

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0140.2
(123 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
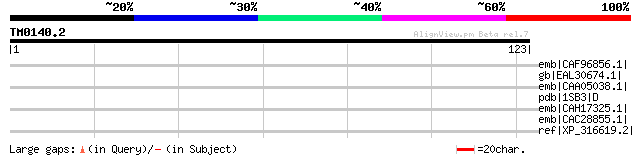
emb|CAF96856.1| unnamed protein product [Tetraodon nigroviridis] 34 0.93
gb|EAL30674.1| GA11551-PA [Drosophila pseudoobscura] 33 1.6
emb|CAA05038.1| 4-Hydroxybenzoyl-CoA reductase alpha-subunit [Th... 32 2.7
pdb|1SB3|D Chain D, Structure Of 4-Hydroxybenzoyl-Coa Reductase ... 32 2.7
emb|CAH17325.1| hypothetical protein [Legionella pneumophila str... 32 4.6
emb|CAC28855.1| conserved hypothetical protein [Neurospora crass... 32 4.6
ref|XP_316619.2| ENSANGP00000002585 [Anopheles gambiae str. PEST... 31 6.0
>emb|CAF96856.1| unnamed protein product [Tetraodon nigroviridis]
Length = 909
Score = 33.9 bits (76), Expect = 0.93
Identities = 17/47 (36%), Positives = 24/47 (50%), Gaps = 1/47 (2%)
Query: 52 CVIMIASTPPSQHHHSCKPTVNRAPGWFPEKRGVFPEP-PKTALHNV 97
C+ + + P Q H SC P RAP P K V P+P P+ H++
Sbjct: 187 CIKHMINVDPKQGHLSCSPLPTRAPSPTPRKDTVSPQPHPQPHSHSL 233
>gb|EAL30674.1| GA11551-PA [Drosophila pseudoobscura]
Length = 824
Score = 33.1 bits (74), Expect = 1.6
Identities = 20/62 (32%), Positives = 27/62 (43%), Gaps = 3/62 (4%)
Query: 48 PRTPCVIMIASTPPSQHHHSCKPTVNRAPGWFPEKRGVFPEPPKTALHNVTLLQHLRNTT 107
PR + TP + H SC PT R P RG P PP + + + Q LR +T
Sbjct: 649 PRAAQITASQLTPTTSHASSCTPTRMRGT---PRGRGELPTPPASTHSSASPYQILRAST 705
Query: 108 QS 109
+
Sbjct: 706 DT 707
>emb|CAA05038.1| 4-Hydroxybenzoyl-CoA reductase alpha-subunit [Thauera aromatica]
gi|13431532|sp|O33819|HCRA_THAAR 4-hydroxybenzoyl-CoA
reductase alpha subunit
Length = 769
Score = 32.3 bits (72), Expect = 2.7
Identities = 23/77 (29%), Positives = 32/77 (40%), Gaps = 5/77 (6%)
Query: 45 NLMPRTPCVIMIASTPPSQHHHSCKPTVNRAPGWFPEKRGVFPEPPKTALHNVTLLQHLR 104
N++P+ P V M A S C V A GW E++G P+ + L H
Sbjct: 391 NMLPQIPYVTMYAQRVMSYGVPECLEKVKAASGW-EERKGKLPKGRGLGI----ALSHFV 445
Query: 105 NTTQSDIHRTGSSDTTV 121
+ T + H TG TV
Sbjct: 446 SGTSTPKHWTGEPHATV 462
>pdb|1SB3|D Chain D, Structure Of 4-Hydroxybenzoyl-Coa Reductase From Thauera
Aromatica gi|58176705|pdb|1SB3|A Chain A, Structure Of
4-Hydroxybenzoyl-Coa Reductase From Thauera Aromatica
gi|58176617|pdb|1RM6|D Chain D, Structure Of
4-Hydroxybenzoyl-Coa Reductase From Thauera Aromatica
gi|58176614|pdb|1RM6|A Chain A, Structure Of
4-Hydroxybenzoyl-Coa Reductase From Thauera Aromatica
Length = 769
Score = 32.3 bits (72), Expect = 2.7
Identities = 23/77 (29%), Positives = 32/77 (40%), Gaps = 5/77 (6%)
