

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0110a.13
(1389 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
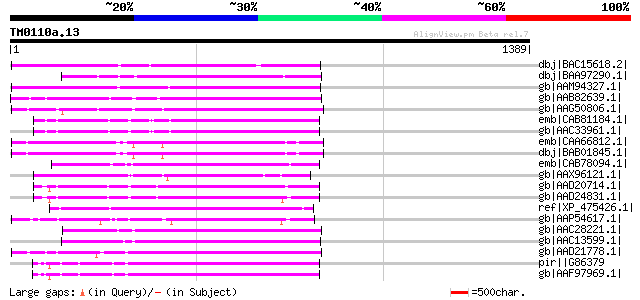
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAC15618.2| unnamed protein product [Oryza sativa (japonica ... 360 2e-97
dbj|BAA97290.1| non-LTR retroelement reverse transcriptase-like ... 359 4e-97
gb|AAM94327.1| putative reverse transcriptase [Sorghum bicolor] 354 1e-95
gb|AAB82639.1| putative non-LTR retroelement reverse transcripta... 352 4e-95
gb|AAG50806.1| unknown protein [Arabidopsis thaliana] gi|2540301... 352 4e-95
emb|CAB81184.1| putative protein [Arabidopsis thaliana] gi|45393... 343 2e-92
gb|AAC33961.1| contains similarity to reverse trancriptase (Pfam... 343 2e-92
emb|CAA66812.1| non-ltr retrotransposon reverse transcriptase-li... 343 3e-92
dbj|BAB01845.1| non-LTR retroelement reverse transcriptase-like ... 342 4e-92
emb|CAB78094.1| RNA-directed DNA polymerase-like protein [Arabid... 340 2e-91
gb|AAX96121.1| retrotransposon protein, putative, unclassified [... 335 5e-90
gb|AAD20714.1| putative non-LTR retroelement reverse transcripta... 335 6e-90
gb|AAD24831.1| putative non-LTR retroelement reverse transcripta... 335 6e-90
ref|XP_475426.1| putative non-LTR retroelement reverse transcrip... 333 2e-89
gb|AAP54617.1| putative non-LTR retroelement reverse transcripta... 326 3e-87
gb|AAC28221.1| similar to reverse transcriptases (PFam: rvt.hmm,... 326 3e-87
gb|AAC13599.1| similar to reverse transcriptase (Pfam: transcrip... 325 9e-87
gb|AAD21778.1| putative non-LTR retroelement reverse transcripta... 323 2e-86
pir||G86379 protein F5A9.24 [imported] - Arabidopsis thaliana gi... 323 2e-86
gb|AAF97969.1| F21J9.30 [Arabidopsis thaliana] 323 2e-86
>dbj|BAC15618.2| unnamed protein product [Oryza sativa (japonica cultivar-group)]
Length = 1197
Score = 360 bits (924), Expect = 2e-97
Identities = 256/839 (30%), Positives = 414/839 (48%), Gaps = 28/839 (3%)
Query: 6 WNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKLDSSRVKTVEAFANAVKMDYVSVDAVG 65
WNVRGLG K V ++S P VL+QE+KL + + + +Y V + G
Sbjct: 10 WNVRGLGDKNKCDIVKNSISDCNPSFVLLQETKLQNIPPEKAKTILPPNLANYDFVGSYG 69
Query: 66 SAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYDFFNLL 125
S GG+++ WN A + S + + + + + N+YGP ++ DF + L
Sbjct: 70 SRGGMLTAWN-ADFTKTSFISRHFSLTISFTSTFTDLSFSLTNIYGPAELDQKQDFLHEL 128
Query: 126 SEALSLNNCFCKIMGGDFNAILNEGERIG--IQMGVESSFSDFVADSNLIDLPLQSDKFT 183
