

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0121b.2
(253 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
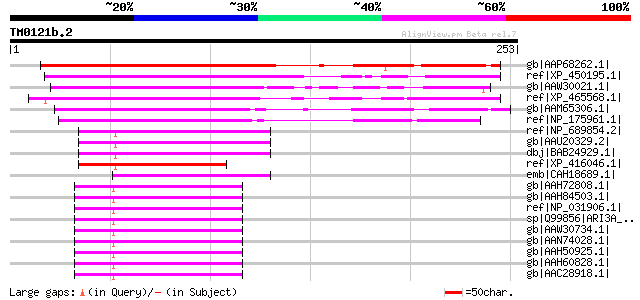
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP68262.1| At1g76110 [Arabidopsis thaliana] gi|20466328|gb|A... 207 2e-52
ref|XP_450195.1| glutathione S-transferase GST 16 - like protein... 169 7e-41
gb|AAW30021.1| At1g04880 [Arabidopsis thaliana] gi|56236040|gb|A... 140 3e-32
ref|XP_465568.1| glutathione S-transferase GST16-like protein [O... 112 1e-23
gb|AAM65306.1| unknown [Arabidopsis thaliana] gi|22137084|gb|AAM... 112 1e-23
ref|NP_175961.1| high mobility group (HMG1/2) family protein / A... 108 1e-22
ref|NP_689854.2| AT rich interactive domain 2 (ARID, RFX-like) [... 85 2e-15
gb|AAU20329.2| ARID2 [Homo sapiens] 85 2e-15
dbj|BAB24929.1| unnamed protein product [Mus musculus] 85 2e-15
ref|XP_416046.1| PREDICTED: similar to hypothetical protein [Gal... 75 2e-12
emb|CAH18689.1| hypothetical protein [Homo sapiens] 70 7e-11
gb|AAH72808.1| MGC80148 protein [Xenopus laevis] gi|58429893|gb|... 60 4e-08
gb|AAH84503.1| Hypothetical LOC496519 [Xenopus tropicalis] gi|58... 60 4e-08
ref|NP_031906.1| dead ringer homolog 1 [Mus musculus] gi|1401348... 59 1e-07
sp|Q99856|ARI3A_HUMAN AT-rich interactive domain-containing prot... 59 1e-07
gb|AAW30734.1| DRIL3 [Homo sapiens] 59 1e-07
gb|AAN74028.1| E2F binding protein [Homo sapiens] 59 1e-07
gb|AAH50925.1| Dead ringer homolog 1 [Mus musculus] 59 1e-07
gb|AAH60828.1| AT rich interactive domain 3A (BRIGHT- like) prot... 59 1e-07
gb|AAC28918.1| DRIL1 DNA binding protein homolog, partial CDS [H... 59 1e-07
>gb|AAP68262.1| At1g76110 [Arabidopsis thaliana] gi|20466328|gb|AAM20481.1| unknown
protein [Arabidopsis thaliana]
gi|15222957|ref|NP_177738.1| high mobility group
(HMG1/2) family protein / ARID/BRIGHT DNA-binding
domain-containing protein [Arabidopsis thaliana]
gi|25405003|pir||B96789 protein T23E18.4 [imported] -
Arabidopsis thaliana gi|6573729|gb|AAF17649.1| T23E18.4
[Arabidopsis thaliana]
Length = 338
Score = 207 bits (528), Expect = 2e-52
Identities = 117/233 (50%), Positives = 142/233 (60%), Gaps = 43/233 (18%)
Query: 16 KFYPPPLASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGY 75
K YP PLA HEV+V D+ +F DTLR+FH M +K+MIPVIGG++LDLH LYVEVTRR GY
Sbjct: 25 KEYPEPLALHEVVVKDSSVFWDTLRRFHSIMSTKFMIPVIGGKELDLHVLYVEVTRRGGY 84
Query: 76 EKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTVL 135
EKVV EKKWREVG VF FS TTTSAS+VL+KHY NLL+++EQVH F +GP+ P T
Sbjct: 85 EKVVVEKKWREVGGVFRFSATTTSASFVLRKHYLNLLFHYEQVHLFTARGPLLHPIAT-- 142
Query: 136 NSFTLCFMEMDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAP---KHANNG 192
HA + S ++ALVEY P ++ N
Sbjct: 143 -------------------FHA--------------NPSTSKEMALVEYTPPSIRYHNTH 169
Query: 193 PESNVEAKGYPRYGEGRIEEKFDCGYLVSVKLGSEVLRGVIFHPEETVPPPSS 245
