

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
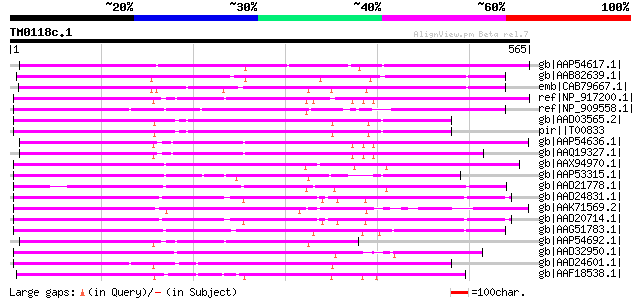
Query= TM0118c.1
(565 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP54617.1| putative non-LTR retroelement reverse transcripta... 261 6e-68
gb|AAB82639.1| putative non-LTR retroelement reverse transcripta... 248 3e-64
emb|CAB79667.1| putative protein [Arabidopsis thaliana] gi|49720... 241 3e-62
ref|NP_917200.1| P0707D10.17 [Oryza sativa (japonica cultivar-gr... 236 1e-60
ref|NP_909558.1| putative reverse transcriptase [Oryza sativa] g... 229 2e-58
gb|AAD03565.2| putative non-LTR retroelement reverse transcripta... 227 7e-58
pir||T00833 RNA-directed DNA polymerase homolog T13L16.7 - Arabi... 227 7e-58
gb|AAP54636.1| putative reverse transcriptase [Oryza sativa (jap... 224 6e-57
gb|AAQ19327.1| bZIP-like protein [Oryza sativa (japonica cultiva... 221 5e-56
gb|AAX94970.1| retrotransposon protein, putative, unclassified [... 219 2e-55
gb|AAP53315.1| putative reverse transcriptase [Oryza sativa (jap... 218 5e-55
gb|AAD21778.1| putative non-LTR retroelement reverse transcripta... 214 5e-54
gb|AAD24831.1| putative non-LTR retroelement reverse transcripta... 212 2e-53
gb|AAK71569.2| putative reverse transcriptase [Oryza sativa (jap... 211 4e-53
gb|AAD20714.1| putative non-LTR retroelement reverse transcripta... 209 1e-52
gb|AAG51783.1| reverse transcriptase, putative; 16838-20266 [Ara... 209 1e-52
gb|AAP54692.1| putative reverse transcriptase [Oryza sativa (jap... 203 1e-50
gb|AAD32950.1| putative non-LTR retroelement reverse transcripta... 199 2e-49
gb|AAD24601.1| putative non-LTR retroelement reverse transcripta... 194 5e-48
gb|AAF18538.1| Very similar to retrotransposon reverse transcrip... 189 3e-46
>gb|AAP54617.1| putative non-LTR retroelement reverse transcriptase [Oryza sativa
(japonica cultivar-group)] gi|37536056|ref|NP_922330.1|
putative non-LTR retroelement reverse transcriptase
[Oryza sativa (japonica cultivar-group)]
gi|10140689|gb|AAG13524.1| putative non-LTR retroelement
reverse transcriptase [Oryza sativa (japonica
cultivar-group)]
Length = 1382
Score = 261 bits (666), Expect = 6e-68
Identities = 174/573 (30%), Positives = 268/573 (46%), Gaps = 23/573 (4%)
Query: 11 AIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGFRDF 70
AIP+YVMSCF + +C +++ I+ +WG + + MHW W L PK GG+GFR+F
Sbjct: 814 AIPNYVMSCFRIPVSICEKMKTCIADHWWGFEDGKKKMHWKSWSWLSTPKFLGGMGFREF 873
Query: 71 KAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQKSAWIF 130
FN A+L + WR++T S+ +++ K YFP SF EA + PS+ W SL +
Sbjct: 874 TTFNQAMLGRQCWRLLTDPDSLCSRVLKGRYFPNSSFWEAAQPKSPSFTWRSLLFGRELL 933
Query: 131 QEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGVVHVSDLMVTNGGCWDRNKVE 190
+G W VG+GK I+ D W+P P + ++ VS LM + CWD + +
Sbjct: 934 AKGVRWGVGDGKTIKIFSDNWIPGFRPQLV-TTLSPFPTDATVSCLMNEDARCWDGDLIR 992
Query: 191 FLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEASSSSARV 250
L A+ IL IP+ D WP + G YS R+ Y+ + + S+S +
Sbjct: 993 SLFPVDIAKEILQIPISRHGDADFASWPHDKLGLYSVRSAYNLARSEAFFADQSNSGRGM 1052
Query: 251 MPPAL-----WMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEVEE 305
L W LWK +A + T WRA + L L RR + C C+ ++
Sbjct: 1053 ASRLLESQKDWKGLWKINAPGKMKITLWRAAHECLATGFQLRRRHIPSTDGCVFCNR-DD 1111
Query: 306 TVHHALFSCPLVRMIW--FASPLNLRFDEEG--SVHDFFLSFLATADTNIAGIFLAVLYS 361
TV H CP IW ++ G ++ + FL ++ + +
Sbjct: 1112 TVEHVFLFCPFAAQIWEEIKGKCAVKLGRNGFSTMRQWIFDFLKRGSSHANTLLAVTFWH 1171
Query: 362 VWQARNELLFKWKDSSID------QLLQRASSLRPSRVLLDDGPRA--VQVLPSSWSRPL 413
+W+ARN K + ++ ++L + DG R Q +P W P
Sbjct: 1172 IWEARNNT--KNNNGTVHPQRVVIKILSYVDMILKHNTKTVDGQRGGNTQAIP-RWQPPP 1228
Query: 414 SGVCKMNIDASV-TSSKEAGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRWALRL 472
+ V +N DA++ +SS+ G G ++R+N G+ L A + + V+ P LAEAL +R AL L
Sbjct: 1229 ASVWMINSDAAIFSSSRTMGVGALIRDNTGKCLVACSEMISDVVLPELAEALAIRRALGL 1288
Query: 473 ATKLGFRRIWIETDCLELYTFWNRQGGEFSHLATVVFDCRSLLPNFDVFNFTFVRRTSNG 532
A + G I + +DCL + G + S + V+ D + L F + +F V R SN
Sbjct: 1289 AKEEGLEHIVMASDCLTVIRRIQTSGRDRSGVGCVIEDIKKLASTFVLCSFMHVNRLSNL 1348
Query: 533 AADALARLAFHYGCMIWIEEAPSEITSITQDDV 565
AA +LAR A C ++ P I I DDV
Sbjct: 1349 AAHSLARNAELSTCTVYRSVIPDYIRDILCDDV 1381
>gb|AAB82639.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25408936|pir||A84888 hypothetical protein
At2g45230 [imported] - Arabidopsis thaliana
Length = 1374
Score = 248 bits (634), Expect = 3e-64
Identities = 164/559 (29%), Positives = 269/559 (47%), Gaps = 36/559 (6%)
Query: 8 INLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGF 67
+ +A+P+Y MSCF + +C QIE +++ F+W R +HW W L RPK GG+GF
Sbjct: 796 VAMALPTYTMSCFKIPKTICQQIESVMAEFWWKNKKEGRGLHWKAWCHLSRPKAVGGLGF 855
Query: 68 RDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQKSA 127
++ +AFN ALL K WR++T+ S++AK++K+ YF K L A G RPS+ W S+ ++
Sbjct: 856 KEIEAFNIALLGKQLWRMITEKDSLMAKVFKSRYFSKSDPLNAPLGSRPSFAWKSIYEAQ 915
Query: 128 WIFQEGGLWRVGNGKNIRA*EDKWV----PHGGPLVYRNDVAESLGV--VH-VSDLMVTN 180
+ ++G +GNG+ I D W+ V R+ + +H V DL++ +
Sbjct: 916 VLIKQGIRAVIGNGETINVWTDPWIGAKPAKAAQAVKRSHLVSQYAANSIHVVKDLLLPD 975
Query: 181 GGCWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSE 240
G W+ N V L T IL++ ++ D F W +R G YS ++GY + + ++
Sbjct: 976 GRDWNWNLVSLLFPDNTQENILALRPGGKETRDRFTWEYSRSGHYSVKSGYWVMTEIINQ 1035
Query: 241 GEASSSSARVMPPAL---WMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCC 297
++ V+ P+L + ++WK D P+ WR + L V S L R + C
Sbjct: 1036 ---RNNPQEVLQPSLDPIFQQIWKLDVPPKIHHFLWRCVNNCLSVASNLAYRHLAREKSC 1092
