

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0114.10
(1082 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
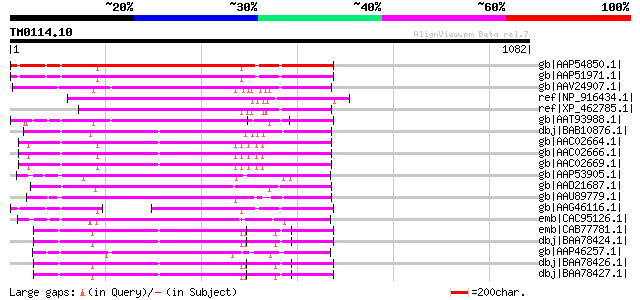
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP54850.1| putative gag-pol polyprotein [Oryza sativa (japon... 565 e-159
gb|AAP51971.1| putative copia-type polyprotein [Oryza sativa (ja... 563 e-158
gb|AAV24907.1| hypothetical protein [Oryza sativa (japonica cult... 513 e-143
ref|NP_916434.1| putative gag/pol polyprotein [Oryza sativa (jap... 490 e-137
ref|XP_462785.1| putative gag/pol polyprotein [Oryza sativa (jap... 459 e-127
gb|AAT93988.1| putative polyprotein [Oryza sativa (japonica cult... 422 e-116
dbj|BAB10876.1| polyprotein [Arabidopsis thaliana] 405 e-111
gb|AAC02664.1| polyprotein [Arabidopsis thaliana] 400 e-109
gb|AAC02666.1| polyprotein [Arabidopsis thaliana] 399 e-109
gb|AAC02669.1| polyprotein [Arabidopsis thaliana] 399 e-109
gb|AAP53905.1| putative pol polyprotein [Oryza sativa (japonica ... 374 e-102
gb|AAD21687.1| Strong similarity to gi|3600044 T12H20.12 proteas... 368 e-100
gb|AAU89779.1| gag-pol polyprotein-like [Solanum tuberosum] 347 2e-93
gb|AAG46116.1| putative copia-like retrotransposon polyprotein [... 342 4e-92
emb|CAC95126.1| gag-pol polyprotein [Populus deltoides] 338 4e-91
emb|CAB77781.1| putative polyprotein of LTR transposon [Arabidop... 317 2e-84
dbj|BAA78424.1| polyprotein [Arabidopsis thaliana] 317 2e-84
gb|AAP46257.1| putative polyprotein [Oryza sativa (japonica cult... 315 5e-84
dbj|BAA78426.1| polyprotein [Arabidopsis thaliana] 315 7e-84
dbj|BAA78427.1| polyprotein [Arabidopsis thaliana] 314 1e-83
>gb|AAP54850.1| putative gag-pol polyprotein [Oryza sativa (japonica
cultivar-group)] gi|37536522|ref|NP_922563.1| putative
gag-pol polyprotein [Oryza sativa (japonica
cultivar-group)] gi|13310887|gb|AAG13591.2| putative
gag/pol polyprotein [Oryza sativa]
Length = 1417
Score = 565 bits (1455), Expect = e-159
Identities = 307/697 (44%), Positives = 419/697 (60%), Gaps = 42/697 (6%)
Query: 1 TSQWTRPSAPPRQPGILGPRPQAHAASTSPVPTDIATAMHTMTLAPPDSTWYMDTGASSH 60
T+Q P+ PP PG+ A S + + + +A T P + W++DTGAS+H
Sbjct: 322 TAQHLLPALPPASPGV----HSTGAWDNSALYSALQSAGVATTTPPSAADWFLDTGASAH 377
Query: 61 VASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSHVLHTPQIV 120
++S+PG L L + + VG+G +P+ + T +PTS L L +VL +P ++
Sbjct: 378 MSSTPGILAHPRPLP-FSSCITVGNGAKLPVTHTASTHIPTSSTD--LHLHNVLVSPPLI 434
Query: 121 KNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLYPVTPSVPFAGLAQSL 180
KNLISV++LT DNNVS+ FDP GFS+ D QT + +RC+S GDLYP+ P A A S
Sbjct: 435 KNLISVKKLTRDNNVSIEFDPTGFSIKDLQTQVVKLRCDSPGDLYPLRLPSPHALSATSS 494
Query: 181 -----WHSRLGHPGSSALQSLRSNKFITYEHLNSPTVCESCVFGKHVRLPFVSSNNVTVM 235
WH RLGHPGS++L + + F + ++P C +C G +VRLPF SS++ T+
Sbjct: 495 PSVEHWHLRLGHPGSASLSKVLGS-FDFQCNKSAPHHCSACHVGTNVRLPFHSSSSQTLF 553
Query: 236 PFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKSQVFEMFTSLTHQIRTHF 295
PF ++H+D+WTSP+ S++G++YYV+FLDDFT ++WTFP+ NKS+VF S T F
Sbjct: 554 PFQLVHTDVWTSPIYSNSGYKYYVVFLDDFTHYIWTFPVRNKSEVFHTVRSFFAYAHTQF 613
Query: 296 SHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLL 355
V LQ DNG E+D+ + + + HG + R SCP++S QNGKAER +RTIN+ +RT+L
Sbjct: 614 GLPVLALQTDNGKEYDSYALRSLLSLHGAVLRLSCPYSSQQNGKAERILRTINDYVRTML 673
Query: 356 AHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVP 415
H++ P SFW ALQ AT+L+N P ++ +L+P QLL P+Y HLRVFGCLCYP
Sbjct: 674 VHSAAPLSFWAEALQTATHLINRRPCRATGSLTPYQLLLGAPPTYDHLRVFGCLCYPNTI 733
Query: 416 STTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVIFDETRFPFATMPTPPPS 475
+T +KL PRS CVF+GYP HRGY+C+D+ R++ SRHV F E FPF P+P PS
Sbjct: 734 ATAPHKLSPRSLACVFIGYPADHRGYRCYDMVSRRVFTSRHVTFVEDVFPFRDAPSPRPS 793
Query: 476 T---YDCFSD--------------PIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTS 518
D D P+ + H SP PSP S P + P+
Sbjct: 794 APPPPDHGDDTIVLLPAPAQHVVTPVGTAPAHDAASP-PSPASSTPSSAAPAH------- 845
Query: 519 SSASPPSSPSPPSPPQPVPPIRTMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLA 578
+PP SP SP PP M TR+ GI KP + ++ T T+SP P + + A
Sbjct: 846 -DVAPPPSPETSSPASASPPRHAMTTRARAGISKPNPRYAMTAT---STLSPTPSSVRAA 901
Query: 579 LSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVG 638
L DPNW++AMQ+EFDAL+ N+TW LVPRP II W+F K A+G ++YKAR V
Sbjct: 902 LRDPNWRAAMQAEFDALLANRTWTLVPRPPGARIITGKWVFKTKLHADGSLDKYKARWVV 961
Query: 639 DGRSQIAGVDCDETFSPVVKPATIRTVLTIALSGLGP 675
G +Q GVD ETFSPVVKPATIRTVLT+ S P
Sbjct: 962 RGFNQRPGVDFGETFSPVVKPATIRTVLTLISSKQWP 998
>gb|AAP51971.1| putative copia-type polyprotein [Oryza sativa (japonica
cultivar-group)] gi|37530764|ref|NP_919684.1| putative
copia-type polyprotein [Oryza sativa (japonica
cultivar-group)] gi|20042923|gb|AAM08751.1| Putative
copia-type polyprotein [Oryza sativa (japonica
cultivar-group)]
Length = 1803
Score = 563 bits (1450), Expect = e-158
Identities = 306/697 (43%), Positives = 418/697 (59%), Gaps = 42/697 (6%)
Query: 1 TSQWTRPSAPPRQPGILGPRPQAHAASTSPVPTDIATAMHTMTLAPPDSTWYMDTGASSH 60
T+Q P+ PP PG+ A S + + + +A T P + W++DTGAS+H
Sbjct: 322 TAQHLLPALPPASPGV----HSTGAWDNSALYSALQSAGVATTTPPSAADWFLDTGASAH 377
Query: 61 VASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSHVLHTPQIV 120
++S+PG L L + + VG+G +P+ + T +PTS L L +VL +P ++
Sbjct: 378 MSSTPGILAHPRPLP-FSSCITVGNGAKLPVTHTASTHIPTSSTD--LHLHNVLVSPPLI 434
Query: 121 KNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLYPVTPSVPFAGLAQSL 180
KNLISV++LT DNNVS+ FDP GFS+ D QT + +RC+S GDLYP+ P A A S
Sbjct: 435 KNLISVKKLTRDNNVSIEFDPTGFSIKDLQTQVVKLRCDSPGDLYPLRLPSPHALSATSS 494
Query: 181 -----WHSRLGHPGSSALQSLRSNKFITYEHLNSPTVCESCVFGKHVRLPFVSSNNVTVM 235
WH RLGHPGS++L + + F + ++P C +C G +VRLPF SS++ T+
Sbjct: 495 PSVEHWHLRLGHPGSASLSKVLGS-FDFQCNKSAPHHCSACHVGTNVRLPFHSSSSQTLF 553
Query: 236 PFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKSQVFEMFTSLTHQIRTHF 295
PF ++H+D+WTSP+ S++G++YYV+FLDDFT ++WTFP+ NKS+VF S T F
Sbjct: 554 PFQLVHTDVWTSPIYSNSGYKYYVVFLDDFTHYIWTFPVRNKSEVFHTVRSFFAYAHTQF 613
Query: 296 SHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLL 355
V LQ DNG E+D+ + + + HG + R SCP++S QNGKAER +RTIN+ +RT+L
Sbjct: 614 GLPVLALQTDNGKEYDSYALRSLLSLHGAVLRLSCPYSSQQNGKAERILRTINDCVRTML 673
Query: 356 AHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVP 415
H++ P SFW ALQ A +L+N P ++ +L P QLL P+Y HLRVFGCLCYP
Sbjct: 674 VHSAAPLSFWAEALQTAMHLINRRPCRATGSLKPYQLLLGAPPTYDHLRVFGCLCYPNTI 733
Query: 416 STTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVIFDETRFPFATMPTPPPS 475
+T +KL PRS CVF+GYP HRGY+C+D+ R++ SRHV F E FPF P+P PS
Sbjct: 734 ATAPHKLSPRSLACVFIGYPADHRGYRCYDMVSRRVFTSRHVTFVEDVFPFRDAPSPRPS 793
Query: 476 T---YDCFSD--------------PIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTS 518
D D P+ + H SP PSP S P + P+
Sbjct: 794 APPPPDHGDDTIVLLPAPAQHVVTPVGTAPAHDAASP-PSPASSTPSSAAPAH------- 845
Query: 519 SSASPPSSPSPPSPPQPVPPIRTMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLA 578
+PP SP SP PP M TR+ GI KP + ++ T T+SP P + ++A
Sbjct: 846 -DVAPPPSPETSSPASASPPRHAMTTRARAGISKPNPRYAMTAT---STLSPTPSSVRVA 901
Query: 579 LSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVG 638
L DPNW++AMQ+EFDAL+ N+TW LVPRP II W+F K A+G ++YKAR V
