

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0133.3
(700 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
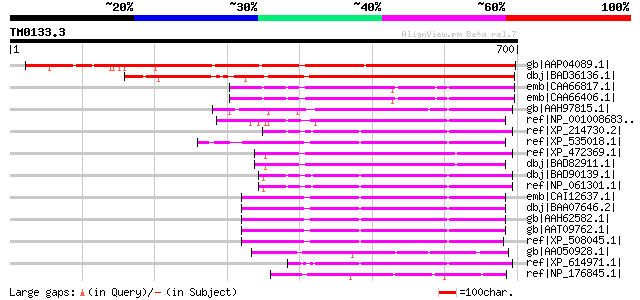
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP04089.1| unknown protein [Arabidopsis thaliana] gi|2897372... 655 0.0
dbj|BAD36136.1| putative SNM1 [Oryza sativa (japonica cultivar-g... 627 e-178
emb|CAA66817.1| hypothetical protein [Arabidopsis thaliana] gi|1... 228 4e-58
emb|CAA66406.1| orf12 [Arabidopsis thaliana] 228 4e-58
gb|AAH97815.1| Unknown (protein for IMAGE:3377284) [Xenopus laevis] 220 1e-55
ref|NP_001008683.1| SNM1A protein [Gallus gallus] gi|47156206|gb... 218 4e-55
ref|XP_214730.2| PREDICTED: similar to SNM1 protein [Rattus norv... 214 1e-53
ref|XP_535018.1| PREDICTED: similar to DNA-crosslink repair gene... 212 4e-53
ref|XP_472369.1| OSJNBb0012E08.8 [Oryza sativa (japonica cultiva... 212 4e-53
dbj|BAD82911.1| Snm1 [Oryza sativa (japonica cultivar-group)] 211 6e-53
dbj|BAD90139.1| mKIAA0086 protein [Mus musculus] 209 2e-52
ref|NP_061301.1| DNA cross-link repair 1A, PSO2 homolog [Mus mus... 208 5e-52
emb|CAI12637.1| DNA cross-link repair 1A (PSO2 homolog, S. cerev... 206 3e-51
dbj|BAA07646.2| KIAA0086 [Homo sapiens] 206 3e-51
gb|AAH62582.1| DNA-crosslink repair gene SNM1 [Homo sapiens] gi|... 206 3e-51
gb|AAT09762.1| DNA cross-link repair 1A (PSO2 homolog, S. cerevi... 206 3e-51
ref|XP_508045.1| PREDICTED: similar to DNA-crosslink repair gene... 202 4e-50
gb|AAO50928.1| similar to Arabidopsis thaliana (Mouse-ear cress)... 196 2e-48
ref|XP_614971.1| PREDICTED: similar to DNA-crosslink repair gene... 189 2e-46
ref|NP_176845.1| ATP dependent DNA ligase family protein [Arabid... 181 6e-44
>gb|AAP04089.1| unknown protein [Arabidopsis thaliana] gi|28973723|gb|AAO64178.1|
unknown protein [Arabidopsis thaliana]
gi|3386625|gb|AAC28555.1| hypothetical protein
[Arabidopsis thaliana] gi|20197051|gb|AAM14896.1|
hypothetical protein [Arabidopsis thaliana]
gi|7485634|pir||T02477 hypothetical protein At2g45700
[imported] - Arabidopsis thaliana
gi|15225548|ref|NP_182094.1| sterile alpha motif (SAM)
domain-containing protein [Arabidopsis thaliana]
Length = 723
Score = 655 bits (1689), Expect = 0.0
Identities = 372/731 (50%), Positives = 464/731 (62%), Gaps = 71/731 (9%)
Query: 22 DDDEFEIPLTQTATIHRPLKKHKLETTAFGGS----------GKENVPPNASSSTIHDDG 71
DDD+F+IP + +I +PL + GKENV P S
Sbjct: 8 DDDDFQIPPSSQLSIRKPLHPTNANNISHRPPNKKPRLCRYPGKENVTPPPSPDPDLFCS 67
Query: 72 YEIENCSLDFIPSTIDSVGTALTDSSSSSSSASVSAFVISSPVKVKMKGTAYFGNSIESK 131
+C LD IPS++D +L D + SS +K+ Y NS+E++
Sbjct: 68 SSTPHCILDCIPSSVDC---SLGDFNGPISSLGEEDKEDKDDC-IKVNREGYLCNSMEAR 123
Query: 132 LVVSRA-----NAINN-----AEAGSEL----NLSDELDG-----------SVRCPLCEV 166
L+ SR + I+ E+ SEL NL E +G S++CPLC +
Sbjct: 124 LLKSRICLGFDSGIHEDDEGFVESNSELDVLINLCSESEGRSGEFSLGKDDSIQCPLCSM 183
Query: 167 DISNLTEEQRHLHTNDCLDRGDDVAVRDDNVDA-------------------QFAPKVSR 207
DIS+L+EEQR +H+N CLD+ + D++ Q +S
Sbjct: 184 DISSLSEEQRQVHSNTCLDKSYNQPSEQDSLRKCENLSSLIKESIDDPVQLPQLVTDLSP 243
Query: 208 VVDWLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHALCEFRKGI 267
V+ WLR LGL KYEDVF+REE+DWDTLQ LTEEDLLS+G+T+LGPRKKIV+AL R
Sbjct: 244 VLKWLRSLGLAKYEDVFIREEIDWDTLQSLTEEDLLSIGITSLGPRKKIVNALSGVRDPF 303
Query: 268 AASNEKPEDALAEPRRNRNQNVKMQPHRPERKVDGTSKPAANKKITEYFPGFATNGKKVS 327
A+S E + + + + + RK KP ANK ITE+FPG AT G K+
Sbjct: 304 ASSAEVQAQSHCT---SGHVTERQRDKSTTRKASEPKKPTANKLITEFFPGQATEGTKIR 360
Query: 328 TAPEEQQEMKSSGSVSGRNHKGKNSSTNRKLRDVPKWCSIEGTPFRVDAFKYLRGDCSHW 387
TAP+ E S S S R + + N K + +P W I GTPFRVDAFKYL DC HW
Sbjct: 361 TAPKPVAEKSPSDSSSRR--AVRRNGNNGKSKVIPHWNCIPGTPFRVDAFKYLTRDCCHW 418
Query: 388 FLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQKVEI 447
FLTHFHLD + L T G + A+LVNM IGIP+++L +L L QKV I
Sbjct: 419 FLTHFHLDHYQGL------TKSFSHGKIYCSLVTAKLVNMKIGIPWERLQVLDLGQKVNI 472
Query: 448 AGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMCPVHTLILDTT 507
+GIDVTC DANHCPGSI+ILF+P NGKAVLHTGDFR+S+E++ L + +LILDTT
Sbjct: 473 SGIDVTCFDANHCPGSIMILFEPANGKAVLHTGDFRYSEEMS--NWLIGSHISSLILDTT 530
Query: 508 YCNPQYDFPKQEAVIQFVIDAIQAEAFNPGTLFLIGSYTIGKERLFLEVARSLRRKVYVN 567
YCNPQYDFPKQEAVIQFV++AIQAEAFNP TLFLIGSYTIGKERLFLEVAR LR K+Y+N
Sbjct: 531 YCNPQYDFPKQEAVIQFVVEAIQAEAFNPKTLFLIGSYTIGKERLFLEVARVLREKIYIN 590
Query: 568 AAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQYASRFSLIVAF 627
AKL+LL+CL F ++D+QWFT+ E ES+IHV P+WTLAS KRLKHV+++Y +R+SLIVAF
Sbjct: 591 PAKLKLLECLGFSKDDIQWFTVKEEESHIHVVPLWTLASFKRLKHVANRYTNRYSLIVAF 650
