGNTQAIPRW 1224
Query: 752 SRPHPGTFKINFDAAM-APTGDAGFGLVARNSRGEVLAAACSSHGPAASSLLAEGLSFRW 810
P + IN DAA+ + + G G + R++ G+ L A LAE L+ R
Sbjct: 1225 QPPPASVWMINSDAAIFSSSRTMGVGALIRDNTGKCLVACSEMISDVVLPELAEALAIRR 1284
Query: 811 ALGLAMDLGFRRVVFEIDCLLLYEAWKRRG-GCSILDSLILDCRRLSSGFDAFDFSFIRR 869
ALGLA + G +V DCL + + G S + +I D ++L+S F F + R
Sbjct: 1285 ALGLAKEEGLEHIVMASDCLTVIRRIQTSGRDRSGVGCVIEDIKKLASTFVLCSFMHVNR 1344
Query: 870 TGNSVADALAKRSFTLGVWVWIEEIPQDFYGLVQSDV 906
N A +LA+ + V+ IP ++ DV
Sbjct: 1345 LSNLAAHSLARNAELSTCTVYRSVIPDYIRDILCDDV 1381
>gb|AAB82639.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25408936|pir||A84888 hypothetical protein
At2g45230 [imported] - Arabidopsis thaliana
Length = 1374
Score = 526 bits (1356), Expect = e-148
Identities = 322/930 (34%), Positives = 464/930 (49%), Gaps = 56/930 (6%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+V+ A F ++P K PG DG+ YQ+FW +GD ++ +NKT + L
Sbjct: 432 EVQRATFSINPHKCPGPDGMNGFLYQQFWETMGDQITEMVQAFFRSGSIEEGMNKTNICL 491
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK+ K FR ISLCNVI+K+I K +ANRLK ILP LI E+Q AFV GRLI+DN L
Sbjct: 492 IPKILKAEKMTDFRPISLCNVIYKVIGKLMANRLKKILPSLISETQAAFVKGRLISDNIL 551
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+A E H + +A+K D+SKAYDRVEWPFL ++ +GF W+ LIM CV
Sbjct: 552 IAHELLHALSSNNKCSEEFIAIKTDISKAYDRVEWPFLEKAMRGLGFADHWIRLIMECVK 611
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V + +++NG P P RGLRQGDPLSPYLF++C E+ ++ A + + G+K+AR
Sbjct: 612 SVRYQVLINGTPHGEIIPSRGLRQGDPLSPYLFVICTEMLVKMLQSAEQKNQITGLKVAR 671
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
AP ISHLLFADDS+ + + + I I+ Y ASGQ +N KS + +++ ++
Sbjct: 672 GAPPISHLLFADDSMFYCKVNDEALGQIIRIIEEYSLASGQRVNYLKSSIYFGKHISEER 731
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+++ L ++ YLGLP SK +++K+R+ KK+ GW+ L G+E+
Sbjct: 732 RCLVKRKLGIEREGGEGVYLGLPESFQGSKVATLSYLKDRLGKKVLGWQSNFLSPGGKEI 791
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
L+KAVA A+P+Y MSCF +P +C QIE +++ F+W RG+H +W L + K G
Sbjct: 792 LLKAVAMALPTYTMSCFKIPKTICQQIESVMAEFWWKNKKEGRGLHWKAWCHLSRPKAVG 851
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGF++ + FN AL+ K WR+ + +SLM KV+++ YF + A G RPS+AW SI
Sbjct: 852 GLGFKEIEAFNIALLGKQLWRMITEKDSLMAKVFKSRYFSKSDPLNAPLGSRPSFAWKSI 911
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHF-------GVSLVSDL 533
+ + G IGNG+++ W+D W+ H + +V DL
Sbjct: 912 YEAQVLIKQGIRAVIGNGETINVWTDPWIGAKPAKAAQAVKRSHLVSQYAANSIHVVKDL 971
Query: 534 MLPGLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFL-- 591
+LP WN L++ + T I+A+ D W +S GHYS KSGY +
Sbjct: 972 LLPDGRDWNWNLVSLLFPDNTQENILALRPGGKETRDRFTWEYSRSGHYSVKSGYWVMTE 1031
Query: 592 ----RHGAEAVVASSSQVPSLPRADWRKLWKADA--------------LLPVRAYLHARG 633
R+ + V+ PSL ++++WK D L V + L R
Sbjct: 1032 IINQRNNPQEVLQ-----PSLDPI-FQQIWKLDVPPKIHHFLWRCVNNCLSVASNLAYRH 1085
Query: 634 LVTDPTCPRCLVALETVQHALLFCPMVQPIW------------FASSLGFRLAHECRVHE 681
L + +C RC ETV H L CP + W +A SL + H VH+
Sbjct: 1086 LAREKSCVRCPSHGETVNHLLFKCPFARLTWAISPLPAPPGGEWAESLFRNMHHVLSVHK 1145
Query: 682 FVSDFMQVADMSSLGAFLTLLYAIWMARNELCFKEKASTLEQILTHAASLTPLPPLEGPM 741
Q + +L+ +W RN+L FK + T Q++ A
Sbjct: 1146 -----SQPEESDHHALIPWILWRLWKNRNDLVFKGREFTAPQVILKATEDMDAWNNRKEP 1200
Query: 742 VPQV-----HRPNTWSRPHPGTFKINFDAAMA-PTGDAGFGLVARNSRGEVLAAACSSHG 795
PQV R W P G K N D A + G+ G G V RN G +L +
Sbjct: 1201 QPQVTSSTRDRCVKWQPPSHGWVKCNTDGAWSKDLGNCGVGWVLRNHTGRLLWLGLRALP 1260
Query: 796 PAASSLLAEGLSFRWALGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRL 855
S L E + RWA+ +RRV+FE D L + L I D R L
Sbjct: 1261 SQQSVLETEVEALRWAVLSLSRFNYRRVIFESDSQYLVSLIQNEMDIPSLAPRIQDIRNL 1320
Query: 856 SSGFDAFDFSFIRRTGNSVADALAKRSFTL 885
F+ F F RR GN+VAD A+ S +L
Sbjct: 1321 LRHFEEVKFQFTRREGNNVADRTARESLSL 1350
>gb|AAP53315.1| putative reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|37533452|ref|NP_921028.1| putative
reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|20279456|gb|AAM18736.1| putative
reverse transcriptase [Oryza sativa (japonica
cultivar-group)]
Length = 1509
Score = 524 bits (1350), Expect = e-147
Identities = 324/858 (37%), Positives = 451/858 (51%), Gaps = 86/858 (10%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+V+EAL + KAPG DG+PA FY+ W ++G+ V+ L+VL G N T +VL
Sbjct: 658 EVKEALDAIGDLKAPGPDGMPAGFYKACWDVVGEKVTVEVLEVLRGGAIPEGWNDTTIVL 717
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPKV LANRLK ILPD+I +Q+AFVPGRLI+DN L
Sbjct: 718 IPKV-------------------------LANRLKKILPDVISPAQSAFVPGRLISDNIL 752
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+A+E HYM+ K SG G A KLDMSKAYDRVEW FL ++ ++GF WV+LIM CVS
Sbjct: 753 IAYEMTHYMRNKRSGQVGYAAFKLDMSKAYDRVEWSFLHDMMLKLGFHTDWVNLIMKCVS 812
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
TV + I +NG E F P RGLRQGDPLSPYLF+LC E FSAL++K LHGI+I +
Sbjct: 813 TVTYRIRVNGELSESFSPERGLRQGDPLSPYLFLLCAEGFSALLSKTEEEGRLHGIRICQ 872
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
AP +SHLLFADDS+I RA EA ++ IL YE+ SGQVIN DKS + S N
Sbjct: 873 GAPSVSHLLFADDSLILCRANGGEAQQLQTILQIYEECSGQVINKDKSAVMFSPNTSSLE 932
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+ L ++ + +KYLGLP +G+S+T+IF+++KER+W++++GWKE+ L RAG+E+
Sbjct: 933 KGAVMAALNMQRETTNEKYLGLPVFVGRSRTKIFSYLKERIWQRIQGWKEKLLSRAGKEI 992
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
L+KAVA IP++ M CF L LC+QI MI++++W MH LSW L K+ G
Sbjct: 993 LIKAVAQVIPTFAMGCFELTKDLCDQISKMIAKYWWSNQEKDNKMHWLSWNKLTLPKNMG 1052
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFRD +FN A++AK WR+ +P+SL +V RA YFP G K+ SY W SI
Sbjct: 1053 GLGFRDIYIFNLAMLAKQGWRLIQDPDSLCSRVLRAKYFPLGDCFRPKQTSNVSYTWRSI 1112
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSLVSDLMLPGLLA 540
+ + G +W +G+G + W+D W+P + + V+ V +L+ P
Sbjct: 1113 QKGLRVLQNGMIWRVGDGSKINIWADPWIPRGWSRKPMTPRGANL-VTKVEELIDPYTGT 1171
Query: 541 WNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHGAEAVVA 600
W+ +L++ AI +IP+ V +DVL W F + G ++ KS Y+ R A
Sbjct: 1172 WDEDLLSQTFWEEDVAAIKSIPVH-VEMEDVLAWHFDARGCFTVKSAYKVQREMERR--A 1228
Query: 601 SSSQVPSLPRAD------WRKLWKADA--------------LLPVRAYLHARGLVTDPTC 640
S + P + + W+KLWK L +RA LH RG+ D C
Sbjct: 1229 SRNGCPGVSNWESGDDDFWKKLWKLGVPGKIKHFLWRMCHNTLALRANLHHRGMDVDTRC 1288
Query: 641 PRCLVALETVQHALLFCPMVQPIWFASSLG--FRLAHECRVHEFVSDFMQVADMSSLGAF 698
C E H C V+ +W A +L + + + V + + +
Sbjct: 1289 VMCGRYNEDAGHLFFKCKPVKKVWQALNLEELRSMLEQQTSGKNVLQSIYCRPENERTSA 1348
Query: 699 LTLLYAIWMARNELCFKEKASTLEQILTHAASLTPLPPLEGPMVPQVHRPNTWSRPHPGT 758
+ L+ W RNE EK+ P+ W RP
Sbjct: 1349 IVCLWQWWKERNE----EKS------------------------PRTGECAVWRRPPLNF 1380
Query: 759 FKINFDAAMAPT-GDAGFGLVARNSRGEVLAAACSSHGPAA---SSLLAEGLSFRWALGL 814
KIN D A + G+G V R+ G VL A GPAA + AE ++ A+
Sbjct: 1381 VKINTDGAYSSNMKQGGWGFVIRDQTGAVLQAGA---GPAAYLQDAFHAEVVACAAAIKT 1437
Query: 815 AMDLGFRRVVFEIDCLLL 832
A + G R+ E D ++L
Sbjct: 1438 ASERGMSRIELETDSMML 1455
>gb|AAG51783.1| reverse transcriptase, putative; 16838-20266 [Arabidopsis thaliana]
gi|25405244|pir||E96519 probable reverse transcriptase,
16838-20266 [imported] - Arabidopsis thaliana
Length = 1142
Score = 517 bits (1331), Expect = e-145
Identities = 317/936 (33%), Positives = 479/936 (50%), Gaps = 49/936 (5%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+V ALF +HP KAPG DG+ ALF+QK W II D+ L + +N T + L
Sbjct: 216 EVRAALFMIHPEKAPGPDGMTALFFQKSWAIIKSDLLSLVNSFLQEGVFDKRLNTTNICL 275
