

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0153b.2
(402 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
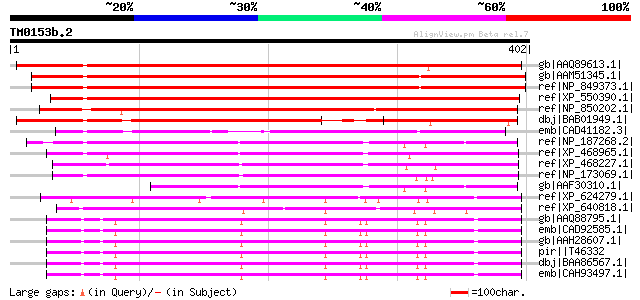
Score E
Sequences producing significant alignments: (bits) Value
gb|AAQ89613.1| At3g24470/MXP5_4 [Arabidopsis thaliana] gi|173812... 571 e-161
gb|AAM51345.1| unknown protein [Arabidopsis thaliana] gi|1660467... 566 e-160
ref|NP_849373.1| TMS membrane family protein / tumour differenti... 563 e-159
ref|XP_550390.1| unknown protein [Oryza sativa (japonica cultiva... 474 e-132
ref|NP_850202.1| TMS membrane family protein / tumour differenti... 325 1e-87
dbj|BAB01949.1| unnamed protein product [Arabidopsis thaliana] 298 2e-79
emb|CAD41182.3| OSJNBb0002J11.6 [Oryza sativa (japonica cultivar... 295 2e-78
ref|NP_187268.2| TMS membrane family protein / tumour differenti... 209 1e-52
ref|XP_468965.1| putative membrane protein [Oryza sativa (japoni... 205 2e-51
ref|XP_468227.1| putative tumor differentially expressed protein... 205 2e-51
ref|NP_173069.1| TMS membrane family protein / tumour differenti... 193 7e-48
gb|AAF30310.1| hypothetical protein [Arabidopsis thaliana] 183 7e-45
ref|XP_624279.1| PREDICTED: similar to ENSANGP00000002975 [Apis ... 149 1e-34
ref|XP_640818.1| hypothetical protein DDB0204014 [Dictyostelium ... 148 3e-34
gb|AAQ88795.1| GSVL396 [Homo sapiens] 141 4e-32
emb|CAD92585.1| RP3-425C14.2 [Homo sapiens] gi|57997080|emb|CAB7... 140 9e-32
gb|AAH28607.1| Tumor differentially expressed 2 [Homo sapiens] 140 9e-32
pir||T46332 hypothetical protein DKFZp434H0413.1 - human (fragment) 140 9e-32
dbj|BAA86567.1| KIAA1253 protein [Homo sapiens] 140 9e-32
emb|CAH93497.1| hypothetical protein [Pongo pygmaeus] gi|5572897... 139 1e-31
>gb|AAQ89613.1| At3g24470/MXP5_4 [Arabidopsis thaliana] gi|17381270|gb|AAL36053.1|
AT3g24470/MXP5_4 [Arabidopsis thaliana]
gi|42565162|ref|NP_189089.3| TMS membrane family protein
/ tumour differentially expressed (TDE) family protein
[Arabidopsis thaliana]
Length = 409
Score = 571 bits (1472), Expect = e-161
Identities = 264/394 (67%), Positives = 319/394 (80%), Gaps = 5/394 (1%)
Query: 6 DNSSNQQGCGVLLKVSSWLSQFRNASNPKMARYVYALIFLVCNLLAWASRDELPGRSVLT 65
+N S + +K SW +QFRN NP MARYVY LIFL+ NLLAWA+RD GR L
Sbjct: 10 NNQSIRNDSYEAIKNGSWFNQFRNGCNPWMARYVYGLIFLIANLLAWAARDY--GRGALR 67
Query: 66 KLKGLKTCKDPKDCLGTNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKI 125
K+ K CK ++CLGT+GVLRVS+GCFLF+ +MF ST SK + +RD WHSGWW +K+
Sbjct: 68 KVTRFKNCKGGENCLGTDGVLRVSLGCFLFYFVMFLSTLGTSKTHSSRDRWHSGWWFVKL 127
Query: 126 VLWVAVTIFPFLLPSELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFASEKYAERC 185
++W A+TI PFLLPS +I LYGE+AHFGAGVFL IQLIS+ISFI WLN+ + S+K AERC
Sbjct: 128 IMWPALTIIPFLLPSSIIHLYGEIAHFGAGVFLLIQLISVISFIQWLNECYQSQKDAERC 187
Query: 186 QIHVMLFATASYFICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKVNA 245
+++VML +T SY +C+VGVILMYIWYAP SCLLNI FIT+TL L+Q+MTS++LHPKVNA
Sbjct: 188 RVYVMLLSTTSYTVCIVGVILMYIWYAPDSSCLLNIFFITWTLFLIQLMTSIALHPKVNA 247
Query: 246 GILSPGLMGLYVVFLCWCAIRSEPEGYECIRKSDSPNKTDWQNIISLVVGILALVIATFS 305
G L+P LMGLYVVF+CWCAIRSEP G C RK+ + N+TDW IIS VV +LA+VIATFS
Sbjct: 248 GYLTPALMGLYVVFICWCAIRSEPVGESCNRKAAASNRTDWLTIISFVVALLAMVIATFS 307
Query: 306 TGIDSKCFQYRKGDKPAE---EDDVPYGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWT 362
TGIDS+CFQ++K + E EDDVPYGYGFFHFVFATGAMYFAMLL+GWN+HH MKKWT
Sbjct: 308 TGIDSQCFQFKKDENDQEEEAEDDVPYGYGFFHFVFATGAMYFAMLLIGWNTHHPMKKWT 367
Query: 363 IDVGWTSAWVRIVNEWLAVCVYLWMLIAPIIWKN 396
IDVGWTS WVR+VNEWLAVCVY+WML+AP+I K+
Sbjct: 368 IDVGWTSTWVRVVNEWLAVCVYIWMLVAPLILKS 401
>gb|AAM51345.1| unknown protein [Arabidopsis thaliana] gi|16604677|gb|AAL24131.1|
unknown protein [Arabidopsis thaliana]
gi|18413990|ref|NP_567403.1| TMS membrane family protein
/ tumour differentially expressed (TDE) family protein
[Arabidopsis thaliana]
Length = 394
Score = 566 bits (1459), Expect = e-160
Identities = 263/382 (68%), Positives = 311/382 (80%), Gaps = 3/382 (0%)
Query: 18 LKVSSWLSQFRNASNPKMARYVYALIFLVCNLLAWASRDELPGRSVLTKLKGLKTCKDPK 77
+K SW QFRN NP MARYVY LIFL+ NLLAWA RD GR LT+++ K CK+
Sbjct: 15 IKNGSWFIQFRNGCNPWMARYVYGLIFLLANLLAWALRDY--GRGALTEMRKFKNCKEGG 72
Query: 78 DCLGTNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFL 137
DCLGT GVLRVS GCFLF+ +MF ST SK + +RD WHSGWW K+ + + +TIFPFL
Sbjct: 73 DCLGTEGVLRVSFGCFLFYFIMFLSTVGTSKTHSSRDKWHSGWWFAKLFMLLGLTIFPFL 132
Query: 138 LPSELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFASEKYAERCQIHVMLFATASY 197
LPS +I YGE+AHFGAGVFL IQLISIISFITWLN+ F ++K AERC +HVML AT +Y