Query: 628 SPTGWTFGKGKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVNNDG 687
SPTGWT GK KKKSPGRR Q GTIIRYEVPYSEH SFTELK+FV+ +SP IIPSVNNDG
Sbjct: 651 SPTGWTSGKTKKKSPGRRLQQGTIIRYEVPYSEHSSFTELKEFVQKVSPEVIIPSVNNDG 710
Query: 688 PESANAMISLM 698
P+SA AM+SL+
Sbjct: 711 PDSAAAMVSLL 721
>dbj|BAD36136.1| putative SNM1 [Oryza sativa (japonica cultivar-group)]
gi|51091343|dbj|BAD36078.1| putative SNM1 [Oryza sativa
(japonica cultivar-group)]
Length = 1024
Score = 627 bits (1618), Expect = e-178
Identities = 319/549 (58%), Positives = 400/549 (72%), Gaps = 40/549 (7%)
Query: 159 VRCPLCEVDISNLTEEQRHLHTNDCLDRGDDVAVRDDNVDAQFAP------KVSRVVDWL 212
V+CPLC +IS+L+EE R +HTN CLD D ++ N D Q P + RV++WL
Sbjct: 500 VQCPLCGSNISDLSEELRLVHTNSCLD--GDKPAKEPNSDNQNEPCGESNVEKRRVMEWL 557
Query: 213 RDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHALCEFRKGIAASNE 272
R+LGL KYE++F++EEVDW+TLQWLTEEDLL MG+T+LGPRKKI HALCE RK +N+
Sbjct: 558 RNLGLSKYEEIFIKEEVDWETLQWLTEEDLLGMGITSLGPRKKIAHALCELRKKNNDAND 617
Query: 273 KPEDALAEPRRNRNQNVKMQPHRPERKVDGTSKPAANKKITEYFPGFATNGK-----KVS 327
D L +N K + + ++G NK ITEYF +++ + KV+
Sbjct: 618 LAADML------NLENTK----KAKIPMNG------NKLITEYFRCPSSDQRQKKACKVN 661
Query: 328 TAPEEQQEMKSSGSVSGRNHKGKNSSTNRKLRDVPKWCSIEGTPFRVDAFKYLRGDCSHW 387
T + KS+ +G K K++D P WC I GTPFRVDAF+YLRGDC HW
Sbjct: 662 TPSNLNSQKKSNAKATGGRRTVKG-----KVKDTPIWCCIPGTPFRVDAFRYLRGDCCHW 716
Query: 388 FLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQKVEI 447
FLTHFH+D + L + G + A LV+ IGIP+D+LH+LPLN+K+ I
Sbjct: 717 FLTHFHVDHYQGLTKSFCH------GKIYCSSVTANLVHYKIGIPWDRLHVLPLNEKITI 770
Query: 448 AGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMCPVHTLILDTT 507
AG+++TC DANHCPG++IILF+P NGKAVLHTGDFRFS E+A N VL+ P+HTLILDTT
Sbjct: 771 AGVNLTCFDANHCPGAVIILFEPSNGKAVLHTGDFRFSSEMANNRVLQSSPIHTLILDTT 830
Query: 508 YCNPQYDFPKQEAVIQFVIDAIQAEAFNPGTLFLIGSYTIGKERLFLEVARSLRRKVYVN 567
YCNP+YDFP QE VIQFVI+AIQAEAFNP TLFLIGSYTIGKERL++EVAR L++K+YV
Sbjct: 831 YCNPRYDFPTQEIVIQFVIEAIQAEAFNPKTLFLIGSYTIGKERLYMEVARLLQKKIYVG 890
Query: 568 AAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQYASRFSLIVAF 627
AAKL++LK L +E M WFT NE ES+IHV PMWTLAS KR+K++S+QYA RF LIVAF
Sbjct: 891 AAKLQILKHLGLPQEIMHWFTANEAESHIHVVPMWTLASFKRMKYLSTQYADRFDLIVAF 950
Query: 628 SPTGWTFGKGKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVNNDG 687
PTGW+FGKGKK++PGR+WQ G IIRYEVPYSEH SFTEL++FV+ ISP +IIPSVNNDG
Sbjct: 951 CPTGWSFGKGKKRTPGRKWQQGAIIRYEVPYSEHSSFTELREFVRFISPEHIIPSVNNDG 1010
Query: 688 PESANAMIS 696
P+SANAM++
Sbjct: 1011 PDSANAMLA 1019
>emb|CAA66817.1| hypothetical protein [Arabidopsis thaliana]
gi|11994302|dbj|BAB01732.1| unnamed protein product
[Arabidopsis thaliana] gi|26449933|dbj|BAC42087.1|
unknown protein [Arabidopsis thaliana]
gi|28827378|gb|AAO50533.1| unknown protein [Arabidopsis
thaliana] gi|15231597|ref|NP_189302.1| DNA cross-link
repair protein-related [Arabidopsis thaliana]
gi|30688447|ref|NP_850635.1| DNA cross-link repair
protein-related [Arabidopsis thaliana]
Length = 484
Score = 228 bits (582), Expect = 4e-58
Identities = 142/403 (35%), Positives = 224/403 (55%), Gaps = 24/403 (5%)
Query: 304 SKPAANKKI--TEYFPGFATNGKKVSTAPEEQQEMKSSGSVSGRNHKGKNSSTNRKLRDV 361
+ P KK+ T+ F + T+P K++ ++ R +S +N R
Sbjct: 88 NSPDTTKKMKQTDLFQSWGLQKPSPFTSPASNSAKKTTSALGKRRRD--SSFSNDSPRPC 145
Query: 362 PKWCSIEGTPFRVDAFKY--LRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFE 419
P + + GTPF VDAF+Y ++G CS +FLTHFH D L T G +
Sbjct: 146 PFYKKLPGTPFTVDAFRYGCVQG-CSAYFLTHFHADHYIGL------TKAWSHGPIYCSS 198
Query: 420 CFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHT 479
+RL+ +++ + +H L L+ + I GI VT ++ANHCPG+ +I F+ +G LHT
Sbjct: 199 LTSRLLRLSLSVNPSSIHPLELDVEYTINGIKVTLIEANHCPGAALIHFRLLDGTCYLHT 258
Query: 480 GDFRFSDELAINPVLRMCPVHTLILDTTYCNPQYDFPKQEAVIQFVI----DAIQAEAFN 535
GDFR S ++ +P+L VH L LDTTYCNP+Y FP +E V+ +V+ D ++ +
Sbjct: 259 GDFRASKQMQTHPLLFNQRVHVLYLDTTYCNPRYKFPSKEDVLSYVVRITKDFLRKQ--- 315
Query: 536 PGTLFLIGSYTIGKERLFLEVARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESN 595
P TL ++GSY+IGKE ++L +A++L K++ NA++ R+L+ +++ T + +
Sbjct: 316 PKTLIVVGSYSIGKECVYLAIAKALGVKIFANASRRRILQSFGWDDISKNLST-DGKATC 374
Query: 596 IHVAPMWTLASLKRLKHVSSQYASRFSLIVAFSPTGWTFGK--GKKKSPGRRWQNGTIIR 653
+HV PM +L ++RL Y ++ ++AF PTGWT+ + G+ + G I
Sbjct: 375 LHVLPMSSL-KVERLDEHLKIYREQYGAVLAFRPTGWTYSEKIGEHLDLIKPTSRGKITI 433
Query: 654 YEVPYSEHCSFTELKDFVKHISPANIIPSVNNDGPESANAMIS 696
Y VPYSEH SFTEL++FV+ + P IIP+VNN + M S
Sbjct: 434 YGVPYSEHSSFTELREFVQFLRPDKIIPTVNNGNAGTREKMQS 476