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK ++P + R ISLCNV +K+I+K L RLK +LP+LI E+Q+AFV GRLI+DN L
Sbjct: 276 IPKTERPTRMTELRPISLCNVGYKVISKILCQRLKTVLPNLISETQSAFVDGRLISDNIL 335
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+A E FH ++ S + MA+K DMSKAYD+VEW F+ ++L++MGF W+S IM C++
Sbjct: 336 IAQEMFHGLRTNSSCKDKFMAIKTDMSKAYDQVEWNFIEALLRKMGFCEKWISWIMWCIT 395
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
TV + +++NG P+ P RGLRQGDPLSPYLFILC EV A I KA + + GIK+A
Sbjct: 396 TVQYKVLINGQPKGLIIPERGLRQGDPLSPYLFILCTEVLIANIRKAERQNLITGIKVAT 455
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
+P +SHLLFADDS+ F +A ++ I IL YE SGQ IN KS + V D
Sbjct: 456 PSPAVSHLLFADDSLFFCKANKEQCGIILEILKQYESVSGQQINFSKSSIQFGHKVEDSI 515
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+++ +L + + YLGLP +G SKT++F+FV++R+ ++ GW + L + G+EV
Sbjct: 516 KADIKLILGIHNLGGMGSYLGLPESLGGSKTKVFSFVRDRLQSRINGWSAKFLSKGGKEV 575
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
++K+VA +P YVMSCF LP + +++ +++F+W + RGMH ++W LC +K +G
Sbjct: 576 MIKSVAATLPRYVMSCFRLPKAITSKLTSAVAKFWWSSNGDSRGMHWMAWDKLCSSKSDG 635
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFR+ FNSAL+AK WR+ + P+SL KV++ YF + + + Y PSY W S+
Sbjct: 636 GLGFRNVDDFNSALLAKQLWRLITAPDSLFAKVFKGRYFRKSNPLDSIKSYSPSYGWRSM 695
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSLVS-DLMLPGLL 539
+ + + G + +G+G S+ W+D W+P +G S+V L + L+
Sbjct: 696 ISARSLVYKGLIKRVGSGASISVWNDPWIPA------QFPRPAKYGGSIVDPSLKVKSLI 749
Query: 540 -----AWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHG 594
WN +L+ + P I A+P+ +D L W F+ G+Y+ KSGY R
Sbjct: 750 DSRSNFWNIDLLKELFDPEDVPLISALPIGNPNMEDTLGWHFTKAGNYTVKSGYHTARLD 809
Query: 595 A-EAVVASSSQVPSLPRADWRK---------LWK-ADALLPVRAYLHARGLVTDPTCPRC 643
E + +L W+ LW+ +PV L RG++ D C C
Sbjct: 810 LNEGTTLIGPDLTTLKAYIWKVQCPPKLRHFLWQILSGCVPVSENLRKRGILCDKGCVSC 869
Query: 644 LVALETVQHALLFCPMVQPIWFASSLGFR---LAHECRVHEFVSDFMQVADMSSLGAFLT 700
+ E++ H L C + IW S + F ++ +
Sbjct: 870 GASEESINHTLFQCHPARQIWALSQIPTAPGIFPSNSIFTNLDHLFWRIPSGVDSAPYPW 929
Query: 701 LLYAIWMARNE---------------LCFKEKASTLE-QILTHAASLTPLPPLEGPMVPQ 744
+++ IW ARNE L KE S E Q+ H+ L V
Sbjct: 930 IIWYIWKARNEKVFENVDKDPMEILLLAVKEAQSWQEAQVELHSERHGSLSIDSRIRVRD 989
Query: 745 VHRPNTWSRPHPGTFKINFDAAMAPTGD-AGFGLVARNSRGEVLAAACSSHGPAASSLLA 803
V + T+S F+ D + + +G G +S GE ++ + S L
Sbjct: 990 VSQDTTFS-----GFRCFIDGSWKASDQFSGTGWFCLSSLGESPTMGAANVRRSLSPLHT 1044
Query: 804 EGLSFRWALGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRLSSGFDAFD 863
E + WA+ + + V F DC L + + + + F F
Sbjct: 1045 EMEALLWAMKCMIGADNQNVAFFTDCSDLVKMVSSPTEWPAFSVYLEELQSDREEFTNFS 1104
Query: 864 FSFIRRTGNSVADALAKRSFTLGVWV-WIEEIPQDF 898
S I R+ N AD LA++ T+ V ++ IP+D+
Sbjct: 1105 LSLISRSANVKADKLARKIRTVPHHVTYVNNIPRDW 1140
>gb|AAD21778.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25410938|pir||G84429 hypothetical protein
At2g01840 [imported] - Arabidopsis thaliana
Length = 1715
Score = 507 bits (1305), Expect = e-142
Identities = 314/922 (34%), Positives = 462/922 (50%), Gaps = 71/922 (7%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLH---GDIS*SIINKTL 57
+V +A+F + +APG DG A FY W +IG+DV CL V H D+ + IN+T
Sbjct: 794 EVRDAVFAIGADRAPGFDGFTAAFYHHLWDLIGNDV---CLMVRHFFESDVMDNQINQTQ 850
Query: 58 LVLIPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITD 117
+ LIPK+ P H + +R ISLC +KII+K L RLK L D+I +SQ AFVPG+ I+D
Sbjct: 851 ICLIPKIIDPKHMSDYRPISLCTASYKIISKILIKRLKQCLGDVISDSQAAFVPGQNISD 910
Query: 118 NALVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMT 177
N LVA E H +K + +G +A+K D+SKAYDRVEW FL V+ Q+GF WV IMT
Sbjct: 911 NVLVAHELLHSLKSRRECQSGYVAVKTDISKAYDRVEWNFLEKVMIQLGFAPRWVKWIMT 970
Query: 178 CVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIK 237
CV++V + +++NG+P P RG+RQGDPLSPYLF+ C EV S ++ KA + +HG+K
Sbjct: 971 CVTSVSYEVLINGSPYGKIFPSRGIRQGDPLSPYLFLFCAEVLSNMLRKAEVNKQIHGMK 1030
Query: 238 IARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVP 297
I + ISHLLFADDS+ F RA+ Q + I YE+ASGQ IN KS + + +P
Sbjct: 1031 ITKDCLAISHLLFADDSLFFCRASNQNIEQLALIFKKYEEASGQKINYAKSSIIFGQKIP 1090
Query: 298 DDCFHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAG 357
L +LL + V KYLGLP +G+ K ++F ++ +V ++ +GW L AG
Sbjct: 1091 TMRRQRLHRLLGIDNVRGGGKYLGLPEQLGRRKVELFEYIVTKVKERTEGWAYNYLSPAG 1150
Query: 358 REVLVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAK 417
+E+++KA+A A+P Y M+CFLLP +CN+I +I+ F+WG +
Sbjct: 1151 KEIVIKAIAMALPVYSMNCFLLPTLICNEINSLITAFWWG------------------KE 1192
Query: 418 DNGRLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAW 477
+ G LGF+D FN AL+AK WRI +NP+SL+ ++Y+ +Y+P T A +G SY W
Sbjct: 1193 NEGDLGFKDLHQFNRALLAKQAWRILTNPQSLLARLYKGLYYPNTTYLRANKGGHASYGW 1252
Query: 478 SSIMRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSL-VSDLMLP 536
+SI + G +G+G++ K W D W+P TL + V+DL
Sbjct: 1253 NSIQEGKLLLQQGLRVRLGDGQTTKIWEDPWLP---TLPPRPARGPILDEDMKVADLWRE 1309
Query: 537 GLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRH--- 593
W+ + + P + ++ L D W ++ + Y+ +SGY H
Sbjct: 1310 NKREWDPVIFEGVLNPEDQQLAKSLYLSNYAARDSYKWAYTRNTQYTVRSGYWVATHVNL 1369
Query: 594 -GAEAVVASSSQVPSLPRADWRK---------LWKA-DALLPVRAYLHARGLVTDPTCPR 642
E + VP L + WR +W+ L L R + DPTC R
Sbjct: 1370 TEEEIINPLEGDVP-LKQEIWRLKITPKIKHFIWRCLSGALSTTTQLRNRNIPADPTCQR 1428
Query: 643 CLVALETVQHALLFCPMVQPIWFASSL--GFRLAHECRVHEFVSDFMQVADMSSLGAF-- 698
C A ET+ H + C Q +W +++ RL + E + +Q +L
Sbjct: 1429 CCNADETINHIIFTCSYAQVVWRSANFSGSNRLCFTDNLEENIRLILQGKKNQNLPILNG 1488
Query: 699 ---LTLLYAIWMARNELCFKE-------KASTLEQILTH---------AASLTPLPPLEG 739
+++ +W +RNE F++ A EQ T A S +
Sbjct: 1489 LMPFWIMWRLWKSRNEYLFQQLDRFPWKVAQKAEQEATEWVETMVNDTAISHNTAQSNDR 1548
Query: 740 PMVPQVHRPNTWSRPHPGTFKINFDAAMAPTGD-AGFGLVARNSRGEVLAAACSSHGPAA 798
P+ R WS P G K NFD+ D G + R+ G VL + C+ +
Sbjct: 1549 PL----SRSKQWSSPPEGFLKCNFDSGYVQGRDYTSTGWILRDCNGRVLHSGCAKLQQSY 1604
Query: 799 SSLLAEGLSFRWALGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRLSSG 858
S+L AE L F AL + G+ V FE D L L + +L++L+ D R +
Sbjct: 1605 SALQAEALGFLHALQMVWIRGYCYVWFEGDNLELTNLINKTEDHHLLETLLYDIRFWMTK 1664
Query: 859 FDAFDFSFIRRTGNSVADALAK 880
++ R N AD L K
Sbjct: 1665 LPFSSIGYVNRERNLAADKLTK 1686
>gb|AAM01179.2| Putative retroelement [Oryza sativa (japonica cultivar-group)]
Length = 1888
Score = 506 bits (1302), Expect = e-141
Identities = 312/909 (34%), Positives = 468/909 (51%), Gaps = 79/909 (8%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
++ ALFQ+ P KAPG DG P FYQ+ W I+ DD+ + S +N+T +VL
Sbjct: 1003 EIANALFQIGPLKAPGPDGFPGRFYQRNWAILKDDIVRAVQEFFSLGTMPSGVNETAIVL 1062
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK ++P FR ISLCNV++KI++K L NRL+ IL DL+ ++Q+AFVPGRLITDNAL
Sbjct: 1063 IPKTEQPQELKDFRPISLCNVVYKIVSKCLVNRLRPILDDLVSQNQSAFVPGRLITDNAL 1122
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+AFE FH++++ + N A KLD+SKAYDRV+W FL + ++GF WV IM C++
Sbjct: 1123 IAFEYFHHIQRNKNPENAYSAYKLDLSKAYDRVDWEFLEQAMVKLGFAHCWVKWIMACIT 1182
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
V +++ LNG F P RGLRQGDPLSP+LF+ + S L+ + V +++ I I R
Sbjct: 1183 LVRYAVKLNGTLLNTFAPSRGLRQGDPLSPFLFLFVADGLSFLLEEKVDQGAMNPIHICR 1242
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
AP + HLLFADD+ +F +A +A+ +K +L +Y +GQ+IN K + ++ P
Sbjct: 1243 HAPAVLHLLFADDTPLFFKAANDQAVVMKEVLETYASCTGQLINPSKCSIMFGQSSPTAV 1302
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
++++Q L + + DKYLG PT G+ F ++ER+WK++ W E+ L G+E+