E L +++ GDFN + ++ G+ ++F+ + +L +LPL +T
Sbjct: 129 LEIDPLIQG-PRLLIGDFNLTRSPSDKNSQNFSTGLAAAFNQTINSLSLFELPLPDRLYT 187
Query: 184 WYSSRAGGLWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSLGAINFGPKPFLF 243
W + R + +R+DR + N S S SD P++L+ K F F
Sbjct: 188 WSNKREVPVLARLDRAFFNNAWNNTLPNTSLSSCTRTTSDHVPLLLTASTSIPKSKTFRF 247
Query: 244 YNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVSRVRINTI 303
NH LLD++F V W + A L KLK ++ IK W + +++
Sbjct: 248 ENHLLLDRDFLPSVMHRWPQQLNMNDPAACLAAKLKQTRSIIKVWMK-----EHRKRSSL 302
Query: 304 EADLQVTMDRLASEEPGIELRKN----RLTILEGLWQEYRKEEYRWRQKARVRWLREGDR 359
D + T+D L E L R + E L+Q + + W+Q+ +V+ L+ G
Sbjct: 303 NEDCKFTIDLLDYIEELRPLNDGERCLRNLVQEKLFQAIQSKVAYWKQRGKVKRLKIGID 362
Query: 360 NTSFF--HSISKIRKSKKLISQLVVGGVSIEDPNMIKDAIFSHFQSFFTSENRLRPKIRC 417
N++FF H+ R S+ I + G I D N + S FQ F S+ +
Sbjct: 363 NSAFFKAHASKNYRNSR--IRSIKHNGAEIADHNGKALLLHSFFQHFLGSDCQPLWDFDL 420
Query: 418 ANLSRLSQANKMSLEKQ---FSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDV 474
+ L + + L Q FS EI +K + N APG DGF +F+ W +KED+
Sbjct: 421 STLYSEDERDYNLLASQGLQFSTEEIITSIKGMNSNSAPGPDGFGPSFYNSAWNSIKEDI 480
Query: 475 VKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQV 534
+K F S ++ +N I LIPK +TV +RPISL + K++SKVLA+RLQ
Sbjct: 481 IKLAASFCSNSVQLERINRAHIVLIPKPGKENTVDGYRPISLQNCSVKILSKVLANRLQK 540
Query: 535 VTPLLISENQFAFAQGRQISDCILIANEVVDFLNKKQGGGFLLKLDFAKDYDNIDWGFLF 594
V +I +Q F +GR IS+ ++ A E++ + ++ ++KLDFAK +D++ W LF
Sbjct: 541 VLTRMIHLDQTGFLKGRCISENLIYATELIQACHARKCKTLIIKLDFAKAFDSVLWTSLF 600
Query: 595 DILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNG 654
IL F + WI+WI++ + T+ ++L+NG + +KGLRQGDPLSP+LF L +
Sbjct: 601 QILAVRGFPNNWISWIKSLLQTSKSAVLLNGIPGKWINCKKGLRQGDPLSPYLFILVADV 660
Query: 655 LSCLLNQLLDDSNFSGVR--IGPGLALNHLQFADDTLLFCENDENQIDLLCNTLLSFLFT 712
L LL + NF +R I +Q+ADDTL+ C +E+ + L +TLL F
Sbjct: 661 LQRLL-----EKNFQ-IRHPIYQDRPCATIQYADDTLVICRAEEDDVLALRSTLLQFSKA 714
Query: 713 SGLKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRRASFWEPMLD 772
+GL++N KST+I ++D + ++ + + LP+ YLG PL + + +P++
Sbjct: 715 TGLQINFAKSTMISLHIDRSKESSLSELLQCKLESLPMSYLGLPLSLHKLTNNDLQPIVV 774
Query: 773 KLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKILRSFLQGG 831
K+ W + +S + RL L+ A L SVP++ M+ FK P KV+ ++K R+F G
Sbjct: 775 KVDSFLTGWEASLLSQAERLILVNAVLSSVPVYAMSAFKLPPKVIEAIDKRRRAFFWTG 833
>dbj|BAA97290.1| non-LTR retroelement reverse transcriptase-like [Arabidopsis
thaliana]
Length = 1072