>emb|CAA66406.1| orf12 [Arabidopsis thaliana]
Length = 484
Score = 228 bits (582), Expect = 4e-58
Identities = 142/403 (35%), Positives = 224/403 (55%), Gaps = 24/403 (5%)
Query: 304 SKPAANKKI--TEYFPGFATNGKKVSTAPEEQQEMKSSGSVSGRNHKGKNSSTNRKLRDV 361
+ P KK+ T+ F + T+P K++ ++ R +S +N R
Sbjct: 88 NSPDTTKKMKQTDLFQSWGLQKPSPFTSPASNSAKKTTSALGKRRRD--SSFSNDSPRPC 145
Query: 362 PKWCSIEGTPFRVDAFKY--LRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFE 419
P + + GTPF VDAF+Y ++G CS +FLTHFH D L T G +
Sbjct: 146 PFYKKLPGTPFTVDAFRYGCVQG-CSAYFLTHFHADHYIGL------TKAWSHGPIYCSS 198
Query: 420 CFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHT 479
+RL+ +++ + +H L L+ + I GI VT ++ANHCPG+ +I F+ +G LHT
Sbjct: 199 LTSRLLRLSLSVNPSSIHPLELDVEYTINGIKVTLIEANHCPGAALIHFRLLDGTCYLHT 258
Query: 480 GDFRFSDELAINPVLRMCPVHTLILDTTYCNPQYDFPKQEAVIQFVI----DAIQAEAFN 535
GDFR S ++ +P+L VH L LDTTYCNP+Y FP +E V+ +V+ D ++ +
Sbjct: 259 GDFRASKQMQTHPLLFNQRVHVLYLDTTYCNPRYKFPSKEDVLSYVVRITKDFLRKQ--- 315
Query: 536 PGTLFLIGSYTIGKERLFLEVARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESN 595
P TL ++GSY+IGKE ++L +A++L K++ NA++ R+L+ +++ T + +
Sbjct: 316 PKTLIVVGSYSIGKECVYLAIAKALGVKIFANASRRRILQSFGWDDISKNLST-DGKATC 374
Query: 596 IHVAPMWTLASLKRLKHVSSQYASRFSLIVAFSPTGWTFGK--GKKKSPGRRWQNGTIIR 653
+HV PM +L ++RL Y ++ ++AF PTGWT+ + G+ + G I
Sbjct: 375 LHVLPMSSL-KVERLDEHLKIYREQYGAVLAFRPTGWTYSEKIGEHLDLIKPTSRGKITI 433
Query: 654 YEVPYSEHCSFTELKDFVKHISPANIIPSVNNDGPESANAMIS 696
Y VPYSEH SFTEL++FV+ + P IIP+VNN + M S
Sbjct: 434 YGVPYSEHSSFTELREFVQFLRPDKIIPTVNNGNAGTREKMQS 476
>gb|AAH97815.1| Unknown (protein for IMAGE:3377284) [Xenopus laevis]
Length = 526
Score = 220 bits (561), Expect = 1e-55
Identities = 144/430 (33%), Positives = 227/430 (52%), Gaps = 34/430 (7%)
Query: 280 EPRRNRNQNVKMQPHRPERKVD----GTSKPAANKKITEYFPGFATNGKKVSTAPEEQQE 335
+P+ + K+ P +P+ + D SKP K+ G + G K A E
Sbjct: 101 KPKAKEDPKTKVPP-KPQNQHDVALPSESKPGQRKR-----KGQGSLGDKDPLADHTNTE 154
Query: 336 MK-SSGSVSGRNHKGKNSSTN---RKLRDVPKWCSIEGTPFRVDAFKYLRGD-CSHWFLT 390
+ ++G+ G + + ST + + P + I GT F VDAF+Y + + CS +FLT
Sbjct: 155 PEGATGNQRGGWKRPRQFSTGGEGKGKKQCPFYKKIPGTGFAVDAFQYGQIEGCSAYFLT 214
Query: 391 HFHLDR----TRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQKVE 446
HFH D T+ I+ + + LV + + + ++ LP+N +
Sbjct: 215 HFHSDHYGGLTKKFRFPIYCSKIT-----------GNLVQNKLRVESEFINTLPMNTECV 263
Query: 447 IAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMCPVHTLILDT 506
+ GI V L+ANHCPG++++LF+ PNG +VLHTGDFR + P L VHTL LDT
Sbjct: 264 VNGIRVVLLEANHCPGAVLLLFRLPNGTSVLHTGDFRADRSMESYPALIGQRVHTLYLDT 323
Query: 507 TYCNPQYDFPKQEAVIQFVID-AIQAEAFNPGTLFLIGSYTIGKERLFLEVARSLRRKVY 565
TYC+P+Y FP Q+ IQF ++ A + P TL + G+Y++GKE++FL +A L KV
Sbjct: 324 TYCSPEYTFPPQQETIQFAVNIAFEMVTLYPCTLVVCGTYSVGKEKVFLAIADVLGCKVC 383
Query: 566 VNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQYASRFSLIV 625
++ K + ++CLE E+ T + H + +HV PM + + K L ++ ++ ++
Sbjct: 384 MSQDKYKTMQCLE-SEDIRSLVTTDWHSTALHVLPMMQV-NFKGLNVHLGKFPGKYDRVL 441
Query: 626 AFSPTGWTFGKGKKKSPGRRWQ-NGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVN 684
AF PTGWT+ + + G + Y +PYSEH S++ELK FV+ + P IIP+VN
Sbjct: 442 AFKPTGWTYSDSSVLVADIKPEIRGKVTVYGIPYSEHSSYSELKRFVQWLKPQKIIPTVN 501
Query: 685 NDGPESANAM 694
S +AM
Sbjct: 502 VGNYNSRSAM 511
>ref|NP_001008683.1| SNM1A protein [Gallus gallus] gi|47156206|gb|AAR27404.1| SNM1A
[Gallus gallus]
Length = 972
Score = 218 bits (556), Expect = 4e-55
Identities = 151/441 (34%), Positives = 225/441 (50%), Gaps = 55/441 (12%)
Query: 286 NQNVKMQPHRPERKVDGTSKPAANKKITEYFPGFA--TNGKKV--STAPE----EQQEMK 337
+ N K P + +++D K+ E G A + GK++ S AP QQ+ K
Sbjct: 520 SMNAKSSPAKELKQMDIGVFFGLKPKVKEESKGEACLSEGKQIPSSVAPSGKRPRQQKRK 579
Query: 338 SSGSV--------------------SGRNHKGKN----SSTN---RKLRDVPKWCSIEGT 370
+ GSV SG K + SST + + P + I GT
Sbjct: 580 AEGSVEDLEAVEESSNKDGGDANVTSGGQRKWRKRFRESSTTDEGARKKQCPFYKKIPGT 639
Query: 371 PFRVDAFKY--LRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFEC---FARLV 425
F VDAF+Y + G C+ +FLTHFH D L N ++ C LV
Sbjct: 640 GFTVDAFQYGEIEG-CTAYFLTHFHSDHYCGLTKN----------FVFPLYCNKITGNLV 688
Query: 426 NMNIGIPYDKLHILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFS 485
+ + +++LP++ + + GI V LDANHCPG+ +ILF P+G A+LHTGDFR
Sbjct: 689 KSKLRVKEQYINVLPMDTECIVNGIKVLLLDANHCPGATMILFYLPSGTAILHTGDFRAD 748
Query: 486 DELAINPVLRMCPVHTLILDTTYCNPQYDFPKQEAVIQFVID-AIQAEAFNPGTLFLIGS 544
+ P L +HTL LDTTYC+P+Y FP Q+ VIQF ++ A + NP TL + G+
Sbjct: 749 PSMERYPALIGQKIHTLYLDTTYCSPEYTFPSQQEVIQFAVNTAFEMVTLNPRTLVVCGT 808
