

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0142.11
(270 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
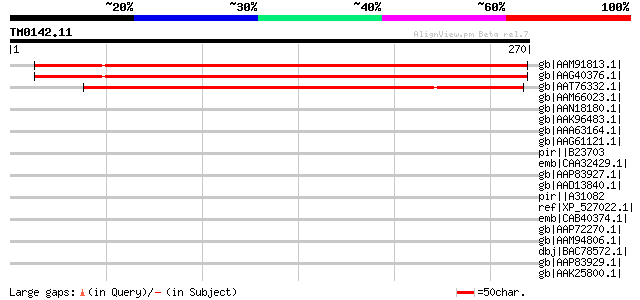
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM91813.1| unknown protein [Arabidopsis thaliana] gi|1433470... 370 e-101
gb|AAG40376.1| AT3g15840 [Arabidopsis thaliana] 370 e-101
gb|AAT76332.1| expressed protein [Oryza sativa (japonica cultiva... 318 8e-86
gb|AAM66023.1| unknown [Arabidopsis thaliana] gi|2642155|gb|AAB8... 40 0.053
gb|AAN18180.1| At2g39730/T5I7.3 [Arabidopsis thaliana] 40 0.053
gb|AAK96483.1| At2g39730/T5I7.3 [Arabidopsis thaliana] 40 0.053
gb|AAA63164.1| ribulose 1,5-bisphosphate carboxylase activase is... 40 0.069
gb|AAG61121.1| ribulose-1,5-bisphosphate carboxylase/oxygenase a... 40 0.069
pir||B23703 ribulose-bisphosphate carboxylase activase (EC 6.3.4... 40 0.069
emb|CAA32429.1| unnamed protein product [Arabidopsis thaliana] 40 0.090
gb|AAP83927.1| Rubisco activase alpha form precursor [Deschampsi... 39 0.12
gb|AAD13840.1| ribulosebisphosphate carboxylase/oxygenase activa... 38 0.26
pir||A31082 ribulose-bisphosphate carboxylase activase (EC 6.3.4... 38 0.26
ref|XP_527022.1| PREDICTED: similar to KIAA1129 protein [Pan tro... 37 0.45
emb|CAB40374.1| Starch synthase isoform SS III [Vigna unguiculata] 36 1.3
gb|AAP72270.1| ribulose-1,5-bisphosphate carboxylase activase [T... 36 1.3
gb|AAM94806.1| rubisco activase alpha [Gossypium hirsutum] 35 1.7
dbj|BAC78572.1| ribulose-bisphosphate carboxylase activase large... 35 2.2
gb|AAP83929.1| Rubisco activase alpha form precursor [Larrea tri... 35 2.9
gb|AAK25800.1| rubisco activase [Zantedeschia aethiopica] 34 3.8
>gb|AAM91813.1| unknown protein [Arabidopsis thaliana] gi|14334704|gb|AAK59530.1|
unknown protein [Arabidopsis thaliana]
gi|11994356|dbj|BAB02315.1| unnamed protein product
[Arabidopsis thaliana] gi|18400924|ref|NP_566528.1|
expressed protein [Arabidopsis thaliana]
Length = 268
Score = 370 bits (951), Expect = e-101
Identities = 174/256 (67%), Positives = 208/256 (80%), Gaps = 1/256 (0%)
Query: 14 PSNPITSRHTFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQEIEEYKIPSWANFEL 73
PS P++S+ +FL S L + K+ + A VA P +EI+EYK+PSWA FE+
Sbjct: 14 PSQPLSSKRSFLYSSRIGPILRRFPRKKLDLQVKA-VATTLAPLEEIKEYKLPSWAMFEM 72
Query: 74 GKASVYWKTMNGLPPTSGEKLKLFYNPTSTQLAPNEEFGIAFNGGFNQPIMCGGEPRAML 133
G A VYWKTMNGLPPTSGEKLKLFYNP +++L NE++G+AFNGGFNQPIMCGGEPRAML
Sbjct: 73 GTAPVYWKTMNGLPPTSGEKLKLFYNPAASKLTLNEDYGVAFNGGFNQPIMCGGEPRAML 132
Query: 134 RKDRGKADAPIYSIQICVPKHALNLIFSFTNGVDWDGPYRLQFQVPKVLQNKPIEFFNEG 193
+KDRGKAD+PIY++QIC+PKHA+NLIFSFTNGVDWDGPYRLQFQVPK QNKPIEFFNEG
