

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0117b.14
(458 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
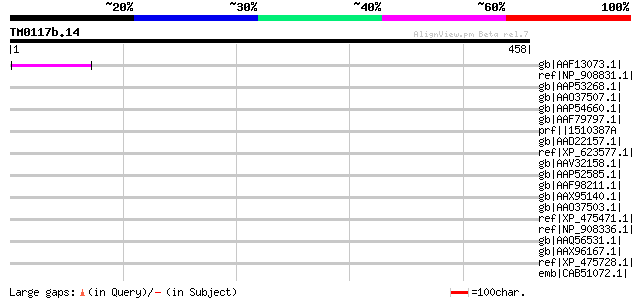
Score E
Sequences producing significant alignments: (bits) Value
gb|AAF13073.1| putative retroelement pol polyprotein [Arabidopsi... 57 1e-06
ref|NP_908831.1| putative polyprotein [Oryza sativa (japonica cu... 47 0.002
gb|AAP53268.1| putative 22 kDa kafirin cluster; Ty3-Gypsy type [... 46 0.002
gb|AAO37507.1| retrotransposon protein, putative, unclassified [... 45 0.006
gb|AAP54660.1| putative plant disease resistance polyprotein [Or... 44 0.014
gb|AAF79797.1| T32E20.30 [Arabidopsis thaliana] 44 0.014
prf||1510387A retrotransposon del1-46 44 0.014
gb|AAD22157.1| hypothetical protein [Sorghum bicolor] 43 0.024
ref|XP_623577.1| PREDICTED: similar to Titin-like protein [Apis ... 43 0.024
gb|AAV32158.1| putative polyprotein [Oryza sativa (japonica cult... 42 0.031
gb|AAP52585.1| putative polyprotein [Oryza sativa (japonica cult... 42 0.040
gb|AAF98211.1| Hypothetical protein [Arabidopsis thaliana] gi|15... 41 0.090
gb|AAX95140.1| retrotransposon protein, putative, Ty3-gypsy sub-... 40 0.20
gb|AAO37503.1| retrotransposon protein, putative, Ty3-gypsy sub-... 39 0.26
ref|XP_475471.1| putative polyprotein [Oryza sativa (japonica cu... 39 0.26
ref|NP_908336.1| putative polyprotein [Oryza sativa (japonica cu... 39 0.26
gb|AAQ56531.1| putative polyprotein [Oryza sativa (japonica cult... 39 0.26
gb|AAX96167.1| retrotransposon protein, putative, Ty3-gypsy sub-... 39 0.34
ref|XP_475728.1| putative polyprotein [Oryza sativa (japonica cu... 39 0.34
emb|CAB51072.1| hypothetical protein [Homo sapiens] 39 0.44
>gb|AAF13073.1| putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1661
Score = 57.0 bits (136), Expect = 1e-06
Identities = 27/71 (38%), Positives = 36/71 (50%)
Query: 2 FPEFDGREAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKATWWE 61
+P ++G A WL +EQ + EEEK A LTG + WW C + R Q TW E
Sbjct: 233 YPAYEGGNADDWLFRLEQCFLSNRTLEEEKLEKAVSCLTGASVTWWRCSKDREQIYTWRE 292
Query: 62 FVEALLRKFEP 72
F E + +F P
Sbjct: 293 FQEKFMLRFRP 303
>ref|NP_908831.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1522
Score = 46.6 bits (109), Expect = 0.002
Identities = 22/70 (31%), Positives = 36/70 (51%)
Query: 2 FPEFDGREAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKATWWE 61
FP+FDG + W+ E + ++E K S A + G+A W+ W++ TW +
Sbjct: 133 FPKFDGTDVCVWVDNCETYFALYQITEGFKVSAASLNMIGDAAHWYQAWKQERGWPTWEQ 192
Query: 62 FVEALLRKFE 71
EA+L +FE
Sbjct: 193 LREAVLNEFE 202
>gb|AAP53268.1| putative 22 kDa kafirin cluster; Ty3-Gypsy type [Oryza sativa
(japonica cultivar-group)] gi|37533358|ref|NP_920981.1|
putative 22 kDa kafirin cluster; Ty3-Gypsy type [Oryza
sativa (japonica cultivar-group)]
gi|21327374|gb|AAM48279.1| Putative 22 kDa kafirin
cluster; Ty3-Gypsy type [Oryza sativa (japonica
cultivar-group)] gi|18767374|gb|AAL79340.1| Putative 22
kDa kafirin cluster; Ty3-Gypsy type [Oryza sativa]
Length = 1230
Score = 46.2 bits (108), Expect = 0.002
Identities = 34/109 (31%), Positives = 49/109 (44%), Gaps = 18/109 (16%)