Query: 545 YTIGKERLFLEVARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTL 604
Y+IGKE++FL +A L K ++ K + L+CLE + T+N + +H+ PM +
Sbjct: 809 YSIGKEKVFLAIAEVLGSKASMSRDKYKTLQCLESAAVN-SLITMNWDGTLLHILPMMQI 867
Query: 605 ASLKRLKHVSSQYASRFSLIVAFSPTGWTFGKGKKKSPGRRWQ-NGTIIRYEVPYSEHCS 663
+ K L+ ++++ F ++AF PTGWT+ + Q G I Y +PYSEH S
Sbjct: 868 -NFKGLQDHLNKFSENFDQVLAFKPTGWTYSDSCLSVMDIKPQTRGNITIYGIPYSEHSS 926
Query: 664 FTELKDFVKHISPANIIPSVN 684
+ E+K FV+ + P IIP+VN
Sbjct: 927 YLEMKRFVQWLKPQKIIPTVN 947
>ref|XP_214730.2| PREDICTED: similar to SNM1 protein [Rattus norvegicus]
Length = 1026
Score = 214 bits (544), Expect = 1e-53
Identities = 135/349 (38%), Positives = 187/349 (52%), Gaps = 15/349 (4%)
Query: 350 KNSSTNRKLRDVPKWCSIEGTPFRVDAFKY--LRGDCSHWFLTHFHLDRTRLLDLNIFDT 407
++S R P + I GT F VDAF+Y + G C+ +FLTHFH D L
Sbjct: 676 ESSGGGESRRTCPFYKRIPGTGFTVDAFQYGEIEG-CTAYFLTHFHSDHYAGLS-----K 729
Query: 408 DLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTCLDANHCPGSIIIL 467
D + Y E L+ + + +H LP++ + + G+ V LDANHCPG+ +IL
Sbjct: 730 DFTRPIYCS--EITGSLLKKKLRVQEQYIHQLPMDTECIVDGVKVVLLDANHCPGATMIL 787
Query: 468 FQPPNGKAVLHTGDFRFSDELAINPVLRMCPVHTLILDTTYCNPQYDFPKQEAVIQFVID 527
FQ PNG LHTGDFR +D +L VHTL LDTTYC+P+Y FP Q+ IQF I+
Sbjct: 788 FQLPNGAVTLHTGDFR-ADPSMERSLLASRKVHTLFLDTTYCSPEYTFPSQQEAIQFAIN 846
Query: 528 -AIQAEAFNPGTLFLIGSYTIGKERLFLEVARSLRRKVYVNAAKLRLLKCLEFEEEDMQW 586
A +A NP L + G+Y IGKE++FL +A L KV ++ K + L+CL E
Sbjct: 847 TAFEAVTLNPRALIVCGTYCIGKEKVFLAIADVLGSKVGMSQEKYKTLQCLNIPEVS-SL 905
Query: 587 FTLNEHESNIHVAPMWTLASLKRLKHVSSQYASRFSLIVAFSPTGWTFGKGKKKSPGRRW 646
T + S +H+ PM + + K L++ + +F I+AF PTGWT
Sbjct: 906 ITTDMCNSLVHLLPMMQI-NFKGLQNHLKKCGGKFDQILAFRPTGWTHSNNITSIADITP 964
Query: 647 Q-NGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVNNDGPESANAM 694
Q G I Y +PYSEH S+ E+K FV+ + P IIP+VN +S N M
Sbjct: 965 QTKGNIAIYGIPYSEHSSYLEMKRFVQWLKPQKIIPTVNVGTFQSRNTM 1013
>ref|XP_535018.1| PREDICTED: similar to DNA-crosslink repair gene SNM1 [Canis
familiaris]
Length = 1227
Score = 212 bits (539), Expect = 4e-53
Identities = 145/430 (33%), Positives = 217/430 (49%), Gaps = 35/430 (8%)
Query: 260 LCEFRKGIAASNEKPEDALAEPRRNRNQNVKMQPHRPERKVDGTSKPAANKKITEYFPGF 319
L E + + SNE+P+ R+ ++ ++ P+++ ++
Sbjct: 805 LNESQPSVEVSNERPQR-----RKRLKKSDSLEEGTPQKRSHHLNR-------------- 845
Query: 320 ATNGKKVSTAPEEQQEMKSSGSVSGRNHKGKNSSTNRKLRD--VPKWCSIEGTPFRVDAF 377
T K V+ ++ + G + N K S +LR P + I GT F VDAF
Sbjct: 846 -TESKTVNLKKDKVSIKSACGRLQRGNMKIPESPNAGQLRKKTCPFYKKIPGTGFTVDAF 904
Query: 378 KY-LRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKL 436
+Y L C+ +FLTHFH D L N + E L+ + + +
Sbjct: 905 QYGLVEGCTAYFLTHFHSDHYAGLSKNFTFP-------VYCSEITGNLLKSKLHVQKQYI 957
Query: 437 HILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRM 496
H LP++ + + G+ V LDANHCPG+++ILF PNG +LHTGDFR +D L
Sbjct: 958 HPLPMDTECIVNGVKVVLLDANHCPGAVMILFYLPNGNVLLHTGDFR-ADPTMERSRLAG 1016
Query: 497 CPVHTLILDTTYCNPQYDFPKQEAVIQFVID-AIQAEAFNPGTLFLIGSYTIGKERLFLE 555
VHTL LDTTYC+P+Y FP Q+ VIQF I+ A +A NP L + G+Y+IGKE++FL
Sbjct: 1017 QKVHTLYLDTTYCSPEYSFPSQQEVIQFAINTAFEAVTRNPRVLVVCGTYSIGKEKVFLA 1076
Query: 556 VARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSS 615
+A L +V ++ K + L+CL + + + T + S +H+ PM + + K L+
Sbjct: 1077 IADVLGSRVAMSQEKYKTLQCLNIPDLN-SFITTDMCNSLVHLLPMMQI-NFKALQSHLK 1134
Query: 616 QYASRFSLIVAFSPTGWTFGKGKKKSPGRRWQ-NGTIIRYEVPYSEHCSFTELKDFVKHI 674
+ F+ I+AF PTGWT Q G I Y +PYSEH S+ ELK FV+ +
Sbjct: 1135 KCGGEFNQILAFRPTGWTHSNQLTNIRDIVPQIKGNISIYGIPYSEHSSYLELKRFVQWL 1194
Query: 675 SPANIIPSVN 684
P IIP+VN
Sbjct: 1195 KPQKIIPTVN 1204
>ref|XP_472369.1| OSJNBb0012E08.8 [Oryza sativa (japonica cultivar-group)]
gi|38345210|emb|CAD40784.2| OSJNBb0012E08.8 [Oryza
sativa (japonica cultivar-group)]
Length = 481
Score = 212 bits (539), Expect = 4e-53
Identities = 131/363 (36%), Positives = 204/363 (56%), Gaps = 15/363 (4%)
Query: 339 SGSVSGRNHKGKN---SSTNRKLRDVPKWCSIEGTPFRVDAFKYLRGD-CSHWFLTHFHL 394
SGS GR + ++ +RK P + I GTPF VDAF+Y + C+ +FL+HFH
Sbjct: 116 SGSWPGRKRRRGGEVEAAADRKPLACPFYKKIPGTPFTVDAFRYGAVEGCNAYFLSHFHH 175
Query: 395 DRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTC 454
D L T G + ARLV M + + + + L L+++ I G+ VT
Sbjct: 176 DHYGGL------TKKWCHGPIYCTALTARLVKMCLSVNPEYICPLELDKEYVIEGVSVTL 229
Query: 455 LDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMCPVHTLILDTTYCNPQYD 514
L+ANHCPG+ +I F+ +GK LHTGDFR S + + P+L+ ++ L LDTTYCNP+Y
Sbjct: 230 LEANHCPGAALIHFRLGDGKKYLHTGDFRASKSMQLYPLLQRGQINLLYLDTTYCNPKYK 289
