

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0114.1
(661 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
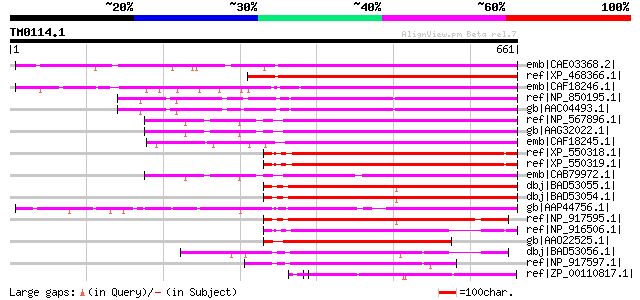
Score E
Sequences producing significant alignments: (bits) Value
emb|CAE03368.2| OSJNBb0065L13.11 [Oryza sativa (japonica cultiva... 369 e-100
ref|XP_468366.1| putative LEUNIG [Oryza sativa (japonica cultiva... 365 2e-99
emb|CAF18246.1| STY-L protein [Antirrhinum majus] 365 2e-99
ref|NP_850195.1| WD-40 repeat family protein [Arabidopsis thaliana] 343 8e-93
gb|AAC04493.1| expressed protein [Arabidopsis thaliana] gi|13605... 341 5e-92
ref|NP_567896.1| WD-40 repeat family protein (LEUNIG) [Arabidops... 327 8e-88
gb|AAG32022.1| LEUNIG [Arabidopsis thaliana] 327 8e-88
emb|CAF18245.1| STYLOSA protein [Antirrhinum majus] 326 1e-87
ref|XP_550318.1| putative transcriptional corepressor LEUNIG [Or... 315 4e-84
ref|XP_550319.1| putative transcriptional corepressor LEUNIG [Or... 315 4e-84
emb|CAB79972.1| putative protein [Arabidopsis thaliana] gi|49144... 306 1e-81
dbj|BAD53055.1| LEUNIG-like [Oryza sativa (japonica cultivar-gro... 299 2e-79
dbj|BAD53054.1| putative LEUNIG [Oryza sativa (japonica cultivar... 299 2e-79
gb|AAP44756.1| putative WD repeat protein [Oryza sativa (japonic... 297 6e-79
ref|NP_917595.1| putative flower development regulator, LEUNI pr... 269 2e-70
ref|NP_916506.1| P0013F10.18 [Oryza sativa (japonica cultivar-gr... 244 6e-63
gb|AAO22525.1| leunig [Brassica rapa subsp. pekinensis] 225 4e-57
dbj|BAD53056.1| putative LEUNIG [Oryza sativa (japonica cultivar... 219 2e-55
ref|NP_917597.1| putative flower development regulator, LEUNI pr... 186 3e-45
ref|ZP_00110817.1| COG2319: FOG: WD40 repeat [Nostoc punctiforme... 139 3e-31
>emb|CAE03368.2| OSJNBb0065L13.11 [Oryza sativa (japonica cultivar-group)]
gi|50926376|ref|XP_473135.1| OSJNBb0065L13.11 [Oryza
sativa (japonica cultivar-group)]
Length = 793
Score = 369 bits (946), Expect = e-100
Identities = 246/770 (31%), Positives = 367/770 (46%), Gaps = 137/770 (17%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPESSCKVPMNMMPAGNSNNASPHRIPQIPM 67
+D+P GFL++WWS+F+++F +R H + + ++ Q+ +
Sbjct: 45 IDAPGGFLFEWWSVFWDIFIARTNEKH----SEIAAAYLEAQQTKAREHQQQMQMQQLQL 100
Query: 68 NEHRPQQFQANSGFNNMIPQPAARLIPQAVFDNGQLGNLAENV-----------EPSLHD 116
+ R Q Q + + + P + L + LA + +
Sbjct: 101 IQQRHAQLQRTNATHPSLNGPISGLNSDGILGPSTASVLAAKMYEERLKHSHPMDSDGSQ 160
Query: 117 LLRENNQKLLSGTNSNHLPHPQDVGKQVQKYVFKVVTQFTSIYTN--ENHKLSGSGVTMR 174
LL + LL ++NH G+ V V T I +N + G
Sbjct: 161 LLDASRLALLKSASTNHS------GQSVPGTPGSVSTTLQQIQARNQQNIDIKSEGNMSV 214
Query: 175 VGRDVPRDPPDVV-QKTMLPLDGPHQTNNNEALNLVP------------QPLNGGPVNTP 221
R +P DP + Q + P G N+ ++ +P +P GG + P
Sbjct: 215 AQRSMPMDPSSLYGQGIIQPKPGLGGGVLNQGVSGLPLKGWPLTGIDQLRPNLGGQMQKP 274
Query: 222 --NSHQQCQVLKTQNQD--------------------------AALTQA--------PAC 245
++ Q Q++ Q Q +ALT++ PA
Sbjct: 275 FLSTQSQFQLMSPQQQQQFLAQAQAQGNLGNSTNLGDMDPRRLSALTRSVLNGKDGQPAG 334
Query: 246 TPNRQTSTVPGTKRKNKHPK--IPKTGSSKNDSQKMEQIISTGEHQCNQDQQIQMQSQNV 303
T TS + + K + + + KT S + Q +Q H Q QQ Q ++
Sbjct: 335 TDGCITSPMQSSSPKVRPDQEYLMKTSSQQTQEQLQQQ------HNQQQQQQTQQGNRKR 388
Query: 304 EKSTKRMITESTRPVESAQDCADAKPA--------------------------------- 330
++ T ST + ++ P+
Sbjct: 389 KQPTSSGAANSTGTANTVGPSTNSPPSTPSTHTPGDGLGMTGNMRHVPKNLMMYGVEGTG 448
Query: 331 -------------------DENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFS 371
D+NVESFL+ D A + F+ ++ A + +KGF+
Sbjct: 449 LPSSSNLDDLEQFGDMGSLDDNVESFLANGDGDAR---DIFAAPEKSPAEPNPVASKGFT 505
Query: 372 LQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVR 431
E+ C + +KV+ CHFSSDGK++ASAGHEKK +WNME FQ +A+ H+ +ITDVR
Sbjct: 506 FSEVNCWRTNNSKVVCCHFSSDGKILASAGHEKKAVLWNMETFQTQYTAEEHAVIITDVR 565
Query: 432 FQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDV 491
F+P S ATSSFDR+++LW+A++P SL GH + SLDFHP++ DLLCS DSN
Sbjct: 566 FRPNSNQLATSSFDRTIKLWNAADPGFSLHTFAGHCSGITSLDFHPKKTDLLCSCDSNGE 625
Query: 492 IRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKD 551
IR WNV+Q CM +GG+ QVRFQP GQ LA A N ++I DVET+ + L+GH+ +
Sbjct: 626 IRYWNVSQLSCMRAMKGGTAQVRFQPNTGQFLAAATENVVSIFDVETNGKKYTLQGHNSE 685
Query: 552 VLSICWDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGG 611
V S+CWD SG +LASVS+D ++WS +S G C E+ S GNKF SC+FHPGY LL+IGG
Sbjct: 686 VQSVCWDSSGQYLASVSQDLVKVWSISS-GDCTHEVSSNGNKFHSCVFHPGYTDLLVIGG 744
Query: 612 YQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
YQSLE W+ + +++ ++ AH+GLIA LA SP+ ++ASASHDNSVK+WK
Sbjct: 745 YQSLELWNMVK-NQSMTVQAHEGLIAALAQSPITGMVASASHDNSVKLWK 793
>ref|XP_468366.1| putative LEUNIG [Oryza sativa (japonica cultivar-group)]
gi|47848544|dbj|BAD22396.1| putative LEUNIG [Oryza
sativa (japonica cultivar-group)]
gi|47847864|dbj|BAD21657.1| putative LEUNIG [Oryza
sativa (japonica cultivar-group)]
Length = 802
Score = 365 bits (938), Expect = 2e-99
Identities = 179/351 (50%), Positives = 243/351 (68%), Gaps = 5/351 (1%)
Query: 311 ITESTRPVESAQDCADAKPADENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGF 370
+ S+ ++ + D ++NVESFL+ +D A + F+ LKR A + +KGF
Sbjct: 457 LASSSNQMDDLEPFGDVGSLEDNVESFLANDDGDAR---DIFAALKRSPAEPNPAASKGF 513
Query: 371 SLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDV 430
+ E+ CL + +KV+ CHFSSDGK++ASAGHEKK +WNM+ FQ +++ HS +ITDV
Sbjct: 514 TFNEVNCLRTNNSKVVCCHFSSDGKILASAGHEKKAVLWNMDTFQSQYTSEEHSLIITDV 573
Query: 431 RFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSND 490
RF+P S+ ATSSFDR+++LW+A++P L GH+ QV SLDFHP++ DLLCS D N
Sbjct: 574 RFRPNSSQLATSSFDRTIKLWNAADPGFCLHTFVGHNVQVTSLDFHPKKTDLLCSCDGNG 633
Query: 491 VIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDK 550
IR WN+ Q CM +GG+ QVRFQP GQ LA A + I DVET S + L+GH+
Sbjct: 634 EIRYWNLTQLSCMRAMKGGTAQVRFQPNTGQFLAAAAETMVAIFDVETHSKKYTLQGHNT 693
Query: 551 DVLSICWDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIG 610