Sbjct: 902 LRDPNWRAAMQAEFDALLANRTWTLVPRPPGARIITGKWVFKTKLHADGSLDKYKARWVV 961
Query: 639 DGRSQIAGVDCDETFSPVVKPATIRTVLTIALSGLGP 675
G +Q GVD ETFSPVVKPATIRTVLT+ S P
Sbjct: 962 RGFNQRPGVDFGETFSPVVKPATIRTVLTLISSKQWP 998
>gb|AAV24907.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|51854440|gb|AAU10819.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1679
Score = 513 bits (1320), Expect = e-143
Identities = 307/768 (39%), Positives = 419/768 (53%), Gaps = 111/768 (14%)
Query: 7 PSAPPRQPGILGPRPQAHAASTSPVPTDIATAMHTMTL-APPDSTWYMDTGASSHVASSP 65
PS PP + P A + +A + +T TL P + WYMDTGA++H+ S
Sbjct: 515 PSQPPTSVALQQPLGWFGGAPSQWDQASLAGSFNTTTLHQPATNDWYMDTGATAHMTSDT 574
Query: 66 GNLT-SYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSHVLHTPQIVKNLI 124
G L+ S+ + + VG+G IP+ G + + H + TL +L +P I+KNLI
Sbjct: 575 GILSLSHPPNPNSPSHIVVGNGSTIPVTSIGHSKI--CHPNCSFTLRDILCSPAIIKNLI 632
Query: 125 SVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLYPVTPSVP-----FAGLAQS 179
SVR+ DN SV FDP+GFSV D +T + R NS G LY + ++P A LA S
Sbjct: 633 SVRRFVIDNWCSVEFDPFGFSVKDLRTRTVIARFNSSGPLYSLHHALPPPPAATALLANS 692
Query: 180 ----LWHSRLGHPGSSALQSLRSNKFITYEHLNSPTVCESCVFGKHVRLPFVSSNNVTVM 235
LWH RLGH G AL L + +T L VC +C G+HVRLPF SS +
Sbjct: 693 STDLLWHRRLGHLGHDALNRLAAVVPMTRGDLTG--VCHACQLGRHVRLPFASSTSRAST 750
Query: 236 PFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKSQVFEMFTSLTHQIRTHF 295
F++ H DLWTSPV+S++G +YY++ LDD + ++WTFPL KS F + ++T F
Sbjct: 751 NFELFHCDLWTSPVVSASGFKYYLVILDDCSHYVWTFPLRFKSDTFTTLSHFFAYVKTQF 810
Query: 296 SHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLL 355
+ N++ +QCDNG EFDN++ + +G+ R SCPHTS QNGKAER +R++NN++R++L
Sbjct: 811 ATNIRSIQCDNGREFDNSAARTFFLTNGVHLRMSCPHTSPQNGKAERILRSLNNIVRSML 870
Query: 356 AHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVP 415
A +P SFW AL AT+L+N P K+L +P LY PSY+HLRVFGC CYP +
Sbjct: 871 FQAKLPGSFWVEALHTATHLINRHPTKTLDRHTPHFALYGTHPSYSHLRVFGCKCYPNLS 930
Query: 416 STTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVIFDETRFPFATM------ 469
+TT +KL PRS+ CVFLGYPL+H+GY+CFD ++IISRHV+FDE FPF +
Sbjct: 931 ATTPHKLAPRSTMCVFLGYPLYHKGYRCFDPLSNRVIISRHVVFDEHSFPFTELTNGVSN 990
Query: 470 -----------------------PTPPPSTYDCFSDPIHP-------------SIIHQWT 493
P P+T S +H S +
Sbjct: 991 ATDLDFLEDFTAPAQAPIGATRRPAVAPTTQTASSPMVHGLERPPPCSPTRPVSTPGGPS 1050
Query: 494 SPD-----PSPTSVV---------PPAVTPSQLPTTSTSSS-----ASPPSSPSPPS--- 531
SPD PSPT + PP+ P+ +T +SS A PP+ P P
Sbjct: 1051 SPDSRLGPPSPTPALIGPASTSPGPPSAGPAPSASTCPASSTWETVARPPTPPGLPRLDG 1110
Query: 532 ---PPQPVPPIR-------------------------TMATRSMRGIYKPRKLFNLSVTI 563
PP P P R M TR+ G +KP NL
Sbjct: 1111 PHLPPAPHVPRRLRSVRATGAPTPLSGLEISPVVNDHVMTTRAKSGHHKPVHRLNLHAA- 1169
Query: 564 DDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKT 623
+S +PK + AL+DP W++AM+ E++AL+ N+TWDLVPRP+ VN++ WIF HK
Sbjct: 1170 ---PLSLVPKTYRAALADPLWRAAMEEEYNALLANRTWDLVPRPAGVNVVTGKWIFKHKF 1226
Query: 624 KANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLTIALS 671
A+G +RYKAR V G +Q GVD DETFSPVVKPAT+RTVL++A+S
Sbjct: 1227 HADGSLDRYKARWVLRGFTQRPGVDFDETFSPVVKPATVRTVLSLAVS 1274
>ref|NP_916434.1| putative gag/pol polyprotein [Oryza sativa (japonica
cultivar-group)]
Length = 1090
Score = 490 bits (1262), Expect = e-137
Identities = 279/653 (42%), Positives = 369/653 (55%), Gaps = 68/653 (10%)
Query: 120 VKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLYPV---TPSVPFAGL 176
V+NL+SVRQ T DN S+ FD +GFSV D QT ++RCNS G+LY + TPS GL
Sbjct: 58 VRNLLSVRQFTRDNKCSIEFDEFGFSVKDLQTRRVILRCNSRGELYTLPAATPSSAAHGL 117
Query: 177 ---AQSLWHSRLGHPGSSALQSLRSNKFITYEHLNSPTVCESCVFGKHVRLPFVSSNNVT 233
+ +LWH RLGHPG +A+ LR+ I+ +++ ++C +C GKH RLPF +S++ T
Sbjct: 118 LATSSTLWHCRLGHPGPAAIHGLRNIASISCNKIDT-SLCHACQLGKHTRLPFHNSSSRT 176
Query: 234 VMPFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKSQVFEMFTSLTHQIRT 293
+PF+++H D+WTSPV+S++G +YY++ LDDF+ F WTF L KS V + T
Sbjct: 177 SVPFELVHCDVWTSPVMSTSGFKYYLVVLDDFSHFCWTFLLRLKSDVHRHIVEFVEYVST 236
Query: 294 HFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRT 353
F +K Q DNG EF NT+ + A+ G R SCP+TS QNGKAER +RTINN IRT
Sbjct: 237 QFGLPLKSFQADNGREFVNTAITTFLASRGTQLRLSCPYTSPQNGKAERMLRTINNSIRT 296
Query: 354 LLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPL 413
LL AS+PPS+W AL ATYLLN P S+ P QLL+R P ++HLRVFGCLCYP
Sbjct: 297 LLIQASMPPSYWAEALATATYLLNRRPSSSIHQSLPFQLLHRTIPDFSHLRVFGCLCYPN 356
Query: 414 VPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVIFDETRFPFATMPTPP 473
+ +TT +KL PRS+ CVFLGYP H+GY+C DLS +IIISRHV+FDE++FPFA P P
Sbjct: 357 LSATTPHKLSPRSTACVFLGYPTSHKGYRCLDLSTHRIIISRHVVFDESQFPFAATP-PA 415
Query: 474 PSTYDCFSDPIHPSIIHQWTSPDPSPTSVVP-------------------PAVTPSQLPT 514
S++D + P+ P P +V P S+ P+
Sbjct: 416 ASSFDFLLQGLSPADAPSLEVEQPRPLTVAPSTEVEQPYLPLPSRRLSAGTVTVASEAPS 475
Query: 515 T------STSSSASPPSSPS---------------------PPSPPQPV------PPIRT 541
++S+ A+PP S + PPS V P +
Sbjct: 476 AGAPLVGTSSADATPPGSATRASTIVSPFRHVYTRRPVTTVPPSSSTAVTNAVAAPQPHS 535
Query: 542 MATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTW 601
M TRS G +P + L+ T SP+P N AL+DPNW++AM E+ L+ N TW
Sbjct: 536 MVTRSQSGSLRP--VDRLTYTATQAAASPVPANYHSALADPNWRAAMADEYKELVDNGTW 593
Query: 602 DLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPAT 661
LV RP NI WIF HK ++G RYKAR V G SQ G+D DETFSPVVK AT
Sbjct: 594 RLVSRPPRANIATGKWIFKHKFHSDGSLARYKARWVVRGYSQQHGIDYDETFSPVVKLAT 653
Query: 662 IRTVLTIALSGLGPF------TS*MFRMLSSMVICMRQSTCISPWVSVTILIL 708
IR VL+IA S P + + L V C + S + P + +L
Sbjct: 654 IRVVLSIAASRAWPIHQLDVKNAFLHGHLKETVYCQQPSGFVDPTAPDAVCLL 706
>ref|XP_462785.1| putative gag/pol polyprotein [Oryza sativa (japonica
cultivar-group)]
Length = 1373
Score = 459 bits (1180), Expect = e-127
Identities = 256/620 (41%), Positives = 346/620 (55%), Gaps = 95/620 (15%)
Query: 143 GFSVHDFQTGIPLMRCNSLGDLYPVTPSVP----FAGLAQSLWHSRLGHPGSSALQSLRS 198
G + D +T + RCNS GDLYP P SLWH RLGH G AL L
Sbjct: 350 GGAALDLETRNVIARCNSSGDLYPFYPPATSTHALLAAPTSLWHRRLGHLGREALSKLIR 409
Query: 199 NKFITYEHLNSPTVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRYY 258
+ I+ + P +C +C G H RLPF SS++ FD++H DLWTSP++S +G++YY
Sbjct: 410 SSVISCTKDDLPHLCHACQLGHHTRLPFSSSSSRASNNFDLIHCDLWTSPIVSVSGYKYY 469
Query: 259 VLFLDDFTDFLWTFPLSNKSQVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAY 318
++ LDD + ++WTFPL KS F + ++T F +K +QCDNG EFDN+ +
Sbjct: 470 LVILDDCSHYIWTFPLRLKSDTFSTIANFFAHVKTQFGTTIKSVQCDNGREFDNSPARTF 529
Query: 319 CAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNI 378
+HG+ FR SCP+TS QNG+AER +RT+NN++R+LL A +PP +W AL AT L+N
Sbjct: 530 FLSHGVAFRMSCPYTSQQNGRAERSLRTLNNILRSLLFQACLPPVYWVEALHTATLLVNR 589
Query: 379 IPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHH 438
IP K+LS+ +P LY P+Y HLRVFGC CYP + ST +KL PRSS CVFLGY H
Sbjct: 590 IPTKTLSSSTPYFHLYSTQPTYDHLRVFGCACYPNMSSTAPHKLAPRSSLCVFLGYSSEH 649
Query: 439 RGYKCFDLSHRKIIISRHVIFDETRFPFATMPTPP--PSTYDCFSD-------PIHPSII 489
+GY+C +L +II SRHV+FDE+ FPFA M T P S D F D P +
Sbjct: 650 KGYRCLELGSNRIITSRHVVFDESFFPFADMSTSPMASSALDIFLDDNELTAQPPRAKFV 709
Query: 490 HQWTS-----------PDPSPTSV--------------------------VPPAVTPSQ- 511
H TS P P+P+S+ P A++P++
Sbjct: 710 HAGTSSAARGAVEPSTPPPAPSSIGPRSPATLAGPEAGPHGGSPAGAATSQPGAISPART 769