Query: 3 PEFDGR----EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQK-- 56
P F G EA W++ +E+ EA G +++EK A L AF WW ++ +
Sbjct: 80 PTFSGTANPLEAEEWIVAMEKSFEAMGCTDKEKIIYATYMLQSSAFEWWDAHKKSYSERI 139
Query: 57 -ATWWEFVEALLRKFEPELEPYMPEPVQDSEEEEI----PGKQEVLEAE 100
TW F EA +K Y PE V+ +E+E G + V E E
Sbjct: 140 FITWELFKEAFYKK-------YFPESVKRMKEKEFLELKQGNKSVAEYE 181
>gb|AAO37507.1| retrotransposon protein, putative, unclassified [Oryza sativa
(japonica cultivar-group)] gi|50916505|ref|XP_468649.1|
putative polyprotein [Oryza sativa (japonica
cultivar-group)]
Length = 1155
Score = 44.7 bits (104), Expect = 0.006
Identities = 22/74 (29%), Positives = 39/74 (51%)
Query: 2 FPEFDGREAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKATWWE 61
FP FDG + WL M E + + +++ K S A ++G A W+ ++ N+ +W +
Sbjct: 124 FPRFDGSDVRIWLDMCETYFDMYQITQNFKVSAAVLHMSGNAAQWYHSYKLVNEVNSWDQ 183
Query: 62 FVEALLRKFEPELE 75
F A+ +FE +E
Sbjct: 184 FRMAVATEFECVVE 197
>gb|AAP54660.1| putative plant disease resistance polyprotein [Oryza sativa
(japonica cultivar-group)] gi|37536142|ref|NP_922373.1|
putative plant disease resistance polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|10122034|gb|AAG13423.1| putative plant disease
resistance polyprotein [Oryza sativa (japonica
cultivar-group)]
Length = 894
Score = 43.5 bits (101), Expect = 0.014
Identities = 21/74 (28%), Positives = 38/74 (50%)
Query: 2 FPEFDGREAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKATWWE 61
FP FDG + WL M E + + +++ K S ++G A W+ ++ N+ +W +
Sbjct: 124 FPRFDGSDVRIWLNMCETYFDMYQITQNFKVSAVVLHMSGNAAQWYHSYKLVNEVNSWDQ 183
Query: 62 FVEALLRKFEPELE 75
F A+ +FE +E
Sbjct: 184 FRMAVATEFEGVVE 197
>gb|AAF79797.1| T32E20.30 [Arabidopsis thaliana]
Length = 1397
Score = 43.5 bits (101), Expect = 0.014
Identities = 20/68 (29%), Positives = 34/68 (49%)
Query: 3 PEFDGREAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKATWWEF 62
P FDG YGW+ VE+ + G +E E+ + +++GEA W+ R +W +
Sbjct: 133 PMFDGSGIYGWIARVERFFRSGGYNEAEQLALVSVSVSGEALSWYNWAISRGDFVSWLKL 192
Query: 63 VEALLRKF 70
L+ +F
Sbjct: 193 KSGLMLRF 200
>prf||1510387A retrotransposon del1-46
Length = 1443
Score = 43.5 bits (101), Expect = 0.014
Identities = 26/72 (36%), Positives = 35/72 (48%), Gaps = 4/72 (5%)
Query: 6 DGREAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKAT----WWE 61
D EA+ W+ V + + GV++E+K A L GEA WW R + T W E
Sbjct: 73 DPLEAHRWIRHVTKILDTLGVTDEQKVILASFQLQGEAEFWWDAKVRSREDDTTQIKWDE 132
Query: 62 FVEALLRKFEPE 73
FVE KF P+
Sbjct: 133 FVEVFTEKFFPD 144
>gb|AAD22157.1| hypothetical protein [Sorghum bicolor]
Length = 416
Score = 42.7 bits (99), Expect = 0.024
Identities = 46/189 (24%), Positives = 80/189 (41%), Gaps = 31/189 (16%)
Query: 10 AYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQK---ATWWEFVEAL 66
A WL +E+ + ++EE A LTG A WW + ++ TW EF EA
Sbjct: 132 AEDWLREIEKKLDLTTCTDEECVGVAAHQLTGAARAWWDSYSDTHENPGGITWAEFTEAF 191
Query: 67 LRKFEPELEPYMPEPVQDSEEEEI----PGKQEVLE-AERRTVVVEAKPEANSVLLAKSE 121
E ++PE V D++ EE G Q+V E A R T + P+ ++
Sbjct: 192 -------REQHVPEGVMDAKVEEFRNISQGTQKVQEYATRFTRTMRYAPDESNT------ 238
