

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0122.7
(125 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
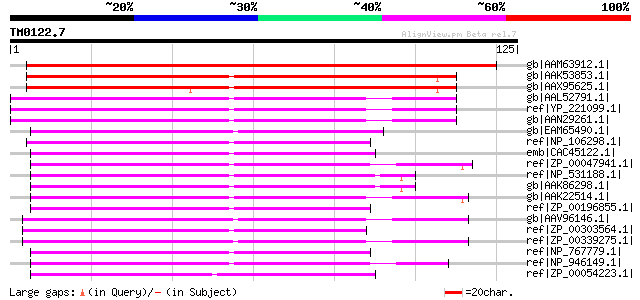
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM63912.1| unknown [Arabidopsis thaliana] gi|30679001|ref|NP... 158 3e-38
gb|AAK53853.1| Hypothetical protein [Oryza sativa] 134 7e-31
gb|AAX95625.1| Protein of unknown function (DUF952) [Oryza sativ... 126 1e-28
gb|AAL52791.1| Hypothetical Cytosolic Protein [Brucella melitens... 67 1e-10
ref|YP_221099.1| hypothetical protein BruAb1_0338 [Brucella abor... 66 2e-10
gb|AAN29261.1| conserved hypothetical protein [Brucella suis 133... 65 3e-10
gb|EAM65490.1| Protein of unknown function DUF952 [Jannaschia sp... 65 4e-10
ref|NP_106298.1| hypothetical protein mll5682 [Mesorhizobium lot... 64 6e-10
emb|CAC45122.1| CONSERVED HYPOTHETICAL PROTEIN [Sinorhizobium me... 64 6e-10
ref|ZP_00047941.1| COG3502: Uncharacterized protein conserved in... 62 3e-09
ref|NP_531188.1| hypothetical protein Atu0485 [Agrobacterium tum... 60 2e-08
gb|AAK86298.1| AGR_C_860p [Agrobacterium tumefaciens str. C58] g... 60 2e-08
gb|AAK22514.1| conserved hypothetical protein [Caulobacter cresc... 59 3e-08
ref|ZP_00196855.1| COG3502: Uncharacterized protein conserved in... 58 5e-08
gb|AAV96146.1| conserved hypothetical protein [Silicibacter pome... 58 6e-08
ref|ZP_00303564.1| COG3502: Uncharacterized protein conserved in... 57 1e-07
ref|ZP_00339275.1| COG3502: Uncharacterized protein conserved in... 55 4e-07
ref|NP_767779.1| hypothetical protein blr1139 [Bradyrhizobium ja... 55 5e-07
ref|NP_946149.1| hypothetical protein RPA0796 [Rhodopseudomonas ... 53 2e-06
ref|ZP_00054223.1| COG3502: Uncharacterized protein conserved in... 52 2e-06
>gb|AAM63912.1| unknown [Arabidopsis thaliana] gi|30679001|ref|NP_849284.1|
expressed protein [Arabidopsis thaliana]
Length = 122
Score = 158 bits (399), Expect = 3e-38
Identities = 72/116 (62%), Positives = 91/116 (78%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
F+YRIST +EWEE + NGS++G +DKS+ + HLSKLDQV+ TL+ F+++ KE LYLLQ+
Sbjct: 6 FIYRISTEQEWEEFKKNGSSYGAEIDKSTCYYHLSKLDQVQLTLKNFFVDVKEYLYLLQV 65
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSDGRFTCCLL 120
D KKLGDGL+YE VD NSFPHFYGP ++F PL LD V KAEKL+ +G FTC L
Sbjct: 66 DPKKLGDGLIYEAVDEVNSFPHFYGPDKTFVPLPLDSVVKAEKLTFINGNFTCSFL 121
>gb|AAK53853.1| Hypothetical protein [Oryza sativa]
Length = 144
Score = 134 bits (336), Expect = 7e-31
Identities = 67/107 (62%), Positives = 79/107 (73%), Gaps = 2/107 (1%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
FVYRISTA EW +LQ G T GG+LD+S+ IHLS L QVR TL+ F++ + +LYLLQ+
Sbjct: 9 FVYRISTADEWAQLQRTGGTLGGDLDRSTGCIHLSDLSQVRKTLKNFFLG-RNDLYLLQV 67
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTK-AEKLSL 110
D KL DGLVYE D SN FPHFYGP RSF+PL LD V K AEK+ L
Sbjct: 68 DTSKLSDGLVYEAADDSNYFPHFYGPGRSFAPLQLDAVIKEAEKIVL 114
>gb|AAX95625.1| Protein of unknown function (DUF952) [Oryza sativa (japonica
cultivar-group)] gi|50901024|ref|XP_462945.1|
