

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0136.17
(83 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
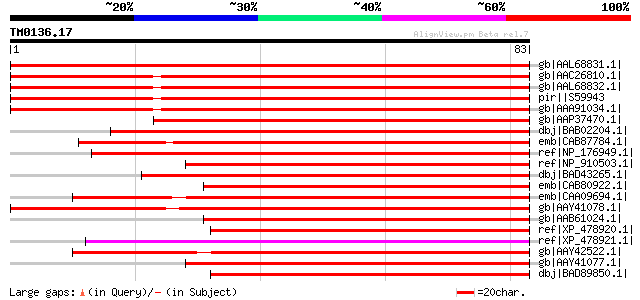
Score E
Sequences producing significant alignments: (bits) Value
gb|AAL68831.1| Enod8.2 [Medicago truncatula] 111 5e-24
gb|AAC26810.1| early nodule-specific protein [Medicago truncatul... 109 2e-23
gb|AAL68832.1| Enod8.1 [Medicago truncatula] 109 2e-23
pir||S59943 early nodulin 8 precursor - alfalfa gi|304037|gb|AAB... 104 5e-22
gb|AAA91034.1| nodulin 104 5e-22
gb|AAP37470.1| ENSP-like protein [Hevea brasiliensis] gi|5131578... 89 4e-17
dbj|BAB02204.1| nodulin-like protein protein [Arabidopsis thalia... 82 3e-15
emb|CAB87784.1| early nodule-specific protein-like [Arabidopsis ... 82 5e-15
ref|NP_176949.1| GDSL-motif lipase/hydrolase family protein [Ara... 82 5e-15
ref|NP_910503.1| putative lanatoside 15'-O-acetylesterase [Oryza... 77 1e-13
dbj|BAD43265.1| ENOD8-like protein [Arabidopsis thaliana] 77 1e-13
emb|CAB80922.1| putative acetyltransferase [Arabidopsis thaliana... 72 3e-12
emb|CAA09694.1| lanatoside 15'-O-acetylesterase [Digitalis lanata] 72 3e-12
gb|AAY41078.1| lanatoside 15-O-acetylesterase [Digitalis grandif... 72 3e-12
gb|AAB61024.1| similar to the GDSL family of lipolytic enzymes [... 72 3e-12
ref|XP_478920.1| putative early nodulin 8 precursor [Oryza sativ... 72 4e-12
ref|XP_478921.1| putative early nodulin 8 precursor [Oryza sativ... 71 6e-12
gb|AAY42522.1| lanatoside 15'-O-acetylesterase [Digitalis lanata] 71 6e-12
gb|AAY41077.1| lanatoside 15-O-acetylesterase [Digitalis lanata] 71 8e-12
dbj|BAD89850.1| hypothetical protein [Zea mays] 70 1e-11
>gb|AAL68831.1| Enod8.2 [Medicago truncatula]
Length = 385
Score = 111 bits (277), Expect = 5e-24
Identities = 52/83 (62%), Positives = 61/83 (72%)
Query: 1 MSLMEFHRHVSFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKS 60
M E RH+S VSL V++ P+FA + C+FPAIF+ GASN DTGGLAAAF P S
Sbjct: 1 MEKFELLRHMSLVSLIVLILCIITPPIFATRNCDFPAIFSFGASNVDTGGLAAAFQAPPS 60
Query: 61 PYGDTYFHRPAGRYSDGRIILDF 83
PYG+TYFHR GR+SDGRIILDF
Sbjct: 61 PYGETYFHRSTGRFSDGRIILDF 83
>gb|AAC26810.1| early nodule-specific protein [Medicago truncatula]
gi|25457244|pir||T52338 early nodule-specific protein
ENOD8 [imported] - barrel medic
Length = 381
Score = 109 bits (272), Expect = 2e-23
Identities = 52/83 (62%), Positives = 63/83 (75%), Gaps = 1/83 (1%)
Query: 1 MSLMEFHRHVSFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKS 60
M+ +E RH+ V+L V++ T P+FA K C+FPAIF+ GASN DTGGLAAAF P S
Sbjct: 1 MAKIELSRHIPLVTLIVLVLCIT-PPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPS 59
Query: 61 PYGDTYFHRPAGRYSDGRIILDF 83
PYG+TYFHR GR+SDGRIILDF
Sbjct: 60 PYGETYFHRSTGRFSDGRIILDF 82
>gb|AAL68832.1| Enod8.1 [Medicago truncatula]
Length = 381
Score = 109 bits (272), Expect = 2e-23
Identities = 52/83 (62%), Positives = 63/83 (75%), Gaps = 1/83 (1%)
Query: 1 MSLMEFHRHVSFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKS 60
M+ +E RH+ V+L V++ T P+FA K C+FPAIF+ GASN DTGGLAAAF P S
Sbjct: 1 MAKIELSRHIPLVTLIVLVLCIT-PPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPS 59
