

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0113.13
(359 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
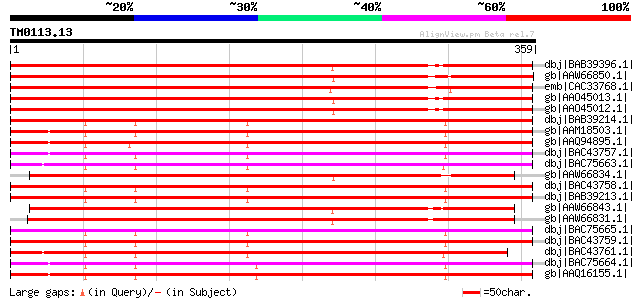
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAB39396.1| S-adenosyl-L-methionine:salicylic acid carboxyl ... 325 1e-87
gb|AAW66850.1| SAMT [Nicotiana tabacum] 322 1e-86
emb|CAC33768.1| S-adenosyl-L-methionine:salicylic acid carboxyl ... 321 2e-86
gb|AAO45013.1| S-adenosyl-L-methionine:benzoic acid/salicylic ac... 318 2e-85
gb|AAO45012.1| S-adenosyl-L-methionine:benzoic acid/salicylic ac... 316 8e-85
dbj|BAB39214.1| theobromine synthase [Coffea arabica] 307 3e-82
gb|AAM18503.1| N-methyltransferase [Coffea arabica] 307 4e-82
gb|AAQ94895.1| putative N-methyltransferase [Coffea liberica var... 306 6e-82
dbj|BAC43757.1| theobromine synthse 2 [Coffea arabica] 306 6e-82
dbj|BAC75663.1| 3,7-dimethylxanthine N-methyltransferase [Coffea... 305 1e-81
gb|AAW66834.1| SAMT [Petunia nyctaginiflora] 305 1e-81
dbj|BAC43758.1| tentative caffeine synthase 3 [Coffea arabica] 304 3e-81
dbj|BAB39213.1| caffeine synthase [Coffea arabica] 303 5e-81
gb|AAW66843.1| SAMT [Juanulloa aurantiaca] 303 5e-81
gb|AAW66831.1| SAMT [Brugmansia sp. Robadey 027] 303 5e-81
dbj|BAC75665.1| Xanthosine N-methyltransferase [Coffea arabica] 303 7e-81
dbj|BAC43759.1| tentative caffeine synthase 4 [Coffea arabica] 303 7e-81
dbj|BAC43761.1| tentative caffeine synthase 7 [Coffea arabica] 303 7e-81
dbj|BAC75664.1| 7-methylxanthine N-methyltransferase [Coffea ara... 302 9e-81
gb|AAQ16155.1| putative caffeine synthase [Coffea canephora] gi|... 302 1e-80
>dbj|BAB39396.1| S-adenosyl-L-methionine:salicylic acid carboxyl methyltransferase
[Atropa belladonna]
Length = 357
Score = 325 bits (834), Expect = 1e-87
Identities = 172/361 (47%), Positives = 235/361 (64%), Gaps = 10/361 (2%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPSCLKVADLGCS 60
M + +VLHMNGG+GD SYANNS QR V+L TK I E +I LYC+ FP L +ADLGCS
Sbjct: 1 MKVVEVLHMNGGNGDISYANNSLVQRKVILMTKPITEQAISDLYCSFFPETLCIADLGCS 60
Query: 61 SGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEK 120
SG N +V S ++ V+ + NL+ F NDL GNDFNT F+SL F + L+++
Sbjct: 61 SGANTFLVVSELVKIVEKERKIHNLQSAGNLFHFNDLPGNDFNTIFQSLGKFQQDLRKQI 120
Query: 121 GHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPLNKGAIYLTKTSPP 180
G +FGPCFFSG PGSFY RLFP++S+HF HSSYSL WLS+V NK IY+ TSPP
Sbjct: 121 GEEFGPCFFSGVPGSFYTRLFPSESLHFVHSSYSLMWLSQVPDLIEKNKENIYIASTSPP 180
Query: 181 AVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRD----ENNELINAWVVIGMVLN 236
+V KAY+ Q+ +DF FL RS EL+ GG MV+T +GR+ + E W ++ M LN
Sbjct: 181 SVIKAYYKQYEKDFSNFLKYRSEELMKGGKMVLTFLGRESEDPSSKECCYIWELLSMALN 240
Query: 237 DMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDI 296
++ E LIE+ K+D+FNIP Y P+ EE++ ++E+EGSF I RLE R W NVS+
Sbjct: 241 ELVLEGLIEEEKVDSFNIPQYTPSPEEVKYIVEKEGSFIINRLEATRVHW----NVSN-- 294
Query: 297 DEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVKLHGVEKLELANLVMHIT 356
+ VAK +RAVAE +L S+F + +M+ +F++++ I EK E N+++ +T
Sbjct: 295 EGINGGYNVAKCMRAVAEPLLVSQFDQKLMNLVFQKYEEIISDCISKEKTEFINVIVSLT 354
Query: 357 K 357
K
Sbjct: 355 K 355
>gb|AAW66850.1| SAMT [Nicotiana tabacum]
Length = 358
Score = 322 bits (825), Expect = 1e-86
Identities = 167/362 (46%), Positives = 234/362 (64%), Gaps = 9/362 (2%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPSCLKVADLGCS 60
M + +VLHMNGG GD SYA NS Q+ V+L TK I E +I LYC+ FP L +ADLGCS
Sbjct: 1 MKVVEVLHMNGGIGDISYAKNSLVQQKVILMTKPITEQAITDLYCSLFPQNLCIADLGCS 60
Query: 61 SGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEK 120
SG N +V S +I V+ + + P F F NDL GNDFNT F+SL F + L+++
Sbjct: 61 SGANTFIVVSELIKIVEKERKKHGFQSPEFHFNFNDLPGNDFNTIFQSLDIFQQDLRKQI 120
Query: 121 GHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPLNKGAIYLTKTSPP 180
G +FGPCFFSG GSFY RLFP++S+HF HSSYSL WLS+V NKG IY+ TSPP
Sbjct: 121 GEEFGPCFFSGVSGSFYTRLFPSNSLHFVHSSYSLMWLSQVPDAVENNKGNIYMASTSPP 180
Query: 181 AVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDE----NNELINAWVVIGMVLN 236