hypothetical protein [Oryza sativa (japonica
cultivar-group)]
Length = 133
Score = 126 bits (316), Expect = 1e-28
Identities = 67/116 (57%), Positives = 79/116 (67%), Gaps = 11/116 (9%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQ---------VRSTLERFYMNC 55
FVYRISTA EW +LQ G T GG+LD+S+ IHLS L Q VR TL+ F++
Sbjct: 9 FVYRISTADEWAQLQRTGGTLGGDLDRSTGCIHLSDLSQGIHNVRKKQVRKTLKNFFLG- 67
Query: 56 KEELYLLQIDAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTK-AEKLSL 110
+ +LYLLQ+D KL DGLVYE D SN FPHFYGP RSF+PL LD V K AEK+ L
Sbjct: 68 RNDLYLLQVDTSKLSDGLVYEAADDSNYFPHFYGPGRSFAPLQLDAVIKEAEKIVL 123
>gb|AAL52791.1| Hypothetical Cytosolic Protein [Brucella melitensis 16M]
gi|17987893|ref|NP_540527.1| Hypothetical Cytosolic
Protein [Brucella melitensis 16M]
gi|25367931|pir||AD3453 hypothetical cytosolic protein
BMEI1610 [imported] - Brucella melitensis (strain 16M)
Length = 117
Score = 66.6 bits (161), Expect = 1e-10
Identities = 38/110 (34%), Positives = 58/110 (52%), Gaps = 7/110 (6%)
Query: 1 MSGIFVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELY 60
MS +Y+I+ + W + + GS G +D + ++IH S QVR+T + + + +L
Sbjct: 1 MSNKIIYKIAPRELWAQAEKAGSFAGAPVDIADSYIHFSTAAQVRATAAKHFAG-QSDLL 59
Query: 61 LLQIDAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSL 110
L+ +DA+ LG L YE+ G FPH Y +PL L V K E L L
Sbjct: 60 LVHVDAQALGQALKYEVSRGGALFPHLY------APLPLTAVVKVEPLPL 103
>ref|YP_221099.1| hypothetical protein BruAb1_0338 [Brucella abortus biovar 1 str.
9-941] gi|62195438|gb|AAX73738.1| conserved hypothetical
protein [Brucella abortus biovar 1 str. 9-941]
Length = 117
Score = 66.2 bits (160), Expect = 2e-10
Identities = 38/110 (34%), Positives = 57/110 (51%), Gaps = 7/110 (6%)
Query: 1 MSGIFVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELY 60
MS +Y+I+ + W + + GS G +D + +IH S QVR+T + + + +L
Sbjct: 1 MSNKIIYKIAPRELWAQAEKAGSFAGAPVDIADGYIHFSTAAQVRATAAKHFAG-QSDLL 59
Query: 61 LLQIDAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSL 110
L+ +DA+ LG L YE+ G FPH Y +PL L V K E L L
Sbjct: 60 LVHVDAQALGQALKYEVSRGGALFPHLY------APLPLTAVVKVEPLPL 103
>gb|AAN29261.1| conserved hypothetical protein [Brucella suis 1330]
gi|23501219|ref|NP_697346.1| hypothetical protein BR0312
[Brucella suis 1330]
Length = 117
Score = 65.5 bits (158), Expect = 3e-10
Identities = 38/110 (34%), Positives = 56/110 (50%), Gaps = 7/110 (6%)
Query: 1 MSGIFVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELY 60
MS +Y+I+ + W + GS G +D + +IH S QVR+T + + + +L
Sbjct: 1 MSNKIIYKIAPRELWARAEKAGSFAGAPVDIADGYIHFSTAAQVRATAAKHFAG-QSDLL 59
Query: 61 LLQIDAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSL 110
L+ +DA+ LG L YE+ G FPH Y +PL L V K E L L
Sbjct: 60 LVHVDAQALGQALKYEVSRGGALFPHLY------APLPLTAVVKVEPLPL 103
>gb|EAM65490.1| Protein of unknown function DUF952 [Jannaschia sp. CCS1]
Length = 111
Score = 65.1 bits (157), Expect = 4e-10
Identities = 31/87 (35%), Positives = 47/87 (53%), Gaps = 1/87 (1%)
Query: 6 VYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQID 65