P S + G IE KFDCGYLV VKLGSE+L GV++H + P PSS
Sbjct: 170 PPSQGSSSFT---AIGTIEGKFDCGYLVKVKLGSEILNGVLYHSAQ--PGPSS 217
>ref|XP_450195.1| glutathione S-transferase GST 16 - like protein [Oryza sativa
(japonica cultivar-group)] gi|32490476|dbj|BAC79159.1|
glutathione S-transferase GST 16 - like protein [Oryza
sativa (japonica cultivar-group)]
Length = 306
Score = 169 bits (428), Expect = 7e-41
Identities = 97/228 (42%), Positives = 134/228 (58%), Gaps = 31/228 (13%)
Query: 18 YPPPLASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEK 77
YPPPL SHE + ND F+DTLR+FH MG+K+MIPVIGG+++DLH LYVEVT R G K
Sbjct: 7 YPPPLLSHEEVANDRAAFMDTLRRFHSLMGTKFMIPVIGGKEMDLHALYVEVTSRGGLAK 66
Query: 78 VVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTVLNS 137
V+ E+KWREV + F+F TTTSASYVL+++Y +LL+++EQV+FF+ G + PA + L
Sbjct: 67 VMEERKWREVMARFSFPATTTSASYVLRRYYLSLLHHYEQVYFFRAHGALLRPAASALTK 126
Query: 138 FTLCFMEMDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAPKHANNGPESNV 197
M S D ++ +GK +AL P+ P S
Sbjct: 127 TPRRKMRGTS------------------DQSPAAAEAGK-RMAL----PERLGGEPCSFS 163
Query: 198 EAKGYPRYGEGRIEEKFDCGYLVSVKLGSEVLRGVIFHPEETVPPPSS 245
G I+ KF+ GYLV+VK+ +E LRGV++ PPP++
Sbjct: 164 VT--------GSIDGKFEHGYLVTVKIAAETLRGVLYRVAPPPPPPAA 203
>gb|AAW30021.1| At1g04880 [Arabidopsis thaliana] gi|56236040|gb|AAV84476.1|
At1g04880 [Arabidopsis thaliana]
gi|15220344|ref|NP_171980.1| high mobility group
(HMG1/2) family protein / ARID/BRIGHT DNA-binding
domain-containing protein [Arabidopsis thaliana]
gi|7211978|gb|AAF40449.1| Contains similarity to the
high mobility group family PF|00505. [Arabidopsis
thaliana] gi|25406818|pir||B86182 hypothetical protein
[imported] - Arabidopsis thaliana
Length = 448
Score = 140 bits (353), Expect = 3e-32
Identities = 85/222 (38%), Positives = 131/222 (58%), Gaps = 30/222 (13%)
Query: 21 PLASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEKVVA 80
P A++E +V D +LF+ +L + H +G+K+M+P+IGGR LDLH L+VEVT R G K++
Sbjct: 21 PEATYEAVVADPRLFMTSLERLHSLLGTKFMVPIIGGRDLDLHKLFVEVTSRGGINKILN 80
Query: 81 EKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTVLNSFTL 140
E++W+EV + F F T T+ASYVL+K+Y++LL N+EQ++FF+ G I PP S
Sbjct: 81 ERRWKEVTATFVFPPTATNASYVLRKYYFSLLNNYEQIYFFRSNGQI-PPDSMQSPSARP 139
Query: 141 CFMEMDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAPKHANNGPESNVEAK 200
CF +QG I E +++ + + +P + E+ + SNV
Sbjct: 140 CF-----IQGA---IRPSQELQAL-------TFTPQPKINTAEFL---GGSLAGSNV--- 178
Query: 201 GYPRYGEGRIEEKFDCGYLVSVKLGSEVLRGVIFH--PEETV 240
G I+ KF+ GYLV+V +GSE L+GV++ P+ TV
Sbjct: 179 ------VGVIDGKFESGYLVTVTIGSEQLKGVLYQLLPQNTV 214
>ref|XP_465568.1| glutathione S-transferase GST16-like protein [Oryza sativa
(japonica cultivar-group)] gi|47497529|dbj|BAD19581.1|
glutathione S-transferase GST16-like protein [Oryza
sativa (japonica cultivar-group)]