Sbjct: 133 KKDRGKADSPIYTMQICIPKHAVNLIFSFTNGVDWDGPYRLQFQVPKRWQNKPIEFFNEG 192
Query: 194 LAEELGKEGACEQAIFPDSNKVITKCAMLGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLA 253
LA EL ++GACE+AIFPDSN V T+C M+ NLTVEGGDRC+L+LV GC D +S +NP A
Sbjct: 193 LANELSQDGACERAIFPDSNVVPTRCTMIANLTVEGGDRCNLDLVPGCMDTNSEHFNPYA 252
Query: 254 NVDDGTCPLDLDSDSE 269
NVDDG+CPL+L E
Sbjct: 253 NVDDGSCPLELSDSDE 268
>gb|AAG40376.1| AT3g15840 [Arabidopsis thaliana]
Length = 268
Score = 370 bits (950), Expect = e-101
Identities = 174/256 (67%), Positives = 208/256 (80%), Gaps = 1/256 (0%)
Query: 14 PSNPITSRHTFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQEIEEYKIPSWANFEL 73
PS P++S+ +FL S L + K+ + A VA P +EI+EYK+PSWA FE+
Sbjct: 14 PSQPLSSKRSFLYSSRIGPILRRFPRKKLDLQVKA-VATTLAPLEEIKEYKLPSWAMFEM 72
Query: 74 GKASVYWKTMNGLPPTSGEKLKLFYNPTSTQLAPNEEFGIAFNGGFNQPIMCGGEPRAML 133
G A VYWKTMNGLPPTSGEKLKLFYNP +++L NE++G+AFNGGFNQPIMCGGEPRAML
Sbjct: 73 GTAPVYWKTMNGLPPTSGEKLKLFYNPAASKLTLNEDYGVAFNGGFNQPIMCGGEPRAML 132
Query: 134 RKDRGKADAPIYSIQICVPKHALNLIFSFTNGVDWDGPYRLQFQVPKVLQNKPIEFFNEG 193
+KDRGKAD+PIY++QIC+PKHA+NLIFSFTNGVDWDGPYRLQFQVPK QNKPIEFFNEG
Sbjct: 133 KKDRGKADSPIYTMQICIPKHAVNLIFSFTNGVDWDGPYRLQFQVPKRRQNKPIEFFNEG 192
Query: 194 LAEELGKEGACEQAIFPDSNKVITKCAMLGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLA 253
LA EL ++GACE+AIFPDSN V T+C M+ NLTVEGGDRC+L+LV GC D +S +NP A
Sbjct: 193 LANELSQDGACERAIFPDSNVVPTRCKMIANLTVEGGDRCNLDLVPGCMDTNSEHFNPYA 252
Query: 254 NVDDGTCPLDLDSDSE 269
NVDDG+CPL+L E
Sbjct: 253 NVDDGSCPLELSDSDE 268
>gb|AAT76332.1| expressed protein [Oryza sativa (japonica cultivar-group)]
Length = 271
Score = 318 bits (816), Expect = 8e-86
Identities = 149/231 (64%), Positives = 184/231 (79%), Gaps = 3/231 (1%)
Query: 39 NVKVTRT-INAAVAVATTPA-QEIEEYKIPSWANFELGKASVYWKTMNGLPPTSGEKLKL 96
+ +V RT AA A AT PA + E +P+WA FELGKA VYWKTMNGLPP++GE L L
Sbjct: 41 SARVVRTGAAAAPAAATAPAVPQTNECSLPTWAEFELGKAPVYWKTMNGLPPSAGEGLIL 100
Query: 97 FYNPTSTQLAPNEEFGIAFNGGFNQPIMCGGEPRAMLRKDRGKADAPIYSIQICVPKHAL 156
FYNP +T++ PN +FGIAFNGGFNQPIMCGGEPR M ++RG AD PIY+I+I VP+HA+
Sbjct: 101 FYNPAATKMTPNAQFGIAFNGGFNQPIMCGGEPRQMTLQERGSADPPIYTIRIRVPQHAM 160
Query: 157 NLIFSFTNGVDWDGPYRLQFQVPKVLQNKPIEFFNEGLAEELGKEGACEQAIFPDSNKVI 216
L+FSFTNGVDWDGPY L+F+VPK NKP+ FFNEGLA+EL +EGAC++AIFPD N VI
Sbjct: 161 TLVFSFTNGVDWDGPYTLKFRVPKPWLNKPLSFFNEGLADELNREGACDRAIFPDENVVI 220
Query: 217 TKCAMLGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTCPLDLDSD 267
T C M G+ EGGDRC L++V GC DP+SH+++PLA VDDG+CP+D DS+
Sbjct: 221 TSCEM-GSYYEEGGDRCKLDIVSGCMDPNSHMFDPLATVDDGSCPMDSDSE 270
>gb|AAM66023.1| unknown [Arabidopsis thaliana] gi|2642155|gb|AAB87122.1| expressed
protein [Arabidopsis thaliana]