Query: 515 FPKQEAVIQFVI-DAIQAEAFNPGTLFLIGSYTIGKERLFLEVARSLRRKVYVNAAKLRL 573
FP +E VI F + A + P TL ++G+Y+IGKE ++L ++++L+ +Y +A++ R+
Sbjct: 290 FPPKEDVIDFAVRTAKRYLQKEPKTLIVVGAYSIGKENVYLAISKALQVPIYTDASRRRI 349
Query: 574 LKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQYASRFSLIVAFSPTGWT 633
L + + + + S++HV P+ +L K++ + RF ++AF PTGWT
Sbjct: 350 LHAFGWSDLS-KMICSDSQSSSLHVLPLSSLRHENLQKYLET-LKQRFLAVLAFRPTGWT 407
Query: 634 FGK--GKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVNNDGPESA 691
F + G + + G I Y VPYSEH SF+EL++FV + P +IP+VN S
Sbjct: 408 FSEETGNQLDLIKPSSRGKITIYGVPYSEHSSFSELREFVMFLRPQKVIPTVNVGNAASR 467
Query: 692 NAM 694
+ M
Sbjct: 468 DKM 470
>dbj|BAD82911.1| Snm1 [Oryza sativa (japonica cultivar-group)]
Length = 485
Score = 211 bits (537), Expect = 6e-53
Identities = 129/353 (36%), Positives = 201/353 (56%), Gaps = 15/353 (4%)
Query: 339 SGSVSGRNHKGKN---SSTNRKLRDVPKWCSIEGTPFRVDAFKYLRGD-CSHWFLTHFHL 394
SGS GR + ++ +RK P + I GTPF VDAF+Y + C+ +FL+HFH
Sbjct: 116 SGSWPGRKRRRGGEVEAAADRKPLACPFYKKIPGTPFTVDAFRYGAVEGCNAYFLSHFHH 175
Query: 395 DRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTC 454
D L T G + ARLV M + + + + L L+++ I G+ VT
Sbjct: 176 DHYGGL------TKKWCHGPIYCTALTARLVKMCLSVNPEYICPLELDKEYVIEGVSVTL 229
Query: 455 LDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMCPVHTLILDTTYCNPQYD 514
L+ANHCPG+ +I F+ +GK LHTGDFR S + + P+L+ ++ L LDTTYCNP+Y
Sbjct: 230 LEANHCPGAALIHFRLGDGKKYLHTGDFRASKSMQLYPLLQRGQINLLYLDTTYCNPKYK 289
Query: 515 FPKQEAVIQFVI-DAIQAEAFNPGTLFLIGSYTIGKERLFLEVARSLRRKVYVNAAKLRL 573
FP +E VI F + A + P TL ++G+Y+IGKE ++L ++++L+ +Y +A++ R+
Sbjct: 290 FPPKEDVIDFAVRTAKRYLQKEPKTLIVVGAYSIGKENVYLAISKALQVPIYTDASRRRI 349
Query: 574 LKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQYASRFSLIVAFSPTGWT 633
L + + + + S++HV P+ +L K++ + RF ++AF PTGWT
Sbjct: 350 LHAFGWSDLS-KMICSDSQSSSLHVLPLSSLRHENLQKYLET-LKQRFLAVLAFRPTGWT 407
Query: 634 FGK--GKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVN 684
F + G + + G I Y VPYSEH SF+EL++FV + P +IP+VN
Sbjct: 408 FSEETGNQLDLIKPSSRGKITIYGVPYSEHSSFSELREFVMFLRPQKVIPTVN 460
>dbj|BAD90139.1| mKIAA0086 protein [Mus musculus]
Length = 984
Score = 209 bits (533), Expect = 2e-52
Identities = 136/360 (37%), Positives = 190/360 (52%), Gaps = 20/360 (5%)
Query: 344 GRNHKG-----KNSSTNRKLRDVPKWCSIEGTPFRVDAFKY--LRGDCSHWFLTHFHLDR 396
GR +G ++S R P + I GT F VDAF+Y + G C+ +FLTHFH D
Sbjct: 623 GRTQRGNMNISESSGAGEVRRTCPFYKRIPGTGFTVDAFQYGEIEG-CTAYFLTHFHSDH 681
Query: 397 TRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTCLD 456
L D + Y E L+ + + + LP++ + + + V LD
Sbjct: 682 YAGLS-----KDFTRPVYCS--EITGNLLKKKLRVQEQYIRQLPMDTECVVDSVKVVLLD 734
Query: 457 ANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMCPVHTLILDTTYCNPQYDFP 516
ANHCPG+ +ILFQ PNG +LHTGDFR +D L VHTL LDTTYC+P+Y FP
Sbjct: 735 ANHCPGATMILFQLPNGAVILHTGDFR-ADPSMERSRLAGRKVHTLFLDTTYCSPEYTFP 793
Query: 517 KQEAVIQFVID-AIQAEAFNPGTLFLIGSYTIGKERLFLEVARSLRRKVYVNAAKLRLLK 575
Q+ VIQF I+ A +A NP L + G+Y IGKE++FL +A L KV ++ K + L+
Sbjct: 794 SQQEVIQFAINTAFEAVTLNPRALVVCGTYCIGKEKVFLAIADVLGSKVGMSQEKYKTLQ 853
Query: 576 CLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQYASRFSLIVAFSPTGWTFG 635
CL E T + +S +H+ PM + + K L+ + ++ I+AF PTGWT
Sbjct: 854 CLNIPEVS-SLITTDMCDSLVHLLPMMQI-NFKGLQSHLKKCGGKYDQILAFRPTGWTHS 911
Query: 636 KGKKKSPGRRWQ-NGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVNNDGPESANAM 694
+ Q G I Y +PYSEH S+ E+K FV+ + P IIP+VN S N M
Sbjct: 912 NNITSTADIIPQTRGNISIYGIPYSEHSSYLEMKRFVQWLKPQKIIPTVNVGSFRSRNTM 971
>ref|NP_061301.1| DNA cross-link repair 1A, PSO2 homolog [Mus musculus]
gi|7595835|gb|AAF64472.1| SNM1 protein [Mus musculus]
Length = 1023
Score = 208 bits (529), Expect = 5e-52
Identities = 135/360 (37%), Positives = 190/360 (52%), Gaps = 20/360 (5%)
Query: 344 GRNHKG-----KNSSTNRKLRDVPKWCSIEGTPFRVDAFKY--LRGDCSHWFLTHFHLDR 396
GR +G ++S R P + I GT F VDAF+Y + G C+ +FLTHFH D
Sbjct: 662 GRTQRGNMNISESSGAGEVRRTCPFYKRIPGTGFTVDAFQYGEIEG-CTAYFLTHFHSDH 720
Query: 397 TRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTCLD 456
L D + Y E L+ + + + LP++ + + + V +D
Sbjct: 721 YAGLS-----KDFTRPVYCS--EITGNLLKKKLRVQEQYIRQLPMDTECVVDSVKVVFVD 773
Query: 457 ANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMCPVHTLILDTTYCNPQYDFP 516
ANHCPG+ +ILFQ PNG +LHTGDFR +D L VHTL LDTTYC+P+Y FP
Sbjct: 774 ANHCPGATMILFQLPNGAVILHTGDFR-ADPSMERSRLAGRKVHTLFLDTTYCSPEYTFP 832
Query: 517 KQEAVIQFVID-AIQAEAFNPGTLFLIGSYTIGKERLFLEVARSLRRKVYVNAAKLRLLK 575
