

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0148.11
(156 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
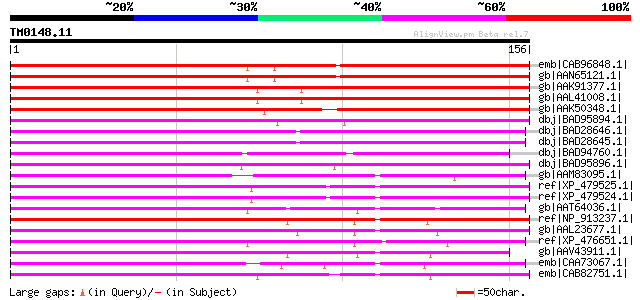
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB96848.1| serine/threonine protein kinase-like protein [Ar... 164 9e-40
gb|AAN65121.1| serine/threonine protein kinase-like protein [Ara... 164 9e-40
gb|AAK91377.1| AT5g25110/T11H3_120 [Arabidopsis thaliana] gi|250... 163 1e-39
gb|AAL41008.1| CBL-interacting protein kinase CIPK25 [Arabidopsi... 163 2e-39
gb|AAK50348.1| CBL-interacting protein kinase 16 [Arabidopsis th... 142 3e-33
dbj|BAD95894.1| Ser/Thr protein kinase [Lotus corniculatus var. ... 135 3e-31
dbj|BAD28646.1| putative CBL-interacting protein kinase [Oryza s... 134 1e-30
dbj|BAD28645.1| putative CBL-interacting protein kinase [Oryza s... 134 1e-30
dbj|BAD94760.1| CBL-interacting protein kinase 20 [Arabidopsis t... 130 1e-29
dbj|BAD95896.1| Ser/Thr protein kinase [Lotus corniculatus var. ... 130 1e-29
gb|AAM83095.1| SOS2-like protein kinase [Glycine max] 115 5e-25
ref|XP_479525.1| putative Serine/threonine Kinase [Oryza sativa ... 113 1e-24
ref|XP_479524.1| putative Serine/threonine Kinase [Oryza sativa ... 113 1e-24
gb|AAT64036.1| putative serine-threonine kinase [Gossypium hirsu... 112 2e-24
ref|NP_913237.1| unnamed protein product [Oryza sativa (japonica... 112 3e-24
gb|AAL23677.1| Serine/threonine Kinase [Persea americana] 111 7e-24
ref|XP_476651.1| putative CBL-interacting protein kinase 23 [Ory... 110 9e-24
gb|AAV43911.1| putative serine/threonine protein kinase [Oryza s... 110 9e-24
emb|CAA73067.1| serine/threonine kinase [Sorghum bicolor] gi|744... 109 2e-23
emb|CAB82751.1| serine/threonine protein kinase ATPK10 [Arabidop... 109 2e-23
>emb|CAB96848.1| serine/threonine protein kinase-like protein [Arabidopsis thaliana]
gi|19424694|gb|AAF86504.2| CBL-interacting protein
kinase 5 [Arabidopsis thaliana]
gi|17065378|gb|AAL32843.1| serine/threonine protein
kinase-like protein [Arabidopsis thaliana]
gi|30683398|ref|NP_568241.2| CBL-interacting protein
kinase 5 (CIPK5) [Arabidopsis thaliana]
gi|11346410|pir||T50802 serine/threonine protein
kinase-like protein - Arabidopsis thaliana
Length = 445
Score = 164 bits (414), Expect = 9e-40
Identities = 84/163 (51%), Positives = 111/163 (67%), Gaps = 8/163 (4%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY LLAGCLPFQ ENLM MY K+ RA+F+FPPWFSPE+++LISK+LV DP+ RI+I +IM
Sbjct: 203 LYVLLAGCLPFQDENLMNMYRKIFRADFEFPPWFSPEARRLISKLLVVDPDRRISIPAIM 262
Query: 61 RVSWFQKGFS--ASIPIPDP-----DESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSM 113
R W +K F+ + I +P ++N + + E T+ + + KF NAFEFISSM
Sbjct: 263 RTPWLRKNFTPPLAFKIDEPICSQSSKNNEEEEEDGDCENQTEPI-SPKFFNAFEFISSM 321
Query: 114 SSGFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