Query: 298 PRCSEVEETVHHALFSCPLVRMIWFASPLNLRFDEEGSV-------HDFFLSFLATADTN 350
RC ETV+H LF CP R+ W SPL E + H + +++
Sbjct: 1093 VRCPSHGETVNHLLFKCPFARLTWAISPLPAPPGGEWAESLFRNMHHVLSVHKSQPEESD 1152
Query: 351 IAGIFLAVLYSVWQARNELLFKWKDSSIDQLLQRASSLRPS---------RVLLDDGPRA 401
+ +L+ +W+ RN+L+FK ++ + Q++ +A+ + +V R
Sbjct: 1153 HHALIPWILWRLWKNRNDLVFKGREFTAPQVILKATEDMDAWNNRKEPQPQVTSSTRDRC 1212
Query: 402 VQVLPSSWSRPLSGVCKMNIDASVTSS-KEAGAGMVMRNNHGEVLAAATTELGPVLSPVL 460
V+ W P G K N D + + G G V+RN+ G +L L S +
Sbjct: 1213 VK-----WQPPSHGWVKCNTDGAWSKDLGNCGVGWVLRNHTGRLLWLGLRALPSQQSVLE 1267
Query: 461 AEALGMRWALRLATKLGFRRIWIETDCLELYTFWNRQGGEFSHLATVVFDCRSLLPNFDV 520
E +RWA+ ++ +RR+ E+D L + + + LA + D R+LL +F+
Sbjct: 1268 TEVEALRWAVLSLSRFNYRRVIFESDSQYLVSLIQNE-MDIPSLAPRIQDIRNLLRHFEE 1326
Query: 521 FNFTFVRRTSNGAADALAR 539
F F RR N AD AR
Sbjct: 1327 VKFQFTRREGNNVADRTAR 1345
>emb|CAB79667.1| putative protein [Arabidopsis thaliana] gi|4972055|emb|CAB43923.1|
putative protein [Arabidopsis thaliana]
gi|67633766|gb|AAY78807.1| putative reverse
transcriptase/RNA-dependent DNA polymerase [Arabidopsis
thaliana] gi|15233451|ref|NP_194638.1| reverse
transcriptase, putative / RNA-dependent DNA polymerase,
putative [Arabidopsis thaliana] gi|7485741|pir||T08964
hypothetical protein F19B15.120 - Arabidopsis thaliana
Length = 575
Score = 241 bits (616), Expect = 3e-62
Identities = 168/556 (30%), Positives = 265/556 (47%), Gaps = 32/556 (5%)
Query: 10 LAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGFRD 69
+A+P+Y M+CFLL +C QI +++ F+W + MHW WD L K GGIGF+D
Sbjct: 1 MALPTYTMACFLLPKTVCKQIISVLADFWWRNKQEAKGMHWKAWDHLSCYKAEGGIGFKD 60
Query: 70 FKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQKSAWI 129
+AFN ALL K WR++++ +S++AK++K+ YF K L A G RPS+VW S+ S I
Sbjct: 61 IEAFNLALLGKQMWRMLSRPESLMAKVFKSRYFHKSDPLNAPLGSRPSFVWKSIHASQEI 120
Query: 130 FQEGGLWRVGNGKNIRA*EDKWV---PHGGPL----VYRNDVAESLGVVHVSDLMVTNGG 182
++G VGNG++I KW+ P L V + A ++ VSDL+ +G
Sbjct: 121 LRQGARAVVGNGEDIIIWRHKWLDSKPASAALRMQRVPPQEYASVSSILKVSDLIDESGR 180
Query: 183 CWDRNKVEFLCWPPTARAILSIPLPLQQRD-DCFFWPETRDGCYSTRTGY----SFI*KK 237
W ++ +E L +P R ++ P +R D + W T G Y+ ++GY I K+
Sbjct: 181 EWRKDVIEML-FPEVERKLIGELRPGGRRILDSYTWDYTSSGDYTVKSGYWVLTQIINKR 239
Query: 238 FSEGEASSSSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCC 297
S E S S ++ K+WK+ P+ W+ + LPV L R + C
Sbjct: 240 SSPQEVSEPSLN----PIYQKIWKSQTSPKIQHFLWKCLSNSLPVAGALAYRHLSKESAC 295
Query: 298 PRCSEVEETVHHALFSCPLVRMIWFAS----PLNLRFDEEGSVHDFFLSFLATAD---TN 350
RC +ETV+H LF C R+ W S PL + + V+ +++ L +
Sbjct: 296 IRCPSCKETVNHLLFKCTFARLTWAISSIPIPLGGEWADSIYVNLYWVFNLGNGNPQWEK 355
Query: 351 IAGIFLAVLYSVWQARNELLFKWKDSSIDQLLQRASS------LRPSRVLLDDGPRAVQV 404
+ + +L+ +W+ RNEL+F+ ++ + ++L+RA +R P+ +
Sbjct: 356 ASQLVPWLLWRLWKNRNELVFRGREFNAQEVLRRAEDDLEEWRIRTEAESCGTKPQVNRS 415
Query: 405 LPSSWSRPLSGVCKMNIDASVTSSKE-AGAGMVMRNNHGEVLAAATTELGPVLSPVLAEA 463
W P K N DA+ E G G V+RN GEV L + S + AE
Sbjct: 416 SCGRWRPPPHQWVKCNTDATWNRDNERCGIGWVLRNEKGEVKWMGARALPKLKSVLEAEL 475
Query: 464 LGMRWALRLATKLGFRRIWIETDCLELYTFWNRQGGEFSHLATVVFDCRSLLPNFDVFNF 523
MRWA+ ++ + + E+D L N + L + D + LL F F
Sbjct: 476 EAMRWAVLSLSRFQYNYVIFESDSQVLIEILNND-EIWPSLKPTIQDLQRLLSQFTEVKF 534
Query: 524 TFVRRTSNGAADALAR 539
F+ R N A+ +AR
Sbjct: 535 VFIPREGNTLAERVAR 550
>ref|NP_917200.1| P0707D10.17 [Oryza sativa (japonica cultivar-group)]
Length = 892
Score = 236 bits (602), Expect = 1e-60
Identities = 165/596 (27%), Positives = 272/596 (44%), Gaps = 45/596 (7%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
I + IP YVM F L + +C ++ ++ F+WG + R HW W+ L RPK GG
Sbjct: 304 IKSVIQTIPVYVMGIFNLPESVCEELTKLTRNFWWGVEKGKRKTHWKSWECLTRPKSNGG 363
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
+GFRDF+ FN ALLA+ WR++ S+ A++ KA YFP S ++ G S VW +++
Sbjct: 364 LGFRDFRLFNQALLARQAWRLIVNPDSLCARVLKAKYFPNGSLVDTSFGGNASPVWKAIE 423
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHG---GPLVYRNDVAESLGVVHVSDLMVTNG 181
+ +EG +WR+GNGK++R D W+P P+ + + + VS+L+ N
Sbjct: 424 YGLSLLKEGIIWRIGNGKSVRIWRDPWLPRDFSRRPITRKGNCR----LKWVSELLSEN- 478
Query: 182 GCWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEG 241
G WD +V + P A AI SI + +Q DD W + G +S R+ Y
Sbjct: 479 GAWDETRVNQVFLPVDAEAICSIRVSSRQEDDFVAWHPDKHGKFSVRSAYGLA-CNLVNM 537
Query: 242 EASSSSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCS 301
E SSSS+ +W +W+ +A + AWRA + L +R + + C C
Sbjct: 538 ETSSSSSSARCKRMWNLIWQTNAPQKVRIFAWRAATNCLATMENKAKRKLEQSDICLVCG 597
Query: 302 EVEETVHHALFSCPLVRMIW----FASPLNLR----FDEEGSVHDFFLSFLATADTNIAG 353
+E H L CP R +W A ++L ++ G + D L T
Sbjct: 598 LEKEDTKHVLCRCPDARNLWGAMCEAGNISLNGGNLCNDPGWIFDQLEILLDYERT---- 653
Query: 354 IFLAVLYSVWQARNELLF-----------KWKDSSIDQLLQ-----RASSLRPSRVL--- 394
+FL +L+ +W RNE++ ++ +S + LL+ +A+ + V+
Sbjct: 654 VFLLILWRIWFVRNEIVHGKQPPPIEASKRFLESYVKSLLEIRQHPQANVEKGKHVISFV 713
Query: 395 ---LDDGPRAVQVLPSSWSRPLSGVCKMNIDASV-TSSKEAGAGMVMRNNHGEVLAAATT 450
+ R + W +P G K+N+D S + G G+VMR+ G+V+ A
Sbjct: 714 NGKTNPARRPPEDKNQKWRKPCEGWMKVNVDGSFDAQTGSGGIGVVMRDWSGKVIFACCK 773
Query: 451 ELGPVLSPVLAEALGMRWALRLATKLGFRRIWIETDCLELYTFWNRQGGEFSHLATVVFD 510
L S + +E L + + + + R+ +ETDCL + + E S +A +V +
Sbjct: 774 HLPRCGSALESEILACKEGILVGLQWSLRQFVVETDCLMVVQMLESKEREMSEVAFLVKE 833
Query: 511 CRSLLPNFDVFNFTFVRRTSNGAADALARLAFHYGCMI-WIEEAPSEITSITQDDV 565
LL + R+ N + LA G + W++ S I+ + DD+
Sbjct: 834 ATELLSGDREIQLRKIHRSQNSVSHLLANKGRCEGLSVFWLDHECSCISRLVLDDL 889
>ref|NP_909558.1| putative reverse transcriptase [Oryza sativa]
gi|14018103|gb|AAK52166.1| putative reverse transcriptase
[Oryza sativa]
Length = 1185
