

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0140.7
(571 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
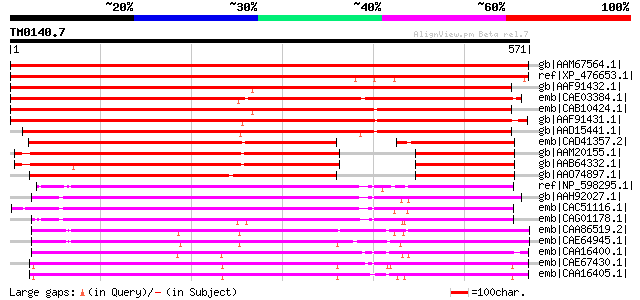
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM67564.1| unknown protein [Arabidopsis thaliana] gi|1837775... 858 0.0
ref|XP_476653.1| putative proton myo-inositol transporter [Oryza... 790 0.0
gb|AAF91432.1| putative Na+/myo-inositol symporter [Mesembryanth... 694 0.0
emb|CAE03384.1| OSJNBa0004N05.8 [Oryza sativa (japonica cultivar... 690 0.0
emb|CAB10424.1| membrane transporter like protein [Arabidopsis t... 685 0.0
gb|AAF91431.1| putative Na+/myo-inositol symporter [Mesembryanth... 668 0.0
gb|AAD15441.1| putative sugar transporter [Arabidopsis thaliana]... 636 0.0
emb|CAD41357.2| OSJNBa0076N16.21 [Oryza sativa (japonica cultiva... 405 e-111
gb|AAM20155.1| putative membrane transporter protein [Arabidopsi... 397 e-109
gb|AAB64332.1| putative membrane transporter [Arabidopsis thalia... 388 e-106
gb|AAO74897.1| putative Na+/myo-inositol symporter [Mesembryanth... 379 e-103
ref|NP_598295.1| solute carrier family 2 (facilitated glucose tr... 376 e-103
gb|AAH92027.1| Unknown (protein for MGC:84927) [Xenopus laevis] 376 e-102
emb|CAC51116.1| proton myo-inositol transporter [Homo sapiens] g... 374 e-102
emb|CAG01178.1| unnamed protein product [Tetraodon nigroviridis] 369 e-100
emb|CAA86519.2| Hypothetical protein M01F1.5 [Caenorhabditis ele... 332 2e-89
emb|CAE64945.1| Hypothetical protein CBG09776 [Caenorhabditis br... 330 1e-88
emb|CAA16400.1| Hypothetical protein Y51A2D.4 [Caenorhabditis el... 328 3e-88
emb|CAE67430.1| Hypothetical protein CBG12920 [Caenorhabditis br... 327 6e-88
emb|CAA16405.1| Hypothetical protein Y51A2D.5 [Caenorhabditis el... 327 8e-88
>gb|AAM67564.1| unknown protein [Arabidopsis thaliana] gi|18377759|gb|AAL67029.1|
unknown protein [Arabidopsis thaliana]
gi|15220697|ref|NP_174313.1| sugar transporter family
protein [Arabidopsis thaliana]
gi|12320850|gb|AAG50560.1| hypothetical protein
[Arabidopsis thaliana] gi|25309027|pir||D86426
hypothetical protein F12P21.2 - Arabidopsis thaliana
Length = 580
Score = 858 bits (2218), Expect = 0.0
Identities = 412/575 (71%), Positives = 497/575 (85%), Gaps = 5/575 (0%)
Query: 1 MEGGVPE--ADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDF 58
MEGG+ AD SAF+EC SL+WKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDF
Sbjct: 1 MEGGIIHGGADESAFKECFSLTWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDF 60
Query: 59 KAVDRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPN 118
K+VDR TWLQE IVS A+AGAI+GAA+GGW ND+ GRR AI++AD LFL+G++IMAAAPN
Sbjct: 61 KSVDRNTWLQEMIVSMAVAGAIVGAAIGGWANDKLGRRSAILMADFLFLLGAIIMAAAPN 120
Query: 119 PAILLFGRVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAF 178
P++L+ GRVFVG GVGMASM +PLYISEASP ++RGALVS N FLITGGQFLSYLINLAF
Sbjct: 121 PSLLVVGRVFVGLGVGMASMTAPLYISEASPAKIRGALVSTNGFLITGGQFLSYLINLAF 180
Query: 179 TNAPGTWRWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAE 238
T+ GTWRWMLG+A PA++Q VLM +LPESPRWLYRKGREEE+K+IL++IY+ EDV+ E
Sbjct: 181 TDVTGTWRWMLGIAGIPALLQFVLMFTLPESPRWLYRKGREEEAKAILRRIYSAEDVEQE 240
Query: 239 IEALKESVESEI-EESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPT 297
I ALK+SVE+EI EE + KI+M+ L K TVRRGL AG+GLQ FQQFVGINTVMYYSPT
Sbjct: 241 IRALKDSVETEILEEGSSEKINMIKLCKAKTVRRGLIAGVGLQVFQQFVGINTVMYYSPT 300
Query: 298 IVQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTV 357
IVQLAGFASNRTALLLSL+T+GLNAFGSI+SIYFID+ GRKKL +ISL GV++SL +LT
Sbjct: 301 IVQLAGFASNRTALLLSLVTAGLNAFGSIISIYFIDRIGRKKLLIISLFGVIISLGILTG 360
Query: 358 AFRESELHAPTVSAIETSHYNN-TCPDFRAAVNPGRWTCMTCLKA-SPSCGFCAASPNKL 415
F E+ HAP +S++ET +NN +CPD+++A+N W CMTCLKA SPSCG+C++ K
Sbjct: 361 VFYEAATHAPAISSLETQRFNNISCPDYKSAMNTNAWDCMTCLKASSPSCGYCSSPIGKE 420
Query: 416 LPGACLISDDTTKDMCGNDHRAWYTRGCPSKFGWAALVGLALYIITFSPGMGTVPWVINS 475
PGAC ISDD+ KD+C N++R WYTRGCPS FGW AL+GL LYII FSPGMGTVPW++NS
Sbjct: 421 HPGACWISDDSVKDLCHNENRLWYTRGCPSNFGWFALLGLGLYIIFFSPGMGTVPWIVNS 480
Query: 476 EIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVF 535
EIYPLR+RG+CGGIA+T WISNLIV+QSFLSLT+ IGT+WTF++FG+++V+A+ FV+V
Sbjct: 481 EIYPLRFRGICGGIAATANWISNLIVAQSFLSLTEAIGTSWTFLIFGVISVIALLFVMVC 540
Query: 536 VPETKGVPMEEVEKMLEQRSLQFKFWQKRDSGSEK 570
VPETKG+PMEE+EKMLE+RS++FKFW+K+ EK
Sbjct: 541 VPETKGMPMEEIEKMLERRSMEFKFWKKKSKLVEK 575
>ref|XP_476653.1| putative proton myo-inositol transporter [Oryza sativa (japonica
cultivar-group)] gi|33146705|dbj|BAC79509.1| putative
proton myo-inositol transporter [Oryza sativa (japonica
cultivar-group)] gi|50509781|dbj|BAD31907.1| putative
proton myo-inositol transporter [Oryza sativa (japonica
cultivar-group)]
Length = 596
Score = 790 bits (2039), Expect = 0.0
Identities = 399/592 (67%), Positives = 468/592 (78%), Gaps = 22/592 (3%)
Query: 1 MEGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKA 60
MEGGV E D S FREC SLSW+NPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDF +
Sbjct: 1 MEGGVHEFDGSTFRECFSLSWRNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFPS 60
Query: 61 VDRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPA 120
VD+ TWLQE IVS A+AGAIIGAA+GGW NDR+GRR +I++AD LF G+ +MA+A PA
Sbjct: 61 VDKNTWLQEMIVSMAVAGAIIGAAIGGWANDRYGRRTSILVADALFFAGAAVMASATGPA 120
Query: 121 ILLFGRVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTN 180
L+ GRVFVG GVG ASM SPLYISEASP R+RGALVS N LITGGQFLSYLINLAFT
Sbjct: 121 QLVVGRVFVGLGVGTASMTSPLYISEASPARIRGALVSTNGLLITGGQFLSYLINLAFTK 180
Query: 181 APGTWRWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIE 240
APGTWRWMLGVAA PA++Q LML LPESPRWLYRKGREEE+++IL+KIY+ E+V+ E E
Sbjct: 181 APGTWRWMLGVAAIPAVVQFFLMLFLPESPRWLYRKGREEEAEAILRKIYSAEEVEREKE 240
Query: 241 ALKESVESEI-EESKTSKISMMTLLKTT-TVRRGLYAGMGLQFFQQFVGINTVMYYSPTI 298
LKESVE+E E S + K S++ LL TT TVRRGL AG+GLQ FQQ VGINTVMYYSPTI
Sbjct: 241 ELKESVEAEARERSSSEKTSLVALLMTTATVRRGLVAGVGLQVFQQLVGINTVMYYSPTI 300
Query: 299 VQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVA 358