DV S+CWD SG +LASVS+D ++WS +S G+CI E+ S GNKF SC+FHP Y +LL+IG
Sbjct: 694 DVQSVCWDSSGEYLASVSQDLVKVWSISS-GECIHEVSSNGNKFHSCVFHPSYANLLVIG 752
Query: 611 GYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
GYQSLE W+ + +++ +I AH+GLIA LA SPV +ASASHDNSVK+WK
Sbjct: 753 GYQSLELWNMVK-NQSMTIQAHEGLIAALAQSPVNGTVASASHDNSVKLWK 802
>emb|CAF18246.1| STY-L protein [Antirrhinum majus]
Length = 777
Score = 365 bits (938), Expect = 2e-99
Identities = 251/756 (33%), Positives = 360/756 (47%), Gaps = 125/756 (16%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGP------------ESSCKVPMNMMPAGNSN 55
+D+P GFL++WWS+F+++F +R H E ++ M +
Sbjct: 45 IDAPGGFLFEWWSVFWDIFIARTNDKHSEAAAAYIETQQIKAREQQQQMQMQQLQLLQQR 104
Query: 56 NASPHRIPQIPMNEHRPQQFQANS-GFNNMIPQPAARLIPQAVFDNGQLGNLAENVEPSL 114
NA R H P NS MI QP+A ++ +++ + + E S
Sbjct: 105 NAQLQRRDP----NHPPLGGPMNSMNSEGMIGQPSASVLAMKMYEERMKHPHSMDSETS- 159
Query: 115 HDLLRENNQKLLSGTNSNHLPHPQDVGKQVQKYVFKVVTQFTSIYTNENHKLSGSGVTMR 174
L+ N LL ++ +Q Q + ++ + + +
Sbjct: 160 PGLIDANRMALLKSASN----------QQGQLMQGNTGSMSAALQQMQGRPQMANDIKGE 209
Query: 175 VG-----RDVPRDPPDVVQKTMLP----LDGPHQTNNNEALNLVPQPLNGGP-------- 217
VG + +P DP + + +L L G L L PL G
Sbjct: 210 VGLGSTQKSLPMDPSSIYGQAILQSKSGLGGAGLNQGVTGLPLKGWPLTGIDQLRPSLGL 269
Query: 218 -VNTPNSHQQCQVLKTQNQDAALTQAPAC-----TPNRQTSTVPGTKRKNKHPKIPKTGS 271
V PN Q Q L Q L QA A +PN +P K + P+
Sbjct: 270 QVQKPNLQTQNQFLLASQQQQVLAQAQAQGSLGNSPNYGYGGLPRGNSNAKDGQPPRNDG 329
Query: 272 S----------KNDSQKMEQIISTGEHQCNQDQQIQMQS--------------------- 300
S K +M+Q S + Q Q QQ Q+
Sbjct: 330 SICSPVQANSPKMKMAQMQQASSQQQDQLQQQQQQLQQNNRKRKTHSSSGPANSTGTGNT 389
Query: 301 ------------------------QNVEKSTKRM----------ITESTRPVESAQDCAD 326
Q+V +K M I ST ++ ++ D
Sbjct: 390 VGPSPGTPQSTHTPGDGMATASSLQHVNSVSKSMMMYGADAAGGIASSTNQLDDLENFGD 449
Query: 327 AKPADENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVL 386
++NVESFLS + D + + +LK+ +KGFS E+GC+ ++ NKV
Sbjct: 450 VGSLEDNVESFLSHD---GDGNL--YGSLKQTLTEHKTETSKGFSFGEVGCI-RTRNKVT 503
Query: 387 TCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDR 446
CHFSSDGK++ASAGH+KK +WNM+ Q + + H +LITDVRF+P ST AT+SFD+
Sbjct: 504 CCHFSSDGKLLASAGHDKKAVLWNMDTLQTETTPEEHQYLITDVRFRPNSTQLATASFDK 563
Query: 447 SVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTT 506
SVRLWDA+NP L TGH VMSLDFHP++ DL C DSN+ IR W+++ C +
Sbjct: 564 SVRLWDAANPSYCLNAYTGHASHVMSLDFHPKKNDLFCFCDSNNEIRYWSISPFACTRIS 623
Query: 507 R-GGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLA 565
+ GGS QVRFQP G LLA A ++I DVE D H +GH V +CWD +G+ LA
Sbjct: 624 KQGGSAQVRFQPITGHLLAAASDKVVSIYDVENDRQTHSFQGHSGVVNYLCWDLNGDLLA 683
Query: 566 SVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
SVSEDS ++WS AS G+CI EL+S GN+F SC+FHP Y +LL+IGG +SLE W+ E +K
Sbjct: 684 SVSEDSIKVWSLAS-GECIHELNSNGNQFHSCVFHPSYSALLVIGGLRSLELWNMVE-NK 741
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+ +++AH+ +IA LA SP+ ++ASASHD+SVK+WK
Sbjct: 742 SMTVSAHENIIAALAQSPLTGMVASASHDSSVKLWK 777
>ref|NP_850195.1| WD-40 repeat family protein [Arabidopsis thaliana]
Length = 785
Score = 343 bits (881), Expect = 8e-93
Identities = 205/542 (37%), Positives = 301/542 (54%), Gaps = 42/542 (7%)
Query: 141 GKQVQKYVFKVVTQFTSIYTNENHKLSG--------SGVTMRVGRDVPRDPPDVVQKTML 192
G QVQK + +QF + H++ + M G PR + + +
Sbjct: 265 GPQVQKSFLQNQSQFQLSPQQQQHQMLAQVQAQGNMTNSPMYGGDMDPRRFTGLPRGNLN 324
Query: 193 PLDGPHQTNNNEALNLVPQPLNGG--------PVNTPNSHQQCQVLKTQNQ-DAALTQAP 243
P DG N+ + P+ PV +S QQ +L Q+Q + + P
Sbjct: 325 PKDGQQNANDGS----IGSPMQSSSSKHISMPPVQQSSSQQQDHLLSQQSQQNNRKRKGP 380
Query: 244 ACT-PNRQTSTVPGTKRKNKHPKIPKTGSSKNDSQKMEQIISTGEHQCNQDQQIQMQ--S 300
+ + P T T N P P T + + I+ H N + M S
Sbjct: 381 SSSGPANSTGTGNTVGPSNSQPSTPSTHTPVDGVA-----IAGNMHHVNSMPKGPMMYGS 435
Query: 301 QNVEKSTKRMITESTRPVESAQDCADAKPADENVESFLSFEDEPADNRIEPFSNLKRISA 360
+ + S ++ D ++NVESFLS +D + F LKR S+
Sbjct: 436 DGIGG-----LASSANQLDDMDQFGDVGALEDNVESFLSQDDGDGGSL---FGTLKRNSS 487
Query: 361 TCSRNENKGFSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASA 420
+ +K FS E+ C+ KS +KV+ C FS DGK++ASAGH+KKVFIWNME Q ++
Sbjct: 488 VHTET-SKPFSFNEVSCIRKSASKVICCSFSYDGKLLASAGHDKKVFIWNMETLQVESTP 546
Query: 421 DTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRM 480
+ H+H+ITDVRF+P ST ATSSFD+++++WDAS+P L ++GH VMS+DFHP++
Sbjct: 547 EEHAHIITDVRFRPNSTQLATSSFDKTIKIWDASDPGYFLRTISGHAAPVMSIDFHPKKT 606
Query: 481 DLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDS 540
+LLCS DSN+ IR W++N S C+ +G S QVRFQP+ GQ LA A N+++I D+E ++
Sbjct: 607 ELLCSCDSNNDIRFWDINAS-CVRAVKGASTQVRFQPRTGQFLAAASENTVSIFDIENNN 665
Query: 541 -HLHDLKGHDKDVLSICWDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIF 599
++ KGH +V S+CW +G +ASVSED+ ++WS +S G CI EL + GNKF S +F
Sbjct: 666 KRVNIFKGHSSNVHSVCWSPNGELVASVSEDAVKLWSLSS-GDCIHELSNSGNKFHSVVF 724
Query: 600 HPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKV 659
HP Y LL+IGGYQ++E W+ E +K ++A H+ +I+ LA SP ++ASASHD SVK+
Sbjct: 725 HPSYPDLLVIGGYQAIELWNTME-NKCMTVAGHECVISALAQSPSTGVVASASHDKSVKI 783
Query: 660 WK 661
WK
Sbjct: 784 WK 785
>gb|AAC04493.1| expressed protein [Arabidopsis thaliana] gi|13605815|gb|AAK32893.1|
At2g32700/F24L7.16 [Arabidopsis thaliana]
gi|25090107|gb|AAN72230.1| At2g32700/F24L7.16
[Arabidopsis thaliana] gi|7486097|pir||T00798
hypothetical protein At2g32700 [imported] - Arabidopsis
thaliana gi|30685392|ref|NP_850192.1| WD-40 repeat
family protein [Arabidopsis thaliana]
gi|30685401|ref|NP_850194.1| WD-40 repeat family protein
[Arabidopsis thaliana] gi|30685398|ref|NP_850193.1|
WD-40 repeat family protein [Arabidopsis thaliana]
gi|18403052|ref|NP_565749.1| WD-40 repeat family protein
[Arabidopsis thaliana]
Length = 787
Score = 341 bits (874), Expect = 5e-92
Identities = 204/541 (37%), Positives = 303/541 (55%), Gaps = 38/541 (7%)
Query: 141 GKQVQKYVFKVVTQFTSIYTNENHKLSG--------SGVTMRVGRDVPRDPPDVVQKTML 192
G QVQK + +QF + H++ + M G PR + + +