Score = 229 bits (584), Expect = 2e-58
Identities = 155/546 (28%), Positives = 244/546 (44%), Gaps = 43/546 (7%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
I I AIP + MS F + +C I I+RF+WG D +HR +HW W +C PK GG
Sbjct: 611 IKSIAQAIPVFAMSVFRIPKKICDCISEAIARFWWGDDENHRRIHWKAWWRMCIPKQRGG 670
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
+GFRD FN A+LAK WR + S+ A++ + Y+P + L A+ SY W S+
Sbjct: 671 LGFRDLYCFNRAMLAKQIWRFLCDPDSLCARVMRCKYYPDGNILRARPKKGSSYAWQSIL 730
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGVVHVSDLMVTNGGCW 184
+ F +G +WR+G+G + ED +P ++ + V +L+ G W
Sbjct: 731 SGSECF-KGYIWRIGDGSQVNIWEDSLIPSSSNGKVLTPRGRNI-ITKVEELINPVSGGW 788
Query: 185 DRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*K-KFSEGEA 243
D + ++ L WP A IL IP+ R+D W TR G +S R+ Y KF E
Sbjct: 789 DEDLIDTLFWPIDANRILQIPIS-AGREDFVAWHHTRSGIFSVRSAYHCQWNAKFGSKER 847
Query: 244 SSSSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEV 303
A +W LWK + P+ AWR ++P + IL + + CP CS
Sbjct: 848 LPDGASSSSRPVWDNLWKLEIPPKIKIFAWRMLHGVIPCKGILADKHIAVQGGCPVCSSG 907
Query: 304 EETVHHALFSCPLVRMIW-------FASPLNLRFDEEGS--VHDFFLSFLATADTNIAGI 354
E + H +F+C ++IW P+ L FD GS V + + N G+
Sbjct: 908 VEDIKHMVFTCRRAKLIWKQLGIWSRIQPI-LGFDRSGSVLVEEMIRKGGKVSHLNAIGM 966
Query: 355 FLAVLYSVWQARNELLFKWKDSSIDQLLQRASSLRPSRVLLDDGPRAVQVLPSSWSRPLS 414
+L W + W + QL+Q P+R +
Sbjct: 967 TELILTGAW-------YIWWERR--QLVQGERIQNPARSAM------------------- 998
Query: 415 GVCKMNIDASVTSSK-EAGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRWALRLA 473
+ + + +++ K G G+V+R++ G +AA+ L VL +AEA +R L LA
Sbjct: 999 SIATLTTNYMLSNKKGSGGIGVVLRDHLGACVAASQAFLPHVLDAPMAEAFALRDGLSLA 1058
Query: 474 TKLGFRRIWIETDCLELYTFWNRQGGEFSHLATVVFDCRSLLPNFDVFNFTFVRRTSNGA 533
+G + + ++TDC+++ + G + +V DC+ + F + R SN
Sbjct: 1059 QHIGAKNVIVQTDCMQVVSTMMDGGFSATAAMSVFDDCKLMWDGFGTISIEHCNRNSNQV 1118
Query: 534 ADALAR 539
A LAR
Sbjct: 1119 AHELAR 1124
>gb|AAD03565.2| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25411819|pir||H84557 hypothetical protein
At2g17910 [imported] - Arabidopsis thaliana
Length = 1344
Score = 227 bits (579), Expect = 7e-58
Identities = 146/491 (29%), Positives = 231/491 (46%), Gaps = 20/491 (4%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
+ I LA+P YVMSCF L LC ++ ++ F+W R +HWL W L PK GG
Sbjct: 771 LKSIALALPVYVMSCFKLPKNLCQKLTTVMMDFWWNSMQQKRKIHWLSWQRLTLPKDQGG 830
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
GF+D + FN ALLAK WR++ + S+ ++++++ YF FL A +G RPSY W S+
Sbjct: 831 FGFKDLQCFNQALLAKQAWRVLQEKGSLFSRVFQSRYFSNSDFLSATRGSRPSYAWRSIL 890
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGG---PLVYRNDVAESLGVVHVSDLMVTNG 181
+ +G +GNG+ DKW+ G PL R + L V + D N
Sbjct: 891 FGRELLMQGLRTVIGNGQKTFVWTDKWLHDGSNRRPLNRRRFINVDLKVSQLIDPTSRN- 949
Query: 182 GCWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEG 241
W+ N + L +P I+ PL ++D F W + +G YS +TGY F+ K+
Sbjct: 950 --WNLNMLRDL-FPWKDVEIILKQRPLFFKEDSFCWLHSHNGLYSVKTGYEFLSKQVHHR 1006
Query: 242 EASSSSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCS 301
+ + +L+ K+W P+ W+A +PV L RG+ + C C
Sbjct: 1007 LYQEAKVKPSVNSLFDKIWNLHTAPKIRIFLWKALHGAIPVEDRLRTRGIRSDDGCLMCD 1066
Query: 302 EVEETVHHALFSCPLVRMIWFASPLNLRFDE-EGSVHDFFLSFLATADTN-----IAGIF 355
ET++H LF CPL R +W + L+ E SV+ + N + +
Sbjct: 1067 TENETINHILFECPLARQVWAITHLSSAGSEFSNSVYTNMSRLIDLTQQNDLPHHLRFVS 1126
Query: 356 LAVLYSVWQARNELLFKWKDSSIDQLLQRASSLR----PSRVLLDDGPRAVQVLPSSWSR 411
+L+ +W+ RN LLF+ K S L+ +A ++ + + + +++ + W
Sbjct: 1127 PWILWFLWKNRNALLFEGKGSITTTLVDKAYEAYHEWFSAQTHMQNDEKHLKI--TKWCP 1184
Query: 412 PLSGVCKMNIDASVTSSKE-AGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRWAL 470
PL G K NI + + +GA V+R++ G+VL + V SP A+ WAL
Sbjct: 1185 PLPGELKCNIGFAWSKQHHFSGASWVVRDSQGKVLLHSRRSFNEVHSPYSAKIRSWEWAL 1244
Query: 471 RLATKLGFRRI 481
T F R+
Sbjct: 1245 ESMTHHHFDRV 1255
>pir||T00833 RNA-directed DNA polymerase homolog T13L16.7 - Arabidopsis thaliana
(fragment)
Length = 1365
Score = 227 bits (579), Expect = 7e-58
Identities = 146/491 (29%), Positives = 231/491 (46%), Gaps = 20/491 (4%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
+ I LA+P YVMSCF L LC ++ ++ F+W R +HWL W L PK GG
Sbjct: 792 LKSIALALPVYVMSCFKLPKNLCQKLTTVMMDFWWNSMQQKRKIHWLSWQRLTLPKDQGG 851
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
GF+D + FN ALLAK WR++ + S+ ++++++ YF FL A +G RPSY W S+
Sbjct: 852 FGFKDLQCFNQALLAKQAWRVLQEKGSLFSRVFQSRYFSNSDFLSATRGSRPSYAWRSIL 911
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGG---PLVYRNDVAESLGVVHVSDLMVTNG 181
+ +G +GNG+ DKW+ G PL R + L V + D N
Sbjct: 912 FGRELLMQGLRTVIGNGQKTFVWTDKWLHDGSNRRPLNRRRFINVDLKVSQLIDPTSRN- 970
Query: 182 GCWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEG 241
W+ N + L +P I+ PL ++D F W + +G YS +TGY F+ K+
Sbjct: 971 --WNLNMLRDL-FPWKDVEIILKQRPLFFKEDSFCWLHSHNGLYSVKTGYEFLSKQVHHR 1027
Query: 242 EASSSSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCS 301
+ + +L+ K+W P+ W+A +PV L RG+ + C C
Sbjct: 1028 LYQEAKVKPSVNSLFDKIWNLHTAPKIRIFLWKALHGAIPVEDRLRTRGIRSDDGCLMCD 1087
Query: 302 EVEETVHHALFSCPLVRMIWFASPLNLRFDE-EGSVHDFFLSFLATADTN-----IAGIF 355
ET++H LF CPL R +W + L+ E SV+ + N + +
Sbjct: 1088 TENETINHILFECPLARQVWAITHLSSAGSEFSNSVYTNMSRLIDLTQQNDLPHHLRFVS 1147
Query: 356 LAVLYSVWQARNELLFKWKDSSIDQLLQRASSLR----PSRVLLDDGPRAVQVLPSSWSR 411
+L+ +W+ RN LLF+ K S L+ +A ++ + + + +++ + W
Sbjct: 1148 PWILWFLWKNRNALLFEGKGSITTTLVDKAYEAYHEWFSAQTHMQNDEKHLKI--TKWCP 1205
Query: 412 PLSGVCKMNIDASVTSSKE-AGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRWAL 470
PL G K NI + + +GA V+R++ G+VL + V SP A+ WAL