SSGFDLS FE KR+ SVFTS+ S +E+ KIE K + +
Sbjct: 322 SSGFDLSSLFESKRKVQSVFTSRSSATEVMEKIETVTKEMNMK 364
>gb|AAN65121.1| serine/threonine protein kinase-like protein [Arabidopsis thaliana]
Length = 435
Score = 164 bits (414), Expect = 9e-40
Identities = 84/163 (51%), Positives = 111/163 (67%), Gaps = 8/163 (4%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY LLAGCLPFQ ENLM MY K+ RA+F+FPPWFSPE+++LISK+LV DP+ RI+I +IM
Sbjct: 203 LYVLLAGCLPFQDENLMNMYRKIFRADFEFPPWFSPEARRLISKLLVVDPDRRISIPAIM 262
Query: 61 RVSWFQKGFS--ASIPIPDP-----DESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSM 113
R W +K F+ + I +P ++N + + E T+ + + KF NAFEFISSM
Sbjct: 263 RTPWLRKNFTPPLAFKIDEPICSQSSKNNEEEEEDGDCENQTEPI-SPKFFNAFEFISSM 321
Query: 114 SSGFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
SSGFDLS FE KR+ SVFTS+ S +E+ KIE K + +
Sbjct: 322 SSGFDLSSLFESKRKVQSVFTSRSSATEVMEKIETVTKEMNMK 364
>gb|AAK91377.1| AT5g25110/T11H3_120 [Arabidopsis thaliana]
gi|25090076|gb|AAN72221.1| At5g25110/T11H3_120
[Arabidopsis thaliana]
Length = 487
Score = 163 bits (413), Expect = 1e-39
Identities = 84/160 (52%), Positives = 110/160 (68%), Gaps = 4/160 (2%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY LLAG LPFQ ENLM MY K+ ++EF++PPWFSPESK+LISK+LV DPN RI+I +IM
Sbjct: 233 LYVLLAGFLPFQDENLMKMYRKIFKSEFEYPPWFSPESKRLISKLLVVDPNKRISIPAIM 292
Query: 61 RVSWFQKGFSASI--PIPDPDESNFNSD--LNSSSEQSTKVVAAAKFINAFEFISSMSSG 116
R WF+K ++ I I + + N + +++ +T + KF NAFEFISSMSSG
Sbjct: 293 RTPWFRKNINSPIEFKIDELEIQNVEDETPTTTATTATTTTPVSPKFFNAFEFISSMSSG 352
Query: 117 FDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
FDLS FE KR+ S+FTS+ S SEI K+EG K + +
Sbjct: 353 FDLSSLFESKRKLRSMFTSRWSASEIMGKLEGIGKEMNMK 392
>gb|AAL41008.1| CBL-interacting protein kinase CIPK25 [Arabidopsis thaliana]
gi|18420882|ref|NP_568466.1| CBL-interacting protein
kinase 25 (CIPK25) [Arabidopsis thaliana]
Length = 488
Score = 163 bits (412), Expect = 2e-39
Identities = 84/161 (52%), Positives = 110/161 (68%), Gaps = 5/161 (3%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY LLAG LPFQ ENLM MY K+ ++EF++PPWFSPESK+LISK+LV DPN RI+I +IM
Sbjct: 233 LYVLLAGFLPFQDENLMKMYRKIFKSEFEYPPWFSPESKRLISKLLVVDPNKRISIPAIM 292
Query: 61 RVSWFQKGFSASI--PIPDPDESNFNSD---LNSSSEQSTKVVAAAKFINAFEFISSMSS 115
R WF+K ++ I I + + N + +++ +T + KF NAFEFISSMSS
Sbjct: 293 RTPWFRKNINSPIEFKIDELEIQNVEDETPTTTATTATTTTTPVSPKFFNAFEFISSMSS 352
Query: 116 GFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
GFDLS FE KR+ S+FTS+ S SEI K+EG K + +
Sbjct: 353 GFDLSSLFESKRKLRSMFTSRWSASEIMGKLEGIGKEMNMK 393
>gb|AAK50348.1| CBL-interacting protein kinase 16 [Arabidopsis thaliana]
gi|15224679|ref|NP_180081.1| CBL-interacting protein
kinase 16 (CIPK16) [Arabidopsis thaliana]
gi|25387041|pir||B84644 probable protein kinase
[imported] - Arabidopsis thaliana