Query: 61 PYGDTYFHRPAGRYSDGRIILDF 83
PYG+TYFHR GR+SDGRIILDF
Sbjct: 60 PYGETYFHRSTGRFSDGRIILDF 82
>pir||S59943 early nodulin 8 precursor - alfalfa gi|304037|gb|AAB41547.1|
early nodulin [Medicago sativa]
Length = 381
Score = 104 bits (260), Expect = 5e-22
Identities = 50/83 (60%), Positives = 62/83 (74%), Gaps = 1/83 (1%)
Query: 1 MSLMEFHRHVSFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKS 60
M+ +E RH+ V+L V++ TT P+FA C+FPAIF+ GASN DTGGLAAAF P S
Sbjct: 1 MARIELSRHMPLVTLIVLVLCTT-PPIFASTHCDFPAIFSFGASNVDTGGLAAAFRAPPS 59
Query: 61 PYGDTYFHRPAGRYSDGRIILDF 83
PYG TYF+R GR+SDGRII+DF
Sbjct: 60 PYGQTYFNRSTGRFSDGRIIIDF 82
>gb|AAA91034.1| nodulin
Length = 381
Score = 104 bits (260), Expect = 5e-22
Identities = 50/83 (60%), Positives = 62/83 (74%), Gaps = 1/83 (1%)
Query: 1 MSLMEFHRHVSFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKS 60
M+ +E RH+ V+L V++ TT P+FA C+FPAIF+ GASN DTGGLAAAF P S
Sbjct: 1 MARIELSRHMPLVTLIVLVLCTT-PPIFASTHCDFPAIFSFGASNVDTGGLAAAFRAPPS 59
Query: 61 PYGDTYFHRPAGRYSDGRIILDF 83
PYG TYF+R GR+SDGRII+DF
Sbjct: 60 PYGQTYFNRSTGRFSDGRIIIDF 82
>gb|AAP37470.1| ENSP-like protein [Hevea brasiliensis]
gi|51315784|sp|Q7Y1X1|EST_HEVBR Esterase precursor
(Early nodule-specific protein homolog) (Latex allergen
Hev b 13)
Length = 391
Score = 88.6 bits (218), Expect = 4e-17
Identities = 39/60 (65%), Positives = 47/60 (78%)
Query: 24 LNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAGRYSDGRIILDF 83
L+ +A + C+FPAIFN G SNSDTGG AAAF P PYG+T+FHR GRYSDGR+I+DF
Sbjct: 21 LSLAYASETCDFPAIFNFGDSNSDTGGKAAAFYPLNPPYGETFFHRSTGRYSDGRLIIDF 80
>dbj|BAB02204.1| nodulin-like protein protein [Arabidopsis thaliana]
gi|20259934|gb|AAM13314.1| unknown protein [Arabidopsis
thaliana] gi|17064918|gb|AAL32613.1| Unknown protein
[Arabidopsis thaliana] gi|15231558|ref|NP_189274.1|
GDSL-motif lipase/hydrolase family protein [Arabidopsis
thaliana]
Length = 380
Score = 82.0 bits (201), Expect = 3e-15
Identities = 37/67 (55%), Positives = 48/67 (71%)
Query: 17 VILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAGRYSD 76
V+L + ++P CNFPAIFN G SNSDTGGL+A+F P G T+FH P+GR+SD
Sbjct: 11 VLLASCLIHPRACSPSCNFPAIFNFGDSNSDTGGLSASFGQAPYPNGQTFFHSPSGRFSD 70
Query: 77 GRIILDF 83
GR+I+DF
Sbjct: 71 GRLIIDF 77
>emb|CAB87784.1| early nodule-specific protein-like [Arabidopsis thaliana]
gi|26451820|dbj|BAC43003.1| putative early
nodule-specific protein [Arabidopsis thaliana]
gi|28950957|gb|AAO63402.1| At5g14450 [Arabidopsis
thaliana] gi|15241404|ref|NP_196949.1| GDSL-motif
lipase/hydrolase family protein [Arabidopsis thaliana]
gi|11357276|pir||T48618 early nodule-specific
protein-like - Arabidopsis thaliana
Length = 389
Score = 81.6 bits (200), Expect = 5e-15
Identities = 39/72 (54%), Positives = 49/72 (67%), Gaps = 1/72 (1%)
Query: 12 FVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPA 71
F L + LF T + V + C FPAI+N G SNSDTGG++AAF P + PYG +FHRP
Sbjct: 17 FSWLLLCLFAVTTS-VSVQPTCTFPAIYNFGDSNSDTGGISAAFEPIRDPYGQGFFHRPT 75
Query: 72 GRYSDGRIILDF 83
GR SDGR+ +DF
Sbjct: 76 GRDSDGRLTIDF 87
>ref|NP_176949.1| GDSL-motif lipase/hydrolase family protein [Arabidopsis thaliana]
gi|11072007|gb|AAG28886.1| F12A21.4 [Arabidopsis