Sbjct: 1303 QNQIKQTLQI-SNSFEDKYLGFPTPEGRMTKGKFQSLQERLWKRIMLWGEKFLSTGGKEI 1361
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
L+KAV A+P YVM F LP+ +C ++ ++ F+WG + R H SW L K G
Sbjct: 1362 LIKAVLQAMPVYVMGLFKLPESVCEELTKLVRNFWWGAEKGMRRTHWKSWDCLIAQKLKG 1421
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
+GFRDF++FN AL+A+ WR+ P+SL ++ +A+Y+P G++ G S W +I
Sbjct: 1422 GMGFRDFRLFNQALLARQAWRLLDRPDSLCARILKAIYYPNGSLIDTSFGGNASPGWRAI 1481
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYS---VGLAQHFGVSLVSDLMLPG 537
E+ G +W IGNGKSV+ W D W+P + YS + ++ + VSDL +
Sbjct: 1482 EYGLELLKKGIIWRIGNGKSVRLWRDPWIPRS----YSRRPISAKRNCRLRWVSDL-IDQ 1536
Query: 538 LLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHGAEA 597
+W+ + I+ P A AI+ I L DD + W G +S +S Y A
Sbjct: 1537 SGSWDTDKISQHFLPMDAEAILNIRLSSRLEDDFIAWHPDKLGRFSVRSAYHLAVALAHV 1596
Query: 598 VVASSSQVPSLPRADWRKLWKADALLPVRAYLHARGLVTDPTCPRCLVALETVQHALLFC 657
SSS RA W LWK +A P+ L L +++
Sbjct: 1597 DEGSSSSGNGNSRA-WNALWKCNA-------------------PQKLNPLSNIRNG---- 1632
Query: 658 PMVQPIWFASSLGFRLAHECRVHEFVSDFMQVADMSSLGAFLTLLYAIWMARNELCFKEK 717
P + + D ++ + L +L+ IW RNE+ +
Sbjct: 1633 ---DP------------------KVILDLLEQSQPEDQVMLLMVLWRIWHTRNEIVHGKP 1671
Query: 718 A-------STLEQILTHAASLTPLP---PLEGPMVPQVHR-------------PNTWSRP 754
A +E + A + P P +G V V + P+ W +P
Sbjct: 1672 APGMLVSKRFIESYVLSLAEIKQHPQASPEKGKHVVDVVQKKLHSIKRSREPVPDKWLKP 1731
Query: 755 HPGTFKINFDAAMAPT-GDAGFGLVARNSRGEVLAAACSSHGPAASSLLAEGLSFRWALG 813
PG+ K+N D + + G G G V RN GEV+ AAC +S+L E L+ R L
Sbjct: 1732 LPGSMKLNVDGSFQESDGKGGIGAVLRNCTGEVIFAACGHVDCCSSALETELLACRDGLA 1791
Query: 814 LAMDLGFRRVVFEIDCLLLYEAWK-RRGGCSILDSLILDCRRLSSGFDAFDFSFIRRTGN 872
LA+ +V E DCL + ++ G S L LI + L G FS R+ N
Sbjct: 1792 LALQWTLLPIVIETDCLAMIHLFRDATGAKSELAFLIREIDSLLVGNRDISFSKCLRSQN 1851
Query: 873 SVADALAKR 881
++ LA +
Sbjct: 1852 HISHYLANK 1860
>gb|AAD03565.2| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25411819|pir||H84557 hypothetical protein
At2g17910 [imported] - Arabidopsis thaliana
Length = 1344
Score = 481 bits (1238), Expect = e-134
Identities = 306/862 (35%), Positives = 439/862 (50%), Gaps = 51/862 (5%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SII----NKT 56
+V A+F ++ APG DG ALF+Q+ W D V H L + G ++ N T
Sbjct: 410 EVYNAVFSINKESAPGPDGFTALFFQQHW----DLVKHQILTEIFGFFETGVLPQDWNHT 465
Query: 57 LLVLIPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLIT 116
+ LIPK+ P + R ISLC+V++KII+K L RLK LP ++ +Q+AFVP RLI+
Sbjct: 466 HICLIPKITSPQRMSDLRPISLCSVLYKIISKILTQRLKKHLPAIVSTTQSAFVPQRLIS 525
Query: 117 DNALVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIM 176
DN LVA E H ++ MA K DMSKAYDRVEWPFL +++ +GF W+S IM
Sbjct: 526 DNILVAHEMIHSLRTNDRISKEHMAFKTDMSKAYDRVEWPFLETMMTALGFNNKWISWIM 585
Query: 177 TCVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGI 236
CV++V +S+++NG P P RG+RQGDPLSP LF+LC E ++NKA A + GI
Sbjct: 586 NCVTSVSYSVLINGQPYGHIIPTRGIRQGDPLSPALFVLCTEALIHILNKAEQAGKITGI 645
Query: 237 KIARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNV 296
+ ++HLLFADD+++ +AT QE + L+ Y Q SGQ+INL+KS ++ +NV
Sbjct: 646 QFQDKKVSVNHLLFADDTLLMCKATKQECEELMQCLSQYGQLSGQMINLNKSAITFGKNV 705
Query: 297 PDDCFHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRA 356
++ + KYLGLP + SK +F F+KE++ +L GW ++L +
Sbjct: 706 DIQIKDWIKSRSGISLEGGTGKYLGLPECLSGSKRDLFGFIKEKLQSRLTGWYAKTLSQG 765
Query: 357 GREVLVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKA 416
G+EVL+K++A A+P YVMSCF LP LC ++ ++ F+W +R +H LSW+ L
Sbjct: 766 GKEVLLKSIALALPVYVMSCFKLPKNLCQKLTTVMMDFWWNSMQQKRKIHWLSWQRLTLP 825
Query: 417 KDNGRLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYA 476
KD G GF+D + FN AL+AK WR+ SL +V+++ YF A RG RPSYA
Sbjct: 826 KDQGGFGFKDLQCFNQALLAKQAWRVLQEKGSLFSRVFQSRYFSNSDFLSATRGSRPSYA 885
Query: 477 WSSIMRSGEIFATGGMWNIGNGKSVKAWSDKWV-PGTDTLVYSVGLAQHFGVSL-VSDLM 534
W SI+ E+ G IGNG+ W+DKW+ G++ + + V L VS L+
Sbjct: 886 WRSILFGRELLMQGLRTVIGNGQKTFVWTDKWLHDGSNR--RPLNRRRFINVDLKVSQLI 943
Query: 535 LPGLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFL--- 591
P WN ++ + P I+ PL +D W S +G YS K+GY+FL
Sbjct: 944 DPTSRNWNLNMLRDL-FPWKDVEIILKQRPLFFKEDSFCWLHSHNGLYSVKTGYEFLSKQ 1002
Query: 592 -RHGAEAVVASSSQVPSLPRADWRK---------LWKA-DALLPVRAYLHARGLVTDPTC 640
H V SL W LWKA +PV L RG+ +D C
Sbjct: 1003 VHHRLYQEAKVKPSVNSLFDKIWNLHTAPKIRIFLWKALHGAIPVEDRLRTRGIRSDDGC 1062
Query: 641 PRCLVALETVQHALLFCPMVQPIW---FASSLGFRLAHECRVHEFVSDFMQVADMSSLGA 697
C ET+ H L CP+ + +W SS G ++ V+ +S + + + L
Sbjct: 1063 LMCDTENETINHILFECPLARQVWAITHLSSAGSEFSNS--VYTNMSRLIDLTQQNDLPH 1120
Query: 698 FLT-----LLYAIWMARNELCFKEKASTLEQILTHAASLTPLPPLEGPMVPQVHRPN--- 749
L +L+ +W RN L F+ K S ++ A Q H N
Sbjct: 1121 HLRFVSPWILWFLWKNRNALLFEGKGSITTTLVDKA-----YEAYHEWFSAQTHMQNDEK 1175
Query: 750 -----TWSRPHPGTFKINFDAAMAPTGD-AGFGLVARNSRGEVLAAACSSHGPAASSLLA 803
W P PG K N A + +G V R+S+G+VL + S S A
Sbjct: 1176 HLKITKWCPPLPGELKCNIGFAWSKQHHFSGASWVVRDSQGKVLLHSRRSFNEVHSPYSA 1235
Query: 804 EGLSFRWALGLAMDLGFRRVVF 825
+ S+ WAL F RV+F
Sbjct: 1236 KIRSWEWALESMTHHHFDRVIF 1257
>pir||T00833 RNA-directed DNA polymerase homolog T13L16.7 - Arabidopsis thaliana
(fragment)
Length = 1365
Score = 481 bits (1238), Expect = e-134
Identities = 306/862 (35%), Positives = 439/862 (50%), Gaps = 51/862 (5%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SII----NKT 56
+V A+F ++ APG DG ALF+Q+ W D V H L + G ++ N T
Sbjct: 431 EVYNAVFSINKESAPGPDGFTALFFQQHW----DLVKHQILTEIFGFFETGVLPQDWNHT 486
Query: 57 LLVLIPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLIT 116
+ LIPK+ P + R ISLC+V++KII+K L RLK LP ++ +Q+AFVP RLI+
Sbjct: 487 HICLIPKITSPQRMSDLRPISLCSVLYKIISKILTQRLKKHLPAIVSTTQSAFVPQRLIS 546
Query: 117 DNALVAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIM 176
DN LVA E H ++ MA K DMSKAYDRVEWPFL +++ +GF W+S IM
Sbjct: 547 DNILVAHEMIHSLRTNDRISKEHMAFKTDMSKAYDRVEWPFLETMMTALGFNNKWISWIM 606
Query: 177 TCVSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGI 236
CV++V +S+++NG P P RG+RQGDPLSP LF+LC E ++NKA A + GI
Sbjct: 607 NCVTSVSYSVLINGQPYGHIIPTRGIRQGDPLSPALFVLCTEALIHILNKAEQAGKITGI 666
Query: 237 KIARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNV 296
+ ++HLLFADD+++ +AT QE + L+ Y Q SGQ+INL+KS ++ +NV
Sbjct: 667 QFQDKKVSVNHLLFADDTLLMCKATKQECEELMQCLSQYGQLSGQMINLNKSAITFGKNV 726
Query: 297 PDDCFHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRA 356
++ + KYLGLP + SK +F F+KE++ +L GW ++L +
Sbjct: 727 DIQIKDWIKSRSGISLEGGTGKYLGLPECLSGSKRDLFGFIKEKLQSRLTGWYAKTLSQG 786
Query: 357 GREVLVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKA 416
G+EVL+K++A A+P YVMSCF LP LC ++ ++ F+W +R +H LSW+ L
Sbjct: 787 GKEVLLKSIALALPVYVMSCFKLPKNLCQKLTTVMMDFWWNSMQQKRKIHWLSWQRLTLP 846
Query: 417 KDNGRLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYA 476
KD G GF+D + FN AL+AK WR+ SL +V+++ YF A RG RPSYA
Sbjct: 847 KDQGGFGFKDLQCFNQALLAKQAWRVLQEKGSLFSRVFQSRYFSNSDFLSATRGSRPSYA 906
Query: 477 WSSIMRSGEIFATGGMWNIGNGKSVKAWSDKWV-PGTDTLVYSVGLAQHFGVSL-VSDLM 534
W SI+ E+ G IGNG+ W+DKW+ G++ + + V L VS L+
Sbjct: 907 WRSILFGRELLMQGLRTVIGNGQKTFVWTDKWLHDGSNR--RPLNRRRFINVDLKVSQLI 964
Query: 535 LPGLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFL--- 591
P WN ++ + P I+ PL +D W S +G YS K+GY+FL
Sbjct: 965 DPTSRNWNLNMLRDL-FPWKDVEIILKQRPLFFKEDSFCWLHSHNGLYSVKTGYEFLSKQ 1023
Query: 592 -RHGAEAVVASSSQVPSLPRADWRK---------LWKA-DALLPVRAYLHARGLVTDPTC 640
H V SL W LWKA +PV L RG+ +D C
