

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0126.7
(542 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
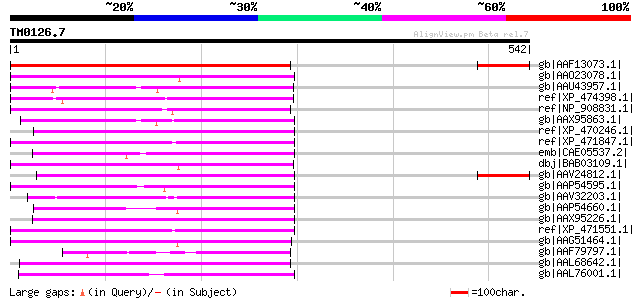
Sequences producing significant alignments: (bits) Value
gb|AAF13073.1| putative retroelement pol polyprotein [Arabidopsi... 233 9e-60
gb|AAO23078.1| polyprotein [Glycine max] 177 1e-42
gb|AAU43957.1| putative polyprotein [Oryza sativa (japonica cult... 176 2e-42
ref|XP_474398.1| OSJNBa0032F06.19 [Oryza sativa (japonica cultiv... 168 4e-40
ref|NP_908831.1| putative polyprotein [Oryza sativa (japonica cu... 167 1e-39
gb|AAX95863.1| retrotransposon protein, putative, unclassified [... 166 2e-39
ref|XP_470246.1| Putative plant disease resistance polyprotein [... 160 1e-37
ref|XP_471847.1| OSJNBb0062H02.17 [Oryza sativa (japonica cultiv... 158 4e-37
emb|CAE05537.2| OSJNBa0053B21.11 [Oryza sativa (japonica cultiva... 158 4e-37
dbj|BAB03109.1| retroelement pol polyprotein [Arabidopsis thalia... 157 1e-36
gb|AAV24812.1| putative polyprotein [Oryza sativa (japonica cult... 156 2e-36
gb|AAP54595.1| putative gag-pol polyprotein [Oryza sativa (japon... 155 3e-36
gb|AAV32203.1| unknown protein [Oryza sativa (japonica cultivar-... 155 4e-36
gb|AAP54660.1| putative plant disease resistance polyprotein [Or... 154 9e-36
gb|AAX95226.1| retrotransposon protein, putative, unclassified [... 154 9e-36
ref|XP_471551.1| OSJNBa0019J05.12 [Oryza sativa (japonica cultiv... 148 4e-34
gb|AAG51464.1| gypsy/Ty3 element polyprotein, putative [Arabidop... 148 5e-34
gb|AAF79797.1| T32E20.30 [Arabidopsis thaliana] 147 8e-34
gb|AAL68642.1| polyprotein [Oryza sativa (japonica cultivar-group)] 147 1e-33
gb|AAL76001.1| putative gag-pol polyprotein [Zea mays] 147 1e-33
>gb|AAF13073.1| putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1661
Score = 233 bits (595), Expect = 9e-60
Identities = 127/297 (42%), Positives = 191/297 (63%), Gaps = 4/297 (1%)
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVET-PYMVE 59
++L S+ + + +S+K+ G +G +V++LVD G + NFIS+ L+ + V +T + V+
Sbjct: 499 MTLSSLNDESQEQSMKMRGYIGNTKVVLLVDSGATCNFISEALVREKGWLVTQTRSFGVK 558
Query: 60 LEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELS 119
+ G ++ GKC D+ L +Q I Q+++LF+LG LD+VLG W A LG RAN+ +L
Sbjct: 559 VGGGRIIKSSGKCVDIPLEVQGIEFVQDYYLFDLGDLDLVLGFSWLAGLGETRANWRDLR 618
Query: 120 LRFHVGEK-VLLRGELGLIRKNASLSSLVKALQQQAEGFWVEFQHLAEV--QEELVAVPE 176
+ + +G V L G+ L R S+ S+ + ++ + +E L E QEE A+
Sbjct: 619 ISWQIGRTWVSLYGDPDLCRGQISMRSMERVIKYTGTAYLLELASLFESKKQEEQTALQP 678
Query: 177 QLQGLMGDYSELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEM 236
+Q L+ Y +F T LPP R ++HAI LQEG++ NIRPYR QK+EIEKLV+EM
Sbjct: 679 AIQRLLDQYQGVFQTPQLLPPVRNREHAITLQEGSSPVNIRPYRYSFAQKNEIEKLVREM 738
Query: 237 LDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKL 293
L+ I RPS+S +SS V+LVKKKDG WRFC+DYRA ++ TIP+K+PI +I+ELL +L
Sbjct: 739 LNAQIIRPSVSPYSSPVLLVKKKDGGWRFCVDYRALNEATIPDKYPIPVIEELLDEL 795
Score = 50.4 bits (119), Expect = 1e-04
Identities = 26/54 (48%), Positives = 35/54 (64%)
Query: 489 GVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFIIR 542
G VL Q R +AF S S QG+ +SVYEREL+A+V+A+ +W L FII+
Sbjct: 1022 GAVLSQNKRLIAFLSQAFSSQGRIRSVYERELLAIVKAVTKWKHYLSSKEFIIK 1075
>gb|AAO23078.1| polyprotein [Glycine max]
Length = 1552
Score = 177 bits (448), Expect = 1e-42
Identities = 109/305 (35%), Positives = 171/305 (55%), Gaps = 8/305 (2%)
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVETPYM-VE 59
LSL ++ +++ G++G + V +LVD G S NFI + PV P + V
Sbjct: 377 LSLNAMRGSNGVGTIRFTGQVGGIAVKILVDGGSSDNFIQPRVAQVLKLPVEPAPNLRVL 436
Query: 60 LEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELS 119
+ +G + +G + + L IQ ++ +L + G DV+LG W A+LG A++ L+
Sbjct: 437 VGNGQILSAEGIVQQLPLHIQGQEVKVPVYLLQISGADVILGSTWLATLGPHVADYAALT 496
Query: 120 LRFHVGEK-VLLRGELGLIRKNASLSSLVKALQQQA--EGFWVEFQHLAEVQEELVAVPE 176
L+F +K + L+GE A L + ++ E F ++ ++ L +P
Sbjct: 497 LKFFQNDKFITLQGEGNSEATQAQLHHFRRLQNTKSIEECFAIQLIQKEVPEDTLKDLPT 556