Sbjct: 133 LPSSIIQFYGEIAHFGAGVFLLIQLISIISFITWLNECFQAQKDAERCHVHVMLLATTAY 192
Query: 198 FICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKVNAGILSPGLMGLYV 257
+C++GVILMYIWY P+PSCLLNI FIT+TL L+Q+MTS+SLHPK+NAG L+P LMGLYV
Sbjct: 193 TVCILGVILMYIWYVPEPSCLLNIFFITWTLFLIQLMTSISLHPKINAGFLTPALMGLYV 252
Query: 258 VFLCWCAIRSEPEGYECIRKSDSPNKTDWQNIISLVVGILALVIATFSTGIDSKCFQYRK 317
VF+CWCAIRSEP G C RK++ ++TDW IIS VV +LA+VIATFSTG+DS+CFQ+RK
Sbjct: 253 VFICWCAIRSEPVGETCNRKAEGSSRTDWLTIISFVVALLAMVIATFSTGVDSQCFQFRK 312
Query: 318 GDKPAEEDDVPYGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNE 377
D+ EED +PYGYGFFHFVFATGAMYFAMLLVGWN HHSMKKWTIDVGWTS WVRIVNE
Sbjct: 313 -DENHEEDAIPYGYGFFHFVFATGAMYFAMLLVGWNIHHSMKKWTIDVGWTSTWVRIVNE 371
Query: 378 WLAVCVYLWMLIAPIIWKNIHT 399
WLAV VY+WML+AP++ K+ T
Sbjct: 372 WLAVGVYIWMLVAPMVLKSRQT 393
>ref|NP_849373.1| TMS membrane family protein / tumour differentially expressed (TDE)
family protein [Arabidopsis thaliana]
Length = 394
Score = 563 bits (1451), Expect = e-159
Identities = 261/382 (68%), Positives = 310/382 (80%), Gaps = 3/382 (0%)
Query: 18 LKVSSWLSQFRNASNPKMARYVYALIFLVCNLLAWASRDELPGRSVLTKLKGLKTCKDPK 77
+K SW QFRN NP MARYVY LIFL+ NLLAWA RD GR LT+++ K CK+
Sbjct: 15 IKNGSWFIQFRNGCNPWMARYVYGLIFLLANLLAWALRDY--GRGALTEMRKFKNCKEGG 72
Query: 78 DCLGTNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFL 137
DCLGT GVLRVS GCFLF+ +MF ST SK + +RD WHSGWW K+ + + +TIFPFL
Sbjct: 73 DCLGTEGVLRVSFGCFLFYFIMFLSTVGTSKTHSSRDKWHSGWWFAKLFMLLGLTIFPFL 132
Query: 138 LPSELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFASEKYAERCQIHVMLFATASY 197
LPS +I YGE+AHFGAGVFL IQLISIISFITWLN+ F ++K AERC +HVML AT +Y
Sbjct: 133 LPSSIIQFYGEIAHFGAGVFLLIQLISIISFITWLNECFQAQKDAERCHVHVMLLATTAY 192
Query: 198 FICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKVNAGILSPGLMGLYV 257
+C++GVILMYIWY P+PSCLLNI FIT+TL L+Q+MTS+SLHPK+NAG L+P LMGLYV
Sbjct: 193 TVCILGVILMYIWYVPEPSCLLNIFFITWTLFLIQLMTSISLHPKINAGFLTPALMGLYV 252
Query: 258 VFLCWCAIRSEPEGYECIRKSDSPNKTDWQNIISLVVGILALVIATFSTGIDSKCFQYRK 317
VF+CWCAIR +P G C RK++ ++TDW IIS VV +LA+VIATFSTG+DS+CFQ+RK
Sbjct: 253 VFICWCAIRRQPVGETCNRKAEGSSRTDWLTIISFVVALLAMVIATFSTGVDSQCFQFRK 312
Query: 318 GDKPAEEDDVPYGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNE 377
D+ EED +PYGYGFFHFVFATGAMYFAMLLVGWN HHSMKKWTIDVGWTS WVRIVNE
Sbjct: 313 -DENHEEDAIPYGYGFFHFVFATGAMYFAMLLVGWNIHHSMKKWTIDVGWTSTWVRIVNE 371
Query: 378 WLAVCVYLWMLIAPIIWKNIHT 399
WLAV VY+WML+AP++ K+ T
Sbjct: 372 WLAVGVYIWMLVAPMVLKSRQT 393
>ref|XP_550390.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|55296297|dbj|BAD68077.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|55296119|dbj|BAD67838.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 421
Score = 474 bits (1219), Expect = e-132
Identities = 220/366 (60%), Positives = 275/366 (75%), Gaps = 4/366 (1%)
Query: 32 NPKMARYVYALIFLVCNLLAWASRDELPGRSVLTKLKGLK-TCKDPKDCLGTNGVLRVSM 90
NP MARYVYA +FL NLLAW RD G VL +L+ L+ +C+ CLG GVLRVS+
Sbjct: 47 NPMMARYVYAFVFLATNLLAWTLRDF--GHPVLAELRRLRGSCQGAGYCLGAEGVLRVSL 104
Query: 91 GCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPSELIDLYGEVA 150
GCFLFF +MF ST R K ++ R++WHS WW KIVLW+ T+ PF LPS LI LYG++A
Sbjct: 105 GCFLFFFVMFLSTVRTRKTHDRRNSWHSEWWPAKIVLWMGFTVVPFFLPSPLIQLYGKIA 164
Query: 151 HFGAGVFLFIQLISIISFITWLNDFFASEKYAERCQIHVMLFATASYFICMVGVILMYIW 210
HFGAG FL IQL+S+ FITWLND SE +RC + V + + A+Y ++GV+LMY+W
Sbjct: 165 HFGAGAFLVIQLVSVTRFITWLNDCCRSETNLKRCHMQVQVVSIAAYVGSILGVVLMYVW 224
Query: 211 YAPQPSCLLNISFITFTLVLLQIMTSVSLHPKVNAGILSPGLMGLYVVFLCWCAIRSEPE 270
YAP+PSC LNI FIT TLVL+QIMT VSL KV AG L+PGLMG+Y+VFLCW AIRSEP
Sbjct: 225 YAPRPSCKLNILFITVTLVLVQIMTGVSLSSKVKAGYLAPGLMGVYIVFLCWTAIRSEPH 284
Query: 271 GYECIRKSDSPNKTDWQNIISLVVGILALVIATFSTGIDSKCFQYRKGD-KPAEEDDVPY 329
C +K++ DW NI S V+ ++ +V ATF+TGIDSKC Q++K + + E+DD+PY
Sbjct: 285 TEICNKKAEVATSADWVNIASFVIAVIVIVTATFATGIDSKCLQFKKAESEQPEDDDIPY 344
Query: 330 GYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLI 389
G+GFFHFVFA GAMYFAML VGWN++ +M+KWTIDVGW S WVR+VNEWLA VY+WM+I
Sbjct: 345 GFGFFHFVFAMGAMYFAMLFVGWNANQTMEKWTIDVGWASTWVRVVNEWLAAIVYIWMVI 404
Query: 390 APIIWK 395
API+WK
Sbjct: 405 APIVWK 410
>ref|NP_850202.1| TMS membrane family protein / tumour differentially expressed (TDE)
family protein [Arabidopsis thaliana]
Length = 421
Score = 325 bits (834), Expect = 1e-87
Identities = 164/374 (43%), Positives = 235/374 (61%), Gaps = 11/374 (2%)
Query: 24 LSQFRNASNPKMARYVYALIFLVCNLLAWASRDELPGRSVLTKLKGLKTCKDPKDCLGTN 83
L ++ + ARY Y IFL+ NL AW RD K L D K+ + +
