

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0150.12
(266 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
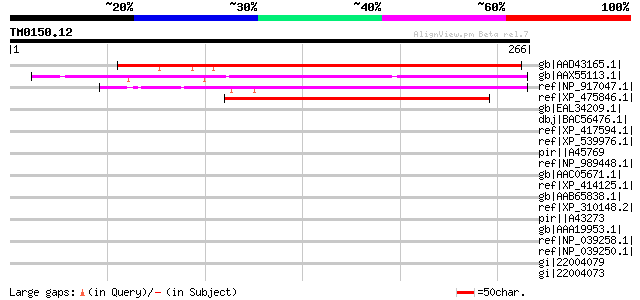
Score E
Sequences producing significant alignments: (bits) Value
gb|AAD43165.1| Hypothetical Protein [Arabidopsis thaliana] gi|15... 202 1e-50
gb|AAX55113.1| hypothetical protein At2g16190 [Arabidopsis thali... 174 3e-42
ref|NP_917047.1| P0034C09.31 [Oryza sativa (japonica cultivar-gr... 163 5e-39
ref|XP_475846.1| unknown protein [Oryza sativa (japonica cultiva... 149 8e-35
gb|EAL34209.1| GA18222-PA [Drosophila pseudoobscura] 39 0.11
dbj|BAC56476.1| similar to heregulin-beta1 [Bos taurus] 39 0.15
ref|XP_417594.1| PREDICTED: similar to atrophin-1 like protein [... 38 0.25
ref|XP_539976.1| PREDICTED: hypothetical protein XP_539976 [Cani... 38 0.33
pir||A45769 acetylcholine receptor synthesis stimulator ARIA-1 p... 37 0.43
ref|NP_989448.1| neuregulin 1 [Gallus gallus] gi|2961135|gb|AAC0... 37 0.43
gb|AAC05671.1| neuregulin beta-2a [Gallus gallus] 37 0.43
ref|XP_414125.1| PREDICTED: similar to KIAA1474 protein [Gallus ... 37 0.43
gb|AAB65838.1| Kruppel 37 0.43
ref|XP_310148.2| ENSANGP00000020042 [Anopheles gambiae str. PEST... 37 0.57
pir||A43273 heregulin precursor, splice form alpha - human 37 0.57
gb|AAA19953.1| neu differentiation factor 37 0.57
ref|NP_039258.1| neuregulin 1 isoform HRG-alpha [Homo sapiens] g... 37 0.57
ref|NP_039250.1| neuregulin 1 isoform HRG-beta1 [Homo sapiens] g... 37 0.57
gi|22004079 TPA: neuregulin 1 isoform HRG-alpha [Homo sapiens] 37 0.57
gi|22004073 TPA: neuregulin 1 isoform HRG-beta2 [Homo sapiens] 37 0.57
>gb|AAD43165.1| Hypothetical Protein [Arabidopsis thaliana]
gi|15222106|ref|NP_175358.1| hydroxyproline-rich
glycoprotein family protein [Arabidopsis thaliana]
gi|25372894|pir||E96529 hypothetical protein F13F21.24
[imported] - Arabidopsis thaliana
Length = 331
Score = 202 bits (513), Expect = 1e-50
Identities = 107/221 (48%), Positives = 142/221 (63%), Gaps = 14/221 (6%)
Query: 56 PVHTTPTYM-NPFHVSLSPPS------PPSTFEPPLAS-TNDSID---SSSSRRVRRRV- 103
P H P +M N F + +PP PPS PP ++ T + + S R R R
Sbjct: 100 PSHQIPLWMSNYFQQTPNPPQLVTHFFPPSGLAPPSSNLTPPPVKRPVTGSVRIYRSRST 159
Query: 104 --EKSEIIPPPFPWATDRRARVYTLSHLLQNQIFTITGEVQCKRCERKFEMGFDLKKNFS 161
+KS+ I PPFPWAT+RR + +L +L NQI TITGEVQC+ CE+ +++ ++L++ F+
Sbjct: 160 VSKKSDTISPPFPWATNRRGEIQSLEYLESNQITTITGEVQCRHCEKVYQVSYNLRERFA 219