+Y+I A+EW L+ G T G +D + ++H S Q T +++ + L L +D
Sbjct: 3 IYKILRAEEWAALEGAGETPGAPIDVADGYVHFSTAAQAAETAAKYFAGV-DGLVLAAMD 61
Query: 66 AKKLGDGLVYEMVDGSNSFPHFYGPSR 92
A LGD L +E+ G + FPH YGP R
Sbjct: 62 ANALGDALKWEVSRGGDKFPHLYGPLR 88
>ref|NP_106298.1| hypothetical protein mll5682 [Mesorhizobium loti MAFF303099]
gi|14025484|dbj|BAB52084.1| mll5682 [Mesorhizobium loti
MAFF303099]
Length = 116
Score = 64.3 bits (155), Expect = 6e-10
Identities = 33/85 (38%), Positives = 46/85 (53%), Gaps = 1/85 (1%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
F+Y+I+ W E +A+G +D + FIH S QVR T + + + +L L+ I
Sbjct: 4 FIYKITPQALWREAEASGRFTDAPIDVADGFIHFSTAAQVRETAAKHFSG-QTDLLLVAI 62
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYG 89
D LGDGL YE+ G FPH YG
Sbjct: 63 DDASLGDGLKYEISRGDALFPHLYG 87
>emb|CAC45122.1| CONSERVED HYPOTHETICAL PROTEIN [Sinorhizobium meliloti]
gi|15964303|ref|NP_384656.1| hypothetical protein
SMc02246 [Sinorhizobium meliloti 1021]
Length = 115
Score = 64.3 bits (155), Expect = 6e-10
Identities = 34/85 (40%), Positives = 47/85 (55%), Gaps = 1/85 (1%)
Query: 6 VYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQID 65
+Y+I A W+E + G+ G +D + FIH S DQV T R + +++L L+ +D
Sbjct: 5 IYKIVPALLWQEARRTGAFAGAPVDLADGFIHFSTGDQVVETAARHFEG-QDDLLLVAVD 63
Query: 66 AKKLGDGLVYEMVDGSNSFPHFYGP 90
A LGD LVYE G FPH Y P
Sbjct: 64 ATALGDKLVYEPSRGGALFPHLYAP 88
>ref|ZP_00047941.1| COG3502: Uncharacterized protein conserved in bacteria
[Magnetospirillum magnetotacticum MS-1]
Length = 120
Score = 62.0 bits (149), Expect = 3e-09
Identities = 39/110 (35%), Positives = 55/110 (49%), Gaps = 8/110 (7%)
Query: 6 VYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQID 65
VY+I W+E QA G G +D++ FIH S QV T R + +++L L+ ++
Sbjct: 4 VYKICPRPLWQEAQALGRFIGAPVDRADGFIHFSTAGQVAETAARHFAG-QDDLLLVAVE 62
Query: 66 AKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSL-SDGR 114
A+ LG+ L +E G FPH YG L L V L L +DGR
Sbjct: 63 AEALGEALRFEPSRGGAPFPHLYG------ELPLSAVRSVVDLPLGTDGR 106
>ref|NP_531188.1| hypothetical protein Atu0485 [Agrobacterium tumefaciens str. C58]
gi|17738832|gb|AAL41504.1| conserved hypothetical
protein [Agrobacterium tumefaciens str. C58]
gi|25367929|pir||AB2636 conserved hypothetical protein
Atu0485 [imported] - Agrobacterium tumefaciens (strain
C58, Dupont)
Length = 117
Score = 59.7 bits (143), Expect = 2e-08
Identities = 38/101 (37%), Positives = 49/101 (47%), Gaps = 8/101 (7%)
Query: 6 VYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQID 65
+Y+I W +A G G +D + FIH S QV T R + + +L L+ +D
Sbjct: 8 IYKIVPETLWTAARAEGMFEGAAIDLTDGFIHFSTAKQVAETAARHFSG-QVDLLLIAVD 66
Query: 66 AKKLGDGLVYEMVDGSNSFPHFYGPSRSFS------PLSLD 100
LGD LVYE G + FPH Y P FS PLSLD
Sbjct: 67 GAALGDKLVYEPSRGGDLFPHLYAP-LPFSAVLWETPLSLD 106
>gb|AAK86298.1| AGR_C_860p [Agrobacterium tumefaciens str. C58]
gi|25518878|pir||A97418 hypothetical protein AGR_C_860
[imported] - Agrobacterium tumefaciens (strain C58,
Cereon) gi|15887832|ref|NP_353513.1| hypothetical
protein AGR_C_860 [Agrobacterium tumefaciens str. C58]