VQLAGFASN+TAL LSL+T+GLNA GS++SIYFID+TGR+KL +ISL GV+LSLALL+
Sbjct: 301 VQLAGFASNQTALALSLVTAGLNAAGSLVSIYFIDRTGRRKLLVISLAGVILSLALLSAV 360
Query: 359 FRESELHAPTVSAIETSHYNN---TCPDFRAAVNPGRWTCMTCLK---ASPSCGFCAA-S 411
F E+ H+P V A ET+H++ TCPD+ + + W C CLK AS CGFCAA
Sbjct: 361 FHEATSHSPPVGAAETAHFHGGALTCPDYSSRSSSSFWDCTRCLKAAAASAGCGFCAAGG 420
Query: 412 PNKLLPGACL-----ISDDTTKDMCGNDHRAWYTRGCPSKFGWAALVGLALYIITFSPGM 466
+KL GACL S+ T +D C + R WYTRGCPS++GW A+ GLALYI FSPGM
Sbjct: 421 GDKLRAGACLAAAAAASNATARDACRGEGREWYTRGCPSRYGWLAMAGLALYIAAFSPGM 480
Query: 467 GTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAV 526
GTVPW++NSE+YPLR+RGVCGG A+T W+SNL V+QSFLSLT IG AWTF++FG L+V
Sbjct: 481 GTVPWIVNSEVYPLRHRGVCGGAAATANWVSNLAVAQSFLSLTDAIGAAWTFLIFGGLSV 540
Query: 527 VAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQKR--------DSGSEK 570
A+ FV+V VPETKG+P+EEVEKMLE R L+ +FW KR D G EK
Sbjct: 541 AALAFVLVCVPETKGLPIEEVEKMLEGRELRLRFWAKRRHHHGGDGDGGGEK 592
>gb|AAF91432.1| putative Na+/myo-inositol symporter [Mesembryanthemum crystallinum]
Length = 581
Score = 694 bits (1790), Expect = 0.0
Identities = 337/559 (60%), Positives = 430/559 (76%), Gaps = 7/559 (1%)
Query: 1 MEGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKA 60
+EGG+ +AD + F EC K PY+LRLAFSAGIGGLLFGYDTGVISGALLYI++DFK
Sbjct: 2 VEGGIVKADKTEFTECFRTIGKTPYILRLAFSAGIGGLLFGYDTGVISGALLYIKEDFKE 61
Query: 61 VDRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPA 120
V+RKTWLQE IV+ A+AGAIIGA VGG++ND+FGR+ AIIIAD LF IG++IM+ AP P
Sbjct: 62 VERKTWLQETIVAMAVAGAIIGAGVGGYLNDKFGRKPAIIIADILFFIGAIIMSLAPAPW 121
Query: 121 ILLFGRVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTN 180
+++ GR+FVG GVGMASM SPLYISE SPTR+R ALVS N LITG QFLSYLINL FT
Sbjct: 122 MIILGRIFVGLGVGMASMTSPLYISETSPTRIRSALVSTNGLLITGSQFLSYLINLGFTR 181
Query: 181 APGTWRWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIE 240
GTWRWMLGVAA PA +Q++LMLSLPESPRWLYRK + E+++IL +IY PE+V+ E+
Sbjct: 182 VKGTWRWMLGVAAVPAFVQLLLMLSLPESPRWLYRKNKVVEAEAILARIYPPEEVEEEMR 241
Query: 241 ALKESVESEI-EESKTSKISMMTLLK----TTTVRRGLYAGMGLQFFQQFVGINTVMYYS 295
ALK S+E E+ EE + SM++ ++ VRRGLYAG+ +Q QQFVGINTVMYYS
Sbjct: 242 ALKASIEYEMAEEGEIGGGSMLSKVRKAWGNKIVRRGLYAGITVQVAQQFVGINTVMYYS 301
Query: 296 PTIVQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALL 355
PTIVQLAGFASN TAL LSL+TSGLNA GSI+S+ F+D+ GR++L +IS+ G++ L +L
Sbjct: 302 PTIVQLAGFASNSTALALSLVTSGLNAIGSIVSMMFVDRHGRRRLMIISMFGIITCLIVL 361
Query: 356 TVAFRESELHAPTVSAIETSHY--NNTCPDFRAAVNPGRWTCMTCLKASPSCGFCAASPN 413
+ F ++ HAP +S E++H+ N+TCP + NP W CMTCL+A+ C FC N
Sbjct: 362 AIGFFQAAAHAPKISHAESTHFGLNSTCPAYTTTRNPATWNCMTCLQAASECAFCTNKGN 421
Query: 414 KLLPGACLISDDTTKDMCGNDHRAWYTRGCPSKFGWAALVGLALYIITFSPGMGTVPWVI 473
+LLPG C+ D K C + R ++T GCPSKFG+ A++ L YII++SPGMGTVPW++
Sbjct: 422 QLLPGGCVSRTDAMKVACHGEKRVYFTEGCPSKFGFLAVILLGAYIISYSPGMGTVPWIV 481
Query: 474 NSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFFVI 533
NSEIYPLRYRGV GGIA+ + W SNLIVS++FL+LT+ +G A TF++F + + + F+
Sbjct: 482 NSEIYPLRYRGVGGGIAAVSNWTSNLIVSETFLTLTEALGAAGTFLLFAGFSAIGLVFIY 541
Query: 534 VFVPETKGVPMEEVEKMLE 552
+ VPETKG+P+EEVE MLE
Sbjct: 542 LLVPETKGLPIEEVEHMLE 560
>emb|CAE03384.1| OSJNBa0004N05.8 [Oryza sativa (japonica cultivar-group)]
gi|50926394|ref|XP_473144.1| OSJNBa0004N05.8 [Oryza
sativa (japonica cultivar-group)]
Length = 581
Score = 690 bits (1780), Expect = 0.0
Identities = 340/571 (59%), Positives = 434/571 (75%), Gaps = 15/571 (2%)
Query: 1 MEGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKA 60
MEGG AD + F+ECL L+W PY+L+L FSAGIGGLLFGYDTGVISGALLYIRDDF A
Sbjct: 1 MEGGATLADKAEFKECLRLTWSQPYILQLVFSAGIGGLLFGYDTGVISGALLYIRDDFTA 60
Query: 61 VDRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPA 120
V++ T L+E IVS A+AGAI+GA GGW+ND+FGR+ +I+IAD+LFL G++IMA AP P
Sbjct: 61 VEKSTVLRETIVSMAVAGAIVGAGFGGWMNDKFGRKPSILIADSLFLAGALIMALAPTPF 120
Query: 121 ILLFGRVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTN 180
+++ GR+FVG GVGMASM +PLYISEASP R+RGALVS N LITGGQF++YLINLAFT
Sbjct: 121 VIIIGRIFVGLGVGMASMTAPLYISEASPARIRGALVSTNGLLITGGQFMAYLINLAFTK 180
Query: 181 APGTWRWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIE 240
GTWRWMLG+A PA IQ +LM LPESPRWLYR+ R+EE+++IL+KIY +V+ EI+
Sbjct: 181 VKGTWRWMLGIAGLPAFIQFILMCMLPESPRWLYRQDRKEEAEAILRKIYPAAEVEEEID 240
Query: 241 ALKESVESEI-------EESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMY 293
+++ S+E E E+S K++ L + VRRGL AG+ Q QQFVGINTVMY
Sbjct: 241 SMRRSIEHEKQLEGSIGEQSLVGKLT--KALSSKVVRRGLMAGVIAQVAQQFVGINTVMY 298
Query: 294 YSPTIVQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLA 353
YSPTIVQLAGFASN TA+ LSLITSGLNA GSI+S++F+D+ GR++L +ISL G+VL LA
Sbjct: 299 YSPTIVQLAGFASNNTAMALSLITSGLNAIGSIVSMFFVDRAGRRRLMIISLVGIVLWLA 358
Query: 354 LLTVAFRESELHAPTVSAIETSHY-NNTCPDFRAAVNPGRWTCMTCLKASPSCGFCAASP 412
+L F + HAP VS +ET + N TCP++ + RW CM CLKA +CGFCA
Sbjct: 359 VLGGTFLGAAHHAPPVSDLETRVFANQTCPEYSPS---ARWNCMNCLKAQSTCGFCAHGG 415
Query: 413 NKLLPGACLISDDTTKDMCGNDHRAWYTRGCPSKFGWAALVGLALYIITFSPGMGTVPWV 472
NKLLPGACL + + ++ C +R +YT GCP+ FGW ALV L YI+++SPGMGTVPW+
Sbjct: 416 NKLLPGACLAAGEASRRTCHAGNREFYTEGCPNNFGWLALVALGAYIVSYSPGMGTVPWI 475
Query: 473 INSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFFV 532
+NSEIYPLR+RGVCGGIA+ W+SNLIV+Q+FLSLT+ +GT+ TF +F ++ A+ V
Sbjct: 476 VNSEIYPLRFRGVCGGIAAVANWVSNLIVTQTFLSLTKALGTSATFFLFCAVSFFALVVV 535
Query: 533 IVFVPETKGVPMEEVEKMLEQRSLQFKFWQK 563
VPETKG+ EEVEKML ++ +K W++
Sbjct: 536 FFTVPETKGLQFEEVEKMLGEK--DYKPWKR 564
>emb|CAB10424.1| membrane transporter like protein [Arabidopsis thaliana]
gi|7268398|emb|CAB78690.1| membrane transporter like
protein [Arabidopsis thaliana]
gi|28973605|gb|AAO64127.1| putative membrane transporter
[Arabidopsis thaliana] gi|28393478|gb|AAO42160.1|
putative membrane transporter [Arabidopsis thaliana]
gi|15235767|ref|NP_193381.1| sugar transporter family
protein [Arabidopsis thaliana] gi|7446726|pir||F71431
hypothetical protein - Arabidopsis thaliana
Length = 582
Score = 685 bits (1767), Expect = 0.0