Sbjct: 1206 PLPGELKCNIGFAWSKQHHFSGASWVVRDSQGKVLLHSRRSFNEVHSPYSAKIRSWEWAL 1265
Query: 471 RLATKLGFRRI 481
T F R+
Sbjct: 1266 ESMTHHHFDRV 1276
>gb|AAP54636.1| putative reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|37536094|ref|NP_922349.1| putative
reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|13786450|gb|AAK39575.1| putative
reverse transcriptase [Oryza sativa]
Length = 791
Score = 224 bits (571), Expect = 6e-57
Identities = 166/585 (28%), Positives = 263/585 (44%), Gaps = 37/585 (6%)
Query: 11 AIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGFRDF 70
AIP YVM F L + + + ++ F+W R HW WD+L +PK GG+GFRD+
Sbjct: 209 AIPVYVMGIFKLPESVIDDLTKLTKNFWWDSMNGQRKTHWKAWDSLTKPKSLGGLGFRDY 268
Query: 71 KAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQKSAWIF 130
FN ALLA+ WR++T S+ A++ KA YFP S ++ G S W S++ +
Sbjct: 269 WLFNQALLARQAWRLITYPDSLCARVLKAKYFPHGSLIDTSFGSNSSPAWRSIEYGLDLL 328
Query: 131 QEGGLWRVGNGKNIRA*EDKWVPHG---GPLVYRNDVAESLGVVHVSDLMVTNGGCWDRN 187
++G +WRVGNG +IR D W+P P+ + + + VSDL +T G WD
Sbjct: 329 KKGIIWRVGNGNSIRIWRDPWLPRDHSRRPITGK----ANCRLKWVSDL-ITEDGSWDVP 383
Query: 188 KVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEASSSS 247
K+ A IL+I + + +D W ++G +S R+ Y + + E+SSS
Sbjct: 384 KIHQYFHNLDAEVILNICISSRSEEDFIAWHPDKNGMFSVRSAYRLAAQLVNIEESSSSG 443
Query: 248 ARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEVEETV 307
+ A W +WK + AWR + L +R + + C C E
Sbjct: 444 TNNINKA-WEMIWKCKVPQKVKIFAWRVASNCLATMVNKKKRKLEQSDMCQICDRENEDD 502
Query: 308 HHALFSCPLVRMIW--FASPLNLRFDEEGSVHDFFLSF--LATADTNIAGIFLAVLYSVW 363
HAL C +W ++ D + SV F F L +FL L+ W
Sbjct: 503 AHALCRCIQASQLWSCMHKSGSVSVDIKASVLGRFWLFDCLEKIPEYEQAMFLMTLWRNW 562
Query: 364 QARNELLF-----------KWKDSSIDQLLQ-----RASSLRPSRVL----LDDGP--RA 401
RNEL+ ++ S +D L Q +A ++ V+ L GP R
Sbjct: 563 YVRNELIHGKSAPPTETSQRFIQSYVDLLFQIRQAPQADLVKGKHVVRTVPLKGGPKYRV 622
Query: 402 VQVLPSSWSRPLSGVCKMNIDASV-TSSKEAGAGMVMRNNHGEVLAAATTELGPVLSPVL 460
+ W RP G K+N+D S SS + G GM++RN+ G+++ + L +P+
Sbjct: 623 LNNHQPCWERPKDGWMKLNVDGSFDASSGKGGLGMILRNSAGDIIFTSCKPLERCNNPLE 682
Query: 461 AEALGMRWALRLATKLGFRRIWIETDCLELYTFWNRQGGEFSHLATVVFDCRSLLPNFDV 520
+E L+LA I +ETDC + G +FS LA ++ + R LL V
Sbjct: 683 SELRACVEGLKLAIHWTLLPIQVETDCASVVQLLQGTGRDFSVLANIIHEARHLLVGERV 742
Query: 521 FNFTFVRRTSNGAADALARLA-FHYGCMIWIEEAPSEITSITQDD 564
+ + R+ N + LA A + W+ E+ + I+ + ++
Sbjct: 743 ISIKKICRSQNCISHHLANTARADFLAGFWLGESCNLISHLLSEN 787
>gb|AAQ19327.1| bZIP-like protein [Oryza sativa (japonica cultivar-group)]
Length = 2367
Score = 221 bits (563), Expect = 5e-56
Identities = 157/535 (29%), Positives = 243/535 (45%), Gaps = 36/535 (6%)
Query: 11 AIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGFRDF 70
AIP YVM F L + + + ++ F+W R HW WD+L +PK GG+GFRD+
Sbjct: 1533 AIPVYVMGIFKLPESVIDDLTKLTKNFWWDSMNGQRKTHWKAWDSLTKPKSLGGLGFRDY 1592
Query: 71 KAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQKSAWIF 130
+ FN ALLA+ WR++T S+ A++ KA YFP S ++ G S W S++ +
Sbjct: 1593 RLFNQALLARQAWRLITYPDSLCARVLKAKYFPHGSLIDTSFGSNSSPAWRSIEYGLDLL 1652
Query: 131 QEGGLWRVGNGKNIRA*EDKWVPHG---GPLVYRNDVAESLGVVHVSDLMVTNGGCWDRN 187
++G +WRVGNG +IR D W+P P+ + + + VSDL +T G WD
Sbjct: 1653 KKGIIWRVGNGNSIRIWRDSWLPRDHSRRPITGK----ANCRLKWVSDL-ITEDGSWDVP 1707
Query: 188 KVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEASSSS 247
K+ A IL+I + + +D W ++G +S R+ Y + + E+SSS
Sbjct: 1708 KIHQYFHNLDAEVILNICISSRSEEDFIAWHPDKNGMFSVRSAYRLAAQLVNIEESSSSG 1767
Query: 248 ARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEVEETV 307
+ A W +WK + AWR + L +R + + C C E
Sbjct: 1768 TNNINKA-WEMIWKCKVPQKVKIFAWRVASNCLATMVNKKKRKLEQSDMCQICDRENEDD 1826
Query: 308 HHALFSCPLVRMIW--FASPLNLRFDEEGSVHDFFLSF--LATADTNIAGIFLAVLYSVW 363
HAL C +W ++ D + SV F F L +FL L+ W
Sbjct: 1827 AHALCRCIQASQLWSCMHKSGSVSVDIKASVLGRFWLFDCLEKIPEYEQAMFLMTLWRNW 1886
Query: 364 QARNELLF-----------KWKDSSIDQLLQ-----RASSLRPSRVL----LDDGP--RA 401
RNEL+ ++ S +D L Q +A ++ V+ L GP R
Sbjct: 1887 YVRNELIHGKSAPPTETSQRFIQSYVDLLFQIRQAPQADLVKGKHVVRTVPLKGGPKYRV 1946
Query: 402 VQVLPSSWSRPLSGVCKMNIDASV-TSSKEAGAGMVMRNNHGEVLAAATTELGPVLSPVL 460
+ W RP G K+N+D S SS + G GM++RN+ G+++ + L +P+
Sbjct: 1947 LNNHQPCWERPKDGWMKLNVDGSFDASSGKGGLGMILRNSAGDIIFTSCKPLERCNNPLE 2006
Query: 461 AEALGMRWALRLATKLGFRRIWIETDCLELYTFWNRQGGEFSHLATVVFDCRSLL 515
+E L+LA I +ETDC + G +FS LA ++ + R LL
Sbjct: 2007 SELRACVEGLKLAIHWTLLPIQVETDCASVVQLLQGIGRDFSVLANIIHEARHLL 2061
>gb|AAX94970.1| retrotransposon protein, putative, unclassified [Oryza sativa
(japonica cultivar-group)]
Length = 816
Score = 219 bits (557), Expect = 2e-55
Identities = 153/580 (26%), Positives = 246/580 (42%), Gaps = 32/580 (5%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
I + A+P+YVM+ F L G C +MI +F+WG + + R HW W L +PK GG
Sbjct: 63 IKSVIQALPTYVMNVFKLPSGFCEDYMKMIRKFWWGEEKNRRKTHWTSWQQLIKPKSKGG 122
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
IGF+D K FN ALLA+ WR++ L S+ AK+ KA YFP ++ S VW +
Sbjct: 123 IGFKDLKIFNQALLARQAWRLIQFLDSLCAKLMKARYFPNGHLIDTVFPTDSSEVWRGIC 182
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGVVHVSDLMVTNGGCW 184
+ ++G +WR+G G + D W+P L + + V+DL+ N W
Sbjct: 183 HGLELLKKGIIWRIGKGDQLHIWRDNWIPRDHQLRVTGKRTRTC-LKWVADLVSPNNQEW 241
Query: 185 DRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSF-I*KKFSEGEA 243
D + + +PP IL I LP DD W R G ++ R+ Y + ++ EA
Sbjct: 242 DEGLIRQIFYPPYVDEILKIKLPESPTDDFLAWHPERLGVFTVRSAYKLGLAEEHRANEA 301
Query: 244 SSSSARVM-PPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSE 302
+ S+ LW +W A + C AWR ++ LP + +R +V + C C