Sbjct: 265 GPQVQKSFLQNQSQFQLSPQQQQHQMLAQVQAQGNMTNSPMYGGDMDPRRFTGLPRGNLN 324
Query: 193 PLDGPHQTNNNEALNLVPQPLNGG--------PVNTPNSHQQCQVLKTQNQ-DAALTQAP 243
P DG N+ + P+ PV +S QQ +L Q+Q + + P
Sbjct: 325 PKDGQQNANDGS----IGSPMQSSSSKHISMPPVQQSSSQQQDHLLSQQSQQNNRKRKGP 380
Query: 244 ACT-PNRQTSTVPGTKRKNKHPKIPKTGSSKNDSQKMEQIISTGEHQCNQDQQIQMQSQN 302
+ + P T T N P P T + + I+ H N + M +
Sbjct: 381 SSSGPANSTGTGNTVGPSNSQPSTPSTHTPVDGVA-----IAGNMHHVNSMPKGPMMYGS 435
Query: 303 VEKSTKRMITESTRPVESAQD-CADAKPADENVESFLSFEDEPADNRIEPFSNLKRISAT 361
+ + + + ++ D D ++NVESFLS +D + F LKR S+
Sbjct: 436 --DGIGGLASSANQLLQDDMDQFGDVGALEDNVESFLSQDDGDGGSL---FGTLKRNSSV 490
Query: 362 CSRNENKGFSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASAD 421
+ +K FS E+ C+ KS +KV+ C FS DGK++ASAGH+KKVFIWNME Q ++ +
Sbjct: 491 HTET-SKPFSFNEVSCIRKSASKVICCSFSYDGKLLASAGHDKKVFIWNMETLQVESTPE 549
Query: 422 THSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMD 481
H+H+ITDVRF+P ST ATSSFD+++++WDAS+P L ++GH VMS+DFHP++ +
Sbjct: 550 EHAHIITDVRFRPNSTQLATSSFDKTIKIWDASDPGYFLRTISGHAAPVMSIDFHPKKTE 609
Query: 482 LLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDS- 540
LLCS DSN+ IR W++N S C+ +G S QVRFQP+ GQ LA A N+++I D+E ++
Sbjct: 610 LLCSCDSNNDIRFWDINAS-CVRAVKGASTQVRFQPRTGQFLAAASENTVSIFDIENNNK 668
Query: 541 HLHDLKGHDKDVLSICWDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFH 600
++ KGH +V S+CW +G +ASVSED+ ++WS +S G CI EL + GNKF S +FH
Sbjct: 669 RVNIFKGHSSNVHSVCWSPNGELVASVSEDAVKLWSLSS-GDCIHELSNSGNKFHSVVFH 727
Query: 601 PGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
P Y LL+IGGYQ++E W+ E +K ++A H+ +I+ LA SP ++ASASHD SVK+W
Sbjct: 728 PSYPDLLVIGGYQAIELWNTME-NKCMTVAGHECVISALAQSPSTGVVASASHDKSVKIW 786
Query: 661 K 661
K
Sbjct: 787 K 787
>ref|NP_567896.1| WD-40 repeat family protein (LEUNIG) [Arabidopsis thaliana]
gi|30580400|sp|Q9FUY2|LEUNG_ARATH Transcriptional
corepressor LEUNIG
Length = 931
Score = 327 bits (838), Expect = 8e-88
Identities = 187/498 (37%), Positives = 265/498 (52%), Gaps = 37/498 (7%)
Query: 176 GRDVPRDPPDVVQKTMLPLDGPHQTNNNEALNLVPQPLNGGPVNTPNSHQQC-----QVL 230
G + P+ P L L P ++N +++ + GG + S QVL
Sbjct: 459 GGNPPQPQPQPQPLNQLALTNPQPQSSNHSIHQQEKLGGGGSITMDGSISNSFRGNEQVL 518
Query: 231 KTQNQDAALTQAPACTPNRQTSTVPGTKRKNKHPKIPKTGSSKNDSQKMEQIISTGE--H 288
K Q+ + P + T N P + S + +IS H
Sbjct: 519 KNQSGRKRKQPVSSSGPANSSGTA------NTAGPSPSSAPSTPSTHTPGDVISMPNLPH 572
Query: 289 QCNQDQQIQM---QSQNVEKSTKRMITESTRPVESAQDCADAKPADENVESFLSFEDEPA 345
+ + M + S + + R VE D+NVESFLS ED
Sbjct: 573 SGGSSKSMMMFGTEGTGTLTSPSNQLADMDRFVEDGS-------LDDNVESFLSQEDGD- 624
Query: 346 DNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKK 405
+R + T + +KGF+ E+ + S KV CHFSSDGK++ASAGH+KK
Sbjct: 625 ----------QRDAVTRCMDVSKGFTFTEVNSVRASTTKVTCCHFSSDGKMLASAGHDKK 674
Query: 406 VFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTG 465
+W + + + + H+ +ITD+RF P ATSSFD++VR+WDA N SL G
Sbjct: 675 AVLWYTDTMKPKTTLEEHTAMITDIRFSPSQLRLATSSFDKTVRVWDADNKGYSLRTFMG 734
Query: 466 HDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLAT 525
H V SLDFHP + DL+CS D+++ IR W++N C +GGS Q+RFQP+ G+ LA
Sbjct: 735 HSSMVTSLDFHPIKDDLICSCDNDNEIRYWSINNGSCTRVYKGGSTQIRFQPRVGKYLAA 794
Query: 526 AIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSARIWS--AASDGKC 583
+ N + ++DVET + H L+GH + S+CWD SG+FLASVSED ++W+ S+G+C
Sbjct: 795 SSANLVNVLDVETQAIRHSLQGHANPINSVCWDPSGDFLASVSEDMVKVWTLGTGSEGEC 854
Query: 584 IGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSP 643
+ EL GNKFQSC+FHP Y SLL+IG YQSLE W+ +E +KT ++ AH+GLI LA S
Sbjct: 855 VHELSCNGNKFQSCVFHPAYPSLLVIGCYQSLELWNMSE-NKTMTLPAHEGLITSLAVST 913
Query: 644 VGELIASASHDNSVKVWK 661
L+ASASHD VK+WK
Sbjct: 914 ATGLVASASHDKLVKLWK 931
Score = 43.1 bits (100), Expect = 0.028
Identities = 19/69 (27%), Positives = 33/69 (47%), Gaps = 3/69 (4%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPESSCKVPMNMMPAGNSNNASPHRIPQIPM 67
+D+P GFL++WWS+F+++F +R H E + M + PQ+
Sbjct: 45 IDAPGGFLFEWWSVFWDIFIARTNEKH---SEVAASYIETQMIKAREQQLQQSQHPQVSQ 101
Query: 68 NEHRPQQFQ 76
+ + QQ Q
Sbjct: 102 QQQQQQQQQ 110
>gb|AAG32022.1| LEUNIG [Arabidopsis thaliana]
Length = 931
Score = 327 bits (838), Expect = 8e-88
Identities = 187/498 (37%), Positives = 265/498 (52%), Gaps = 37/498 (7%)
Query: 176 GRDVPRDPPDVVQKTMLPLDGPHQTNNNEALNLVPQPLNGGPVNTPNSHQQC-----QVL 230
G + P+ P L L P ++N +++ + GG + S QVL
Sbjct: 459 GGNPPQPQPQPQPLNQLALTNPQPQSSNHSIHQQEKLGGGGSITMDGSISNSFRGNEQVL 518
Query: 231 KTQNQDAALTQAPACTPNRQTSTVPGTKRKNKHPKIPKTGSSKNDSQKMEQIISTGE--H 288
K Q+ + P + T N P + S + +IS H
Sbjct: 519 KNQSGRKRKQPVSSSGPANSSGTA------NTAGPSPSSAPSTPSTHTPGDVISMPNLPH 572
Query: 289 QCNQDQQIQM---QSQNVEKSTKRMITESTRPVESAQDCADAKPADENVESFLSFEDEPA 345
+ + M + S + + R VE D+NVESFLS ED
Sbjct: 573 SGGSSKSMMMFGTEGTGTLTSPSNQLADMDRFVEDGS-------LDDNVESFLSQEDGD- 624
Query: 346 DNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKK 405
+R + T + +KGF+ E+ + S KV CHFSSDGK++ASAGH+KK
Sbjct: 625 ----------QRDAVTRCMDVSKGFTFTEVNSVRASTTKVTCCHFSSDGKMLASAGHDKK 674
Query: 406 VFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTG 465
+W + + + + H+ +ITD+RF P ATSSFD++VR+WDA N SL G
Sbjct: 675 AVLWYTDTMKPKTTLEEHTAMITDIRFSPSQLRLATSSFDKTVRVWDADNKGYSLRTFMG 734
Query: 466 HDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLAT 525
H V SLDFHP + DL+CS D+++ IR W++N C +GGS Q+RFQP+ G+ LA
Sbjct: 735 HSSMVTSLDFHPIKDDLICSCDNDNEIRYWSINNGSCTRVYKGGSTQIRFQPRVGKYLAA 794
Query: 526 AIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSARIWS--AASDGKC 583
+ N + ++DVET + H L+GH + S+CWD SG+FLASVSED ++W+ S+G+C
Sbjct: 795 SSANLVNVLDVETQAIRHSLQGHANPINSVCWDPSGDFLASVSEDMVKVWTLGTGSEGEC 854
Query: 584 IGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSP 643
+ EL GNKFQSC+FHP Y SLL+IG YQSLE W+ +E +KT ++ AH+GLI LA S
Sbjct: 855 VHELSCNGNKFQSCVFHPAYPSLLVIGCYQSLELWNMSE-NKTMTLPAHEGLITSLAVST 913
Query: 644 VGELIASASHDNSVKVWK 661
L+ASASHD VK+WK
Sbjct: 914 ATGLVASASHDKLVKLWK 931