Length = 244
Score = 59.7 bits (143), Expect = 2e-08
Identities = 38/101 (37%), Positives = 49/101 (47%), Gaps = 8/101 (7%)
Query: 6 VYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQID 65
+Y+I W +A G G +D + FIH S QV T R + + +L L+ +D
Sbjct: 135 IYKIVPETLWTAARAEGMFEGAAIDLTDGFIHFSTAKQVAETAARHFSG-QVDLLLIAVD 193
Query: 66 AKKLGDGLVYEMVDGSNSFPHFYGPSRSFS------PLSLD 100
LGD LVYE G + FPH Y P FS PLSLD
Sbjct: 194 GAALGDKLVYEPSRGGDLFPHLYAP-LPFSAVLWETPLSLD 233
>gb|AAK22514.1| conserved hypothetical protein [Caulobacter crescentus CB15]
gi|16124782|ref|NP_419346.1| hypothetical protein CC0527
[Caulobacter crescentus CB15] gi|25367928|pir||F87314
conserved hypothetical protein CC0527 [imported] -
Caulobacter crescentus
Length = 114
Score = 58.9 bits (141), Expect = 3e-08
Identities = 36/109 (33%), Positives = 57/109 (52%), Gaps = 8/109 (7%)
Query: 6 VYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQID 65
+Y+I + EW+ +A G G +D + FIHLS +Q + T +++ + L LL ++
Sbjct: 4 IYKILSRAEWDAAKAQGRFEGSAVDLADGFIHLSAGEQAQETAAKWFRG-QANLVLLAVE 62
Query: 66 AKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSL-SDG 113
A+ LG+ L +E G FPH Y PL + VT+ L L +DG
Sbjct: 63 AEPLGEDLKWEASRGGARFPHLY------RPLLVSEVTREADLDLDADG 105
>ref|ZP_00196855.1| COG3502: Uncharacterized protein conserved in bacteria
[Mesorhizobium sp. BNC1]
Length = 115
Score = 58.2 bits (139), Expect = 5e-08
Identities = 29/84 (34%), Positives = 43/84 (50%), Gaps = 1/84 (1%)
Query: 6 VYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQID 65
+Y+I A W E + G G +D FIH S Q+ T + + + +L L+ I+
Sbjct: 5 IYKICPASLWREAEERGFFIGAPVDLQDGFIHFSTGAQIEETAAKHFSG-ERDLLLIAIE 63
Query: 66 AKKLGDGLVYEMVDGSNSFPHFYG 89
+KLG L YE+ G + FPH YG
Sbjct: 64 EEKLGQELKYEVSRGGDLFPHLYG 87
>gb|AAV96146.1| conserved hypothetical protein [Silicibacter pomeroyi DSS-3]
gi|56697743|ref|YP_168113.1| hypothetical protein
SPO2905 [Silicibacter pomeroyi DSS-3]
Length = 112
Score = 57.8 bits (138), Expect = 6e-08
Identities = 35/110 (31%), Positives = 53/110 (47%), Gaps = 7/110 (6%)
Query: 4 IFVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQ 63
+ +Y+I A EW L+ +G + G +D + F+H S +Q T + + E L LL
Sbjct: 1 MLIYKIFRADEWAALERDGQSVGAPVDLADGFVHFSTAEQAGETAAKHFAGA-EGLVLLA 59
Query: 64 IDAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSDG 113
+A +GD L +E+ G FPH Y L L V ++ L L DG
Sbjct: 60 CEADAMGDVLKWEVSRGGALFPHLY------RELRLSDVLWSQPLPLVDG 103
>ref|ZP_00303564.1| COG3502: Uncharacterized protein conserved in bacteria
[Novosphingobium aromaticivorans DSM 12444]
Length = 117
Score = 57.0 bits (136), Expect = 1e-07
Identities = 26/85 (30%), Positives = 49/85 (57%), Gaps = 1/85 (1%)
Query: 4 IFVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQ 63
+ Y++ TA + EL+A G+ G +D + +IHLS Q+ T+++ + +++L+++
Sbjct: 6 LLAYKVLTASQMAELEAEGTFAGAPIDLTDGYIHLSTDAQLTETVDKHFSG-QDDLHVVA 64
Query: 64 IDAKKLGDGLVYEMVDGSNSFPHFY 88
+D LGD + +E G FPH Y
Sbjct: 65 VDLAVLGDAVKWEPSRGGQDFPHVY 89
>ref|ZP_00339275.1| COG3502: Uncharacterized protein conserved in bacteria