Query: 512 -LPTTSTSSSASPPSSP------SPPS----------PPQ-------------------- 534
P+ +TS++ + S+P + PS PP+
Sbjct: 770 AAPSAATSTTRAVTSAPRAATSGTTPSLSPLAGTAAPPPRAEVAASSTAATGRTLATRPV 829
Query: 535 ---PVPPIRTMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSE 591
PV +M TR G+ +P NL +SP+P++ + ALSDPNW++AMQ+E
Sbjct: 830 SIAPVDNAHSMRTRGKAGMAQPVDRLNLHAA----PLSPVPRSVREALSDPNWRAAMQAE 885
Query: 592 FDALIRNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDE 651
FDAL+ N TW LVPRP VN++ WIF HK ++G +RYKAR V G +Q GVD DE
Sbjct: 886 FDALLANDTWSLVPRPRGVNLVTGKWIFRHKLHSDGSLDRYKARWVLRGFTQRPGVDYDE 945
Query: 652 TFSPVVKPATIRTVLTIALS 671
TFSPVVKPAT+R VL++ALS
Sbjct: 946 TFSPVVKPATVRVVLSLALS 965
>gb|AAT93988.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1480
Score = 422 bits (1085), Expect = e-116
Identities = 253/623 (40%), Positives = 348/623 (55%), Gaps = 57/623 (9%)
Query: 2 SQWTRPSAPPRQPGILGPRPQAHAASTS-----------PVP--------TDIATAMHTM 42
S W PS P G+LGP PQAH A S P+P + A++ +
Sbjct: 291 STWRPPSNPSNSAGLLGPYPQAHTAFASSYVSPPTPGWPPMPQVQPNWDQAGLVAALNQL 350
Query: 43 TLAPPDSTWYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTS 102
++ P S W +DTGA+SH++S+ G L + SH + VG+G IP+ G + LP
Sbjct: 351 SVNSP-SPWVLDTGATSHMSSTDGILVTRLPNSHTF--ITVGNGHTIPVICRGTSFLPIG 407
Query: 103 HKHKPLTLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLG 162
L ++L P +V+NL+ VRQ T DN S FD +GFSV D T ++RCNS G
Sbjct: 408 TTR--FALKNILVAPSLVRNLLFVRQFTRDNKCSFEFDEFGFSVKDLPTRRVILRCNSRG 465
Query: 163 DLYPVTPSVP------FAGLAQSLWHSRLGHPGSSALQSLRSNKFITYEHLNSPTVCESC 216
DLY + +VP F + +LWH RLGHP +A+Q+L ++ N+ +C +C
Sbjct: 466 DLYTLPTTVPAITAHSFLAKSSTLWHHRLGHPSPAAVQTLHKLAILSCTRSNNK-LCHAC 524
Query: 217 VFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSN 276
GKH RL F S++ T PF+++H D+WTSPVLS +G +YY++ LDDFT F WTFPL +
Sbjct: 525 HLGKHTRLSFSKSSSSTSSPFELVHCDVWTSPVLSLSGFKYYLVVLDDFTHFCWTFPLRH 584
Query: 277 KSQVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQ 336
KS V + ++T FS ++C Q DNG +F N + ++ A+ G++ R SCP+TS Q
Sbjct: 585 KSDVHQHLLEFVAYVKTQFSLPIRCFQADNGTKFVNHATTSFFASRGIVLRLSCPYTSPQ 644
Query: 337 NGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRR 396
NGKAER +RTIN IRTLL AS+PPS+W AL ATYLLN P S+ N P QLL+ +
Sbjct: 645 NGKAERVLRTINKSIRTLLIQASMPPSYWAEALATATYLLNRRPSTSVRNSIPYQLLHNK 704
Query: 397 DPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRH 456
P Y++L+VFGCLCYP + + T +KL PRS+P VFLGY H+G++C D+S R++ ISRH
Sbjct: 705 LPDYSNLQVFGCLCYPNLSAMTSHKLSPRSAPYVFLGYSASHKGFRCLDISTRRLYISRH 764
Query: 457 VIFDETRFPFATMPTPPPSTYDC----FSDPIHPSI---IHQWTSPDPSP--TSVVPPAV 507
V+FDE FPFA +P S++D FS + PS +++S PSP +P
Sbjct: 765 VVFDEKTFPFAAIP-QDASSFDFLLQGFSIAVAPSSEVERPRFSSMTPSPEVEQPIPDDD 823
Query: 508 TPS----QLPTTSTSSSASPPSSPSPPSPPQPVPPIRTMATRSMRGIYKPRKLFNLSVTI 563
T QL SS+A P + SP P A S NLS +
Sbjct: 824 TSGTELFQLLPGLRSSAAGRPLAGSPVDARLPGGCANDAANGSSSS--------NLSPVM 875
Query: 564 DDPTIS---PLP-KNPKLALSDP 582
D P S P P + P +L P
Sbjct: 876 DPPAASVVRPAPSEGPTTSLISP 898
Score = 142 bits (359), Expect = 5e-32
Identities = 87/198 (43%), Positives = 111/198 (55%), Gaps = 22/198 (11%)
Query: 496 DPSPTSVVPPAVTPSQLPTTSTSSSASPP----SSPSPPSPPQPVPPIR----------- 540
DP SVV PA PS+ PTTS S S P+P + +P+ R
Sbjct: 876 DPPAASVVRPA--PSEGPTTSLISPYRHTYLRRSQPAPTAIHRPIRASRAFHSATDQQQQ 933
Query: 541 ---TMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIR 597
TM TRS G +P + F + T D +SP+P N + AL+DPNW++A +E+ AL+
Sbjct: 934 TGHTMVTRSQTGHLRPIQRFTYTATHD--VVSPVPSNYRSALADPNWRAATANEYKALVD 991
Query: 598 NKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVV 657
N TW LVPRP N++ WIF HK ++G R+KAR V G SQ G+D DETFSPVV
Sbjct: 992 NNTWRLVPRPPGANVVTGKWIFKHKFHSDGTLARHKARWVVRGYSQQHGIDYDETFSPVV 1051
Query: 658 KPATIRTVLTIALSGLGP 675
KPATI VL+IA S P
Sbjct: 1052 KPATIHVVLSIAASRSWP 1069
>dbj|BAB10876.1| polyprotein [Arabidopsis thaliana]
Length = 1429
Score = 405 bits (1040), Expect = e-111
Identities = 254/736 (34%), Positives = 356/736 (47%), Gaps = 102/736 (13%)
Query: 29 SPVPTDIATAMHTMTLAPPDSTWYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQG 88
SP P + APP + W +D+GA+ H+ S NL+ + + +++ + G G
Sbjct: 292 SPSPMAPWQPRANIATAPPFNPWVLDSGATHHLTSDLANLSMHQPYTG-GEEVTIADGSG 350
Query: 89 IPIQGSGDTTLPTSHKHKPLTLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHD 148
+PI +G LPT + L L +L+ P + KNLISV +L N VSV F P F V D
Sbjct: 351 LPISHTGSALLPTPSRS--LALKDILYVPNVSKNLISVYRLCNANQVSVEFFPAHFQVKD 408
Query: 149 FQTGIPLMRCNSLGDLYP---------VTPSVPFAGLAQSLWHSRLGHPGSSALQSLRSN 199
TG L++ + +LY + + P S WH RLGHP L+ + S+
Sbjct: 409 LNTGARLLQGRTRNELYEWPVNQKSITILTASPSPKTDLSSWHQRLGHPALPILKDVVSH 468
Query: 200 KFITYEH-LNSPTVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRYY 258
+ + + C C K +LPF ++ V+ P + L++D+WTSP +S ++YY
Sbjct: 469 FHLPLSNTIPKQLPCSDCSINKSHKLPFFTNTIVSSQPLEYLYTDVWTSPCISVDNYKYY 528
Query: 259 VLFLDDFTDFLWTFPLSNKSQVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAY 318
++ +D FT + W +PL KSQV ++F + + F ++ L DNG EF +
Sbjct: 529 LVIVDHFTRYTWMYPLKQKSQVKDVFVAFKALVENRFQSRIRTLYSDNGGEF--IGLRPF 586
Query: 319 CAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNI 378
AAHG+ S PHT NG AERK R I LL HAS+P +FW +A A YL+N
Sbjct: 587 LAAHGISHLTSPPHTPEHNGLAERKHRHIVETGLALLTHASLPKTFWTYAFATAVYLINR 646
Query: 379 IPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHH 438
+P + L SP L++ P+Y LRVFGCLCYP + NKL+ RS+ CVFLGY L
Sbjct: 647 MPTEVLQGTSPYVKLFQMSPNYLKLRVFGCLCYPWLRPYNTNKLEARSTMCVFLGYSLTQ 706
Query: 439 RGYKCFDLSHRKIIISRHVIFDETRFPFATMPTPPPSTYDCFSDPIHPSII--------- 489
Y C D++ +I SRHV F E+ FPFA+ T + S P ++I
Sbjct: 707 SAYLCLDIATNRIYTSRHVQFVESSFPFASPRTSETDSTQTMSQPTTTNVIPLLQRPPHI 766
Query: 490 ----------------HQWTSPDPSPTSVVP----------------------------- 504
H +SP P+ VP
Sbjct: 767 APPTALPLCPIFHSPPHSPSSPASPPSEHVPLTAASSSSNAINDDNISSTGQVSVSGPTS 826
Query: 505 --PAVTPSQLPT-----------TSTSSSASPPSSPS-------------PPSPPQPVPP 538
P TP+ T T+ S +++PP+SP+ P +PP P PP
Sbjct: 827 QSPHTTPTNQNTSPLSKSPNPTNTNQSQNSTPPTSPTTSVHQHSPTPSPLPQNPPLPPPP 886
Query: 539 --IRTMATRSMRGIYKPRKLFNL--SVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDA 594
M TR+ I KP+ FNL S+T PTI P AL DPNW++AM E +A
Sbjct: 887 QNDHPMRTRAKNQITKPKTKFNLTTSLTSSKPTI---PTTVAQALKDPNWRNAMSEEINA 943
Query: 595 LIRNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFS 654
++N TWDLV ++I C WIF K +G RYKARLV G +Q G+D ETFS
Sbjct: 944 QMKNHTWDLVSPEEAKHVISCKWIFTLKYNVDGSIARYKARLVARGFNQQYGIDYSETFS 1003
Query: 655 PVVKPATIRTVLTIAL 670
PV+K TIRTVL +A+
Sbjct: 1004 PVIKSTTIRTVLEVAV 1019
>gb|AAC02664.1| polyprotein [Arabidopsis thaliana]
Length = 1451
Score = 400 bits (1027), Expect = e-109
Identities = 264/753 (35%), Positives = 363/753 (48%), Gaps = 105/753 (13%)
Query: 19 PRPQAHAASTSPVPTDIAT-----AMHTMTLAPP--DSTWYMDTGASSHVASSPGNLTSY 71
P+ Q HA S + + A+ M A P W +D+GA+ H+ S+ NL +
Sbjct: 294 PQLQQHAGSYASNQSSSASYAPWQPRANMVSATPYNSGNWLLDSGATHHLTSNLNNLALH 353
Query: 72 SNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSHVLHTPQIVKNLISVRQLTT 131
+ ++++ + G G+PI SG LPT + L L VL+ P I KNLISV ++
Sbjct: 354 QPYNG-DEEVTIADGSGLPISHSGSALLPTPTRS--LALKDVLYVPDIQKNLISVYRMCN 410