Identities = 337/562 (59%), Positives = 429/562 (75%), Gaps = 12/562 (2%)
Query: 1 MEGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKA 60
+EGG+ +AD + F EC +WK PY++RLA SAGIGGLLFGYDTGVISGALL+I++DF
Sbjct: 2 VEGGIAKADKTEFTECWRTTWKTPYIMRLALSAGIGGLLFGYDTGVISGALLFIKEDFDE 61
Query: 61 VDRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPA 120
VD+KTWLQ IVS A+AGAI+GAAVGGWIND+FGRR++I+IAD LFLIG+++MA AP P
Sbjct: 62 VDKKTWLQSTIVSMAVAGAIVGAAVGGWINDKFGRRMSILIADVLFLIGAIVMAFAPAPW 121
Query: 121 ILLFGRVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTN 180
+++ GR+FVGFGVGMASM SPLYISEASP R+RGALVS N LITGGQF SYLINLAF +
Sbjct: 122 VIIVGRIFVGFGVGMASMTSPLYISEASPARIRGALVSTNGLLITGGQFFSYLINLAFVH 181
Query: 181 APGTWRWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIE 240
PGTWRWMLGVA PAI+Q VLMLSLPESPRWLYRK R ES++IL++IY ++V+AE+E
Sbjct: 182 TPGTWRWMLGVAGVPAIVQFVLMLSLPESPRWLYRKDRIAESRAILERIYPADEVEAEME 241
Query: 241 ALKESVESEIEESKTSKISMMTLLK----TTTVRRGLYAGMGLQFFQQFVGINTVMYYSP 296
ALK SVE+E + S LK VRRGL AG+ +Q QQFVGINTVMYYSP
Sbjct: 242 ALKLSVEAEKADEAIIGDSFSAKLKGAFGNPVVRRGLAAGITVQVAQQFVGINTVMYYSP 301
Query: 297 TIVQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLT 356
+IVQ AG+ASN+TA+ LSLITSGLNA GSI+S+ F+D+ GR+KL +IS+ G++ L +L
Sbjct: 302 SIVQFAGYASNKTAMALSLITSGLNALGSIVSMMFVDRYGRRKLMIISMFGIIACLIILA 361
Query: 357 VAFRESELHAPTVSAIETSHY--NNTCPDFR--AAVN--PGRWTCMTCLKASPSCGFCAA 410
F ++ +HAP + A E+ + N TC + AA N P RW CM CL++ CGFCA+
Sbjct: 362 TVFSQAAIHAPKIDAFESRTFAPNATCSAYAPLAAENAPPSRWNCMKCLRS--ECGFCAS 419
Query: 411 SPNKLLPGACLISDDTTKDMCGNDHRAWYTRGCPSKFGWAALVGLALYIITFSPGMGTVP 470
PGAC++ D K C + R ++ GCPSKFG+ A+V L LYI+ ++PGMGTVP
Sbjct: 420 GVQPYAPGACVVLSDDMKATCSSRGRTFFKDGCPSKFGFLAIVFLGLYIVVYAPGMGTVP 479
Query: 471 WVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIF 530
W++NSEIYPLRYRG+ GGIA+ + W+SNLIVS+SFLSLT +G++ TF++F + + +F
Sbjct: 480 WIVNSEIYPLRYRGLGGGIAAVSNWVSNLIVSESFLSLTHALGSSGTFLLFAGFSTIGLF 539
Query: 531 FVIVFVPETKGVPMEEVEKMLE 552
F+ + VPETKG+ EEVEK+LE
Sbjct: 540 FIWLLVPETKGLQFEEVEKLLE 561
>gb|AAF91431.1| putative Na+/myo-inositol symporter [Mesembryanthemum crystallinum]
Length = 581
Score = 668 bits (1723), Expect = 0.0
Identities = 320/576 (55%), Positives = 432/576 (74%), Gaps = 13/576 (2%)
Query: 1 MEGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKA 60
+EGGV + D + F EC K PY++RLAFSAGIGGLLFGYDTGVISGALLYI++DFK
Sbjct: 2 VEGGVVKVDKTEFTECFRTVGKTPYIMRLAFSAGIGGLLFGYDTGVISGALLYIKEDFKE 61
Query: 61 VDRKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPA 120
V +KTWLQE IV+ A+AGAI+GA +GG++ND+FGR+ A+I+AD LFL G++IM+ AP P
Sbjct: 62 VAQKTWLQETIVAMAVAGAIVGAGLGGFLNDKFGRKPAMIVADILFLTGAIIMSVAPAPW 121
Query: 121 ILLFGRVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTN 180
+++ GR+ VG GVGMASM +PLYISE SP ++RGAL + N LITGGQF+SYL+NL FT
Sbjct: 122 VIIIGRIVVGLGVGMASMTAPLYISETSPAKIRGALGATNGLLITGGQFVSYLVNLGFTR 181
Query: 181 APGTWRWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIE 240
GTWRWMLGVAA PA IQ+VLML+LPESPRWLYR+ + E++ IL +IY PE V E++
Sbjct: 182 VKGTWRWMLGVAAVPAAIQVVLMLTLPESPRWLYRQNKISEAEEILGRIYPPEQVKEEMD 241
Query: 241 ALKESVESEIEESK-----TSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYS 295
+LK S+E+E+ + K + + VRRGL AG+ + QQFVGINTVMYYS
Sbjct: 242 SLKTSIENEMADRKAVGEGNAFVRAKRAWDNKVVRRGLIAGISVLVAQQFVGINTVMYYS 301
Query: 296 PTIVQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALL 355
PTI+QLAGFASN TAL LSL+TSGLNA GSI+S+ F+D+ GR++L +IS+ ++ L +L
Sbjct: 302 PTIIQLAGFASNSTALALSLVTSGLNAVGSIVSMMFVDRFGRRRLMIISMFAIITCLVVL 361
Query: 356 TVAFRESELHAPTVSAIETSHY--NNTCPDFRAAVNPGRWTCMTCLKASPSCGFCAASPN 413
+ F + AP +S +E+SH+ N+TCP F +A +P RW CMTCLKAS C FC+ S +
Sbjct: 362 SGLFYGAAQAAPKISQLESSHFGANSTCPAFASATSPDRWNCMTCLKAS-DCAFCSNSAS 420
Query: 414 KLLPGACLISDDTTKDMCGNDHRAWYTRGCPSKFGWAALVGLALYIITFSPGMGTVPWVI 473
+ PGAC+ T K+ C + R ++T GCPSKFG+ A++ L LYIIT+SPGMGTVPW++
Sbjct: 421 EFHPGACVAQTSTMKNACLGEKRIYFTEGCPSKFGFMAIIVLGLYIITYSPGMGTVPWIL 480
Query: 474 NSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFFVI 533
NSEIYPLRYRG+CGGI + T+W +NLIVS++FL+LT+ +G++ TF+++ +++ + +
Sbjct: 481 NSEIYPLRYRGICGGIGAVTLWCANLIVSETFLTLTEALGSSGTFLLYAGFSLIGLIVIF 540
Query: 534 VFVPETKGVPMEEVEKMLEQRSLQFKFWQKRDSGSE 569
+ VPETKG+P+E++EKMLE+ FW G++
Sbjct: 541 LLVPETKGLPIEDIEKMLEK-----GFWPSLCGGNK 571
>gb|AAD15441.1| putative sugar transporter [Arabidopsis thaliana]
gi|15227479|ref|NP_181117.1| sugar transporter family
protein [Arabidopsis thaliana] gi|25309025|pir||D84772
probable sugar transporter [imported] - Arabidopsis
thaliana
Length = 580
Score = 636 bits (1640), Expect = 0.0
Identities = 309/548 (56%), Positives = 406/548 (73%), Gaps = 12/548 (2%)
Query: 15 ECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVST 74
E + +W+ PY++RLA SAGIGGLLFGY+TGVI+GALLYI+++F VD KTWLQE IVS
Sbjct: 15 EVWTTTWETPYIMRLALSAGIGGLLFGYNTGVIAGALLYIKEEFGEVDNKTWLQEIIVSM 74
Query: 75 AIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVG 134
+AGAI+GAA+GGW ND+FGRR++++IAD LFL+G+++M A P +++ GR+ VGFGVG
Sbjct: 75 TVAGAIVGAAIGGWYNDKFGRRMSVLIADVLFLLGALVMVIAHAPWVIILGRLLVGFGVG 134
Query: 135 MASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGTWRWMLGVAAA 194
MASM SPLYISE SP R+RGALVS N LITGGQFLSYLINLAF + PGTWRWMLGV+A
Sbjct: 135 MASMTSPLYISEMSPARIRGALVSTNGLLITGGQFLSYLINLAFVHTPGTWRWMLGVSAI 194
Query: 195 PAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEE-- 252
PAIIQ LML+LPESPRWLYR R+ ES+ IL++IY E V+AEI ALKESV +E +
Sbjct: 195 PAIIQFCLMLTLPESPRWLYRNDRKAESRDILERIYPAEMVEAEIAALKESVRAETADED 254
Query: 253 --SKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTA 310
T + L VR GL AG+ +Q QQFVGINTVMYYSPTI+Q AG+ASN+TA
Sbjct: 255 IIGHTFSDKLRGALSNPVVRHGLAAGITVQVAQQFVGINTVMYYSPTILQFAGYASNKTA 314
Query: 311 LLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVAFRESELHAPTVS 370
+ L+LITSGLNA GS++S+ F+D+ GR+KL +IS+ G++ L +L F E+ HAP +
Sbjct: 315 MALALITSGLNAVGSVVSMMFVDRYGRRKLMIISMFGIITCLVILAAVFNEASNHAPKID 374
Query: 371 AIETSHY--NNTCPDF----RAAVNPGRWTCMTCLKASPSCGFCAASPNKLLPGACLISD 424