+V KAY+ Q+ +DF FL RS EL+ GG MV+T +GR+ + E W ++ M LN
Sbjct: 181 SVIKAYYKQYEKDFSNFLKYRSEELMKGGKMVLTFLGRESEDPTSKECCYIWELLAMALN 240
Query: 237 DMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDI 296
++ E LIE+ K+D+FNIP Y P+ +++ V+E+EGSF I +LE R W N +D
Sbjct: 241 ELVVEGLIEEEKVDSFNIPQYTPSPADVKYVVEKEGSFTINQLEATRVHW----NACNDK 296
Query: 297 DEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVKLHGVEKLELANLVMHIT 356
++ +V++ +RAVAE +L S+FGE +MD +F +++ I + + E N+++ +T
Sbjct: 297 YKNV-GYSVSRCMRAVAEPLLVSQFGEELMDLVFHKYEQIISECMSKAQTEFTNVIVSLT 355
Query: 357 KS 358
K+
Sbjct: 356 KT 357
>emb|CAC33768.1| S-adenosyl-L-methionine:salicylic acid carboxyl methyltransferase
[Stephanotis floribunda]
Length = 366
Score = 321 bits (822), Expect = 2e-86
Identities = 169/369 (45%), Positives = 226/369 (60%), Gaps = 17/369 (4%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPSCLKVADLGCS 60
M + +VLHMNGG GDASYA+NS Q+ V+L TK I E++I LY FP + +AD+GCS
Sbjct: 1 MEVVEVLHMNGGTGDASYASNSLLQKKVILLTKPITEEAITELYTRLFPKSICIADMGCS 60
Query: 61 SGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEK 120
SGPN + S +I V+ +L E P +Q LNDL NDFNT F+SLP F K ++
Sbjct: 61 SGPNTFLAVSELIKNVEKKRTSLGHESPEYQIHLNDLPSNDFNTIFRSLPSFQKSFSKQM 120
Query: 121 GHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPLNKGAIYLTKTSPP 180
G FG CFF+G PGSFYGRLFPN S+HF HSSYSL WLS+V + +NKG IYL+ TSP
Sbjct: 121 GSGFGHCFFTGVPGSFYGRLFPNKSLHFVHSSYSLMWLSRVPDLEEVNKGNIYLSSTSPL 180
Query: 181 AVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGR----DENNELINAWVVIGMVLN 236
+V +AY QF+ DF FL R+ EL+PGG MV+TL+GR E A ++ LN
Sbjct: 181 SVIRAYLKQFQRDFTTFLQCRAEELVPGGVMVLTLMGRKGEDHSGKESGYALELLARALN 240
Query: 237 DMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDI 296
++ SE IE+ +LD FN+P Y P+ E++ +EEEGSF I RLE W + D
Sbjct: 241 ELVSEGQIEEEQLDCFNVPQYTPSPAEVKYFVEEEGSFSITRLEATTIHW-----TAYDH 295
Query: 297 DEDT--------RAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVKLHGVEKLEL 348
D T +++ +RAV E +L FGEAIMDE+F R++ + EK+E
Sbjct: 296 DHVTGHHHAFKDGGYSLSNCVRAVVEPLLVRHFGEAIMDEVFHRYREILTNCMTKEKIEF 355
Query: 349 ANLVMHITK 357
N+ + + +
Sbjct: 356 INVTVSMKR 364
>gb|AAO45013.1| S-adenosyl-L-methionine:benzoic acid/salicylic acid carboxyl
methyltransferase [Petunia x hybrida]
Length = 357
Score = 318 bits (815), Expect = 2e-85
Identities = 168/361 (46%), Positives = 230/361 (63%), Gaps = 10/361 (2%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPSCLKVADLGCS 60
M + +VLHMNGG+GD+SYANNS Q+ V+L TK I E ++I LY + FP L +ADLGCS
Sbjct: 1 MEVVEVLHMNGGNGDSSYANNSLVQQKVILMTKPITEQAMIDLYSSLFPETLCIADLGCS 60
Query: 61 SGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEK 120
G N +V S ++ V+ + + P F F NDL GNDFNT F+SL F + L++
Sbjct: 61 LGANTFLVVSQLVKIVEKERKKHGFKSPEFYFHFNDLPGNDFNTLFQSLGAFQEDLRKHI 120
Query: 121 GHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPLNKGAIYLTKTSPP 180
G FGPCFFSG PGSFY RLFP+ S+HF +SSYSL WLS+V NKG IY+ +TSP
Sbjct: 121 GESFGPCFFSGVPGSFYTRLFPSKSLHFVYSSYSLMWLSQVPNGIENNKGNIYMARTSPL 180
Query: 181 AVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDE----NNELINAWVVIGMVLN 236
+V KAY+ Q+ DF FL RS EL+ GG MV+TL+GR+ + E W ++ M LN
Sbjct: 181 SVIKAYYKQYEIDFSNFLKYRSEELMKGGKMVLTLLGRESEDPTSKECCYIWELLAMALN 240
Query: 237 DMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDI 296
+ E LI++ K+DAFNIP Y P+ E++ ++E+EGSF I RLET R W N S+
Sbjct: 241 KLVEEGLIKEEKVDAFNIPQYTPSPAEVKYIVEKEGSFTINRLETSRVHW----NASN-- 294
Query: 297 DEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVKLHGVEKLELANLVMHIT 356
+E V++ +RAVAE +L S F + +MD +F +++ I EK E N+++ +T
Sbjct: 295 NEKNGGYNVSRCMRAVAEPLLVSHFDKELMDLVFHKYEEIISDCMSKEKTEFINVIVSLT 354
Query: 357 K 357
K
Sbjct: 355 K 355
>gb|AAO45012.1| S-adenosyl-L-methionine:benzoic acid/salicylic acid carboxyl
methyltransferase [Petunia x hybrida]
Length = 357
Score = 316 bits (809), Expect = 8e-85
Identities = 166/361 (45%), Positives = 229/361 (62%), Gaps = 10/361 (2%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPSCLKVADLGCS 60
M + +VLHMNGG+GD+SYANNS Q+ V+L TK I E ++I LY + FP L +ADLGCS