Length = 469
Score = 142 bits (358), Expect = 3e-33
Identities = 74/165 (44%), Positives = 102/165 (60%), Gaps = 13/165 (7%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LYALLAG LPF EN+MT+Y K+ +AE +FPPWFS ESK+L+S++LV DP RI++S I
Sbjct: 214 LYALLAGFLPFIDENVMTLYTKIFKAECEFPPWFSLESKELLSRLLVPDPEQRISMSEIK 273
Query: 61 RVSWFQKGFSASIPI---------PDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFIS 111
+ WF+K F+ S+ P+P DLN + A+ + NAF+FI+
Sbjct: 274 MIPWFRKNFTPSVAFSIDETIPSPPEPPTKKKKKDLNEKEDDG----ASPRSFNAFQFIT 329
Query: 112 SMSSGFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
SMSSGFDLS FE KR+ +FTSK + ++E AA+ + R
Sbjct: 330 SMSSGFDLSNLFEIKRKPKRMFTSKFPAKSVKERLETAAREMDMR 374
>dbj|BAD95894.1| Ser/Thr protein kinase [Lotus corniculatus var. japonicus]
Length = 426
Score = 135 bits (340), Expect = 3e-31
Identities = 74/165 (44%), Positives = 98/165 (58%), Gaps = 9/165 (5%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LYALL+G LPFQ EN+M +Y K +AE++FP W SP++K LIS +LVADP R +I I+
Sbjct: 188 LYALLSGYLPFQGENVMRIYRKAFKAEYEFPEWISPQAKNLISNLLVADPEKRYSIPEII 247
Query: 61 RVSWFQKGFSASIPIPDPD----ESNFNSDLNSSSEQSTKVVA-----AAKFINAFEFIS 111
WFQ GF + + E + E+ ++VA A F NAFE IS
Sbjct: 248 SDPWFQYGFMRPLAFSMKESAVGEDSTIDHFGGEEEEVPELVAAMGKPARPFYNAFEIIS 307
Query: 112 SMSSGFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
S+S GFDL FE ++R S+F SK S S + +K+EG AK L FR
Sbjct: 308 SLSHGFDLRSLFETRKRSPSMFISKYSASAVVAKLEGMAKKLNFR 352
>dbj|BAD28646.1| putative CBL-interacting protein kinase [Oryza sativa (japonica
cultivar-group)]
Length = 456
Score = 134 bits (336), Expect = 1e-30
Identities = 65/155 (41%), Positives = 93/155 (59%), Gaps = 1/155 (0%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY LL G LPFQHEN MY K+ +AE+Q PPW S ++++LI ++LV DP RI+I IM
Sbjct: 213 LYVLLCGFLPFQHENYAKMYQKIFKAEYQVPPWVSGDARRLIVRLLVVDPAKRISIPEIM 272
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 120
R WF+KGF +P + D + + + + NAF+ ISSMSSGFDLS
Sbjct: 273 RTPWFKKGFVPPVPTSPVSPKKWEED-DVLLDGGDSGAMSPRTCNAFQLISSMSSGFDLS 331
Query: 121 GFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRF 155
G FE +++ +VFTS+ + + K+E + L +
Sbjct: 332 GMFESEQKAATVFTSRAPAATVIQKLEAVGRSLGY 366
>dbj|BAD28645.1| putative CBL-interacting protein kinase [Oryza sativa (japonica
cultivar-group)]
Length = 348
Score = 134 bits (336), Expect = 1e-30
Identities = 65/155 (41%), Positives = 93/155 (59%), Gaps = 1/155 (0%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY LL G LPFQHEN MY K+ +AE+Q PPW S ++++LI ++LV DP RI+I IM
Sbjct: 150 LYVLLCGFLPFQHENYAKMYQKIFKAEYQVPPWVSGDARRLIVRLLVVDPAKRISIPEIM 209
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 120
R WF+KGF +P + D + + + + NAF+ ISSMSSGFDLS
Sbjct: 210 RTPWFKKGFVPPVPTSPVSPKKWEED-DVLLDGGDSGAMSPRTCNAFQLISSMSSGFDLS 268