thaliana]
Length = 372
Score = 81.6 bits (200), Expect = 5e-15
Identities = 37/70 (52%), Positives = 50/70 (70%)
Query: 14 SLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAGR 73
SLF + ++L+P +C+FPAIFN G SNSDTGGL+AAF P+G ++F PAGR
Sbjct: 7 SLFALSLLSSLSPSTHAHQCHFPAIFNFGDSNSDTGGLSAAFGQAGPPHGSSFFGSPAGR 66
Query: 74 YSDGRIILDF 83
Y DGR+++DF
Sbjct: 67 YCDGRLVIDF 76
>ref|NP_910503.1| putative lanatoside 15'-O-acetylesterase [Oryza sativa (japonica
cultivar-group)] gi|5295941|dbj|BAA81842.1| putative
lanatoside 15'-O-acetylesterase [Oryza sativa (japonica
cultivar-group)]
Length = 379
Score = 77.0 bits (188), Expect = 1e-13
Identities = 34/55 (61%), Positives = 42/55 (75%)
Query: 29 AKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAGRYSDGRIILDF 83
A +C FPA+FN G SNSDTGG AAF ++P+G TYF RPAGR SDGR+++DF
Sbjct: 23 ATAQCRFPAVFNFGDSNSDTGGFWAAFPAQQAPFGMTYFRRPAGRASDGRLVVDF 77
>dbj|BAD43265.1| ENOD8-like protein [Arabidopsis thaliana]
Length = 364
Score = 76.6 bits (187), Expect = 1e-13
Identities = 34/62 (54%), Positives = 46/62 (73%)
Query: 22 TTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAGRYSDGRIIL 81
++L+P +C+FPAIFN G SNSDTGGL+AAF P+G ++F PAGRY DGR+++
Sbjct: 7 SSLSPSTHAHQCHFPAIFNFGDSNSDTGGLSAAFGQAGPPHGSSFFGSPAGRYCDGRLVI 66
Query: 82 DF 83
DF
Sbjct: 67 DF 68
>emb|CAB80922.1| putative acetyltransferase [Arabidopsis thaliana]
gi|15234074|ref|NP_192022.1| acetylesterase, putative
[Arabidopsis thaliana] gi|25407083|pir||H85014 probable
acetyltransferase [imported] - Arabidopsis thaliana
Length = 382
Score = 72.4 bits (176), Expect = 3e-12
Identities = 33/52 (63%), Positives = 39/52 (74%)
Query: 32 RCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAGRYSDGRIILDF 83
+C+F AIFN G SNSDTGG AAF P+G TYF +PAGR SDGR+I+DF
Sbjct: 29 KCDFEAIFNFGDSNSDTGGFWAAFPAQSGPWGMTYFKKPAGRASDGRLIIDF 80
>emb|CAA09694.1| lanatoside 15'-O-acetylesterase [Digitalis lanata]
Length = 386
Score = 72.4 bits (176), Expect = 3e-12
Identities = 36/73 (49%), Positives = 48/73 (65%), Gaps = 2/73 (2%)
Query: 11 SFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRP 70
+F+ V+L ++ ++ +C+F AIFN G SNSDTGG AAF P G TYF RP
Sbjct: 11 NFMVYVVVLMEVSVRS--SESKCDFKAIFNFGDSNSDTGGFWAAFPAENPPNGMTYFKRP 68
Query: 71 AGRYSDGRIILDF 83
AGR +DGR+I+DF
Sbjct: 69 AGRVTDGRLIIDF 81
>gb|AAY41078.1| lanatoside 15-O-acetylesterase [Digitalis grandiflora]
Length = 386
Score = 72.4 bits (176), Expect = 3e-12
Identities = 38/83 (45%), Positives = 51/83 (60%), Gaps = 2/83 (2%)
Query: 1 MSLMEFHRHVSFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKS 60
M + F +F+ V+L ++ ++ +C+F AIFN G SNSDTGG AAF
Sbjct: 1 MKTIPFSTLRNFMVYVVVLMGVSVR--MSEAKCDFKAIFNFGDSNSDTGGFWAAFPAENP 58
Query: 61 PYGDTYFHRPAGRYSDGRIILDF 83
P G TYF RPAGR +DGR+I+DF
Sbjct: 59 PNGMTYFKRPAGRAADGRLIIDF 81
>gb|AAB61024.1| similar to the GDSL family of lipolytic enzymes [Arabidopsis
thaliana] gi|7485222|pir||T01727 hypothetical protein
A_IG002N01.17 - Arabidopsis thaliana
Length = 367
Score = 72.4 bits (176), Expect = 3e-12
Identities = 33/52 (63%), Positives = 39/52 (74%)
Query: 32 RCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAGRYSDGRIILDF 83