gi|15810188|gb|AAL06995.1| At2g39730/T5I7.3_
[Arabidopsis thaliana] gi|11762250|gb|AAG40401.1|
At2g39730 [Arabidopsis thaliana]
gi|12643259|sp|P10896|RCA_ARATH Ribulose bisphosphate
carboxylase/oxygenase activase, chloroplast precursor
(RuBisCO activase) (RA) gi|18405145|ref|NP_565913.1|
ribulose bisphosphate carboxylase/oxygenase activase /
RuBisCO activase [Arabidopsis thaliana]
gi|166834|gb|AAA20202.1| ribulose bisphosphate
carboxylase/oxygenase activase
Length = 474
Score = 40.4 bits (93), Expect = 0.053
Identities = 27/91 (29%), Positives = 42/91 (45%), Gaps = 9/91 (9%)
Query: 176 FQVPKVLQNKPIEFFNEGLAEELG------KEGACEQAIFPDSNKVITKCAMLGNLTVEG 229
F+ P++ K +E+ N + E+ E QA D+N G +G
Sbjct: 383 FEQPEMTYEKLMEYGNMLVMEQENVKRVQLAETYLSQAALGDAN---ADAIGRGTFYGKG 439
Query: 230 GDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
+ +L + EGCTDP + ++P A DDGTC
Sbjct: 440 AQQVNLPVPEGCTDPVAENFDPTARSDDGTC 470
>gb|AAN18180.1| At2g39730/T5I7.3 [Arabidopsis thaliana]
Length = 474
Score = 40.4 bits (93), Expect = 0.053
Identities = 27/91 (29%), Positives = 42/91 (45%), Gaps = 9/91 (9%)
Query: 176 FQVPKVLQNKPIEFFNEGLAEELG------KEGACEQAIFPDSNKVITKCAMLGNLTVEG 229
F+ P++ K +E+ N + E+ E QA D+N G +G
Sbjct: 383 FEQPEMTYEKLMEYGNMLVMEQENVKRVQLAETYLSQAALGDAN---ADAIGRGTFYGKG 439
Query: 230 GDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
+ +L + EGCTDP + ++P A DDGTC
Sbjct: 440 AQQVNLPVPEGCTDPVAENFDPTARSDDGTC 470
>gb|AAK96483.1| At2g39730/T5I7.3 [Arabidopsis thaliana]
Length = 474
Score = 40.4 bits (93), Expect = 0.053
Identities = 27/91 (29%), Positives = 42/91 (45%), Gaps = 9/91 (9%)
Query: 176 FQVPKVLQNKPIEFFNEGLAEELG------KEGACEQAIFPDSNKVITKCAMLGNLTVEG 229
F+ P++ K +E+ N + E+ E QA D+N G +G
Sbjct: 383 FEQPEMTYEKLMEYGNMLVMEQENVKRVQLAETYLSQAALGDAN---ADAIGRGTFYGKG 439
Query: 230 GDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
+ +L + EGCTDP + ++P A DDGTC
Sbjct: 440 AQQVNLPVPEGCTDPVAENFDPTARSDDGTC 470
>gb|AAA63164.1| ribulose 1,5-bisphosphate carboxylase activase isoform 2 [Hordeum
vulgare subsp. vulgare] gi|12643756|sp|Q40073|RCAA_HORVU
Ribulose bisphosphate carboxylase/oxygenase activase A,
chloroplast precursor (RuBisCO activase A) (RA A)
gi|7438147|pir||T06176 ribulose-bisphosphate carboxylase
activase (EC 6.3.4.-) A2 - barley
Length = 464
Score = 40.0 bits (92), Expect = 0.069
Identities = 29/98 (29%), Positives = 48/98 (48%), Gaps = 11/98 (11%)
Query: 169 DGPYRLQFQVPKVLQNKPIEFFNEGLAEELGKEGA------CEQAIFPDSNKVITKCAML 222
DGP + F+ PK+ K +E+ + + E+ + QA D+N+ K
Sbjct: 368 DGP--VTFEQPKMTVEKLLEYGHMLVQEQDNVKRVQLADTYMSQAALGDANQDAMKT--- 422
Query: 223 GNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