Sbjct: 1 MEVVEVLHMNGGNGDSSYANNSLVQQKVILMTKPITEQAMIDLYSSLFPETLCIADLGCS 60
Query: 61 SGPNALMVASNIINTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEK 120
G N +V S ++ V+ + + P F F NDL GNDFNT F+SL F + L++
Sbjct: 61 LGANTFLVVSQLVKIVEKERKKHGFKSPEFYFHFNDLPGNDFNTLFQSLGAFQEDLRKHI 120
Query: 121 GHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPLNKGAIYLTKTSPP 180
G FGPCFFSG PGSFY RLFP+ S+HF +SSYSL WLS+V NKG IY+ +TSP
Sbjct: 121 GESFGPCFFSGVPGSFYTRLFPSKSLHFVYSSYSLMWLSQVPNGIENNKGNIYMARTSPL 180
Query: 181 AVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDE----NNELINAWVVIGMVLN 236
+V KAY+ Q+ DF FL RS EL+ GG MV+TL+GR+ + E W ++ M LN
Sbjct: 181 SVIKAYYKQYEIDFSNFLKYRSEELMKGGKMVLTLLGRESEDPTSKECCYIWELLAMALN 240
Query: 237 DMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDI 296
+ E LI++ K+DAFNIP Y P+ E++ ++E+EGSF I RLET R W N S+
Sbjct: 241 KLVEEGLIKEEKVDAFNIPQYTPSPAEVKYIVEKEGSFTINRLETSRVHW----NASN-- 294
Query: 297 DEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVKLHGVEKLELANLVMHIT 356
+E V++ +RAVAE +L S F + +MD +F +++ + E E N+++ +T
Sbjct: 295 NEKNGGYNVSRCMRAVAEPLLVSHFDKELMDLVFHKYEEIVSDCMSKENTEFINVIISLT 354
Query: 357 K 357
K
Sbjct: 355 K 355
>dbj|BAB39214.1| theobromine synthase [Coffea arabica]
Length = 385
Score = 307 bits (787), Expect = 3e-82
Identities = 165/378 (43%), Positives = 231/378 (60%), Gaps = 21/378 (5%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPS---CLKVADL 57
M +++VLHMNGG+GDASYA NSSF + V+ K +LE + L A P+ C+KVADL
Sbjct: 1 MELQEVLHMNGGEGDASYAKNSSFNQLVLAKVKPVLEQCVGELLRANLPNINKCIKVADL 60
Query: 58 GCSSGPNALMVASNIINTVDAVSQNLN--LEQPVFQFFLNDLFGNDFNTTFKSLPDFYKR 115
GC+SGPN L+ +I+ ++D V Q + LE+P Q FL DLF NDFN+ F LP FY++
Sbjct: 61 GCASGPNTLLTVRDIVQSIDKVRQEMKNELERPTIQVFLTDLFQNDFNSVFMLLPSFYRK 120
Query: 116 LQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLNK 169
L++E G K G C + PGSF+GRLFP +S+HF HSSYSL +LS+V NK
Sbjct: 121 LEKENGRKIGSCLIAAMPGSFHGRLFPEESMHFLHSSYSLQFLSQVPSGLVTELGITANK 180
Query: 170 GAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAWV 229
+IY +K SPP V KAY QF +DF FL RS ELL G M++T I + + + N
Sbjct: 181 RSIYSSKASPPPVQKAYLDQFTKDFTTFLRIRSEELLSRGRMLLTCICKGDEFDGPNTMD 240
Query: 230 VIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKN 289
++ M +ND+ E +E+ KLD+FN+P Y + EE++ ++EEEGSF+I LET + +
Sbjct: 241 LLEMAINDLVVEGHLEEEKLDSFNVPIYAASVEELKCIVEEEGSFEILYLETFKLRYDAG 300
Query: 290 INVSDDI----------DEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVK 339
++ DD DE RA VA +R+V E IL + FGEAI+ ++F RF K
Sbjct: 301 FSIDDDCQVRSHSPEYSDEHARAAHVASLLRSVYEPILANHFGEAIIPDIFHRFATNAAK 360
Query: 340 LHGVEKLELANLVMHITK 357
+ + K NL++ + K
Sbjct: 361 VIRLGKGFYNNLIISLAK 378
>gb|AAM18503.1| N-methyltransferase [Coffea arabica]
Length = 378
Score = 307 bits (786), Expect = 4e-82
Identities = 168/372 (45%), Positives = 226/372 (60%), Gaps = 16/372 (4%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPS---CLKVADL 57
M +++VLHMNGG+GD SYA NSS+ + K +LE I L A P+ C+KVADL
Sbjct: 1 MELQEVLHMNGGEGDTSYAKNSSYNL-ALAKVKPVLEQCIRELLRANLPNINNCIKVADL 59
Query: 58 GCSSGPNALMVASNIINTVDAVSQNLN--LEQPVFQFFLNDLFGNDFNTTFKSLPDFYKR 115
GC+SGPN L+ +I+ ++D V Q LE+P Q FLNDLF NDFN+ FK LP FY++
Sbjct: 60 GCASGPNTLLTVRDIVQSIDKVGQEEKNELERPTIQIFLNDLFQNDFNSVFKLLPSFYRK 119
Query: 116 LQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLNK 169
L++E G K G C S PGSFYGRLFP +S+HF HS YS HWLS+V NK
Sbjct: 120 LEKENGRKIGSCLISAMPGSFYGRLFPEESMHFIHSCYSFHWLSQVPSGLVIELGISANK 179
Query: 170 GAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAWV 229
G+IY +K P V KAY QF +DF FL S EL G M++T I + + + N
Sbjct: 180 GSIYSSKGCRPPVQKAYLDQFTKDFTTFLRIHSKELFSRGRMLLTCICKVDEFDEPNPLD 239
Query: 230 VIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKN 289
++ M +ND+ E L+E+ KLD+FNIP + P+AEE++ ++EEEGS +I LET + +
Sbjct: 240 LLDMAINDLIVEGLLEEEKLDSFNIPFFTPSAEEVKCIVEEEGSCEILYLETFKAHYDAA 299
Query: 290 INVSDDI----DEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVKLHGVEK 345
++ DD E +AE VA IR+V E IL S FGEAIM +LF R K+ + K