+C+F AIFN G SNSDTGG AAF P+G TYF +PAGR SDGR+I+DF
Sbjct: 29 KCDFEAIFNFGDSNSDTGGFWAAFPAQSGPWGMTYFKKPAGRASDGRLIIDF 80
>ref|XP_478920.1| putative early nodulin 8 precursor [Oryza sativa (japonica
cultivar-group)] gi|33147015|dbj|BAC80099.1| putative
early nodulin 8 precursor [Oryza sativa (japonica
cultivar-group)]
Length = 405
Score = 72.0 bits (175), Expect = 4e-12
Identities = 32/51 (62%), Positives = 38/51 (73%)
Query: 33 CNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAGRYSDGRIILDF 83
C+FPAIFN G SNSDTGGL+A P+G TYF PAGR+SDGR+ +DF
Sbjct: 45 CDFPAIFNFGDSNSDTGGLSALIAVVPPPFGRTYFGMPAGRFSDGRLTIDF 95
>ref|XP_478921.1| putative early nodulin 8 precursor [Oryza sativa (japonica
cultivar-group)] gi|33147016|dbj|BAC80100.1| putative
early nodulin 8 precursor [Oryza sativa (japonica
cultivar-group)]
Length = 391
Score = 71.2 bits (173), Expect = 6e-12
Identities = 34/71 (47%), Positives = 43/71 (59%)
Query: 13 VSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAG 72
V+ +L + V A C FPA+FN G SNSDTGGL+A F P G T+F P G
Sbjct: 12 VATVAVLVLVSQVSVAAGADCRFPAVFNFGDSNSDTGGLSATFGAAPPPNGRTFFGMPVG 71
Query: 73 RYSDGRIILDF 83
RY DGR+++DF
Sbjct: 72 RYCDGRLVIDF 82
>gb|AAY42522.1| lanatoside 15'-O-acetylesterase [Digitalis lanata]
Length = 386
Score = 71.2 bits (173), Expect = 6e-12
Identities = 38/73 (52%), Positives = 46/73 (62%), Gaps = 2/73 (2%)
Query: 11 SFVSLFVILFTTTLNPVFAKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRP 70
+F+ V+L ++ AK C F AIFN G SNSDTGG AAF P G TYF RP
Sbjct: 11 NFMVYVVVLMEVSVRSSEAK--CYFKAIFNFGDSNSDTGGFWAAFPAENPPNGMTYFKRP 68
Query: 71 AGRYSDGRIILDF 83
AGR +DGR+I+DF
Sbjct: 69 AGRVTDGRLIIDF 81
>gb|AAY41077.1| lanatoside 15-O-acetylesterase [Digitalis lanata]
Length = 386
Score = 70.9 bits (172), Expect = 8e-12
Identities = 33/55 (60%), Positives = 40/55 (72%)
Query: 29 AKKRCNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAGRYSDGRIILDF 83
++ +C+F AIFN G SNSDTGG AAF P G TYF RPAGR +DGR+I+DF
Sbjct: 27 SEAKCDFNAIFNFGDSNSDTGGFWAAFPAENPPNGMTYFKRPAGRVTDGRLIIDF 81
>dbj|BAD89850.1| hypothetical protein [Zea mays]
Length = 394
Score = 70.1 bits (170), Expect = 1e-11
Identities = 30/51 (58%), Positives = 38/51 (73%)
Query: 33 CNFPAIFNLGASNSDTGGLAAAFLPPKSPYGDTYFHRPAGRYSDGRIILDF 83
C+FPA+FN G SNSDTGGL++ F P G T+F PAGRY DGR+++DF
Sbjct: 36 CHFPAVFNFGDSNSDTGGLSSLFGAAPPPNGRTFFGMPAGRYCDGRLVIDF 86
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.328 0.143 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 150,915,302
Number of Sequences: 2540612
Number of extensions: 5682838
Number of successful extensions: 12294
Number of sequences better than 10.0: 214
Number of HSP's better than 10.0 without gapping: 150
Number of HSP's successfully gapped in prelim test: 64
Number of HSP's that attempted gapping in prelim test: 12052
Number of HSP's gapped (non-prelim): 220
length of query: 83
length of database: 863,360,394
effective HSP length: 59
effective length of query: 24
effective length of database: 713,464,286
effective search space: 17123142864
effective search space used: 17123142864
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0136.17