G+ +G + L + EGCTD ++ Y+P A DDG+C
Sbjct: 423 GSFYGKGAQQGTLPVPEGCTDQNAKNYDPTARSDDGSC 460
>gb|AAG61121.1| ribulose-1,5-bisphosphate carboxylase/oxygenase activase 2
[Gossypium hirsutum]
Length = 435
Score = 40.0 bits (92), Expect = 0.069
Identities = 27/92 (29%), Positives = 44/92 (47%), Gaps = 11/92 (11%)
Query: 176 FQVPKVLQNKPIEFFNEGLAEE-------LGKEGACEQAIFPDSNKVITKCAMLGNLTVE 228
F+ PK+ K +E+ N +AE+ L + E A+ + I + G +
Sbjct: 344 FEQPKMTIEKLLEYGNMLVAEQENVKRVQLADKYLSEAALGEANEDSINRGTFYGKAAQQ 403
Query: 229 GGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
G + + EGCTDP++ ++P A DDGTC
Sbjct: 404 VG----VPVPEGCTDPNADNFDPTARSDDGTC 431
>pir||B23703 ribulose-bisphosphate carboxylase activase (EC 6.3.4.-) A long form
precursor - barley (fragment) gi|167089|gb|AAA62701.1|
ribulose 1,5-bisphosphate carboxylase activase
Length = 426
Score = 40.0 bits (92), Expect = 0.069
Identities = 29/98 (29%), Positives = 48/98 (48%), Gaps = 11/98 (11%)
Query: 169 DGPYRLQFQVPKVLQNKPIEFFNEGLAEELGKEGA------CEQAIFPDSNKVITKCAML 222
DGP + F+ PK+ K +E+ + + E+ + QA D+N+ K
Sbjct: 330 DGP--VTFEQPKMTVEKLLEYGHMLVQEQDNVKRVQLADTYMSQAALGDANQDAMKT--- 384
Query: 223 GNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
G+ +G + L + EGCTD ++ Y+P A DDG+C
Sbjct: 385 GSFYGKGAQQGTLPVPEGCTDQNAKNYDPTARSDDGSC 422
>emb|CAA32429.1| unnamed protein product [Arabidopsis thaliana]
Length = 473
Score = 39.7 bits (91), Expect = 0.090
Identities = 27/91 (29%), Positives = 41/91 (44%), Gaps = 9/91 (9%)
Query: 176 FQVPKVLQNKPIEFFNEGLAEELG------KEGACEQAIFPDSNKVITKCAMLGNLTVEG 229
F+ P++ K +E+ N + E+ E QA D+N G +G
Sbjct: 382 FEQPEMTYEKLMEYGNMLVMEQENVKRVQLAETYLSQAALGDAN---ADAIGRGTFYGKG 438
Query: 230 GDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
+L + EGCTDP + ++P A DDGTC
Sbjct: 439 AHEVNLPVPEGCTDPVAENFDPTARSDDGTC 469
>gb|AAP83927.1| Rubisco activase alpha form precursor [Deschampsia antarctica]
Length = 465
Score = 39.3 bits (90), Expect = 0.12
Identities = 29/98 (29%), Positives = 47/98 (47%), Gaps = 11/98 (11%)
Query: 169 DGPYRLQFQVPKVLQNKPIEFFNEGLAEELGKEGA------CEQAIFPDSNKVITKCAML 222
DGP + F+ PK+ K +E+ + + E+ + QA D+NK K
Sbjct: 369 DGP--VTFEQPKMTVEKLLEYGHMLVQEQDNVKRVQLADTYMSQAALGDANKDAMKT--- 423
Query: 223 GNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
G+ +G + L + EGCTD + ++P A DDG+C
Sbjct: 424 GSFYGKGAQQGTLPVPEGCTDRDAKNFDPTARSDDGSC 461
>gb|AAD13840.1| ribulosebisphosphate carboxylase/oxygenase activase [Spinacia
oleracea] gi|12643998|sp|P10871|RCA_SPIOL Ribulose
bisphosphate carboxylase/oxygenase activase, chloroplast
precursor (RuBisCO activase) (RA)
Length = 472
Score = 38.1 bits (87), Expect = 0.26
