

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111a.4
(74 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
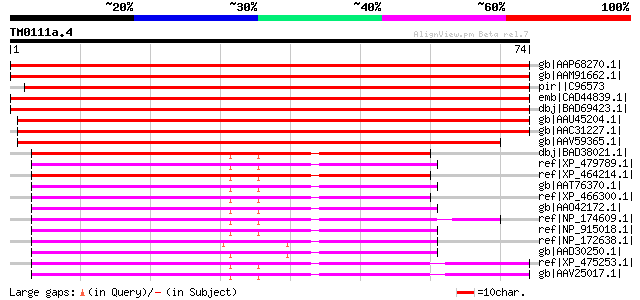
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP68270.1| At1g53290 [Arabidopsis thaliana] gi|22135994|gb|A... 137 5e-32
gb|AAM91662.1| putative galactosyltransferase [Arabidopsis thali... 137 5e-32
pir||C96573 protein F12M16.19 [imported] - Arabidopsis thaliana ... 133 1e-30
emb|CAD44839.1| beta 1,3-glycosyltransferase-like protein III [O... 129 2e-29
dbj|BAD69423.1| putative Avr9 elicitor response protein [Oryza s... 129 2e-29
gb|AAU45204.1| At2g26100 [Arabidopsis thaliana] gi|51536430|gb|A... 112 2e-24
gb|AAC31227.1| unknown protein [Arabidopsis thaliana] gi|7487205... 112 2e-24
gb|AAV59365.1| 'putative galactosyl transferase, PF01762' [Oryza... 89 4e-17
dbj|BAD38021.1| putative Avr9 elicitor response protein [Oryza s... 50 2e-05
ref|XP_479789.1| putative avr9 elicitor response protein [Oryza ... 50 2e-05
ref|XP_464214.1| putative Avr9 elicitor response protein [Oryza ... 49 4e-05
gb|AAT76370.1| putative glycosyltransferase [Oryza sativa (japon... 48 7e-05
ref|XP_466300.1| putative avr9 elicitor response protein [Oryza ... 47 2e-04
gb|AAO42172.1| unknown protein [Arabidopsis thaliana] gi|3831453... 47 2e-04
ref|NP_174609.1| galactosyltransferase family protein [Arabidops... 47 2e-04
ref|NP_915018.1| putative elicitor response protein [Oryza sativ... 46 2e-04
ref|NP_172638.1| galactosyltransferase family protein [Arabidops... 46 3e-04
gb|AAD30250.1| Strong similarity to gb|AJ006228 Avr9 elicitor re... 46 3e-04
ref|XP_475253.1| putative galactosyltransferase [Oryza sativa (j... 45 4e-04
gb|AAV25017.1| putative galactosyltransferase [Oryza sativa (jap... 45 4e-04
>gb|AAP68270.1| At1g53290 [Arabidopsis thaliana] gi|22135994|gb|AAM91579.1| unknown
protein [Arabidopsis thaliana]
gi|30695469|ref|NP_175736.2| galactosyltransferase
family protein [Arabidopsis thaliana]
Length = 345
Score = 137 bits (346), Expect = 5e-32
Identities = 60/74 (81%), Positives = 68/74 (91%)
Query: 1 FRMFSNEDVTIGAWMLAMNVNHENNHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELH 60
FRMF+NEDVTIGAWMLAMNVNHEN+H LC PEC+ +S+AVWDIPKCSGLCNPEKRMLELH
Sbjct: 270 FRMFNNEDVTIGAWMLAMNVNHENHHILCEPECSPSSVAVWDIPKCSGLCNPEKRMLELH 329
Query: 61 QMESCTQSPTAESD 74
+ ESC++SPT SD
Sbjct: 330 KQESCSKSPTLPSD 343
>gb|AAM91662.1| putative galactosyltransferase [Arabidopsis thaliana]
gi|17979337|gb|AAL49894.1| putative
galactosyltransferase [Arabidopsis thaliana]
gi|8777479|dbj|BAA97059.1| unnamed protein product
[Arabidopsis thaliana] gi|15232447|ref|NP_188114.1|
galactosyltransferase family protein [Arabidopsis
thaliana]
Length = 343
Score = 137 bits (346), Expect = 5e-32
Identities = 62/74 (83%), Positives = 66/74 (88%)