Score = 43.1 bits (100), Expect = 0.028
Identities = 19/69 (27%), Positives = 33/69 (47%), Gaps = 3/69 (4%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPESSCKVPMNMMPAGNSNNASPHRIPQIPM 67
+D+P GFL++WWS+F+++F +R H E + M + PQ+
Sbjct: 45 IDAPGGFLFEWWSVFWDIFIARTNEKH---SEVAASYIETQMIKAREQQLQQSQHPQVSQ 101
Query: 68 NEHRPQQFQ 76
+ + QQ Q
Sbjct: 102 QQQQQQQQQ 110
>emb|CAF18245.1| STYLOSA protein [Antirrhinum majus]
Length = 915
Score = 326 bits (836), Expect = 1e-87
Identities = 198/511 (38%), Positives = 277/511 (53%), Gaps = 42/511 (8%)
Query: 179 VPRDPPDVVQK---TMLPLDGPHQTNNNEALNLVPQPLNGG-PVNTPNSHQQCQVLKTQN 234
+PR P+++ K L Q N+N+ L+G P ++ ++ QQ +++ T +
Sbjct: 419 LPRADPEMLMKLKIAQLQQQQQQQQNSNQTQQQQHHTLSGQQPQSSNHNLQQDKMMGTSS 478
Query: 235 QDAALTQAPACTPNRQTSTVPGTKRKNKHP-----KIPKTGSSKNDSQKMEQIISTGEHQ 289
+ + + N Q S T RK K P +G++ ST
Sbjct: 479 AAGEGSMSNSFRGNDQASKNQ-TGRKRKQPVSSSGPANSSGTANTAGPSPSSAPSTPSTH 537
Query: 290 CNQDQQIQMQSQNVEKSTKRMI-------TESTRPVESAQDCADAKPAD---------EN 333
D + S+K ++ T P D D PAD +N
Sbjct: 538 TPGDVMSMPALPHSGSSSKPLMMFGADNNATLTSPSNQLWDDKDLVPADMDRFVDDVEDN 597
Query: 334 VESFLSFED-EPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHFSS 392
VESFLS +D +P D + + +KGF+ E+ + S +KV+ CHFS
Sbjct: 598 VESFLSNDDADPRD------------AVGRCMDVSKGFTFTEVSYVRASASKVVCCHFSP 645
Query: 393 DGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWD 452
DGK++AS GH+KK +W + + + + HS LITDVRF P ATSSFD++VR+WD
Sbjct: 646 DGKLLASGGHDKKAVLWYTDTLKPKTTLEEHSSLITDVRFSPSMARLATSSFDKTVRVWD 705
Query: 453 ASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQ 512
A NP S+ TGH VMSLDFHP + DL+CS D + IR W++ C +GG+ Q
Sbjct: 706 ADNPGYSIRTFTGHSAGVMSLDFHPVKEDLICSCDGDGEIRYWSIKNGSCARVFKGGTAQ 765
Query: 513 VRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSA 572
VRFQP+ G+ LA A N ++I+D ET + H LKGH K + S+CWD SG LASVSEDS
Sbjct: 766 VRFQPRLGRYLAAAAENVVSILDSETLACRHSLKGHTKPIHSVCWDPSGELLASVSEDSV 825
Query: 573 RIWS--AASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIA 630
R+W+ + S+G C+ EL GNKF SC+FHP Y SLL+IG YQSLE W+ +E +KT +++
Sbjct: 826 RVWTLRSGSEGDCLHELSCNGNKFHSCVFHPTYSSLLVIGCYQSLELWNMSE-NKTMTLS 884
Query: 631 AHQGLIAGLADSPVGELIASASHDNSVKVWK 661
AH+GLIA LA S L+ASASHD VK+WK
Sbjct: 885 AHEGLIASLAVSTGAGLVASASHDKIVKLWK 915
Score = 42.7 bits (99), Expect = 0.037
Identities = 22/84 (26%), Positives = 38/84 (45%), Gaps = 8/84 (9%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPESSCKVPMNMMPAGNSNNASPHRIPQIPM 67
+D+P GFL++WWS+F+++F +R H S + + A + Q P
Sbjct: 45 IDAPGGFLFEWWSVFWDIFIARTNEKHSEVAASYIETQL--------MKAREQQQQQQPQ 96
Query: 68 NEHRPQQFQANSGFNNMIPQPAAR 91
+PQQ Q ++ Q A+
Sbjct: 97 QSQQPQQQQQQLHMQQLLLQRQAQ 120
>ref|XP_550318.1| putative transcriptional corepressor LEUNIG [Oryza sativa (japonica
cultivar-group)] gi|55295951|dbj|BAD67819.1| putative
transcriptional corepressor LEUNIG [Oryza sativa
(japonica cultivar-group)]
Length = 877
Score = 315 bits (806), Expect = 4e-84
Identities = 161/333 (48%), Positives = 220/333 (65%), Gaps = 17/333 (5%)
Query: 331 DENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHF 390
+++V+SFLS +D AD R + R+ +T KGF +E+ + S NKV+ CHF
Sbjct: 560 EDHVDSFLSHDD--ADRR-----DGSRMEST------KGFIFREVSSVQASTNKVVCCHF 606
Query: 391 SSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRL 450
SSDGK++A+ GH+KKV +W+ E + + + HS LITDVRF P ATSSFD++VR+
Sbjct: 607 SSDGKLLATGGHDKKVVLWHAETLKQKSVLEEHSLLITDVRFSPSIPRLATSSFDKTVRV 666
Query: 451 WDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGS 510
WDA N S+ TGH VMSLDFHP + DL+CS D ++ IR W++N + +GGS
Sbjct: 667 WDADNQGYSIRTFTGHSASVMSLDFHPNKDDLICSCDGDNEIRFWSINNGNIVRIFKGGS 726
Query: 511 KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED 570
Q+RFQP++G LA A N+++I+DVET + L +GH K V S+CWD SG ++ SVSED
Sbjct: 727 SQLRFQPRHGGYLAVASENAVSILDVETQACLRRFEGHTKHVDSVCWDPSGEYVVSVSED 786
Query: 571 SARIWS--AASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWS 628
+ ++WS A SD +C+ EL G+KF SC FHP Y S+LIIG YQSLE W +E ++T +
Sbjct: 787 TVKVWSVNAGSDDRCVQELSCTGSKFHSCAFHPSYSSMLIIGCYQSLELWDMSE-NRTMT 845
Query: 629 IAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+AAH LI LA S G L+AS SHD VK+WK
Sbjct: 846 LAAHDSLITALASSSSG-LVASTSHDKFVKLWK 877
Score = 39.7 bits (91), Expect = 0.31
Identities = 20/83 (24%), Positives = 37/83 (44%), Gaps = 11/83 (13%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPESSCKV-----------PMNMMPAGNSNN 56
+D+P GFL +WWS+F+++F +R H S + + A + +
Sbjct: 50 IDAPGGFLLEWWSVFWDIFIARTNEKHSDVAASYIETQSIKAREQQPSQLQQQEAHSQQS 109
Query: 57 ASPHRIPQIPMNEHRPQQFQANS 79
+ ++ Q+ + H QQ Q S
Sbjct: 110 SQQIQMQQLLLQRHAQQQQQQQS 132
>ref|XP_550319.1| putative transcriptional corepressor LEUNIG [Oryza sativa (japonica
cultivar-group)] gi|55295950|dbj|BAD67818.1| putative
transcriptional corepressor LEUNIG [Oryza sativa
(japonica cultivar-group)]
Length = 875
Score = 315 bits (806), Expect = 4e-84
Identities = 161/333 (48%), Positives = 220/333 (65%), Gaps = 17/333 (5%)
Query: 331 DENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHF 390
+++V+SFLS +D AD R + R+ +T KGF +E+ + S NKV+ CHF
Sbjct: 558 EDHVDSFLSHDD--ADRR-----DGSRMEST------KGFIFREVSSVQASTNKVVCCHF 604
Query: 391 SSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRL 450
SSDGK++A+ GH+KKV +W+ E + + + HS LITDVRF P ATSSFD++VR+
Sbjct: 605 SSDGKLLATGGHDKKVVLWHAETLKQKSVLEEHSLLITDVRFSPSIPRLATSSFDKTVRV 664
Query: 451 WDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGS 510
WDA N S+ TGH VMSLDFHP + DL+CS D ++ IR W++N + +GGS
Sbjct: 665 WDADNQGYSIRTFTGHSASVMSLDFHPNKDDLICSCDGDNEIRFWSINNGNIVRIFKGGS 724
Query: 511 KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED 570
Q+RFQP++G LA A N+++I+DVET + L +GH K V S+CWD SG ++ SVSED
Sbjct: 725 SQLRFQPRHGGYLAVASENAVSILDVETQACLRRFEGHTKHVDSVCWDPSGEYVVSVSED 784
Query: 571 SARIWS--AASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWS 628