Sbjct: 37 LYYYQEKNKSLRARYTYGTIFLIINLCAWFIRD------YAQKALALLPFVDQKEVVVSI 90
Query: 84 GV--LRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPSE 141
+ +S+ +F+ +MF ST KL+E +++WHS W K L V V + F +P
Sbjct: 91 RLEYSALSLAMKIFYFVMFLSTWNTMKLHEAQNSWHSDNWIFKFFLLVIVMVASFFIPQL 150
Query: 142 LIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFASEKYAERCQIHVMLFATASYFICM 201
I +YGE+A GAG+FL +QL+S+I FITW N+++ + +++ ++ + Y +
Sbjct: 151 YIQIYGEIARVGAGIFLGLQLVSVIEFITWWNNYWMPQNQSKQSCSFGLVMSIVFYIGSV 210
Query: 202 VGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKV-NAGILSPGLMGLYVVFL 260
G+ +MY +Y +C LNI FI++T++LL +M +SLH KV N G+LS G+M Y+VFL
Sbjct: 211 CGIAVMYYFYGASTACGLNIFFISWTVILLIVMMVISLHSKVKNRGLLSSGIMASYIVFL 270
Query: 261 CWCAIRSEPEGYECIRKSDSPNKTDWQNIISLVVGILALVIATFSTGIDSKCFQYR-KGD 319
CW AIRSEP +C + + + TDW I+S ++ I A+V+ATFSTGIDS+ F++ + D
Sbjct: 271 CWSAIRSEPSHTKCNAHTQNSH-TDWTTILSFLIAIGAIVMATFSTGIDSESFRFEFRKD 329
Query: 320 KPAEEDDVPYGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWL 379
+ EEDD+PY YGFFH VF+ GAMYFAML + WN HS +KW+IDVGWTS WV+IVNEW
Sbjct: 330 EAKEEDDIPYSYGFFHLVFSLGAMYFAMLFISWNLSHSTEKWSIDVGWTSTWVKIVNEWF 389
Query: 380 AVCVYLWMLIAPII 393
A +YLW LIAPI+
Sbjct: 390 AAAIYLWKLIAPIV 403
>dbj|BAB01949.1| unnamed protein product [Arabidopsis thaliana]
Length = 528
Score = 298 bits (764), Expect = 2e-79
Identities = 151/284 (53%), Positives = 194/284 (68%), Gaps = 31/284 (10%)
Query: 6 DNSSNQQGCGVLLKVSSWLSQFRNASNPKMARYVYALIFLVCNLLAWASRDELPGRSVLT 65
+N S + +K SW +QFRN NP MARYVY LIFL+ NLLAWA+RD GR L
Sbjct: 10 NNQSIRNDSYEAIKNGSWFNQFRNGCNPWMARYVYGLIFLIANLLAWAARDY--GRGALR 67
Query: 66 KLKGLKTCKDPKDCLGTNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKI 125
K+ K CK ++CLGT+GVLR LF+ +MF ST SK + +RD WHSGWW +K+
Sbjct: 68 KVTRFKNCKGGENCLGTDGVLR------LFYFVMFLSTLGTSKTHSSRDRWHSGWWFVKL 121
Query: 126 VLWVAVTIFPFLLPSELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFASEKYAERC 185
++W A+TI PFLLPS +I LYGE+AHFGAGVFL IQLIS+ISFI WLN+ + S+K AERC
Sbjct: 122 IMWPALTIIPFLLPSSIIHLYGEIAHFGAGVFLLIQLISVISFIQWLNECYQSQKDAERC 181
Query: 186 QIHVMLFATASYFICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKVNA 245
+++VML +T SY +C+VGVILMYIWYAP SCLLNI FIT+TL L+Q+MTS++LHPK
Sbjct: 182 RVYVMLLSTTSYTVCIVGVILMYIWYAPDSSCLLNIFFITWTLFLIQLMTSIALHPK--- 238
Query: 246 GILSPGLMGLYVVFLCWCAIRSEPEGYECIRKSDSPNKTDWQNI 289
V LC+C + K+D+P + D +N+
Sbjct: 239 ------------VCLCFCQHQQ--------FKTDNPVRLDLKNL 262
Score = 177 bits (450), Expect = 4e-43
Identities = 93/162 (57%), Positives = 110/162 (67%), Gaps = 19/162 (11%)
Query: 242 KVNAGILSPGLMGLYVVFLCWCAIRSEPEGYECIRKSDSPNKTDWQNII-SLVVGILALV 300
+ + GI P G +VF+ GY +K + N NI+ S VV +LA+V
Sbjct: 373 ETSLGIPYPMGNGRVIVFV---------NGYLTRKKDNVSNMLILNNIVQSFVVALLAMV 423
Query: 301 IATFSTGIDSKCFQYRKGDKPAEE---DDVPYGYGFFHFVFATGAMYFAMLLVGWNSHHS 357
IATFSTGIDS+CFQ++K + EE DDVPYGYGFFHFVFATGAMYFAMLL+GWN+HH
Sbjct: 424 IATFSTGIDSQCFQFKKDENDQEEEAEDDVPYGYGFFHFVFATGAMYFAMLLIGWNTHHP 483
Query: 358 MKKWTIDVGWTSAWVRIVNEWLAVCVYL------WMLIAPII 393
MKKWTIDVGWTS WVR+VNEWLAVCVY W LI I+
Sbjct: 484 MKKWTIDVGWTSTWVRVVNEWLAVCVYKQKTNANWDLIGKIV 525
>emb|CAD41182.3| OSJNBb0002J11.6 [Oryza sativa (japonica cultivar-group)]
gi|50926300|ref|XP_473097.1| OSJNBb0002J11.6 [Oryza
sativa (japonica cultivar-group)]
Length = 477
Score = 295 bits (754), Expect = 2e-78
Identities = 156/350 (44%), Positives = 206/350 (58%), Gaps = 40/350 (11%)
Query: 36 ARYVYALIFLVCNLLAWASRDELPGRSVLTKLKGLKTC-KDPKDCLGTNGVLRVSMGCFL 94
ARY Y LIF NLLAW RD G +L L + C C + GVLR+
Sbjct: 27 ARYAYGLIFFATNLLAWFVRDY--GAKLLRGLHHVPVCGAGDSKCFQSGGVLRI------ 78
Query: 95 FFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPSELIDLYGEVAHFGA 154
FF +MF +T KL+E R++WHSG W +K +++ I PF++P+ I LYGE+A GA
Sbjct: 79 FFWVMFATTFGTRKLHEVRNSWHSGCWILKFLVYAVSIIIPFIVPNIFIQLYGEIARMGA 138
Query: 155 GVFLFIQLISIISFITWLNDFFASEKYAERCQIHVMLFATASYFICMVGVILMYIWYAPQ 214
G LF +S ISFI AS+ G+ ++Y+ Y P
Sbjct: 139 G-GLFGLFLSTISFI-------------------------ASF----AGIAVLYVLYVPN 168
Query: 215 PSCLLNISFITFTLVLLQIMTSVSLHPKVNAGILSPGLMGLYVVFLCWCAIRSEPEGYEC 274
SC NI IT+T L+ +M +VSLH KVN G+LS G+MGLY+VFLCW A+ SEP+ +C
Sbjct: 169 SSCAFNIFTITWTATLVAVMMAVSLHSKVNEGLLSSGIMGLYIVFLCWSALHSEPQTGKC 228
Query: 275 IRKSDSPNKTDWQNIISLVVGILALVIATFSTGIDSKCFQYRKGDKPAEEDDVPYGYGFF 334
+ N DW I+S ++ I A+V+ATFSTGID++ FQ+R D+ EDDVPY Y F
Sbjct: 229 HTRLIFANDGDWATIVSFIIAICAIVMATFSTGIDTRSFQFR-NDEDQLEDDVPYSYEIF 287
Query: 335 HFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVY 384
H VFA GAMYFAML + W +H +KW+IDVGW S WV+I+NEW A +Y
Sbjct: 288 HIVFAMGAMYFAMLFINWELNHPTRKWSIDVGWVSTWVKIINEWFAASIY 337