Sbjct: 300 FSIDDDYPVRSHEQIKAEYVASLIRSVYEPILASHFGEAIMPDLFHRLAKHAAKVLHMGK 359
Query: 346 LELANLVMHITK 357
NL++ + K
Sbjct: 360 GCYNNLIISLAK 371
>gb|AAQ94895.1| putative N-methyltransferase [Coffea liberica var. dewevrei]
Length = 384
Score = 306 bits (784), Expect = 6e-82
Identities = 167/378 (44%), Positives = 228/378 (60%), Gaps = 22/378 (5%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPS---CLKVADL 57
M +++VLHMNGG+GD SYA NSS+ + K +LE I L A P+ C+KVADL
Sbjct: 1 MELQEVLHMNGGEGDTSYAKNSSYNL-ALAKVKPVLEQCIRELLRANLPNINNCIKVADL 59
Query: 58 GCSSGPNALMVASNIINTVDAVS--QNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKR 115
GC+SGPN L+ +I+ ++D V + LE+P Q FLNDLF NDFN+ FK LP FY++
Sbjct: 60 GCASGPNTLLTVRDIVQSIDKVGLEEKNELERPTVQIFLNDLFQNDFNSVFKLLPSFYRK 119
Query: 116 LQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLNK 169
L++E G K G C S PGSF+GRLFP +S+HF HS YS+HWLS+V NK
Sbjct: 120 LEKENGRKIGSCLISAMPGSFHGRLFPEESMHFLHSCYSIHWLSQVPSGLVIELGISANK 179
Query: 170 GAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAWV 229
G+IY +K S P V KAY QF +DF FL S EL G M++T I + + + N
Sbjct: 180 GSIYSSKASRPPVQKAYLDQFTKDFTTFLRIHSKELFSRGRMLLTCICKVDEFDEPNPLD 239
Query: 230 VIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKN 289
++ M +ND+ E +E+ KL +FN+P Y P+AEE++ ++EEEGSF+I LET + +
Sbjct: 240 LLDMAINDLVVEGHLEEEKLASFNLPFYTPSAEEVKCIVEEEGSFEILYLETFKAHYDAG 299
Query: 290 INVSDDI----------DEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVK 339
++ DD DE +AE VA IR+V E IL S FGEAIM +LF R K
Sbjct: 300 FSIDDDYPVRSHFQGYGDEHIKAEYVASLIRSVYEPILASHFGEAIMPDLFHRLAKHAAK 359
Query: 340 LHGVEKLELANLVMHITK 357
+ + K NL++ + K
Sbjct: 360 VLRLGKGCYNNLIISLAK 377
>dbj|BAC43757.1| theobromine synthse 2 [Coffea arabica]
Length = 384
Score = 306 bits (784), Expect = 6e-82
Identities = 168/378 (44%), Positives = 226/378 (59%), Gaps = 22/378 (5%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPS---CLKVADL 57
M ++ VLHMNGG+GD SYA NSS+ + K +LE I L A P+ C+KVADL
Sbjct: 1 MELQAVLHMNGGEGDTSYAKNSSYNL-ALAKVKPVLEQCIRELLRANLPNINNCIKVADL 59
Query: 58 GCSSGPNALMVASNIINTVDAVSQNLN--LEQPVFQFFLNDLFGNDFNTTFKSLPDFYKR 115
GC+SGPN L+ +I+ ++D V Q LE+P Q FLNDLF NDFN+ FK LP FY++
Sbjct: 60 GCASGPNTLLTVRDIVQSIDKVGQEEKNELERPTIQIFLNDLFQNDFNSVFKLLPSFYRK 119
Query: 116 LQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLNK 169
L++E G K G C S PGSFYGRLFP +S+HF HS YS HWLS+V NK
Sbjct: 120 LEKENGRKIGSCLISAMPGSFYGRLFPEESMHFIHSCYSFHWLSQVPSGLVIELGISANK 179
Query: 170 GAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAWV 229
G+IY +K S P V KAY QF +DF FL S EL G M++T I + + + N
Sbjct: 180 GSIYSSKASRPPVQKAYLDQFTKDFTTFLRIHSKELFSRGRMLLTCICKVDEYDEPNPLD 239
Query: 230 VIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKN 289
++ M +ND+ E +E+ KL +FN+P + P+AEE++ ++EEEGSF+I LET + +
Sbjct: 240 LLDMAINDLIVEGHLEEEKLASFNLPFFTPSAEEVKCIVEEEGSFEILYLETFKAHYDAG 299
Query: 290 INVSDDI----------DEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVK 339
++ DD DE +AE VA IR+V E IL S FGEAIM +LF R K
Sbjct: 300 FSIDDDYPVRSHFQVYGDEHIKAEYVASLIRSVYEPILASHFGEAIMPDLFHRLAKHAAK 359
Query: 340 LHGVEKLELANLVMHITK 357
+ + K NL++ + K
Sbjct: 360 VLHLGKGCYNNLIISLAK 377
>dbj|BAC75663.1| 3,7-dimethylxanthine N-methyltransferase [Coffea arabica]
Length = 384
Score = 305 bits (782), Expect = 1e-81
Identities = 165/378 (43%), Positives = 226/378 (59%), Gaps = 22/378 (5%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPS---CLKVADL 57
M +++VLHMNGG+GD SYA NS F ++ K ILE I L A P+ C+KVADL
Sbjct: 1 MELQEVLHMNGGEGDTSYAKNS-FYNLFLIRVKPILEQCIQELLRANLPNINKCIKVADL 59
Query: 58 GCSSGPNALMVASNIINTVDAVSQNLN--LEQPVFQFFLNDLFGNDFNTTFKSLPDFYKR 115
GC+SGPN L+ +I+ ++D V Q LE+P Q FLNDLF NDFN+ FKSLP FY++
Sbjct: 60 GCASGPNTLLTVRDIVQSIDKVGQEKKNELERPTIQIFLNDLFQNDFNSVFKSLPSFYRK 119
Query: 116 LQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLNK 169
L++E G K G C PGSFYGRLFP +S+HF HS Y LHWLS+V NK
Sbjct: 120 LEKENGRKIGSCLIGAMPGSFYGRLFPEESMHFLHSCYCLHWLSQVPSGLVTELGISANK 179
Query: 170 GAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAWV 229
G IY +K S P + KAY QF +DF FL S EL+ G M++T I +++ E N+