Query: 1 FRMFSNEDVTIGAWMLAMNVNHENNHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELH 60
FRMFSNEDVTIGAWMLAMNVNHEN H LC PEC+ SIAVWDIPKCSGLCNPEKRMLELH
Sbjct: 268 FRMFSNEDVTIGAWMLAMNVNHENLHTLCEPECSPYSIAVWDIPKCSGLCNPEKRMLELH 327
Query: 61 QMESCTQSPTAESD 74
+ESC++SPT SD
Sbjct: 328 MLESCSKSPTLPSD 341
>pir||C96573 protein F12M16.19 [imported] - Arabidopsis thaliana
gi|7769857|gb|AAF69535.1| F12M16.19 [Arabidopsis
thaliana]
Length = 353
Score = 133 bits (335), Expect = 1e-30
Identities = 58/72 (80%), Positives = 66/72 (91%)
Query: 3 MFSNEDVTIGAWMLAMNVNHENNHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELHQM 62
MF+NEDVTIGAWMLAMNVNHEN+H LC PEC+ +S+AVWDIPKCSGLCNPEKRMLELH+
Sbjct: 280 MFNNEDVTIGAWMLAMNVNHENHHILCEPECSPSSVAVWDIPKCSGLCNPEKRMLELHKQ 339
Query: 63 ESCTQSPTAESD 74
ESC++SPT SD
Sbjct: 340 ESCSKSPTLPSD 351
>emb|CAD44839.1| beta 1,3-glycosyltransferase-like protein III [Oryza sativa]
Length = 207
Score = 129 bits (324), Expect = 2e-29
Identities = 56/74 (75%), Positives = 65/74 (87%)
Query: 1 FRMFSNEDVTIGAWMLAMNVNHENNHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELH 60
FRMFSNEDVTIG+WMLAMNVNHEN H LC+PECT +SIAVWDIPKCSGLC+PE +MLELH
Sbjct: 130 FRMFSNEDVTIGSWMLAMNVNHENTHALCSPECTESSIAVWDIPKCSGLCHPEVKMLELH 189
Query: 61 QMESCTQSPTAESD 74
+ + CT P+A S+
Sbjct: 190 RRKECTGGPSAVSE 203
>dbj|BAD69423.1| putative Avr9 elicitor response protein [Oryza sativa (japonica
cultivar-group)]
Length = 368
Score = 129 bits (324), Expect = 2e-29
Identities = 56/74 (75%), Positives = 65/74 (87%)
Query: 1 FRMFSNEDVTIGAWMLAMNVNHENNHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELH 60
FRMFSNEDVTIG+WMLAMNVNHEN H LC+PECT +SIAVWDIPKCSGLC+PE +MLELH
Sbjct: 291 FRMFSNEDVTIGSWMLAMNVNHENTHALCSPECTESSIAVWDIPKCSGLCHPEVKMLELH 350
Query: 61 QMESCTQSPTAESD 74
+ + CT P+A S+
Sbjct: 351 RRKECTGGPSAVSE 364
>gb|AAU45204.1| At2g26100 [Arabidopsis thaliana] gi|51536430|gb|AAU05453.1|
At2g26100 [Arabidopsis thaliana]
gi|30683005|ref|NP_180179.2| galactosyltransferase
family protein [Arabidopsis thaliana]
Length = 371
Score = 112 bits (281), Expect = 2e-24
Identities = 48/73 (65%), Positives = 60/73 (81%)
Query: 2 RMFSNEDVTIGAWMLAMNVNHENNHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELHQ 61
RMF+NEDVTIG+WMLAM+V+HE+N LC P C+ SIAVWDIPKCSGLC+PE R+ ELH+
Sbjct: 295 RMFNNEDVTIGSWMLAMDVHHEDNRALCDPHCSPKSIAVWDIPKCSGLCDPESRLKELHK 354
Query: 62 MESCTQSPTAESD 74
+ C++SPT D
Sbjct: 355 TDMCSKSPTLPPD 367
>gb|AAC31227.1| unknown protein [Arabidopsis thaliana] gi|7487205|pir||T02614
hypothetical protein At2g26100 [imported] - Arabidopsis
thaliana
Length = 333
Score = 112 bits (281), Expect = 2e-24
Identities = 48/73 (65%), Positives = 60/73 (81%)
Query: 2 RMFSNEDVTIGAWMLAMNVNHENNHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELHQ 61
RMF+NEDVTIG+WMLAM+V+HE+N LC P C+ SIAVWDIPKCSGLC+PE R+ ELH+