Query: 122 IKPYSPENSEMEASRKDSEFAMAAEQTAEVSRCQPVTVVDYIDCTLTAKGEKIDAEGEIK 181
E +M +K ++ +S T+ + I+ L + ++++A+ + K
Sbjct: 239 ------EKKKMYFFKK----GLSTRLKVALSGHTCYTLREMINKALEMERDRLEADAQYK 288
Query: 182 IKRLQEGSS 190
K+ + SS
Sbjct: 289 EKKRRSESS 297
>ref|XP_623577.1| PREDICTED: similar to Titin-like protein [Apis mellifera]
Length = 9787
Score = 42.7 bits (99), Expect = 0.024
Identities = 42/173 (24%), Positives = 76/173 (43%), Gaps = 20/173 (11%)
Query: 73 ELEPYMPEPVQDSEE----------EEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEI 122
ELE Y+PE ++SE+ G+ + +R+ + EA E + + E
Sbjct: 7703 ELEKYVPEEHEESEKPPPEPIETSFRRSNGRNYTQKTKRKRKIEEAADEVTIKKIVEEET 7762
Query: 123 KPYSPENSEMEASRKDSEFAMAAEQTAEVSRCQPVTVVDYIDCT----LTAKGEKIDAEG 178
K E E R S+ E+ EV+ +PV + D T ++ + E++ E
Sbjct: 7763 K--EEEAPEEVVLRPKSKRKSIVEEEEEVTEKKPVVEEEVADVTVKKLISTEAEEVPEEF 7820
Query: 179 EIKIKRLQEGSSM--EKWQICSIHGRKKVQPIVLPQISDLGFDWERRPPRKPP 229
IK K+ E E+ + +I +K +P ++ + F +R+PP++PP
Sbjct: 7821 TIKKKKKPEKKPTVPEEVEETAITIKKVREPEEKEEVQE--FTVKRKPPKQPP 7871
>gb|AAV32158.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1131
Score = 42.4 bits (98), Expect = 0.031
Identities = 24/67 (35%), Positives = 32/67 (46%), Gaps = 3/67 (4%)
Query: 9 EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRR---NQKATWWEFVEA 65
EA WL VE+ + +E+EK S A L G A WW +R+ + TWWEF A
Sbjct: 73 EAGDWLHAVEKKLDLIQCTEQEKVSFASHQLHGPAAEWWDHFRQNRAAGEPITWWEFTAA 132
Query: 66 LLRKFEP 72
+ P
Sbjct: 133 FKKTHIP 139
>gb|AAP52585.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|37531992|ref|NP_920298.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|22857594|gb|AAN09868.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1802
Score = 42.0 bits (97), Expect = 0.040
Identities = 28/89 (31%), Positives = 39/89 (43%), Gaps = 8/89 (8%)
Query: 9 EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRR---RNQKATWWEFVEA 65
+A WL V + + S+EEK A L G A +WW ++ Q TW F A
Sbjct: 419 DALDWLHAVGKKLDTVQCSDEEKVIFAAHQLQGPASLWWDHFQATQPEGQPITWARFTAA 478
Query: 66 LLRKFEP-----ELEPYMPEPVQDSEEEE 89
R P L Y PE V++ EE++
Sbjct: 479 FRRTHVPAGVFNNLARYAPEDVREDEEKQ 507
>gb|AAF98211.1| Hypothetical protein [Arabidopsis thaliana]
gi|15219809|ref|NP_176872.1| hypothetical protein
[Arabidopsis thaliana] gi|25404659|pir||H96693
hypothetical protein F1O19.7 [imported] - Arabidopsis
thaliana
Length = 659
Score = 40.8 bits (94), Expect = 0.090
Identities = 30/125 (24%), Positives = 49/125 (39%), Gaps = 7/125 (5%)
Query: 3 PEFDGREAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKATWWEF 62
P FDG Y W VE+ + +K +L G A W+ + W F
Sbjct: 113 PVFDGSGVYEWFSKVERFFRVGRYQDSDKLDLVALSLEGVALKWFLREMSTLEFRDWNSF 172
Query: 63 VEALLRKFEPELEPYMPEPVQD-------SEEEEIPGKQEVLEAERRTVVVEAKPEANSV 115
+ LL +F+P PE + + E E +E++E + +V ++ PE +
Sbjct: 173 EQRLLARFDPVKIHSSPEVLMPTTLIHVVAHETESRCTEELIEEDESSVRKKSNPEIVLI 232
Query: 116 LLAKS 120
KS
Sbjct: 233 TQKKS 237
>gb|AAX95140.1| retrotransposon protein, putative, Ty3-gypsy sub-class [Oryza
sativa (japonica cultivar-group)]
Length = 1313
Score = 39.7 bits (91), Expect = 0.20
Identities = 35/133 (26%), Positives = 56/133 (41%), Gaps = 22/133 (16%)