Sbjct: 180 GCIYSSKASRPPIQKAYLDQFTKDFTTFLRIHSEELISRGRMLLTWICKEDEFENPNSID 239
Query: 230 VIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKN 289
++ M +ND+ E +E+ KLD+FN+P Y P+ EE++ ++EEEGSF+I LET + +
Sbjct: 240 LLEMSINDLVIEGHLEEEKLDSFNVPIYAPSTEEVKCIVEEEGSFEILYLETFKVPYDAG 299
Query: 290 INVSDD----------IDEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVK 339
++ DD DE RA VA +R++ E I+ S FGEAIM +L R K
Sbjct: 300 FSIDDDYQGRSHSPVSCDEHARAAHVASVVRSIFEPIVASHFGEAIMPDLSHRIAKNAAK 359
Query: 340 LHGVEKLELANLVMHITK 357
+ K +L++ + K
Sbjct: 360 VLRSGKGFYDSLIISLAK 377
>gb|AAW66834.1| SAMT [Petunia nyctaginiflora]
Length = 332
Score = 305 bits (781), Expect = 1e-81
Identities = 159/336 (47%), Positives = 214/336 (63%), Gaps = 10/336 (2%)
Query: 14 GDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPSCLKVADLGCSSGPNALMVASNII 73
GD SYANNS Q V+L TK I+E ++ LY + FP L +ADLGCSSG N +V S ++
Sbjct: 2 GDMSYANNSLVQAKVILMTKPIIEQAMKDLYSSLFPETLCIADLGCSSGANTFLVVSELV 61
Query: 74 NTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKGHKFGPCFFSGTP 133
++ +N + P F F NDL GNDFNT F+SL F + L ++ G FGPCFFSG P
Sbjct: 62 KIIEKERKNHGFKSPEFYFHFNDLPGNDFNTIFQSLGPFQEDLTKQIGESFGPCFFSGVP 121
Query: 134 GSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPLNKGAIYLTKTSPPAVHKAYFAQFRED 193
GSFY RLFP++S++F HSSYSL WLS+V NKG IY+ +TSPP+V KAY+ Q+ D
Sbjct: 122 GSFYTRLFPSNSLNFIHSSYSLMWLSQVPVAVESNKGNIYMARTSPPSVIKAYYKQYEID 181
Query: 194 FKLFLGSRSCELLPGGAMVITLIGRDE----NNELINAWVVIGMVLNDMASENLIEKAKL 249
F FL RS EL+ GG MV+TL+GR+ + E W ++ M LN++ E LIE+ KL
Sbjct: 182 FSNFLKYRSEELMKGGRMVLTLLGRESEDPTSKECCYIWELLAMALNELVEEGLIEEEKL 241
Query: 250 DAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDIDEDTRAEAVAKYI 309
DAFNIP Y P+ EE++ V+E+EGSF I RLET R W + NV + V++ +
Sbjct: 242 DAFNIPQYTPSPEEVKYVVEKEGSFTINRLETSRVHWNASNNVKNG------GYNVSRCM 295
Query: 310 RAVAESILKSEFGEAIMDELFRRFKNKIVKLHGVEK 345
RAVAE +L S F + +MD +F +++ + EK
Sbjct: 296 RAVAEPLLVSHFDKELMDLVFHKYEEIVSDCMSKEK 331
>dbj|BAC43758.1| tentative caffeine synthase 3 [Coffea arabica]
Length = 385
Score = 304 bits (778), Expect = 3e-81
Identities = 164/378 (43%), Positives = 230/378 (60%), Gaps = 21/378 (5%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPS---CLKVADL 57
M +++VLHMNGG+GDASYA NSSF + V+ K +LE + L A P+ C+KVADL
Sbjct: 1 MELQEVLHMNGGEGDASYAKNSSFNQLVLAKVKPVLEQCVGELLRANLPNINKCIKVADL 60
Query: 58 GCSSGPNALMVASNIINTVDAVSQNLN--LEQPVFQFFLNDLFGNDFNTTFKSLPDFYKR 115
GC+SGPN L+ +I+ ++D V Q + LE+P Q FL DLF NDFN+ F LP FY++
Sbjct: 61 GCASGPNTLLTVRDIVQSIDKVRQEMKNELERPTIQVFLTDLFQNDFNSVFMLLPSFYRK 120
Query: 116 LQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLNK 169
L++E G K G C + PGSF+GRLFP +S+HF HSSYSL +LS+V NK
Sbjct: 121 LEKENGRKIGSCLIAAMPGSFHGRLFPEESMHFLHSSYSLQFLSQVPSGLVTELGITANK 180
Query: 170 GAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAWV 229
+IY +K SPP V KA QF +DF FL RS ELL G M++T I + + + N
Sbjct: 181 RSIYSSKASPPPVQKANLDQFTKDFTTFLRIRSEELLSRGRMLLTCICKGDEFDGPNTMD 240
Query: 230 VIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKN 289
++ M +ND+ E +E+ KLD+FN+P Y + EE++ ++EEEGSF+I LET + +
Sbjct: 241 LLEMAINDLVVEGHLEEEKLDSFNVPIYAASVEELKCIVEEEGSFEILYLETFKLRYDAG 300
Query: 290 INVSDDI----------DEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVK 339
++ DD DE RA VA +R+V E IL + FGEAI+ ++F RF K
Sbjct: 301 FSIDDDCQVRSHSPEYSDEHARAAHVASLLRSVYEPILANHFGEAIIPDIFHRFATNAAK 360
Query: 340 LHGVEKLELANLVMHITK 357
+ + K NL++ + K
Sbjct: 361 VIRLGKGFYNNLIISLAK 378
>dbj|BAB39213.1| caffeine synthase [Coffea arabica]
Length = 385
Score = 303 bits (776), Expect = 5e-81
Identities = 163/378 (43%), Positives = 230/378 (60%), Gaps = 21/378 (5%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPS---CLKVADL 57
M +++VLHMNGG+G+ASYA NSSF + V+ K +LE + L A P+ C+KVADL
Sbjct: 1 MELQEVLHMNGGEGEASYAKNSSFNQLVLAKVKPVLEQCVRELLRANLPNINKCIKVADL 60
Query: 58 GCSSGPNALMVASNIINTVDAVSQNLN--LEQPVFQFFLNDLFGNDFNTTFKSLPDFYKR 115
GC+SGPN L+ + + ++D V Q + LE+P Q FL DLF NDFN+ F LP FY++
Sbjct: 61 GCASGPNTLLTVWDTVQSIDKVKQEMKNELERPTIQVFLTDLFQNDFNSVFMLLPSFYRK 120