Query: 162 EVSSFIAKYECTMHDRAPTIWLNPKLPTCKHCGQENSVKPVMAEKKRSVNWLFLLLGQML 221
EV F + M DRA W P+ C+ CG+E +VKPV+AE+K +NWLFLLLGQ L
Sbjct: 220 EVVKFYLTEKRKMRDRAHKDWAYPEQRRCELCGREKAVKPVIAERKSQINWLFLLLGQTL 279
Query: 222 GCCTLEQLRYFCKHTKNHRTGAKDRVLYRAYVELCKQLCPE 262
G CTLEQL+ FCKH+KNHRTGAKDRVLY Y+ LCK L P+
Sbjct: 280 GFCTLEQLKNFCKHSKNHRTGAKDRVLYLTYMGLCKMLQPK 320
>gb|AAX55113.1| hypothetical protein At2g16190 [Arabidopsis thaliana]
gi|52354251|gb|AAU44446.1| hypothetical protein
AT2G16190 [Arabidopsis thaliana]
gi|4678214|gb|AAD26960.1| hypothetical protein
[Arabidopsis thaliana] gi|15226704|ref|NP_179215.1|
hypothetical protein [Arabidopsis thaliana]
gi|25372895|pir||F84537 hypothetical protein At2g16190
[imported] - Arabidopsis thaliana
Length = 303
Score = 174 bits (440), Expect = 3e-42
Identities = 105/267 (39%), Positives = 141/267 (52%), Gaps = 18/267 (6%)
Query: 12 QLICSSISSSTKSHPFFLSQMEMKKAENVKNNINDPL-ALSLSVEPVHT---TPTYMNPF 67
QL+ S +T+ P +M N N++ P AL +V P + TP P
Sbjct: 40 QLLTSDPPQNTQPSP--PQPNDMTSFANGTNHVIVPTQALEQAVPPPNVSVRTPLPYQPS 97
Query: 68 HVSLSPPSPPSTFEPPLASTNDSIDSSSSRR---------VRRRVEKSEIIPPPFPWATD 118
L PP LA+ R V R V EI+PP +PWAT
Sbjct: 98 EEVLPPPQLNQVATVALATPRRGRPPGGQARRNSKRPVAGVERNVGDREIVPP-YPWATK 156
Query: 119 RRARVYTLSHLLQNQIFTITGEVQCKRCERKFEMGFDLKKNFSEVSSFIAKYECTMHDRA 178
+ ++ + L N I I+G+V CK C+R + ++L++ FSE+ +I + M RA
Sbjct: 157 KPGKIQSFRDLSSNNINVISGQVHCKTCDRTDTVEYNLEEKFSELYGYIKVNKEEMRHRA 216
Query: 179 PTIWLNPKLPTCKHCGQENSVKPVMAEKKRSVNWLFLLLGQMLGCCTLEQLRYFCKHTKN 238
P W PKL C+ C E +KPVM+E+K +NWLFLLLGQMLGCCTL+QLRYFC+
Sbjct: 217 PGSWSTPKLIPCRTCKSE--MKPVMSERKEEINWLFLLLGQMLGCCTLDQLRYFCQLNSK 274
Query: 239 HRTGAKDRVLYRAYVELCKQLCPEWHF 265
HRTG+KDRV+Y Y+ LCKQL PE F
Sbjct: 275 HRTGSKDRVVYITYLSLCKQLDPEGPF 301
>ref|NP_917047.1| P0034C09.31 [Oryza sativa (japonica cultivar-group)]
gi|18565431|dbj|BAB84618.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 316
Score = 163 bits (412), Expect = 5e-39
Identities = 95/255 (37%), Positives = 130/255 (50%), Gaps = 41/255 (16%)
Query: 47 PLALSLSVEPVHTTPTYMNPFHVSLSPPSPPSTFEPPLASTNDSIDSSSSRRVRRRVEKS 106
P+ L PV + M P H L P PPS+ + P +S + S+ + S R R +S
Sbjct: 65 PMLLPSPTRPVVFS---MQP-HFDLVPALPPSSPQVPQSSLS-SLSAPGSTRHYRNSPRS 119
Query: 107 EIIPPP----------------------------------FPWATDRRARVY--TLSHLL 130
++ PP FPW T V TL +L