gi|47497414|dbj|BAD19471.1| glutathione S-transferase
GST16-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 467
Score = 112 bits (279), Expect = 1e-23
Identities = 76/241 (31%), Positives = 122/241 (50%), Gaps = 42/241 (17%)
Query: 10 SGGDDGK-----FYPPPLASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHT 64
SGG G YP +A ++ +V DA +F L H MG+K +P+IGG+ LDLH
Sbjct: 51 SGGSGGGRRTLVAYPARVAGYKDVVADAAVFRRALEGLHAQMGTKLKVPIIGGKDLDLHQ 110
Query: 65 LYVEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQ 124
L+ EVT R G +KV ++ +WREV + F F T T+AS++LKK+Y +LLY+FE+++ F+ Q
Sbjct: 111 LFKEVTSRGGIDKVKSDNRWREVTASFIFPATATNASFMLKKYYMSLLYHFERLYLFEAQ 170
Query: 125 GPITPPAGTVLNSFTLCFMEMDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEY 184
G + E DS S ++ + +S K
Sbjct: 171 G---------------WYQETDS------------RSISCIEMKAEGQASRK-------- 195
Query: 185 APKHANNGPESNVEAKGYPRYGEGRIEEKFDCGYLVSVKLGSEVLRGVIFHPEETVPPPS 244
K +N S++ A + I+ KF+ GY+V+V +GS+ + V+++ E P+
Sbjct: 196 -RKRGSNSCSSDL-AASLDNDVQVIIDGKFEHGYIVTVIMGSKSTKAVLYNCTEEPAVPT 253
Query: 245 S 245
+
Sbjct: 254 A 254
>gb|AAM65306.1| unknown [Arabidopsis thaliana] gi|22137084|gb|AAM91387.1|
At3g13350/MDC11_14 [Arabidopsis thaliana]
gi|9294541|dbj|BAB02804.1| high mobility group
protein-like [Arabidopsis thaliana]
gi|13605513|gb|AAK32750.1| AT3g13350/MDC11_14
[Arabidopsis thaliana] gi|18399977|ref|NP_566454.1| high
mobility group (HMG1/2) family protein / ARID/BRIGHT
DNA-binding domain-containing protein [Arabidopsis
thaliana]
Length = 319
Score = 112 bits (279), Expect = 1e-23
Identities = 73/228 (32%), Positives = 114/228 (49%), Gaps = 49/228 (21%)
Query: 23 ASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEK 82
A ++ +V ++ LF + LR F +P +GG LDLH L++EVT R G E+VV ++
Sbjct: 34 AKYDDLVRNSALFWEKLRAFLGLTSKTLKVPTVGGNTLDLHRLFIEVTSRGGIERVVKDR 93
Query: 83 KWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTVLNSFTLCF 142
KW+EV F+F TT TSAS+VL+K+Y L+ E V++ ++ P++
Sbjct: 94 KWKEVIGAFSFPTTITSASFVLRKYYLKFLFQLEHVYY--LEKPVS-------------- 137
Query: 143 MEMDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAPKHANNGPESNVEAKGY 202
SLQ S+ LA P+ + P+ E +G+
Sbjct: 138 ----SLQ---------------------STDEALKSLANESPNPEEGIDEPQVGYEVQGF 172
Query: 203 PRYGEGRIEEKFDCGYLVSVKLGSEVLRGVIFHPEETVPPPSSRVVGT 250
I+ KFD GYLV++KLGS+ L+GV++H +T P S + + T
Sbjct: 173 -------IDGKFDSGYLVTMKLGSQELKGVLYHIPQT-PSQSQQTMET 212
>ref|NP_175961.1| high mobility group (HMG1/2) family protein / ARID/BRIGHT
DNA-binding domain-containing protein [Arabidopsis
thaliana]
Length = 337
Score = 108 bits (270), Expect = 1e-22
Identities = 67/211 (31%), Positives = 105/211 (49%), Gaps = 48/211 (22%)
Query: 25 HEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKW 84
++ IV + +LF + LR FH K+ IP++GG+ LDLH L+ EVT R G EKV+ +++
Sbjct: 30 YQDIVRNPELFWEMLRDFHESSDKKFKIPIVGGKSLDLHRLFNEVTSRGGLEKVIKDRRC 89