Query: 177 ----QLQGLMGDYSELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKL 232
+L L+ Y+++F+ LPP R QDHAI L++G+ +RPYR PH QK +IEK+
Sbjct: 557 NIDPELAILLHTYAQVFAVPASLPPQREQDHAIPLKQGSGPVKVRPYRYPHTQKDQIEKM 616
Query: 233 VKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYK 292
++EML GI +PS S FS ++LVKKKDGSWRFC DYRA + IT+ + FP+ +DELL +
Sbjct: 617 IQEMLVQGIIQPSNSPFSLPILLVKKKDGSWRFCTDYRALNAITVKDSFPMPTVDELLDE 676
Query: 293 LGEAK 297
L A+
Sbjct: 677 LHGAQ 681
Score = 42.4 bits (98), Expect = 0.038
Identities = 21/54 (38%), Positives = 35/54 (63%)
Query: 489 GVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFIIR 542
G VL Q P+A++S ++ + Q++S Y REL+A+ A+ ++ L G+ FIIR
Sbjct: 904 GAVLGQNGHPIAYFSKKLAPRMQKQSAYTRELLAITEALSKFRHYLLGNKFIIR 957
>gb|AAU43957.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1263
Score = 176 bits (445), Expect = 2e-42
Identities = 107/307 (34%), Positives = 172/307 (55%), Gaps = 17/307 (5%)
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQEL---MSSAHFPVVETPYM 57
+S+ ++ S+ ++++ G++ L +++LVD G S +F+S +L ++ F + P
Sbjct: 454 MSVHAVQGTDSQSTIRLQGKIQGLNILMLVDSGSSASFVSDQLADRLTGVQF--LAHPLS 511
Query: 58 VELEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEE 117
V++ +G + CQ + + + F + LGG D +LG+ W + +++
Sbjct: 512 VKVANGQVLHCQKEMPSAEWFVSGQKFLTTFKVVALGGYDAILGMDWLTKYSPMHIDWQL 571
Query: 118 LSLRFHV-GEKVLLRGELGLIRKNASLSSLVKALQQQ-------AEGFWVEFQHLAEVQE 169
S+ V G +VLLRG ++ N S +L+ + Q G L++ +
Sbjct: 572 KSIHLIVMGHEVLLRG----VQSNTSDCALISSQHLQHLHSIDSVAGLVHLCAALSDSTD 627
Query: 170 ELVAVPEQLQGLMGDYSELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEI 229
+L ++P+ +Q L+ ++S LF LPP+R DH I L GA N+RPYR QK EI
Sbjct: 628 QLSSIPDAVQSLVQEFSHLFEEPSGLPPARDYDHTIPLIPGAVPVNVRPYRYTPAQKDEI 687
Query: 230 EKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDEL 289
EK VK+ML GI +PS S FSS V+LVKKKDG+WRFC+DYR + IT+ NK+P+ IIDEL
Sbjct: 688 EKQVKDMLQKGIIQPSASPFSSPVLLVKKKDGTWRFCVDYRHLNAITVKNKYPLPIIDEL 747
Query: 290 LYKLGEA 296
+ +L A
Sbjct: 748 MDELAGA 754
Score = 39.3 bits (90), Expect = 0.32
Identities = 20/54 (37%), Positives = 30/54 (55%)
Query: 489 GVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFIIR 542
G VL+Q P+AF S + + Q S YE+E +A++ A+ W L F+IR
Sbjct: 896 GAVLMQDQHPVAFLSKALGPRTQVLSTYEKESLAIILAVDHWRSYLQHEEFLIR 949
>ref|XP_474398.1| OSJNBa0032F06.19 [Oryza sativa (japonica cultivar-group)]
gi|38345562|emb|CAE03436.2| OSJNBa0032F06.19 [Oryza
sativa (japonica cultivar-group)]
Length = 1575
Score = 168 bits (426), Expect = 4e-40
Identities = 109/302 (36%), Positives = 168/302 (55%), Gaps = 9/302 (2%)
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVE---TPYM 57
+SL++I S +L++ G + V++LVD G S +F+S ++ H V+ TP
Sbjct: 482 ISLQAIQGTVSAHTLRLQGSIQGFDVLILVDSGSSCSFLSSVVLP--HLTGVKSLLTPIQ 539
Query: 58 VELEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEE 117
V++ +G + C + + +Q F L L D++LG+ W + N++
Sbjct: 540 VKVANGTVLYCTSELPAAKWEVQGHQFSTNFKLLPLDNYDMILGMDWLEKYSPMDINWQA 599
Query: 118 LSLRFHVGEK-VLLRGELGLIRKN--ASLSSLVKALQQQAEGFWVEFQHLAEVQEELVAV 174
+++F + +K V L+G + + K S+ L Q A V+ ++ V
Sbjct: 600 KTIQFCLHDKSVELKGVVPELDKCDLVSIHQLQLLHNQGAVESLVQLT-ASDGSSSSQNV 658
Query: 175 PEQLQGLMGDYSELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVK 234
P +Q ++ +++++F+ LPPSR DH I L GA + NIRPYR QK+EIEK V+
Sbjct: 659 PSSIQSILDEFADIFTEPEGLPPSRSYDHTIPLIAGAQLVNIRPYRYTPDQKNEIEKQVQ 718
Query: 235 EMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKLG 294
EML GI RPS S FSS V+LVKKKDG+WRFC+DYR + ITI NK+P+ +IDELL +L
Sbjct: 719 EMLKKGIIRPSSSPFSSPVLLVKKKDGTWRFCVDYRHLNAITIKNKYPLPVIDELLDELA 778
Query: 295 EA 296
A
Sbjct: 779 GA 780
>ref|NP_908831.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1522
Score = 167 bits (422), Expect = 1e-39
Identities = 111/301 (36%), Positives = 164/301 (53%), Gaps = 12/301 (3%)
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVETPYM-VE 59
LSL+++A ++ SL++ G VI+LVD G SH+FI+ +L S + P V+