Query: 132 DNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY--PVTPSVPFAGLAQSL-------WH 182
N VSV F P F V D TG L++ + +LY PV S+ + A WH
Sbjct: 411 TNGVSVEFFPAHFQVKDLSTGARLLQGKTKNELYEWPVNSSIATSMFASPTPKTDLPSWH 470
Query: 183 SRLGHPGSSALQSLRSNKFITYEH-LNSPTVCESCVFGKHVRLPFVSSNNVTVMPFDILH 241
+RLGHP L++L S + H L + +C C K +LPF S+ + P + L+
Sbjct: 471 ARLGHPSLPILKALISKFSLPISHSLQNQLLCSDCSINKSHKLPFYSNTIASSHPLEYLY 530
Query: 242 SDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKSQVFEMFTSLTHQIRTHFSHNVKC 301
+D+WTSP+ S ++YY++ +D +T + W +PL KSQV EMF + T + F +
Sbjct: 531 TDVWTSPITSIDNYKYYLVIVDHYTRYTWLYPLRKKSQVREMFITFTALVENKFKFKIGT 590
Query: 302 LQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVP 361
L DNG EF + ++ A+HG+ + PHT NG +ERK R I TLL+ AS+P
Sbjct: 591 LYSDNGGEF--IAMRSFLASHGISHMTTPPHTPELNGISERKHRHIVETGLTLLSTASMP 648
Query: 362 PSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSTTINK 421
+W +A A YL+N + L N SP L+ + P+Y LR+FGCLC+P + T +K
Sbjct: 649 KEYWSYAFATAVYLINRMLTPVLGNESPYVKLFGQPPNYLKLRIFGCLCFPWLRPYTAHK 708
Query: 422 LQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVIFDETRFPFAT------------- 468
L RS PCV LGY L Y C D + ++ SRHV F E+ FPF+T
Sbjct: 709 LDNRSVPCVLLGYSLSQSAYLCLDRATGRVYTSRHVQFAESSFPFSTTSPSVTPPSDPPL 768
Query: 469 ---------------MPTPPPSTYDCFSD--------------PIHPSIIHQWTSP---- 495
+ T PPS+ C + P+ PS+ TSP
Sbjct: 769 SQDTRPVSVPLLARPLTTAPPSSPSCSAPHRSPSQSENLSPPAPLQPSLSLSPTSPITSP 828
Query: 496 ------------DPSPTSVVPPAVTPSQLP------TTSTSSSASPP------------- 524
+ PT PP Q P +TS SS+ P
Sbjct: 829 SLSEESLVGHNSETGPTGSSPPLSPQPQRPQPQSPQSTSPHSSSPQPNSPNPQHSPRSLT 888
Query: 525 ----SSPSPPSPPQPVPP--IRTMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLA 578
SSPSP PP P PP TM TRS I KP F T P +PK A
Sbjct: 889 PTLTSSPSPSPPPNPNPPPIQHTMRTRSKNNIVKPNPKFANLATKPTPLKPIIPKTVVEA 948
Query: 579 LSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVG 638
L DPNW+ AM E +A RN T+DLVP + N++ C W+F K +NG +RYKARLV
Sbjct: 949 LLDPNWRQAMCDEINAQTRNGTFDLVPPAPNQNVVGCKWVFTLKYLSNGVLDRYKARLVA 1008
Query: 639 DGRSQIAGVDCDETFSPVVKPATIRTVLTIALS 671
G Q G D ETFSPV+K T+R+VL IA+S
Sbjct: 1009 KGFHQQYGHDFKETFSPVIKSTTVRSVLHIAVS 1041
>gb|AAC02666.1| polyprotein [Arabidopsis thaliana]
Length = 1451
Score = 399 bits (1026), Expect = e-109
Identities = 264/753 (35%), Positives = 362/753 (48%), Gaps = 105/753 (13%)
Query: 19 PRPQAHAASTSPVPTDIAT-----AMHTMTLAPP--DSTWYMDTGASSHVASSPGNLTSY 71
P+ Q HA S + + A+ M A P W +D+GA+ H+ S NL +
Sbjct: 294 PQLQQHAGSYASNQSSSASYAPWQPRANMVSATPYNSGNWLLDSGATHHLTSDLNNLALH 353
Query: 72 SNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSHVLHTPQIVKNLISVRQLTT 131
+ ++++ + G G+PI SG LPT + L L VL+ P I KNLISV ++
Sbjct: 354 QPYNG-DEEVTIADGSGLPISHSGSALLPTPTRS--LALKDVLYVPDIQKNLISVYRMCN 410
Query: 132 DNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY--PVTPSVPFAGLAQSL-------WH 182
N VSV F P F V D TG L++ + +LY PV S+ + A WH
Sbjct: 411 TNGVSVEFFPAHFQVKDLSTGARLLQGKTKNELYEWPVNSSIATSMFASPTPKTDLPSWH 470
Query: 183 SRLGHPGSSALQSLRSNKFITYEH-LNSPTVCESCVFGKHVRLPFVSSNNVTVMPFDILH 241
+RLGHP L++L S + H L + +C C K +LPF S+ + P + L+
Sbjct: 471 ARLGHPSLPILKALISKFSLPISHSLQNQLLCSDCSINKSHKLPFYSNTIASSHPLEYLY 530
Query: 242 SDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKSQVFEMFTSLTHQIRTHFSHNVKC 301
+D+WTSP+ S ++YY++ +D +T + W +PL KSQV EMF + T + F +
Sbjct: 531 TDVWTSPITSIDNYKYYLVIVDHYTRYTWLYPLRKKSQVREMFITFTALVENKFKFKIGT 590
Query: 302 LQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVP 361
L DNG EF + ++ A+HG+ + PHT NG +ERK R I TLL+ AS+P
Sbjct: 591 LYSDNGGEF--IAMRSFLASHGISHMTTPPHTPELNGISERKHRHIVETGLTLLSTASMP 648
Query: 362 PSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSTTINK 421
+W +A A YL+N + L N SP L+ + P+Y LR+FGCLC+P + T +K
Sbjct: 649 KEYWSYAFATAVYLINRMLTPVLGNESPYVKLFGQPPNYLKLRIFGCLCFPWLRPYTAHK 708
Query: 422 LQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVIFDETRFPFAT------------- 468
L RS PCV LGY L Y C D + ++ SRHV F E+ FPF+T
Sbjct: 709 LDNRSVPCVLLGYSLSQSAYLCLDRATGRVYTSRHVQFAESSFPFSTTSPSVTPPSDPPL 768
Query: 469 ---------------MPTPPPSTYDCFSD--------------PIHPSIIHQWTSP---- 495
+ T PPS+ C + P+ PS+ TSP
Sbjct: 769 SQDTRPVSVPLLARPLTTAPPSSPSCSAPHRSPSQSENLSPPAPLQPSLSLSPTSPITSP 828
Query: 496 ------------DPSPTSVVPPAVTPSQLP------TTSTSSSASPP------------- 524
+ PT PP Q P +TS SS+ P
Sbjct: 829 SLSEESLVGHNSETGPTGSSPPLSPQPQRPQPQSPQSTSPHSSSPQPNSPNPQHSPRSLT 888
Query: 525 ----SSPSPPSPPQPVPP--IRTMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLA 578
SSPSP PP P PP TM TRS I KP F T P +PK A
Sbjct: 889 PTLTSSPSPSPPPNPNPPPIQHTMRTRSKNNIVKPNPKFANLATKPTPLKPIIPKTVVEA 948
Query: 579 LSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVG 638
L DPNW+ AM E +A RN T+DLVP + N++ C W+F K +NG +RYKARLV
Sbjct: 949 LLDPNWRQAMCDEINAQTRNGTFDLVPPAPNQNVVGCKWVFTLKYLSNGVLDRYKARLVA 1008
Query: 639 DGRSQIAGVDCDETFSPVVKPATIRTVLTIALS 671
G Q G D ETFSPV+K T+R+VL IA+S
Sbjct: 1009 KGFHQQYGHDFKETFSPVIKSTTVRSVLHIAVS 1041
>gb|AAC02669.1| polyprotein [Arabidopsis thaliana]
Length = 1451
Score = 399 bits (1024), Expect = e-109
Identities = 264/753 (35%), Positives = 362/753 (48%), Gaps = 105/753 (13%)
Query: 19 PRPQAHAASTSPVPTDIAT-----AMHTMTLAPP--DSTWYMDTGASSHVASSPGNLTSY 71
P+ Q HA S + + A+ M A P W +D+GA+ H+ S NL +
Sbjct: 294 PQLQQHAGSYASNQSSSASYAPWQPRANMVSATPYNSGNWLLDSGATHHLTSDLNNLALH 353
Query: 72 SNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSHVLHTPQIVKNLISVRQLTT 131
+ ++++ + G G+PI SG LPT + L L VL+ P I KNLISV ++
Sbjct: 354 QPYNG-DEEVTIADGSGLPISHSGSALLPTPTRS--LALKDVLYVPDIQKNLISVYRMCN 410
Query: 132 DNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY--PVTPSVPFAGLAQSL-------WH 182
N VSV F P F V D TG L++ + +LY PV S+ + A WH
Sbjct: 411 TNGVSVEFFPAHFQVKDLSTGARLLQGKTKNELYEWPVNSSIATSMFASPTPKTDLPSWH 470
Query: 183 SRLGHPGSSALQSLRSNKFITYEH-LNSPTVCESCVFGKHVRLPFVSSNNVTVMPFDILH 241
+RLGHP L++L S + H L + +C C K +LPF S+ + P + L+
Sbjct: 471 ARLGHPSLPILKALISKFSLPISHSLQNQLLCSDCSINKSHKLPFYSNTIASSHPLEYLY 530
Query: 242 SDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKSQVFEMFTSLTHQIRTHFSHNVKC 301
+D+WTSP+ S ++YY++ +D +T + W +PL KSQV EMF + T + F +
Sbjct: 531 TDVWTSPITSIDNYKYYLVIVDHYTRYTWLYPLRKKSQVREMFITFTALVENKFKFKIGT 590
Query: 302 LQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVP 361
L DNG EF + ++ A+HG+ + PHT NG +ERK R I TLL+ AS+P
Sbjct: 591 LYSDNGGEF--IAMRSFLASHGISHMTTPPHTPELNGISERKHRHIVETGLTLLSTASMP 648
Query: 362 PSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSTTINK 421
+W +A A YL+N + L N SP L+ + P+Y LR+FGCLC+P + T +K
Sbjct: 649 KEYWSYAFATAVYLINRMLTPVLGNESPYVKLFGQPPNYLKLRIFGCLCFPWLRPYTAHK 708
Query: 422 LQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVIFDETRFPFAT------------- 468
L RS PCV LGY L Y C D + ++ SRHV F E+ FPF+T
Sbjct: 709 LDNRSVPCVLLGYSLSQSAYLCLDRATGRVYTSRHVQFAESSFPFSTTSPSVTPPSDPPL 768
Query: 469 ---------------MPTPPPSTYDCFSD--------------PIHPSIIHQWTSP---- 495
+ T PPS+ C + P+ PS+ TSP
Sbjct: 769 SQDTRPVSVPLLARPLTTAPPSSPSCSAPHRSPSQSENLSPPAPLQPSLSLSPTSPITSP 828
Query: 496 ------------DPSPTSVVPPAVTPSQLP------TTSTSSSASPP------------- 524
+ PT PP Q P +TS SS+ P
Sbjct: 829 SLSEESLVGHNSETGPTGSSPPLSPQPQRPQPQSPQSTSPHSSSPQPNSPNPQHSPRSLT 888