Query: 45 NLMPRTPCVIMIASTPPSQHHHSCKPTVNRAPGWFPEKRGVFPEPPKTALHNVTLLQHLR 104
N++P+ P V M A S C V A GW E++G P+ + L H
Sbjct: 391 NMLPQIPYVTMYAQRVMSYGVPECLEKVKAASGW-EERKGKLPKGRGLGI----ALSHFV 445
Query: 105 NTTQSDIHRTGSSDTTV 121
+ T + H TG TV
Sbjct: 446 SGTSTPKHWTGEPHATV 462
>emb|CAH17325.1| hypothetical protein [Legionella pneumophila str. Lens]
gi|54292913|ref|YP_122300.1| hypothetical protein
plpl0006 [Legionella pneumophila str. Lens]
Length = 387
Score = 31.6 bits (70), Expect = 4.6
Identities = 25/84 (29%), Positives = 35/84 (40%), Gaps = 16/84 (19%)
Query: 16 YLASMSTNARTLSPHILHLYQPMLVCHLDNLMPRTPCVIMIASTPPS----QHHHSCKPT 71
Y+ + TNAR + LYQ L+ ++ C I+ PPS + H C
Sbjct: 263 YIQFIETNARPPGIGLNKLYQRKYSISLETIL----CCIVCGVEPPSMVENKSHFVC--- 315
Query: 72 VNRAPGWFPEKRGVFPEPPKTALH 95
G+FP K GV + K LH
Sbjct: 316 -----GYFPTKMGVIKKINKPELH 334
>emb|CAC28855.1| conserved hypothetical protein [Neurospora crassa]
gi|32404802|ref|XP_323014.1| hypothetical protein
[Neurospora crassa] gi|28923034|gb|EAA32252.1|
hypothetical protein [Neurospora crassa]
Length = 479
Score = 31.6 bits (70), Expect = 4.6
Identities = 24/95 (25%), Positives = 38/95 (39%), Gaps = 28/95 (29%)
Query: 37 PMLV-CHLDNLMPRTPCVIMIASTPPS----------QHHHSCKPTVNRAPGWFPEKRGV 85
P++V CH + +++ TPPS QHHHS + + + +F K+ V
Sbjct: 65 PVIVPCHSQMTLDDFAAALLVPVTPPSSPLPNNSNNPQHHHSHRSVPSGSGSYFSSKQNV 124
Query: 86 -----------------FPEPPKTALHNVTLLQHL 103
+PP T + NV L Q+L
Sbjct: 125 PGSSSSAHPPPPSQQQQQQQPPATQIANVVLAQNL 159
>ref|XP_316619.2| ENSANGP00000002585 [Anopheles gambiae str. PEST]
gi|55239345|gb|EAA11397.2| ENSANGP00000002585 [Anopheles
gambiae str. PEST]
Length = 880
Score = 31.2 bits (69), Expect = 6.0
Identities = 15/47 (31%), Positives = 20/47 (41%)
Query: 30 HILHLYQPMLVCHLDNLMPRTPCVIMIASTPPSQHHHSCKPTVNRAP 76
H++ LY P +V P P + PS HH+ TV R P
Sbjct: 211 HVMGLYLPAMVRSTPTPSPSPPAPATVGGHVPSAKHHALHQTVLREP 257
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.131 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 213,351,193
Number of Sequences: 2540612
Number of extensions: 8709483
Number of successful extensions: 18948
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 18944
Number of HSP's gapped (non-prelim): 8
length of query: 123
length of database: 863,360,394
effective HSP length: 99
effective length of query: 24
effective length of database: 611,839,806
effective search space: 14684155344
effective search space used: 14684155344
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0140.2