Query: 121 GFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRF 155
G FE +++ +VFTS+ + + K+E + L +
Sbjct: 269 GMFESEQKAATVFTSRAPAATVIQKLEAVGRSLGY 303
>dbj|BAD94760.1| CBL-interacting protein kinase 20 [Arabidopsis thaliana]
gi|9758929|dbj|BAB09310.1| serine/threonine protein
kinase [Arabidopsis thaliana] gi|14486384|gb|AAK61493.1|
CBL-interacting protein kinase 20 [Arabidopsis thaliana]
gi|15242509|ref|NP_199394.1| CBL-interacting protein
kinase 20 (CIPK20) [Arabidopsis thaliana]
Length = 439
Score = 130 bits (327), Expect = 1e-29
Identities = 74/150 (49%), Positives = 90/150 (59%), Gaps = 3/150 (2%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY LLAG LPF +NL+ MY K+ + EF+ P WF PE KKL+S+IL +PN RI I IM
Sbjct: 202 LYVLLAGFLPFHEQNLVEMYRKITKGEFKCPNWFPPEVKKLLSRILDPNPNSRIKIEKIM 261
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 120
SWFQKGF I P ES+ L S + V + NAF+ ISS+S GFDLS
Sbjct: 262 ENSWFQKGFK-KIETPKSPESHQIDSLISDVHAAFSVKPMS--YNAFDLISSLSQGFDLS 318
Query: 121 GFFEEKRRGGSVFTSKCSVSEIASKIEGAA 150
G FE++ R S FT+K EI SK E A
Sbjct: 319 GLFEKEERSESKFTTKKDAKEIVSKFEEIA 348
>dbj|BAD95896.1| Ser/Thr protein kinase [Lotus corniculatus var. japonicus]
Length = 397
Score = 130 bits (326), Expect = 1e-29
Identities = 74/168 (44%), Positives = 94/168 (55%), Gaps = 12/168 (7%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LYALL G LPFQ EN+M +Y+K +A++ P W SPE+K LISK+LV DP R +I IM
Sbjct: 155 LYALLCGYLPFQGENVMRIYSKSFKADYAIPEWISPEAKNLISKLLVVDPEKRYSIDEIM 214
Query: 61 RVSWFQKG------FSASIPIPDPDESNFNSDLNSSSEQSTK------VVAAAKFINAFE 108
R WF+ G FS D +FN D+ + + V AA NAFE
Sbjct: 215 RDPWFEIGFMRPLAFSMKASAVDDTVIDFNDDVVEDLDPKSDVGGGEFVEAARPSYNAFE 274
Query: 109 FISSMSSGFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
ISS+S GFDL FE ++R S F SK S S + +K+E AK L R
Sbjct: 275 IISSLSHGFDLRNLFETRKRSPSTFISKLSASAVVAKLEAVAKKLNLR 322
>gb|AAM83095.1| SOS2-like protein kinase [Glycine max]
Length = 438
Score = 115 bits (287), Expect = 5e-25
Identities = 65/157 (41%), Positives = 92/157 (58%), Gaps = 9/157 (5%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
LY LLAG LPFQ +NL+ +Y K+ R +F+ PPWFS E+++LI+K+L +PN RITIS IM
Sbjct: 211 LYVLLAGFLPFQDDNLVALYKKIYRGDFKCPPWFSSEARRLITKLLDPNPNTRITISKIM 270
Query: 61 RVSWFQKGFSASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFDLS 120
SWF+K P+P +L+ + + +NAF I S+S GFDLS
Sbjct: 271 DSSWFKK------PVPKNLMGKKREELDLEEKIKQHEQEVSTTMNAFHII-SLSEGFDLS 323
Query: 121 GFFEEKRRGGSV--FTSKCSVSEIASKIEGAAKGLRF 155
FEEK+R F + S + S++E AK ++F
Sbjct: 324 PLFEEKKREEKELRFATTRPASSVISRLEDLAKAVKF 360
>ref|XP_479525.1| putative Serine/threonine Kinase [Oryza sativa (japonica
cultivar-group)] gi|33146432|dbj|BAC79540.1| putative
Serine/threonine Kinase [Oryza sativa (japonica
cultivar-group)]
Length = 356
Score = 113 bits (283), Expect = 1e-24