>ref|NP_187268.2| TMS membrane family protein / tumour differentially expressed (TDE)
family protein [Arabidopsis thaliana]
Length = 409
Score = 209 bits (532), Expect = 1e-52
Identities = 127/404 (31%), Positives = 200/404 (49%), Gaps = 39/404 (9%)
Query: 14 CGVLLKVSSWLSQFRNASNPKMARYVYALIFLVCNLLAWASRDELPGRSVLTKLKGLKTC 73
CG+ V+S +S+ K AR Y +F +++W R+ G +L KL + T
Sbjct: 14 CGLCSSVASGISR-------KSARIAYCGLFGASLVVSWILRET--GAPLLEKLPWINTS 64
Query: 74 KD-PKDCLGTNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVT 132
K+ VLRVS G FLFF + N+ RD+WH G W +K+++W +
Sbjct: 65 DSYTKEWYQQQAVLRVSFGNFLFFAIYALIMIGVKDQNDRRDSWHHGGWGLKMIVWFLLV 124
Query: 133 IFPFLLPSELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFASEKYAERCQIHVMLF 192
+ F +P+ ++ LYG ++ FGAG FL +Q++ ++ ND + EK ++ I +++
Sbjct: 125 VLMFFVPNVIVSLYGTLSKFGAGAFLLVQVVLLLDATHNWNDSWV-EKDEKKWYIALLVI 183
Query: 193 ATASYFICMVGVILMYIWYAPQ-PSCLLNISFITFTLVLLQIMTSVSLHPKVNAGILSPG 251
+ Y +++IW+ P C LN+ FI ++L + ++LHP VN +L
Sbjct: 184 SIVCYIATYTFSGILFIWFNPSGQDCGLNVFFIVMPMILAFVFAIIALHPAVNGSLLPAS 243
Query: 252 LMGLYVVFLCWCAIRSEPEGYECIRKSDSPNKTDWQNIISLVVGILALVIATF------- 304
++ +Y ++C+ + SEP Y C + NK+ N +L++G+L V++
Sbjct: 244 VISVYCAYVCYTGLSSEPHDYVC----NGLNKSKAVNASTLILGMLTTVLSVLYSALRAG 299
Query: 305 -STGIDSKCFQYRKGDK--------------PAEEDDVPYGYGFFHFVFATGAMYFAMLL 349
ST S R G K AE V Y Y FFH +FA +MY AMLL
Sbjct: 300 SSTTFLSPPSSPRSGVKDALLGDPEDGKKSGEAEARPVSYSYSFFHIIFALASMYAAMLL 359
Query: 350 VGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLIAPII 393
GW + S IDVGWTS WV+I W+ +Y+W LIAP+I
Sbjct: 360 SGW-TDSSESATLIDVGWTSVWVKICTGWVTAGLYIWTLIAPLI 402
>ref|XP_468965.1| putative membrane protein [Oryza sativa (japonica cultivar-group)]
gi|29244652|gb|AAO73245.1| putative membrane protein
[Oryza sativa (japonica cultivar-group)]
Length = 417
Score = 205 bits (521), Expect = 2e-51
Identities = 114/394 (28%), Positives = 195/394 (48%), Gaps = 34/394 (8%)
Query: 29 NASNPKMARYVYALIFLVCNLLAWASRDELPGRSVLTKLKGLKTCK--DPKDCLGTNGVL 86
+A + + AR Y +F +L++ R +L ++ + T P + N VL
Sbjct: 24 SAISRRSARLAYCGLFAASLILSFLMRQF--ATPLLKQIPWINTFDYTQPDEWFQMNAVL 81
Query: 87 RVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPSELIDLY 146
RVS+G FLFF + N+ RD WH G W KIV+WV + + F +P+ +I +Y
Sbjct: 82 RVSLGNFLFFAIFALMMIGVKDQNDRRDAWHHGGWIAKIVVWVVLIVLMFCVPNVVITIY 141
Query: 147 GEVAHFGAGVFLFIQLISIISFITWLNDFFASEKYAERCQIHVMLFATASYFICMVGVIL 206
++ FG+G+FL +Q++ ++ F ND + EK ++ +I +++ Y L
Sbjct: 142 EVLSKFGSGLFLLVQVVMLLDFTNNWNDSWI-EKDEQKWEIALLVVTVVCYLSTFAFSGL 200
Query: 207 MYIWYAPQP-SCLLNISFITFTLVLLQIMTSVSLHPKVNAGILSPGLMGLYVVFLCWCAI 265
++ W+ P C LN+ FIT T++L ++LHP+VN ++ ++ +Y +LC+ ++
Sbjct: 201 LFTWFNPSGHDCGLNVFFITMTIILAFAFAIIALHPQVNGSVMPASVISVYCAYLCYTSL 260
Query: 266 RSEPEGYECIRKSDSPNKTDWQNIISLVVGILALVIATFSTGI----------------- 308
SEP+ Y C + + ++ +L++G+L V++ + +
Sbjct: 261 SSEPDDYAC---NGLHRHSKQVSMSALILGMLTTVLSVVYSAVRAGSSTTFLSPPSSPRS 317
Query: 309 --------DSKCFQYRKGDKPAEEDDVPYGYGFFHFVFATGAMYFAMLLVGWNSHHSMKK 360
D + K + V Y Y FFH +FA +MY AMLL GW S S
Sbjct: 318 GIKNPLLGDDNVEAGKSNSKEIDARPVSYSYTFFHVIFALASMYSAMLLTGWTSAASDSS 377
Query: 361 WTIDVGWTSAWVRIVNEWLAVCVYLWMLIAPIIW 394
+DVGWT+ WVRI EW +Y+W L+AP+++
Sbjct: 378 ELMDVGWTTVWVRICTEWATAALYIWTLVAPLLF 411
>ref|XP_468227.1| putative tumor differentially expressed protein 1 [Oryza sativa
(japonica cultivar-group)] gi|51979391|ref|XP_507542.1|
PREDICTED OJ1249_F12.26 gene product [Oryza sativa
(japonica cultivar-group)] gi|51979389|ref|XP_507541.1|
PREDICTED OJ1249_F12.26 gene product [Oryza sativa
(japonica cultivar-group)] gi|51964476|ref|XP_507023.1|
PREDICTED OJ1249_F12.26 gene product [Oryza sativa
(japonica cultivar-group)] gi|47497137|dbj|BAD19186.1|
putative tumor differentially expressed protein 1 [Oryza
sativa (japonica cultivar-group)]
gi|47497584|dbj|BAD19654.1| putative tumor
differentially expressed protein 1 [Oryza sativa
(japonica cultivar-group)]
Length = 414
Score = 205 bits (521), Expect = 2e-51
Identities = 113/386 (29%), Positives = 194/386 (49%), Gaps = 30/386 (7%)
Query: 34 KMARYVYALIFLVCNLLAWASRD-ELPGRSVLTKLKGLKTCKDPKDCLGTNGVLRVSMGC 92
+ AR Y +F + +WA R+ P + + D ++ T+ VLRVS+G
Sbjct: 29 RSARIAYCGLFALSLFASWALREVAAPLLQSIPWINHFHKTPD-REWFETDAVLRVSLGN 87
Query: 93 FLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPSELIDLYGEVAHF 152
F+FF ++ A + RD H G W KI WV + F +P+ ++ Y ++ F
Sbjct: 88 FVFFTILAIIMAGIKDQKDPRDKIHHGGWMAKIFCWVVIVFLMFFVPNGVVSFYESISKF 147
Query: 153 GAGVFLFIQLISIISFITWLNDFFASEKYAERCQIHVMLFATASYFICMVGVILMYIWYA 212