Sbjct: 1024 VHHRLYQEAKVKPSVNSLFDKIWNLHTAPKIRIFLWKALHGAIPVEDRLRTRGIRSDDGC 1083
Query: 641 PRCLVALETVQHALLFCPMVQPIW---FASSLGFRLAHECRVHEFVSDFMQVADMSSLGA 697
C ET+ H L CP+ + +W SS G ++ V+ +S + + + L
Sbjct: 1084 LMCDTENETINHILFECPLARQVWAITHLSSAGSEFSNS--VYTNMSRLIDLTQQNDLPH 1141
Query: 698 FLT-----LLYAIWMARNELCFKEKASTLEQILTHAASLTPLPPLEGPMVPQVHRPN--- 749
L +L+ +W RN L F+ K S ++ A Q H N
Sbjct: 1142 HLRFVSPWILWFLWKNRNALLFEGKGSITTTLVDKA-----YEAYHEWFSAQTHMQNDEK 1196
Query: 750 -----TWSRPHPGTFKINFDAAMAPTGD-AGFGLVARNSRGEVLAAACSSHGPAASSLLA 803
W P PG K N A + +G V R+S+G+VL + S S A
Sbjct: 1197 HLKITKWCPPLPGELKCNIGFAWSKQHHFSGASWVVRDSQGKVLLHSRRSFNEVHSPYSA 1256
Query: 804 EGLSFRWALGLAMDLGFRRVVF 825
+ S+ WAL F RV+F
Sbjct: 1257 KIRSWEWALESMTHHHFDRVIF 1278
>gb|AAD24831.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25408166|pir||G84721 hypothetical protein
At2g31520 [imported] - Arabidopsis thaliana
Length = 1524
Score = 476 bits (1224), Expect = e-132
Identities = 306/929 (32%), Positives = 459/929 (48%), Gaps = 61/929 (6%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
++ +A+ Q+ KAPG DGL A FY+ W I+G DV + IN T + +
Sbjct: 588 EIYDAICQIGDDKAPGPDGLTARFYKNCWDIVGYDVILEVKKFFETSFMKPSINHTNICM 647
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK+ P + +R I+LCNV++K+I+K L NRLK L ++ +SQ AF+PGR+I DN +
Sbjct: 648 IPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPGRIINDNVM 707
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+A E H +K + MA+K D+SKAYDRVEW FL + ++ GF W+ IM V
Sbjct: 708 IAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWIGWIMAAVK 767
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V +S+++NG+P P RG+RQGDPLSPYLFILCG++ S LIN + L G++I
Sbjct: 768 SVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASSGDLRGVRIGN 827
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
AP I+HL FADDS+ F +A V+ ++K + YE SGQ IN+ KS ++ V
Sbjct: 828 GAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITFGSRVYGST 887
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+L+Q+L + KYLGLP G+ K ++F ++ +RV K+ W R L AG+E+
Sbjct: 888 QSKLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWSARFLSPAGKEI 947
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
++K+VA A+P Y MSCF LP G+ ++IE ++ F+W +RG+ ++WK L +K G
Sbjct: 948 MLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRLQYSKKEG 1007
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFRD FN AL+AK WR+ P SL +V +A YF +I AK + SY W+S+
Sbjct: 1008 GLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKDVSILDAKVRKQQSYGWASL 1067
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYS-----VGLAQHFGVSLVSDLM- 534
+ + G IG+G++++ G D +V S + + + +++L
Sbjct: 1068 LDGIALLKKGTRHLIGDGQNIRI-------GLDNIVDSHPPRPLNTEETYKEMTINNLFE 1120
Query: 535 -LPGLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRH 593
W+ I+ I I L D + W +++ G Y+ +SGY L H
Sbjct: 1121 RKGSYYFWDDSKISQFVDQSDHGFIHRIYLAKSKKPDKIIWNYNTTGEYTVRSGYWLLTH 1180
Query: 594 GAEAVVASSSQ----------------VPSLPRADWRKLWKADALLPVRAYLHARGLVTD 637
+ + + +P L WR L +A L L RG+ D
Sbjct: 1181 DPSTNIPAINPPHGSIDLKTRIWNLPIMPKLKHFLWRALSQA---LATTERLTTRGMRID 1237
Query: 638 PTCPRCLVALETVQHALLFCPMVQPIWFASSLGFRLAHECRVHEF------VSDFMQVAD 691
P CPRC E++ HAL CP W+ S + ++ ++F + +F+Q
Sbjct: 1238 PICPRCHRENESINHALFTCPFATMAWWLSDSSL-IRNQLMSNDFEENISNILNFVQDTT 1296
Query: 692 MSSLGAFLT--LLYAIWMARNELCFKE------------KASTLEQI-LTHAASLTPLPP 736
MS L L++ IW ARN + F + KA T + + T + TP P
Sbjct: 1297 MSDFHKLLPVWLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNATQSHKKTPSPT 1356
Query: 737 LEGPMVPQVHRPNTWSRPHPGTFKINFDAAM-APTGDAGFGLVARNSRGEVLAAACSSHG 795
+ W P K NFDA +A G + RN G ++
Sbjct: 1357 RQ-----IAENKIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLA 1411
Query: 796 PAASSLLAEGLSFRWALGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRL 855
++ L AE + AL G+ +V E DC L S L + + D
Sbjct: 1412 HTSNPLEAETKALLAALQQTWIRGYTQVFMEGDCQTLINLINGISFHSSLANHLEDISFW 1471
Query: 856 SSGFDAFDFSFIRRTGNSVADALAKRSFT 884
++ F + F FIRR GN +A LAK T
Sbjct: 1472 ANKFASIQFGFIRRKGNKLAHVLAKYGCT 1500
>gb|AAD20714.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25412331|pir||G84649 hypothetical protein
At2g25550 [imported] - Arabidopsis thaliana
Length = 1750
Score = 473 bits (1218), Expect = e-132
Identities = 305/929 (32%), Positives = 458/929 (48%), Gaps = 61/929 (6%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
++ +A+ Q+ KAPG DGL A FY+ W I+G DV + IN T + +
Sbjct: 814 EIYDAICQIGDDKAPGPDGLTARFYKNCWDIVGYDVILEVKKFFETSFMKPSINHTNICM 873
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK+ P + +R I+LCNV++K+I+K L NRLK L ++ +SQ AF+PGR+I DN +
Sbjct: 874 IPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPGRIINDNVM 933
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+A E H +K + MA+K D+SKAYDRVEW FL + ++ GF W+ IM V
Sbjct: 934 IAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCNKWIGWIMAAVK 993
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V +S+++NG+P P RG+RQGDPLSPYLFILCG++ S LIN + L G++I
Sbjct: 994 SVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASSGDLRGVRIGN 1053
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
AP I+HL FADDS+ F +A V+ ++K + YE SGQ IN+ KS ++ V
Sbjct: 1054 GAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSMITFGSRVYGST 1113
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
L+Q+L + KYLGLP G+ K ++F ++ +RV K+ W R L AG+E+
Sbjct: 1114 QSRLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWSARFLSPAGKEI 1173
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
++K+VA A+P Y MSCF LP G+ ++IE ++ F+W +RG+ ++WK L +K G
Sbjct: 1174 MLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRLQYSKKEG 1233
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFRD FN AL+AK WR+ P SL +V +A YF +I AK + SY W+S+
Sbjct: 1234 GLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKDVSILDAKVRKQQSYGWASL 1293
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYS-----VGLAQHFGVSLVSDLM- 534
+ + G IG+G++++ G D +V S + + + +++L
Sbjct: 1294 LDGIALLKKGTRHLIGDGQNIRI-------GLDNIVDSHPPRPLNTEETYKEMTINNLFE 1346
Query: 535 -LPGLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRH 593
W+ I+ I I L D + W +++ G Y+ +SGY L H
Sbjct: 1347 RKGSYYFWDDSKISQFVDQSDHGFIHRIYLAKSKKPDKIIWNYNTTGEYTVRSGYWLLTH 1406
Query: 594 GAEAVVASSSQ----------------VPSLPRADWRKLWKADALLPVRAYLHARGLVTD 637
+ + + +P L WR L +A L L RG+ D
Sbjct: 1407 DPSTNIPAINPPHGSIDLKTRIWNLPIMPKLKHFLWRALSQA---LATTERLTTRGMRID 1463
Query: 638 PTCPRCLVALETVQHALLFCPMVQPIWFASSLGFRLAHECRVHEF------VSDFMQVAD 691
P+CPRC E++ HAL CP W S + ++ ++F + +F+Q
Sbjct: 1464 PSCPRCHRENESINHALFTCPFATMAWRLSDSSL-IRNQLMSNDFEENISNILNFVQDTT 1522
Query: 692 MSSLGAFLT--LLYAIWMARNELCFKE------------KASTLEQI-LTHAASLTPLPP 736
MS L L++ IW ARN + F + KA T + + T + TP P
Sbjct: 1523 MSDFHKLLPVWLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNATQSHKKTPSPT 1582
Query: 737 LEGPMVPQVHRPNTWSRPHPGTFKINFDAAM-APTGDAGFGLVARNSRGEVLAAACSSHG 795
+ W P K NFDA +A G + RN G ++
Sbjct: 1583 RQ-----IAENKIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLA 1637
Query: 796 PAASSLLAEGLSFRWALGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRL 855
++ L AE + AL G+ +V E DC L S L + + D
Sbjct: 1638 HTSNPLEAETKALLAALQQTWIRGYTQVFMEGDCQTLINLINGISFHSSLANHLEDISFW 1697
Query: 856 SSGFDAFDFSFIRRTGNSVADALAKRSFT 884
++ F + F FIR+ GN +A LAK T
Sbjct: 1698 ANKFASIQFGFIRKKGNKLAHVLAKYGCT 1726
>gb|AAQ19327.1| bZIP-like protein [Oryza sativa (japonica cultivar-group)]