Query: 85 REVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTVLNSFTLCFME 144
+EV FNF TT T++++VL+K Y +L+ FE +++F Q P+
Sbjct: 90 KEVIDAFNFKTTITNSAFVLRKSYLKMLFEFEHLYYF--QAPL----------------- 130
Query: 145 MDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAPKHANNGPESNVEAKGYPR 204
S+ + + AL K AN +S G
Sbjct: 131 ---------------------------STFWEKEKALKLLIEKSANRDKDSQELKPG--T 161
Query: 205 YGEGRIEEKFDCGYLVSVKLGSEVLRGVIFH 235
G I+ KF+ GYL+S K+GSE L+G+++H
Sbjct: 162 VITGIIDGKFESGYLISTKVGSEKLKGMLYH 192
>ref|NP_689854.2| AT rich interactive domain 2 (ARID, RFX-like) [Homo sapiens]
Length = 1835
Score = 84.7 bits (208), Expect = 2e-15
Identities = 41/97 (42%), Positives = 58/97 (59%), Gaps = 1/97 (1%)
Query: 35 FLDTLRQFHFHMGSKYM-IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNF 93
FLD LRQFH GS + IP +GG++LDLH LY VT G+ KV + +W E+ FNF
Sbjct: 19 FLDELRQFHHSRGSPFKKIPAVGGKELDLHGLYTRVTTLGGFAKVSEKNQWGEIVEEFNF 78
Query: 94 STTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPP 130
+ ++A++ LK++Y L +E+VH F PP
Sbjct: 79 PRSCSNAAFALKQYYLRYLEKYEKVHHFGEDDDEVPP 115
>gb|AAU20329.2| ARID2 [Homo sapiens]
Length = 1113
Score = 84.7 bits (208), Expect = 2e-15
Identities = 41/97 (42%), Positives = 58/97 (59%), Gaps = 1/97 (1%)
Query: 35 FLDTLRQFHFHMGSKYM-IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNF 93
FLD LRQFH GS + IP +GG++LDLH LY VT G+ KV + +W E+ FNF
Sbjct: 19 FLDELRQFHHSRGSPFKKIPAVGGKELDLHGLYTRVTTLGGFAKVSEKNQWGEIVEEFNF 78
Query: 94 STTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPP 130
+ ++A++ LK++Y L +E+VH F PP
Sbjct: 79 PRSCSNAAFALKQYYLRYLEKYEKVHHFGEDDDEVPP 115
>dbj|BAB24929.1| unnamed protein product [Mus musculus]
Length = 145
Score = 84.7 bits (208), Expect = 2e-15
Identities = 41/97 (42%), Positives = 58/97 (59%), Gaps = 1/97 (1%)
Query: 35 FLDTLRQFHFHMGSKYM-IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNF 93
FLD LRQFH GS + IP +GG++LDLH LY VT G+ KV + +W E+ FNF
Sbjct: 19 FLDELRQFHHSRGSPFKKIPAVGGKELDLHGLYTRVTTLGGFAKVSEKNQWGEIVEEFNF 78
Query: 94 STTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPP 130
+ ++A++ LK++Y L +E+VH F PP
Sbjct: 79 PRSCSNAAFALKQYYLRYLEKYEKVHHFGEDDDEVPP 115
>ref|XP_416046.1| PREDICTED: similar to hypothetical protein [Gallus gallus]
Length = 2143
Score = 75.1 bits (183), Expect = 2e-12
Identities = 35/75 (46%), Positives = 50/75 (66%), Gaps = 1/75 (1%)
Query: 35 FLDTLRQFHFHMGSKYM-IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNF 93
FLD LRQFH GS + IPV+GG++LDLH LY VT G+ KV + +W E+ FNF
Sbjct: 19 FLDELRQFHHSRGSPFKKIPVVGGKELDLHALYTRVTTLGGFGKVSEKNQWGEIVEEFNF 78
Query: 94 STTTTSASYVLKKHY 108
+ ++A++ LK++Y
Sbjct: 79 PRSCSNAAFALKQYY 93
>emb|CAH18689.1| hypothetical protein [Homo sapiens]
Length = 1756
Score = 69.7 bits (169), Expect = 7e-11
Identities = 31/79 (39%), Positives = 47/79 (59%)