G+G+FL +Q++ ++ F+ N+ + + K + + +++ + Y + L++ W+
Sbjct: 148 GSGLFLLVQVVLLLDFVHGWNENWVA-KDEQFWYMALLVVSVVCYIVTFSFSGLLFHWFT 206
Query: 213 PQP-SCLLNISFITFTLVLLQIMTSVSLHPKVNAGILSPGLMGLYVVFLCWCAIRSEPEG 271
P C +N+ FI FTL+L+ + V+LHPK+N +L ++ LY +LC+ + SEP
Sbjct: 207 PSGHDCGINLFFIVFTLILVFVFAIVALHPKINGSLLPASVIALYCTYLCYSGLSSEPRD 266
Query: 272 YECIRKSDSPNKTDWQNIISLVVGILALVIATFST-------------------GIDSKC 312
YEC + N + + SL +G+L +++ + G D
Sbjct: 267 YEC---NGLHNHSKAVSTGSLSLGLLTTILSVVYSAVRAGSSATVLSAPDSPRAGADKPL 323
Query: 313 FQYRKGDKPAEEDDVP----YGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWT 368
+ K D+ AE+ DVP Y Y FFH +F+ +MY AMLL GW++ +DVGW
Sbjct: 324 LPFSKADEEAEKKDVPRPVTYSYSFFHLIFSLASMYSAMLLTGWSTSVGESGKLVDVGWP 383
Query: 369 SAWVRIVNEWLAVCVYLWMLIAPIIW 394
S WVRI +W +Y+W L+AP+++
Sbjct: 384 SVWVRIATQWATAGLYIWSLVAPLLF 409
>ref|NP_173069.1| TMS membrane family protein / tumour differentially expressed (TDE)
family protein [Arabidopsis thaliana]
gi|6587821|gb|AAF18512.1| Contains similarity to
gb|AF181686 membrane protein TMS1d from Drosophila
melanogaster. ESTs gb|R64994, gb|AI994832, gb|Z47674
come from this gene. [Arabidopsis thaliana]
gi|25354033|pir||F86296 hypothetical protein T24D18.26 -
Arabidopsis thaliana
Length = 412
Score = 193 bits (491), Expect = 7e-48
Identities = 119/383 (31%), Positives = 189/383 (49%), Gaps = 26/383 (6%)
Query: 34 KMARYVYALIFLVCNLLAWASRDELPGRSVLTKLKGLKTC-KDP-KDCLGTNGVLRVSMG 91
+ AR Y +F + +++W R+ ++ KL + K P ++ T+ VLRVS+G
Sbjct: 29 RSARIAYCGLFALSLIVSWILREV--AAPLMEKLPWINHFHKTPDREWFETDAVLRVSLG 86
Query: 92 CFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPSELIDLYGEVAH 151
FLFF ++ + RD H G W +KI+ W + IF F LP+E+I Y ++
Sbjct: 87 NFLFFSILSVMMIGVKNQKDPRDGIHHGGWMMKIICWCILVIFMFFLPNEIISFYESMSK 146
Query: 152 FGAGVFLFIQLISIISFITWLNDFFASEKYAERCQIHVMLFAT-ASYFICMVGVILMYIW 210
FGAG FL +Q++ ++ F+ ND + Y E+ +L + Y V ++ W
Sbjct: 147 FGAGFFLLVQVVLLLDFVHGWNDTWVG--YDEQFWYAALLVVSLVCYLATFVFSGFLFHW 204
Query: 211 YAPQ-PSCLLNISFITFTLVLLQIMTSVSLHPKVNAGILSPGLMGLYVVFLCWCAIRSEP 269
+ P C LN FI TL+ + + V LHP V IL ++ LY ++LC+ + SEP
Sbjct: 205 FTPSGHDCGLNTFFIIMTLIFVFVFAIVVLHPTVGGSILPASVISLYCMYLCYSGLASEP 264
Query: 270 EGYECI-RKSDSPNKTDWQNIISLVVGILALVIATFSTGIDSKCF---QYRKGDKP---- 321
YEC + S + I L+ +L++V + G + + +KP
Sbjct: 265 RDYECNGLHNHSKAVSTGTMTIGLLTTVLSVVYSAVRAGSSTTLLSPPDSPRAEKPLLPI 324
Query: 322 ---AEEDD-------VPYGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAW 371
AEE + V Y Y FFH +F+ +MY AMLL GW++ +DVGW S W
Sbjct: 325 DGKAEEKEEKENKKPVSYSYAFFHIIFSLASMYSAMLLTGWSTSVGESGKLVDVGWPSVW 384
Query: 372 VRIVNEWLAVCVYLWMLIAPIIW 394
VR+V W +++W L+API++
Sbjct: 385 VRVVTSWATAGLFIWSLVAPILF 407
>gb|AAF30310.1| hypothetical protein [Arabidopsis thaliana]
Length = 315
Score = 183 bits (465), Expect = 7e-45
Identities = 99/307 (32%), Positives = 159/307 (51%), Gaps = 29/307 (9%)
Query: 110 NETRDTWHSGWWSIKIVLWVAVTIFPFLLPSELIDLYGEVAHFGAGVFLFIQLISIISFI 169
N+ RD+WH G W +K+++W + + F +P+ ++ LYG ++ FGAG FL +Q++ ++
Sbjct: 8 NDRRDSWHHGGWGLKMIVWFLLVVLMFFVPNVIVSLYGTLSKFGAGAFLLVQVVLLLDAT 67
Query: 170 TWLNDFFASEKYAERCQIHVMLFATASYFICMVGVILMYIWYAPQ-PSCLLNISFITFTL 228
ND + EK ++ I +++ + Y +++IW+ P C LN+ FI +
Sbjct: 68 HNWNDSWV-EKDEKKWYIALLVISIVCYIATYTFSGILFIWFNPSGQDCGLNVFFIVMPM 126
Query: 229 VLLQIMTSVSLHPKVNAGILSPGLMGLYVVFLCWCAIRSEPEGYECIRKSDSPNKTDWQN 288
+L + ++LHP VN +L ++ +Y ++C+ + SEP Y C + NK+ N
Sbjct: 127 ILAFVFAIIALHPAVNGSLLPASVISVYCAYVCYTGLSSEPHDYVC----NGLNKSKAVN 182
Query: 289 IISLVVGILALVIATF--------STGIDSKCFQYRKGDK--------------PAEEDD 326
+L++G+L V++ ST S R G K AE
Sbjct: 183 ASTLILGMLTTVLSVLYSALRAGSSTTFLSPPSSPRSGVKDALLGDPEDGKKSGEAEARP 242
Query: 327 VPYGYGFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLW 386
V Y Y FFH +FA +MY AMLL GW + S IDVGWTS WV+I W+ +Y+W
Sbjct: 243 VSYSYSFFHIIFALASMYAAMLLSGW-TDSSESATLIDVGWTSVWVKICTGWVTAGLYIW 301
Query: 387 MLIAPII 393
LIAP+I
Sbjct: 302 TLIAPLI 308
>ref|XP_624279.1| PREDICTED: similar to ENSANGP00000002975 [Apis mellifera]
Length = 445
Score = 149 bits (377), Expect = 1e-34
Identities = 115/423 (27%), Positives = 188/423 (44%), Gaps = 56/423 (13%)
Query: 25 SQFRNASNPKMARYVYALIFLV-----CNLLAWASRDELPGRSVLTKLKGLKTCKDPKDC 79
SQ N R +YAL+ ++ C LA +D L + K DC
Sbjct: 24 SQCPTCRNSTSTRIMYALLLMLGTIAACITLAPGLQDALKKVPFCSNSSNYVVSKFTVDC 83
Query: 80 LGTNGVLRVSMGCF---LFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPF 136
G L V CF L+F +M R + R +G+W+IK +L + I F
Sbjct: 84 ESAVGYLAVYRICFIIALYFFLMSIIMIRVRSSRDPRAPIQNGFWAIKYLLIIGGIIGAF 143
Query: 137 LLPSELIDL----YGEVAHFGAGVFLFIQLISIISFI-TWLNDFFASEKYAERCQIHVML 191