Sbjct: 120 SLLAPPSNRRRLNNPDEGQSPRGRGEEANGDNGVLVMATSFPWVTSADLPVLHCTLESML 179
Query: 131 QNQIFTITGEVQCKRCERKFEMGFDLKKNFSEVSSFIAKYECTMHDRAPTIWLNPKLPTC 190
I ++ G+ C RC + + +DL F EV ++A M DRAP W+ P+LP C
Sbjct: 180 LKGITSVEGKATCNRCSAEVPIAYDLDAKFREVRDYVAANIHIMDDRAPEHWMCPRLPDC 239
Query: 191 KHCGQENSVKPVMAEKKRSVNWLFLLLGQMLGCCTLEQLRYFCKHTKNHRTGAKDRVLYR 250
CG++ + P + +KR +NWLFL LGQMLGCCTLE L++FCK+TKNH TGAK RVLY
Sbjct: 240 GSCGKKACMWPQIPNEKREINWLFLFLGQMLGCCTLEGLKFFCKNTKNHCTGAKTRVLYY 299
Query: 251 AYVELCKQLCPEWHF 265
AY+E+C+QL P+ F
Sbjct: 300 AYIEMCRQLDPQGPF 314
>ref|XP_475846.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|47900492|gb|AAT39249.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 315
Score = 149 bits (376), Expect = 8e-35
Identities = 64/136 (47%), Positives = 91/136 (66%)
Query: 111 PPFPWATDRRARVYTLSHLLQNQIFTITGEVQCKRCERKFEMGFDLKKNFSEVSSFIAKY 170
PP+PWAT+ A+ ++L L + I I GE +C+RC+ + + +++ F EVS + +
Sbjct: 171 PPYPWATNEVAKHHSLVELARRDIININGEARCRRCDTRKMIVYNIATKFREVSDYFRQN 230
Query: 171 ECTMHDRAPTIWLNPKLPTCKHCGQENSVKPVMAEKKRSVNWLFLLLGQMLGCCTLEQLR 230
M+DRA W+NP +P C CG E ++PV+A +K +NWLFLLLG+ LG CTL+QL+
Sbjct: 231 YQHMNDRAQARWMNPVVPNCDSCGHERCMRPVIAAEKERINWLFLLLGETLGLCTLDQLK 290
Query: 231 YFCKHTKNHRTGAKDR 246
YFC HT HRTGAKDR
Sbjct: 291 YFCAHTNRHRTGAKDR 306
>gb|EAL34209.1| GA18222-PA [Drosophila pseudoobscura]
Length = 514
Score = 39.3 bits (90), Expect = 0.11
Identities = 35/122 (28%), Positives = 51/122 (41%), Gaps = 18/122 (14%)
Query: 51 SLSVEPVHTTPTYMNPFHVSLSPP----SPPSTFEPPLASTNDSIDSSSSRRVRRRVEKS 106
+L+V V P + P V + PP PP+T PPL + S +SSSS +
Sbjct: 301 TLTVTLVPIIPQHTPPAPVVVVPPPPPIQPPATARPPLPRSCSSANSSSSLS-----NDA 355
Query: 107 EIIPPPFPWATDRRA-------RVYTLSHLLQNQIFTITGE--VQCKRCERKFEMGFDLK 157
+ PP + RR+ R + H L+ TGE C +C + F LK
Sbjct: 356 TVTAPPMRPSRSRRSYPCPLCHRPFGTRHNLKRHYMIHTGEKPFSCTKCRKPFRECSTLK 415
Query: 158 KN 159
K+
Sbjct: 416 KH 417
>dbj|BAC56476.1| similar to heregulin-beta1 [Bos taurus]
Length = 167
Score = 38.9 bits (89), Expect = 0.15
Identities = 26/73 (35%), Positives = 34/73 (45%), Gaps = 3/73 (4%)
Query: 59 TTPTYMNP--FHVSLSPPSPPSTFEPPLASTNDSIDSSS-SRRVRRRVEKSEIIPPPFPW 115
TTP M+P FH SP SPPS PP++ST S+ S + S V + PP
Sbjct: 68 TTPARMSPVDFHTPSSPKSPPSEMSPPVSSTTVSMPSMAVSPFVEEERPLLLVTPPRLRE 127
Query: 116 ATDRRARVYTLSH 128
D A+ + H
Sbjct: 128 KYDHHAQQFNSFH 140
>ref|XP_417594.1| PREDICTED: similar to atrophin-1 like protein [Gallus gallus]