Query: 52 IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNL 111
IP +GG++LDLH LY VT G+ KV + +W E+ FNF + ++A++ LK++Y
Sbjct: 10 IPAVGGKELDLHGLYTRVTTLGGFAKVSEKNQWGEIVEEFNFPRSCSNAAFALKQYYLRY 69
Query: 112 LYNFEQVHFFKVQGPITPP 130
L +E+VH F PP
Sbjct: 70 LEKYEKVHHFGEDDDEVPP 88
>gb|AAH72808.1| MGC80148 protein [Xenopus laevis] gi|58429893|gb|AAW78333.1| Dril1
[Xenopus laevis]
Length = 539
Score = 60.5 bits (145), Expect = 4e-08
Identities = 33/85 (38%), Positives = 48/85 (55%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL+ LYV VT + G +V+ +K WRE+
Sbjct: 213 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLYMLYVLVTEKGGLVEVINKKLWREITKGL 272
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 273 NLPTSITSAAFTLRTQYMKYLYPYE 297
>gb|AAH84503.1| Hypothetical LOC496519 [Xenopus tropicalis]
gi|58331845|ref|NP_001011106.1| hypothetical protein
LOC496519 [Xenopus tropicalis]
Length = 541
Score = 60.5 bits (145), Expect = 4e-08
Identities = 33/85 (38%), Positives = 48/85 (55%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL+ LYV VT + G +V+ +K WRE+
Sbjct: 216 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLYMLYVLVTEKGGLVEVINKKLWREITKGL 275
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 276 NLPTSITSAAFTLRTQYMKYLYPYE 300
>ref|NP_031906.1| dead ringer homolog 1 [Mus musculus] gi|1401348|gb|AAB03416.1|
Bright [Mus musculus] gi|12230032|sp|Q62431|ARI3A_MOUSE
AT-rich interactive domain-containing protein 3A (ARID
domain-containing protein 3A) (Dead ringer like-1
protein) (B-cell regulator of IgH transcription)
(Bright)
Length = 601
Score = 59.3 bits (142), Expect = 1e-07
Identities = 33/85 (38%), Positives = 47/85 (54%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL LYV VT + G +V+ +K WRE+
Sbjct: 247 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLFMLYVLVTEKGGLVEVINKKLWREITKGL 306
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 307 NLPTSITSAAFTLRTQYMKYLYPYE 331
>sp|Q99856|ARI3A_HUMAN AT-rich interactive domain-containing protein 3A (ARID
domain-containing protein 3A) (B-cell regulator of IgH
transcription) (Bright) (E2F binding protein 1)
gi|3002816|gb|AAC69994.1| dead ringer like 1 protein
[Homo sapiens] gi|2529688|gb|AAC32888.1| DNA binding
protein homolog [Homo sapiens]
gi|4885193|ref|NP_005215.1| AT rich interactive domain
3A (BRIGHT- like) protein [Homo sapiens]
Length = 593
Score = 59.3 bits (142), Expect = 1e-07
Identities = 33/85 (38%), Positives = 47/85 (54%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL LYV VT + G +V+ +K WRE+
Sbjct: 242 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLFMLYVLVTEKGGLVEVINKKLWREITKGL 301
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 302 NLPTSITSAAFTLRTQYMKYLYPYE 326
>gb|AAW30734.1| DRIL3 [Homo sapiens]
Length = 589
Score = 59.3 bits (142), Expect = 1e-07