Query: 525 ----SSPSPPSPPQPVPP--IRTMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKLA 578
SSPSP PP P PP TM TRS I KP F T P +PK A
Sbjct: 889 PTLTSSPSPSPPPNPNPPPIQHTMRTRSKNNIVKPNPKFANLATKPTPLKPIIPKTVVEA 948
Query: 579 LSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVG 638
L DPNW+ AM E +A RN T+DLVP + N++ C W+F K +NG +RYKARLV
Sbjct: 949 LLDPNWRQAMCDEINAQTRNGTFDLVPPAPNQNVVGCKWVFTLKYLSNGVLDRYKARLVA 1008
Query: 639 DGRSQIAGVDCDETFSPVVKPATIRTVLTIALS 671
G Q G D ETFSPV+K T+R+VL IA+S
Sbjct: 1009 KGFHQQYGHDFKETFSPVIKLTTVRSVLHIAVS 1041
>gb|AAP53905.1| putative pol polyprotein [Oryza sativa (japonica cultivar-group)]
gi|37534632|ref|NP_921618.1| putative pol polyprotein
[Oryza sativa (japonica cultivar-group)]
Length = 1688
Score = 374 bits (960), Expect = e-102
Identities = 260/726 (35%), Positives = 348/726 (47%), Gaps = 97/726 (13%)
Query: 15 GILGPRPQAHAASTSPVPTDIATAMHTMTLAPPDSTWYMDTGASSHVASSPGNLTSYSNL 74
GILGP P A AS S + T A W +D+GAS H++ LTS L
Sbjct: 157 GILGPYPCAGIASCSSISTVTPIA---------SQPWILDSGASFHMSFDDSWLTS-CRL 206
Query: 75 SHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSHVLHTPQIVKNLISVRQLTTDNN 134
++ +G + G + P T+ +V P++ NLISV QLT D N
Sbjct: 207 VKNGATVHTANGTLCKVTHQGSISSPQ------FTVPNVSLVPKLSMNLISVGQLT-DTN 259
Query: 135 VSVCFDPYGFSVHDFQTG--------------IPLMRCNSLGDLYPVTPSV--PFAGLAQ 178
V FD V D TG + ++ SL TPSV P A
Sbjct: 260 CFVGFDDTSCFVQDRHTGAVIGTGHRQKRSCGLYILDSLSLPSSSTNTPSVYSPMCSTAC 319
Query: 179 SL---WHSRLGHPGSSALQSLRSNKFITYEHLNSPTVCESCVFGKHVRLPFVSSNNVTVM 235
WH RLGH S L +L + + +++ VC+ C GK V+LP+ SS + +
Sbjct: 320 KSFPQWHHRLGHLCGSRLATLINQGVLGSVPVDTTFVCKGCKLGKQVQLPYPSSTSRSSR 379
Query: 236 PFDILHSDLW-TSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKSQVFEMFTSLTHQIRTH 294
PFD++HSD+W SP S GH YYV+F+DD++ + W + + ++SQ+ ++ S I T
Sbjct: 380 PFDLVHSDVWGKSPFPSKGGHNYYVIFVDDYSRYTWIYFMKHRSQLISIYQSFAQMIHTQ 439
Query: 295 FSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTL 354
FS ++ + D+G E+ + +F + + G + + SCP +QNG AERK R I RTL
Sbjct: 440 FSSAIRIFRSDSGGEYMSNAFREFLVSQGTLPQLSCPGAHAQNGVAERKHRHIIETARTL 499
Query: 355 LAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLV 414
L + VP FW A+ A YL+N+ P SL SP ++L+ P Y HLRVFGC CY L+
Sbjct: 500 LIASFVPAHFWAEAISTAVYLINMQPSSSLQGRSPGEVLFGSPPRYDHLRVFGCTCYVLL 559
Query: 415 PSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVIFDETRFPFATMPT--- 471
KL +S CVFLGY L H+GY+C+D S R+I ISR V FDE + PF T
Sbjct: 560 APRERTKLTAQSVECVFLGYSLEHKGYRCYDPSARRIRISRDVTFDENK-PFFYSSTNQP 618
Query: 472 -------------PPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTS 518
P PS S PI P SP P P SV P P P+ S S
Sbjct: 619 SSPENSISFLYLPPIPSPESLPSSPITP-------SPSPIPPSVPSPTYVPPPPPSPSPS 671
Query: 519 SSASPPSSPSPPSPPQPVPPIRTMATRSMRGIYKPRKLFNLS----VTIDDPTI-----S 569
+ PPS S P VP T+ T +P K+ N S T++DPT S
Sbjct: 672 PVSPPPSHIPASSSPPHVPSTITLDTFPFHYSRRP-KIPNESQPSQPTLEDPTCSVDDSS 730
Query: 570 PLPK---NPKLALSDPN-----------------------WKSAMQSEFDALIRNKTWDL 603
P P+ + AL PN WK AM E AL R TWD+
Sbjct: 731 PAPRYNLRARDALRAPNRDDFVVGVVFEPSTYQEAIVLPHWKLAMSEELAALERTNTWDV 790
Query: 604 VPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIR 663
VP PS I C W++ KTK++G ERYKARLV G Q G D DETF+PV T+R
Sbjct: 791 VPLPSHAVPITCKWVYKVKTKSDGQVERYKARLVARGFQQAHGRDYDETFAPVAHMTTVR 850
Query: 664 TVLTIA 669
T++ +A
Sbjct: 851 TLIAVA 856
>gb|AAD21687.1| Strong similarity to gi|3600044 T12H20.12 protease homolog from
Arabidopsis thaliana BAC gb|AF080119 and is a member of
the reverse transcriptase family PF|00078
gi|25301706|pir||C86438 hypothetical protein F28K20.17 -
Arabidopsis thaliana
Length = 1415
Score = 368 bits (944), Expect = e-100
Identities = 233/659 (35%), Positives = 325/659 (48%), Gaps = 40/659 (6%)
Query: 43 TLAPPDST---WYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTL 99
TL D T W+ D+ A++HV SS L S + + + VG G +PI +G TT+
Sbjct: 311 TLRVSDDTGKEWHPDSAATAHVTSSTNGLQSATEYEG-DDAVLVGDGTYLPITHTGSTTI 369
Query: 100 PTSHKHKPLTLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCN 159
+S+ PL + VL P I K+L+SV +L D V FD + D QT +
Sbjct: 370 KSSNGKIPL--NEVLVVPNIQKSLLSVSKLCDDYPCGVYFDANKVCIIDLQTQKVVTTGP 427
Query: 160 SLGDLYPVTPSVPFAGL--------AQSLWHSRLGHPGSSALQSLRSNKFITYEHLNSPT 211
LY V + F L + +WH RLGH S ALQ L+++K I +
Sbjct: 428 RRNGLY-VLENQEFVALYSNRQCAATEEVWHHRLGHANSKALQHLQNSKAIQINKSRTSP 486
Query: 212 VCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLW-TSPVLSSAGHRYYVLFLDDFTDFLW 270
VCE C GK RLPF+ S++ + P D +H DLW SPV+S+ G +YY +F+DD++ + W
Sbjct: 487 VCEPCQMGKSSRLPFLISDSRVLHPLDRIHCDLWGPSPVVSNQGLKYYAIFVDDYSRYSW 546
Query: 271 TFPLSNKSQVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSC 330
+PL NKS+ +F S + + +K Q D G EF + + + HG+ R SC
Sbjct: 547 FYPLHNKSEFLSVFISFQKLVENQLNTKIKVFQSDGGGEFVSNKLKTHLSEHGIHHRISC 606
Query: 331 PHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPT 390
P+T QNG AERK R + + ++L H+ P FW + A Y++N +P L NLSP
Sbjct: 607 PYTPQQNGLAERKHRHLVELGLSMLFHSHTPQKFWVESFFTANYIINRLPSSVLKNLSPY 666
Query: 391 QLLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRK 450
+ L+ P Y+ LRVFG CYP + NK PRS CVFLGY ++GY+CF K
Sbjct: 667 EALFGEKPDYSSLRVFGSACYPCLRPLAQNKFDPRSLQCVFLGYNSQYKGYRCFYPPTGK 726
Query: 451 IIISRHVIFDETRFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPS 510
+ ISR+VIF+E+ PF S +S P+ + H S P + V P
Sbjct: 727 VYISRNVIFNESELPF---KEKYQSLVPQYSTPLLQAWQHNKISEISVPAAPVQLFSKPI 783
Query: 511 QLPTTSTSSSASPPSSPSPPS----PPQPVPPI--------------RTMATRSMRGIYK 552
L T + S + P P S + V P+ M TRS GI K
Sbjct: 784 DLNTYAGSQVTEQLTDPEPTSNNEGSDEEVNPVAEEIAANQEQVINSHAMTTRSKAGIQK 843
Query: 553 PRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNI 612
P + L I + PK A+ P W A+ E + + TW LVP D+NI
Sbjct: 844 PNTRYAL---ITSRMNTAEPKTLASAMKHPGWNEAVHEEINRVHMLHTWSLVPPTDDMNI 900
Query: 613 IRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLTIALS 671
+ W+F K +G ++ KARLV G Q GVD ETFSPVV+ ATIR VL ++ S
Sbjct: 901 LSSKWVFKTKLHPDGSIDKLKARLVAKGFDQEEGVDYLETFSPVVRTATIRLVLDVSTS 959
>gb|AAU89779.1| gag-pol polyprotein-like [Solanum tuberosum]
Length = 1212
Score = 347 bits (889), Expect = 2e-93
Identities = 221/660 (33%), Positives = 339/660 (50%), Gaps = 56/660 (8%)
Query: 35 IATAMHTMTLAPPDST---WYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPI 91
I +A + L D T W +D+GAS+H+ +S L + +Q + + +G +PI
Sbjct: 301 IVSAFSALGLQGNDVTSNFWIVDSGASNHMTNSTSILKNVRKYQGPSQ-IQIANGSNLPI 359
Query: 92 QGSGDTTLPTSHKHKPLTLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQT 151
GD T T +V +P++ +LISV QL DNN V F G V D +
Sbjct: 360 TKVGDITP---------TFKNVFVSPKLSTSLISVGQLV-DNNCDVNFSRNGCLVQDQVS 409
Query: 152 GIPLMRCNSLGDLYPVTPSVP----FAGLAQS----LWHSRLGHPGSSALQSLRSNKFIT 203
G + + +G L+P+ S+P FA + + +WH RLGHP S L + ++ +
Sbjct: 410 GTIIAKGPKVGRLFPIHFSIPPVLSFACTSTASKTEVWHKRLGHPNSVVLSHISNSGLLG 469
Query: 204 YEHLNSPTV--CESCVFGKHVRLPFVSSNNVTVMPFDILHSDLW-TSPVLSSAGHRYYVL 260
++ S C +C GK LPF + + FD++HSD+W SP++S A +Y++
Sbjct: 470 NKNKFSVASIDCSTCKLGKSKTLPFPNFGSRATKCFDVIHSDVWGISPIISHAHFKYFMT 529
Query: 261 FLDDFTDFLWTFPLSNKSQVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCA 320
F+DD++ F W + L +KS+VF MF + I T FS +K L+ D+G E+ + F +
Sbjct: 530 FIDDYSRFTWVYFLRSKSEVFSMFKTFLAYIETQFSTCIKLLRSDSGGEYMSYEFKKFLL 589