Q+ VIQF I+ A +A NP L + G+Y IGKE++FL +A L KV ++ K + L+
Sbjct: 833 SQQEVIQFAINTAFEAVTLNPRALVVCGTYCIGKEKVFLAIADVLGSKVGMSQEKYKTLQ 892
Query: 576 CLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQYASRFSLIVAFSPTGWTFG 635
CL E T + +S +H+ PM + + K L+ + ++ I+AF PTGWT
Sbjct: 893 CLNIPEVS-SLITTDMCDSLVHLLPMMQI-NFKGLQSHLKKCGGKYDQILAFRPTGWTHS 950
Query: 636 KGKKKSPGRRWQ-NGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVNNDGPESANAM 694
+ Q G I Y +PYSEH S+ E+K FV+ + P IIP+VN S N M
Sbjct: 951 NNITSTADIIPQTRGNISIYGIPYSEHSSYLEMKRFVQWLKPQKIIPTVNVGSFRSRNTM 1010
>emb|CAI12637.1| DNA cross-link repair 1A (PSO2 homolog, S. cerevisiae) [Homo sapiens]
gi|16753254|gb|AAH13124.1| DNA-crosslink repair gene SNM1
[Homo sapiens]
Length = 1040
Score = 206 bits (523), Expect = 3e-51
Identities = 132/369 (35%), Positives = 197/369 (52%), Gaps = 15/369 (4%)
Query: 321 TNGKKVSTAPEEQQEMKSSGSVSGRNHKGKNSST--NRKLRDVPKWCSIEGTPFRVDAFK 378
T + V+ + + + G + N K SS + + P + I GT F VDAF+
Sbjct: 659 TESEAVNLSKVKVFTKSAHGGLQRGNKKIPESSNVGGSRKKTCPFYKKIPGTGFTVDAFQ 718
Query: 379 Y-LRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLH 437
Y + C+ +FLTHFH D L + + E L+ + + +H
Sbjct: 719 YGVVEGCTAYFLTHFHSDHYAGLSKHFTFP-------VYCSEITGNLLKNKLHVQEQYIH 771
Query: 438 ILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMC 497
LPL+ + + G+ V LDANHCPG+++ILF PNG +LHTGDFR +D +L
Sbjct: 772 PLPLDTECIVNGVKVVLLDANHCPGAVMILFYLPNGTVILHTGDFR-ADPSMERSLLADQ 830
Query: 498 PVHTLILDTTYCNPQYDFPKQEAVIQFVID-AIQAEAFNPGTLFLIGSYTIGKERLFLEV 556
VH L LDTTYC+P+Y FP Q+ VI+F I+ A +A NP L + G+Y+IGKE++FL +
Sbjct: 831 KVHMLYLDTTYCSPEYTFPSQQEVIRFAINTAFEAVTLNPHALVVCGTYSIGKEKVFLAI 890
Query: 557 ARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQ 616
A L KV ++ K + L+CL E + T + S +H+ PM + + K L+ +
Sbjct: 891 ADVLGSKVGMSQEKYKTLQCLNIPEIN-SLITTDMCSSLVHLLPMMQI-NFKGLQSHLKK 948
Query: 617 YASRFSLIVAFSPTGWTF-GKGKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHIS 675
+++ I+AF PTGWT K + + G I Y +PYSEH S+ E+K FV+ +
Sbjct: 949 CGGKYNQILAFRPTGWTHSNKFTRIADVIPQTKGNISIYGIPYSEHSSYLEMKRFVQWLK 1008
Query: 676 PANIIPSVN 684
P IIP+VN
Sbjct: 1009 PQKIIPTVN 1017
>dbj|BAA07646.2| KIAA0086 [Homo sapiens]
Length = 1044
Score = 206 bits (523), Expect = 3e-51
Identities = 132/369 (35%), Positives = 197/369 (52%), Gaps = 15/369 (4%)
Query: 321 TNGKKVSTAPEEQQEMKSSGSVSGRNHKGKNSST--NRKLRDVPKWCSIEGTPFRVDAFK 378
T + V+ + + + G + N K SS + + P + I GT F VDAF+
Sbjct: 663 TESEAVNLSKVKVFTKSAHGGLQRGNKKIPESSNVGGSRKKTCPFYKKIPGTGFTVDAFQ 722
Query: 379 Y-LRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLH 437
Y + C+ +FLTHFH D L + + E L+ + + +H
Sbjct: 723 YGVVEGCTAYFLTHFHSDHYAGLSKHFTFP-------VYCSEITGNLLKNKLHVQEQYIH 775
Query: 438 ILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMC 497
LPL+ + + G+ V LDANHCPG+++ILF PNG +LHTGDFR +D +L
Sbjct: 776 PLPLDTECIVNGVKVVLLDANHCPGAVMILFYLPNGTVILHTGDFR-ADPSMERSLLADQ 834
Query: 498 PVHTLILDTTYCNPQYDFPKQEAVIQFVID-AIQAEAFNPGTLFLIGSYTIGKERLFLEV 556
VH L LDTTYC+P+Y FP Q+ VI+F I+ A +A NP L + G+Y+IGKE++FL +
Sbjct: 835 KVHMLYLDTTYCSPEYTFPSQQEVIRFAINTAFEAVTLNPHALVVCGTYSIGKEKVFLAI 894
Query: 557 ARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQ 616
A L KV ++ K + L+CL E + T + S +H+ PM + + K L+ +
Sbjct: 895 ADVLGSKVGMSQEKYKTLQCLNIPEIN-SLITTDMCSSLVHLLPMMQI-NFKGLQSHLKK 952
Query: 617 YASRFSLIVAFSPTGWTF-GKGKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHIS 675
+++ I+AF PTGWT K + + G I Y +PYSEH S+ E+K FV+ +
Sbjct: 953 CGGKYNQILAFRPTGWTHSNKFTRIADVIPQTKGNISIYGIPYSEHSSYLEMKRFVQWLK 1012
Query: 676 PANIIPSVN 684
P IIP+VN
Sbjct: 1013 PQKIIPTVN 1021
>gb|AAH62582.1| DNA-crosslink repair gene SNM1 [Homo sapiens]
gi|42734319|ref|NP_055696.2| DNA-crosslink repair gene
SNM1 [Homo sapiens]
Length = 1040
Score = 206 bits (523), Expect = 3e-51
Identities = 132/369 (35%), Positives = 197/369 (52%), Gaps = 15/369 (4%)
Query: 321 TNGKKVSTAPEEQQEMKSSGSVSGRNHKGKNSST--NRKLRDVPKWCSIEGTPFRVDAFK 378
T + V+ + + + G + N K SS + + P + I GT F VDAF+
Sbjct: 659 TESEAVNLSKVKVFTKSAHGGLQRGNKKIPESSNVGGSRKKTCPFYKKIPGTGFTVDAFQ 718
Query: 379 Y-LRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLH 437
Y + C+ +FLTHFH D L + + E L+ + + +H
Sbjct: 719 YGVVEGCTAYFLTHFHSDHYAGLSKHFTFP-------VYCSEITGNLLKNKLHVQEQYIH 771
Query: 438 ILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMC 497
LPL+ + + G+ V LDANHCPG+++ILF PNG +LHTGDFR +D +L
Sbjct: 772 PLPLDTECIVNGVKVVLLDANHCPGAVMILFYLPNGTVILHTGDFR-ADPSMERSLLADQ 830
Query: 498 PVHTLILDTTYCNPQYDFPKQEAVIQFVID-AIQAEAFNPGTLFLIGSYTIGKERLFLEV 556