Sbjct: 257 RMFNNEDVTIGSWMLAMDVHHEDNRALCDPHCSPKSIAVWDIPKCSGLCDPESRLKELHK 316
Query: 62 MESCTQSPTAESD 74
+ C++SPT D
Sbjct: 317 TDMCSKSPTLPPD 329
>gb|AAV59365.1| 'putative galactosyl transferase, PF01762' [Oryza sativa (japonica
cultivar-group)] gi|50933149|ref|XP_476102.1| 'putative
galactosyl transferase, PF01762' [Oryza sativa (japonica
cultivar-group)] gi|53981367|gb|AAV24921.1| unknown
protein [Oryza sativa (japonica cultivar-group)]
Length = 390
Score = 88.6 bits (218), Expect = 4e-17
Identities = 40/69 (57%), Positives = 47/69 (67%)
Query: 2 RMFSNEDVTIGAWMLAMNVNHENNHELCAPECTSTSIAVWDIPKCSGLCNPEKRMLELHQ 61
RMF EDVTIG+WMLAMNV HE+N +C CT TSIAVWD KCS CN + + LH
Sbjct: 314 RMFDYEDVTIGSWMLAMNVKHEDNRAMCDSACTPTSIAVWDSKKCSNSCNTTEIVKALHN 373
Query: 62 MESCTQSPT 70
C++SPT
Sbjct: 374 TTLCSKSPT 382
>dbj|BAD38021.1| putative Avr9 elicitor response protein [Oryza sativa (japonica
cultivar-group)]
Length = 393
Score = 50.1 bits (118), Expect = 2e-05
Identities = 23/66 (34%), Positives = 42/66 (62%), Gaps = 10/66 (15%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA---PECT------STSIAVWDIPKCSGLCNPEK 54
++NEDV++GAW + ++V H ++ ++C P+C + +A +D KCSG+CNP +
Sbjct: 314 YANEDVSLGAWFIGLDVEHIDDRDMCCGTPPDCEWKAQAGNICVASFDW-KCSGVCNPVE 372
Query: 55 RMLELH 60
R+ +H
Sbjct: 373 RLKYVH 378
>ref|XP_479789.1| putative avr9 elicitor response protein [Oryza sativa (japonica
cultivar-group)] gi|51964628|ref|XP_507098.1| PREDICTED
P0470F10.14 gene product [Oryza sativa (japonica
cultivar-group)] gi|50725628|dbj|BAD33095.1| putative
avr9 elicitor response protein [Oryza sativa (japonica
cultivar-group)]
Length = 388
Score = 49.7 bits (117), Expect = 2e-05
Identities = 24/67 (35%), Positives = 40/67 (58%), Gaps = 10/67 (14%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA---PECT------STSIAVWDIPKCSGLCNPEK 54
+SNEDV+ G+W++ ++V H + LC P+C + A +D C+G+CNP +
Sbjct: 309 YSNEDVSFGSWLIGLDVEHVDERSLCCGTPPDCEWKAQAGNPCAASFDW-NCTGICNPVE 367
Query: 55 RMLELHQ 61
RM E+H+
Sbjct: 368 RMEEVHR 374
>ref|XP_464214.1| putative Avr9 elicitor response protein [Oryza sativa (japonica
cultivar-group)] gi|49388048|dbj|BAD25162.1| putative
Avr9 elicitor response protein [Oryza sativa (japonica
cultivar-group)]
Length = 400
Score = 48.5 bits (114), Expect = 4e-05
Identities = 24/66 (36%), Positives = 40/66 (60%), Gaps = 10/66 (15%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA---PECT------STSIAVWDIPKCSGLCNPEK 54
+ NEDV++G+W + ++V H ++ LC P+C +T A +D +CSG+CN E
Sbjct: 321 YINEDVSLGSWFIGLDVEHIDDRRLCCGTPPDCEWKAQAGNTCAASFDW-RCSGICNSEG 379
Query: 55 RMLELH 60
R+ E+H
Sbjct: 380 RIWEVH 385
>gb|AAT76370.1| putative glycosyltransferase [Oryza sativa (japonica
cultivar-group)]
Length = 406
Score = 47.8 bits (112), Expect = 7e-05
Identities = 24/67 (35%), Positives = 38/67 (55%), Gaps = 10/67 (14%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA---PECT------STSIAVWDIPKCSGLCNPEK 54