++ ++ N TCP F + P W CM CL+ CGFC+ + PGAC++
Sbjct: 375 KRDSRNFAKNATCPAFAPFTASRSPPSNWNCMKCLQY--DCGFCSNGAQEYAPGACIVQS 432
Query: 425 DTTKDMCGNDHRAWYTRGCPSKFGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRG 484
K +C + R ++ GCPSKFG+ A+V L LYII ++PGMGTVPW++NSEIYPLRYRG
Sbjct: 433 ADMKALCHSKGRTFFKDGCPSKFGYLAIVFLGLYIIVYAPGMGTVPWIVNSEIYPLRYRG 492
Query: 485 VCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPM 544
+ GGIA+ + W+SNL+VS++FL+LT +G++ TF++F + V +FF+ + VPETKG+
Sbjct: 493 LAGGIAAVSNWMSNLVVSETFLTLTNAVGSSGTFLLFAGSSAVGLFFIWLLVPETKGLQF 552
Query: 545 EEVEKMLE 552
EEVEK+LE
Sbjct: 553 EEVEKLLE 560
>emb|CAD41357.2| OSJNBa0076N16.21 [Oryza sativa (japonica cultivar-group)]
gi|50925717|ref|XP_472996.1| OSJNBa0076N16.21 [Oryza
sativa (japonica cultivar-group)]
Length = 506
Score = 405 bits (1040), Expect = e-111
Identities = 205/339 (60%), Positives = 256/339 (75%), Gaps = 2/339 (0%)
Query: 21 WKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAI 80
+ N YVL L +AGIGG LFGYDTGVISGALLYIRDDF AV +LQE IVS A+ GAI
Sbjct: 26 FSNRYVLALTGAAGIGGFLFGYDTGVISGALLYIRDDFPAVRDNYFLQETIVSMALVGAI 85
Query: 81 IGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMAS 140
IGAA GGWIND +GRR + ++AD LF +GS++M AA P IL+ GR+ VG GVG+AS+ +
Sbjct: 86 IGAAGGGWINDTYGRRKSTLVADMLFALGSLVMCAAGGPYILILGRLLVGLGVGIASVTA 145
Query: 141 PLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGTWRWMLGVAAAPAIIQI 200
P+YI+EA+P+ +RG LVS N +ITGGQF SYLINL FT PGTWRWMLGVAA PAI+Q
Sbjct: 146 PVYIAEAAPSEIRGGLVSTNVLMITGGQFFSYLINLGFTEVPGTWRWMLGVAAVPAILQF 205
Query: 201 VLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEESKTSKISM 260
VLML LPESPRWL+ K + ++ S+L+KIY + ++ E+E L S E + T S
Sbjct: 206 VLMLFLPESPRWLFWKDEKAKAISVLEKIYDSDRLEEEVELLASSSMHEFQSDGTG--SY 263
Query: 261 MTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGL 320
+ + K+ +R +AG GLQ FQQF GINTVMYYSPTIVQ+AGF SN+ ALLLSLI +G+
Sbjct: 264 LDIFKSKELRLAFFAGAGLQAFQQFTGINTVMYYSPTIVQMAGFTSNKLALLLSLIVAGM 323
Query: 321 NAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVAF 359
NA G+I+ IY ID+ GR++LAL SL GVV+SLA+L +AF
Sbjct: 324 NAAGTIVGIYLIDRCGRRRLALTSLAGVVVSLAILAMAF 362
Score = 139 bits (350), Expect = 2e-31
Identities = 64/130 (49%), Positives = 90/130 (69%), Gaps = 4/130 (3%)
Query: 426 TTKDMCGNDHRAWYTRGCPSKFGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGV 485
++ D+C N C GW A+ GLALYI FSPGMG VPW +NSEIYP YRG+
Sbjct: 366 SSSDICSNALNG----ACQGALGWFAVAGLALYIAFFSPGMGPVPWAVNSEIYPEAYRGM 421
Query: 486 CGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPME 545
CGG+++T W+SNLIV+Q+FLS+ +GT TF++ +AV+A FV ++VPETKG+ E
Sbjct: 422 CGGMSATVNWVSNLIVAQTFLSIVGLVGTGLTFLIIAGIAVLAFIFVALYVPETKGLSFE 481
Query: 546 EVEKMLEQRS 555
+VE + ++R+
Sbjct: 482 QVELLWKERA 491
>gb|AAM20155.1| putative membrane transporter protein [Arabidopsis thaliana]
gi|17380890|gb|AAL36257.1| putative membrane transporter
protein [Arabidopsis thaliana]
gi|30689342|ref|NP_850393.1| sugar transporter family
protein [Arabidopsis thaliana]
Length = 509
Score = 397 bits (1020), Expect = e-109
Identities = 201/359 (55%), Positives = 264/359 (72%), Gaps = 11/359 (3%)
Query: 6 PEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKT 65
PE MS F N Y+L L +AGIGGLLFGYDTGVISGALLYI+DDF+ V + +
Sbjct: 19 PERRMSYFG--------NSYILGLTVTAGIGGLLFGYDTGVISGALLYIKDDFEVVKQSS 70
Query: 66 WLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFG 125
+LQE IVS A+ GA+IGAA GGWIND +GR+ A + AD +F G+++MAAAP+P +L+ G
Sbjct: 71 FLQETIVSMALVGAMIGAAAGGWINDYYGRKKATLFADVVFAAGAIVMAAAPDPYVLISG 130
Query: 126 RVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGTW 185
R+ VG GVG+AS+ +P+YI+EASP+ VRG LVS N +ITGGQFLSYL+N AFT PGTW
Sbjct: 131 RLLVGLGVGVASVTAPVYIAEASPSEVRGGLVSTNVLMITGGQFLSYLVNSAFTQVPGTW 190
Query: 186 RWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKES 245
RWMLGV+ PA+IQ +LML +PESPRWL+ K R+ E+ +L + Y ++ EI+ L +
Sbjct: 191 RWMLGVSGVPAVIQFILMLFMPESPRWLFMKNRKAEAIQVLARTYDISRLEDEIDHLSAA 250
Query: 246 VESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFA 305
E E + +T + + + ++ +R AG GLQ FQQF GINTVMYYSPTIVQ+AGF
Sbjct: 251 EEEEKQRKRT--VGYLDVFRSKELRLAFLAGAGLQAFQQFTGINTVMYYSPTIVQMAGFH 308
Query: 306 SNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVA-FRESE 363
SN+ AL LSLI + +NA G+++ IYFID GRKKLAL SL GV++SL +L+V+ F++SE
Sbjct: 309 SNQLALFLSLIVAAMNAAGTVVGIYFIDHCGRKKLALSSLFGVIISLLILSVSFFKQSE 367
Score = 143 bits (360), Expect = 2e-32
Identities = 61/109 (55%), Positives = 89/109 (80%)
Query: 447 FGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFL 506
+GW A++GLALYI+ F+PGMG VPW +NSEIYP +YRG+CGG+++T WISNLIV+Q+FL
Sbjct: 375 YGWLAVLGLALYIVFFAPGMGPVPWTVNSEIYPQQYRGICGGMSATVNWISNLIVAQTFL 434
Query: 507 SLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRS 555
++ + GT TF++ +AV+A+ FVIVFVPET+G+ EVE++ ++R+
Sbjct: 435 TIAEAAGTGMTFLILAGIAVLAVIFVIVFVPETQGLTFSEVEQIWKERA 483
>gb|AAB64332.1| putative membrane transporter [Arabidopsis thaliana]
gi|25308955|pir||G84864 probable membrane transporter
[imported] - Arabidopsis thaliana
Length = 521
Score = 388 bits (997), Expect = e-106
Identities = 201/371 (54%), Positives = 264/371 (70%), Gaps = 23/371 (6%)
Query: 6 PEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKT 65
PE MS F N Y+L L +AGIGGLLFGYDTGVISGALLYI+DDF+ V + +
Sbjct: 19 PERRMSYFG--------NSYILGLTVTAGIGGLLFGYDTGVISGALLYIKDDFEVVKQSS 70
Query: 66 WLQ------------EAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIM 113
+LQ E IVS A+ GA+IGAA GGWIND +GR+ A + AD +F G+++M
Sbjct: 71 FLQVYNVSSFTSSKLETIVSMALVGAMIGAAAGGWINDYYGRKKATLFADVVFAAGAIVM 130
Query: 114 AAAPNPAILLFGRVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYL 173
AAAP+P +L+ GR+ VG GVG+AS+ +P+YI+EASP+ VRG LVS N +ITGGQFLSYL
Sbjct: 131 AAAPDPYVLISGRLLVGLGVGVASVTAPVYIAEASPSEVRGGLVSTNVLMITGGQFLSYL 190
Query: 174 INLAFTNAPGTWRWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPE 233
+N AFT PGTWRWMLGV+ PA+IQ +LML +PESPRWL+ K R+ E+ +L + Y
Sbjct: 191 VNSAFTQVPGTWRWMLGVSGVPAVIQFILMLFMPESPRWLFMKNRKAEAIQVLARTYDIS 250