Query: 321 AHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIP 380
G++ + SCP+T QNG AERK R + ++ RTLL +SVP +W AL A YL+N +P
Sbjct: 590 DKGIVSQHSCPYTPQQNGVAERKNRHLLDVTRTLLIESSVPSKYWVEALSTAVYLINRLP 649
Query: 381 RKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRG 440
K L+ SP LY ++P+Y+ FGC+C+ +P + NKL +S+ C F+GY +G
Sbjct: 650 SKVLNLESPYFRLYHQNPNYSDFHTFGCVCFVHLPPSQCNKLSVQSTKCAFMGYSTSQKG 709
Query: 441 YKCFDLSHRKIIISRHVIFDETRFPFATM-----PTPPPSTYDCFSDP---IHPSIIHQW 492
+ C+D K ISR+V+F E ++ F T+ +P T++ S P +++
Sbjct: 710 FICYDPCSHKFRISRNVVFFENQYFFPTIVDLSSVSPLLPTFEDLSSSFKRFKPGFVYER 769
Query: 493 TSPD-PSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSPPSPPQPVPPIRTMATRSMRGIY 551
P P P + PP P S +SS S P P TR +
Sbjct: 770 RRPTLPYPNTDPPPETAPQ---LESENSSRSGPLEP----------------TRRSTRVS 810
Query: 552 KPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVN 611
+ + S T+ + ++ P A W+ AM+ E AL N TWD+V PS+V
Sbjct: 811 RTPNWYGFSSTLSNISV---PSCYSQASKHECWQKAMEEELLALKENDTWDIVSCPSNVR 867
Query: 612 IIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLTIALS 671
I C W++ K ++G +RYKARLV G Q GVD +ETF+PV K T+RT++ IA S
Sbjct: 868 PIGCKWVYSIKLHSDGTLDRYKARLVVLGNRQEYGVDYEETFAPVAKMTTVRTIIAIAAS 927
>gb|AAG46116.1| putative copia-like retrotransposon polyprotein [Oryza sativa]
Length = 1302
Score = 342 bits (877), Expect = 4e-92
Identities = 183/398 (45%), Positives = 236/398 (58%), Gaps = 29/398 (7%)
Query: 295 FSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTL 354
F V LQ DNG E+D+ + + + HG + R SCP++S QNGKAER +RTIN+ +RT+
Sbjct: 515 FGLPVLALQTDNGKEYDSYALRSLLSLHGAVLRLSCPYSSQQNGKAERILRTINDYVRTM 574
Query: 355 LAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLV 414
L H++ P SFW ALQ AT+L+N P ++ +L+P QLL P+Y HLRVFGCLCYP
Sbjct: 575 LVHSAAPLSFWAEALQTATHLINRRPCRATGSLTPYQLLLGAPPTYDHLRVFGCLCYPNT 634
Query: 415 PSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVIFDETRFPFATMPTPPP 474
+T +KL PRS CVF+GYP HRGY+C+D+ R++ SRHV F E FPF P+P P
Sbjct: 635 IATAPHKLSPRSLACVFIGYPADHRGYRCYDMVSRRVFTSRHVTFVEDVFPFRDAPSPRP 694
Query: 475 ST---YDCFSD--------------PIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTST 517
S D D P+ + H SP PSP S P + P+
Sbjct: 695 SAPPPPDHGDDTIVLLPAPAQHVVTPVGTAPAHDAASP-PSPASSTPSSAAPAH------ 747
Query: 518 SSSASPPSSPSPPSPPQPVPPIRTMATRSMRGIYKPRKLFNLSVTIDDPTISPLPKNPKL 577
+PP SP SP PP M TR+ GI KP + ++ T T+SP P + +
Sbjct: 748 --DVAPPPSPETSSPASASPPRHAMTTRARAGISKPNPRYAMTAT---STLSPTPSSVRA 802
Query: 578 ALSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLV 637
AL DPNW++AMQ+EFDAL+ N+TW LVPRP II W+F K A+G ++YKAR V
Sbjct: 803 ALRDPNWRAAMQAEFDALLANRTWTLVPRPPGARIITGKWVFKTKLHADGSLDKYKARWV 862
Query: 638 GDGRSQIAGVDCDETFSPVVKPATIRTVLTIALSGLGP 675
G +Q GVD ETFSPVVKPATIRTVLT+ S P
Sbjct: 863 VRGFNQRPGVDFGETFSPVVKPATIRTVLTLISSKQWP 900
Score = 137 bits (344), Expect = 3e-30
Identities = 81/198 (40%), Positives = 116/198 (57%), Gaps = 12/198 (6%)
Query: 1 TSQWTRPSAPPRQPGILGPRPQAHAASTSPVPTDIATAMHTMTLAPPDSTWYMDTGASSH 60
T+Q P+ PP PG+ A S + + + +A T P + W++DTGAS+H
Sbjct: 322 TAQHLLPALPPASPGV----HSTGAWDNSALYSALQSAGVATTTPPSAADWFLDTGASAH 377
Query: 61 VASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSHVLHTPQIV 120
++S+PG L L + + VG+G +P+ + T +PTS L L +VL +P ++
Sbjct: 378 MSSTPGILAHPRPLP-FSSCITVGNGAKLPVTHTASTHIPTSSTD--LHLHNVLVSPPLI 434
Query: 121 KNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLYPVTPSVPFAGLAQSL 180
KNLISV++LT DNNVS+ FDP GFS+ D QT + +RC+S GDLYP+ P A A S
Sbjct: 435 KNLISVKKLTRDNNVSIEFDPTGFSIKDLQTQVVKLRCDSPGDLYPLRLPSPHALSATSS 494
Query: 181 -----WHSRLGHPGSSAL 193
WH RLGHPGS++L
Sbjct: 495 PSVEHWHLRLGHPGSASL 512
>emb|CAC95126.1| gag-pol polyprotein [Populus deltoides]
Length = 1382
Score = 338 bits (868), Expect = 4e-91
Identities = 235/681 (34%), Positives = 337/681 (48%), Gaps = 51/681 (7%)
Query: 17 LGPRPQAHAAST-SPVPTDIATAMHTMTLAPPDSTWYMDTGASSHVASSPGNLTSYSNLS 75
L +PQA +AS+ +P + H S W +D+GAS H++ + TS S LS
Sbjct: 315 LSLQPQAMSASSIGQLPHSSSGISH--------SEWVLDSGASHHMSPDSSSFTSVSPLS 366
Query: 76 HLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPLTLSHVLHTPQIVKNLISVRQLTTDNNV 135
+ V + G P+ +G ++ T H L+L +V P++ NL S+ Q+ +
Sbjct: 367 SIP----VMTADGTPMPLAGVGSVVTLH----LSLPNVYLIPKLKLNLASIGQICDSGDY 418
Query: 136 SVCFDPYGFSVHDFQTGIPLMRCNSLGDLY-------PV-----TPSVPFAGLAQS---- 179
V F V D Q+ + LY PV T + F L+ S
Sbjct: 419 LVMFSGSFCCVQDLQSQKLIGTGRRENGLYILDELKVPVVVAATTVDLSFFRLSLSSSSF 478
Query: 180 -LWHSRLGHPGSSALQSLRSNKFITYEHLNSPTVCESCVFGKHVRLPFVSSNNVTVMPFD 238
LWHSRLGH SS L+ L S + + C C K LPF S +V+ PFD
Sbjct: 479 YLWHSRLGHVSSSRLRFLASTGALGNLKTCDISDCSGCKLAKFSALPFNRSTSVSSSPFD 538
Query: 239 ILHSDLW-TSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKSQVFEMFTSLTHQIRTHFSH 297
++HSD+W SPV + G RYYV F+DD T + W + + ++S+ FE++ + I+T S
Sbjct: 539 LIHSDVWGPSPVSTKGGSRYYVSFIDDHTRYCWVYLMKHRSEFFEIYAAFRALIKTQHSA 598
Query: 298 NVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAH 357
+KC +CD G E+ + F A G I + SC T QNG AERK R I R+LL
Sbjct: 599 VIKCFRCDLGGEYTSNKFCQMLALDGTIHQTSCTDTPEQNGVAERKHRHIVETARSLLLS 658
Query: 358 ASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDPSYTHLRVFGCLCYPLVPST 417
A V FW A+ A L+N IP S LSP + LY P Y+ RVFGC + L P
Sbjct: 659 AFVLSEFWGEAVLTAVSLINTIPSSHSSGLSPFEKLYGHVPDYSSFRVFGCTYFVLHPHV 718
Query: 418 TINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVIFDETRFPFATMPTPPPSTY 477
NKL RS+ CVFLGY +GY+CFD +K+ +S HV+F E PF ++P+ S
Sbjct: 719 ERNKLSSRSAICVFLGYGEGKKGYRCFDPITQKLYVSHHVVFLE-HIPFFSIPSTTHSLT 777
Query: 478 DCFSDPIHPSIIHQWTSPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSPPSPPQPVP 537
SD IH I ++ + TS ++ T T S +P +S S +P
Sbjct: 778 K--SDLIH---IDPFSEDSGNDTSPYVRSICTHNSAGTGTLLSGTPEASFSSTAPQASSE 832
Query: 538 PIRTMATRSMRGIYKPRKLFNLSVTIDDPTISPL---------PKNPKLALSDPNWKSAM 588
+ +S+R I K KL + + + + + P + K A+ DP + AM
Sbjct: 833 IVDPPPRQSIR-IRKSTKLPDFAYSCYSSSFTSFLAYIHCLFEPSSYKEAILDPLGQQAM 891
Query: 589 QSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVD 648
E AL + TWDLVP P +++ C W++ KT ++G ERYKARLV G SQ G+D
Sbjct: 892 DEELSALHKTDTWDLVPLPPGKSVVGCRWVYKIKTNSDGSIERYKARLVAKGYSQQYGMD 951
Query: 649 CDETFSPVVKPATIRTVLTIA 669
+ETF+P+ K TIRT++ +A
Sbjct: 952 YEETFAPIAKMTTIRTLIAVA 972
>emb|CAB77781.1| putative polyprotein of LTR transposon [Arabidopsis thaliana]
gi|3924609|gb|AAC79110.1| putative polyprotein of LTR
transposon [Arabidopsis thaliana] gi|7444420|pir||T01397
LTR gag/pol polyprotein homolog T4I9.16 - Arabidopsis
thaliana
Length = 1456
Score = 317 bits (811), Expect = 2e-84
Identities = 202/579 (34%), Positives = 291/579 (49%), Gaps = 45/579 (7%)
Query: 49 STWYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPL 108
+ W +D+GA+ H+ S NL+ + + + + G IPI +G +LPTS + L
Sbjct: 308 NNWLLDSGATHHITSDFNNLSFHQPYTG-GDDVMIADGSTIPITHTGSASLPTS--SRSL 364
Query: 109 TLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY--P 166
L+ VL+ P I KNLISV +L N VSV F P F V D TG+PL++ + +LY P
Sbjct: 365 DLNKVLYVPNIHKNLISVYRLCNTNRVSVEFFPASFQVKDLNTGVPLLQGKTKDELYEWP 424
Query: 167 VTPS-------VPFAGLAQSLWHSRLGHPGSSALQSLRSNKFITYEHLNSPTV-CESCVF 218
+ S P + S WHSRLGHP + L S+ SN + + + + C C