+P + +G + F +F+ IQLI I+ F +W N + + + E + L
Sbjct: 144 FIPEKSFGTTWMYFGMIGGF---LFIIIQLILIVDFAHSWANAWVGNYEETESKGWYAAL 200
Query: 192 FATA--SYFICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKV-----N 244
+Y + + G++L++I++ SC LN FI+F L+L I + +S P V
Sbjct: 201 LGATLFNYIVSITGIVLLFIYFTHANSCDLNKFFISFNLILCVIASIISTLPNVQEYNPR 260
Query: 245 AGILSPGLMGLYVVFLCWCAIRSEPEGYEC--------IRKSDSPNKT--DWQNIISLVV 294
+G+L ++ LYVV+L W I + P+ +EC + +DS N+ D ++II L++
Sbjct: 261 SGLLQSSVVSLYVVYLTWSGISNNPD-HECNPGFLEIILNDADSRNRVAFDKESIIGLII 319
Query: 295 GILALVIATFSTGIDSKCFQY------------RKGDKPA---------EEDDVPYGYGF 333
++ ++ ST S R D + EED V Y + F
Sbjct: 320 WFSCVLYSSLSTASKSSKITMSENILVKDNGAGRNADSESGNEAKVWDNEEDSVAYNWSF 379
Query: 334 FHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLIAPII 393
FH +FA +Y M L W +S T++ S WV+I++ W+ + +Y+W LIAP +
Sbjct: 380 FHLMFALATLYVMMTLTNWYQPNSNLD-TLNANTASMWVKIISSWMCLGLYIWSLIAPAV 438
Query: 394 WKN 396
N
Sbjct: 439 LPN 441
>ref|XP_640818.1| hypothetical protein DDB0204014 [Dictyostelium discoideum]
gi|60468846|gb|EAL66846.1| hypothetical protein
DDB0204014 [Dictyostelium discoideum]
Length = 417
Score = 148 bits (373), Expect = 3e-34
Identities = 106/396 (26%), Positives = 184/396 (45%), Gaps = 41/396 (10%)
Query: 37 RYVYALIFLVCNLLAWASRDELPGRSVLTKLKGLKTC-KDPKDCLGTNGVLRVSMGCFLF 95
R VY + FL+ +++A+ S L LK C K +C G V R++ G L+
Sbjct: 23 RLVYVVFFLLVSIVAYIL--SYWTFSWFNNLDVLKICSKGDNECKGALVVYRLTFGLALY 80
Query: 96 FMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPSELIDLYGEVAHFGAG 155
+++ ++R G+W +KI+L + F +P+ +Y ++ F A
Sbjct: 81 HILLGLVMINVKSAGDSRAKLQDGYWPLKILLLGVLIFVSFFIPNSFFRVYTWISIFSAA 140
Query: 156 VFLFIQLISIISFITWLNDFFASE-----KYAERCQIHVMLFATASYFICMVGVILMYIW 210
+F+FIQL+ +I LN+ + ++ + + + + S + + G +LM ++
Sbjct: 141 IFIFIQLVLLIECAYSLNESCVRKIEDEGHSGKKWYVLLCVLSFGSIALAVAGTVLMLVF 200
Query: 211 YAPQPSCLLNISFITFTLVLLQIMTSVSLHPKV-----NAGILSPGLMGLYVVFLCWCAI 265
Y + SC +N +I F L + I+ +S+ KV ++G+ G++ LY +L + AI
Sbjct: 201 YG-RGSCSINQFYIVFNLGICLIVGVLSISEKVREYRPSSGLFQSGVVMLYCTYLIYSAI 259
Query: 266 RSEPEGYECIRKSDSPNKTDWQNIISLVVGILALVIATFSTGIDSKC------FQYRKGD 319
SEP G + SP ++ II V I+++ + F + ++ + D
Sbjct: 260 NSEPPGTCSSNNTSSPKES--TIIIGAVFTIISVCYSAFRSSDSTELLGNHNHYSSIPTD 317
Query: 320 KPAEEDDV--------PYGYGFFHFVFATGAMYFAMLLVGW-----------NSHHSMKK 360
AE V Y Y FFHF FA GAMY + LL W ++ S
Sbjct: 318 PNAETTGVADDECECTAYNYSFFHFTFACGAMYLSALLTNWATMTSTDITSSSTSSSNST 377
Query: 361 WTIDVGWTSAWVRIVNEWLAVCVYLWMLIAPIIWKN 396
++D G S WV++V+ W+ V +YLW LI PI+ +N
Sbjct: 378 ISVDSGMVSVWVKVVSSWVVVLLYLWTLIGPILLRN 413
>gb|AAQ88795.1| GSVL396 [Homo sapiens]
Length = 453
Score = 141 bits (355), Expect = 4e-32
Identities = 105/425 (24%), Positives = 192/425 (44%), Gaps = 63/425 (14%)
Query: 29 NASNPKMARYVYALIFLVCNLLAWASRDELPG-RSVLTKLKGLKTCKDPKDCL------G 81
+ +N + R +YAL LV +A +PG L K+ G C++ K + G
Sbjct: 31 SGNNSTVTRLIYALFLLVGVCVACVML--IPGMEEQLNKIPGF--CENEKGVVPCNILVG 86
Query: 82 TNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPS- 140
V R+ G +F++++ + ++ R H+G+W K +A+ I F +P
Sbjct: 87 YKAVYRLCFGLAMFYLLLSLLMIKVKSSSDPRAAVHNGFWFFKFAAAIAIIIGAFFIPEG 146
Query: 141 ELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFAS--EKYAERCQIHVMLFATA-SY 197
++ V GA F+ IQL+ +I F N+ + E+ RC +L ATA +Y
Sbjct: 147 TFTTVWFYVGMAGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNSRCWYAALLSATALNY 206
Query: 198 FICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKV-----NAGILSPGL 252
+ +V ++L +++Y SC N +FI+ ++L + +S+ PK+ +G+L +
Sbjct: 207 LLSLVAIVLFFVYYTHPASCSENKAFISVNMLLCVGASVMSILPKIQESQPRSGLLQSSV 266
Query: 253 MGLYVVFLCWCAIRSEPE-----------GYECI----RKSDSPNKTDWQNIISLVVGIL 297
+ +Y ++L W A+ +EPE GY ++ S Q II L++ +L
Sbjct: 267 ITVYTMYLTWSAMTNEPETNCNPSLLSIIGYNTTSTVPKEGQSVQWWHAQGIIGLILFLL 326
Query: 298 ALVIATFSTGIDSKCFQ---------------------YRKGDK-----PAEEDDVPYGY 331
+ ++ T +S+ + GD E D V Y Y
Sbjct: 327 CVFYSSIRTSNNSQVNKLTLTSDESTLIEDGGARSDGSLEDGDDVHRAVDNERDGVTYSY 386
Query: 332 GFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLIAP 391
FFHF+ ++Y M L W+ + ++ + WT+ WV+I + W+ + +Y+W L+AP
Sbjct: 387 SFFHFMLFLASLYIMMTLTNWSRYEPSRE--MKSQWTAVWVKISSSWIGIVLYVWTLVAP 444
Query: 392 IIWKN 396
++ N
Sbjct: 445 LVLTN 449
>emb|CAD92585.1| RP3-425C14.2 [Homo sapiens] gi|57997080|emb|CAB70662.2|
hypothetical protein [Homo sapiens]
gi|21542576|gb|AAH33029.1| Tumor differentially
expressed 2 [Homo sapiens]
gi|25453298|sp|Q9NRX5|TDE2_HUMAN Tumor differentially
expressed protein 2 (Tumor differentially expressed 1
protein like) gi|24308213|ref|NP_065806.1| tumor