Query: 234 DVDAEIEALKESVESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMY 293
++ EI+ L + E E + +T + + + ++ +R AG GLQ FQQF GINTVMY
Sbjct: 251 RLEDEIDHLSAAEEEEKQRKRT--VGYLDVFRSKELRLAFLAGAGLQAFQQFTGINTVMY 308
Query: 294 YSPTIVQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLA 353
YSPTIVQ+AGF SN+ AL LSLI + +NA G+++ IYFID GRKKLAL SL GV++SL
Sbjct: 309 YSPTIVQMAGFHSNQLALFLSLIVAAMNAAGTVVGIYFIDHCGRKKLALSSLFGVIISLL 368
Query: 354 LLTVA-FRESE 363
+L+V+ F++SE
Sbjct: 369 ILSVSFFKQSE 379
Score = 143 bits (360), Expect = 2e-32
Identities = 61/109 (55%), Positives = 89/109 (80%)
Query: 447 FGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFL 506
+GW A++GLALYI+ F+PGMG VPW +NSEIYP +YRG+CGG+++T WISNLIV+Q+FL
Sbjct: 387 YGWLAVLGLALYIVFFAPGMGPVPWTVNSEIYPQQYRGICGGMSATVNWISNLIVAQTFL 446
Query: 507 SLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRS 555
++ + GT TF++ +AV+A+ FVIVFVPET+G+ EVE++ ++R+
Sbjct: 447 TIAEAAGTGMTFLILAGIAVLAVIFVIVFVPETQGLTFSEVEQIWKERA 495
>gb|AAO74897.1| putative Na+/myo-inositol symporter [Mesembryanthemum crystallinum]
Length = 498
Score = 379 bits (974), Expect = e-103
Identities = 190/337 (56%), Positives = 247/337 (72%), Gaps = 3/337 (0%)
Query: 23 NPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAIIG 82
N +VL L +AGIGGLLFGYDTGVISGALLYI+D+F AV ++LQE IVS A+ GA+IG
Sbjct: 26 NKFVLALTVTAGIGGLLFGYDTGVISGALLYIKDEFPAVKNSSFLQETIVSMALVGAMIG 85
Query: 83 AAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPL 142
+A GWIND +GR+ A ++AD +F IG+V+MAAAP+P IL+ GR VG GVG+AS+ +P+
Sbjct: 86 SATAGWINDVYGRKKATLLADFIFAIGAVVMAAAPDPYILIVGRFLVGLGVGLASVCAPV 145
Query: 143 YISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGTWRWMLGVAAAPAIIQIVL 202
YI+EASPT VRG LVS N +IT GQF+SY +NLAFT PGTWRWMLGV+ PA++Q
Sbjct: 146 YIAEASPTEVRGGLVSTNVLMITFGQFVSYCVNLAFTEVPGTWRWMLGVSGVPAVLQFGF 205
Query: 203 MLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEESKTSKISMMT 262
ML LPESPRWLY K + ++ ++L KIY P ++ E++ L +E EE + +
Sbjct: 206 MLLLPESPRWLYLKHEKSKAAAVLAKIYDPFRLEDELDLL---AAAEEEEKNKPAVHISD 262
Query: 263 LLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNA 322
+ +R AG GL FQQ GINTVMYYSPTIVQ+AGF+SN+ ALL+SLI + +NA
Sbjct: 263 VFTKRELRYAFIAGGGLLAFQQLAGINTVMYYSPTIVQMAGFSSNQLALLISLIVAAMNA 322
Query: 323 FGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVAF 359
G++L IY ID GR+KLAL SL GV ++L +LT++F
Sbjct: 323 VGTVLGIYLIDHMGRRKLALTSLSGVFVALVMLTISF 359
Score = 131 bits (329), Expect = 7e-29
Identities = 59/109 (54%), Positives = 78/109 (71%)
Query: 447 FGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFL 506
+ W A++GLALYI F+PGMG VPW INSEIYP YRG+CGG+ +T WI NL VS++FL
Sbjct: 371 YSWLAVLGLALYIAFFAPGMGPVPWAINSEIYPQAYRGLCGGMGATICWIVNLFVSETFL 430
Query: 507 SLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRS 555
S+ IGT TF++ + +VA FV+ FVPETK + EEV++M R+
Sbjct: 431 SIADAIGTGPTFLIIAGIVIVAFVFVVCFVPETKALTFEEVDQMFMDRA 479
>ref|NP_598295.1| solute carrier family 2 (facilitated glucose transporter), member
13 [Rattus norvegicus] gi|15211931|emb|CAC51117.1|
proton myo-inositol transporter [Rattus norvegicus]
gi|51316137|sp|Q921A2|MYCT_RAT Proton myo-inositol
cotransporter (H(+)-myo-inositol cotransporter) (Hmit)
Length = 618
Score = 376 bits (966), Expect = e-103
Identities = 213/545 (39%), Positives = 316/545 (57%), Gaps = 38/545 (6%)
Query: 30 AFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAIIGAAVGGWI 89
AFSA +GG LFGYDTGV+SGA+L +R + W QE +VS A+ A + A GG +
Sbjct: 56 AFSA-LGGFLFGYDTGVVSGAMLLLRRQMRL--GAMW-QELLVSGAVGAAAVAALAGGAL 111
Query: 90 NDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPLYISEASP 149
N GRR AI++A L +GS ++AAA N LL GR+ VG G+G+ASM P+YI+E SP
Sbjct: 112 NGALGRRSAILLASALCTVGSAVLAAAANKETLLAGRLVVGLGIGIASMTVPVYIAEVSP 171
Query: 150 TRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGT-WRWMLGVAAAPAIIQIVLMLSLPE 208
+RG LV++N+ ITGGQF + +++ AF+ WR+MLG+AA PA+IQ + L LPE
Sbjct: 172 PNLRGRLVTINTLFITGGQFFASVVDGAFSYLQKDGWRYMLGLAAIPAVIQFLGFLFLPE 231
Query: 209 SPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEESKTSKISMMTLLKTTT 268
SPRWL +KG+ ++++ IL ++ + +D E ++++ S+E E +E+ + + +L
Sbjct: 232 SPRWLIQKGQTQKARRILSQMRGNQTIDEEYDSIRNSIEEEEKEASAAGPIICRMLSYPP 291
Query: 269 VRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAFGSILS 328
RR L G GLQ FQQ GINT+MYYS TI+Q++G +R A+ L+ IT+ N +++
Sbjct: 292 TRRALAVGCGLQMFQQLSGINTIMYYSATILQMSGVEDDRLAIWLASITAFTNFIFTLVG 351
Query: 329 IYFIDKTGRKKLALISLCGVVLSLALLTVAFRESELHAPTVSAIET--SHYNNTCPDFRA 386
++ ++K GR+KL SL G ++L +L + F S +P V+ T S N TC ++
Sbjct: 352 VWLVEKVGRRKLTFGSLAGTTVALTILALGFLLSAQVSPRVTFRPTAPSGQNATCTEYS- 410
Query: 387 AVNPGRWTCMTCLKASPSCGFC-----------------AASPNKLLPGACLISDDTTKD 429
C C+ P CGFC AS N+ G C ++ TK
Sbjct: 411 -------YCNECM-LDPDCGFCYKINSSAVIDSSCVPVNKASTNEAAWGRC---ENETKF 459
Query: 430 MCGNDHRAWYTRGCPSKFGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGI 489
+ H W CP+ + W ALVGL LY++ F+PGMG +PW +NSEIYPL R
Sbjct: 460 KAEDVH--WAYSFCPTPYSWTALVGLVLYLVFFAPGMGPMPWTVNSEIYPLWARSTGNAC 517
Query: 490 ASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEK 549
++ WI N++VS +FL + + F ++ A V + FV +PETKG +EE+E
Sbjct: 518 SAGINWIFNVLVSLTFLHTAEYLTYYGAFFLYAGFAAVGLLFVYGCLPETKGKKLEEIES 577
Query: 550 MLEQR 554
+ + R
Sbjct: 578 LFDHR 582
>gb|AAH92027.1| Unknown (protein for MGC:84927) [Xenopus laevis]
Length = 604
Score = 376 bits (965), Expect = e-102
Identities = 203/554 (36%), Positives = 326/554 (58%), Gaps = 27/554 (4%)
Query: 25 YVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAIIGAA 84
+V ++ + +GG LFGYDTGV+SGA+L ++ + ++ QE +VS+ + A + A
Sbjct: 61 FVYVVSVFSALGGFLFGYDTGVVSGAMLLLK---REMNLSALWQELLVSSTVGAAAVSAL 117
Query: 85 VGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPLYI 144
GG +N GRR I++A LF G+VI+AAA + LL GRV VG G+G+ASM P+YI
Sbjct: 118 AGGGLNGVLGRRPCILMASLLFTAGAVILAAARDKETLLGGRVVVGLGIGIASMTVPVYI 177
Query: 145 SEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTN-APGTWRWMLGVAAAPAIIQIVLM 203
+EA+P +RG LV++N+ ITGGQF + +++ AF+ A WR+MLG++A PA++Q +