Sbjct: 302 GACSSNGNGDRVLWKNIWSAQVPQKVCIFAWRLAVERLPTQHNKQKRNIVPSARCEVCGA 361
Query: 303 VEETVHHALFSCPLVRMI------WFASPLNLRFDEEGSVHDFFLSFLATADTNIAGIFL 356
E HA C + + ++ P +F +G D+ L L T D L
Sbjct: 362 PREDGFHATVVCTKAKALRDELRAFWPIPPEDKFVRQGP--DWLLLLLDTLDMRNKAQML 419
Query: 357 AVLYSVWQARNELLFKWK----DSSIDQLLQRASSLRPSRVLLDDGPRAVQVLPSS---- 408
+L+ W RN + SI L L P R D + +P
Sbjct: 420 LMLWRAWHLRNNSVHDTGTLTISGSIRFLQSYVQLLFPIRQKEDPKGKCAAQVPGRLNQN 479
Query: 409 -----------WSRPLSGVCKMNIDASVT-SSKEAGAGMVMRNNHGEVLAAATTELGPVL 456
W P G K+N+D + + + G G++ R++ G+ L ++ L
Sbjct: 480 PGVTKADCLQIWEPPPEGWAKINVDGAFSMTDNTGGIGVIARDSEGKALLSSWKYLHRCA 539
Query: 457 SPVLAEALGMRWALRLATKLGFRRIWIETDCLELYTFWNRQGGEFSHLATVVFDCRSLLP 516
E L ++LA + + I +E+DC+ + + E S ++ ++++
Sbjct: 540 DAEQVEILACYEGMKLAAEWIRKPIILESDCVTVIGRMTAEDEERSRWTFLIRSAKAVMR 599
Query: 517 NFDVFNFTFVRRTSNGAADALARLAFHYG-CMIWIEEAPS 555
+ +R N A LA+LA C +W + PS
Sbjct: 600 SLQEVRIQHRKRECNRVAHELAQLAKRTAHCAVWRDHVPS 639
>gb|AAP53315.1| putative reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|37533452|ref|NP_921028.1| putative
reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|20279456|gb|AAM18736.1| putative
reverse transcriptase [Oryza sativa (japonica
cultivar-group)]
Length = 1509
Score = 218 bits (554), Expect = 5e-55
Identities = 143/496 (28%), Positives = 217/496 (42%), Gaps = 44/496 (8%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
I + IP++ M CF L LC QI +MI++++W MHWL W+ L PK GG
Sbjct: 994 IKAVAQVIPTFAMGCFELTKDLCDQISKMIAKYWWSNQEKDNKMHWLSWNKLTLPKNMGG 1053
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
+GFRD FN A+LAK WR++ S+ +++ +A YFP K+ SY W S+Q
Sbjct: 1054 LGFRDIYIFNLAMLAKQGWRLIQDPDSLCSRVLRAKYFPLGDCFRPKQTSNVSYTWRSIQ 1113
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGVVHVSDLMVTNGGCW 184
K + Q G +WRVG+G I D W+P G +L V V +L+ G W
Sbjct: 1114 KGLRVLQNGMIWRVGDGSKINIWADPWIPRGWSRKPMTPRGANL-VTKVEELIDPYTGTW 1172
Query: 185 DRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEAS 244
D + + W AI SIP+ ++ +D W GC++ ++ Y ++ E AS
Sbjct: 1173 DEDLLSQTFWEEDVAAIKSIPVHVEM-EDVLAWHFDARGCFTVKSAYKV--QREMERRAS 1229
Query: 245 S------SSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCP 298
S+ W KLWK + WR C + L +R+ LH RG+ + C
Sbjct: 1230 RNGCPGVSNWESGDDDFWKKLWKLGVPGKIKHFLWRMCHNTLALRANLHHRGMDVDTRCV 1289
Query: 299 RCSEVEETVHHALFSCPLVRMIWFA---SPLNLRFDEEGSVHDFFLSFLATADTNIAGIF 355
C E H F C V+ +W A L +++ S + S + N
Sbjct: 1290 MCGRYNEDAGHLFFKCKPVKKVWQALNLEELRSMLEQQTSGKNVLQSIYCRPE-NERTSA 1348
Query: 356 LAVLYSVWQARNELLFKWKDSSIDQLLQRASSLRPSRVLLDDGPRAVQVLPSSWSRPLSG 415
+ L+ W+ RNE + PR + + W RP
Sbjct: 1349 IVCLWQWWKERNE---------------------------EKSPRTGEC--AVWRRPPLN 1379
Query: 416 VCKMNIDASVTSS-KEAGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRWALRLAT 474
K+N D + +S+ K+ G G V+R+ G VL A + AE + A++ A+
Sbjct: 1380 FVKINTDGAYSSNMKQGGWGFVIRDQTGAVLQAGAGPAAYLQDAFHAEVVACAAAIKTAS 1439
Query: 475 KLGFRRIWIETDCLEL 490
+ G RI +ETD + L
Sbjct: 1440 ERGMSRIELETDSMML 1455
>gb|AAD21778.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25410938|pir||G84429 hypothetical protein
At2g01840 [imported] - Arabidopsis thaliana
Length = 1715
Score = 214 bits (546), Expect = 5e-54
Identities = 158/559 (28%), Positives = 260/559 (46%), Gaps = 47/559 (8%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
I I +A+P Y M+CFLL +C++I +I+ F+WG + G
Sbjct: 1155 IKAIAMALPVYSMNCFLLPTLICNEINSLITAFWWGKENE------------------GD 1196
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
+GF+D FN ALLAK WRI+T +S+LA++YK +Y+P ++L A KG SY W S+Q
Sbjct: 1197 LGFKDLHQFNRALLAKQAWRILTNPQSLLARLYKGLYYPNTTYLRANKGGHASYGWNSIQ 1256
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGVVHVSDLMVTNGGCW 184
+ + Q+G R+G+G+ + ED W+P P R + + + V+DL N W
Sbjct: 1257 EGKLLLQQGLRVRLGDGQTTKIWEDPWLPTLPPRPARGPILDE--DMKVADLWRENKREW 1314
Query: 185 DRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*K-KFSEGEA 243
D E + P + S+ L D + W TR+ Y+ R+GY +E E
Sbjct: 1315 DPVIFEGVLNPEDQQLAKSLYLSNYAARDSYKWAYTRNTQYTVRSGYWVATHVNLTEEEI 1374
Query: 244 SSSSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEV 303
+ +P L ++W+ P+ WR L + L R + +P C RC
Sbjct: 1375 INPLEGDVP--LKQEIWRLKITPKIKHFIWRCLSGALSTTTQLRNRNIPADPTCQRCCNA 1432
Query: 304 EETVHHALFSCPLVRMIW----FASPLNLRFDEEGSVHDFFLSFLATADTNIA---GIF- 355
+ET++H +F+C +++W F+ L F + + L + N+ G+
Sbjct: 1433 DETINHIIFTCSYAQVVWRSANFSGSNRLCF-TDNLEENIRLILQGKKNQNLPILNGLMP 1491
Query: 356 LAVLYSVWQARNELLFKWKDSSIDQLLQRASSLRPSRV--LLDDGPRAVQVLPSS----- 408
+++ +W++RNE LF+ D ++ Q+A V +++D + S+
Sbjct: 1492 FWIMWRLWKSRNEYLFQQLDRFPWKVAQKAEQEATEWVETMVNDTAISHNTAQSNDRPLS 1551
Query: 409 ----WSRPLSGVCKMNIDASVTSSKE-AGAGMVMRNNHGEVLAAATTELGPVLSPVLAEA 463
WS P G K N D+ ++ G ++R+ +G VL + +L S + AEA
Sbjct: 1552 RSKQWSSPPEGFLKCNFDSGYVQGRDYTSTGWILRDCNGRVLHSGCAKLQQSYSALQAEA 1611
Query: 464 LGMRWALRLATKLGFRRIWIETDCLELYTFWNRQGGEFSH-LATVVFDCRSLLPNFDVFN 522
LG AL++ G+ +W E D LEL N+ E H L T+++D R + +
Sbjct: 1612 LGFLHALQMVWIRGYCYVWFEGDNLELTNLINKT--EDHHLLETLLYDIRFWMTKLPFSS 1669
Query: 523 FTFVRRTSNGAADALARLA 541
+V R N AAD L + A
Sbjct: 1670 IGYVNRERNLAADKLTKYA 1688
>gb|AAD24831.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25408166|pir||G84721 hypothetical protein
At2g31520 [imported] - Arabidopsis thaliana
Length = 1524
Score = 212 bits (540), Expect = 2e-53
Identities = 164/567 (28%), Positives = 261/567 (45%), Gaps = 41/567 (7%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
+ + LA+P Y MSCF L G+ S+IE ++ F+W + R + W+ W L K GG
Sbjct: 949 LKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRLQYSKKEGG 1008