differentially expressed 2 [Homo sapiens]
gi|8895091|gb|AAF80758.1| Diff33 protein homolog [Homo
sapiens]
Length = 453
Score = 140 bits (352), Expect = 9e-32
Identities = 105/425 (24%), Positives = 191/425 (44%), Gaps = 63/425 (14%)
Query: 29 NASNPKMARYVYALIFLVCNLLAWASRDELPG-RSVLTKLKGLKTCKDPKDCL------G 81
+ +N + R +YAL LV +A +PG L K+ G C++ K + G
Sbjct: 31 SGNNSTVTRLIYALFLLVGVCVACVML--IPGMEEQLNKIPGF--CENEKGVVPCNILVG 86
Query: 82 TNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPS- 140
V R+ G +F++++ + ++ R H+G+W K +A+ I F +P
Sbjct: 87 YKAVYRLCFGLAMFYLLLSLLMIKVKSSSDPRAAVHNGFWFFKFAAAIAIIIGAFFIPEG 146
Query: 141 ELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFAS--EKYAERCQIHVMLFATA-SY 197
++ V GA F+ IQL+ +I F N+ + E+ RC +L ATA +Y
Sbjct: 147 TFTTVWFYVGMAGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNSRCWYAALLSATALNY 206
Query: 198 FICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKV-----NAGILSPGL 252
+ +V ++L +++Y SC N +FI+ ++L + +S+ PK+ +G+L +
Sbjct: 207 LLSLVAIVLFFVYYTHPASCSENKAFISVNMLLCVGASVMSILPKIQESQPRSGLLQSSV 266
Query: 253 MGLYVVFLCWCAIRSEPE-----------GYECI----RKSDSPNKTDWQNIISLVVGIL 297
+ +Y ++L W A+ +EPE GY ++ S Q II L++ +L
Sbjct: 267 ITVYTMYLTWSAMTNEPETNCNPSLLSIIGYNTTSTVPKEGQSVQWWHAQGIIGLILFLL 326
Query: 298 ALVIATFSTGIDSKCFQ---------------------YRKGDK-----PAEEDDVPYGY 331
+ ++ T +S+ + GD E D V Y Y
Sbjct: 327 CVFYSSIRTSNNSQVNKLTLTSDESTLIEDGGARSDGSLEDGDDVHRAVDNERDGVTYSY 386
Query: 332 GFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLIAP 391
FFHF+ ++Y M L W + ++ + WT+ WV+I + W+ + +Y+W L+AP
Sbjct: 387 SFFHFMLFLASLYIMMTLTNWYRYEPSRE--MKSQWTAVWVKISSSWIGIVLYVWTLVAP 444
Query: 392 IIWKN 396
++ N
Sbjct: 445 LVLTN 449
>gb|AAH28607.1| Tumor differentially expressed 2 [Homo sapiens]
Length = 453
Score = 140 bits (352), Expect = 9e-32
Identities = 105/425 (24%), Positives = 191/425 (44%), Gaps = 63/425 (14%)
Query: 29 NASNPKMARYVYALIFLVCNLLAWASRDELPG-RSVLTKLKGLKTCKDPKDCL------G 81
+ +N + R +YAL LV +A +PG L K+ G C++ K + G
Sbjct: 31 SGNNSTVTRLIYALFLLVGVCVACVML--IPGMEEQLNKIPGF--CENEKGVVPCNILVG 86
Query: 82 TNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPS- 140
V R+ G +F++++ + ++ R H+G+W K +A+ I F +P
Sbjct: 87 YKSVYRLCFGLAMFYLLLSLLMIKVKSSSDPRAAVHNGFWFFKFAAAIAIIIGAFFIPEG 146
Query: 141 ELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFAS--EKYAERCQIHVMLFATA-SY 197
++ V GA F+ IQL+ +I F N+ + E+ RC +L ATA +Y
Sbjct: 147 TFTTVWFYVGMAGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNSRCWYAALLSATALNY 206
Query: 198 FICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKV-----NAGILSPGL 252
+ +V ++L +++Y SC N +FI+ ++L + +S+ PK+ +G+L +
Sbjct: 207 LLSLVAIVLFFVYYTHPASCSENKAFISVNMLLCVGASVMSILPKIQESQPRSGLLQSSV 266
Query: 253 MGLYVVFLCWCAIRSEPE-----------GYECI----RKSDSPNKTDWQNIISLVVGIL 297
+ +Y ++L W A+ +EPE GY ++ S Q II L++ +L
Sbjct: 267 ITVYTMYLTWSAMTNEPETNCNPSLLSIIGYNTTSTVPKEGQSVQWWHAQGIIGLILFLL 326
Query: 298 ALVIATFSTGIDSKCFQ---------------------YRKGDK-----PAEEDDVPYGY 331
+ ++ T +S+ + GD E D V Y Y
Sbjct: 327 CVFYSSIRTSNNSQVNKLTLTSDESTLIEDGGARSDGSLEDGDDVHRAVDNERDGVTYSY 386
Query: 332 GFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLIAP 391
FFHF+ ++Y M L W + ++ + WT+ WV+I + W+ + +Y+W L+AP
Sbjct: 387 SFFHFMLFLASLYIMMTLTNWYRYEPSRE--MKSQWTAVWVKISSSWIGIVLYVWTLVAP 444
Query: 392 IIWKN 396
++ N
Sbjct: 445 LVLTN 449
>pir||T46332 hypothetical protein DKFZp434H0413.1 - human (fragment)
Length = 457
Score = 140 bits (352), Expect = 9e-32
Identities = 105/425 (24%), Positives = 191/425 (44%), Gaps = 63/425 (14%)
Query: 29 NASNPKMARYVYALIFLVCNLLAWASRDELPG-RSVLTKLKGLKTCKDPKDCL------G 81
+ +N + R +YAL LV +A +PG L K+ G C++ K + G
Sbjct: 35 SGNNSTVTRLIYALFLLVGVCVACVML--IPGMEEQLNKIPGF--CENEKGVVPCNILVG 90
Query: 82 TNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPS- 140
V R+ G +F++++ + ++ R H+G+W K +A+ I F +P
Sbjct: 91 YKAVYRLCFGLAMFYLLLSLLMIKVKSSSDPRAAVHNGFWFFKFAAAIAIIIGAFFIPEG 150
Query: 141 ELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFAS--EKYAERCQIHVMLFATA-SY 197
++ V GA F+ IQL+ +I F N+ + E+ RC +L ATA +Y
Sbjct: 151 TFTTVWFYVGMAGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNSRCWYAALLSATALNY 210
Query: 198 FICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKV-----NAGILSPGL 252
+ +V ++L +++Y SC N +FI+ ++L + +S+ PK+ +G+L +
Sbjct: 211 LLSLVAIVLFFVYYTHPASCSENKAFISVNMLLCVGASVMSILPKIQESQPRSGLLQSSV 270
Query: 253 MGLYVVFLCWCAIRSEPE-----------GYECI----RKSDSPNKTDWQNIISLVVGIL 297
+ +Y ++L W A+ +EPE GY ++ S Q II L++ +L
Sbjct: 271 ITVYTMYLTWSAMTNEPETNCNPSLLSIIGYNTTSTVPKEGQSVQWWHAQGIIGLILFLL 330