Sbjct: 178 AEAAPPHLRGRLVTINTLFITGGQFFAAVVDGAFSYLARDGWRYMLGLSAVPAVLQFLGF 237
Query: 204 LSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESEIEESKTSKISMMTL 263
L LPESPRWL +KG+ ++++ +L +I + +D E +++K S++ E +E T + +
Sbjct: 238 LFLPESPRWLIQKGQTQKARRVLSQIRGNQTIDEEYDSIKNSIDEEEKEGGTGGPIIYRM 297
Query: 264 LKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAF 323
L RR L G GLQ FQQ GINTVMYY+ TI+Q++G +R A+ L+ +T+ N
Sbjct: 298 LIYPPTRRALIVGCGLQMFQQLAGINTVMYYNATILQMSGVEDDRLAIWLAAVTAFTNFS 357
Query: 324 GSILSIYFIDKTGRKKLALISLCGVVLSLALLTVAFRESELHAPTVSAI--ETSHYNNTC 381
++L ++ ++K GR+KL L SL G ++L +L + F S +P V+ + S N+TC
Sbjct: 358 FTLLGVWLVEKLGRRKLTLGSLTGTTVALFVLALGFLLSAQVSPPVTFTPGDPSGQNSTC 417
Query: 382 PDFRAAVNPGRWTCMTCLKASPSCGFC-AASPNKLLPGACLISDDTTKD-----MCGNDH 435
+ C C+ P+CGFC + + ++ +C+ + + D C N+
Sbjct: 418 TKYS--------YCNKCM-LDPNCGFCFKKNGSSVVDSSCVPVNPESTDRAAWGRCMNES 468
Query: 436 RA------WYTRGCPSKFGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRYRGVCGGI 489
R+ W CP+ + W LVGL LY++ F+PGMG +PW +NSEIYPL R
Sbjct: 469 RSKAEGSIWAYNYCPTPYSWTVLVGLILYLVFFAPGMGPMPWTVNSEIYPLWARSTGNAC 528
Query: 490 ASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEK 549
++ WI N+++S +FL + + F ++ A V + F+ +PETKG +EE+E
Sbjct: 529 SAGVNWIFNVLISLTFLHTAEFLTYYGAFFLYAGFACVGLIFIYGCLPETKGKKLEEIES 588
Query: 550 MLEQRSLQFKFWQK 563
+ E + + W +
Sbjct: 589 LFESKLYADRMWHR 602
>emb|CAC51116.1| proton myo-inositol transporter [Homo sapiens]
gi|20177982|sp|Q96QE2|MYCT_HUMAN Proton myo-inositol
cotransporter (H(+)-myo-inositol cotransporter) (Hmit)
gi|16418395|ref|NP_443117.1| solute carrier family 2
(facilitated glucose transporter), member 13 [Homo
sapiens]
Length = 629
Score = 374 bits (960), Expect = e-102
Identities = 207/567 (36%), Positives = 325/567 (56%), Gaps = 28/567 (4%)
Query: 3 GGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVD 62
GGV + + +A R+ +V +A + +GG LFGYDTGV+SGA+L ++ + +
Sbjct: 40 GGVGDLERAARRQ-FQQDETPAFVYVVAVFSALGGFLFGYDTGVVSGAMLLLK---RQLS 95
Query: 63 RKTWLQEAIVSTAIAGAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAIL 122
QE +VS+ + A + A GG +N FGRR AI++A LF GS ++AAA N L
Sbjct: 96 LDALWQELLVSSTVGAAAVSALAGGALNGVFGRRAAILLASALFTAGSAVLAAANNKETL 155
Query: 123 LFGRVFVGFGVGMASMASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAP 182
L GR+ VG G+G+ASM P+YI+E SP +RG LV++N+ ITGGQF + +++ AF+
Sbjct: 156 LAGRLVVGLGIGIASMTVPVYIAEVSPPNLRGRLVTINTLFITGGQFFASVVDGAFSYLQ 215
Query: 183 GT-WRWMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEA 241
WR+MLG+A PA+IQ L LPESPRWL +KG+ ++++ IL ++ + +D E ++
Sbjct: 216 KDGWRYMLGLAXVPAVIQFFGFLFLPESPRWLIQKGQTQKARRILSQMRGNQTIDEEYDS 275
Query: 242 LKESVESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQL 301
+K ++E E +E ++ + +L RR L G GLQ FQQ GINT+MYYS TI+Q+
Sbjct: 276 IKNNIEEEEKEVGSAGPVICRMLSYPPTRRALIVGCGLQMFQQLSGINTIMYYSATILQM 335
Query: 302 AGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVAFRE 361
+G +R A+ L+ +T+ N +++ ++ ++K GR+KL SL G ++L +L + F
Sbjct: 336 SGVEDDRLAIWLASVTAFTNFIFTLVGVWLVEKVGRRKLTFGSLAGTTVALIILALGFVL 395
Query: 362 SELHAP--TVSAIETSHYNNTCPDFRAAVNPGRWTCMTCLKASPSCGFCA-ASPNKLLPG 418
S +P T I S N TC + C C+ P CGFC + + ++
Sbjct: 396 SAQVSPRITFKPIAPSGQNATCTRYS--------YCNECM-LDPDCGFCXKMNKSTVIDS 446
Query: 419 ACL-----ISDDTTKDMCGNDHR------AWYTRGCPSKFGWAALVGLALYIITFSPGMG 467
+C+ +++ C N+ + W CP+ + W AL+GL LY++ F+PGMG
Sbjct: 447 SCVPVNKASTNEAAWGRCENETKFKTEDIFWAYNFCPTPYSWTALLGLILYLVFFAPGMG 506
Query: 468 TVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVV 527
+PW +NSEIYPL R +S WI N++VS +FL + + F ++ A V
Sbjct: 507 PMPWTVNSEIYPLWARSTGNACSSGINWIFNVLVSLTFLHTAEYLTYYGAFFLYAGFAAV 566
Query: 528 AIFFVIVFVPETKGVPMEEVEKMLEQR 554
+ F+ +PETKG +EE+E + + R
Sbjct: 567 GLLFIYGCLPETKGKKLEEIESLFDNR 593
>emb|CAG01178.1| unnamed protein product [Tetraodon nigroviridis]
Length = 614
Score = 369 bits (946), Expect = e-100
Identities = 211/572 (36%), Positives = 324/572 (55%), Gaps = 56/572 (9%)
Query: 25 YVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAIIGAA 84
YVL AFSA +GG LFGYDTGVISGA+L ++ K ++ QE ++S+ +A A + A
Sbjct: 21 YVLA-AFSA-MGGFLFGYDTGVISGAMLLLK---KELELSALWQELLISSTVAAAALSAL 75
Query: 85 VGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPLYI 144
+GG++N FGRRV I++A F +G ++++ AP +LL GR+ VG G+G+A M P+YI
Sbjct: 76 LGGFLNGLFGRRVCILLASFFFTVGGIVLSTAPGKEVLLAGRLIVGVGLGVACMTVPVYI 135
Query: 145 SEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGT-WRWMLGVAAAPAIIQIVLM 203
+EASP +RG LV++N+ ITGGQF + L++ AF+ WR+MLG++ PA +Q +
Sbjct: 136 AEASPPHLRGQLVTVNTLFITGGQFTASLVDGAFSYLQHDGWRYMLGLSVLPAALQFIGF 195
Query: 204 LSLPESPRWLYRKGREEESKSILKKIYAPEDVDAEIEALKESVESE-------------- 249
L LPESPRWL ++G ++++ +L +I +++D E +++K S++ E
Sbjct: 196 LFLPESPRWLIQRGLTQKARRVLSQIRGNQNIDEEYDSIKNSLDEEESGGGNAASGWLEI 255
Query: 250 ------IEESKTSKIS--------MMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYS 295
+E S++ + +L RR L G GL FQQ GINT+MYYS
Sbjct: 256 SRWVECMEMKSKSQMGDDYGDGPVIWRMLTYPPTRRALLVGCGLHMFQQVSGINTIMYYS 315
Query: 296 PTIVQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALL 355
TI+Q++G ++ A+ L+ +T+ N ++L ++ +++ GR+KL L S+ G LSL+LL
Sbjct: 316 ATILQMSGVRDDKLAIWLAGLTTLTNFLFTLLGVWLVERVGRRKLTLGSILGTCLSLSLL 375
Query: 356 TVAFRESELHAPTVSAIET--SHYNNTCPDFRAAVNPGRWTCMTCLKASPSCGFC-AASP 412
V F S H+P V+ T S N+TC ++ C C+ P CGFC +
Sbjct: 376 AVGFLMSAQHSPPVTLHPTHPSLANSTCSKYQL--------CEPCM-LDPECGFCYRENG 426
Query: 413 NKLLPGACLISDDTTKDM-----CGN-----DHRAWYTRGCPSKFGWAALVGLALYIITF 462
+ L +C+ D + + C N D W CP+ + W L+GL LY+ F
Sbjct: 427 SALFASSCVPVDKASTERAAWGRCSNTSLMRDQTFWAYNYCPTSYSWLVLLGLVLYLAAF 486
Query: 463 SPGMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFG 522
+PGMG +PW INSEIYPL R A+ W N++VS +FL L Q F ++
Sbjct: 487 APGMGPMPWTINSEIYPLWARSTGNACAAGVNWTFNILVSLTFLHLAQYFTYYGAFFLYS 546
Query: 523 ILAVVAIFFVIVFVPETKGVPMEEVEKMLEQR 554
+A++ FF+ +PETK +EE+E + E R
Sbjct: 547 SMALLGFFFIYGCLPETKARRLEEIEALFENR 578
>emb|CAA86519.2| Hypothetical protein M01F1.5 [Caenorhabditis elegans]
Length = 604