VH L LDTTYC+P+Y FP Q+ VI+F I+ A +A NP L + G+Y+IGKE++FL +
Sbjct: 831 KVHMLYLDTTYCSPEYTFPSQQEVIRFAINTAFEAVTLNPHALVVCGTYSIGKEKVFLAI 890
Query: 557 ARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQ 616
A L KV ++ K + L+CL E + T + S +H+ PM + + K L+ +
Sbjct: 891 ADVLGSKVGMSQEKYKTLQCLNIPEIN-SLITTDMCSSLVHLLPMMQI-NFKGLQSHLKK 948
Query: 617 YASRFSLIVAFSPTGWTF-GKGKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHIS 675
+++ I+AF PTGWT K + + G I Y +PYSEH S+ E+K FV+ +
Sbjct: 949 CGGKYNQILAFRPTGWTHSNKFTRIADVIPQTKGNISIYGIPYSEHSSYLEMKRFVQWLK 1008
Query: 676 PANIIPSVN 684
P IIP+VN
Sbjct: 1009 PQKIIPTVN 1017
>gb|AAT09762.1| DNA cross-link repair 1A (PSO2 homolog, S. cerevisiae) [Homo sapiens]
Length = 1040
Score = 206 bits (523), Expect = 3e-51
Identities = 132/369 (35%), Positives = 197/369 (52%), Gaps = 15/369 (4%)
Query: 321 TNGKKVSTAPEEQQEMKSSGSVSGRNHKGKNSST--NRKLRDVPKWCSIEGTPFRVDAFK 378
T + V+ + + + G + N K SS + + P + I GT F VDAF+
Sbjct: 659 TESEAVNLSKVKVFTKSAHGGLQRGNKKIPESSNVGGSRKKTCPFYKKIPGTGFTVDAFQ 718
Query: 379 Y-LRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLH 437
Y + C+ +FLTHFH D L + + E L+ + + +H
Sbjct: 719 YGVVEGCTAYFLTHFHSDHYAGLSKHFTFP-------VYCSEITGNLLKNKLHVQEQYIH 771
Query: 438 ILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMC 497
LPL+ + + G+ V LDANHCPG+++ILF PNG +LHTGDFR +D +L
Sbjct: 772 PLPLDTECIVNGVKVVLLDANHCPGAVMILFYLPNGTVILHTGDFR-ADPSMERSLLADQ 830
Query: 498 PVHTLILDTTYCNPQYDFPKQEAVIQFVID-AIQAEAFNPGTLFLIGSYTIGKERLFLEV 556
VH L LDTTYC+P+Y FP Q+ VI+F I+ A +A NP L + G+Y+IGKE++FL +
Sbjct: 831 KVHMLYLDTTYCSPEYTFPSQQEVIRFAINTAFEAVTLNPHALVVCGTYSIGKEKVFLAI 890
Query: 557 ARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQ 616
A L KV ++ K + L+CL E + T + S +H+ PM + + K L+ +
Sbjct: 891 ADVLGSKVGMSQEKYKTLQCLNIPEIN-SLITTDMCSSLVHLLPMMQI-NFKGLQSHLKK 948
Query: 617 YASRFSLIVAFSPTGWTF-GKGKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHIS 675
+++ I+AF PTGWT K + + G I Y +PYSEH S+ E+K FV+ +
Sbjct: 949 CGGKYNQILAFRPTGWTHSNKFTRIADVIPQTKGNISIYGIPYSEHSSYLEMKRFVQWLK 1008
Query: 676 PANIIPSVN 684
P IIP+VN
Sbjct: 1009 PQKIIPTVN 1017
>ref|XP_508045.1| PREDICTED: similar to DNA-crosslink repair gene SNM1 [Pan
troglodytes]
Length = 1190
Score = 202 bits (513), Expect = 4e-50
Identities = 130/367 (35%), Positives = 195/367 (52%), Gaps = 15/367 (4%)
Query: 321 TNGKKVSTAPEEQQEMKSSGSVSGRNHKGKNSST--NRKLRDVPKWCSIEGTPFRVDAFK 378
T + V+ + + + G + N K SS + + P + I GT F VDAF+
Sbjct: 708 TESEAVNLSKVKVFTKSAHGGLQRGNKKIPESSNVGGSRKKTCPFYKKIPGTGFTVDAFQ 767
Query: 379 Y-LRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLH 437
Y + C+ +FLTHFH D L + + E L+ + + +H
Sbjct: 768 YGVVEGCTAYFLTHFHSDHYAGLSKHFTFP-------VYCSEITGNLLKNKLHVQEQYIH 820
Query: 438 ILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMC 497
LPL+ + + G+ V LDANHCPG+++ILF PNG +LHTGDFR +D +L
Sbjct: 821 PLPLDTECIVNGVKVVLLDANHCPGAVMILFYLPNGTVILHTGDFR-ADPSMERSLLADQ 879
Query: 498 PVHTLILDTTYCNPQYDFPKQEAVIQFVID-AIQAEAFNPGTLFLIGSYTIGKERLFLEV 556
VH L LDTTYC+P+Y FP Q+ VI+F I+ A +A NP L + G+Y+IGKE++FL +
Sbjct: 880 KVHMLYLDTTYCSPEYTFPSQQEVIRFAINTAFEAVTLNPHALVVCGTYSIGKEKVFLAI 939
Query: 557 ARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQ 616
A L KV ++ K + L+CL E + T + S +H+ PM + + K L+ +
Sbjct: 940 ADVLGSKVGMSREKYKTLQCLNIPEIN-SLITTDMCSSLVHLLPMMQI-NFKGLQSHLKK 997
Query: 617 YASRFSLIVAFSPTGWTF-GKGKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHIS 675
+++ I+AF PTGWT K + + G I Y +PYSEH S+ E+K FV+ +
Sbjct: 998 CGGKYNQILAFRPTGWTHSNKFTRIADVIPQTKGNISIYGIPYSEHSSYLEMKRFVQWLK 1057
Query: 676 PANIIPS 682
P IIP+
Sbjct: 1058 PQKIIPT 1064
>gb|AAO50928.1| similar to Arabidopsis thaliana (Mouse-ear cress). At2g45700
protein (Hypothetical 81.0 kDa protein) [Dictyostelium
discoideum] gi|66817282|ref|XP_642494.1| hypothetical
protein DDB0169391 [Dictyostelium discoideum]
gi|60470572|gb|EAL68551.1| hypothetical protein
DDB0169391 [Dictyostelium discoideum]
Length = 920
Score = 196 bits (499), Expect = 2e-48
Identities = 128/365 (35%), Positives = 187/365 (51%), Gaps = 25/365 (6%)
Query: 335 EMKSSGSVSGRNHKGKNSSTNRKLRDVP-KWCSIEGTPFRVDAFKYLRGDCSHWFLTHFH 393
++K +G + + T K + +P + I+GT F VD F+Y D +H+FLTHFH
Sbjct: 233 DLKKTGKEEAKKRGYQVRKTPTKEKKIPPSFKVIDGTNFLVDGFQYKSEDFTHYFLTHFH 292
Query: 394 LDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVT 453
D + T G + E +LV+ +G+ + N+ +EI G+ V
Sbjct: 293 SDHY------VGITKTWSFGNIYCTEETGKLVSHKLGVDQRYIVKCEWNKLIEIQGVKVA 346
Query: 454 CLDANHCPGSIIILFQPP---------NGKAVLHTGDFRFSDELAINPVLRMCPVHTLIL 504