Sbjct: 383 LSLQAVAGTSATHSLQLRALAGNQVVIILVDSGSSHSFINADLCSKLNLSTDPIPVSTVK 442
Query: 60 LEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELS 119
+ +G ++C+ + + IQ + ++GG D+VLG+ W + + N+ +
Sbjct: 443 VANGETLQCEAQVSEFSWWIQGCTFVSSLKVLSMGGHDIVLGMDWLSKFSPMTCNWADRW 502
Query: 120 LRF-HVGEKVLLRG-ELGLIRKNASLSSLVKALQQQAEGFWVEFQHLAEVQ---EELV-- 172
L F + G+ V L+G + ++ + +SS Q W LA V L
Sbjct: 503 LEFTYQGKLVKLQGIQSHQLQYLSEVSSDQMMKWQVGNDIWA----LAVVDCLSSPLCDS 558
Query: 173 AVPEQLQGLMGDYSELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKL 232
AVP +Q ++ YS +F LPP R DH+I L GA N RPYR QK EIEK
Sbjct: 559 AVPASIQDVLQKYSSVFQPSAGLPPHRELDHSIPLVPGAIPVNSRPYRYSPAQKDEIEKQ 618
Query: 233 VKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYK 292
V+EML+ G+ S S F+S V+LVKKKD +WRFC+DYR + ITI NKFP+ I+DELL +
Sbjct: 619 VQEMLESGLISRSCSPFASPVLLVKKKDNTWRFCVDYRRLNAITIKNKFPLPIVDELLDE 678
Query: 293 L 293
L
Sbjct: 679 L 679
Score = 42.0 bits (97), Expect = 0.050
Identities = 20/54 (37%), Positives = 31/54 (57%)
Query: 489 GVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFIIR 542
G VL Q P+AF+S + Q+ S+YE+E +A++ AI +W L F I+
Sbjct: 907 GAVLSQEGHPIAFYSKALGMANQKLSIYEKEFLAIMMAIDKWRSYLLRGPFTIK 960
>gb|AAX95863.1| retrotransposon protein, putative, unclassified [Oryza sativa
(japonica cultivar-group)]
Length = 1126
Score = 166 bits (419), Expect = 2e-39
Identities = 104/294 (35%), Positives = 159/294 (53%), Gaps = 13/294 (4%)
Query: 12 RRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVET-PYMVELEDGHKVECQG 70
+R++++ G + V++L+D G S +F++Q L+ S V+ET P V + DG ++C
Sbjct: 440 KRTVRLQGLIQNQEVLILIDSGSSSSFVAQHLVDSLSLAVLETAPAQVVIADGGVLQCNK 499
Query: 71 KCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELSLRF-HVGEKVL 129
V Q + + LGG DV+LG+ W + N++ LRF + G+++
Sbjct: 500 VATKVDWWCQGHSFTSDLKVLQLGGYDVILGMDWLEHFSPMWINWQRKKLRFTYQGKRIT 559
Query: 130 LRGELGLIRKNASLSSLVKALQ------QQAEGFWVEFQHLAEVQEELVAVPEQLQGLMG 183
L G +++N S V A Q +A V+ + E++ +P ++ ++
Sbjct: 560 LTG----VKENLSDCKSVTAKQLKGMFKSRAIAQVVQLCSIEEIETN-EQLPPEISAVLQ 614
Query: 184 DYSELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVGIFR 243
+ + F LPP R DH I L A N +PYR QK EIE+ VK ML+ GI +
Sbjct: 615 ESQDQFQEPSRLPPHRDFDHNIPLMPNAKPVNAKPYRYAPKQKDEIERQVKVMLEQGIIK 674
Query: 244 PSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKLGEAK 297
S S F+S V+LVKKKDGSWRFC+DYR + ITI NKFP+ ++DELL +L AK
Sbjct: 675 DSSSPFASPVLLVKKKDGSWRFCVDYRGLNDITIKNKFPMPVVDELLDELAGAK 728
Score = 40.8 bits (94), Expect = 0.11
Identities = 20/53 (37%), Positives = 31/53 (57%)
Query: 489 GVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFII 541
G VL Q P+A+ S + + Q S YE+E +A++ A+Q+W L S F+I
Sbjct: 954 GAVLTQNGHPIAYLSKALGVKAQALSTYEKECLAILMAVQKWRAYLQHSEFVI 1006
>ref|XP_470246.1| Putative plant disease resistance polyprotein [Oryza sativa
(japonica cultivar-group)] gi|21397270|gb|AAM51834.1|
Putative plant disease resistance polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 624
Score = 160 bits (405), Expect = 1e-37
Identities = 99/276 (35%), Positives = 148/276 (52%), Gaps = 4/276 (1%)
Query: 26 VIVLVDCGVSHNFISQELMSSAHFPVVETPY-MVELEDGHKVECQGKCRDVQLSIQHIPI 84
+++LVD G SHNFI P+ P +V++ +G + CQ IQ
Sbjct: 210 LLLLVDSGSSHNFIDYNFAQMLGLPLKPIPSTLVKVANGDCLPCQYVVPGFFWWIQGHTF 269
Query: 85 EQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELSLRF-HVGEKVLLRGELGLIRKNASL 143
+ + LGG D +LG+ W G + ++ +L+F + G + L+G +
Sbjct: 270 TYDMRVVTLGGHDAILGMDWLEQWGEMSCHWANKTLKFQYQGSWISLQGVQDAAVPSEIA 329
Query: 144 SSLVKALQQQAEGFWVEFQHLAEVQEELV--AVPEQLQGLMGDYSELFSTIWDLPPSRVQ 201
+ + L + +G + L E ++ V +P +Q L+ D+ ++F LPP RV
Sbjct: 330 AVTLPQLLKWHKGCEIWAAALLEHSQDDVQHVIPSSVQQLLTDFDDVFHDPKCLPPHRVF 389
Query: 202 DHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDG 261
DHAI L A N RPYR QK EIEK VKEM+D G+ PS+S ++S V+LVKKKDG
Sbjct: 390 DHAISLVPDATPVNYRPYRYSPLQKDEIEKQVKEMIDAGLITPSMSPYASPVLLVKKKDG 449
Query: 262 SWRFCIDYRAFHKITIPNKFPILIIDELLYKLGEAK 297
SWRFC+DYR + +TI +KFP+ ++DELL +L K
Sbjct: 450 SWRFCVDYRRLNALTIKSKFPMPVVDELLDELSGTK 485