Identities = 66/161 (40%), Positives = 97/161 (59%), Gaps = 7/161 (4%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ LLAG LPF +NLM MY K+ +AEF+ P WF+ + ++L+ +IL +P+ RI++ IM
Sbjct: 149 LFVLLAGYLPFHDKNLMDMYKKIGKAEFKCPSWFNTDVRRLLLRILDPNPSTRISMDKIM 208
Query: 61 RVSWFQKGFSA-----SIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSS 115
WF+KG A ++ D + ++D +S + T + +NAF+ I S+S+
Sbjct: 209 ENPWFRKGLDAKLLRYNLQPKDAIPVDMSTDFDSFNSAPT-LEKKPSNLNAFDII-SLST 266
Query: 116 GFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
G DLSG FEE + S FTS + S I SKIE AKGLR +
Sbjct: 267 GLDLSGMFEESDKKESKFTSTSTASTIISKIEDIAKGLRLK 307
>ref|XP_479524.1| putative Serine/threonine Kinase [Oryza sativa (japonica
cultivar-group)] gi|33146431|dbj|BAC79539.1| putative
Serine/threonine Kinase [Oryza sativa (japonica
cultivar-group)]
Length = 443
Score = 113 bits (283), Expect = 1e-24
Identities = 66/161 (40%), Positives = 97/161 (59%), Gaps = 7/161 (4%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ LLAG LPF +NLM MY K+ +AEF+ P WF+ + ++L+ +IL +P+ RI++ IM
Sbjct: 203 LFVLLAGYLPFHDKNLMDMYKKIGKAEFKCPSWFNTDVRRLLLRILDPNPSTRISMDKIM 262
Query: 61 RVSWFQKGFSA-----SIPIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSS 115
WF+KG A ++ D + ++D +S + T + +NAF+ I S+S+
Sbjct: 263 ENPWFRKGLDAKLLRYNLQPKDAIPVDMSTDFDSFNSAPT-LEKKPSNLNAFDII-SLST 320
Query: 116 GFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
G DLSG FEE + S FTS + S I SKIE AKGLR +
Sbjct: 321 GLDLSGMFEESDKKESKFTSTSTASTIISKIEDIAKGLRLK 361
>gb|AAT64036.1| putative serine-threonine kinase [Gossypium hirsutum]
Length = 478
Score = 112 bits (281), Expect = 2e-24
Identities = 72/174 (41%), Positives = 96/174 (54%), Gaps = 22/174 (12%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ L+AG LPFQ +N+M MY K+ + EF+ P WFSPE +L++K+L +P RITI IM
Sbjct: 218 LFVLMAGYLPFQDQNIMAMYKKIYKGEFRCPRWFSPELIRLLTKLLDTNPETRITIPEIM 277
Query: 61 RVSWFQKGFS-----------ASIPIPDPDESNFNSDLNSSSEQSTKVVAAAKF------ 103
WF+KGF S+ D D + SD +S SE T++ +
Sbjct: 278 EKRWFKKGFKHIKFYIEDDKLCSVEDDDNDVGSC-SDQSSMSESETELETRKRVGTLPRP 336
Query: 104 --INAFEFISSMSSGFDLSGFFEEKRRGGSVFTSKCSVSEIASKIEGAAKGLRF 155
+NAF+ I S S GF+LSG FEE GS F S VS I SK+E AK + F
Sbjct: 337 ASLNAFDLI-SFSPGFNLSGLFEEGEE-GSRFVSGAPVSTIISKLEEIAKVVSF 388
>ref|NP_913237.1| unnamed protein product [Oryza sativa (japonica cultivar-group)]
gi|20804455|dbj|BAB92151.1| putative CBL-interacting
protein kinase 2 [Oryza sativa (japonica
cultivar-group)] gi|7340882|dbj|BAA92972.1| putative
CBL-interacting protein kinase 2 [Oryza sativa (japonica
cultivar-group)]
Length = 461
Score = 112 bits (280), Expect = 3e-24
Identities = 60/168 (35%), Positives = 103/168 (60%), Gaps = 13/168 (7%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ LLAG LPF NLM MY K+ + + +FP WF+ + ++L+S++L +PN+RIT+ ++