Sbjct: 425 IASSQAVSMFASPCSKATHSSWHSRLGHPSLAILNSVISNHSLPVLNPSHKLLSCSDCFI 484
Query: 219 GKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKS 278
K ++PF +S + P + ++SD+W+SP+LS +RYYV+F+D FT + W +PL KS
Sbjct: 485 NKSHKVPFSNSTITSSKPLEYIYSDVWSSPILSIDNYRYYVIFVDHFTRYTWLYPLKQKS 544
Query: 279 QVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNG 338
QV + F + F + L DNG EF Y + HG+ S PHT NG
Sbjct: 545 QVKDTFIIFKSLVENRFQTRIGTLYSDNGGEF--VVLRDYLSQHGISHFTSPPHTPEHNG 602
Query: 339 KAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDP 398
+ERK R I M TLL+HASVP ++W +A +A YL+N +P L SP Q L+ + P
Sbjct: 603 LSERKHRHIVEMGLTLLSHASVPKTYWPYAFSVAVYLINRLPTPLLQLQSPFQKLFGQPP 662
Query: 399 SYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVI 458
+Y L+VFGC CYP + +KL+ +S C F+GY L Y C + ++ SRHV
Sbjct: 663 NYEKLKVFGCACYPWLRPYNRHKLEDKSKQCAFMGYSLTQSAYLCLHIPTGRLYTSRHVQ 722
Query: 459 FDETRFPFATMPTPPPSTYDCFSD-----------PIHPSII--------HQWTSPDP-- 497
FDE FPF+T ++ + SD P P ++ H TSP P
Sbjct: 723 FDERCFPFSTTNFGVSTSQEQRSDSAPNWPSHTTLPTTPLVLPAPPCLGPHLDTSPRPPS 782
Query: 498 SPTSVVPPAVTPSQLPTTSTSS-SASPPSSPSPPSPPQPVPPIRTMATRSMRGIY----- 551
SP+ + V+ S LP++S SS S+S P++PS P P +T + S I
Sbjct: 783 SPSPLCTTQVSSSNLPSSSISSPSSSEPTAPSHNGPQPTAQPHQTQNSNSNSPILNNPNP 842
Query: 552 ---KPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSA 587
P S P SP P ++S+PN S+
Sbjct: 843 NSPSPNSPNQNSPLPQSPISSPHIPTPSTSISEPNSPSS 881
Score = 123 bits (308), Expect = 4e-26
Identities = 75/189 (39%), Positives = 107/189 (55%), Gaps = 11/189 (5%)
Query: 494 SPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSPPSPP----QPVPPIRT--MATRSM 547
SP SP + P+ + S+ + S+SS+++PP P P+PP P+ T MATR+
Sbjct: 859 SPISSP-HIPTPSTSISEPNSPSSSSTSTPPLPPVLPAPPIIQVNAQAPVNTHSMATRAK 917
Query: 548 RGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRP 607
GI KP + ++ + ++ + P+ A+ D W+ AM SE +A I N TWDLVP P
Sbjct: 918 DGIRKPNQKYSYATSL---AANSEPRTAIQAMKDDRWRQAMGSEINAQIGNHTWDLVPPP 974
Query: 608 S-DVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVL 666
V I+ C WIF K ++G RYKARLV G +Q G+D ETFSPV+K +IR VL
Sbjct: 975 PPSVTIVGCRWIFTKKFNSDGSLNRYKARLVAKGYNQRPGLDYAETFSPVIKSTSIRIVL 1034
Query: 667 TIALSGLGP 675
+A+ P
Sbjct: 1035 GVAVDRSWP 1043
>dbj|BAA78424.1| polyprotein [Arabidopsis thaliana]
Length = 1330
Score = 317 bits (811), Expect = 2e-84
Identities = 202/579 (34%), Positives = 291/579 (49%), Gaps = 45/579 (7%)
Query: 49 STWYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPL 108
+ W +D+GA+ H+ S NL+ + + + + G IPI +G +LPTS + L
Sbjct: 182 NNWLLDSGATHHITSDFNNLSFHQPYTG-GDDVMIADGSTIPITHTGSASLPTS--SRSL 238
Query: 109 TLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY--P 166
L+ VL+ P I KNLISV +L N VSV F P F V D TG+PL++ + +LY P
Sbjct: 239 DLNKVLYVPNIHKNLISVYRLCNTNRVSVEFFPASFQVKDLNTGVPLLQGKTKDELYEWP 298
Query: 167 VTPS-------VPFAGLAQSLWHSRLGHPGSSALQSLRSNKFITYEHLNSPTV-CESCVF 218
+ S P + S WHSRLGHP + L S+ SN + + + + C C
Sbjct: 299 IASSQAVSMFASPCSKATHSSWHSRLGHPSLAILNSVISNHSLPVLNPSHKLLSCSDCFI 358
Query: 219 GKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKS 278
K ++PF +S + P + ++SD+W+SP+LS +RYYV+F+D FT + W +PL KS
Sbjct: 359 NKSHKVPFSNSTITSSKPLEYIYSDVWSSPILSIDNYRYYVIFVDHFTRYTWLYPLKQKS 418
Query: 279 QVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNG 338
QV + F + F + L DNG EF Y + HG+ S PHT NG
Sbjct: 419 QVKDTFIIFKSLVENRFQTRIGTLYSDNGGEF--VVLRDYLSQHGISHFTSPPHTPEHNG 476
Query: 339 KAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDP 398
+ERK R I M TLL+HASVP ++W +A +A YL+N +P L SP Q L+ + P
Sbjct: 477 LSERKHRHIVEMGLTLLSHASVPKTYWPYAFSVAVYLINRLPTPLLQLQSPFQKLFGQPP 536
Query: 399 SYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVI 458
+Y L+VFGC CYP + +KL+ +S C F+GY L Y C + ++ SRHV
Sbjct: 537 NYEKLKVFGCACYPWLRPYNRHKLEDKSKQCAFMGYSLTQSAYLCLHIPTGRLYTSRHVQ 596
Query: 459 FDETRFPFATMPTPPPSTYDCFSD-----------PIHPSII--------HQWTSPDP-- 497
FDE FPF+T ++ + SD P P ++ H TSP P
Sbjct: 597 FDERCFPFSTTNFGVSTSQEQRSDSAPNWPSHTTLPTTPLVLPAPPCLGPHLDTSPRPPS 656
Query: 498 SPTSVVPPAVTPSQLPTTSTSS-SASPPSSPSPPSPPQPVPPIRTMATRSMRGIY----- 551
SP+ + V+ S LP++S SS S+S P++PS P P +T + S I
Sbjct: 657 SPSPLCTTQVSSSNLPSSSISSPSSSEPTAPSHNGPQPTAQPHQTQNSNSNSPILNNPNP 716
Query: 552 ---KPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSA 587
P S P SP P ++S+PN S+
Sbjct: 717 NSPSPNSPNQNSPLPQSPISSPHIPTPSTSISEPNSPSS 755
Score = 123 bits (308), Expect = 4e-26
Identities = 75/189 (39%), Positives = 107/189 (55%), Gaps = 11/189 (5%)
Query: 494 SPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSPPSPP----QPVPPIRT--MATRSM 547
SP SP + P+ + S+ + S+SS+++PP P P+PP P+ T MATR+
Sbjct: 733 SPISSP-HIPTPSTSISEPNSPSSSSTSTPPLPPVLPAPPIIQVNAQAPVNTHSMATRAK 791
Query: 548 RGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRP 607
GI KP + ++ + ++ + P+ A+ D W+ AM SE +A I N TWDLVP P
Sbjct: 792 DGIRKPNQKYSYATSL---AANSEPRTAIQAMKDDRWRQAMGSEINAQIGNHTWDLVPPP 848
Query: 608 S-DVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVL 666
V I+ C WIF K ++G RYKARLV G +Q G+D ETFSPV+K +IR VL
Sbjct: 849 PPSVTIVGCRWIFTKKFNSDGSLNRYKARLVAKGYNQRPGLDYAETFSPVIKSTSIRIVL 908
Query: 667 TIALSGLGP 675
+A+ P
Sbjct: 909 GVAVDRSWP 917
>gb|AAP46257.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|50919599|ref|XP_470160.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1335
Score = 315 bits (807), Expect = 5e-84
Identities = 204/648 (31%), Positives = 327/648 (49%), Gaps = 53/648 (8%)
Query: 48 DSTWYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKP 107
D W +D+G ++H+A+ P + H K+++G+G +G G + T+ P
Sbjct: 323 DDVWVIDSGCTNHMAADPNLFREMDSSYHA--KIHMGNGSIAQSEGKGTVAVQTADG--P 378
Query: 108 LTLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCN---SLGDL 164
+ VL P + +NL+S+ QL ++ +V F+ + + D + + + N + L
Sbjct: 379 KFIKDVLLVPDLKQNLLSIGQLL-EHGYAVYFEDFSCKILDRKNNRLVAKINMEKNRNFL 437
Query: 165 YPVTPSVPFAGLAQ----SLWHSRLGHPGSSALQSLRSN------KFITYEHLNSPTVCE 214
+ + A ++ LWH R+GH AL+ LR+ FIT + + P CE
Sbjct: 438 LRMNHTTQMALRSEVDISDLWHKRMGHLNYRALKLLRTKGMVQGLPFITLK--SDP--CE 493
Query: 215 SCVFGKHVRLPFVSSNNVTVM-PFDILHSDLWTS-PVLSSAGHRYYVLFLDDFTDFLWTF 272
CVFGK +R F S P +++H+D+ P +S G+ Y++ F+DD+T +W +
Sbjct: 494 GCVFGKQIRASFPHSGAWRASAPLELVHADIVGKVPTISEGGNWYFITFIDDYTRMIWVY 553
Query: 273 PLSNKSQVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPH 332
L KS E+F + + +K L+ D G E+ + F YC G+ + + +
Sbjct: 554 FLKEKSAALEIFKKFKAMVENQSNRKIKVLRSDQGREYISKEFEKYCENAGIRRQLTAGY 613
Query: 333 TSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQL 392
++ QNG AERK RTIN+M ++L +P SFW A+ A Y+LN P K+++N +P +
Sbjct: 614 SAQQNGVAERKNRTINDMANSMLQDKGMPKSFWAEAVNTAVYILNRSPTKAVTNRTPFEA 673
Query: 393 LYRRDPSYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKII 452
Y + P H+RVFGC+CY VP+ K +S C+F+GY +GY+ ++L +KII
Sbjct: 674 WYGKKPVIGHMRVFGCICYAQVPAQKRVKFDNKSDRCIFVGYADGIKGYRLYNLEKKKII 733
Query: 453 ISRHVIFDET-----RFPFATMPTPPPSTYDCFSDPIHPSIIHQWTSPDPSPTSVVPPAV 507
ISR IFDE+ + P A+ P+T P H H+ PSP
Sbjct: 734 ISRDAIFDESATWNWKSPEASSTPLLPTTTITLGQP-HMHGTHEVEDHTPSP-------- 784
Query: 508 TPSQLPTTSTSSSASPPSSPSPPSPPQPVP-PIRTM-----ATRSMRGIYKPRKLFNLSV 561
PS ++S++SS S PSS S P+ P +R+M +T RG + + N SV
Sbjct: 785 QPSSPMSSSSASSDSSPSSEEQISTPESAPRRVRSMVELLESTSQQRG-SEQHEFCNYSV 843