Query: 116 LQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLNK 169
L++E G K G C + PGSF+GRLFP +S+HF HSSYSL +LS+V NK
Sbjct: 121 LEKENGRKIGSCLIAAMPGSFHGRLFPEESMHFLHSSYSLQFLSQVPSGLVTELGITANK 180
Query: 170 GAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAWV 229
+IY +K SPP V KAY QF +DF FL RS ELL G M++T I + + + N
Sbjct: 181 RSIYSSKASPPPVQKAYLDQFTKDFTTFLRMRSEELLSRGRMLLTCICKGDECDGPNTMD 240
Query: 230 VIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKN 289
++ M +ND+ +E + + KLD+FN+P Y + EE++ ++EEEGSF+I L+T + +
Sbjct: 241 LLEMAINDLVAEGRLGEEKLDSFNVPIYTASVEEVKCMVEEEGSFEILYLQTFKLRYDAG 300
Query: 290 INVSDDI----------DEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVK 339
++ DD DE RA VA IR+V E IL S FGEAI+ ++F RF K
Sbjct: 301 FSIDDDCQVRSHSPVYSDEHARAAHVASLIRSVYEPILASHFGEAIIPDIFHRFATNAAK 360
Query: 340 LHGVEKLELANLVMHITK 357
+ + K NL++ + K
Sbjct: 361 VIRLGKGFYNNLIISLAK 378
>gb|AAW66843.1| SAMT [Juanulloa aurantiaca]
Length = 334
Score = 303 bits (776), Expect = 5e-81
Identities = 162/336 (48%), Positives = 213/336 (63%), Gaps = 8/336 (2%)
Query: 14 GDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPSCLKVADLGCSSGPNALMVASNII 73
GD SYANNS QR V+L TK I E +I LYC FP L +ADLGCSSG N +V S +I
Sbjct: 2 GDISYANNSLVQRKVILMTKPITEQAISDLYCTLFPETLCIADLGCSSGANTFLVVSELI 61
Query: 74 NTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKGHKFGPCFFSGTP 133
++ + NL+ P F F NDL GNDFNT F+SL F + L+++ G FG CFFSG
Sbjct: 62 KIIEKERKKHNLQSPEFHFNFNDLPGNDFNTIFQSLGGFEQDLRKQIGEGFGSCFFSGVA 121
Query: 134 GSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPLNKGAIYLTKTSPPAVHKAYFAQFRED 193
GSFY RLFP++S+HF HSSYSL WLS+V NKG IY+ TSP +V KAY+ Q+ +D
Sbjct: 122 GSFYTRLFPSNSLHFVHSSYSLMWLSQVPDLIEKNKGNIYMASTSPASVIKAYYEQYEKD 181
Query: 194 FKLFLGSRSCELLPGGAMVITLIGRD----ENNELINAWVVIGMVLNDMASENLIEKAKL 249
F FL RS EL+ GG MV+T +GR+ + E W ++ M LN++ E LIE+ K+
Sbjct: 182 FSNFLKYRSEELMKGGKMVLTFLGRESEDPSSKECCYIWELLSMALNELVVEGLIEEGKV 241
Query: 250 DAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDIDEDTRAEAVAKYI 309
DAFNIP Y P+ E++ V+E+EGSF I RLE R W N S+D + V++ +
Sbjct: 242 DAFNIPQYTPSPTEVKYVVEKEGSFTINRLEATRVHW---NNASND-ENINGGYNVSRCM 297
Query: 310 RAVAESILKSEFGEAIMDELFRRFKNKIVKLHGVEK 345
RAVAE +L S+FG +MD +F+++K I EK
Sbjct: 298 RAVAEPLLASQFGPKLMDLVFQKYKEIIFDCMSKEK 333
>gb|AAW66831.1| SAMT [Brugmansia sp. Robadey 027]
Length = 335
Score = 303 bits (776), Expect = 5e-81
Identities = 158/337 (46%), Positives = 216/337 (63%), Gaps = 7/337 (2%)
Query: 13 DGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPSCLKVADLGCSSGPNALMVASNI 72
+GD SYANNS QR V+L TK I E +I LY + FP L VADLGCSSG N +V + +
Sbjct: 1 NGDISYANNSLVQRKVILMTKPITEQAISDLYSSLFPETLYVADLGCSSGANTFLVVTEL 60
Query: 73 INTVDAVSQNLNLEQPVFQFFLNDLFGNDFNTTFKSLPDFYKRLQQEKGHKFGPCFFSGT 132
+ ++ + NL+ P F F NDL GNDFNT F+SL F + L+++ G FG CFFSG
Sbjct: 61 VKIIEKERKKHNLQSPEFHFNFNDLPGNDFNTIFQSLGKFQQDLRKQIGEGFGSCFFSGV 120
Query: 133 PGSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPLNKGAIYLTKTSPPAVHKAYFAQFRE 192
GSFY RLFP+ S+HF HSSYSL WLS+V NKG IY+ TSPP+V KAY+ Q+ +
Sbjct: 121 AGSFYNRLFPSKSLHFVHSSYSLMWLSQVPNLSEKNKGNIYIASTSPPSVIKAYYKQYEK 180
Query: 193 DFKLFLGSRSCELLPGGAMVITLIGRD----ENNELINAWVVIGMVLNDMASENLIEKAK 248
DF FL RS EL+ GG MV+T +GR+ + E W ++ + LN++ E LIE+ K
Sbjct: 181 DFSNFLKYRSEELMKGGKMVLTFLGRESEDPSSKECCCIWELLSIALNELVVEGLIEEEK 240
Query: 249 LDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKNINVSDDIDEDTRAEAVAKY 308
+++FNIP Y P+ E++ ++E+EGSF I +LET R W N S+D + V+K
Sbjct: 241 VNSFNIPQYTPSPGEVKYIVEKEGSFIINQLETTRVHW---NNASNDHENINGGYNVSKC 297
Query: 309 IRAVAESILKSEFGEAIMDELFRRFKNKIVKLHGVEK 345
+RAVAE +L S+FG MD +F+++++ I G EK
Sbjct: 298 MRAVAEPLLVSQFGPKFMDLVFQKYEDNISDCMGKEK 334
>dbj|BAC75665.1| Xanthosine N-methyltransferase [Coffea arabica]
Length = 385
Score = 303 bits (775), Expect = 7e-81
Identities = 165/378 (43%), Positives = 225/378 (58%), Gaps = 21/378 (5%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPS---CLKVADL 57
M +++VL MNGG+GD SYA NS++ + V+ K +LE + L A P+ C+KVADL