Length = 2367
Score = 473 bits (1216), Expect = e-131
Identities = 296/899 (32%), Positives = 450/899 (49%), Gaps = 48/899 (5%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
++ +A+FQ+ P K+PG DG PA FYQ+ W I D+ + + +N T +VL
Sbjct: 1166 EISQAIFQIGPLKSPGPDGFPARFYQRNWGTIKADIIGAVRRFFQTGLMPEGVNDTAIVL 1225
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK ++P+ FR ISLCNV++K+++K L NRL+ IL DL+ Q+AFV GR+ITDNAL
Sbjct: 1226 IPKKEQPVDLRDFRPISLCNVVYKVVSKCLVNRLRPILDDLVSVEQSAFVQGRMITDNAL 1285
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+AFECFH M+K + A KLD+SKAYDRV+W FL + ++GF WV+ IM CV+
Sbjct: 1286 LAFECFHAMQKNKKANHAACAYKLDLSKAYDRVDWRFLEMAMNKLGFARRWVNWIMKCVT 1345
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V + + NG + F P RGLRQGDPL P+LF+ + S L+ + V +SL K+ R
Sbjct: 1346 SVRYMVKFNGTLLQSFAPTRGLRQGDPLLPFLFLFVADGLSLLLKEKVAQNSLTPFKVCR 1405
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
AP ISHLLFADD+++F +A +EA +K +L+SY +GQ+IN K +
Sbjct: 1406 AAPGISHLLFADDTLLFFKAHQREAEVVKEVLSSYAMGTGQLINPAKCSILMGGASTPAV 1465
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+ ++L V+ D+YLG PT G+ F ++ ++WK++ W E L G+EV
Sbjct: 1466 SEAISEILQVERDRFEDRYLGFPTPEGRMHKGRFQSLQAKIWKRVIQWGENHLSTGGKEV 1525
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
L+KAV AIP YVM F LP+ + + + + F+W +R H +W L K K G
Sbjct: 1526 LIKAVIQAIPVYVMGIFKLPESVIDDLTKLTKNFWWDSMNGQRKTHWKAWDSLTKPKSLG 1585
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFRD+++FN AL+A+ WR+ + P+SL +V +A YFP G++ G S AW SI
Sbjct: 1586 GLGFRDYRLFNQALLARQAWRLITYPDSLCARVLKAKYFPHGSLIDTSFGSNSSPAWRSI 1645
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSLVSDLMLPGLLA 540
++ G +W +GNG S++ W D W+P + G A + + VSDL+ +
Sbjct: 1646 EYGLDLLKKGIIWRVGNGNSIRIWRDSWLPRDHSRRPITGKA-NCRLKWVSDLITED-GS 1703
Query: 541 WNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHGAEAVVA 600
W+ I+ A I+ I + +D + W +G +S +S Y+ +
Sbjct: 1704 WDVPKIHQYFHNLDAEVILNICISSRSEEDFIAWHPDKNGMFSVRSAYRLAAQLVNIEES 1763
Query: 601 SSSQVPSLPRADWRKLWKADALLPVRAYL--------------HARGLVTDPTCPRCLVA 646
SSS ++ +A W +WK V+ + R L C C
Sbjct: 1764 SSSGTNNINKA-WEMIWKCKVPQKVKIFAWRVASNCLATMVNKKKRKLEQSDMCQICDRE 1822
Query: 647 LETVQHALLFCPMVQPIWF----ASSLGFRLAHECRVHEFVSDFMQVADMSSLGAFLTLL 702
E HAL C +W + S+ + ++ D ++ FL L
Sbjct: 1823 NEDDAHALCRCIQASQLWSCMHKSGSVSVDIKASVLGRFWLFDCLEKIPEYEQAMFLMTL 1882
Query: 703 YAIWMARNELCFKEKASTLE--------------QI-------LTHAASLTPLPPLEGPM 741
+ W RNEL + A E QI L + PL+G
Sbjct: 1883 WRNWYVRNELIHGKSAPPTETSQRFIQSYVDLLFQIRQAPQADLVKGKHVVRTVPLKGGP 1942
Query: 742 VPQV---HRPNTWSRPHPGTFKINFDAAM-APTGDAGFGLVARNSRGEVLAAACSSHGPA 797
+V H+P W RP G K+N D + A +G G G++ RNS G+++ +C
Sbjct: 1943 KYRVLNNHQP-CWERPKDGWMKLNVDGSFDASSGKGGLGMILRNSAGDIIFTSCKPLERC 2001
Query: 798 ASSLLAEGLSFRWALGLAMDLGFRRVVFEIDCLLLYEAWKRRG-GCSILDSLILDCRRL 855
+ L +E + L LA+ + E DC + + + G S+L ++I + R L
Sbjct: 2002 NNPLESELRACVEGLKLAIHWTLLPIQVETDCASVVQLLQGIGRDFSVLANIIHEARHL 2060
>gb|AAC33961.1| contains similarity to reverse trancriptase (Pfam: rvt.hmm, score:
42.57) [Arabidopsis thaliana] gi|7486711|pir||T01893
hypothetical protein F8M12.22 - Arabidopsis thaliana
Length = 1662
Score = 470 bits (1209), Expect = e-130
Identities = 294/894 (32%), Positives = 443/894 (48%), Gaps = 90/894 (10%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
++ A+ + KAPG DGL A FY+ W I+G DV IN T + +
Sbjct: 814 EIYNAICHIGDDKAPGPDGLTARFYKSCWEIVGPDVIKEVKIFFRTSYMKQSINHTNICM 873
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK+ P + +R I+LCNV++KII+K L RLK L ++ +SQ AF+PGRL+ DN +
Sbjct: 874 IPKITNPETLSDYRPIALCNVLYKIISKCLVERLKGHLDAIVSDSQAAFIPGRLVNDNVM 933
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+A E H +K + MA+K D+SKAYDRVEW FL + ++ GF +W+ IM V
Sbjct: 934 IAHEMMHSLKTRKRVSQSYMAVKTDVSKAYDRVEWNFLETTMRLFGFSETWIKWIMGAVK 993
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V++S+++NG P QP RG+RQGDPLSPYLFILC ++ + LI V + GI+I
Sbjct: 994 SVNYSVLVNGIPHGTIQPQRGIRQGDPLSPYLFILCADILNHLIKNRVAEGDIRGIRIGN 1053
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
P ++HL FADDS+ F ++ V+ ++K + YE SGQ IN+ KS ++ V
Sbjct: 1054 GVPGVTHLQFADDSLFFCQSNVRNCQALKDVFDVYEYYSGQKINMSKSMITFGSRVHGTT 1113
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+ L+ +L +++ KYLGLP G+ K +FN++ ERV K+ W + L AG+E+
Sbjct: 1114 QNRLKNILGIQSHGGGGKYLGLPEQFGRKKRDMFNYIIERVKKRTSSWSAKYLSPAGKEI 1173
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
++K+VA ++P Y MSCF LP + ++IE ++ F+W + +R + ++WK L +K G
Sbjct: 1174 MLKSVAMSMPVYAMSCFKLPLNIVSEIEALLMNFWWEKNAKKREIPWIAWKRLQYSKKEG 1233
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFRD FN AL+AK WR+ +NP SL ++ +A YF +I AKR SY W+S+
Sbjct: 1234 GLGFRDLAKFNDALLAKQVWRMINNPNSLFARIMKARYFREDSILDAKRQRYQSYGWTSM 1293
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSLVSDLMLPGLLA 540
+ ++ G + +G+GK T + Y
Sbjct: 1294 LAGLDVIKKGSRFIVGDGK------------TGSYRY----------------------- 1318
Query: 541 WNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHGAEAVVA 600
WN LI+ + P R +M L + H D L W +SS G Y+ +
Sbjct: 1319 WNAHLISQLVSPDDHRFVMNHHLSRIVHQDKLVWNYSSSGDYT---------------LW 1363
Query: 601 SSSQVPSLPRADWRKLWKADALLPVRAYLHARGLVTDPTCPRCLVALETVQHALLFCPMV 660
+P + WR + KA LP R+ L RG+ DP CPRC ET+ H L CP
Sbjct: 1364 KLPIIPKIKYMLWRTISKA---LPTRSRLLTRGMDIDPHCPRCPTEEETINHVLFTCPYA 1420
Query: 661 QPIWFASSLGFRLAHECR--VHEFVSDFMQVADMSSLG-----AFLTLLYAIWMARNELC 713
IW S+ + H E +S + ++L A L++ +W ARN L
Sbjct: 1421 ASIWGLSNFPWLPGHTFSQDTEENISFLINSFSNNTLNTEQRLAPFWLIWRLWKARNNLV 1480
Query: 714 FKEKASTLEQILTHAAS-----LTPLPPLEGPMV--PQVHRPNT-WSRPHPGTFKINFDA 765
F + + + +++T + L + E V HR N W +P K NFDA
Sbjct: 1481 FNKFSESCSRVVTQTEAEVNEWLQSVNRREDASVLTRSSHRANVRWKKPVFPLVKCNFDA 1540
Query: 766 AMAPTGDAGFGLVARNSRGEVLAAACSSHGPAASSLLAEGLSFRWALGLAMDLGFRRVVF 825
G +++ G ++ P +LL A+ A G++ V F
Sbjct: 1541 GFT-------GNNTQSTGGWII--------PETKALLI-------AMQQAWVRGYKCVQF 1578
Query: 826 EIDCLLLYEAWKRRGGCSILDSLILDCRRLSSGFDAFDFSFIRRTGNSVADALA 879
E DC +L +A + SL+ D +S F F+F R N+ A LA
Sbjct: 1579 EGDCEILIKAINGAISRCEITSLLRDVDFWASRFSTVVFTFTNRLCNNTAHLLA 1632
>emb|CAE04866.2| OSJNBa0086O06.14 [Oryza sativa (japonica cultivar-group)]
gi|50928373|ref|XP_473714.1| OSJNBa0086O06.14 [Oryza
sativa (japonica cultivar-group)]
Length = 1205
Score = 469 bits (1207), Expect = e-130
Identities = 289/805 (35%), Positives = 423/805 (51%), Gaps = 40/805 (4%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
++ +ALFQ+ P KAPG DG PA F+Q+ W ++ DV + +N T++V+
Sbjct: 420 EISDALFQIGPLKAPGPDGFPARFFQRNWGVLKRDVIEGVREFFETGEWKEGMNDTVIVM 479
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK P+ FR +SLCNVI+K++ K L NRL+ +L ++I E+Q+AFVPGR+ITDNAL
Sbjct: 480 IPKTNAPVEMKDFRPVSLCNVIYKVVAKCLVNRLRPLLQEIISETQSAFVPGRMITDNAL 539
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
VAFECFH + K ALKLD+SKAYDRV+W FL LQ++GF W IM+CV+
Sbjct: 540 VAFECFHSIHKCTRESQDFCALKLDLSKAYDRVDWGFLDGALQKLGFGNIWRKWIMSCVT 599
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V +S+ LNG EPF P RGLR+GDPL+PYLF+ + S ++ + + +K+ R
Sbjct: 600 SVRYSVRLNGNMLEPFYPTRGLREGDPLNPYLFLFIADGLSNILQRRRDERQIQPLKVCR 659
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
+AP +SHLLFADDS++F +A V +A IK L YE+ +GQ+IN + L S P +
Sbjct: 660 SAPGVSHLLFADDSLLFFKAEVIQATRIKEALDLYERCTGQLINPKECSLLFSALCPQER 719
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
++ +L V+ DK LGLPT G+ K + F +KER K+L W ER L AG+E