Query: 562 TIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRPSDVNIIRCMWIFHH 621
P++ + A NW AM+ E + +N TW+LV RP D +I W++
Sbjct: 844 V--------EPQSFQEAEKHDNWIKAMEDEIHMIEKNNTWELVDRPRDREVIGVKWVYKT 895
Query: 622 KTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLTIA 669
K +G ++YKARLV G Q G+D ET++PV + TIRT++ +A
Sbjct: 896 KLNPDGSVQKYKARLVAKGFKQKPGIDYYETYAPVARLETIRTIIALA 943
>dbj|BAA78426.1| polyprotein [Arabidopsis thaliana]
Length = 1475
Score = 315 bits (806), Expect = 7e-84
Identities = 201/579 (34%), Positives = 290/579 (49%), Gaps = 45/579 (7%)
Query: 49 STWYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPL 108
+ W +D+GA+ H+ S NL+ + + + + G IPI +G +LPTS + L
Sbjct: 327 NNWLLDSGATHHITSDFNNLSFHQPYTG-GDDVMIADGSTIPITHTGSASLPTS--SRSL 383
Query: 109 TLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY--P 166
L+ VL+ P I KNLISV +L N VSV F P F V D TG+PL++ + +LY P
Sbjct: 384 DLNKVLYVPNINKNLISVYRLCNTNRVSVEFFPASFQVKDLNTGVPLLQGKTKDELYEWP 443
Query: 167 VTPS-------VPFAGLAQSLWHSRLGHPGSSALQSLRSNKFITYEHLNSPTV-CESCVF 218
+ S P + S WHSRLGHP + L S+ SN + + + + C C
Sbjct: 444 IASSQAVSMFASPCSKATHSSWHSRLGHPSLAILNSVISNHSLPVLNPSHKLLSCSDCFI 503
Query: 219 GKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKS 278
K ++PF +S + P + ++SD+W+SP+LS +RYYV+F+D FT + W +PL KS
Sbjct: 504 NKSHKVPFSNSTITSSKPLEYIYSDVWSSPILSIDNYRYYVIFVDHFTRYTWLYPLKQKS 563
Query: 279 QVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNG 338
QV + F + F + L DNG EF Y + HG+ S PHT NG
Sbjct: 564 QVKDTFIIFKSLVENRFQTRIGTLYSDNGGEF--VVLRDYLSQHGISHFTSPPHTPEHNG 621
Query: 339 KAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDP 398
+ERK R I M TLL+HASVP ++W +A +A YL+N +P L SP Q L+ + P
Sbjct: 622 LSERKHRHIVEMGLTLLSHASVPKTYWPYAFSVAVYLINRLPTPLLQLQSPFQKLFGQPP 681
Query: 399 SYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVI 458
+Y L+VFGC CYP + +KL+ +S C F+GY L Y C + ++ SRHV
Sbjct: 682 NYEKLKVFGCACYPWLRPYNRHKLEDKSKQCAFMGYSLTQSAYLCLHIPTGRLYTSRHVQ 741
Query: 459 FDETRFPFATMPTPPPSTYDCFSD-----------PIHPSII--------HQWTSPDPS- 498
FDE FPF+T ++ + SD P P ++ H TSP P
Sbjct: 742 FDERCFPFSTTNFGVSTSQEQRSDSAPNWPSHTTLPTTPLVLPAPPCLGPHLDTSPRPPS 801
Query: 499 -PTSVVPPAVTPSQLPTTSTSS-SASPPSSPSPPSPPQPVPPIRTMATRSMRGIY----- 551
P+ + V+ S LP++S SS S+S P++PS P P +T + S I
Sbjct: 802 LPSPLCTTQVSSSNLPSSSISSPSSSEPTAPSHNGPQPTAQPHQTQNSNSNSPILNNPNP 861
Query: 552 ---KPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSA 587
P S P SP P ++S+PN S+
Sbjct: 862 NSPSPNSPNQNSPLPQSPISSPHIPTPSTSISEPNSPSS 900
Score = 121 bits (304), Expect = 1e-25
Identities = 74/189 (39%), Positives = 107/189 (56%), Gaps = 11/189 (5%)
Query: 494 SPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSPPSPP----QPVPPIRT--MATRSM 547
SP SP + P+ + S+ + S+SS+++PP P P+PP P+ T MATR+
Sbjct: 878 SPISSP-HIPTPSTSISEPNSPSSSSTSTPPLPPVLPAPPIIQVNAQAPVNTHSMATRAK 936
Query: 548 RGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRP 607
GI KP + ++ + ++ + P+ A+ D W+ AM S+ +A I N TWDLVP P
Sbjct: 937 DGIRKPNQKYSYATSL---AANSEPRTAIQAMKDDRWRQAMGSKINAQIGNHTWDLVPPP 993
Query: 608 S-DVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVL 666
V I+ C WIF K ++G RYKARLV G +Q G+D ETFSPV+K +IR VL
Sbjct: 994 PPSVTIVGCRWIFTKKFNSDGSLNRYKARLVAKGYNQRPGLDYAETFSPVIKSTSIRIVL 1053
Query: 667 TIALSGLGP 675
+A+ P
Sbjct: 1054 GVAVDRSWP 1062
>dbj|BAA78427.1| polyprotein [Arabidopsis thaliana]
Length = 1421
Score = 314 bits (804), Expect = 1e-83
Identities = 201/579 (34%), Positives = 290/579 (49%), Gaps = 45/579 (7%)
Query: 49 STWYMDTGASSHVASSPGNLTSYSNLSHLNQKLYVGSGQGIPIQGSGDTTLPTSHKHKPL 108
+ W +D+GA+ H+ S NL+ + + + + G IPI +G +LPTS + L
Sbjct: 327 NNWLLDSGATHHITSDFNNLSFHQPYTG-GDDVMIADGSTIPITHTGSASLPTS--SRSL 383
Query: 109 TLSHVLHTPQIVKNLISVRQLTTDNNVSVCFDPYGFSVHDFQTGIPLMRCNSLGDLY--P 166
L+ VL+ P I KNLISV +L N VSV F P F V D TG+PL++ + +LY P
Sbjct: 384 DLNKVLYVPNIHKNLISVYRLCNTNRVSVEFFPASFQVKDLNTGVPLLQGKTKDELYEWP 443
Query: 167 VTPS-------VPFAGLAQSLWHSRLGHPGSSALQSLRSNKFITYEHLNSPTV-CESCVF 218
+ S P + WHSRLGHP + L S+ SN + + + + C C
Sbjct: 444 IASSQAVSMFASPCSKATHCSWHSRLGHPSLAILNSVISNHSLPVLNPSHKLLSCSDCFI 503
Query: 219 GKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRYYVLFLDDFTDFLWTFPLSNKS 278
K ++PF +S + P + ++SD+W+SP+LS +RYYV+F+D FT + W +PL KS
Sbjct: 504 NKSHKVPFSNSTITSSKPLEYIYSDVWSSPILSIDNYRYYVIFVDHFTRYTWLYPLKQKS 563
Query: 279 QVFEMFTSLTHQIRTHFSHNVKCLQCDNGPEFDNTSFHAYCAAHGLIFRFSCPHTSSQNG 338
QV + F + F + L DNG EF Y + HG+ S PHT NG
Sbjct: 564 QVKDTFIIFKSLVENRFQTRIGTLYSDNGGEF--VVLRDYLSQHGISHFTSPPHTPEHNG 621
Query: 339 KAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKSLSNLSPTQLLYRRDP 398
+ERK R I M TLL+HASVP ++W +A +A YL+N +P L SP Q L+ + P
Sbjct: 622 LSERKHRHIVEMGLTLLSHASVPKTYWLYAFSVAVYLINRLPTPLLQLQSPFQKLFGQPP 681
Query: 399 SYTHLRVFGCLCYPLVPSTTINKLQPRSSPCVFLGYPLHHRGYKCFDLSHRKIIISRHVI 458
+Y L+VFGC CYP + +KL+ +S C F+GY L Y C + ++ SRHV
Sbjct: 682 NYEKLKVFGCACYPWLRPYNRHKLEDKSKQCAFMGYSLTQSAYLCLHIPTGRLYTSRHVQ 741
Query: 459 FDETRFPFATMPTPPPSTYDCFSD-----------PIHPSII--------HQWTSPDP-- 497
FDE FPF+T ++ + SD P P ++ H TSP P
Sbjct: 742 FDERCFPFSTTNFGVSTSQEQRSDSASNWPSHTTLPTTPLVLPAPPCLGPHLDTSPRPPS 801
Query: 498 SPTSVVPPAVTPSQLPTTSTSS-SASPPSSPSPPSPPQPVPPIRTMATRSMRGIY----- 551
SP+ + V+ S LP++S SS S+S P++PS P P +T + S I
Sbjct: 802 SPSPLCTTQVSSSNLPSSSISSPSSSEPTAPSYNGPQPTTQPHQTQNSNSNSPILNNPNP 861
Query: 552 ---KPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSA 587
P S P SP P ++S+PN S+
Sbjct: 862 NSPSPNSPNQNSPLPQSPISSPHIPTPSTSISEPNSPSS 900
Score = 123 bits (308), Expect = 4e-26
Identities = 75/189 (39%), Positives = 107/189 (55%), Gaps = 11/189 (5%)
Query: 494 SPDPSPTSVVPPAVTPSQLPTTSTSSSASPPSSPSPPSPP----QPVPPIRT--MATRSM 547
SP SP + P+ + S+ + S+SS+++PP P P+PP P+ T MATR+
Sbjct: 878 SPISSP-HIPTPSTSISEPNSPSSSSTSTPPLPPVLPAPPIIQVNAQAPVNTHSMATRAK 936
Query: 548 RGIYKPRKLFNLSVTIDDPTISPLPKNPKLALSDPNWKSAMQSEFDALIRNKTWDLVPRP 607
GI KP + ++ + ++ + P+ A+ D W+ AM SE +A I N TWDLVP P
Sbjct: 937 DGIRKPNQKYSYATSL---AANSEPRTAIQAMKDDRWRQAMGSEINAQIGNHTWDLVPPP 993
Query: 608 S-DVNIIRCMWIFHHKTKANGCFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVL 666
V I+ C WIF K ++G RYKARLV G +Q G+D ETFSPV+K +IR VL
Sbjct: 994 PPSVTIVGCRWIFTKKFNSDGSLNRYKARLVAKGYNQRPGLDYAETFSPVIKSTSIRIVL 1053
Query: 667 TIALSGLGP 675
+A+ P
Sbjct: 1054 GVAVDRSWP 1062
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.329 0.140 0.451
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,911,938,973
Number of Sequences: 2540612
Number of extensions: 86835327
Number of successful extensions: 764871
Number of sequences better than 10.0: 5097
Number of HSP's better than 10.0 without gapping: 1855
Number of HSP's successfully gapped in prelim test: 3517
Number of HSP's that attempted gapping in prelim test: 645706
Number of HSP's gapped (non-prelim): 42128
length of query: 1082
length of database: 863,360,394
effective HSP length: 139
effective length of query: 943
effective length of database: 510,215,326
effective search space: 481133052418
effective search space used: 481133052418
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 81 (35.8 bits)
Lotus: description of TM0114.10