>ref|XP_471847.1| OSJNBb0062H02.17 [Oryza sativa (japonica cultivar-group)]
gi|38347666|emb|CAE05600.2| OSJNBa0054D14.1 [Oryza
sativa (japonica cultivar-group)]
gi|38346992|emb|CAD40278.2| OSJNBb0062H02.17 [Oryza
sativa (japonica cultivar-group)]
Length = 1629
Score = 158 bits (400), Expect = 4e-37
Identities = 107/304 (35%), Positives = 168/304 (55%), Gaps = 11/304 (3%)
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVETPYMVEL 60
+SL+++ + +++ G++ +++LVD G S +FIS+ + SS V+E P V++
Sbjct: 464 VSLQAVQGTETGGCMRMLGQIQGKEILILVDSGSSASFISKRVASSL-MGVLEQPVHVQV 522
Query: 61 --EDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEEL 118
G K+ C + + + +IQ + L D++LG+ W + ++
Sbjct: 523 MVAGGAKLHCCSEILNCEWTIQGHVFFTNLKVLELNNYDMILGMDWLMQHSPMTVDWTTK 582
Query: 119 SLRF-HVGEKVLLRGELGLIRKNASLSS--LVKALQQQAEGFWVEFQHL--AEVQEELVA 173
SL + G ++ L G + A +SS L + + A V+F + E QE+
Sbjct: 583 SLIIAYAGTQIQLYGVRSDTEQCAHISSKQLRELNDRTAVSNLVQFCSVFALEYQEQ--- 639
Query: 174 VPEQLQGLMGDYSELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLV 233
+PE +Q ++ ++S +F LPP R DH I L GA N+RPYR QK+EIE V
Sbjct: 640 IPEVVQTVLTEFSSVFDEPKGLPPIRQFDHTIPLLPGAGPVNVRPYRYTPIQKNEIESQV 699
Query: 234 KEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKL 293
+EML GI +PS S FSS V+LVKKKDGSWRFC+DYR + IT+ NK+P+ +IDELL +L
Sbjct: 700 QEMLSKGIIQPSSSPFSSPVLLVKKKDGSWRFCVDYRHLNAITVKNKYPLPVIDELLDEL 759
Query: 294 GEAK 297
A+
Sbjct: 760 AGAQ 763
Score = 35.0 bits (79), Expect = 6.1
Identities = 20/54 (37%), Positives = 28/54 (51%)
Query: 489 GVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFIIR 542
G VL+Q PLAF S + + S YE+E +A++ A+ W L F IR
Sbjct: 987 GAVLMQNNHPLAFLSRALGLRHPGLSTYEKESLAIMLAVDHWRPYLQHDEFFIR 1040
>emb|CAE05537.2| OSJNBa0053B21.11 [Oryza sativa (japonica cultivar-group)]
gi|50923861|ref|XP_472291.1| OSJNBa0053B21.11 [Oryza
sativa (japonica cultivar-group)]
Length = 1251
Score = 158 bits (400), Expect = 4e-37
Identities = 91/281 (32%), Positives = 154/281 (54%), Gaps = 13/281 (4%)
Query: 25 RVIVLVDCGVSHNFISQELMSSAHFPVVETPYM-VELEDGHKVECQGKCRDVQLSIQHIP 83
R + LVD G +++F++ + + ++E + + G + K + ++Q P
Sbjct: 327 RAVALVDSGSTNSFMNYDFAIKSGCEILEAGNRRILVAGGGMISSSTKTEQLSYTVQGHP 386
Query: 84 IEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELSL-------RFHVGEKVLLRGELGL 136
E+ F L L G DV+LG W I + ++ L + + L +G++ +
Sbjct: 387 FERVFQLIPLKGFDVILGADWIYDHSPISLDLKQRVLWVTCKGQQVKFVDHTLPQGKIMI 446
Query: 137 IRKNASLSSLVKALQQQAEGFWVEFQHLAEVQEELVAVPEQLQGLMGDYSELFSTIWDLP 196
K L + GF++E E ++E +PEQ+ L+ +++++F+ LP
Sbjct: 447 -----KAGKFGKMLDKGIMGFFIEVLSKTEEKKEETIIPEQIAVLLQEFADVFADPKGLP 501
Query: 197 PSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILV 256
P R DHAI L+ GA P++RPYR PH QK+ +E+++KE+LD RPS+S +SS ++V
Sbjct: 502 PKRDCDHAIPLKVGAEPPSLRPYRVPHQQKNAMEEIIKELLDSKEIRPSLSPYSSPAVMV 561
Query: 257 KKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKLGEAK 297
+KKDG+WR C+DYR + T+ NKFP+ II++LL +L AK
Sbjct: 562 RKKDGNWRMCVDYRQLNAKTVKNKFPMPIIEDLLDELNGAK 602
>dbj|BAB03109.1| retroelement pol polyprotein [Arabidopsis thaliana]
gi|12322008|gb|AAG51046.1| gypsy/Ty-3 retroelement
polyprotein; 69905-74404 [Arabidopsis thaliana]
Length = 1499
Score = 157 bits (396), Expect = 1e-36
Identities = 97/309 (31%), Positives = 165/309 (53%), Gaps = 13/309 (4%)
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVETPYM-VE 59
+S+ +++ ++++V G + + +L+D G +HNF+ + V V
Sbjct: 376 ISVNAVSGIAGYKTMRVKGTYDKKIIFILIDSGSTHNFLDPNTAAKLGCKVDTAGLTRVS 435
Query: 60 LEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELS 119
+ DG K+ +GK D +Q + + L L G+D+VLG+ W +LG I F++L
Sbjct: 436 VADGRKLRVEGKVTDFSWKLQTTTFQSDILLIPLQGIDMVLGVQWLETLGRISWEFKKLE 495
Query: 120 LRFHVG-EKVLLRGELGLIRKNASLSSLVKALQQQAEGFWVEFQHLAEVQE-ELVAV--- 174
+RF +KVLL G + L K + Q + + Q ++E E EL +
Sbjct: 496 MRFKFNNQKVLLHGLTSGSVREVKAQKLQKLQEDQVQLAMLCVQEVSESTEGELCTINAL 555
Query: 175 ------PEQLQGLMGDYSELFSTIWDLPPSRVQ-DHAIHLQEGAAIPNIRPYRCPHYQKS 227
++ ++ +Y ++F LPP R + +H I L EG+ N RPYR +QK+