Sbjct: 202 LFVLLAGYLPFHDSNLMEMYRKISKGDVKFPQWFTTDVRRLLSRLLDPNPNIRITVEKLV 261
Query: 61 RVSWFQKGFSASIPIPDPDESN--------FNSDLNSSSEQSTKVVAAAK--FINAFEFI 110
WF+KG+ ++ + P+ESN F++D + ++ + ++ K +NAF+ I
Sbjct: 262 EHPWFKKGYKPAVMLSQPNESNNLKDVHTAFSADHKDNEGKAKEPASSLKPVSLNAFDII 321
Query: 111 SSMSSGFDLSGFFE--EKRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
S+S GFDLSG FE ++++ S F ++ S I SK+E A+ F+
Sbjct: 322 -SLSKGFDLSGLFENDKEQKADSRFMTQKPASAIVSKLEQIAETESFK 368
>gb|AAL23677.1| Serine/threonine Kinase [Persea americana]
Length = 453
Score = 111 bits (277), Expect = 7e-24
Identities = 69/165 (41%), Positives = 96/165 (57%), Gaps = 10/165 (6%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ LLAG LPF NLM +Y KV +AE++ P WF PE ++L+++IL +PN RI+I+ I
Sbjct: 202 LFVLLAGYLPFHDSNLMELYRKVGKAEYKCPNWFPPEVRRLLARILDPNPNTRISIAKIK 261
Query: 61 RVSWFQKGFSASIPIPDPDESNFNS-DLNSSSEQSTKVVAAAK-------FINAFEFISS 112
SWF++GF A + D +++ S VAA K +NAF+ I S
Sbjct: 262 DNSWFKRGFEAKFAKGETKSKELAPLDTDAAFASSDTGVAARKPELVKPQHLNAFDII-S 320
Query: 113 MSSGFDLSGFFEEKR-RGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
+S+GFDLSG FE+ R + F SK S I SK+E AK LR +
Sbjct: 321 LSAGFDLSGLFEDTNCRKEARFASKQPASAIISKLEDIAKRLRLK 365
>ref|XP_476651.1| putative CBL-interacting protein kinase 23 [Oryza sativa (japonica
cultivar-group)] gi|34393400|dbj|BAC82911.1| putative
CBL-interacting protein kinase 23 [Oryza sativa
(japonica cultivar-group)]
Length = 450
Score = 110 bits (276), Expect = 9e-24
Identities = 65/164 (39%), Positives = 96/164 (57%), Gaps = 10/164 (6%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ L+AG LPF+ NLM++Y K+ +A+F P WFS +KKLI KIL +P+ RITI+ ++
Sbjct: 204 LFVLMAGYLPFEDSNLMSLYKKIFKADFSCPSWFSTSAKKLIKKILDPNPSTRITIAELI 263
Query: 61 RVSWFQKGFS-ASIPIPDPDESNFNSDLNSSSEQSTKVVAAAK----FINAFEFISSMSS 115
WF+KG+ D + + NS N S +Q+ VV + +NAFE IS+ S
Sbjct: 264 NNEWFKKGYQPPRFETADVNLDDINSIFNESGDQTQLVVERREERPSVMNAFELIST-SQ 322
Query: 116 GFDLSGFFEEKRRGG----SVFTSKCSVSEIASKIEGAAKGLRF 155
G +L FE++ +G + F S+ +EI SKIE AA + F
Sbjct: 323 GLNLGTLFEKQSQGSVKRETRFASRLPANEILSKIEAAAGPMGF 366
>gb|AAV43911.1| putative serine/threonine protein kinase [Oryza sativa (japonica
cultivar-group)] gi|55167966|gb|AAV43835.1| putative
serine/threonine protein kinase [Oryza sativa (japonica
cultivar-group)]
Length = 457
Score = 110 bits (276), Expect = 9e-24
Identities = 67/164 (40%), Positives = 99/164 (59%), Gaps = 15/164 (9%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ L+AG LPFQ NLM MY K+ +AEF+ P WFS + +KL+S+IL +P R+ I+ IM
Sbjct: 205 LFVLMAGYLPFQDSNLMEMYRKIGKAEFKCPAWFSSDVRKLVSRILDPNPRSRMPITKIM 264
Query: 61 RVSWFQKGFSASIPIPDPDESN-----------FNSDLNSSSEQSTKVVAAAKF--INAF 107
WF+KG + + + + + + F+S +SSS+++ + A K +NAF
Sbjct: 265 ETYWFKKGLDSKLILKNVETNEPVTALADVNVVFSSMGSSSSKKTEEKQDAGKLTNLNAF 324