Query: 9 EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWW---FCWRRRNQKATWWEFVEA 65
EA WL +E+ +++EK + A L G A WW R K TW EF +
Sbjct: 2 EANDWLRAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNYVVTRPDATKGTWVEFCQN 61
Query: 66 LLRKFEPELEPYMPEPVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEIKPY 125
+ P +PE + ++ E L+ RT V E E N ++L Y
Sbjct: 62 FRK-------PQIPEGIMAQQKREF----HALQQGTRT-VTEYLHEFNRLVL-------Y 102
Query: 126 SPENSEMEASRKD 138
+PE+ +A +++
Sbjct: 103 APEDVRTDAEKQE 115
>gb|AAO37503.1| retrotransposon protein, putative, Ty3-gypsy sub-class [Oryza
sativa (japonica cultivar-group)]
gi|50916491|ref|XP_468642.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1079
Score = 39.3 bits (90), Expect = 0.26
Identities = 35/133 (26%), Positives = 57/133 (42%), Gaps = 22/133 (16%)
Query: 9 EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWW---FCWRRRNQKATWWEFVEA 65
EA+ WL +E+ +E+EK + A L G A +WW R + TW EF +
Sbjct: 112 EAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYMVTRPAGAEVTWTEFRHS 171
Query: 66 LLRKFEPELEPYMPEPVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEIKPY 125
+ +PE + ++ E Q + V+E E N LLA+ Y
Sbjct: 172 FNK-------AQVPEGIVAQKKREFRSLQ-----QGTRTVIEYLHEFN--LLAR-----Y 212
Query: 126 SPENSEMEASRKD 138
+PE+ +A R++
Sbjct: 213 APEDVRTDAERQE 225
>ref|XP_475471.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|50080316|gb|AAT69650.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1458
Score = 39.3 bits (90), Expect = 0.26
Identities = 35/133 (26%), Positives = 57/133 (42%), Gaps = 22/133 (16%)
Query: 9 EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWW---FCWRRRNQKATWWEFVEA 65
EA+ WL +E+ +E+EK + A L G A +WW R + TW EF +
Sbjct: 112 EAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYMVTRPAGAEVTWTEFRHS 171
Query: 66 LLRKFEPELEPYMPEPVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEIKPY 125
+ +PE + ++ E Q + V+E E N LLA+ Y
Sbjct: 172 FNK-------AQVPEGIVAQKKREFRSLQ-----QGTRTVIEYLHEFN--LLAR-----Y 212
Query: 126 SPENSEMEASRKD 138
+PE+ +A R++
Sbjct: 213 APEDVRTDAERQE 225
>ref|NP_908336.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|52353682|gb|AAU44248.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|52353613|gb|AAU44179.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|20804440|dbj|BAB92137.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|15128451|dbj|BAB62635.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1524
Score = 39.3 bits (90), Expect = 0.26
Identities = 35/133 (26%), Positives = 57/133 (42%), Gaps = 22/133 (16%)
Query: 9 EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWW---FCWRRRNQKATWWEFVEA 65
EA+ WL +E+ +E+EK + A L G A +WW R + TW EF +
Sbjct: 112 EAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYMVTRPAGAEVTWTEFRHS 171
Query: 66 LLRKFEPELEPYMPEPVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEIKPY 125
+ +PE + ++ E Q + V+E E N LLA+ Y
Sbjct: 172 FNK-------AQVPEGIVAQKKREFRSLQ-----QGTRTVIEYLHEFN--LLAR-----Y 212
Query: 126 SPENSEMEASRKD 138
+PE+ +A R++
Sbjct: 213 APEDVRTDAERQE 225
>gb|AAQ56531.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1289