+ ++WS A SD +C+ EL G+KF SC FHP Y S+LIIG YQSLE W +E ++T +
Sbjct: 785 TVKVWSVNAGSDDRCVQELSCTGSKFHSCAFHPSYSSMLIIGCYQSLELWDMSE-NRTMT 843
Query: 629 IAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+AAH LI LA S G L+AS SHD VK+WK
Sbjct: 844 LAAHDSLITALASSSSG-LVASTSHDKFVKLWK 875
Score = 40.0 bits (92), Expect = 0.24
Identities = 22/81 (27%), Positives = 38/81 (46%), Gaps = 9/81 (11%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPES---SCKV------PMNMMPAGNSNNAS 58
+D+P GFL +WWS+F+++F +R H S S K + A + ++
Sbjct: 50 IDAPGGFLLEWWSVFWDIFIARTNEKHSDVAASYIESIKAREQQPSQLQQQEAHSQQSSQ 109
Query: 59 PHRIPQIPMNEHRPQQFQANS 79
++ Q+ + H QQ Q S
Sbjct: 110 QIQMQQLLLQRHAQQQQQQQS 130
>emb|CAB79972.1| putative protein [Arabidopsis thaliana] gi|4914452|emb|CAB43692.1|
putative protein [Arabidopsis thaliana]
gi|7486768|pir||T08588 hypothetical protein L23H3.30 -
Arabidopsis thaliana
Length = 930
Score = 306 bits (784), Expect = 1e-81
Identities = 181/498 (36%), Positives = 258/498 (51%), Gaps = 45/498 (9%)
Query: 176 GRDVPRDPPDVVQKTMLPLDGPHQTNNNEALNLVPQPLNGGPVNTPNSHQQC-----QVL 230
G + P+ P L L P ++N +++ + GG + S QVL
Sbjct: 466 GGNPPQPQPQPQPLNQLALTNPQPQSSNHSIHQQEKLGGGGSITMDGSISNSFRGNEQVL 525
Query: 231 KTQNQDAALTQAPACTPNRQTSTVPGTKRKNKHPKIPKTGSSKNDSQKMEQIISTGE--H 288
K Q+ + P + T N P + S + +IS H
Sbjct: 526 KNQSGRKRKQPVSSSGPANSSGTA------NTAGPSPSSAPSTPSTHTPGDVISMPNLPH 579
Query: 289 QCNQDQQIQM---QSQNVEKSTKRMITESTRPVESAQDCADAKPADENVESFLSFEDEPA 345
+ + M + S + + R VE D+NVESFLS ED
Sbjct: 580 SGGSSKSMMMFGTEGTGTLTSPSNQLADMDRFVEDGS-------LDDNVESFLSQEDGD- 631
Query: 346 DNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKK 405
+R + T + +KGF+ E+ + S KV CHFSSDGK++ASAGH+KK
Sbjct: 632 ----------QRDAVTRCMDVSKGFTFTEVNSVRASTTKVTCCHFSSDGKMLASAGHDKK 681
Query: 406 VFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTG 465
+W + + + + H+ +ITD+RF P ATSSFD++ SL G
Sbjct: 682 AVLWYTDTMKPKTTLEEHTAMITDIRFSPSQLRLATSSFDKTKGY--------SLRTFMG 733
Query: 466 HDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLAT 525
H V SLDFHP + DL+CS D+++ IR W++N C +GGS Q+RFQP+ G+ LA
Sbjct: 734 HSSMVTSLDFHPIKDDLICSCDNDNEIRYWSINNGSCTRVYKGGSTQIRFQPRVGKYLAA 793
Query: 526 AIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSARIWS--AASDGKC 583
+ N + ++DVET + H L+GH + S+CWD SG+FLASVSED ++W+ S+G+C
Sbjct: 794 SSANLVNVLDVETQAIRHSLQGHANPINSVCWDPSGDFLASVSEDMVKVWTLGTGSEGEC 853
Query: 584 IGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSP 643
+ EL GNKFQSC+FHP Y SLL+IG YQSLE W+ +E +KT ++ AH+GLI LA S
Sbjct: 854 VHELSCNGNKFQSCVFHPAYPSLLVIGCYQSLELWNMSE-NKTMTLPAHEGLITSLAVST 912
Query: 644 VGELIASASHDNSVKVWK 661
L+ASASHD VK+WK
Sbjct: 913 ATGLVASASHDKLVKLWK 930
Score = 43.1 bits (100), Expect = 0.028
Identities = 19/69 (27%), Positives = 33/69 (47%), Gaps = 3/69 (4%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPESSCKVPMNMMPAGNSNNASPHRIPQIPM 67
+D+P GFL++WWS+F+++F +R H E + M + PQ+
Sbjct: 52 IDAPGGFLFEWWSVFWDIFIARTNEKH---SEVAASYIETQMIKAREQQLQQSQHPQVSQ 108
Query: 68 NEHRPQQFQ 76
+ + QQ Q
Sbjct: 109 QQQQQQQQQ 117
>dbj|BAD53055.1| LEUNIG-like [Oryza sativa (japonica cultivar-group)]
gi|53792191|dbj|BAD52824.1| LEUNIG-like [Oryza sativa
(japonica cultivar-group)]
Length = 538
Score = 299 bits (766), Expect = 2e-79
Identities = 162/338 (47%), Positives = 213/338 (62%), Gaps = 18/338 (5%)
Query: 331 DENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHF 390
DENVESFLS +D ++P +L R S + +KGF E+ S KV CHF
Sbjct: 212 DENVESFLSQDD------MDPRDSLGR-----SMDASKGFGFAEVAKARASATKVTCCHF 260
Query: 391 SSDGKVMASAGHEKKVFIWNMEN-FQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
SSDGK++A+ GH+KKV +W E + +S + HS LITDVRF P + ATSSFD++VR
Sbjct: 261 SSDGKLLATGGHDKKVLLWCTEPALKPTSSLEEHSALITDVRFSPSMSRLATSSFDKTVR 320
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECM---HTT 506
+WDA N SL TGH VMSLDFHP + D++CS D + +R W++N C+
Sbjct: 321 VWDADNTSYSLRTFTGHSASVMSLDFHPNKEDMICSCDGDGEVRSWSINNGSCLTFVKVF 380
Query: 507 RGGSKQVRFQPQYGQLLATAIGNSITIIDVETD-SHLHDLKGHDKDVLSICWDRSGNFLA 565
+GG+ Q+RFQPQ G+ LA A +I I+D ET + + L+GH K++ S+CWD +G+ LA
Sbjct: 381 KGGATQMRFQPQKGKYLAAASEKAIYILDGETQLACRNPLQGHTKNIHSLCWDSTGDNLA 440
Query: 566 SVSEDSARIWSAA--SDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEG 623
SVSEDS RIWS A DG+ + EL GNKFQSC+FHP Y LL+IG Y+SLE W E
Sbjct: 441 SVSEDSVRIWSFAPGHDGEFVNELSCSGNKFQSCVFHPSYPYLLVIGCYESLELWDIREK 500
Query: 624 SKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+ +AH GL+A LA S +AS SHD VK+WK
Sbjct: 501 NAMTVHSAHDGLVAALAASSATGKVASVSHDRFVKLWK 538
>dbj|BAD53054.1| putative LEUNIG [Oryza sativa (japonica cultivar-group)]
gi|53792190|dbj|BAD52823.1| putative LEUNIG [Oryza
sativa (japonica cultivar-group)]
Length = 876
Score = 299 bits (766), Expect = 2e-79
Identities = 162/338 (47%), Positives = 213/338 (62%), Gaps = 18/338 (5%)
Query: 331 DENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHF 390
DENVESFLS +D ++P +L R S + +KGF E+ S KV CHF
Sbjct: 550 DENVESFLSQDD------MDPRDSLGR-----SMDASKGFGFAEVAKARASATKVTCCHF 598
Query: 391 SSDGKVMASAGHEKKVFIWNMEN-FQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
SSDGK++A+ GH+KKV +W E + +S + HS LITDVRF P + ATSSFD++VR
Sbjct: 599 SSDGKLLATGGHDKKVLLWCTEPALKPTSSLEEHSALITDVRFSPSMSRLATSSFDKTVR 658
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECM---HTT 506
+WDA N SL TGH VMSLDFHP + D++CS D + +R W++N C+
Sbjct: 659 VWDADNTSYSLRTFTGHSASVMSLDFHPNKEDMICSCDGDGEVRSWSINNGSCLTFVKVF 718
Query: 507 RGGSKQVRFQPQYGQLLATAIGNSITIIDVETD-SHLHDLKGHDKDVLSICWDRSGNFLA 565
+GG+ Q+RFQPQ G+ LA A +I I+D ET + + L+GH K++ S+CWD +G+ LA
Sbjct: 719 KGGATQMRFQPQKGKYLAAASEKAIYILDGETQLACRNPLQGHTKNIHSLCWDSTGDNLA 778
Query: 566 SVSEDSARIWSAA--SDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEG 623
SVSEDS RIWS A DG+ + EL GNKFQSC+FHP Y LL+IG Y+SLE W E
Sbjct: 779 SVSEDSVRIWSFAPGHDGEFVNELSCSGNKFQSCVFHPSYPYLLVIGCYESLELWDIREK 838