Query: 298 ALVIATFSTGIDSKCFQ---------------------YRKGDK-----PAEEDDVPYGY 331
+ ++ T +S+ + GD E D V Y Y
Sbjct: 331 CVFYSSIRTSNNSQVNKLTLTSDESTLIEDGGARSDGSLEDGDDVHRAVDNERDGVTYSY 390
Query: 332 GFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLIAP 391
FFHF+ ++Y M L W + ++ + WT+ WV+I + W+ + +Y+W L+AP
Sbjct: 391 SFFHFMLFLASLYIMMTLTNWYRYEPSRE--MKSQWTAVWVKISSSWIGIVLYVWTLVAP 448
Query: 392 IIWKN 396
++ N
Sbjct: 449 LVLTN 453
>dbj|BAA86567.1| KIAA1253 protein [Homo sapiens]
Length = 472
Score = 140 bits (352), Expect = 9e-32
Identities = 105/425 (24%), Positives = 191/425 (44%), Gaps = 63/425 (14%)
Query: 29 NASNPKMARYVYALIFLVCNLLAWASRDELPG-RSVLTKLKGLKTCKDPKDCL------G 81
+ +N + R +YAL LV +A +PG L K+ G C++ K + G
Sbjct: 50 SGNNSTVTRLIYALFLLVGVCVACVML--IPGMEEQLNKIPGF--CENEKGVVPCNILVG 105
Query: 82 TNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPS- 140
V R+ G +F++++ + ++ R H+G+W K +A+ I F +P
Sbjct: 106 YKAVYRLCFGLAMFYLLLSLLMIKVKSSSDPRAAVHNGFWFFKFAAAIAIIIGAFFIPEG 165
Query: 141 ELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFAS--EKYAERCQIHVMLFATA-SY 197
++ V GA F+ IQL+ +I F N+ + E+ RC +L ATA +Y
Sbjct: 166 TFTTVWFYVGMAGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNSRCWYAALLSATALNY 225
Query: 198 FICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKV-----NAGILSPGL 252
+ +V ++L +++Y SC N +FI+ ++L + +S+ PK+ +G+L +
Sbjct: 226 LLSLVAIVLFFVYYTHPASCSENKAFISVNMLLCVGASVMSILPKIQESQPRSGLLQSSV 285
Query: 253 MGLYVVFLCWCAIRSEPE-----------GYECI----RKSDSPNKTDWQNIISLVVGIL 297
+ +Y ++L W A+ +EPE GY ++ S Q II L++ +L
Sbjct: 286 ITVYTMYLTWSAMTNEPETNCNPSLLSIIGYNTTSTVPKEGQSVQWWHAQGIIGLILFLL 345
Query: 298 ALVIATFSTGIDSKCFQ---------------------YRKGDK-----PAEEDDVPYGY 331
+ ++ T +S+ + GD E D V Y Y
Sbjct: 346 CVFYSSIRTSNNSQVNKLTLTSDESTLIEDGGARSDGSLEDGDDVHRAVDNERDGVTYSY 405
Query: 332 GFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLIAP 391
FFHF+ ++Y M L W + ++ + WT+ WV+I + W+ + +Y+W L+AP
Sbjct: 406 SFFHFMLFLASLYIMMTLTNWYRYEPSRE--MKSQWTAVWVKISSSWIGIVLYVWTLVAP 463
Query: 392 IIWKN 396
++ N
Sbjct: 464 LVLTN 468
>emb|CAH93497.1| hypothetical protein [Pongo pygmaeus] gi|55728978|emb|CAH91227.1|
hypothetical protein [Pongo pygmaeus]
Length = 453
Score = 139 bits (351), Expect = 1e-31
Identities = 105/425 (24%), Positives = 191/425 (44%), Gaps = 63/425 (14%)
Query: 29 NASNPKMARYVYALIFLVCNLLAWASRDELPG-RSVLTKLKGLKTCKDPKDCL------G 81
+ +N + R +YAL LV +A +PG L K+ G C++ K + G
Sbjct: 31 SGNNSTVTRLIYALFLLVGVCVACVML--IPGMEEQLNKIPGF--CENEKGVVPCNILVG 86
Query: 82 TNGVLRVSMGCFLFFMMMFWSTARASKLNETRDTWHSGWWSIKIVLWVAVTIFPFLLPS- 140
V R+ G +F++++ + ++ R H+G+W K +A+ I F +P
Sbjct: 87 YKAVYRLCFGLAMFYLLLSLLMIKVKSSSDPRAAVHNGFWFFKFAAAIAIIIGAFFIPEG 146
Query: 141 ELIDLYGEVAHFGAGVFLFIQLISIISFITWLNDFFAS--EKYAERCQIHVMLFATA-SY 197
++ V GA F+ IQL+ +I F N+ + E+ RC +L ATA +Y
Sbjct: 147 TFTTVWFYVGMAGAFCFILIQLVLLIDFAHSWNESWVEKMEEGNSRCWYAALLSATALNY 206
Query: 198 FICMVGVILMYIWYAPQPSCLLNISFITFTLVLLQIMTSVSLHPKV-----NAGILSPGL 252
+ +V ++L +++Y SC N +FI+ ++L + +S+ PK+ +G+L +
Sbjct: 207 LLSLVAIVLFFVYYTHPASCSENKAFISVNMLLCIGASVMSILPKIQESQPRSGLLQSSV 266
Query: 253 MGLYVVFLCWCAIRSEPE-----------GYECI----RKSDSPNKTDWQNIISLVVGIL 297
+ +Y ++L W A+ +EPE GY ++ S Q II L++ +L
Sbjct: 267 ITVYTMYLTWSAMTNEPETNCNPSLLSIIGYNTTSTVPKEGQSVQWWHAQGIIGLILFLL 326
Query: 298 ALVIATFSTGIDSKCFQ---------------------YRKGDK-----PAEEDDVPYGY 331
+ ++ T +S+ + GD E D V Y Y
Sbjct: 327 CVFYSSIRTSNNSQVNKLTLTSDESTLIEDGGARSDGSLEDGDDVHRAVDNERDGVTYSY 386
Query: 332 GFFHFVFATGAMYFAMLLVGWNSHHSMKKWTIDVGWTSAWVRIVNEWLAVCVYLWMLIAP 391
FFHF+ ++Y M L W + ++ + WT+ WV+I + W+ + +Y+W L+AP
Sbjct: 387 SFFHFMLFLASLYIMMTLTNWYRYEPSRE--MKSQWTAVWVKISSSWIGIVLYVWTLVAP 444
Query: 392 IIWKN 396
++ N
Sbjct: 445 LVLTN 449
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.328 0.140 0.466
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 723,062,867
Number of Sequences: 2540612
Number of extensions: 29920894
Number of successful extensions: 92226
Number of sequences better than 10.0: 177
Number of HSP's better than 10.0 without gapping: 125
Number of HSP's successfully gapped in prelim test: 52
Number of HSP's that attempted gapping in prelim test: 91617
Number of HSP's gapped (non-prelim): 284
length of query: 402
length of database: 863,360,394
effective HSP length: 130
effective length of query: 272
effective length of database: 533,080,834
effective search space: 144997986848
effective search space used: 144997986848
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0153b.2