Sbjct: 556 TSELGEESVVEEVLNEYPDIFIEPTALPPFREKHNHKIKLLEGSNPVNQRPYRYSIHQKN 615
Query: 228 EIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIID 287
EI+KLV+++L G + S S ++S V+LVKKKDG+WR C+DYR + +T+ + FPI +I+
Sbjct: 616 EIDKLVEDLLTNGTVQASSSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDSFPIPLIE 675
Query: 288 ELLYKLGEA 296
+L+ +LG A
Sbjct: 676 DLMDELGGA 684
Score = 47.8 bits (112), Expect = 0.001
Identities = 25/67 (37%), Positives = 40/67 (59%)
Query: 476 QSLIFNNFCY*ETGVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLF 535
Q ++ + C G VL+Q PLA+ S + + S+YE+EL+AV+ A+++W L
Sbjct: 895 QFVVETDACGQGIGAVLMQEGHPLAYISRQLKGKQLHLSIYEKELLAVIFAVRKWRHYLL 954
Query: 536 GSHFIIR 542
SHFII+
Sbjct: 955 QSHFIIK 961
>gb|AAV24812.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1475
Score = 156 bits (394), Expect = 2e-36
Identities = 92/273 (33%), Positives = 147/273 (53%), Gaps = 4/273 (1%)
Query: 29 LVDCGVSHNFISQELMSSAHFPVVETPYM-VELEDGHKVECQGKCRDVQLSIQHIPIEQE 87
LVD G + F+ Q+ ++P+ T V + G +++ D+ IQ +
Sbjct: 384 LVDSGSTTTFMDQDYALRNYYPLKNTDTKKVVVAGGGELKTDVMVPDISYEIQGECFTNQ 443
Query: 88 FFLFNLGGLDVVLGL*WFASLGHIRANFEELSLRFHVGEKVLLRGELGLIRKNASLSS-- 145
F L L G D++LG W + I + ++ L G KV+L + K+ +S
Sbjct: 444 FKLLPLKGYDIILGADWIYNYSPISLDLKQRILGITKGNKVILLQDFTKPNKHFQISGKR 503
Query: 146 LVKALQQQAEGFWVEFQHLAE-VQEELVAVPEQLQGLMGDYSELFSTIWDLPPSRVQDHA 204
L K L++ A G ++ ++E V+EE +PE + ++ + + LPP R DH
Sbjct: 504 LEKMLKKGALGMVIQVNVMSETVEEEGHVIPEDISDIIQQFPAVLKEPKGLPPKRECDHV 563
Query: 205 IHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWR 264
I+LQ GA PNIRPYR PHYQK +E ++ E+++ + S S +SS ++V+KKDGSWR
Sbjct: 564 INLQSGAVPPNIRPYRVPHYQKEAMENIINELIESKEIQTSDSPYSSPAVMVRKKDGSWR 623
Query: 265 FCIDYRAFHKITIPNKFPILIIDELLYKLGEAK 297
C+DYR + T+ NKFP+ II++LL +L A+
Sbjct: 624 MCVDYRQLNAQTVKNKFPMPIIEDLLDELNGAR 656
Score = 48.1 bits (113), Expect = 7e-04
Identities = 21/54 (38%), Positives = 36/54 (65%)
Query: 489 GVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFIIR 542
G VL+Q RP+AF+S + + +S+YE+E MA++ A+++W + GS II+
Sbjct: 879 GAVLMQGGRPIAFFSKALGPKAAGQSIYEKEAMAILEALKKWRHYVLGSKLIIK 932
>gb|AAP54595.1| putative gag-pol polyprotein [Oryza sativa (japonica
cultivar-group)] gi|37536012|ref|NP_922308.1| putative
gag-pol polyprotein [Oryza sativa (japonica
cultivar-group)] gi|10140673|gb|AAG13508.1| putative
gag-pol polyprotein [Oryza sativa (japonica
cultivar-group)]
Length = 1608
Score = 155 bits (392), Expect = 3e-36
Identities = 103/306 (33%), Positives = 161/306 (51%), Gaps = 16/306 (5%)
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSS-AHFPVVETPYMVE 59
+S +++ S +S+++ G + +++LVD G +H+FI +L + + V+
Sbjct: 425 ISFQALNGTDSSKSIRLRGWVQNTELLMLVDSGSTHSFIDAKLGAQLCGLQKLNQAIKVQ 484
Query: 60 LEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELS 119
+ DG ++ C + Q +F L LG D +LG+ W ++ ++
Sbjct: 485 VADGSQLFCDSFLPNCSWWSQGHSFTSDFRLLPLGSYDAILGMDWLEQFSPMQVDWVHKW 544
Query: 120 LRF-HVGEKVLLRGELGLIRKNASLSSLVKALQQQAEGFWVE-----FQHLAEVQEELVA 173
+ F H G+ V L+G + LS+ Q +G + HL V E L A
Sbjct: 545 IAFQHHGQAVQLQGI------HPQLSTCFPISNDQLQGMSKKGAVMCLVHL-NVAETLTA 597
Query: 174 --VPEQLQGLMGDYSELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEK 231
VPE +Q ++ ++ E+FS +LPP R DH I L EGA N+RPYR K EIE+
Sbjct: 598 TTVPEIVQPILNEFQEIFSEPTELPPKRNCDHHIPLVEGAKPVNLRPYRYKPALKDEIER 657
Query: 232 LVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLY 291
V EML G+ +PS S FSS +LVKKKDG+WR CIDYR + +T+ +K+P+ +IDELL
Sbjct: 658 QVAEMLRSGVIQPSSSPFSSPALLVKKKDGTWRLCIDYRQLNDVTVKSKYPVPVIDELLD 717
Query: 292 KLGEAK 297
+L +K
Sbjct: 718 ELAGSK 723
Score = 36.6 bits (83), Expect = 2.1
Identities = 19/53 (35%), Positives = 29/53 (53%)
Query: 489 GVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFII 541
G VL Q P+A+ S + + + S YE+E MA++ A++ W L FII
Sbjct: 947 GAVLSQEKHPIAYLSRALGPKTRGLSTYEKEYMAIILAVEHWRPYLQQGEFII 999
>gb|AAV32203.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 434
Score = 155 bits (391), Expect = 4e-36