Score = 332 bits (851), Expect = 2e-89
Identities = 216/573 (37%), Positives = 311/573 (53%), Gaps = 33/573 (5%)
Query: 25 YVLRLAFSAGIGGLLFGYDTGVISGALLYIRD--DFKAVDRKTWLQEAIVSTAIAGAIIG 82
+V LAFSA IGG LFGYDTG++S A+LY+ + K +D W QE IVS A IG
Sbjct: 27 FVYMLAFSAVIGGFLFGYDTGIVSAAMLYVPNASGIKPLD-SVW-QEIIVSITPGVAAIG 84
Query: 83 AAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPL 142
+ G +D GR+ II A F IG++I AAA ILL GR+ +G +G ASM P+
Sbjct: 85 SLCSGPGSDFLGRKKIIIGASVTFTIGAIICAAAWTKIILLIGRILLGLAIGFASMIVPI 144
Query: 143 YISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPGT---WRWMLGVAAAPAIIQ 199
Y+SEASP+ +RG LV+ +IT G ++ +I AF+ WR M AA PAIIQ
Sbjct: 145 YVSEASPSHIRGKLVTGFQLMITVGLVIANIIGGAFSYVDPDQVGWRLMFAFAAVPAIIQ 204
Query: 200 IVLMLSLPESPRWLYRKGREEESKSILKKIYA--PEDVDAEIEALKESVESEIE---ESK 254
V L LPESPRWLY GR E++ +L +IY E VD EI + S E E+ E
Sbjct: 205 FVCFLFLPESPRWLYEHGRTVEAREVLTRIYNGHTEWVDYEINEISFSYEEELRAKAEHA 264
Query: 255 TSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLS 314
+ +++ +LKT VR+ + G LQ FQQ GINTVMYY+ I++ AG N T + +S
Sbjct: 265 GNGPTIIRILKTPHVRKAMIIGSLLQMFQQLSGINTVMYYTGNIIRSAGVKDNHTTIWIS 324
Query: 315 LITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVAFRESELHAPTVSAIET 374
+ TS +N G+ + I +++ GR+ L L+S+ GV+L L + V+F ++ ++ +
Sbjct: 325 VGTSAINFLGTFIPIALVERVGRRVLLLVSMIGVILFLIAMGVSFLL--INNDSLRTFDQ 382
Query: 375 SHYN-NTCPDFRAAVNPGRWTCMTCLKASPSCGFCAASPNKLLPGACL------ISDDTT 427
+Y N P + A +++ +CGFC + K G CL SD +
Sbjct: 383 QNYTLNYNPSVKHAAKCLKYSNCDFCVTDENCGFCESKVAK--KGYCLPFPSDDSSDYSA 440
Query: 428 KDMC--------GNDHRAWYTRGCPSKFGWAALVGLALYIITFSPGMGTVPWVINSEIYP 479
+C G D W C +KF ++ + Y+++FS G +PWV+N+E YP
Sbjct: 441 TGICEFSNLTNNGTDFE-WEDTYCHTKFTVLPIIIMVFYLLSFSAGYAPLPWVLNAEFYP 499
Query: 480 LRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPET 539
L R +++ WI NLIVS +FLSL+Q TF ++ +VA+ FV FVPET
Sbjct: 500 LWARSTAVSVSTACNWIFNLIVSLTFLSLSQAATKYGTFFIYCGCTMVALVFVFFFVPET 559
Query: 540 KGVPMEEVEKMLEQRSLQFKFWQKRDSGSE-KH 571
KG ++EVE + + + K + D E KH
Sbjct: 560 KGYSIDEVEMLFMTKEERRKAQKVLDESKEGKH 592
>emb|CAE64945.1| Hypothetical protein CBG09776 [Caenorhabditis briggsae]
Length = 613
Score = 330 bits (845), Expect = 1e-88
Identities = 218/588 (37%), Positives = 311/588 (52%), Gaps = 48/588 (8%)
Query: 25 YVLRLAFSAGIGGLLFGYDTGVISGALLYIRD--DFKAVDRKTWLQEAIVSTAIAGAIIG 82
+V LAFSA IGG LFGYDTG++S A+LY+ D K +D W QE IVS A IG
Sbjct: 21 FVYMLAFSAVIGGFLFGYDTGIVSAAMLYVPDAAGIKPLD-SVW-QEIIVSVTPGVAAIG 78
Query: 83 AAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPL 142
+ G +D GR+ II A F +G+VI AAA +LL GR+ +G +G ASM P+
Sbjct: 79 SLCSGPGSDFLGRKKIIIGASVTFAVGAVICAAAWTKIVLLIGRILLGAAIGFASMIVPI 138
Query: 143 YISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNA-PGTWR--------------- 186
Y+SEASP+ +RG LV+ +IT G ++ ++ AF+ P WR
Sbjct: 139 YVSEASPSHIRGKLVTGFQLMITVGLVIANIVGGAFSYVDPFGWRSFLNTSNPSKNLPFR 198
Query: 187 WMLGVAAAPAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYA--PEDVDAEIEALKE 244
M AA PAIIQ V L LPESPRWLY GR E++ +L +IY E VD EI +
Sbjct: 199 LMFAFAAVPAIIQFVCFLFLPESPRWLYEHGRTVEAREVLTRIYNGHTEWVDYEINEISF 258
Query: 245 SVESEIE---ESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQL 301
S E E+ E + +++ +LKT VR+ L G LQ FQQ GINTVMYY+ I++
Sbjct: 259 SYEEELRAKAEHAGNGPTIIRILKTPHVRKALIIGSLLQMFQQLSGINTVMYYTGNIIRS 318
Query: 302 AGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVAF-- 359
AG + T + +S+ TS +N G+ + I +++ GR+ L L+S+ GV+L L + V+F
Sbjct: 319 AGVKNKHTTIWISVGTSAINFIGTFIPIALVERLGRRVLLLVSMIGVILFLIAMGVSFLL 378
Query: 360 --RESELHAPTVSAIETSHYNNTCPDFRAAVNPGRWTCMTCLKASPSCGFCAASPNKLLP 417
+S L P T ++N P + A+ +++ CGFC +K
Sbjct: 379 INNDSSLTYPQNEYAATPNFN---PSVKDAIKCMKYSNCDFCVTDEKCGFCENKASKT-- 433
Query: 418 GACLISDDTTK------------DMCGNDHR-AWYTRGCPSKFGWAALVGLALYIITFSP 464
G CL D+ ++ GN W C +KF ++ + Y+++FS
Sbjct: 434 GYCLPFPDSDSADFSATGICQFSNLTGNGKTYEWEDSYCHTKFTVLPIIIMVFYLLSFSA 493
Query: 465 GMGTVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGIL 524
G +PWV+N+E YPL R I++ WI NLIVS +FLSL+Q TF ++
Sbjct: 494 GYAPLPWVLNAEFYPLWARSTAVSISTAFNWIFNLIVSLTFLSLSQAATKYGTFFIYCGC 553
Query: 525 AVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQKRDSGSE-KH 571
+VA+ FV FVPETKG ++EVE + + + K + D E KH
Sbjct: 554 TIVALIFVFFFVPETKGYSIDEVEMLFMTKEEREKTQKVLDESKEGKH 601
>emb|CAA16400.1| Hypothetical protein Y51A2D.4 [Caenorhabditis elegans]
gi|17565976|ref|NP_507623.1| general substrate
transporter family member (66.7 kD) (5T3)
[Caenorhabditis elegans] gi|7510110|pir||T27072
hypothetical protein Y51A2D.4 - Caenorhabditis elegans
Length = 606
Score = 328 bits (841), Expect = 3e-88
Identities = 202/568 (35%), Positives = 315/568 (54%), Gaps = 33/568 (5%)
Query: 25 YVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIAGAIIGAA 84
+V LA ++ IGG LFGYDT V+S A+LY+ D T QE +VS + A +G+
Sbjct: 24 FVYILAAASVIGGFLFGYDTSVVSAAMLYMPDAPGLKPMDTVWQEVLVSISPGMAAVGSL 83
Query: 85 VGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMASMASPLYI 144
+ G +D GRR I+ A +F IG+++ AA+ N +LL GRV +G +G ASM P+Y+
Sbjct: 84 MSGTSSDYIGRRKVILGASAIFTIGALVCAASVNKIMLLVGRVLLGIAIGFASMIVPVYL 143
Query: 145 SEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTNAPG---TWRWMLGVAAAPAIIQIV 201
E +PT VRG LV+ + +I+ GQ ++ + AF+ WR M AA P+IIQ V
Sbjct: 144 GETAPTHVRGMLVAAFALMISFGQVVANITGGAFSYIDPYNVGWRLMFAFAAVPSIIQFV 203
Query: 202 LMLSLPESPRWLYRKGREEESKSILKKIYA-----PEDVDAEIEALKESVESEIEESKTS 256
+ LPE+PRWLY G E E++ +L+K+Y E AEI A E E E++ S
Sbjct: 204 CFMFLPETPRWLYENGFETETREVLEKVYNGDKEWVEYEMAEIIAFNEDQAKENEKAHAS 263
Query: 257 KISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLI 316
+ +LKT V + + G LQ FQQ GINT++YY+ I++ +G ++N T + +S++
Sbjct: 264 GPVIWRILKTPHVLKACFIGSMLQAFQQLAGINTILYYTADIIRSSGISNNHTTIWISVL 323
Query: 317 TSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVAF-RESELHAPTVSAIE-T 374
S N G + + I+K GR+ + L S VVLSL + VAF + A T+ A +
Sbjct: 324 LSLCNFIGPFVPMSLIEKVGRRIIFLFSCGLVVLSLVFIGVAFLLVNHDSAATLPANQYG 383
Query: 375 SHYNNTCPDFRAAVNPGRWTCMTCLKASPSCGFCAASPNKLLPGACLISDDTTKDM---- 430