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
+GFRD FN ALLAK WR++ S+ A++ KA YF S L+AK + SY W SL
Sbjct: 1009 LGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKDVSILDAKVRKQQSYGWASLL 1068
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGVVHVSDLMVTNGG-- 182
+ ++G +G+G+NIR D V P + E+ + +++L G
Sbjct: 1069 DGIALLKKGTRHLIGDGQNIRIGLDNIVDSHPPRPLNTE--ETYKEMTINNLFERKGSYY 1126
Query: 183 CWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGE 242
WD +K+ I I L ++ D W G Y+ R+GY + +
Sbjct: 1127 FWDDSKISQFVDQSDHGFIHRIYLAKSKKPDKIIWNYNTTGEYTVRSGYWLL-----THD 1181
Query: 243 ASSSSARVMPP----ALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCP 298
S++ + PP L ++W +P+ WRA L L RG+ +P CP
Sbjct: 1182 PSTNIPAINPPHGSIDLKTRIWNLPIMPKLKHFLWRALSQALATTERLTTRGMRIDPICP 1241
Query: 299 RCSEVEETVHHALFSCPLVRMIWFASPLNLRFDEEGSVHDF------FLSFLATADTNIA 352
RC E+++HALF+CP M W+ S +L ++ S +DF L+F+ DT ++
Sbjct: 1242 RCHRENESINHALFTCPFATMAWWLSDSSLIRNQLMS-NDFEENISNILNFV--QDTTMS 1298
Query: 353 GIF----LAVLYSVWQARNELLF-KWKDSSIDQLLQRAS------SLRPSRVLLDDGPRA 401
+ +++ +W+ARN ++F K+++S +L + + S R
Sbjct: 1299 DFHKLLPVWLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNATQSHKKTPSPTRQ 1358
Query: 402 VQVLPSSWSRPLSGVCKMNIDASVTSSK-EAGAGMVMRNNHGEVLAAATTELGPVLSPVL 460
+ W P + K N DA K EA G ++RN++G ++ + +L +P+
Sbjct: 1359 IAENKIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLAHTSNPLE 1418
Query: 461 AEALGMRWALRLATKLGFRRIWIETDCLELYTFWNRQGGEF-SHLATVVFDCRSLLPNFD 519
AE + AL+ G+ ++++E DC L N G F S LA + D F
Sbjct: 1419 AETKALLAALQQTWIRGYTQVFMEGDCQTLINLIN--GISFHSSLANHLEDISFWANKFA 1476
Query: 520 VFNFTFVRRTSNGAADALARLAFHYGC 546
F F+RR N A LA+ YGC
Sbjct: 1477 SIQFGFIRRKGNKLAHVLAK----YGC 1499
>gb|AAK71569.2| putative reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|50918647|ref|XP_469720.1| putative
reverse transcriptase [Oryza sativa (japonica
cultivar-group)]
Length = 1833
Score = 211 bits (538), Expect = 4e-53
Identities = 154/577 (26%), Positives = 242/577 (41%), Gaps = 76/577 (13%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
I + A+P+Y+M F + D C RM+ F+WG D R +HW+ W+ + PK GG
Sbjct: 705 IKSVAQALPTYIMGVFKMTDKFCDDYTRMVRNFWWGHDNGERKVHWIAWEKILYPKSHGG 764
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
+GFRD K FN ALLA+ WR++T S+ A+++KA Y+P + ++ S VW +Q
Sbjct: 765 LGFRDMKCFNQALLARQAWRLLTSPDSLCARVFKAKYYPNGNLVDTAFPTVSSSVWKGIQ 824
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLG---VVHVSDLMVTNG 181
+ ++G +WRVGNG NI+ KWVPHG + +V E G +++V++L+
Sbjct: 825 HGLDLLKQGTIWRVGNGHNIKIWRHKWVPHGEHI----NVLEKKGRNRLIYVNELINEEN 880
Query: 182 GCWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEG 241
CW+ V + A +L I LP Q DD W + G ++ ++ Y +G
Sbjct: 881 RCWNETLVRHVLKEEDANEVLKIRLPNHQMDDFPAWHHEKSGLFTVKSAYKLAWNLSGKG 940
Query: 242 --EASSSSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPR 299
++SSS+A +W ++W A + W+ D LP RR + N CP
Sbjct: 941 VVQSSSSTATSGERKIWSRVWNAKVQAKVKIFIWKLAQDKLPTWENKRRRKIEMNGTCPV 1000
Query: 300 CSEVEETVHHALFSC-------PLVRMIWFASPLNLRFDEEGSVHDFFLSFLATADTNIA 352
C E +HA C +R +W P +F G D+ L L +
Sbjct: 1001 CGTKGENSYHATVECTKARALREALRAVWHL-PGEDKFLWTGP--DWLLILLDGVNEEQR 1057
Query: 353 GIFLAVLYSVWQARNELLFKWKDSSIDQLLQRASS----LRPSRVLLDDGPRAVQVLPSS 408
+ +L+ W RN+L+ SI + +S L P+R + DD
Sbjct: 1058 THIMYMLWRAWYLRNDLIHGDGRCSIAGSVSFLTSYEEVLLPNRQMPDD---------IK 1108
Query: 409 WSRPLSGVCKMNIDASVTSSKEAGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRW 468
+P+S EA A L L + PV+ E
Sbjct: 1109 GKKPIS-----------AEQAEALAS----------LEGIRCALQWIHMPVILETDNAEV 1147
Query: 469 ALRLATKLGFRRIWIETDCLELYTFWNRQGGEFSHLATVVFDCRSLLPNFDVFNFTFVRR 528
RL TK R +W V+ + ++ + ++R
Sbjct: 1148 VARLKTKHSSRSVW----------------------EGVIMEAKAAMQGLQAVEVAHIKR 1185
Query: 529 TSNGAADALARLAFHYG-CMIWIEEAPSEITSITQDD 564
SN A LA++A G C+ W AP+EI + +
Sbjct: 1186 DSNKVAHTLAQMALSSGNCLEWRLCAPAEILELLNQE 1222
>gb|AAD20714.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25412331|pir||G84649 hypothetical protein
At2g25550 [imported] - Arabidopsis thaliana
Length = 1750
Score = 209 bits (533), Expect = 1e-52
Identities = 163/567 (28%), Positives = 260/567 (45%), Gaps = 41/567 (7%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
+ + LA+P Y MSCF L G+ S+IE ++ F+W + R + W+ W L K GG
Sbjct: 1175 LKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQRGIPWVAWKRLQYSKKEGG 1234
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
+GFRD FN ALLAK WR++ S+ A++ KA YF S L+AK + SY W SL
Sbjct: 1235 LGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKDVSILDAKVRKQQSYGWASLL 1294
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGVVHVSDLMVTNGG-- 182
+ ++G +G+G+NIR D V P + E+ + +++L G
Sbjct: 1295 DGIALLKKGTRHLIGDGQNIRIGLDNIVDSHPPRPLNTE--ETYKEMTINNLFERKGSYY 1352
Query: 183 CWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGE 242
WD +K+ I I L ++ D W G Y+ R+GY + +
Sbjct: 1353 FWDDSKISQFVDQSDHGFIHRIYLAKSKKPDKIIWNYNTTGEYTVRSGYWLL-----THD 1407
Query: 243 ASSSSARVMPP----ALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCP 298
S++ + PP L ++W +P+ WRA L L RG+ +P CP
Sbjct: 1408 PSTNIPAINPPHGSIDLKTRIWNLPIMPKLKHFLWRALSQALATTERLTTRGMRIDPSCP 1467
Query: 299 RCSEVEETVHHALFSCPLVRMIWFASPLNLRFDEEGSVHDF------FLSFLATADTNIA 352
RC E+++HALF+CP M W S +L ++ S +DF L+F+ DT ++
Sbjct: 1468 RCHRENESINHALFTCPFATMAWRLSDSSLIRNQLMS-NDFEENISNILNFV--QDTTMS 1524
Query: 353 GIF----LAVLYSVWQARNELLF-KWKDSSIDQLLQRAS------SLRPSRVLLDDGPRA 401
+ +++ +W+ARN ++F K+++S +L + + S R
Sbjct: 1525 DFHKLLPVWLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNATQSHKKTPSPTRQ 1584
Query: 402 VQVLPSSWSRPLSGVCKMNIDASVTSSK-EAGAGMVMRNNHGEVLAAATTELGPVLSPVL 460
+ W P + K N DA K EA G ++RN++G ++ + +L +P+
Sbjct: 1585 IAENKIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPISWGSMKLAHTSNPLE 1644
Query: 461 AEALGMRWALRLATKLGFRRIWIETDCLELYTFWNRQGGEF-SHLATVVFDCRSLLPNFD 519
AE + AL+ G+ ++++E DC L N G F S LA + D F
Sbjct: 1645 AETKALLAALQQTWIRGYTQVFMEGDCQTLINLIN--GISFHSSLANHLEDISFWANKFA 1702
Query: 520 VFNFTFVRRTSNGAADALARLAFHYGC 546
F F+R+ N A LA+ YGC
Sbjct: 1703 SIQFGFIRKKGNKLAHVLAK----YGC 1725
>gb|AAG51783.1| reverse transcriptase, putative; 16838-20266 [Arabidopsis thaliana]
gi|25405244|pir||E96519 probable reverse transcriptase,
16838-20266 [imported] - Arabidopsis thaliana
Length = 1142
Score = 209 bits (533), Expect = 1e-52
Identities = 148/554 (26%), Positives = 254/554 (45%), Gaps = 28/554 (5%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
I + +P YVMSCF L + S++ +++F+W + R MHW+ WD LC K GG
Sbjct: 577 IKSVAATLPRYVMSCFRLPKAITSKLTSAVAKFWWSSNGDSRGMHWMAWDKLCSSKSDGG 636
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
+GFR+ FN+ALLAK WR++T S+ AK++K YF K + L++ K Y PSY W S+
Sbjct: 637 LGFRNVDDFNSALLAKQLWRLITAPDSLFAKVFKGRYFRKSNPLDSIKSYSPSYGWRSMI 696
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVP--HGGPLVYRNDVAESLGVVHVSDLMVTNGG 182
+ + +G + RVG+G +I D W+P P Y + + + V L+ +
Sbjct: 697 SARSLVYKGLIKRVGSGASISVWNDPWIPAQFPRPAKYGGSIVDP--SLKVKSLIDSRSN 754
Query: 183 CWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGE 242
W+ + ++ L P I ++P+ +D W T+ G Y+ ++GY +EG
Sbjct: 755 FWNIDLLKELFDPEDVPLISALPIGNPNMEDTLGWHFTKAGNYTVKSGYHTARLDLNEG- 813
Query: 243 ASSSSARVMPPALWMK--LWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRC 300
+ + P +K +WK P+ W+ +PV L +RG++ + C C
Sbjct: 814 ----TTLIGPDLTTLKAYIWKVQCPPKLRHFLWQILSGCVPVSENLRKRGILCDKGCVSC 869
Query: 301 SEVEETVHHALFSCPLVRMIWFASPLNLR---FDEEGSVHDFFLSFLATADTNIAGIFLA 357
EE+++H LF C R IW S + F + F + +
Sbjct: 870 GASEESINHTLFQCHPARQIWALSQIPTAPGIFPSNSIFTNLDHLFWRIPSGVDSAPYPW 929
Query: 358 VLYSVWQARNELLFKWKDSSIDQLL----QRASSLRPSRVLLDDGPRA-------VQVLP 406
+++ +W+ARNE +F+ D ++L + A S + ++V L ++V
Sbjct: 930 IIWYIWKARNEKVFENVDKDPMEILLLAVKEAQSWQEAQVELHSERHGSLSIDSRIRVRD 989
Query: 407 SSWSRPLSGVCKMNIDASVTSSKE-AGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALG 465
S SG + ID S +S + +G G ++ GE + LSP+ E
Sbjct: 990 VSQDTTFSGF-RCFIDGSWKASDQFSGTGWFCLSSLGESPTMGAANVRRSLSPLHTEMEA 1048
Query: 466 MRWALRLATKLGFRRIWIETDCLELYTFWNRQGGEFSHLATVVFDCRSLLPNFDVFNFTF 525
+ WA++ + + TDC +L + E+ + + + +S F F+ +
Sbjct: 1049 LLWAMKCMIGADNQNVAFFTDCSDLVKMVS-SPTEWPAFSVYLEELQSDREEFTNFSLSL 1107
Query: 526 VRRTSNGAADALAR 539
+ R++N AD LAR
Sbjct: 1108 ISRSANVKADKLAR 1121
>gb|AAP54692.1| putative reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|37536206|ref|NP_922405.1| putative
reverse transcriptase [Oryza sativa (japonica
cultivar-group)] gi|27311287|gb|AAO00713.1|
retrotransposon protein, putative, unclassified [Oryza
sativa (japonica cultivar-group)]
Length = 1557
Score = 203 bits (516), Expect = 1e-50
Identities = 115/376 (30%), Positives = 187/376 (49%), Gaps = 13/376 (3%)
Query: 11 AIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGFRDF 70
AIP YVM F L + +C + + F+WG + R HW W +L + K GG+GF+D
Sbjct: 1168 AIPVYVMGIFKLPESVCEDLTSLTRNFWWGAEKGKRKTHWKAWKSLTKSKSLGGLGFKDI 1227
Query: 71 KAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQKSAWIF 130
+ FN ALLA+ WR++ S+ A++ KA Y+P S ++ G S W +++ +
Sbjct: 1228 RLFNQALLARQAWRLIDNPDSLCARVLKAKYYPNGSIVDTSFGGNASPGWQAIEHGLELV 1287
Query: 131 QEGGLWRVGNGKNIRA*EDKWVPHG---GPLVYRNDVAESLGVVHVSDLMVTNGGCWDRN 187
++G +WR+GNG+++R +D W+P P+ +N+ + V+DLM+ N G WD N
Sbjct: 1288 KKGIIWRIGNGRSVRVWQDPWLPRDLSRRPITPKNNCR----IKWVADLMLDN-GMWDAN 1342
Query: 188 KVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEASSSS 247
K+ + P IL + + +D W + G +S RT Y + +++ EASSSS
Sbjct: 1343 KINQIFLPVDVEIILKLRTSSRDEEDFIAWHPDKLGNFSVRTAYRLA-ENWAKEEASSSS 1401
Query: 248 ARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEVEETV 307
+ V W LWK + + WRA + LP +R + + C C +E
Sbjct: 1402 SDVNIRKAWELLWKCNVPSKVKIFTWRATSNCLPTWDNKKKRNLEISDTCVICGMEKEDT 1461
Query: 308 HHALFSCPLVRMIWFA----SPLNLRFDEEGSVHDFFLSFLATADTNIAGIFLAVLYSVW 363
HAL CP + +W A + L+LR D+ + + LA + +FL VL+ +W
Sbjct: 1462 MHALCRCPQAKHLWLAMKESNDLSLRMDDHLLGPSWLFNRLALLPDHEQPMFLMVLWRIW 1521
Query: 364 QARNELLFKWKDSSID 379
NE++ SI+
Sbjct: 1522 FVHNEIIHGKPSPSIE 1537
>gb|AAD32950.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25411805|pir||C84554 hypothetical protein
At2g17610 [imported] - Arabidopsis thaliana
Length = 773
Score = 199 bits (507), Expect = 2e-49
Identities = 149/525 (28%), Positives = 235/525 (44%), Gaps = 36/525 (6%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
I I AIP+Y MSCFLL L QI + F+W + W+ W L PK GG
Sbjct: 248 IKSIVTAIPTYSMSCFLLPMRLIHQITSAMRWFWWSNTKVKHKIPWVAWSKLNDPKKMGG 307
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
+ RD K FN ALLAK WRI+ + S++A+++KA YFPKE L+AK + SY W S+
Sbjct: 308 LAIRDLKDFNIALLAKQSWRILQQPFSLMARVFKAKYFPKERLLDAKATSQSSYAWKSIL 367
Query: 125 KSAWIFQEGGLWRVGNGKNIRA*EDKWVPHGGPLVYRNDVAESLGVVHVSDLMVTNGGCW 184
+ G + GNG NI+ +D W+P P + VSDL++ G W
Sbjct: 368 HGTKLISRGLKYIAGNGNNIQLWKDNWLPLNPPRPPVGTCDSIYSQLKVSDLLIE--GRW 425
Query: 185 DRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEAS 244
+ + + L I +I + +D W T DG YS ++GY + K + AS
Sbjct: 426 NEDLLCKLIHQNDIPHIRAIRPSITGANDAITWIYTHDGNYSVKSGYHLLRKLSQQQHAS 485
Query: 245 SSSAR-VMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCSEV 303
S V ++ +WK +A P+ WR+ + LP L RR ++ + C RC E