Query: 624 SKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+ +AH GL+A LA S +AS SHD VK+WK
Sbjct: 839 NAMTVHSAHDGLVAALAASSATGKVASVSHDRFVKLWK 876
Score = 42.7 bits (99), Expect = 0.037
Identities = 35/138 (25%), Positives = 58/138 (41%), Gaps = 10/138 (7%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPESSCKVPMNMMPAGNSNNASPHRIPQIPM 67
+D+P GFL++WWS+F+++F +R H ++ + M A + PQ
Sbjct: 45 IDAPGGFLFEWWSVFWDIFIARTNEKH--SDVAASYIETQHMKAREQQQQQQQQPPQ--Q 100
Query: 68 NEHRPQQFQANSGFNNMIPQPAARLIPQAVFDNGQLGNLAENVEPSLHDLLRENNQKLLS 127
+ +PQ Q M+ Q AA+ Q Q + + R + LL+
Sbjct: 101 RQQQPQHIQ----MQQMLLQRAAQQ-QQQQQQQQQQQQQQQQQQQQQQQQQRRDGTHLLN 155
Query: 128 GTNSNHLPHPQDVGKQVQ 145
GT S LP + +Q Q
Sbjct: 156 GTASG-LPGNNPLMRQNQ 172
>gb|AAP44756.1| putative WD repeat protein [Oryza sativa (japonica cultivar-group)]
gi|50920301|ref|XP_470511.1| putative WD repeat protein
[Oryza sativa (japonica cultivar-group)]
Length = 684
Score = 297 bits (761), Expect = 6e-79
Identities = 226/689 (32%), Positives = 348/689 (49%), Gaps = 84/689 (12%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPESSCKVPMNMMPAGNSNNASPHRIPQIPM 67
+D+P GFL++WWSIF+++F +R P P +P + R+ +
Sbjct: 45 IDAPGGFLFEWWSIFWDIFDARTR----DKPPQPQPQPPPPIPIDIKSREQQMRLQLLQQ 100
Query: 68 NEHRPQQFQ---ANSGFNNMIPQPAARLIPQAVFDNGQLGNLAENVEPSLHDLLRENNQK 124
++ QQ + N+ + + +A L + + D + N ++ + H LL N
Sbjct: 101 QQNAHQQRRDPPLNAAMDALNSDVSAVLASKMMQDRMRNPNPTDS--DASHQLLDANRIA 158
Query: 125 LLS-GTN---------SNHLPHPQDVGKQVQK--------------YVFKVVTQFTSIYT 160
LL TN S ++ Q + + Q+ Y ++T +I+
Sbjct: 159 LLKPATNQTGQLVQGASVNMSALQQIHSRNQQPVIPFHLLKLHWFMYRLSLLTLCPTIFK 218
Query: 161 NENHKLSGSGVTMRVGRDVPRDPPDVVQKTML-PLDGPHQTNNNEALNLVPQPLNGGPVN 219
+ + G + R +P DP + M+ P G T N+ + VP L G P+
Sbjct: 219 D----MKGDAAMSQ--RSMPTDPSTLYGSGMMQPKSGLVSTGLNQGVGSVP--LKGWPL- 269
Query: 220 TPNSHQQCQVLKTQNQDAALTQAPACTPNRQTSTVPGT-KRKNKHPKIPKTGSSKNDSQK 278
T + C +LK D + +Q P + + ++ S+ND +
Sbjct: 270 TKSLPTSC-LLKVPGIDQLRSNLGV---QKQLMASPNQFQLLSPQQQLIAQAQSQNDLAR 325
Query: 279 MEQIISTGEHQCNQDQQIQMQ----SQNVEKSTKRMITESTRPVESAQDCADA-KPADEN 333
M +G + D+ M +Q + S R++ + + QD D+ DEN
Sbjct: 326 MGSPAPSGSPKVRPDESDYMMKLKMAQMQQPSGHRLMELQQQQQQLQQDNLDSFVDFDEN 385
Query: 334 VESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHFSSD 393
V+SFLS +D D R + F++LK+ S+ ++ KG SL E G S NKV+ CHFS+D
Sbjct: 386 VDSFLSNDD--GDGR-DIFASLKKGSS--EQDSLKGLSLSEFGNNRTSNNKVVCCHFSTD 440
Query: 394 GKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDA 453
GK++ASAGHEKKVF+WNM+N + H++ ITD+RF+P ST ATSS D +VRLW+A
Sbjct: 441 GKLLASAGHEKKVFLWNMDNLNMDTKIEEHTNFITDIRFKPNSTQLATSSSDGTVRLWNA 500
Query: 454 SNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQV 513
+ I T + ++ +L V + + + +GG+ +V
Sbjct: 501 ------IEIFTRNLQRFYAL-----------------VTTMEKFVSGKLVKMKQGGTGRV 537
Query: 514 RFQPQYGQLLATAIGNSITIIDVETDSHLHDL-KGHDKDVLSICWDRSGNFLASVSEDSA 572
RFQPQ GQLLA A G+ + I+DVE ++ LH L K H +V ICWD G +ASVS+D+
Sbjct: 538 RFQPQIGQLLAVATGSIVNIVDVEKEASLHSLPKVHTNEVNCICWDEKGERVASVSQDTV 597
Query: 573 RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAH 632
++WS AS G CI EL S GN++QSCIFHP Y ++LI+GGYQ++E WS ++ + ++ AH
Sbjct: 598 KVWSVAS-GACIHELRSHGNQYQSCIFHPRYPNVLIVGGYQTMELWSLSDNHRN-TVQAH 655
Query: 633 QGLIAGLADSPVGELIASASHDNSVKVWK 661
+GLIA LA S +IASASHD SVK+WK
Sbjct: 656 EGLIAALAHSQFTGMIASASHDRSVKLWK 684
>ref|NP_917595.1| putative flower development regulator, LEUNI protein [Oryza sativa
(japonica cultivar-group)]
Length = 883
Score = 269 bits (688), Expect = 2e-70
Identities = 152/327 (46%), Positives = 200/327 (60%), Gaps = 24/327 (7%)
Query: 331 DENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHF 390
DENVESFLS +D ++P +L R S + +KGF E+ S KV CHF
Sbjct: 564 DENVESFLSQDD------MDPRDSLGR-----SMDASKGFGFAEVAKARASATKVTCCHF 612
Query: 391 SSDGKVMASAGHEKKVFIWNMEN-FQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
SSDGK++A+ GH+KKV +W E + +S + HS LITDVRF P + ATSSFD++VR
Sbjct: 613 SSDGKLLATGGHDKKVLLWCTEPALKPTSSLEEHSALITDVRFSPSMSRLATSSFDKTVR 672
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECM---HTT 506
+WDA N SL TGH VMSLDFHP + D++CS D + +R W++N C+
Sbjct: 673 VWDADNTSYSLRTFTGHSASVMSLDFHPNKEDMICSCDGDGEVRSWSINNGSCLTFVKVF 732
Query: 507 RGGSKQVRFQPQYGQLLATAIGNSITIIDVETD-SHLHDLKGHDKDVLSICWDRSGNFLA 565
+GG+ Q+RFQPQ G+ LA A +I I+D ET + + L+GH K++ S+CWD +G+ LA
Sbjct: 733 KGGATQMRFQPQKGKYLAAASEKAIYILDGETQLACRNPLQGHTKNIHSLCWDSTGDNLA 792
Query: 566 SVSEDSARIWSAA--SDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEG 623
SVSEDS RIWS A DG+ + EL GNKFQSC+FHP Y LL SLE W E
Sbjct: 793 SVSEDSVRIWSFAPGHDGEFVNELSCSGNKFQSCVFHPSYPYLL------SLELWDIREK 846
Query: 624 SKTWSIAAHQGLIAGLADSPVGELIAS 650
+ +AH GL+A LA S +AS
Sbjct: 847 NAMTVHSAHDGLVAALAASSATGKVAS 873
Score = 42.0 bits (97), Expect = 0.063
Identities = 36/141 (25%), Positives = 58/141 (40%), Gaps = 11/141 (7%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPESSCKV---PMNMMPAGNSNNASPHRIPQ 64
+D+P GFL++WWS+F+++F +R H S ++ M A + PQ
Sbjct: 54 IDAPGGFLFEWWSVFWDIFIARTNEKHSDVAASYIEIHHKQTQHMKAREQQQQQQQQPPQ 113
Query: 65 IPMNEHRPQQFQANSGFNNMIPQPAARLIPQAVFDNGQLGNLAENVEPSLHDLLRENNQK 124
+ +PQ Q M+ Q AA+ Q Q + + R +
Sbjct: 114 --QRQQQPQHIQ----MQQMLLQRAAQQ-QQQQQQQQQQQQQQQQQQQQQQQQQRRDGTH 166
Query: 125 LLSGTNSNHLPHPQDVGKQVQ 145
LL+GT S LP + +Q Q
Sbjct: 167 LLNGTASG-LPGNNPLMRQNQ 186
>ref|NP_916506.1| P0013F10.18 [Oryza sativa (japonica cultivar-group)]
Length = 816
Score = 244 bits (623), Expect = 6e-63
Identities = 136/331 (41%), Positives = 190/331 (57%), Gaps = 54/331 (16%)
Query: 331 DENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHF 390
+++V+SFLS +D AD R + R+ +T GF +E+ + S NKV+ CHF
Sbjct: 540 EDHVDSFLSHDD--ADRR-----DGSRMEST-----KAGFIFREVSSVQASTNKVVCCHF 587
Query: 391 SSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRL 450
SSDGK++A+ GH+KKV +W+ E + + + HS LITDVRF P ATSSFD++VR+
Sbjct: 588 SSDGKLLATGGHDKKVVLWHAETLKQKSVLEEHSLLITDVRFSPSIPRLATSSFDKTVRV 647