Identities = 29/103 (28%), Positives = 47/103 (45%), Gaps = 13/103 (12%)
Query: 169 DGPYRLQFQVPKVLQNKPIEFFNEGLAEELGKEGA------CEQAIFPDSNK-VITKCAM 221
DGP F+ P++ K +E+ N + E+ + A D+NK I +
Sbjct: 376 DGPP--VFEQPEMTLQKLMEYGNMLVQEQENVKRVQLADQYMSSAALGDANKDAIDRGTF 433
Query: 222 LGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTCPLDL 264
G + + L + +GCTDP + Y+P A DDG+C +L
Sbjct: 434 FG----KAAQQVSLPVAQGCTDPEAKNYDPTARSDDGSCTYNL 472
>pir||A31082 ribulose-bisphosphate carboxylase activase (EC 6.3.4.-) precursor -
spinach gi|170129|gb|AAA34038.1| rubisco activase
precursor
Length = 472
Score = 38.1 bits (87), Expect = 0.26
Identities = 29/103 (28%), Positives = 47/103 (45%), Gaps = 13/103 (12%)
Query: 169 DGPYRLQFQVPKVLQNKPIEFFNEGLAEELGKEGA------CEQAIFPDSNK-VITKCAM 221
DGP F+ P++ K +E+ N + E+ + A D+NK I +
Sbjct: 376 DGPP--VFEQPEMTLQKLMEYGNMLVQEQENVKRVQLADQYMSSAALGDANKDAIDRGTF 433
Query: 222 LGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTCPLDL 264
G + + L + +GCTDP + Y+P A DDG+C +L
Sbjct: 434 FG----KAAQQVSLPVAQGCTDPEAKNYDPTARSDDGSCTYNL 472
>ref|XP_527022.1| PREDICTED: similar to KIAA1129 protein [Pan troglodytes]
Length = 1066
Score = 37.4 bits (85), Expect = 0.45
Identities = 23/67 (34%), Positives = 33/67 (48%), Gaps = 9/67 (13%)
Query: 188 EFFNEGLAEELGKEGACEQAIFPDSNKVITKCAMLGNLTVEGGDRCDLNLVEGCTDPSSH 247
+F+ ++ +LGK C Q P+S +V CA+ G L V GGD NL S
Sbjct: 741 DFYRTQMSTDLGKSSGCHQCPLPNSFRV---CAVNGLLYVVGGDDGSCNLA------SVE 791
Query: 248 LYNPLAN 254
YNP+ +
Sbjct: 792 YYNPVTD 798
>emb|CAB40374.1| Starch synthase isoform SS III [Vigna unguiculata]
Length = 1147
Score = 35.8 bits (81), Expect = 1.3
Identities = 48/195 (24%), Positives = 84/195 (42%), Gaps = 30/195 (15%)
Query: 6 DLLFFSVPPSNPIT-------SRHTFLNVSLPRC-YLVKE-----RNVKVTRTINAAV-- 50
D +F PP N + H + ++ P Y V+E R ++ R + V
Sbjct: 501 DWVFADGPPQNAVVYDNNRMQDFHAIVPMATPDAQYWVEEEQLIYRKLQEERKLKEEVIR 560
Query: 51 AVATTPAQEIEEYKIPSWANFELGKASVYWKTMNGLPPTSGEKLKLFYNPTSTQLAPNEE 110
A A AQ E K + F L + + + L +G + +FYNP++T L E
Sbjct: 561 AKAEKTAQMKAETKEKTLKKFLLSQKHIVYT--EPLDIQAGSTVTVFYNPSNTNLNGRPE 618
Query: 111 FGIAFNGGFNQPIMCGG--EPRAMLRKDRG---KADAPIYSIQICVPKHALNLIFSFT-N 164
+ F G FN+ G P+ ML + G KA S+++ + + ++ +FS + N
Sbjct: 619 --VWFRGSFNRWSHRNGPLPPQRMLPAESGTHVKA-----SVKVPLDAYMMDFVFSESEN 671
Query: 165 GVDWDGPYRLQFQVP 179
G +D + + + +P
Sbjct: 672 GGVFDNKFGMDYHIP 686
>gb|AAP72270.1| ribulose-1,5-bisphosphate carboxylase activase [Triticum aestivum]
Length = 201
Score = 35.8 bits (81), Expect = 1.3
Identities = 27/98 (27%), Positives = 46/98 (46%), Gaps = 11/98 (11%)
Query: 169 DGPYRLQFQVPKVLQNKPIEFFNEGLAEELGKEGA------CEQAIFPDSNKVITKCAML 222
DGP + F+ PK+ K +E+ + + E+ + QA D+N+ K