[Silicibacter sp. TM1040]
Length = 112
Score = 55.1 bits (131), Expect = 4e-07
Identities = 34/110 (30%), Positives = 52/110 (46%), Gaps = 7/110 (6%)
Query: 4 IFVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQ 63
+ +Y+I EW +A G T G +D + +IH S Q T + + E L+LL
Sbjct: 1 MLIYKIFRDDEWRAFEAAGETDGAPVDLADGYIHFSTAAQATETAAKHFAGA-EGLWLLA 59
Query: 64 IDAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSDG 113
+A ++GD L +E G FPH Y L+L + A+ L L +G
Sbjct: 60 FEADQMGDALKWEPSRGGALFPHLY------RRLTLSEMLWAKPLPLVEG 103
>ref|NP_767779.1| hypothetical protein blr1139 [Bradyrhizobium japonicum USDA 110]
gi|27349390|dbj|BAC46404.1| blr1139 [Bradyrhizobium
japonicum USDA 110]
Length = 115
Score = 54.7 bits (130), Expect = 5e-07
Identities = 29/84 (34%), Positives = 39/84 (45%), Gaps = 1/84 (1%)
Query: 6 VYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQID 65
+Y+I A W E + G G D FIH S QV TL + Y + L+L+++D
Sbjct: 5 IYKICPASAWREAERQGVYRGSADDARDGFIHFSTAAQVPETLRKHYFG-QRALFLVEVD 63
Query: 66 AKKLGDGLVYEMVDGSNSFPHFYG 89
LG L +E FPH YG
Sbjct: 64 GDALGSELRWERSRNDELFPHLYG 87
>ref|NP_946149.1| hypothetical protein RPA0796 [Rhodopseudomonas palustris CGA009]
gi|39647720|emb|CAE26240.1| conserved unknown protein
[Rhodopseudomonas palustris CGA009]
Length = 196
Score = 52.8 bits (125), Expect = 2e-06
Identities = 34/103 (33%), Positives = 46/103 (44%), Gaps = 7/103 (6%)
Query: 6 VYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQID 65
+Y+I A W + G G D FIH S Q+ TL + Y + L L+ +D
Sbjct: 86 IYKICDAAAWRGAEQAGRYGGSADDARDGFIHFSTAPQLAGTLAKHYAG-QTGLKLIAVD 144
Query: 66 AKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKL 108
+ LGD L +E G FPH YG L+L VT + L
Sbjct: 145 VEALGDRLRWEPSRGGELFPHLYG------ELALSAVTGVQDL 181
>ref|ZP_00054223.1| COG3502: Uncharacterized protein conserved in bacteria
[Magnetospirillum magnetotacticum MS-1]
Length = 117
Score = 52.4 bits (124), Expect = 2e-06
Identities = 25/85 (29%), Positives = 44/85 (51%), Gaps = 1/85 (1%)
Query: 6 VYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQID 65
+Y + EW+ Q G G + D++ FIH S QV++++ + + ++ L ++ +
Sbjct: 7 IYHVCRRSEWQYAQTTGVYGGSSQDEADGFIHFSGPTQVKASVAK-HRAGQDGLVIIAVA 65
Query: 66 AKKLGDGLVYEMVDGSNSFPHFYGP 90
LG L +E+ G FPH YGP
Sbjct: 66 TPSLGPALKWEVSRGGQLFPHLYGP 90
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.136 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 207,078,733
Number of Sequences: 2540612
Number of extensions: 7788960
Number of successful extensions: 15599
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 28
Number of HSP's successfully gapped in prelim test: 34
Number of HSP's that attempted gapping in prelim test: 15538
Number of HSP's gapped (non-prelim): 62
length of query: 125
length of database: 863,360,394
effective HSP length: 101
effective length of query: 24
effective length of database: 606,758,582
effective search space: 14562205968
effective search space used: 14562205968
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0122.7