Sbjct: 720 QDGIKAVLQVERTCFDDKCLGLPTPDGRMKAEQFQPIKERFEKRLTDWSERFLSLAGKEA 779
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
L+K+VA A+P+Y M F +P+ C + E ++ F+WG + + +H ++W+ L K G
Sbjct: 780 LIKSVAQALPTYTMGVFKMPERFCEEYEQLVRNFWWGHEKGEKKVHWIAWEKLTSPKLLG 839
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFRD + FN AL+A+ WR+ +P+SL +V +A Y+P GTI S W I
Sbjct: 840 GLGFRDIRCFNQALLARQAWRLIESPDSLCARVLKAKYYPNGTITDTAFPSVSSPTWKGI 899
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSLVSDLMLPGLLA 540
+ E+ G +W IG+G K W + WV + L + V V +L++
Sbjct: 900 VHGLELLKKGLIWRIGDGSKTKIWRNHWVAHGENLKILEKKTWN-RVIYVRELIVTDTKT 958
Query: 541 WNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHGAE--AV 598
WN LI I A I+ I +P +D W + G +S +S Y+ + A +
Sbjct: 959 WNEPLIRHIIREEDADEILKIRIPQREEEDFPAWHYEKTGIFSVRSVYRLAWNLARKTSE 1018
Query: 599 VASSSQVPSLPRADWRKLWKADALLPVRAYL--------------HARGLVTDPTCPRCL 644
ASSS + R W +WKA+ VR + R + TCP C
Sbjct: 1019 QASSSSGGADGRKIWDNVWKANVQPKVRVFAWKLAQDRLATWENKKKRKIEMFGTCPICG 1078
Query: 645 VALETVQHALLFCPMVQPIWFASSLGFRLAHECRVHEFVSDFMQVADMSSLGAFLTLLYA 704
ET HA + C +L L R H + D + M+ L LL
Sbjct: 1079 QKEETGFHATVEC----------TLAKALRASLREHWTLPD-ESLFSMTGPDWLLVLL-- 1125
Query: 705 IWMARNELCFKEKAS-TLEQILTHAASLTPL--PPLEGPMVPQVH-RPNTWSRPHPGTFK 760
+ L ++KA + + P+ P + Q++ WS P G+ K
Sbjct: 1126 -----DRLSSEKKAQLPARKHMEDIKGKGPMFQDPCQKEQTCQLNAEKEKWSCPPDGSAK 1180
Query: 761 INFDAA-MAPTGDAGFGLVARNSRG 784
+N DAA TG+A G++ R+ RG
Sbjct: 1181 LNVDAAYRTETGEASAGIIIRDCRG 1205
>pir||G96509 protein F27F5.21 [imported] - Arabidopsis thaliana
gi|7767672|gb|AAF69169.1| F27F5.21 [Arabidopsis
thaliana]
Length = 1023
Score = 464 bits (1193), Expect = e-129
Identities = 258/644 (40%), Positives = 363/644 (56%), Gaps = 23/644 (3%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+V ALF MHP KAPG DG+ ALF+QK W I D+ L + +N T + L
Sbjct: 282 EVRAALFLMHPEKAPGPDGMTALFFQKPWATIKMDMLFLVNSFLQDGVFDKQLNTTNICL 341
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK ++P + R ISLCNV +KII+K L RLK ILP LI E+ +AFV GRLI+DN L
Sbjct: 342 IPKTERPTRMTELRPISLCNVGYKIISKILCQRLKKILPSLISETHSAFVEGRLISDNIL 401
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+A E FH ++ S MA+K DMSKAYDRVEW F+ +L++MGF W+S IM C++
Sbjct: 402 IAQEMFHGLRTNPSCKGKFMAIKTDMSKAYDRVEWNFIEELLKKMGFCEKWISWIMWCIT 461
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
TV + ++LNG P+ P RGLRQGDPLSPYLFILC EV A I KA + GIK+A
Sbjct: 462 TVQYRVLLNGQPRGLIVPKRGLRQGDPLSPYLFILCTEVLIANIRKAEQEKLITGIKVAT 521
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
+P +SHLLFA+DS+ F +A ++ I IL YE SGQ+IN KS + V DD
Sbjct: 522 ASPSVSHLLFANDSLFFCKANKEQCGVILGILKQYEAVSGQMINFSKSSIQFGHKVGDDI 581
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
E++ L + ++ YLGLP +G SKT+IF+FV++R+ ++ GW + L + G+EV
Sbjct: 582 KAEIKSALGIHSIGGMGSYLGLPESLGGSKTRIFSFVRDRLQTRINGWSAKFLSKGGKEV 641
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
++K+V +P+YVMSCF LP + +++ +++F+W D RGMH ++W LC +K +G
Sbjct: 642 MIKSVVAVLPTYVMSCFRLPKTITSKLTSAVAKFWWSSDGESRGMHWMAWNKLCSSKADG 701
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFR FNSAL+AK WR+ + P+SL KV++ YF + + Y PSY W SI
Sbjct: 702 GLGFRSVNDFNSALLAKQLWRLITVPDSLFAKVFKGRYFRKSNPLDKLKSYSPSYGWRSI 761
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSL-VSDLMLPGLL 539
+ + + G + +G+G S+ W D W+P ++ SL V++L+
Sbjct: 762 ISARSLVKKGLINRVGSGASISVWEDPWIPAQFPRP-AISNGSVIDPSLKVNNLIDSRSN 820
Query: 540 AWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHGAEAVV 599
WN + +N I P I A+ L +D L K+ +A +
Sbjct: 821 FWNIDFLNEIFVPEDVLLISAMHLGSSTKEDSL----------DIKT--------LKAHI 862
Query: 600 ASSSQVPSLPRADWRKLWKADALLPVRAYLHARGLVTDPTCPRC 643
P L W+ L +PV L RG+ D C RC
Sbjct: 863 WKVQCPPKLRHFLWQVL---SGCVPVTENLRKRGINCDSGCARC 903
>gb|AAP52395.1| putative retroelement [Oryza sativa (japonica cultivar-group)]
gi|37531612|ref|NP_920108.1| putative retroelement [Oryza
sativa (japonica cultivar-group)]
Length = 1694
Score = 463 bits (1192), Expect = e-128
Identities = 247/631 (39%), Positives = 367/631 (58%), Gaps = 10/631 (1%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
++ ALFQ+ P KAPG DG P FYQ+ W I+ DD+ + S +N+T +VL
Sbjct: 1046 EIANALFQIGPLKAPGPDGFPGRFYQRNWAILKDDIVRAVQEFFSLGTMPSGVNETAIVL 1105
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK ++P FR ISLCNV++KI++K L NRL+ IL DL+ ++Q+AFVPGRLITDNAL
Sbjct: 1106 IPKTEQPQELKDFRPISLCNVVYKIVSKCLVNRLRPILDDLVSQNQSAFVPGRLITDNAL 1165
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+AFE FH++++ + N A KLD+SKAYDRV+W FL + ++GF WV IM C++
Sbjct: 1166 IAFEYFHHIQRNKNPENAYSAYKLDLSKAYDRVDWEFLEQAMVKLGFAHCWVKWIMACIT 1225
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
V +++ LNG F P RGLRQGDPLSP+LF+ + S L+ + V +++ I I R
Sbjct: 1226 LVRYAVKLNGTLLNTFAPSRGLRQGDPLSPFLFLFVADGLSFLLEEKVDQGAMNPIHICR 1285
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
AP + HLLFADD+ +F +A +A+ +K +L +Y +GQ+IN K + ++ P
Sbjct: 1286 HAPAVLHLLFADDTPLFFKAANDQAVVMKEVLETYASCTGQLINPSKCSIMFGQSSPTAV 1345
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
++++Q L + + DKYLG PT G+ F ++ER+WK++ W E+ L G+E+
Sbjct: 1346 QNQIKQTLQI-SNSFEDKYLGFPTPEGRMTKGKFQSLQERLWKRIMLWGEKFLSTGGKEI 1404
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
L+KAV A+P YVM F LP+ +C ++ ++ F+WG + R H SW L K G
Sbjct: 1405 LIKAVLQAMPVYVMGLFKLPESVCEELTKLVRNFWWGAEKGMRRTHWKSWDCLIAQKLKG 1464
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
+GFRDF++FN AL+A+ WR+ P+SL ++ +A+Y+P G++ G S W +I
Sbjct: 1465 GMGFRDFRLFNQALLARQAWRLLDRPDSLCARILKAIYYPNGSLIDTSFGGNASPGWRAI 1524
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYS---VGLAQHFGVSLVSDLMLPG 537
E+ G +W IGNGKSV+ W D W+P + YS + ++ + VSDL +
Sbjct: 1525 EYGLELLKKGIIWRIGNGKSVRLWRDPWIPRS----YSRRPISAKRNCRLRWVSDL-IDQ 1579
Query: 538 LLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHGAEA 597
+W+ + I+ P A AI+ I L DD + W G +S +S Y A
Sbjct: 1580 SGSWDTDKISQHFLPMDAEAILNIRLSSRLEDDFIAWHPDKLGRFSVRSAYHLAVALAHV 1639
Query: 598 VVASSSQVPSLPRADWRKLWKADALLPVRAY 628
SSS RA W LWK +A V+ +
Sbjct: 1640 DEGSSSSGNGNSRA-WNALWKCNAPQKVKIF 1669
>emb|CAB75484.1| putative protein [Arabidopsis thaliana] gi|11357921|pir||T47495
hypothetical protein F9K21.130 - Arabidopsis thaliana
Length = 851
Score = 462 bits (1189), Expect = e-128
Identities = 276/799 (34%), Positives = 406/799 (50%), Gaps = 38/799 (4%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
++ EA+ Q+ KAPG DGL A FY++ W I+G+DV + +N T + +
Sbjct: 53 EIFEAICQIGDDKAPGPDGLTARFYKQCWDIVGNDVIKEVKLFFESSHMKTSVNHTNICM 112
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK++ P + +R I+LCNV++K+I+K + NRLK L ++ +SQ AF+PGR+I DN +
Sbjct: 113 IPKIQNPQTLSDYRPIALCNVLYKVISKCMVNRLKAHLNSIVSDSQAAFIPGRIINDNVM 172
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+A E H +K + MA+K D+SKAYDRVEW FL + ++ GF W+ IM V
Sbjct: 173 IAHEIMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTMRLFGFCDKWIGWIMAAVK 232
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V +S+++NG+P P RG+RQGDPLSPYLFILCG++ S LI + + G++I
Sbjct: 233 SVHYSVLINGSPHGYISPTRGIRQGDPLSPYLFILCGDILSHLIKVKASSGDIRGVRIGN 292
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
AP I+HL FADDS+ F +A V+ ++K + YE SGQ IN+ KS ++ V
Sbjct: 293 GAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSLITFGSRVYGST 352
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
L+ LL + KYLGLP G+ K ++FN++ +RV ++ W + L AG+E+