Identities = 101/286 (35%), Positives = 158/286 (54%), Gaps = 11/286 (3%)
Query: 19 GELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVETPYMVELE--DGHKVECQGKCRDVQ 76
G++ L +++LVD G S +F++ + + V P + ++ +G+ + C + Q
Sbjct: 3 GQIQGLEILILVDSGSSVSFLNSTIADNLK-NVQRKPQQISVQVANGNILHCTSAITNCQ 61
Query: 77 LSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELSLRFHV-GEKVLLRGELG 135
S Q + +L D++LG+ W + N+ L + G+ +LL+G
Sbjct: 62 WSTQGYVFNTTLRVLSLNSYDLILGMDWLTEHSPMTVNWATKQLTVSLKGQFILLQGINS 121
Query: 136 LIRK-NASLSSLVKALQ-QQAEGFWVEF--QHLAEVQEELVAVPEQLQGLMGDYSELFST 191
+ + N L +K LQ +QA V+ H + +++E + P+ +Q ++ ++S +F
Sbjct: 122 QVSQCNLLLGHQMKQLQARQAVAHLVQLCVSH-SNIEQEFI--PDSVQQVLQEFSGIFDE 178
Query: 192 IWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSS 251
LPPSR DH I L GA ++RPYR QK EIE V EMLD GI +PS S FSS
Sbjct: 179 PSGLPPSRSFDHTIPLLPGAQPVSVRPYRYTPAQKDEIESEVTEMLDKGIIQPSASPFSS 238
Query: 252 LVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKLGEAK 297
V+LVKKKDG+WRFC+DYR + IT+ NK+P+ IIDELL +L A+
Sbjct: 239 PVLLVKKKDGTWRFCVDYRHLNAITVKNKYPLPIIDELLDELSGAQ 284
>gb|AAP54660.1| putative plant disease resistance polyprotein [Oryza sativa
(japonica cultivar-group)] gi|37536142|ref|NP_922373.1|
putative plant disease resistance polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|10122034|gb|AAG13423.1| putative plant disease
resistance polyprotein [Oryza sativa (japonica
cultivar-group)]
Length = 894
Score = 154 bits (388), Expect = 9e-36
Identities = 97/281 (34%), Positives = 140/281 (49%), Gaps = 39/281 (13%)
Query: 26 VIVLVDCGVSHNFISQELMSSAHFPVVETP-YMVELEDGHKVECQGKCRDVQLSIQHIPI 84
+++LVD G SH+FI P+ P +V++ +G + CQ IQ
Sbjct: 404 LLLLVDYGSSHSFIDYNFAQMLGLPLKPIPSTLVKVANGDCLPCQYVVPGFSWWIQGHTF 463
Query: 85 EQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELSLRFHVGEKVLLRGELGLIRKNASLS 144
+ + LG D +LG+ W G + ++ +L+F
Sbjct: 464 TYDMRVVTLGVHDAILGMDWLEQWGEMSCHWANKTLKF---------------------- 501
Query: 145 SLVKALQQQAEGFWVEFQHL--AEVQEELVA------VPEQLQGLMGDYSELFSTIWDLP 196
Q +G W+ Q + A V E+ A +P +Q L+ D+ ++F LP
Sbjct: 502 --------QYQGSWISLQGVQDAAVPSEIAADDVQHVIPSSVQQLLTDFDDVFHDPKCLP 553
Query: 197 PSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILV 256
P RV DHAI L A N RPYR QK EIEK VKEM+D G+ PS+S ++S V+LV
Sbjct: 554 PHRVFDHAIFLVPDATPVNCRPYRYSPLQKDEIEKQVKEMIDAGLITPSMSPYASPVLLV 613
Query: 257 KKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKLGEAK 297
KKKDGSWRFC+DYR + +TI +KFP+ ++DELL +L K
Sbjct: 614 KKKDGSWRFCVDYRRLNALTIKSKFPMPVVDELLDELSGTK 654
>gb|AAX95226.1| retrotransposon protein, putative, unclassified [Oryza sativa
(japonica cultivar-group)]
Length = 1513
Score = 154 bits (388), Expect = 9e-36
Identities = 91/278 (32%), Positives = 149/278 (52%), Gaps = 5/278 (1%)
Query: 25 RVIVLVDCGVSHNFISQELMSSAHFPVVETPYM-VELEDGHKVECQGKCRDVQLSIQHIP 83
R + LVD G F+ QE P+V T V + G +++ + + D+ IQ
Sbjct: 449 RAVGLVDSGSPSTFMDQEYAIRNDCPLVSTDSKKVVVAGGGELKSEVQVPDILYQIQGET 508
Query: 84 IEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELSLRFHVGEKVLLRGELGLIRKNASL 143
+F L L G D++LG W I + ++ L GEKV+ + K+ +
Sbjct: 509 FSNKFNLIPLKGYDIILGADWIYKYSPITLDLKKRELGITKGEKVITLQDFTNTGKHLLV 568
Query: 144 SS--LVKALQQQAEGFWVEFQHLAEVQ--EELVAVPEQLQGLMGDYSELFSTIWDLPPSR 199
S + K +++ G+ + +AE EE +++P+ +Q ++ + + LPP R
Sbjct: 569 DSSKVDKIIRKGGLGYLFQITQVAEESDVEESISIPDDIQDILQRFPTVLEKPKGLPPQR 628
Query: 200 VQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQFSSLVILVKKK 259
DH I L++GA PN+RPYR PH+QK +EK++ E+++ + S S +SS ++V+KK
Sbjct: 629 GCDHVITLKDGAIPPNLRPYRVPHHQKEAMEKIIAELIESKEIQVSNSPYSSPAVMVRKK 688
Query: 260 DGSWRFCIDYRAFHKITIPNKFPILIIDELLYKLGEAK 297
DGSWR C+DYR + T+ NKFP+ II++LL +L AK
Sbjct: 689 DGSWRLCVDYRQLNAQTVKNKFPMPIIEDLLDELNGAK 726
Score = 47.8 bits (112), Expect = 0.001
Identities = 20/54 (37%), Positives = 35/54 (64%)
Query: 489 GVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFIIR 542
G VL+Q RP+AF+S + + +S+YE+E MA++ A+++W GS +I+