++NEDV++GAW + ++V H ++ LC P+C + A +D CSG+C
Sbjct: 327 YANEDVSLGAWFIGLDVEHVDDRRLCCGTQPDCEWKAQAGNVCAASFDW-SCSGICKSAD 385
Query: 55 RMLELHQ 61
RM E+HQ
Sbjct: 386 RMKEVHQ 392
>ref|XP_466300.1| putative avr9 elicitor response protein [Oryza sativa (japonica
cultivar-group)] gi|46806681|dbj|BAD17751.1| putative
avr9 elicitor response protein [Oryza sativa (japonica
cultivar-group)]
Length = 400
Score = 46.6 bits (109), Expect = 2e-04
Identities = 22/66 (33%), Positives = 38/66 (57%), Gaps = 10/66 (15%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA---PECT------STSIAVWDIPKCSGLCNPEK 54
F+NEDV++G+W + + VNH + +C P+C + +A +D CSG+C +
Sbjct: 321 FANEDVSLGSWFIGLEVNHIDERNMCCGTPPDCEWKGQAGNVCVASFDW-SCSGICKSVE 379
Query: 55 RMLELH 60
R+ E+H
Sbjct: 380 RIKEVH 385
>gb|AAO42172.1| unknown protein [Arabidopsis thaliana] gi|3831453|gb|AAC69935.1|
unknown protein [Arabidopsis thaliana]
gi|15225684|ref|NP_180802.1| galactosyltransferase
family protein [Arabidopsis thaliana]
gi|25370616|pir||A84733 hypothetical protein At2g32430
[imported] - Arabidopsis thaliana
Length = 409
Score = 46.6 bits (109), Expect = 2e-04
Identities = 23/67 (34%), Positives = 39/67 (57%), Gaps = 10/67 (14%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA---PECT------STSIAVWDIPKCSGLCNPEK 54
++NEDVT+GAW + ++V H ++ LC P+C + +A +D CSG+C
Sbjct: 330 YANEDVTLGAWFIGLDVTHIDDRRLCCGTPPDCEWKAQAGNICVASFDW-TCSGICRSAD 388
Query: 55 RMLELHQ 61
R+ E+H+
Sbjct: 389 RIKEVHK 395
>ref|NP_174609.1| galactosyltransferase family protein [Arabidopsis thaliana]
gi|12322375|gb|AAG51207.1| elicitor response protein,
putative; 49810-48196 [Arabidopsis thaliana]
gi|25370623|pir||A86458 probasble elicitor response
protein - Arabidopsis thaliana
Length = 395
Score = 46.6 bits (109), Expect = 2e-04
Identities = 25/76 (32%), Positives = 40/76 (51%), Gaps = 12/76 (15%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA---PECT------STSIAVWDIPKCSGLCNPEK 54
++NEDV++GAWML + V H + +C P+C + A +D CSG+C
Sbjct: 314 YANEDVSLGAWMLGLEVEHVDERSMCCGTPPDCQWKAQAGNVCAASFDW-SCSGICKSVD 372
Query: 55 RMLELHQMESCTQSPT 70
RM +H+ +C + T
Sbjct: 373 RMARVHR--ACAEGDT 386
>ref|NP_915018.1| putative elicitor response protein [Oryza sativa (japonica
cultivar-group)] gi|22202663|dbj|BAC07321.1| putative
Avr9 elicitor response protein [Oryza sativa (japonica
cultivar-group)]
Length = 408
Score = 46.2 bits (108), Expect = 2e-04
Identities = 22/67 (32%), Positives = 38/67 (55%), Gaps = 10/67 (14%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA---PECT------STSIAVWDIPKCSGLCNPEK 54
F+NEDV++GAW++ + V H ++ +C P+C + +A +D CSG+C
Sbjct: 329 FANEDVSLGAWLIGLEVEHVDDRSMCCATPPDCEWKKRAGNVCVASFDW-SCSGVCKSVD 387
Query: 55 RMLELHQ 61
RM +H+
Sbjct: 388 RMKHIHR 394
>ref|NP_172638.1| galactosyltransferase family protein [Arabidopsis thaliana]