Sbjct: 1 MELQEVLRMNGGEGDTSYAKNSAYNQLVLAKVKPVLEQCVRELLRANLPNINKCIKVADL 60
Query: 58 GCSSGPNALMVASNIINTVDAVSQNLN--LEQPVFQFFLNDLFGNDFNTTFKSLPDFYKR 115
GC+SGPN L+ +I+ ++D V Q LE+P Q FLNDLF NDFN+ FK LP FY++
Sbjct: 61 GCASGPNTLLTVRDIVQSIDKVGQEKKNELERPTIQIFLNDLFPNDFNSVFKLLPSFYRK 120
Query: 116 LQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLNK 169
L++E G K G C PGSFY RLFP +S+HF HS Y L WLS+V NK
Sbjct: 121 LEKENGRKIGSCLIGAMPGSFYSRLFPEESMHFLHSCYCLQWLSQVPSGLVTELGISTNK 180
Query: 170 GAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAWV 229
G+IY +K S V KAY QF +DF FL S EL G M++T I + + NA
Sbjct: 181 GSIYSSKASRLPVQKAYLDQFTKDFTTFLRIHSEELFSHGRMLLTCICKGVELDARNAID 240
Query: 230 VIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKN 289
++ M +ND+ E +E+ KLD+FN+P Y P+AEE++ ++EEEGSF+I LET + +
Sbjct: 241 LLEMAINDLVVEGHLEEEKLDSFNLPVYIPSAEEVKCIVEEEGSFEILYLETFKVLYDAG 300
Query: 290 INVSDDI----------DEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVK 339
++ DD DE +AE VA IR+V E IL S FGEAIM +LF R K
Sbjct: 301 FSIDDDYPVRSHFQVYGDEHIKAEYVASLIRSVYEPILASHFGEAIMPDLFHRLAKHAAK 360
Query: 340 LHGVEKLELANLVMHITK 357
+ + K NL++ + K
Sbjct: 361 VLHLGKGFYNNLIISLAK 378
>dbj|BAC43759.1| tentative caffeine synthase 4 [Coffea arabica]
Length = 385
Score = 303 bits (775), Expect = 7e-81
Identities = 163/378 (43%), Positives = 230/378 (60%), Gaps = 21/378 (5%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPS---CLKVADL 57
M +++VLHMNGG+G+ASYA NSSF + V+ K +LE + L A P+ C+KVADL
Sbjct: 1 MELQEVLHMNGGEGEASYAKNSSFNQLVLAKVKPVLEQCVRELLRANLPNINKCIKVADL 60
Query: 58 GCSSGPNALMVASNIINTVDAVSQNLN--LEQPVFQFFLNDLFGNDFNTTFKSLPDFYKR 115
GC+SGPN L+ + + ++D V Q + LE+P Q FL DLF NDFN+ F LP FY++
Sbjct: 61 GCASGPNTLLTVWDTVQSIDKVRQEMKNELERPTIQVFLTDLFQNDFNSVFMLLPSFYRK 120
Query: 116 LQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLNK 169
L++E G K G C + PGSF+GRLFP +S+HF HSSYSL +LS+V NK
Sbjct: 121 LEKENGRKIGSCLIAAMPGSFHGRLFPEESMHFLHSSYSLQFLSQVPSGLVTELGITANK 180
Query: 170 GAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAWV 229
+IY +K SPP V KAY QF +DF FL RS ELL G M++T I + + + N
Sbjct: 181 RSIYSSKASPPPVQKAYLDQFTKDFTTFLRMRSEELLSRGRMLLTCICKGDECDGPNTMD 240
Query: 230 VIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKN 289
++ M +ND+ +E + + KLD+FN+P Y + EE++ ++EEEGSF+I L+T + +
Sbjct: 241 LLEMAINDLVAEGRLGEEKLDSFNVPIYTASVEEVKCMVEEEGSFEILYLQTFKLRYDAG 300
Query: 290 INVSDDI----------DEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVK 339
++ DD DE RA VA IR+V E IL S FGEAI+ ++F RF K
Sbjct: 301 FSIDDDCQVRSHSPVYSDEHARAAHVASLIRSVYEPILASHFGEAIIPDIFHRFATNAAK 360
Query: 340 LHGVEKLELANLVMHITK 357
+ + K NL++ + K
Sbjct: 361 VIRLGKGFYNNLIISLAK 378
>dbj|BAC43761.1| tentative caffeine synthase 7 [Coffea arabica]
Length = 384
Score = 303 bits (775), Expect = 7e-81
Identities = 161/361 (44%), Positives = 219/361 (60%), Gaps = 22/361 (6%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPS---CLKVADL 57
M +++VLHMNGG+GD SYA NS F ++ K ILE I L A P+ C+KVADL
Sbjct: 1 MELQEVLHMNGGEGDTSYAKNS-FYNLFLIRVKPILEQCIQELLRANLPNINKCIKVADL 59
Query: 58 GCSSGPNALMVASNIINTVDAVSQNLN--LEQPVFQFFLNDLFGNDFNTTFKSLPDFYKR 115
GC+SGPN L+ +I+ ++D V Q LE+P Q FLNDLF NDFN+ FKSLP FY++
Sbjct: 60 GCASGPNTLLTVRDIVQSIDKVGQEKKNELERPTIQIFLNDLFQNDFNSVFKSLPSFYRK 119
Query: 116 LQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKV------NYTKPLNK 169
L++E G K G C PGSFYGRLFP +S+HF HS Y LHWLS+V NK
Sbjct: 120 LEKENGCKIGSCLIGAMPGSFYGRLFPEESMHFLHSCYCLHWLSQVPSGLVTELGISANK 179
Query: 170 GAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAWV 229
G IY +K S P + KAY QF +DF FL S EL+ G M++T I +++ E N+
Sbjct: 180 GCIYSSKASRPPIQKAYLDQFTKDFTTFLRIHSEELISRGRMLLTWICKEDEFENPNSID 239
Query: 230 VIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKN 289
++ M +ND+ E +E+ KLD+FN+P Y P+ EE++ ++EEEGSF+I LET + +
Sbjct: 240 LLEMSINDLVIEGHLEEEKLDSFNVPIYAPSTEEVKCIVEEEGSFEILYLETFKVPYDAG 299
Query: 290 INVSDD----------IDEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVK 339
++ DD DE RA VA +R++ E I+ S FGEAI+ +L R K