Sbjct: 949 GAVLMQGDRPIAFYSKSLGPKAAAQSIYEKEAMAILEALKKWRHYFLGSKLVIK 1002
>ref|XP_471551.1| OSJNBa0019J05.12 [Oryza sativa (japonica cultivar-group)]
gi|38346427|emb|CAD40214.2| OSJNBa0019J05.12 [Oryza
sativa (japonica cultivar-group)]
Length = 1817
Score = 148 bits (374), Expect = 4e-34
Identities = 95/301 (31%), Positives = 161/301 (52%), Gaps = 6/301 (1%)
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQEL-MSSAHFPVVETPYMVE 59
+S ++I S S ++ G + +++LVD G +H+F+ + + + A + + V+
Sbjct: 583 ISQQAIWGTESPNSFRLRGWVQGTELLMLVDSGSTHSFMEESIGLKLAGVKPLRSKLSVK 642
Query: 60 LEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELS 119
L DG + C + + + +Q F L NLGG D++LG+ W ++ N+ +
Sbjct: 643 LADGETLSCSYEVTNCRWWMQGHNFINNFRLLNLGGYDIILGMDWLEQFSPMQVNWVDKW 702
Query: 120 LRFHV-GEKVLLRGELGLIRKNASLSS--LVKALQQQAEGFWVEFQHLAEVQEELVAVPE 176
+ + G+ V L+G + + + +S+ + ++Q + + V+ V +E +P
Sbjct: 703 MDITISGQPVRLQGIVSQVSHCSPISTEQCLGMVRQGSLMYLVQLTSSDPVTKE--PIPA 760
Query: 177 QLQGLMGDYSELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEM 236
+Q L+ + +F+ LPP R DH I L GAA N+RPYR K EIE+ + +M
Sbjct: 761 AVQQLIDSFQAVFAEPNGLPPKRYCDHTIPLVPGAAPVNLRPYRFNPALKDEIEQQISDM 820
Query: 237 LDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKLGEA 296
L G+ +PS S FSS +LVKKKDG+WR IDYR + IT+ +P+ +IDELL +L A
Sbjct: 821 LKSGVIQPSHSAFSSPALLVKKKDGTWRLVIDYRKLNAITVKGTYPMPVIDELLDELAHA 880
Query: 297 K 297
K
Sbjct: 881 K 881
>gb|AAG51464.1| gypsy/Ty3 element polyprotein, putative [Arabidopsis thaliana]
gi|25403233|pir||G86474 probable protein gypsy/Ty3
element polyprotein [imported] - Arabidopsis thaliana
Length = 1447
Score = 148 bits (373), Expect = 5e-34
Identities = 92/307 (29%), Positives = 167/307 (53%), Gaps = 13/307 (4%)
Query: 1 LSLKSIAEFTSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAHFPVVETPYM-VE 59
+S+ +++ + +++ V G + + + +L+D G +HNFI + + V V
Sbjct: 355 ISVNAVSGISGYKTMGVKGTVDKRDLFILIDSGSTHNFIDSTVAAKLGCHVESAGLTKVA 414
Query: 60 LEDGHKVECQGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELS 119
+ DG K+ G+ + +Q + + L L G+D+VLG+ W +LG I F++L
Sbjct: 415 VADGRKLNVDGQIKGFTWKLQSTTFQSDILLIPLQGVDMVLGVQWLETLGRISWEFKKLE 474
Query: 120 LRF-HVGEKVLLRGELGLIRKNASLSSLVKALQQQAEGFWVEFQHLAEVQEELVA----- 173
++F + ++V L G + ++ L K Q + V + + +E+ +
Sbjct: 475 MQFFYKNQRVWLHGIITGSVRDIKAHKLQKTQADQIQLAMVCVREVVSDEEQEIGSISAL 534
Query: 174 ---VPEQ--LQGLMGDYSELFSTIWDLPPSRVQ-DHAIHLQEGAAIPNIRPYRCPHYQKS 227
V E+ +Q ++ ++ ++F+ DLPP R + DH I L EGA N RPYR +QK
Sbjct: 535 TSDVVEESVVQNIVEEFPDVFAEPTDLPPFREKHDHKIKLLEGANPVNQRPYRYVVHQKD 594
Query: 228 EIEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIID 287
EI+K+V++M+ G + S S F+S V+LVKKKDG+WR C+DY + +T+ ++F I +I+
Sbjct: 595 EIDKIVQDMIKSGTIQVSSSPFASPVVLVKKKDGTWRLCVDYTELNGMTVKDRFLIPLIE 654
Query: 288 ELLYKLG 294
+L+ +LG
Sbjct: 655 DLMDELG 661
Score = 43.1 bits (100), Expect = 0.022
Identities = 23/67 (34%), Positives = 38/67 (56%)
Query: 476 QSLIFNNFCY*ETGVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLF 535
Q ++ + C VL+Q PLA+ S + + S+YE+EL+A + A+++W L
Sbjct: 866 QFMVETDACGQGIRAVLMQKGHPLAYISRQLKGKQLHLSIYEKELLAFIFAVRKWRHYLL 925
Query: 536 GSHFIIR 542
SHFII+
Sbjct: 926 PSHFIIK 932
>gb|AAF79797.1| T32E20.30 [Arabidopsis thaliana]
Length = 1397
Score = 147 bits (371), Expect = 8e-34
Identities = 97/245 (39%), Positives = 128/245 (51%), Gaps = 33/245 (13%)
Query: 56 YMVELEDGHKVECQGKCRDVQLSI-----QHIPIEQEFFLFNLGGLDVVLGL*WFASLGH 110
+ + L G V+ G C V +++ Q + + F +LG +DV+LG+ W +LG
Sbjct: 390 FHIRLGTGITVQGLGLCDKVTMTLPVGCGQELELTTHFITLDLGPVDVILGIAWLRTLGD 449
Query: 111 IRANFE--ELSLRFHVGEKVLLRGELGLIRKNASLSSLVKALQQQAEGFWVEFQHLAEVQ 168
+ N+E ELS +H G V LRG+ L SL S + Q
Sbjct: 450 CKVNWERHELSFLYH-GRTVTLRGDPELDTFKMSLKSFSTKFRLQ--------------N 494
Query: 169 EELVAVPEQLQGLMGDYSELFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSE 228
+EL Q L G LPP + +HAI L G ++RPYR PH K
Sbjct: 495 KELEVSLNSHQNLKG-----------LPPIKGNEHAISLLPGTRAISVRPYRYPHAHKEA 543
Query: 229 IEKLVKEMLDVGIFRPSISQFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDE 288