Sbjct: 353 QTRLKTLLNIPNQGGGGKYLGLPEQFGRKKKEMFNYIIDRVKERTASWSAKFLSPAGKEI 412
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
L+K+VA A+P Y MSCF LP G+ ++IE ++ F+W +RG+ ++WK L +K G
Sbjct: 413 LLKSVALAMPVYAMSCFKLPQGIVSEIESLLMNFWWEKASNKRGIPWVAWKRLQYSKKEG 472
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFRD FN AL+AK WRI P SL +V +A YF +I AK + SY WSS+
Sbjct: 473 GLGFRDLAKFNDALLAKQAWRIIQYPNSLFARVMKARYFKDNSIIDAKTRSQQSYGWSSL 532
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSLVSDLMLPG-LL 539
+ + G + IG+GK+++ D V QH G+SL + G
Sbjct: 533 LSGIALLRKGTRYVIGDGKTIRLGIDNVVDSHPPRPLLTD-EQHNGLSLDNLFQHRGHSR 591
Query: 540 AWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHGAEAVV 599
W+ + I I L D L W ++S G Y+ +SGY H +
Sbjct: 592 CWDNAKLQTFVDQSDHDYIKRIYLSTRSKTDRLIWSYNSTGDYTVRSGYWLSTHDPSNTI 651
Query: 600 ASSSQ----------------VPSLPRADWRKLWKADALLPVRAYLHARGLVTDPTCPRC 643
+ ++ +P L WR L KA LP L RG+ DP CPRC
Sbjct: 652 PTMAKPHGSVDLKTKIWNLPIMPKLKHFLWRILSKA---LPTTDRLTTRGMRIDPGCPRC 708
Query: 644 LVALETVQHALLFCPMVQPIWFASSLGFRLAH--ECRVHEFVSDFM------QVADMSSL 695
E++ HAL CP W S + + + +S+ + + D L
Sbjct: 709 RRENESINHALFTCPFATMAWRLSDTPLYRSSILSNNIEDNISNILLLLQNTTITDSQKL 768
Query: 696 GAFLTLLYAIWMARNELCFKEKASTLEQILTHAASLT--------PLPPLEGPMVPQVHR 747
F LL+ IW ARN + F + + A + T P P
Sbjct: 769 IPF-WLLWRIWKARNNVVFNNLRESPSITVVRAKAETNEWLNATQTQGPRRLPKRTTAAG 827
Query: 748 PNTWSRPHPGTFKINFDAA 766
TW +P K NFDA+
Sbjct: 828 NTTWVKPQMPYIKCNFDAS 846
>gb|AAP54692.1| putative reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|37536206|ref|NP_922405.1| putative
reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|27311287|gb|AAO00713.1|
retrotransposon protein, putative, unclassified [Oryza
sativa (japonica cultivar-group)]
Length = 1557
Score = 462 bits (1189), Expect = e-128
Identities = 257/740 (34%), Positives = 392/740 (52%), Gaps = 40/740 (5%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
++ ALFQ+ P KAPG DG A FYQ+ W I +D+ + H I +N T +VL
Sbjct: 820 EIANALFQIGPLKAPGTDGFLARFYQRNWGCIKNDIVRAVKEFFHSGIMLEGVNDTAIVL 879
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK+++P FR ISLCNVI+K+++K L NRL+ L +LI ++Q+AFVP R+ITDNAL
Sbjct: 880 IPKIEQPRDLRDFRPISLCNVIYKVLSKCLVNRLRPFLDELISKNQSAFVPRRMITDNAL 939
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+AFECFH++++ + + A KLD+SKAYDRV+W FL + + GF WVS +MTC+S
Sbjct: 940 LAFECFHFIQRNKNPRSAACAYKLDLSKAYDRVDWRFLEQSMYKWGFSHCWVSWVMTCIS 999
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
TV +G P+LF+ + S L+ + V ++ I++
Sbjct: 1000 TV-------------------CDKGTRCDPFLFLFVADGLSLLLEEKVEQGAISPIRVCH 1040
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
AP ISHLLFADD+++F +A + ++ +IK + SY ++GQ+IN K + ++P
Sbjct: 1041 QAPGISHLLFADDTLLFFKADLSQSQAIKEVFGSYATSTGQLINPTKCSILFGNSLPIAS 1100
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+ L + + E DKYLGLPT G+ F ++ER+WKK+ W E L G+EV
Sbjct: 1101 RDAITNCLQIASTEFEDKYLGLPTPGGRMHKGRFQSLRERIWKKILQWGENYLSSGGKEV 1160
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
L+KAV AIP YVM F LP+ +C + + F+WG + +R H +WK L K+K G
Sbjct: 1161 LIKAVIQAIPVYVMGIFKLPESVCEDLTSLTRNFWWGAEKGKRKTHWKAWKSLTKSKSLG 1220
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGF+D ++FN AL+A+ WR+ NP+SL +V +A Y+P G+I G S W +I
Sbjct: 1221 GLGFKDIRLFNQALLARQAWRLIDNPDSLCARVLKAKYYPNGSIVDTSFGGNASPGWQAI 1280
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSLVSDLMLPGLLA 540
E+ G +W IGNG+SV+ W D W+P D + + + V+DLML +
Sbjct: 1281 EHGLELVKKGIIWRIGNGRSVRVWQDPWLP-RDLSRRPITPKNNCRIKWVADLMLDNGM- 1338
Query: 541 WNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHGAEAVVA 600
W+ IN I P I+ + +D + W G++S ++ Y+ + A+ +
Sbjct: 1339 WDANKINQIFLPVDVEIILKLRTSSRDEEDFIAWHPDKLGNFSVRTAYRLAENWAKEEAS 1398
Query: 601 SSSQVPSLPRADWRKLWKADA--------------LLPVRAYLHARGLVTDPTCPRCLVA 646
SSS ++ +A W LWK + LP R L TC C +
Sbjct: 1399 SSSSDVNIRKA-WELLWKCNVPSKVKIFTWRATSNCLPTWDNKKKRNLEISDTCVICGME 1457
Query: 647 LETVQHALLFCPMVQPIWFA----SSLGFRLAHECRVHEFVSDFMQVADMSSLGAFLTLL 702
E HAL CP + +W A + L R+ ++ + + + FL +L
Sbjct: 1458 KEDTMHALCRCPQAKHLWLAMKESNDLSLRMDDHLLGPSWLFNRLALLPDHEQPMFLMVL 1517
Query: 703 YAIWMARNELCFKEKASTLE 722
+ IW NE+ + + ++E
Sbjct: 1518 WRIWFVHNEIIHGKPSPSIE 1537
>emb|CAB78094.1| RNA-directed DNA polymerase-like protein [Arabidopsis thaliana]
gi|4538901|emb|CAB39638.1| RNA-directed DNA
polymerase-like protein [Arabidopsis thaliana]
gi|7485606|pir||T04018 hypothetical protein F17A8.60 -
Arabidopsis thaliana
Length = 1274
Score = 461 bits (1185), Expect = e-128
Identities = 315/918 (34%), Positives = 448/918 (48%), Gaps = 58/918 (6%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+++EALF + KAPG DG A F+ +W II DVS +N+T + L
Sbjct: 373 EIKEALFSISADKAPGPDGFSASFFHAYWDIIEADVSRDIRSFFVDSCLSPRLNETHVTL 432
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK+ P + +R I+LCNV +KI+ K L RL+ L +LI Q+AFVPGR I DN L
Sbjct: 433 IPKISAPRKVSDYRPIALCNVQYKIVAKILTRRLQPWLSELISLHQSAFVPGRAIADNVL 492
Query: 121 VAFECFHYMKKKISGCNGV--MALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTC 178
+ E H+++ +SG MA+K DMSKAYDR++W FL VL ++GF W+ +M C
Sbjct: 493 ITHEILHFLR--VSGAKKYCSMAIKTDMSKAYDRIKWNFLQEVLMRLGFHDKWIRWVMQC 550
Query: 179 VSTVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKI 238
V TV +S ++NG+PQ P RGLRQGDPLSPYLFILC EV S L KA + GI++
Sbjct: 551 VCTVSYSFLINGSPQGSVVPSRGLRQGDPLSPYLFILCTEVLSGLCRKAQEKGVMVGIRV 610
Query: 239 ARTAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPD 298
AR +P ++HLLFADD++ F + ++ IL YE ASGQ INL KS ++ S P
Sbjct: 611 ARGSPQVNHLLFADDTMFFCKTNPTCCGALSNILKKYELASGQSINLAKSAITFSSKTPQ 670
Query: 299 DCFHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGR 358
D ++ L + KYLGLP G+ K IF+ + +R+ ++ W R L AG+
Sbjct: 671 DIKRRVKLSLRIDNEGGIGKYLGLPEHFGRRKRDIFSSIVDRIRQRSHSWSIRFLSSAGK 730
Query: 359 EVLVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKD 418
++L+KAV ++PSY M CF LP LC QI+ +++RF+W +R M +SW L +
Sbjct: 731 QILLKAVLSSMPSYAMMCFKLPASLCKQIQSVLTRFWWDSKPDKRKMAWVSWDKLTLPIN 790
Query: 419 NGRLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYA-- 476
G LGFR+ + AK WRI P SL+ +V Y + PS+A
Sbjct: 791 EGGLGFREIE-------AKLSWRILKEPHSLLSRVLLGKYCNTSSFMDCSAS--PSFASH 841
Query: 477 -WSSIMRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSL-VSDLM 534
W I+ ++ G W+IG G S+ W++ W+ + +G L V DL+
Sbjct: 842 GWRGILAGRDLLRKGLGWSIGQGDSINVWTEAWLSPSSPQT-PIGPPTETNKDLSVHDLI 900
Query: 535 LPGLLAWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHG 594
+ +WN E I P I I + +P D L W G Y+TK+GY
Sbjct: 901 CHDVKSWNVEAIR-KHLPQYEDQIRKITINALPLQDSLVWLPVKSGEYTTKTGY------ 953
Query: 595 AEAVVASSSQVPSLPRADWRK--------------LWKA-DALLPVRAYLHARGLVTDPT 639
A+ +S S +W+K LWKA LPV L R + + T
Sbjct: 954 --ALAKLNSFPASQLDFNWQKNIWKIHTSPKVKHFLWKAMKGALPVGEALSRRNIEAEVT 1011
Query: 640 CPRCLVALETVQHALLFCPMVQPIWFASSLGF---RLAHECRVHEFVSDFMQVA----DM 692
C RC E+ H +L CP + +W + + F H V VA +
Sbjct: 1012 CKRC-GQTESSLHLMLLCPYAKKVWELAPVLFNPSEATHSSVALLLVDAKRMVALPPTGL 1070
Query: 693 SSLGAFLTLLYAIWMARNELCFKEKASTLEQILTHAASLTPLPPLEGPMVPQVHRPNTWS 752
S + LL+ +W ARN L F + S E+ L A L +E ++ +H P+ S
Sbjct: 1071 GSAPLYPWLLWHLWKARNRLIF-DNHSCSEEGLVLKAILDARAWMEAQLL--IHHPSPIS 1127
Query: 753 -----RPHPGTFKINFDAAMAPTGDAGFGLVARNSRGEVLAAACSSHGPAASSLLAEGLS 807
P+ DAA +G G G ++ + SS S+L+AE L+
Sbjct: 1128 DYPSPTPNLKVTSCFVDAAWTTSGYCGMGWFLQDPYKVKIKENQSSSSFVGSALMAETLA 1187
Query: 808 FRWALGLAMDLGFRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRLSSGFDAFDFSFI 867
AL A+ G R++ DC L L L+ D R LS F F FI
Sbjct: 1188 VHLALVDALSTGVRQLNVFSDCKELISLLNSGKSIVELRGLLHDIRELSVSFTHLCFFFI 1247
Query: 868 RRTGNSVADALAKRSFTL 885
R N VAD+LAK + ++
Sbjct: 1248 PRLSNVVADSLAKSALSV 1265
>gb|AAD29058.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25411151|pir||H84465 hypothetical protein
At2g05200 [imported] - Arabidopsis thaliana
Length = 1229
Score = 451 bits (1161), Expect = e-125
Identities = 296/904 (32%), Positives = 455/904 (49%), Gaps = 39/904 (4%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
+V++A+F ++ +KAPG DG A FY +WHII DV +N+T + L
Sbjct: 327 EVKDAVFSINASKAPGPDGFTAGFYHSYWHIISTDVGREIRLFFTSKNFPRRMNETHIRL 386
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK P +R I+LCN+ +KI+ K + R+++ILP LI E+Q+AFVPGR+I+DN L
Sbjct: 387 IPKDLGPRKVADYRPIALCNIFYKIVAKIMTKRMQLILPKLISENQSAFVPGRVISDNVL 446
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+ E H+++ + + MA+K DMSKAYDRVEW FL VLQ+ GF + W+ ++ CV+
Sbjct: 447 ITHEVLHFLRTSSAKKHCSMAVKTDMSKAYDRVEWDFLKKVLQRFGFHSIWIDWVLECVT 506
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
+V +S ++NG PQ P RGLRQGDPLSP LFILC EV S L +A L G++++
Sbjct: 507 SVSYSFLINGTPQGKVVPTRGLRQGDPLSPCLFILCTEVLSGLCTRAQRLRQLPGVRVSI 566
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
P ++HLLFADD++ F+++ + + IL+ Y +ASGQ IN KS ++ S P
Sbjct: 567 NGPRVNHLLFADDTMFFSKSDPESCNKLSEILSRYGKASGQSINFHKSSVTFSSKTPRSV 626
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
+++++L ++ KYLGLP G+ K IF + +++ +K W R L +AG++V
Sbjct: 627 KGQVKRILKIRKEGGTGKYLGLPEHFGRRKRDIFGAIIDKIRQKSHSWASRFLSQAGKQV 686
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
++KAV ++P Y MSCF LP LC +I+ +++RF+W R ++W L K+ G
Sbjct: 687 MLKAVLASMPLYSMSCFKLPSALCRKIQSLLTRFWWDTKPDVRKTSWVAWSKLTNPKNAG 746
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFRD + N +L+AK WR+ ++PESL+ ++ Y + K +PS+ W SI
Sbjct: 747 GLGFRDIERCNDSLLAKLGWRLLNSPESLLSRILLGKYCHSSSFMECKLPSQPSHGWRSI 806
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLV-YSVGLAQHFGVSLVSDLMLPGLL 539
+ EI G W I NG+ V W+D W+ + LV L +H + VS L+ L
Sbjct: 807 IAGREILKEGLGWLITNGEKVSIWNDPWLSISKPLVPIGPALREHQDLR-VSALINQNTL 865
Query: 540 AWNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHGAEAVV 599
W+ I I P I +P P D L W G Y+++SGY +
Sbjct: 866 QWDWNKIAVI-LPNYENLIKQLPAPSSRGVDKLAWLPVKSGQYTSRSGYG---------I 915
Query: 600 ASSSQVPSLPRA--DWR-KLWKADAL--------------LPVRAYLHARGLVTDPTCPR 642
AS + +P +P+ +W+ LWK L LPV L R + C R
Sbjct: 916 ASVASIP-IPQTQFNWQSNLWKLQTLPKIKHLMWKAAMEALPVGIQLVRRHISPSAACHR 974
Query: 643 CLVALETVQHALLFCPMVQPIWFASSL-GFRLAHECRVHEFVSDFMQVADMSSLGAFLTL 701
C A E+ H C +W + L + + + +S + + G
Sbjct: 975 C-GAPESTTHLFFHCEFAAQVWELAPLQETTVPPGSSMLDALSLLKKAIILPPTGVTSAA 1033
Query: 702 LYAIWMARNELCFKEKASTLEQILTHAASLTPLPPLEGPMVPQVHRPNTWSRPHPGTFKI 761
L+ W+ + L ++ LP + P P S G F
Sbjct: 1034 LFP-WICGIYGKLGTMTRAILDALAWQSAQRCLPKTRNVVHPISQLPVLRS----GYFCF 1088
Query: 762 NFDAAMAPTGDAGFGLVARNSRGEVLAAACSSHG--PAASSLLAEGLSFRWALGLAMDLG 819
A +A + AG G V +++ A S G S+L AE + + AL A+ LG
Sbjct: 1089 VDAAWIAQSSLAGSGWVFQSATALEKETATYSAGCRRLPSALSAEAWAIKSALLHALQLG 1148
Query: 820 FRRVVFEIDCLLLYEAWKRRGGCSILDSLILDCRRLSSGFDAFDFSFIRRTGNSVADALA 879
++ D + +A + + L+++ R L F + F+FI R+ N++ADA A
Sbjct: 1149 RTDLMVLSDSKSVVDALTSNISINEIYGLLMEIRALRVSFHSLCFNFISRSANAIADATA 1208
Query: 880 KRSF 883
K SF
Sbjct: 1209 KLSF 1212
>gb|AAV32224.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|45267888|gb|AAS55787.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 1936
Score = 448 bits (1152), Expect = e-124
Identities = 252/678 (37%), Positives = 360/678 (52%), Gaps = 17/678 (2%)
Query: 1 DVEEALFQMHPTKAPGCDGLPALFYQKFWHIIGDDVSHFCLQVLHGDIS*SIINKTLLVL 60
++ +A+FQ+ P K+P DG PA FYQ+ W + D+ + +N T +VL
Sbjct: 1072 EIAQAIFQIGPLKSPRPDGFPARFYQRNWGTLKSDIILAVRNFFQSGLMPKGVNDTAIVL 1131
Query: 61 IPKVKKPMHANQFRLISLCNVIFKIITKTLANRLKIILPDLICESQTAFVPGRLITDNAL 120
IPK +P+ +R ISLCNV++K+++K L NRL+ IL DL+ + Q+AF+ GR+ITDNAL
Sbjct: 1132 IPKKDQPIDLKDYRPISLCNVVYKVVSKCLVNRLRPILDDLVSKEQSAFIQGRMITDNAL 1191
Query: 121 VAFECFHYMKKKISGCNGVMALKLDMSKAYDRVEWPFLASVLQQMGFPASWVSLIMTCVS 180
+AFECFH ++K + A KLD+SKAYDRV+W FL L ++GF WVS IM CV+
Sbjct: 1192 LAFECFHSIQKNKKANSAACAYKLDLSKAYDRVDWRFLELALNKLGFAHRWVSWIMLCVT 1251
Query: 181 TVDFSIMLNGAPQEPFQPHRGLRQGDPLSPYLFILCGEVFSALINKAVLASSLHGIKIAR 240
TV +S+ NG F P RGLRQG+PLSP+LF+ + S L+ + V +SL +KI R
Sbjct: 1252 TVRYSVKFNGTLLRSFAPTRGLRQGEPLSPFLFLFVADGLSLLLKEKVAQNSLTPLKICR 1311
Query: 241 TAPVISHLLFADDSVIFARATVQEALSIKAILASYEQASGQVINLDKSKLSCSRNVPDDC 300
AP IS+LLFADD+++F +A +EA +K +L +Y Q +GQ+IN K + P
Sbjct: 1312 QAPGISYLLFADDTLLFFKAEKKEAEVVKEVLTNYAQGTGQLINPAKCSILFGEASPSSV 1371
Query: 301 FHELRQLLTVKAVESYDKYLGLPTIIGKSKTQIFNFVKERVWKKLKGWKERSLYRAGREV 360
++R L V+ D+YLG PT G+ F ++ ++ K++ W E L G+E+
Sbjct: 1372 SEDIRNTLQVERDNFEDRYLGFPTPEGRMHKGRFQSLQAKIAKRVIQWGENFLSSGGKEI 1431
Query: 361 LVKAVAHAIPSYVMSCFLLPDGLCNQIEGMISRFYWGGDVTRRGMHCLSWKDLCKAKDNG 420
L+KAV AIP YVM F PD + +++ M F+WG D RR H +W L KAK NG
Sbjct: 1432 LIKAVIQAIPVYVMGLFKFPDSVYDELTKMTRNFWWGADNGRRRTHWRAWDSLTKAKING 1491
Query: 421 RLGFRDFKMFNSALVAKNWWRIKSNPESLMGKVYRAVYFPRGTIRGAKRGYRPSYAWSSI 480
LGFRD+K+FN AL+ + WR+ P SL +V +A YFP G++ S W I
Sbjct: 1492 GLGFRDYKLFNQALLTRQAWRLIEFPNSLCAQVLKAKYFPHGSLTDTTFSANASPTWHGI 1551
Query: 481 MRSGEIFATGGMWNIGNGKSVKAWSDKWVPGTDTLVYSVGLAQHFGVSLVSDLMLPGLLA 540
++ G +W IGNG SV+ W D W+P D V + + VSDL+
Sbjct: 1552 EYGLDLLKKGIIWRIGNGNSVRIWRDPWIP-RDLSRRPVSSKANCRLKWVSDLIAED-GT 1609
Query: 541 WNRELINFICCPPTARAIMAIPLPLVPHDDVLYWPFSSDGHYSTKSGYQFLRHGAEAVVA 600
W+ IN A I I + +D + W G +S +S Y+ A+
Sbjct: 1610 WDSAKINQYFLKIDADIIQKICISARLEEDFIAWHPDKTGRFSVRSAYKLALQLADMNNC 1669
Query: 601 SSSQVPSLPRADWRKLWKADALLPVRAYL--------------HARGLVTDPTCPRCLVA 646
SSS L ++ W +WK + VR + R L C C
Sbjct: 1670 SSSSSSRLNKS-WELIWKCNVPQKVRIFAWRVASNSLATMENKKKRNLERFDVCGICDRE 1728
Query: 647 LETVQHALLFCPMVQPIW 664
E HAL C +W
Sbjct: 1729 KEDAGHALCRCVHANSLW 1746
Score = 45.8 bits (107), Expect = 0.006
Identities = 37/137 (27%), Positives = 62/137 (45%), Gaps = 8/137 (5%)
Query: 751 WSRPHPGTFKINFDAAMAPTGD-AGFGLVARNSRGEVLAAACSSHGPAASSLLAEGLSFR 809
W RP G K+N D + + G G++ RN G V+ ++C S + L AE +
Sbjct: 1777 WERPRNGWMKLNVDGSFDINSEKGGIGMILRNCLGNVIFSSCRSLDSCSGPLEAELHACV 1836
Query: 810 WALGLAMDLGFRRVVFEIDC----LLLYEAWKRRGGCSILDSLILDCRRLSSGFDAFDFS 865
L LA+ + E DC LL K R S+L ++ + + L +G S
Sbjct: 1837 EGLHLALHWTLLPIQVETDCSSVIQLLNHPDKDR---SVLANIAQEAKSLMAGDRQIAIS 1893
Query: 866 FIRRTGNSVADALAKRS 882
++R+ N ++ LA ++
Sbjct: 1894 KVQRSQNVISHFLANKA 1910
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.325 0.139 0.444
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,586,618,282
Number of Sequences: 2540612
Number of extensions: 66892388
Number of successful extensions: 184755
Number of sequences better than 10.0: 1418
Number of HSP's better than 10.0 without gapping: 816
Number of HSP's successfully gapped in prelim test: 602
Number of HSP's that attempted gapping in prelim test: 181420
Number of HSP's gapped (non-prelim): 2218
length of query: 906
length of database: 863,360,394
effective HSP length: 137
effective length of query: 769
effective length of database: 515,296,550
effective search space: 396263046950
effective search space used: 396263046950
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 80 (35.4 bits)
Lotus: description of TM0154.14