Length = 1167
Score = 38.1 bits (87), Expect = 0.25
Identities = 29/99 (29%), Positives = 46/99 (46%), Gaps = 4/99 (4%)
Query: 17 SISSSTKSHPFFLSQMEMKKAENVKNNINDPLALSLS-VEPVHTTPTYMNP---FHVSLS 72
S S+ SHP + +++A+ + P +++ ++P TTP P FH SLS
Sbjct: 519 SHSALQVSHPALPAAAPLQQAQPPREQPLPPAPMAMPHIKPPPTTPIPQLPTPSFHNSLS 578
Query: 73 PPSPPSTFEPPLASTNDSIDSSSSRRVRRRVEKSEIIPP 111
PPS PPL S SS+ + +S+ +PP
Sbjct: 579 THHPPSAHSPPLQLMPQSQPLQSSQAQPPVLTQSQSLPP 617
>ref|XP_539976.1| PREDICTED: hypothetical protein XP_539976 [Canis familiaris]
Length = 1099
Score = 37.7 bits (86), Expect = 0.33
Identities = 21/56 (37%), Positives = 28/56 (49%), Gaps = 2/56 (3%)
Query: 59 TTPTYMNP--FHVSLSPPSPPSTFEPPLASTNDSIDSSSSRRVRRRVEKSEIIPPP 112
TTP M+P FH SP SPPS PP++ST S+ S + ++ PP
Sbjct: 889 TTPARMSPVDFHTPSSPKSPPSETSPPVSSTTVSMPSMAVSPFVEEERPLLLVTPP 944
>pir||A45769 acetylcholine receptor synthesis stimulator ARIA-1 precursor -
chicken gi|9297019|sp|Q05199|NRG1_CHICK Pro-neuregulin-1
precursor (Pro-NRG1) [Contains: Neuregulin-1
(Acetylcholine receptor inducing activity) (ARIA)]
gi|212604|gb|AAA49037.1| aria
Length = 602
Score = 37.4 bits (85), Expect = 0.43
Identities = 19/37 (51%), Positives = 22/37 (59%), Gaps = 2/37 (5%)
Query: 59 TTPTYMNP--FHVSLSPPSPPSTFEPPLASTNDSIDS 93
TTP M+P FH SP SPPS PP++S SI S
Sbjct: 392 TTPARMSPVDFHTPTSPKSPPSEMSPPVSSLTISIPS 428
>ref|NP_989448.1| neuregulin 1 [Gallus gallus] gi|2961135|gb|AAC05670.1| neuregulin
beta-1a [Gallus gallus]
Length = 685
Score = 37.4 bits (85), Expect = 0.43
Identities = 19/37 (51%), Positives = 22/37 (59%), Gaps = 2/37 (5%)
Query: 59 TTPTYMNP--FHVSLSPPSPPSTFEPPLASTNDSIDS 93
TTP M+P FH SP SPPS PP++S SI S
Sbjct: 475 TTPARMSPVDFHTPTSPKSPPSEMSPPVSSLTISIPS 511
>gb|AAC05671.1| neuregulin beta-2a [Gallus gallus]
Length = 677
Score = 37.4 bits (85), Expect = 0.43
Identities = 19/37 (51%), Positives = 22/37 (59%), Gaps = 2/37 (5%)
Query: 59 TTPTYMNP--FHVSLSPPSPPSTFEPPLASTNDSIDS 93
TTP M+P FH SP SPPS PP++S SI S
Sbjct: 467 TTPARMSPVDFHTPTSPKSPPSEMSPPVSSLTISIPS 503
>ref|XP_414125.1| PREDICTED: similar to KIAA1474 protein [Gallus gallus]
Length = 1248
Score = 37.4 bits (85), Expect = 0.43
Identities = 25/98 (25%), Positives = 46/98 (46%), Gaps = 4/98 (4%)
Query: 89 DSIDSSSSRRVRRRVEKSEIIPPPFPWATDRRARVYTLSHLLQNQIFTITGEVQCKRCER 148
DS S S +++ +V +S I+P P ++T ++Y S L IFT + +CK C
Sbjct: 334 DSTVSDSLEQMKAQVSQSRILPEPSLFST---VQLYRQSSKLYGSIFTGASKFRCKDCSA 390