+E LV EMLD GI R S S FSS V+LVKKKD SWRFC+DYRA ++ TIPNKFPI +ID+
Sbjct: 544 MEGLVSEMLDNGIIRASKSPFSSPVLLVKKKDQSWRFCVDYRALNRATIPNKFPIPMIDQ 603
Query: 289 LLYKL 293
LL +L
Sbjct: 604 LLDEL 608
Score = 38.9 bits (89), Expect = 0.42
Identities = 17/36 (47%), Positives = 25/36 (69%)
Query: 506 ISDQGQRKSVYERELMAVVRAIQRW*RDLFGSHFII 541
++ + Q K VYERELMA+V +IQ+W L G F++
Sbjct: 813 LTSKEQLKPVYERELMAIVLSIQKWKHYLMGRRFVL 848
>gb|AAL68642.1| polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 677
Score = 147 bits (370), Expect = 1e-33
Identities = 89/285 (31%), Positives = 149/285 (52%), Gaps = 2/285 (0%)
Query: 11 SRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAH-FPVVETPYMVELEDGHKVECQ 69
S S +V G + ++++L+D G SH+F+ + ++ + + P V++ G + C
Sbjct: 29 STHSFRVKGLIQGTKLLMLIDSGSSHSFLDENIVQTMQGVTSLPQPVKVKVASGEVLICD 88
Query: 70 GKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELSLRFHV-GEKV 128
+ D +Q F L +L G D +LG+ W LG +R + + L + + G V
Sbjct: 89 KQLPDYAWWLQGRCYRTNFKLLSLPGYDAILGIDWLQGLGVMRIIWVQKWLEYEINGSPV 148
Query: 129 LLRGELGLIRKNASLSSLVKALQQQAEGFWVEFQHLAEVQEELVAVPEQLQGLMGDYSEL 188
++G + + +S + FQ +E +P +Q +M +++ +
Sbjct: 149 RIQGVMPKVDACPLISGDKLMGLCKMGALMHLFQITSETDIHKPEIPIMIQEVMNEFNSV 208
Query: 189 FSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVGIFRPSISQ 248
F LPP R DH+I L G+ N+RPY QK EIE+ V++ML G+ PS+S
Sbjct: 209 FEEPQGLPPRRRCDHSIPLIPGSKPVNLRPYGYNPAQKDEIEQQVRDMLKSGVIHPSVSP 268
Query: 249 FSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKL 293
+SSL +LV KKDG+WR C+D+R + ITI +K+P+ +IDELL +L
Sbjct: 269 YSSLALLVNKKDGTWRLCVDHRLLNAITIKSKYPLPVIDELLDEL 313
>gb|AAL76001.1| putative gag-pol polyprotein [Zea mays]
Length = 2396
Score = 147 bits (370), Expect = 1e-33
Identities = 92/290 (31%), Positives = 148/290 (50%), Gaps = 16/290 (5%)
Query: 10 TSRRSLKVWGELGQLRVIVLVDCGVSHNFISQELMSSAH-FPVVETPYMVELEDGHKVEC 68
T R++LK+ G + +++L+D G SH F++ +L + + V++ +G V C
Sbjct: 450 TGRQTLKLNGSIQNHPLLILIDSGSSHTFLNDQLRPHLQGVTSMASTLQVQVANGAMVTC 509
Query: 69 QGKCRDVQLSIQHIPIEQEFFLFNLGGLDVVLGL*WFASLGHIRANFEELSLRF-HVGEK 127
K Q IQ+ + L D+V+G+ W S +R ++ + L + G
Sbjct: 510 HYKLLQAQWQIQNCSFTSDVSFLPLPYYDMVVGMDWLESFSPMRVDWAQKWLIIPYQGSS 569
Query: 128 VLLRGELGLIRKNASLSSLVKALQQQAEGFWVEFQHLAEVQEELVAVPEQLQGLMGDYSE 187
VLL+G + + + L F A +Q L+ +S
Sbjct: 570 VLLQGNTAGVPADTVIELL--------------FMESASSVSSSPDSHPAIQALLQQFSS 615
Query: 188 LFSTIWDLPPSRVQDHAIHLQEGAAIPNIRPYRCPHYQKSEIEKLVKEMLDVGIFRPSIS 247
+F+ LPPSR DHAI L EGA ++RPYR P K +IEK V+EML G+ + S S
Sbjct: 616 VFAEPQGLPPSRDCDHAIPLVEGAQPVSVRPYRYPPALKDKIEKQVQEMLHQGVIQKSNS 675
Query: 248 QFSSLVILVKKKDGSWRFCIDYRAFHKITIPNKFPILIIDELLYKLGEAK 297
F+S V+LVKKKD +WRFC+DYR + +T+ +K+P+ + D+L+ +L +K
Sbjct: 676 SFASPVLLVKKKDMTWRFCVDYRYLNALTLKSKYPVPVFDQLIDELAHSK 725
Score = 43.1 bits (100), Expect = 0.022
Identities = 25/63 (39%), Positives = 34/63 (53%)
Query: 479 IFNNFCY*ETGVVLLQV*RPLAFWSSIISDQGQRKSVYERELMAVVRAIQRW*RDLFGSH 538
I + Y G VL+Q PLAF S + + Q S YE+E MA++ AI +W L +
Sbjct: 939 IHTDASYYGVGAVLMQSGHPLAFLSKALGPKNQGLSTYEKEYMAIILAIAQWRSYLQLAE 998
Query: 539 FII 541
FII
Sbjct: 999 FII 1001
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.349 0.158 0.543
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 789,004,805
Number of Sequences: 2540612
Number of extensions: 29789877
Number of successful extensions: 146790
Number of sequences better than 10.0: 1578
Number of HSP's better than 10.0 without gapping: 1143
Number of HSP's successfully gapped in prelim test: 436
Number of HSP's that attempted gapping in prelim test: 143646
Number of HSP's gapped (non-prelim): 2764
length of query: 542
length of database: 863,360,394
effective HSP length: 133
effective length of query: 409
effective length of database: 525,458,998
effective search space: 214912730182
effective search space used: 214912730182
T: 11
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.8 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0126.7