Length = 384
Score = 45.8 bits (107), Expect = 3e-04
Identities = 22/67 (32%), Positives = 40/67 (58%), Gaps = 10/67 (14%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELC---APECTSTSI------AVWDIPKCSGLCNPEK 54
++NEDV++G+W + +NV H + LC + +C ++ A +D KCSG+C +
Sbjct: 305 YANEDVSLGSWFIGLNVEHVDEKRLCCSTSQDCELKAMMGHVCAASFDW-KCSGICRSAE 363
Query: 55 RMLELHQ 61
RM ++H+
Sbjct: 364 RMADVHE 370
>gb|AAD30250.1| Strong similarity to gb|AJ006228 Avr9 elicitor response protein
from Nicotiana tabacum. EST gb|F15429 comes from this
gene. [Arabidopsis thaliana] gi|25370620|pir||A86251
hypothetical protein [imported] - Arabidopsis thaliana
Length = 401
Score = 45.8 bits (107), Expect = 3e-04
Identities = 23/74 (31%), Positives = 40/74 (53%), Gaps = 17/74 (22%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA----------PECTSTSI------AVWDIPKCS 47
++NEDV++G+W + +NV H + LC P+C ++ A +D KCS
Sbjct: 315 YANEDVSLGSWFIGLNVEHVDEKRLCCSTSQGKELNNPDCELKAMMGHVCAASFDW-KCS 373
Query: 48 GLCNPEKRMLELHQ 61
G+C +RM ++H+
Sbjct: 374 GICRSAERMADVHE 387
>ref|XP_475253.1| putative galactosyltransferase [Oryza sativa (japonica
cultivar-group)] gi|46391132|gb|AAS90659.1| putative
galactosyltransferase [Oryza sativa (japonica
cultivar-group)]
Length = 534
Score = 45.4 bits (106), Expect = 4e-04
Identities = 26/80 (32%), Positives = 42/80 (52%), Gaps = 12/80 (15%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA---PECT------STSIAVWDIPKCSGLCNPEK 54
F+NEDV++GAW++ + V H ++ LC P+C + A +D CSG+C
Sbjct: 450 FANEDVSLGAWLIGLEVEHVDDRSLCCATPPDCEWKKQAGNVCAASFDW-SCSGICKSVD 508
Query: 55 RMLELHQMESCTQSPTAESD 74
RM +H +C + A S+
Sbjct: 509 RMRAIH--SACGEGDGAVSN 526
>gb|AAV25017.1| putative galactosyltransferase [Oryza sativa (japonica
cultivar-group)]
Length = 416
Score = 45.4 bits (106), Expect = 4e-04
Identities = 26/80 (32%), Positives = 42/80 (52%), Gaps = 12/80 (15%)
Query: 4 FSNEDVTIGAWMLAMNVNHENNHELCA---PECT------STSIAVWDIPKCSGLCNPEK 54
F+NEDV++GAW++ + V H ++ LC P+C + A +D CSG+C
Sbjct: 332 FANEDVSLGAWLIGLEVEHVDDRSLCCATPPDCEWKKQAGNVCAASFDW-SCSGICKSVD 390
Query: 55 RMLELHQMESCTQSPTAESD 74
RM +H +C + A S+
Sbjct: 391 RMRAIH--SACGEGDGAVSN 408
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.317 0.127 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 131,192,024
Number of Sequences: 2540612
Number of extensions: 4094737
Number of successful extensions: 7168
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 33
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 7112
Number of HSP's gapped (non-prelim): 44
length of query: 74
length of database: 863,360,394
effective HSP length: 50
effective length of query: 24
effective length of database: 736,329,794
effective search space: 17671915056
effective search space used: 17671915056
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0111a.4