Query: 149 KFEMGFDLKKNFSEVSSFIAKYECTMHDRAPTIWLNPK 186
++ +L + +E + T ++ P W P+
Sbjct: 391 AYDTLVELTVHMNETGHYRDDNHETDNNN-PKRWSKPR 427
>gb|AAB65838.1| Kruppel
Length = 523
Score = 37.4 bits (85), Expect = 0.43
Identities = 43/178 (24%), Positives = 72/178 (40%), Gaps = 14/178 (7%)
Query: 4 HTAAIYSSQLICSSISSSTKSHPFFLSQME-MKKAENVKNNINDPLALSLSVEPVHTTPT 62
H+ S + SS +S+ + H L E +KKA + +S+SV ++ TP
Sbjct: 131 HSPLANSGKHPLSSPNSTPQQHHLGLGLGEPVKKARKLSVKKEFQTEISMSVSDLYHTPG 190
Query: 63 YMNPFHVSLSPPSPPSTFEPPLASTNDSIDSSSSRRVRRRVEKSEIIPPPFPWATDR-RA 121
P SPPS+ P +S +++ ++ P T + +
Sbjct: 191 ---------GPISPPSSGSSPNSSAHEATSGVTAAATAAATAAPAKDPSRDKSFTCKICS 241
Query: 122 RVYTLSHLLQNQIFTITGE--VQCKRCERKFEMGFDLKKNFSEVSSFIAKYECTMHDR 177
R + H+LQN T TGE +C C ++F LK + + + Y C+ DR
Sbjct: 242 RSFGYKHVLQNHERTHTGEKPFECPECHKRFTRDHHLKTHM-RLHTGEKPYHCSHCDR 298
>ref|XP_310148.2| ENSANGP00000020042 [Anopheles gambiae str. PEST]
gi|55244043|gb|EAA05901.3| ENSANGP00000020042 [Anopheles
gambiae str. PEST]
Length = 710
Score = 37.0 bits (84), Expect = 0.57
Identities = 35/102 (34%), Positives = 47/102 (45%), Gaps = 15/102 (14%)
Query: 9 YSSQLICSSISSSTKSHPFFLSQMEMKKAENVKNNINDPLALSLSVEPVHTTPTYMNPFH 68
YS Q I S+ S + SH +EN +N+ P S SV P TP ++P
Sbjct: 86 YSQQSIAFSVISPSSSH-----------SEN-RNHPTPPEVCSGSVRPTSGTPPAISPPT 133
Query: 69 VSLSPPSPPSTFEPPLASTNDSIDSSSSRRVRRRVEKSEIIP 110
+ P PS E AST S+ +S+S R EK EI+P
Sbjct: 134 DNRVPSRAPS--ESSSASTTQSL-ASTSAASGGRSEKDEILP 172
>pir||A43273 heregulin precursor, splice form alpha - human
Length = 640
Score = 37.0 bits (84), Expect = 0.57
Identities = 20/56 (35%), Positives = 27/56 (47%), Gaps = 2/56 (3%)
Query: 59 TTPTYMNP--FHVSLSPPSPPSTFEPPLASTNDSIDSSSSRRVRRRVEKSEIIPPP 112
TTP M+P FH SP SPPS PP++S S+ S + ++ PP
Sbjct: 429 TTPARMSPVDFHTPSSPKSPPSEMSPPVSSMTVSMPSMAVSPFMEEERPLLLVTPP 484
>gb|AAA19953.1| neu differentiation factor
Length = 552
Score = 37.0 bits (84), Expect = 0.57
Identities = 20/56 (35%), Positives = 27/56 (47%), Gaps = 2/56 (3%)
Query: 59 TTPTYMNP--FHVSLSPPSPPSTFEPPLASTNDSIDSSSSRRVRRRVEKSEIIPPP 112
TTP M+P FH SP SPPS PP++S S+ S + ++ PP
Sbjct: 341 TTPARMSPVDFHTPSSPKSPPSEMSPPVSSMTVSMPSMAVSPFMEEERPLLLVTPP 396
>ref|NP_039258.1| neuregulin 1 isoform HRG-alpha [Homo sapiens]
gi|9297018|sp|Q02297|NRG1_HUMAN Pro-neuregulin-1
precursor (Pro-NRG1) [Contains: Neuregulin-1 (Neu