Query: 108 EFISSMSSGFDLSGFFEE-KRRGGSVFTSKCSVSEIASKIEGAA 150
+ I S+S GFDLSG FEE ++ + FTS S S I SK+E A
Sbjct: 325 DII-SLSEGFDLSGLFEETDKKKEARFTSSQSASAIISKLEDVA 367
>emb|CAA73067.1| serine/threonine kinase [Sorghum bicolor] gi|7446371|pir||T14735
probable serine/threonine kinase (EC 2.7.1.-) SNFL1 -
sorghum
Length = 440
Score = 109 bits (273), Expect = 2e-23
Identities = 70/165 (42%), Positives = 95/165 (57%), Gaps = 15/165 (9%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ LLAG LPF+ NLMT+Y K+ AEF FPPW S +K+L+++IL +P RITI I+
Sbjct: 204 LFVLLAGYLPFEDSNLMTLYKKISNAEFTFPPWTSFPAKRLLTRILDPNPMTRITIPEIL 263
Query: 61 RVSWFQKGFSASIPIPDPDE------SNFNSDLNSSSEQ--STKVVAAAKFINAFEFISS 112
WF+KG+ P+ DE + ++ N S E + K +NAFE I S
Sbjct: 264 EDEWFKKGYKR----PEFDEKYDTTLDDVDAVFNDSEEHHVTEKKEEEPVALNAFELI-S 318
Query: 113 MSSGFDLSGFF--EEKRRGGSVFTSKCSVSEIASKIEGAAKGLRF 155
MS+G +L F E++ + + FTSKC EI KIE AAK L F
Sbjct: 319 MSAGLNLGNLFDSEQEFKRETRFTSKCPPKEIVRKIEEAAKPLGF 363
>emb|CAB82751.1| serine/threonine protein kinase ATPK10 [Arabidopsis thaliana]
gi|15241066|ref|NP_195801.1| CBL-interacting protein
kinase 15 (CIPK15) [Arabidopsis thaliana]
gi|56748639|sp|P92937|CPK15_ARATH CBL-interacting
serine/threonine-protein kinase 15
(Serine/threonine-protein kinase ATPK10) (SOS2-like
protein kinase PKS3) (SOS-interacting protein 2)
(SNF1-related kinase 3.1)
Length = 421
Score = 109 bits (273), Expect = 2e-23
Identities = 62/159 (38%), Positives = 96/159 (59%), Gaps = 7/159 (4%)
Query: 1 LYALLAGCLPFQHENLMTMYNKVLRAEFQFPPWFSPESKKLISKILVADPNLRITISSIM 60
L+ LLAG LPF+ NLM +Y K+ +AE +FP W +P +K+L+ +IL +PN R++ IM
Sbjct: 202 LFVLLAGYLPFRDSNLMELYKKIGKAEVKFPNWLAPGAKRLLKRILDPNPNTRVSTEKIM 261
Query: 61 RVSWFQKGFSASI--PIPDPDESNFNSDLNSSSEQSTKVVAAAKFINAFEFISSMSSGFD 118
+ SWF+KG + + + E + ++ N+S+E+ K +NAFE I S+S+GFD
Sbjct: 262 KSSWFRKGLQEEVKESVEEETEVDAEAEGNASAEKEKK---RCINLNAFEII-SLSTGFD 317
Query: 119 LSGFFEE-KRRGGSVFTSKCSVSEIASKIEGAAKGLRFR 156
LSG FE+ + + FTS SEI K+ K L+ +
Sbjct: 318 LSGLFEKGEEKEEMRFTSNREASEITEKLVEIGKDLKMK 356
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.133 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 236,654,088
Number of Sequences: 2540612
Number of extensions: 8270250
Number of successful extensions: 29228
Number of sequences better than 10.0: 3068
Number of HSP's better than 10.0 without gapping: 1693
Number of HSP's successfully gapped in prelim test: 1376
Number of HSP's that attempted gapping in prelim test: 26651
Number of HSP's gapped (non-prelim): 3236
length of query: 156
length of database: 863,360,394
effective HSP length: 117
effective length of query: 39
effective length of database: 566,108,790
effective search space: 22078242810
effective search space used: 22078242810
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0148.11