Query: 451 WDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGS 510
WDA N S+ TGH VMSLDFHP + DL+CS D ++ IR W++N + +GGS
Sbjct: 648 WDADNQGYSIRTFTGHSASVMSLDFHPNKDDLICSCDGDNEIRFWSINNGNIVRIFKGGS 707
Query: 511 KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED 570
Q+RFQP++G LA A N+++I+DVET + L +GH K V S+CWD SG ++ SVSED
Sbjct: 708 SQLRFQPRHGGYLAVASENAVSILDVETQACLRRFEGHTKHVDSVCWDPSGEYVVSVSED 767
Query: 571 SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIA 630
+ + SLE W +E ++T ++A
Sbjct: 768 TVK----------------------------------------SLELWDMSE-NRTMTLA 786
Query: 631 AHQGLIAGLADSPVGELIASASHDNSVKVWK 661
AH LI LA S G L+AS SHD VK+WK
Sbjct: 787 AHDSLITALASSSSG-LVASTSHDKFVKLWK 816
>gb|AAO22525.1| leunig [Brassica rapa subsp. pekinensis]
Length = 249
Score = 225 bits (573), Expect = 4e-57
Identities = 107/245 (43%), Positives = 152/245 (61%), Gaps = 11/245 (4%)
Query: 331 DENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHF 390
D+NVESFLS ED +R + + +KGF+ E+ + S +KV CHF
Sbjct: 16 DDNVESFLSHEDGD-----------QRDTVGRCMDVSKGFTFAEVNSVRASTSKVTCCHF 64
Query: 391 SSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRL 450
S DGK++ASAGH+KK IW+ + + + + H+ +ITDVRF P ATSSFD++VR+
Sbjct: 65 SLDGKMLASAGHDKKAVIWHTDTMKPKTTLEEHTAMITDVRFSPNLPRLATSSFDKTVRV 124
Query: 451 WDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGS 510
WDA N SL GH + S+DFHP + DL+CS D + IR W++N C +GGS
Sbjct: 125 WDADNKGYSLRNFIGHSSMITSVDFHPNKDDLICSCDDDGEIRYWSINNGTCSRVYKGGS 184
Query: 511 KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED 570
Q RFQP+ G+ LA + N ++++DVET + H L+GH + S+CWD SG+FLA+VSED
Sbjct: 185 TQTRFQPRVGKYLAASSANVVSVLDVETQACRHSLQGHTNQINSVCWDASGDFLATVSED 244
Query: 571 SARIW 575
++W
Sbjct: 245 MVKVW 249
>dbj|BAD53056.1| putative LEUNIG [Oryza sativa (japonica cultivar-group)]
gi|53792192|dbj|BAD52825.1| putative LEUNIG [Oryza
sativa (japonica cultivar-group)]
Length = 774
Score = 219 bits (558), Expect = 2e-55
Identities = 150/451 (33%), Positives = 211/451 (46%), Gaps = 76/451 (16%)
Query: 223 SHQQCQVLKTQNQDAALTQAPACTPNRQTSTVPGTKRKNKHPKIPKTGSSKNDSQKMEQI 282
SHQQ Q L + A + + S + K KI + N S +
Sbjct: 369 SHQQAQSLNQLHHQQAQSIGSMLDGSIPNSFGLANRASKKRKKIVSSSERANSSGTSNNV 428
Query: 283 ISTGE--------HQCNQDQQIQMQSQNVEKS----------TKRMITESTRPVESAQDC 324
S+ H + + N KS TK +I+ T P+
Sbjct: 429 GSSSSSAPSTPFTHTPRDEMSMPQLKYNGGKSKSLSMFGYDDTKSLISP-TNPLGDVDQL 487
Query: 325 ADAKPADENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINK 384
+ DENVESFLS ED ++P + + +KGF E+ S NK
Sbjct: 488 QEDGSLDENVESFLSQED------MDPQETMGHCM-----DASKGFGFIEVAKARASTNK 536
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
V CHFSSDGK++A+ GH+KKV +W ++ A + HS +ITDVRF T ATSSF
Sbjct: 537 VDCCHFSSDGKLLATGGHDKKVVLWFTDDLNIKAIFEEHSMIITDVRFSSIMTRLATSSF 596
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D+++R+WDA+NP SL GH V+SLDFHP + D++CS DS+ +R W+++ C++
Sbjct: 597 DKTIRVWDANNPEYSLHTFIGHSTSVVSLDFHPNKEDIICSCDSDGEVRCWSIDNGSCVN 656
Query: 505 TTR---GGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLH--DLKGHDKDVLSICWDR 559
R GG+ Q+RFQP +G+ LA I+I+D ET H++ DL+GH K++ S+CWD
Sbjct: 657 CVRVFNGGAIQLRFQPHHGKYLAVVSEKMISILDAET-LHIYRSDLQGHLKNIHSVCWDA 715
Query: 560 SGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWS 619
+G +LASVSEDS + SLE W
Sbjct: 716 TGGYLASVSEDSIK----------------------------------------SLELWD 735
Query: 620 PTEGSKTWSIAAHQGLIAGLADSPVGELIAS 650
E + AH G+I LA S LIAS
Sbjct: 736 IREKNIVTINNAHDGMIPSLAASNASGLIAS 766
Score = 43.1 bits (100), Expect = 0.028
Identities = 14/30 (46%), Positives = 23/30 (76%)
Query: 5 DSVLDSPDGFLYDWWSIFYEVFASRYGRGH 34
D +D+P GFL++WWSIF+++F ++ R H
Sbjct: 4 DKAIDAPGGFLFEWWSIFWDIFIAQTDREH 33
>ref|NP_917597.1| putative flower development regulator, LEUNI protein [Oryza sativa
(japonica cultivar-group)]
Length = 650
Score = 186 bits (471), Expect = 3e-45
Identities = 105/284 (36%), Positives = 163/284 (56%), Gaps = 21/284 (7%)
Query: 307 TKRMITESTRPVESAQDCADAKPADENVESFLSFEDEPADNRIEPFSNLKRISATCSRNE 366
TK +I+ T P+ + DENVESFLS ED ++P + +
Sbjct: 373 TKSLISP-TNPLGDVDQLQEDGSLDENVESFLSQED------MDPQETMGHCM-----DA 420
Query: 367 NKGFSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHL 426
+KGF E+ S NKV CHFSSDGK++A+ GH+KKV +W ++ A + HS +
Sbjct: 421 SKGFGFIEVAKARASTNKVDCCHFSSDGKLLATGGHDKKVVLWFTDDLNIKAIFEEHSMI 480
Query: 427 ITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSS 486
ITDVRF T ATSSFD+++R+WDA+NP SL GH V+SLDFHP + D++CS
Sbjct: 481 ITDVRFSSIMTRLATSSFDKTIRVWDANNPEYSLHTFIGHSTSVVSLDFHPNKEDIICSC 540
Query: 487 DSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLH--D 544
DS+ +R W+++ C++ RGG+ Q+RFQP +G+ LA I+I+D ET H++ D
Sbjct: 541 DSDGEVRCWSIDNGSCVNCVRGGAIQLRFQPHHGKYLAVVSEKMISILDAET-LHIYRSD 599
Query: 545 LKG------HDKDVLSICWDRSGNFLASVSEDSARIWSAASDGK 582
L+ +K++++I G + + +++ + ++ +GK
Sbjct: 600 LQSLELWDIREKNIVTINNAHDGMIPSLAASNASGLIASTFEGK 643
Score = 43.1 bits (100), Expect = 0.028
Identities = 14/30 (46%), Positives = 23/30 (76%)
Query: 5 DSVLDSPDGFLYDWWSIFYEVFASRYGRGH 34
D +D+P GFL++WWSIF+++F ++ R H
Sbjct: 4 DKAIDAPGGFLFEWWSIFWDIFIAQTDREH 33
>ref|ZP_00110817.1| COG2319: FOG: WD40 repeat [Nostoc punctiforme PCC 73102]
Length = 1833
Score = 139 bits (350), Expect = 3e-31
Identities = 86/301 (28%), Positives = 151/301 (49%), Gaps = 10/301 (3%)
Query: 364 RNENKGFSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTH 423
+ EN+ E+ L + V + +S +G +ASA +K + IW++ + Q + H
Sbjct: 1157 KKENRAI---EVNTLEGHSDWVSSVAYSPNGYQLASASADKTIKIWDVSSGQLLKTLTGH 1213
Query: 424 SHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLL 483
S I + + P ++S D+++++WD S+ + L LTGH V S+ ++P L
Sbjct: 1214 SDRIRSIAYSPNGQQLVSASADKTIKIWDVSSG-KLLKTLTGHTSAVSSVAYNPNGQQLA 1272
Query: 484 CSSDSNDVIRLWNVNQSECMHTTRGGSK---QVRFQPQYGQLLATAIGNSITIIDVETDS 540
+SD N I++W+++ + + T G S V + P QL + + +I I D+ +
Sbjct: 1273 SASDDN-TIKIWDISSGKLLKTLPGHSSVVNSVAYNPNGQQLASASNDKTIKIWDINSGK 1331