LD+NHCPGS +ILF P +++LHTGDFR++ + P+L+ + L L
Sbjct: 347 FLDSNHCPGSALILFIIPLRNKDGEIIGEESILHTGDFRYNQSMNNYPLLKGRTISKLYL 406
Query: 505 DTTYCNPQYDFPKQEAVIQFVIDAIQAEAFNPG-TLFLIGSYTIGKERLFLEVARSLRRK 563
D TYC+PQY FP Q +I+ V ++ E N G TLFL G+Y IGKER+ LE+A+ +
Sbjct: 407 DNTYCDPQYVFPPQPEIIKQVASIVRKE--NDGETLFLFGTYVIGKERILLEIAKQEGKP 464
Query: 564 VYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQYASRFSL 623
V+V+ K +L CL D+ FT NE + M L+ L + S +++
Sbjct: 465 VHVSNEKYAILCCLS-TCLDINKFTTNELITPFRAVTMSMLSYHNMLSLLDSS-NNKYKR 522
Query: 624 IVAFSPTGWTFGKGKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSV 683
++ F PTGWT K R G Y V YSEH SF EL+D + H P IIP+V
Sbjct: 523 VIGFRPTGWTQAKKSITYLNR----GPTTFYSVAYSEHSSFNELRDCIDHFRPTQIIPTV 578
Query: 684 NNDGP 688
+ D P
Sbjct: 579 DCDSP 583
>ref|XP_614971.1| PREDICTED: similar to DNA-crosslink repair gene SNM1, partial [Bos
taurus] gi|61810453|ref|XP_593078.1| PREDICTED: similar
to DNA-crosslink repair gene SNM1, partial [Bos taurus]
Length = 365
Score = 189 bits (481), Expect = 2e-46
Identities = 118/313 (37%), Positives = 168/313 (52%), Gaps = 12/313 (3%)
Query: 384 CSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQ 443
C+ +FLTHFH D L N F + Y K L+ + + +H LP +
Sbjct: 50 CTAYFLTHFHSDHYAGLSKN-FTFPI----YCSKIT--GSLLKSKLHVQEQYIHPLPTDT 102
Query: 444 KVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMCPVHTLI 503
+ + GI V LDANHCPG+++IL PNG +LHTGDFR +D +L VHTL
Sbjct: 103 ECIVNGIKVIFLDANHCPGAVMILLCLPNGHVILHTGDFR-ADPSMERSLLGCHKVHTLY 161
Query: 504 LDTTYCNPQYDFPKQEAVIQFVID-AIQAEAFNPGTLFLIGSYTIGKERLFLEVARSLRR 562
LDTTYC+P+Y FP Q+ VIQF I+ A + NP L + G+Y+IGKE++FL +A L
Sbjct: 162 LDTTYCSPEYSFPSQQEVIQFAINTAFETVTLNPWALVVCGTYSIGKEKIFLAIANVLGS 221
Query: 563 KVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQYASRFS 622
KV ++ K L+C E + T + S +H+ PM + + K L++ + +++
Sbjct: 222 KVGMSKEKYNTLQCFNIPEVS-SFITTDMCNSLVHLLPMMQI-NFKGLQNHLKKCGGKYN 279
Query: 623 LIVAFSPTGWTFGKGKKKSPGRRWQ-NGTIIRYEVPYSEHCSFTELKDFVKHISPANIIP 681
I+AF PTGWT Q G I Y + YSEH S+ E+K FV+ + P IIP
Sbjct: 280 QILAFRPTGWTHSNKLTSLADIILQTKGNISIYGIRYSEHSSYLEMKRFVQWLKPQKIIP 339
Query: 682 SVNNDGPESANAM 694
+VN +S M
Sbjct: 340 TVNVGSLKSRRTM 352
>ref|NP_176845.1| ATP dependent DNA ligase family protein [Arabidopsis thaliana]
gi|12597768|gb|AAG60081.1| DNA ligase I, putative
[Arabidopsis thaliana]
Length = 1417
Score = 181 bits (460), Expect = 6e-44
Identities = 116/338 (34%), Positives = 175/338 (51%), Gaps = 23/338 (6%)
Query: 361 VPKWCSIEGTPFRVDAFKYLRGDCS-HWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFE 419
+P I T F VD F+ S +FL+HFH D L + G +
Sbjct: 54 IPNSKRIPNTNFIVDLFRLPHQSSSVAFFLSHFHSDHYSGLSSSW------SKGIIYCSH 107
Query: 420 CFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTCLDANHCPGSIIILF----QPPNGKA 475
ARLV + +P + LP+NQ V+I G +V ++ANHCPG++ LF + +
Sbjct: 108 KTARLVAEILQVPSQFVFALPMNQMVKIDGSEVVLIEANHCPGAVQFLFKVKLESSGFEK 167
Query: 476 VLHTGDFRFSDELAINPVLR-MCPVHTLILDTTYCNPQYDFPKQEAVIQFVIDAIQAEAF 534
+HTGDFRF DE+ +P L + LDTTYCNP++ FP QE + +V+ I +
Sbjct: 168 YVHTGDFRFCDEMRFDPFLNGFVGCDGVFLDTTYCNPKFVFPSQEESVGYVVSVID-KIS 226
Query: 535 NPGTLFLIGSYTIGKERLFLEVARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHES 594
LFL+ +Y +GKE++ +E+AR +RK+ V+A K+ +L L EE M FT +E+ES
Sbjct: 227 EEKVLFLVATYVVGKEKILVEIARRCKRKIVVDARKMSMLSVLGCGEEGM--FTEDENES 284
Query: 595 NIHV------APMWTLASLKRLKHVSSQYASRFSLIVAFSPTGWTFGKGKKKSPGRRWQN 648
++HV W +K + +V F PTGWT+ + K R +
Sbjct: 285 DVHVVGWNVLGETWPYFRPNFVKMNEIMVEKGYDKVVGFVPTGWTYEVKRNKFAVRFKDS 344
Query: 649 GTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVNND 686
I + VPYSEH ++ EL++F+K + P +IP+V D
Sbjct: 345 MEI--HLVPYSEHSNYDELREFIKFLKPKRVIPTVGVD 380
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.134 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,213,808,826
Number of Sequences: 2540612
Number of extensions: 53691021
Number of successful extensions: 146861
Number of sequences better than 10.0: 684
Number of HSP's better than 10.0 without gapping: 336
Number of HSP's successfully gapped in prelim test: 352
Number of HSP's that attempted gapping in prelim test: 145646
Number of HSP's gapped (non-prelim): 1014
length of query: 700
length of database: 863,360,394
effective HSP length: 135
effective length of query: 565
effective length of database: 520,377,774
effective search space: 294013442310
effective search space used: 294013442310
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 79 (35.0 bits)
Lotus: description of TM0133.3