S++N++ PD + + C C+ + +CGFC + K G CL + ++
Sbjct: 384 SNFNSSYPDAKGCM--AYSNCDYCV-TTDACGFCHDANTK--QGYCLPAGFDNPEVYSST 438
Query: 431 --CGNDHRA------WYTRGCPSKFGWAALVGLALYIITFSPGMGTVPWVINSEIYPLRY 482
C N + + W C +K+ ++ +Y++TFS G ++PWV+NSE YP+
Sbjct: 439 GSCTNSNGSIANNFKWEKYYCDTKYTLLPIIACGVYLLTFSSGFTSLPWVLNSEFYPMWA 498
Query: 483 RGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGV 542
R C I++T+ W+ NLI++ ++LSLTQ IG F ++ L V+A F++ VPETKG
Sbjct: 499 RSTCVAISTTSNWVFNLIIALTYLSLTQVIGKYGAFWLYAGLTVIAFIFILFLVPETKGY 558
Query: 543 PMEEVEKMLEQRSLQFKFWQKRDSGSEK 570
+EEVE + + Q+R++ S +
Sbjct: 559 SIEEVEMLFMNKK------QRREAESRR 580
>emb|CAE67430.1| Hypothetical protein CBG12920 [Caenorhabditis briggsae]
Length = 615
Score = 327 bits (838), Expect = 6e-88
Identities = 208/584 (35%), Positives = 310/584 (52%), Gaps = 38/584 (6%)
Query: 22 KNP----YVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIA 77
KNP +V L +A IGG LFGYDT V+S A+LY+ + T +E IVS
Sbjct: 18 KNPKLGFFVYLLGSAAIIGGFLFGYDTSVVSAAMLYVPEAPGLKPMGTVWKEVIVSITPG 77
Query: 78 GAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMAS 137
A +GA G +DR+GR+ II + +F+ G++I A A ++L GR+F+G G+G AS
Sbjct: 78 MAAVGAWFSGAGSDRYGRKPIIIGSTIIFIAGAIICAVAWTKIVMLIGRIFLGVGIGFAS 137
Query: 138 MASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTN-APGT--WRWMLGVAAA 194
M P+Y+ EASPT VRG LVS + +I+ GQ ++ ++ F+ P T WR M A
Sbjct: 138 MVVPVYLGEASPTHVRGTLVSAFAMMISFGQVVANVMGGIFSYWDPYTIGWRLMFAFAGI 197
Query: 195 PAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAP-----EDVDAEIEALKESVESE 249
PA+IQ V + LPE+PRWLY G+ E +K +L+KIY+ E AEIE E + +
Sbjct: 198 PALIQFVCFIFLPETPRWLYENGQTERAKQVLEKIYSGDAEWIEYELAEIETYAEERKKQ 257
Query: 250 IEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRT 309
+EE K S + +LKT V + + G LQ FQQ GINT++YY+ I++ AG + T
Sbjct: 258 MEEEKKSGPVIWRILKTPHVLKACFIGSMLQAFQQLAGINTILYYTADIIRSAGIENYHT 317
Query: 310 ALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVAF----RESELH 365
+ +S+I S N G + + I+K GR+KL L S GVV+SL L+ V+F +S +
Sbjct: 318 IIWISVILSICNLIGPFIPMTLIEKLGRRKLFLFSCAGVVISLVLIGVSFLLVGNDSAPN 377
Query: 366 APTVSAIETSHYNNTCPDFRAAVNPGRWTCMTCLKASPSCGFCAASPNK---LLP----- 417
S Y+ P+ + C +C+ S CGFC S K LP
Sbjct: 378 FEMSSYSLAGSYDPLHPEAESCRILS--NCDSCV-TSEHCGFCEDSETKTGFCLPVNHND 434
Query: 418 -------GACLISDDTTKDMCGN-DHRAWYTRGCPSKFGWAALVGLALYIITFSPGMGTV 469
G C D + N W C + + +V + +Y++TFS G ++
Sbjct: 435 PTLFSSTGLCTSGVDKSNSSFPNATSFTWQKHHCTTSYTILPIVMMGVYLLTFSSGFTSL 494
Query: 470 PWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVVAI 529
PWV+NSE YP+ R C I++ + W+ NL+VS ++LSLT I F ++ I ++A
Sbjct: 495 PWVLNSEFYPMWARSTCVSISTLSNWVFNLLVSLTYLSLTHAITKYGAFWLYAIFTIIAF 554
Query: 530 FFVIVFVPETKGVPMEEVEKML---EQRSLQFKFWQKRDSGSEK 570
F+ VPET G ++EVE + QR++ + Q + ++K
Sbjct: 555 IFIYFLVPETTGYSIDEVEMLFMNKRQRNIAMQIRQAKLEANDK 598
>emb|CAA16405.1| Hypothetical protein Y51A2D.5 [Caenorhabditis elegans]
gi|17565978|ref|NP_507624.1| general substrate
transporter family member (67.9 kD) (5T9)
[Caenorhabditis elegans] gi|7510111|pir||T27077
hypothetical protein Y51A2D.5 - Caenorhabditis elegans
Length = 613
Score = 327 bits (837), Expect = 8e-88
Identities = 207/587 (35%), Positives = 314/587 (53%), Gaps = 44/587 (7%)
Query: 22 KNP----YVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKAVDRKTWLQEAIVSTAIA 77
KNP +V L +A IGG LFGYDT V+S A+LY+ + T +E IVS
Sbjct: 18 KNPKLGFFVYLLGSAAIIGGFLFGYDTSVVSAAMLYVPEAPGLKPMGTVWKEVIVSITPG 77
Query: 78 GAIIGAAVGGWINDRFGRRVAIIIADTLFLIGSVIMAAAPNPAILLFGRVFVGFGVGMAS 137
A +GA G +DR+GR+ II + +F+ G+VI A A ++L GR+F+G G+G AS
Sbjct: 78 MAAVGAWFSGAGSDRYGRKPIIIGSTLIFVCGAVICAVAWTKIVMLIGRIFLGVGIGFAS 137
Query: 138 MASPLYISEASPTRVRGALVSLNSFLITGGQFLSYLINLAFTN-APGT--WRWMLGVAAA 194
M P+Y+ EASPT VRG LVS + +I+ GQ ++ ++ F+ P T WR M A
Sbjct: 138 MVVPVYLGEASPTHVRGTLVSAFAMMISFGQVVANIMGGVFSYWEPYTIGWRLMFAFAGI 197
Query: 195 PAIIQIVLMLSLPESPRWLYRKGREEESKSILKKIYAP-----EDVDAEIEALKESVESE 249
PA+IQ V + LPE+PRWLY G E+++ +L+KIY E AEI+ E + +
Sbjct: 198 PALIQFVCFIFLPETPRWLYENGHTEQAEQVLEKIYGGNTEWIEYELAEIKTYAEERQKQ 257
Query: 250 IEESKTSKISMMTLLKTTTVRRGLYAGMGLQFFQQFVGINTVMYYSPTIVQLAGFASNRT 309
+EE K S + +LKT V + + G LQ FQQ GINT++YY+ I++ AG + T
Sbjct: 258 MEEEKKSGPVIWRILKTPHVLKACFIGSMLQAFQQLAGINTILYYTADIIRSAGIENYHT 317
Query: 310 ALLLSLITSGLNAFGSILSIYFIDKTGRKKLALISLCGVVLSLALLTVAF----RESELH 365
+ +S+I S N G ++FI+K GR+KL L S GVV+SL L+ V+F +S +
Sbjct: 318 IIWISVILSICNLIGPFAPMFFIEKLGRRKLFLFSCAGVVVSLVLIGVSFLLVGNDSAPN 377
Query: 366 APTVSAIETSHYNNTCPDFRAAVNPGRWTCMTCLKASPSCGFCAASPNKLLPGACLISDD 425
+ + +Y + + +C+T S CGFC S + G CL D
Sbjct: 378 FDRSAYLLAGNYQSNGEAESCLMLSNCDSCVT----SEHCGFCEDSETR--TGFCLPVDH 431
Query: 426 ------TTKDMCGN------------DHRAWYTRGCPSKFGWAALVGLALYIITFSPGMG 467
++ +C N W C + + +V + +Y++TFS G
Sbjct: 432 NDVTLYSSTGLCTNGLDKSNSSFPNATSYVWQKHHCTTSYTILPIVMMGVYLLTFSCGFT 491
Query: 468 TVPWVINSEIYPLRYRGVCGGIASTTVWISNLIVSQSFLSLTQTIGTAWTFMMFGILAVV 527
++PWV+NSE YP+ R C I++ + W+ NLI++ ++LSLT I F ++ I ++
Sbjct: 492 SLPWVLNSEFYPMWARSTCVSISTLSNWVFNLIIALTYLSLTHAITKYGAFWLYAIFTII 551
Query: 528 AIFFVIVFVPETKGVPMEEVEKML---EQRSLQFKFWQ-KRDSGSEK 570
A F+ VPET G ++EVE + QR++ + Q K D+ S+K
Sbjct: 552 AFIFIYFLVPETTGYSIDEVEMLFMNKRQRNIAMQARQAKLDAASDK 598
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.323 0.138 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 937,233,359
Number of Sequences: 2540612
Number of extensions: 38512720
Number of successful extensions: 137282
Number of sequences better than 10.0: 6509
Number of HSP's better than 10.0 without gapping: 3639
Number of HSP's successfully gapped in prelim test: 2870
Number of HSP's that attempted gapping in prelim test: 120840
Number of HSP's gapped (non-prelim): 11649
length of query: 571
length of database: 863,360,394
effective HSP length: 133
effective length of query: 438
effective length of database: 525,458,998
effective search space: 230151041124
effective search space used: 230151041124
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0140.7