Score = 39.3 bits (90), Expect = 0.26
Identities = 23/67 (34%), Positives = 31/67 (45%), Gaps = 3/67 (4%)
Query: 9 EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKA---TWWEFVEA 65
EA WL +E+ + +++EK S A L G AF WW +R A TW EF A
Sbjct: 79 EAGDWLHAIEKKLDLLQCTDQEKVSFASHQLHGSAFEWWDHFRLNRTTAEPITWLEFTAA 138
Query: 66 LLRKFEP 72
+ P
Sbjct: 139 FRKTHIP 145
>gb|AAX96167.1| retrotransposon protein, putative, Ty3-gypsy sub-class [Oryza
sativa (japonica cultivar-group)]
gi|62733812|gb|AAX95921.1| retrotransposon protein,
putative, Ty3-gypsy sub-class [Oryza sativa (japonica
cultivar-group)]
Length = 344
Score = 38.9 bits (89), Expect = 0.34
Identities = 24/67 (35%), Positives = 32/67 (46%), Gaps = 3/67 (4%)
Query: 9 EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRR---RNQKATWWEFVEA 65
EA WL VE+ + +E+EK S A L G A WW +R+ Q TW +F EA
Sbjct: 82 EAGDWLHAVEKKLDLIQCTEQEKVSFASHQLHGPAAEWWDHFRQGRAEGQPITWAKFTEA 141
Query: 66 LLRKFEP 72
+ P
Sbjct: 142 FKKTHIP 148
>ref|XP_475728.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|50080333|gb|AAT69667.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1524
Score = 38.9 bits (89), Expect = 0.34
Identities = 33/133 (24%), Positives = 55/133 (40%), Gaps = 22/133 (16%)
Query: 9 EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWW---FCWRRRNQKATWWEFVEA 65
EA+ WL VE+ +E+EK + A L G A +WW R + TW EF +
Sbjct: 112 EAHDWLHAVEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYMVTRPAGAEVTWTEFCHS 171
Query: 66 LLRKFEPELEPYMPEPVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEIKPY 125
+ +PE + ++ E L+ RTV+ L + + Y
Sbjct: 172 FNK-------AQVPEGIVAQKKREF----RSLQQGTRTVI--------EYLHEFNRLARY 212
Query: 126 SPENSEMEASRKD 138
+PE+ +A R++
Sbjct: 213 APEDVRTDAERQE 225
>emb|CAB51072.1| hypothetical protein [Homo sapiens]
Length = 3261
Score = 38.5 bits (88), Expect = 0.44
Identities = 32/116 (27%), Positives = 53/116 (45%), Gaps = 15/116 (12%)
Query: 71 EPELEPYMPEPVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEIKPYSPENS 130
EP+ + +P Q SEE P +AE+ +A+P+AN A E +P + E+
Sbjct: 1349 EPDSTQPLSKPAQKSEEANEP------KAEKPDATADAEPDANQKAEAAPESQPPASEDL 1402
Query: 131 EME--ASRKDSEFAMAAEQTAEVSRCQPVTVVDYIDCTLTAKGEKIDAEGEIKIKR 184
E++ + KD + + V V ++ +T K E+ID E K+KR
Sbjct: 1403 EVDPPVAAKDKKPNKSKRSKTPVQ----AAAVSIVEKPVTRKSERIDRE---KLKR 1451
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.135 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 801,026,968
Number of Sequences: 2540612
Number of extensions: 33299667
Number of successful extensions: 88588
Number of sequences better than 10.0: 248
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 237
Number of HSP's that attempted gapping in prelim test: 88200
Number of HSP's gapped (non-prelim): 505
length of query: 458
length of database: 863,360,394
effective HSP length: 131
effective length of query: 327
effective length of database: 530,540,222
effective search space: 173486652594
effective search space used: 173486652594
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 77 (34.3 bits)
Lotus: description of TM0117b.14