differentiation factor) (Heregulin) (HRG) (Breast cancer
cell differentiation factor p45) (Acetylcholine receptor
inducing activity) (ARIA) (Sensory and motor
neuron-derived factor) (Glial growth factor)]
gi|183993|gb|AAA58638.1| heregulin-alpha
Length = 640
Score = 37.0 bits (84), Expect = 0.57
Identities = 20/56 (35%), Positives = 27/56 (47%), Gaps = 2/56 (3%)
Query: 59 TTPTYMNP--FHVSLSPPSPPSTFEPPLASTNDSIDSSSSRRVRRRVEKSEIIPPP 112
TTP M+P FH SP SPPS PP++S S+ S + ++ PP
Sbjct: 429 TTPARMSPVDFHTPSSPKSPPSEMSPPVSSMTVSMPSMAVSPFMEEERPLLLVTPP 484
>ref|NP_039250.1| neuregulin 1 isoform HRG-beta1 [Homo sapiens]
gi|183995|gb|AAA58639.1| heregulin-beta1
Length = 645
Score = 37.0 bits (84), Expect = 0.57
Identities = 20/56 (35%), Positives = 27/56 (47%), Gaps = 2/56 (3%)
Query: 59 TTPTYMNP--FHVSLSPPSPPSTFEPPLASTNDSIDSSSSRRVRRRVEKSEIIPPP 112
TTP M+P FH SP SPPS PP++S S+ S + ++ PP
Sbjct: 434 TTPARMSPVDFHTPSSPKSPPSEMSPPVSSMTVSMPSMAVSPFMEEERPLLLVTPP 489
>gi|22004079 TPA: neuregulin 1 isoform HRG-alpha [Homo sapiens]
Length = 640
Score = 37.0 bits (84), Expect = 0.57
Identities = 20/56 (35%), Positives = 27/56 (47%), Gaps = 2/56 (3%)
Query: 59 TTPTYMNP--FHVSLSPPSPPSTFEPPLASTNDSIDSSSSRRVRRRVEKSEIIPPP 112
TTP M+P FH SP SPPS PP++S S+ S + ++ PP
Sbjct: 429 TTPARMSPVDFHTPSSPKSPPSEMSPPVSSMTVSMPSMAVSPFMEEERPLLLVTPP 484
>gi|22004073 TPA: neuregulin 1 isoform HRG-beta2 [Homo sapiens]
Length = 637
Score = 37.0 bits (84), Expect = 0.57
Identities = 20/56 (35%), Positives = 27/56 (47%), Gaps = 2/56 (3%)
Query: 59 TTPTYMNP--FHVSLSPPSPPSTFEPPLASTNDSIDSSSSRRVRRRVEKSEIIPPP 112
TTP M+P FH SP SPPS PP++S S+ S + ++ PP
Sbjct: 426 TTPARMSPVDFHTPSSPKSPPSEMSPPVSSMTVSMPSMAVSPFMEEERPLLLVTPP 481
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.133 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 461,208,388
Number of Sequences: 2540612
Number of extensions: 18349690
Number of successful extensions: 69537
Number of sequences better than 10.0: 93
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 89
Number of HSP's that attempted gapping in prelim test: 69475
Number of HSP's gapped (non-prelim): 134
length of query: 266
length of database: 863,360,394
effective HSP length: 126
effective length of query: 140
effective length of database: 543,243,282
effective search space: 76054059480
effective search space used: 76054059480
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0150.12