Identities = 33/85 (38%), Positives = 47/85 (54%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL LYV VT + G +V+ +K WRE+
Sbjct: 242 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLFMLYVLVTEKGGLVEVINKKLWREITKGL 301
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 302 NLPTSITSAAFTLRTQYMKYLYPYE 326
>gb|AAN74028.1| E2F binding protein [Homo sapiens]
Length = 593
Score = 59.3 bits (142), Expect = 1e-07
Identities = 33/85 (38%), Positives = 47/85 (54%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL LYV VT + G +V+ +K WRE+
Sbjct: 242 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLFMLYVLVTEKGGLVEVINKKLWREITKGL 301
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 302 NLPTSITSAAFTLRTQYMKYLYPYE 326
>gb|AAH50925.1| Dead ringer homolog 1 [Mus musculus]
Length = 599
Score = 59.3 bits (142), Expect = 1e-07
Identities = 33/85 (38%), Positives = 47/85 (54%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL LYV VT + G +V+ +K WRE+
Sbjct: 247 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLFMLYVLVTEKGGLVEVINKKLWREITKGL 306
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 307 NLPTSITSAAFTLRTQYMKYLYPYE 331
>gb|AAH60828.1| AT rich interactive domain 3A (BRIGHT- like) protein [Homo sapiens]
Length = 593
Score = 59.3 bits (142), Expect = 1e-07
Identities = 33/85 (38%), Positives = 47/85 (54%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL LYV VT + G +V+ +K WRE+
Sbjct: 242 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLFMLYVLVTEKGGLVEVINKKLWREITKGL 301
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 302 NLPTSITSAAFTLRTQYMEYLYPYE 326
>gb|AAC28918.1| DRIL1 DNA binding protein homolog, partial CDS [Homo sapiens]
Length = 363
Score = 59.3 bits (142), Expect = 1e-07
Identities = 33/85 (38%), Positives = 47/85 (54%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL LYV VT + G +V+ +K WRE+
Sbjct: 12 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLFMLYVLVTEKGGLVEVINKKLWREITKGL 71
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 72 NLPTSITSAAFTLRTQYMKYLYPYE 96
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.137 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 474,370,413
Number of Sequences: 2540612
Number of extensions: 19858244
Number of successful extensions: 38027
Number of sequences better than 10.0: 313
Number of HSP's better than 10.0 without gapping: 276
Number of HSP's successfully gapped in prelim test: 37
Number of HSP's that attempted gapping in prelim test: 37682
Number of HSP's gapped (non-prelim): 327
length of query: 253
length of database: 863,360,394
effective HSP length: 125
effective length of query: 128
effective length of database: 545,783,894
effective search space: 69860338432
effective search space used: 69860338432
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0121b.2