Sbjct: 105 DGP--VTFEQPKMTVEKLLEYGHMLVQEQDNVKRVQLADTYMSQAALGDANQDAMKT--- 159
Query: 223 GNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
G +G + L + GCTD ++ ++P A DDG+C
Sbjct: 160 GTFYGKGAQQGTLPVPAGCTDQTAKNFDPTARSDDGSC 197
>gb|AAM94806.1| rubisco activase alpha [Gossypium hirsutum]
Length = 76
Score = 35.4 bits (80), Expect = 1.7
Identities = 13/22 (59%), Positives = 17/22 (77%)
Query: 239 EGCTDPSSHLYNPLANVDDGTC 260
EGCTDP++ ++P A DDGTC
Sbjct: 51 EGCTDPNADNFDPTARSDDGTC 72
>dbj|BAC78572.1| ribulose-bisphosphate carboxylase activase large isoform precursor
protein [Oryza sativa (japonica cultivar-group)]
gi|8918359|dbj|BAA97583.1| RuBisCO activase large
isoform precursor [Oryza sativa (japonica
cultivar-group)]
Length = 466
Score = 35.0 bits (79), Expect = 2.2
Identities = 26/93 (27%), Positives = 43/93 (45%), Gaps = 10/93 (10%)
Query: 175 QFQVPKVLQNKPIEFF------NEGLAEELGKEGACEQAIFPDSNKVITKCAMLGNLTVE 228
+F+ PK+ K +E+ E + E +A D+N K G+ +
Sbjct: 373 EFEQPKMTIEKLMEYGYMLVKEQENVKRVQLAEQYLSEAALGDANSDAMKT---GSFYGQ 429
Query: 229 GGDRC-DLNLVEGCTDPSSHLYNPLANVDDGTC 260
G + +L + EGCTDP + ++P A DDG+C
Sbjct: 430 GAQQAGNLPVPEGCTDPVAKNFDPTARSDDGSC 462
>gb|AAP83929.1| Rubisco activase alpha form precursor [Larrea tridentata]
Length = 476
Score = 34.7 bits (78), Expect = 2.9
Identities = 13/22 (59%), Positives = 16/22 (72%)
Query: 239 EGCTDPSSHLYNPLANVDDGTC 260
EGCTDP + Y+P A DDG+C
Sbjct: 451 EGCTDPQATNYDPTARSDDGSC 472
>gb|AAK25800.1| rubisco activase [Zantedeschia aethiopica]
Length = 426
Score = 34.3 bits (77), Expect = 3.8
Identities = 23/91 (25%), Positives = 44/91 (48%), Gaps = 9/91 (9%)
Query: 176 FQVPKVLQNKPIEFFNEGLAEELGKEGA------CEQAIFPDSNKVITKCAMLGNLTVEG 229
F PK+ +K +E+ N + E+ + +A D+N+ K G+ +
Sbjct: 335 FAQPKMTLDKLLEYGNMLVQEQENVKRVQLADKYLSEAALGDANQDAIKT---GSFYGKA 391
Query: 230 GDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
+ ++ + EGCTD ++ ++P A DDG+C
Sbjct: 392 AQQANVPVPEGCTDRNATNFDPTARSDDGSC 422
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.317 0.137 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 505,067,906
Number of Sequences: 2540612
Number of extensions: 22026286
Number of successful extensions: 35877
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 18
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 35858
Number of HSP's gapped (non-prelim): 29
length of query: 270
length of database: 863,360,394
effective HSP length: 126
effective length of query: 144
effective length of database: 543,243,282
effective search space: 78227032608
effective search space used: 78227032608
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0142.11