Sbjct: 300 FSIDDDYQGRSHSPVSCDEHARAAHVASVVRSIFEPIVASHFGEAILPDLSHRIAKNAAK 359
Query: 340 L 340
+
Sbjct: 360 V 360
>dbj|BAC75664.1| 7-methylxanthine N-methyltransferase [Coffea arabica]
Length = 384
Score = 302 bits (774), Expect = 9e-81
Identities = 166/378 (43%), Positives = 227/378 (59%), Gaps = 22/378 (5%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPS---CLKVADL 57
M +++VLHMN G+GD SYA N+S+ + K LE I L A P+ C+KVADL
Sbjct: 1 MELQEVLHMNEGEGDTSYAKNASYNL-ALAKVKPFLEQCIRELLRANLPNINKCIKVADL 59
Query: 58 GCSSGPNALMVASNIINTVDAVSQNLN--LEQPVFQFFLNDLFGNDFNTTFKSLPDFYKR 115
GC+SGPN L+ +I+ ++D V Q LE+P Q FLNDLF NDFN+ FK LP FY++
Sbjct: 60 GCASGPNTLLTVRDIVQSIDKVGQEEKNELERPTIQIFLNDLFQNDFNSVFKLLPSFYRK 119
Query: 116 LQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPL------NK 169
L++E G K G C S PGSFYGRLFP +S+HF HS YS+HWLS+V + NK
Sbjct: 120 LEKENGRKIGSCLISAMPGSFYGRLFPEESMHFLHSCYSVHWLSQVPSGLVIELGIGANK 179
Query: 170 GAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAWV 229
G+IY +K S P V KAY QF +DF FL S EL G M++T I + + + N
Sbjct: 180 GSIYSSKASRPPVQKAYLDQFTKDFTTFLRIHSKELFSRGRMLLTCICKVDEYDEPNPLD 239
Query: 230 VIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKN 289
++ M +ND+ E +E+ KL +FN+P + P+AEE++ ++EEEGSF+I LET + +
Sbjct: 240 LLDMAINDLIVEGHLEEEKLASFNLPFFTPSAEEVKCIVEEEGSFEILYLETFKAHYDAG 299
Query: 290 INVSDDI----------DEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVK 339
++ DD DE +AE VA IR+V E IL S FGEAIM +LF R K
Sbjct: 300 FSIDDDYPVRSHFQVYGDEHIKAEYVASLIRSVYEPILASHFGEAIMPDLFHRLAKHAAK 359
Query: 340 LHGVEKLELANLVMHITK 357
+ + K NL++ + K
Sbjct: 360 VLHLGKGCYNNLIISLAK 377
>gb|AAQ16155.1| putative caffeine synthase [Coffea canephora]
gi|20271024|gb|AAM18504.1| N-methyltransferase [Coffea
canephora] gi|20271018|gb|AAM18501.1|
N-methyltransferase [Coffea arabica]
gi|13365753|dbj|BAB39216.1| 7-methylxanthine
N-methyltransferase [Coffea arabica]
Length = 378
Score = 302 bits (773), Expect = 1e-80
Identities = 166/372 (44%), Positives = 226/372 (60%), Gaps = 16/372 (4%)
Query: 1 MAIEQVLHMNGGDGDASYANNSSFQRNVMLTTKHILEDSIIRLYCATFPS---CLKVADL 57
M +++VLHMN G+GD SYA N+S+ + K LE I L A P+ C+KVADL
Sbjct: 1 MELQEVLHMNEGEGDTSYAKNASYNL-ALAKVKPFLEQCIRELLRANLPNINKCIKVADL 59
Query: 58 GCSSGPNALMVASNIINTVDAVSQNLN--LEQPVFQFFLNDLFGNDFNTTFKSLPDFYKR 115
GC+SGPN L+ +I+ ++D V Q LE+P Q FLNDLF NDFN+ FK LP FY++
Sbjct: 60 GCASGPNTLLTVRDIVQSIDKVGQEEKNELERPTIQIFLNDLFQNDFNSVFKLLPSFYRK 119
Query: 116 LQQEKGHKFGPCFFSGTPGSFYGRLFPNDSVHFFHSSYSLHWLSKVNYTKPL------NK 169
L++E G K G C S PGSFYGRLFP +S+HF HS YS+HWLS+V + NK
Sbjct: 120 LEKENGRKIGSCLISAMPGSFYGRLFPEESMHFLHSCYSVHWLSQVPSGLVIELGIGANK 179
Query: 170 GAIYLTKTSPPAVHKAYFAQFREDFKLFLGSRSCELLPGGAMVITLIGRDENNELINAWV 229
G+IY +K P V KAY QF +DF FL S EL G M++T I + + + N
Sbjct: 180 GSIYSSKGCRPPVQKAYLDQFTKDFTTFLRIHSKELFSRGRMLLTCICKVDEFDEPNPLD 239
Query: 230 VIGMVLNDMASENLIEKAKLDAFNIPSYCPTAEEIRQVIEEEGSFDIQRLETIRTDWIKN 289
++ M +ND+ E L+E+ KLD+FNIP + P+AEE++ ++EEEGS +I LET + +
Sbjct: 240 LLDMAINDLIVEGLLEEEKLDSFNIPFFTPSAEEVKCIVEEEGSCEILYLETFKAHYDAA 299
Query: 290 INVSDDI----DEDTRAEAVAKYIRAVAESILKSEFGEAIMDELFRRFKNKIVKLHGVEK 345
++ DD E +AE VA IR+V E IL S FGEAIM +LF R K+ + K
Sbjct: 300 FSIDDDYPVRSHEQIKAEYVASLIRSVYEPILASHFGEAIMPDLFHRLAKHAAKVLHMGK 359
Query: 346 LELANLVMHITK 357
NL++ + K
Sbjct: 360 GCYNNLIISLAK 371
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.137 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 610,410,908
Number of Sequences: 2540612
Number of extensions: 25683388
Number of successful extensions: 58441
Number of sequences better than 10.0: 179
Number of HSP's better than 10.0 without gapping: 162
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 57649
Number of HSP's gapped (non-prelim): 185
length of query: 359
length of database: 863,360,394
effective HSP length: 129
effective length of query: 230
effective length of database: 535,621,446
effective search space: 123192932580
effective search space used: 123192932580
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0113.13