Query: 541 HLHDLKGHDKDVLSICWDRSGNFLASVSEDSA-RIWSAASDGKCIGELHSVGNKFQSCIF 599
L L GH +V S+ + +G LAS S D+ +IW +S GK + L N S +
Sbjct: 1332 LLKSLTGHSSEVNSVAYSPNGQQLASASFDNTIKIWDISS-GKLLKTLTGHSNVVFSVAY 1390
Query: 600 HPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKV 659
P + L ++++ W + G S+A H ++ +A SP G+ +ASAS D ++KV
Sbjct: 1391 SPNGQHLASASADKTIKIWDVSSGKPLKSLAGHSNVVFSVAYSPNGQQLASASDDKTIKV 1450
Query: 660 W 660
W
Sbjct: 1451 W 1451
Score = 136 bits (343), Expect = 2e-30
Identities = 81/282 (28%), Positives = 143/282 (49%), Gaps = 7/282 (2%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
N V + +S +G+ +ASA +K + +W++ N + S HS + V + P A+
Sbjct: 1425 NVVFSVAYSPNGQQLASASDDKTIKVWDISNGKPLESMTDHSDRVNSVVYSPNGQHLASP 1484
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
S+D+++++W+ S+ + L LTGH +V S+ + P L S+ + I++W+VN +
Sbjct: 1485 SYDKTIKIWNVSSG-KLLKTLTGHSSEVNSVAYSPNGQQL-ASASWDKTIKVWDVNSGKP 1542
Query: 503 MHTTRGGSK---QVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDR 559
+ T G S V + P QL + + N+I + DV + L L GH V S+ +
Sbjct: 1543 LKTLIGHSSVVNSVAYSPNGQQLASASFDNTIKVWDVSSGKLLKTLTGHSNAVSSVAYSP 1602
Query: 560 SGNFLASVSEDSA-RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAW 618
+G LAS S D+ +IW +S K + L + S + P + L +++ W
Sbjct: 1603 NGQQLASASLDNTIKIWDVSS-AKLLKTLTGHSDAVSSVAYSPNGQQLASASDDNTIKIW 1661
Query: 619 SPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ G S++ H + +A SP G+ +ASAS DN++K+W
Sbjct: 1662 DVSSGKLLKSLSGHSNAVYSIAYSPNGQQLASASADNTIKIW 1703
Score = 129 bits (324), Expect = 3e-28
Identities = 78/282 (27%), Positives = 142/282 (49%), Gaps = 7/282 (2%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
++V + +S +G+ +AS ++K + IWN+ + + + HS + V + P A++
Sbjct: 1467 DRVNSVVYSPNGQHLASPSYDKTIKIWNVSSGKLLKTLTGHSSEVNSVAYSPNGQQLASA 1526
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
S+D+++++WD N + L L GH V S+ + P L +S N I++W+V+ +
Sbjct: 1527 SWDKTIKVWDV-NSGKPLKTLIGHSSVVNSVAYSPNGQQLASASFDN-TIKVWDVSSGKL 1584
Query: 503 MHTTRGGSK---QVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDR 559
+ T G S V + P QL + ++ N+I I DV + L L GH V S+ +
Sbjct: 1585 LKTLTGHSNAVSSVAYSPNGQQLASASLDNTIKIWDVSSAKLLKTLTGHSDAVSSVAYSP 1644
Query: 560 SGNFLASVSEDSA-RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAW 618
+G LAS S+D+ +IW +S GK + L N S + P + L +++ W
Sbjct: 1645 NGQQLASASDDNTIKIWDVSS-GKLLKSLSGHSNAVYSIAYSPNGQQLASASADNTIKIW 1703
Query: 619 SPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ G S++ H + + +P G+ +ASAS D ++ +W
Sbjct: 1704 DVSSGKLLKSLSGHSDWVMRVTYNPNGQQLASASVDKTIILW 1745
Score = 128 bits (321), Expect = 7e-28
Identities = 76/283 (26%), Positives = 143/283 (49%), Gaps = 9/283 (3%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
+++ + +S +G+ + SA +K + IW++ + + + H+ ++ V + P A++
Sbjct: 1215 DRIRSIAYSPNGQQLVSASADKTIKIWDVSSGKLLKTLTGHTSAVSSVAYNPNGQQLASA 1274
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSND-VIRLWNVNQSE 501
S D ++++WD S+ + L L GH V S+ ++P L +S SND I++W++N +
Sbjct: 1275 SDDNTIKIWDISSG-KLLKTLPGHSSVVNSVAYNPNGQQL--ASASNDKTIKIWDINSGK 1331
Query: 502 CMHTTRGGSKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWD 558
+ + G S +V + P QL + + N+I I D+ + L L GH V S+ +
Sbjct: 1332 LLKSLTGHSSEVNSVAYSPNGQQLASASFDNTIKIWDISSGKLLKTLTGHSNVVFSVAYS 1391
Query: 559 RSGNFLASVSEDSA-RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEA 617
+G LAS S D +IW +S GK + L N S + P + L ++++
Sbjct: 1392 PNGQHLASASADKTIKIWDVSS-GKPLKSLAGHSNVVFSVAYSPNGQQLASASDDKTIKV 1450
Query: 618 WSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
W + G S+ H + + SP G+ +AS S+D ++K+W
Sbjct: 1451 WDISNGKPLESMTDHSDRVNSVVYSPNGQHLASPSYDKTIKIW 1493
Score = 127 bits (319), Expect = 1e-27
Identities = 73/275 (26%), Positives = 140/275 (50%), Gaps = 7/275 (2%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
++ +G+ +ASA + + IW++ + + + HS ++ V + P A++S D++++
Sbjct: 1264 YNPNGQQLASASDDNTIKIWDISSGKLLKTLPGHSSVVNSVAYNPNGQQLASASNDKTIK 1323
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
+WD N + L LTGH +V S+ + P L +S N I++W+++ + + T G
Sbjct: 1324 IWDI-NSGKLLKSLTGHSSEVNSVAYSPNGQQLASASFDN-TIKIWDISSGKLLKTLTGH 1381
Query: 510 SK---QVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
S V + P L + + +I I DV + L L GH V S+ + +G LAS
Sbjct: 1382 SNVVFSVAYSPNGQHLASASADKTIKIWDVSSGKPLKSLAGHSNVVFSVAYSPNGQQLAS 1441
Query: 567 VSEDSA-RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
S+D ++W S+GK + + ++ S ++ P + L ++++ W+ + G
Sbjct: 1442 ASDDKTIKVWDI-SNGKPLESMTDHSDRVNSVVYSPNGQHLASPSYDKTIKIWNVSSGKL 1500
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
++ H + +A SP G+ +ASAS D ++KVW
Sbjct: 1501 LKTLTGHSSEVNSVAYSPNGQQLASASWDKTIKVW 1535
Score = 45.4 bits (106), Expect = 0.006
Identities = 20/70 (28%), Positives = 38/70 (53%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
N V + +S +G+ +ASA + + IW++ + + S HS + V + P A++
Sbjct: 1677 NAVYSIAYSPNGQQLASASADNTIKIWDVSSGKLLKSLSGHSDWVMRVTYNPNGQQLASA 1736
Query: 443 SFDRSVRLWD 452
S D+++ LWD
Sbjct: 1737 SVDKTIILWD 1746
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.315 0.130 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,207,484,786
Number of Sequences: 2540612
Number of extensions: 53917750
Number of successful extensions: 185934
Number of sequences better than 10.0: 4896
Number of HSP's better than 10.0 without gapping: 1891
Number of HSP's successfully gapped in prelim test: 3044
Number of HSP's that attempted gapping in prelim test: 151486
Number of HSP's gapped (non-prelim): 20594
length of query: 661
length of database: 863,360,394
effective HSP length: 135
effective length of query: 526
effective length of database: 520,377,774
effective search space: 273718709124
effective search space used: 273718709124
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 79 (35.0 bits)
Lotus: description of TM0114.1