Sbjct: 486 LPSPNEVSAQTVFTNIWKQNAPPKIKHFWWRSAHNALPTAGNLKRRRLITDDTCQRCGEA 545
Query: 304 EETVHHALFSCPLVRMIWFASPLNLRFDEEGSVHDFFLSFLATADTNIAG-----IFLAV 358
E V+H LF C + + IW + + L + + F + + N + +F +
Sbjct: 546 SEDVNHLLFQCRVSKEIWEQAHIKLCPGDSLMSNSFNQNLESIQKLNQSARKDVSLFPFI 605
Query: 359 LYSVWQARNELLFKWKDSSIDQLLQRASSLRPSRVLLDDGPRAVQVLPSSWSRPLSGVC- 417
+ +W+ RN+L+F K SI +Q+A L+D W L+ C
Sbjct: 606 GWRIWKMRNDLIFNNKRWSIPDSIQKA--------LIDQ---------QQWKESLN--CN 646
Query: 418 -----KMNIDASVTSSKEAGAGMVMRN-NHGEVLAAATTELGPVLSPVLAEALGMRWALR 471
K + S + G +R + + + + + P +P+ AEA ++ A+
Sbjct: 647 EQQQRKPQLHDSHNNKCYGGHLKTLRTLSKAQFVLRGMSSIPPTSTPLEAEAEALKLAMI 706
Query: 472 LATKLGFRRIWIETDCLELYTFWNR--QGGEFSHLATVVFDCRSL 514
+LG+ I D L+ R E+S +AT + D ++
Sbjct: 707 HLQRLGYEDIIFHGDVAALFQPITRTTHSCEYSPIATFLRDIENI 751
>gb|AAD24601.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25411729|pir||H84542 hypothetical protein
At2g16680 [imported] - Arabidopsis thaliana
Length = 1319
Score = 194 bits (494), Expect = 5e-48
Identities = 134/489 (27%), Positives = 225/489 (45%), Gaps = 17/489 (3%)
Query: 5 ITKINLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGG 64
+ I LA+P Y+M+CF L GLC+++ ++ F+W +HW+ L PK GG
Sbjct: 745 LKSIALALPVYIMTCFRLPKGLCTKLTSVMMDFWWNSMEFSNKIHWIGGKKLTLPKSLGG 804
Query: 65 IGFRDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQ 124
GF+D + FN ALLAK WR+ + KS++++I+K+ YF FL A++G RPSY W S+
Sbjct: 805 FGFKDLQCFNQALLAKQAWRLFSDSKSIVSQIFKSRYFMNTDFLNARQGTRPSYTWRSIL 864
Query: 125 KSAWIFQEGGLWR-VGNGKNIRA*EDKWVPHG---GPLVYRNDVAESLGVVHVSDLMVTN 180
+ GGL R +GNG+ DKW+ G P+ + + + V H+ D + N
Sbjct: 865 YGRELL-NGGLKRLIGNGEQTNVWIDKWLFDGHSRRPMNLHSLMNIHMKVSHLIDPLTRN 923
Query: 181 GGCWDRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSE 240
W+ K+ L + I+ PL +D + W T +G Y+ ++GY ++ +
Sbjct: 924 ---WNLKKLTELFHEKDVQLIMH-QRPLISSEDSYCWAGTNNGLYTVKSGYERSSRETFK 979
Query: 241 GEASSSSARVMPPALWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRC 300
+ L+ K+W + VP+ W+A L V L RG+ C C
Sbjct: 980 NLFKEADVYPSVNPLFDKVWSLETVPKIKVFMWKALKGALAVEDRLRSRGIRTADGCLFC 1039
Query: 301 SEVEETVHHALFSCPLVRMIWFASPLNLRFDEEGSVHDFFLSFLATADTN------IAGI 354
E ET++H LF CP R +W S + G+ ++ + N + +
Sbjct: 1040 KEEIETINHLLFQCPFARQVWALSLIQAPATGFGTSIFSNINHVIQNSQNFGIPRHMRTV 1099
Query: 355 FLAVLYSVWQARNELLFKWKDSSIDQLLQRASSLRPSRV-LLDDGPRAVQVLPSSWSRPL 413
+L+ +W+ RN+ LF+ + +++ +A + + V W+ P
Sbjct: 1100 SPWLLWEIWKNRNKTLFQGTGLTSSEIVAKAYEECNLWINAQEKSSGGVSPSEHKWNPPP 1159
Query: 414 SGVCKMNIDASVTSSKE-AGAGMVMRNNHGEVLAAATTELGPVLSPVLAEALGMRWALRL 472
+G K NI + + K+ AG V+R++ G+VL + V S A+ WAL
Sbjct: 1160 AGELKCNIGVAWSRQKQLAGVSWVLRDSMGQVLLHSRRSYSQVYSLFDAKIKSWDWALES 1219
Query: 473 ATKLGFRRI 481
F ++
Sbjct: 1220 MDHFHFDKV 1228
>gb|AAF18538.1| Very similar to retrotransposon reverse transcriptase [Arabidopsis
thaliana] gi|25518314|pir||A86359 hypothetical protein
F12K8.9 - Arabidopsis thaliana
Length = 1231
Score = 189 bits (479), Expect = 3e-46
Identities = 146/520 (28%), Positives = 233/520 (44%), Gaps = 39/520 (7%)
Query: 8 INLAIPSYVMSCFLLLDGLCSQIERMISRFYWGGDVSHRTMHWLKWDTLCRPKGCGGIGF 67
+ +A+P + MSCF L C+ + ++ F+W + R +HWL W+ LC PK GG+GF
Sbjct: 644 VAMAMPVFAMSCFKLPKTTCTNLASAMADFWWSANDHQRKIHWLSWEKLCLPKEHGGLGF 703
Query: 68 RDFKAFNTALLAKNWWRIMTKLKSMLAKIYKAVYFPKESFLEAKKGYRPSYVWLSLQKSA 127
RD FN ALLAK WR++ + A++ K+ YFP FL++ G RPS+ W S+
Sbjct: 704 RDIGLFNQALLAKQAWRLLQFPDCLFARLIKSRYFPVGEFLDSDVGSRPSFGWRSILHGR 763
Query: 128 WIFQEGGLWRVGNGKNIRA*EDKWVPHGG---PLVYRNDVAESLGVVHVSDLMVTNGGCW 184
+ G + RVGNGK+IR D W+ G P + + L VSDL+ W
Sbjct: 764 DLLCRGLVKRVGNGKSIRVWIDYWLDDNGLRAPWIKNPIINVDL---LVSDLIDYEKRDW 820
Query: 185 DRNKVEFLCWPPTARAILSIPLPLQQRDDCFFWPETRDGCYSTRTGYSFI*KKFSEGEAS 244
+K+E +P I P+ DD + W + G YS + GY ++ + G+ +
Sbjct: 821 RLDKLEEQFFPDDVVKIRE-NRPVVSLDDFWIWKHNKSGDYSVKLGY-WLASNQNLGQVA 878
Query: 245 SSSARVMPPA---LWMKLWKADAVPQCCETAWRACLDILPVRSILHRRGVVGNPCCPRCS 301
+ +M P+ L ++WK P+ W+ +PV +L RG+ + C C
Sbjct: 879 IEA--MMQPSLNDLKTQVWKLQTEPKIKVFLWKVLSGAIPVVDLLSYRGMKLDSRCQTCG 936
Query: 302 EVEETVHHALFSCPLVRMIWFASPLN--LRFDEEGSVHDFFLSFLATAD-----TNIAGI 354
E++ H LFSC R +W S ++ L E GSV+ FL D +
Sbjct: 937 CEGESIQHVLFSCSFPRQVWAMSNIHVPLLGFECGSVYANLYHFLINRDNLKWPVELRRS 996
Query: 355 FLAVLYSVWQARNELLFKWKDSSIDQLL-------------QRASSLRPSRVLLDD---- 397
F +++ +W+ RN F+ K ++ + + Q R + V D
Sbjct: 997 FPWIIWRIWKNRNLFFFEGKRFTVLETILKVRKDVEDWFAAQVVEKERRAEVGQSDQQVF 1056
Query: 398 GPRAVQVLPSSWSRPLSGVCKMNIDASVT-SSKEAGAGMVMRNNHGEVLAAATTELGPVL 456
PR V + W P + K N+ S + ++ AG V+RN+ G VL + +
Sbjct: 1057 SPRNVSPV-VRWLPPPTDWVKCNVGLSWSRRNRLAGVAWVLRNDRGNVLMHSRRAFSNIS 1115
Query: 457 SPVLAEALGMRWALRLATKLGFRRIWIETDCLELYTFWNR 496
S + A+ L + WA+ R+ + L NR
Sbjct: 1116 SFLEAQFLSIVWAVESMVSHHVNRVVFGVEAAVLVGVINR 1155
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.326 0.139 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,009,554,938
Number of Sequences: 2540612
Number of extensions: 42859308
Number of successful extensions: 97117
Number of sequences better than 10.0: 475
Number of HSP's better than 10.0 without gapping: 226
Number of HSP's successfully gapped in prelim test: 249
Number of HSP's that attempted gapping in prelim test: 96157
Number of HSP's gapped (non-prelim): 712
length of query: 565
length of database: 863,360,394
effective HSP length: 133
effective length of query: 432
effective length of database: 525,458,998
effective search space: 226998287136
effective search space used: 226998287136
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0118c.1