Score = 359 bits (921), Expect = 4e-97
Identities = 236/716 (32%), Positives = 372/716 (50%), Gaps = 28/716 (3%)
Query: 139 MGGDFNAILNEGERIGI-QMGVESSFSDF---VADSNLIDLPLQSDKFTWYS-SRAGGLW 193
M GDFN +L E + ++ DF +++ L DL + + FTW++ S +
Sbjct: 1 MLGDFNQVLLPQEHSNPPSLNIDRRMRDFGSCLSEMELSDLVFKGNSFTWWNKSSIRPIA 60
Query: 194 SRIDRWLVSEDAINKFDGISQSVENWGLSDRRP--IILSLGAINFGPKPFLFYNHWLLDK 251
++DR L ++ N + N SD ++L I+ +PF F+N L ++
Sbjct: 61 KKLDRILANDSWCNLYPSSHGLFGNLDFSDHVSCGVVLEANGIS-AKRPFKFFNFLLKNE 119
Query: 252 EFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVSRVRINTIEA-DLQVT 310
+F +V D W S VG S + + +KLK +K IK++ + S + + T EA +L +T
Sbjct: 120 DFLNVVMDNWFSTNVVGSSMYRVSKKLKAMKKPIKDFSRL--NYSGIELRTKEAHELLIT 177
Query: 311 MDRLASEEPGI-----ELRKNRLTILEGLWQEYRKEEYRWRQKARVRWLREGDRNTSFFH 365
L P + EL R +L EE + Q++RV W EGD NT +FH
Sbjct: 178 CQNLTLANPSVSNAALELEAQRKWVLLSC-----AEESFFHQRSRVSWFAEGDSNTHYFH 232
Query: 366 SISKIRKSKKLISQLV-VGGVSIEDPNMIKDAIFSHFQSFFTS-ENRLRPKIRCANLS-- 421
+ RKS I+ LV G+ I+ I D ++++ S E+ + NL
Sbjct: 233 RMVDSRKSFNTINSLVDSNGLLIDSQQGILDHCVTYYERLLGSIESPFSMEQEDMNLLLT 292
Query: 422 -RLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDVVKFFED 480
R SQ LEK F++ EI KS N+ G DG+++ FF+ W I+ +V+ +
Sbjct: 293 YRCSQDQCSELEKSFTDDEIKAAFKSLPRNKTSGPDGYSVEFFRDTWSIIGPEVLAAIHE 352
Query: 481 FHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPLLI 540
F +G+L+K NAT + LIPK + T+S+FRPIS + + YK+ISK+L SRLQ + +I
Sbjct: 353 FFDSGQLLKQWNATTLVLIPKTSNACTISEFRPISCLNTLYKVISKLLTSRLQGLLSAVI 412
Query: 541 SENQFAFAQGRQISDCILIANEVVDFLNKKQ-GGGFLLKLDFAKDYDNIDWGFLFDILKE 599
+Q AF GR +++ +L+A E+V N+ +LK+D K +D++ W F+ L+
Sbjct: 413 GHSQSAFLPGRSLAENVLLATEMVHGYNRLNISPRGMLKVDLKKAFDSVKWEFVTAALRA 472
Query: 600 MNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLL 659
+ +R+INWI C++T + +I VNG++ GFF KGLRQGDPLSP+LF L + S LL
Sbjct: 473 LAIPERYINWIHQCITTPSFTISVNGATGGFFRSTKGLRQGDPLSPYLFVLAMEVFSKLL 532
Query: 660 NQLLDDSNFSGVRIGPGLALNHLQFADDTLLFCENDENQIDLLCNTLLSFLFTSGLKVNL 719
D L+++HL FADD ++F + + + +C TL F SGLKVN
Sbjct: 533 YSRYDSGYIHYHPKAGDLSISHLMFADDVMIFFDGGSSSMHGICETLDDFADWSGLKVNK 592
Query: 720 HKSTIIGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRRASFWEPMLDKLRRKRR 779
KS + +D+ E + + YG+ G PI YLG PL R + + P+L+KL + R
Sbjct: 593 DKSQLFQAGLDLSE-RITSAAYGFPAGTFPIRYLGLPLMCRKLRIADYGPLLEKLSARLR 651
Query: 780 TWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKILRSFLQGGKLHG 835
+W SK +S +GR L+ + + + F M+ F P + ++E + FL G + G
Sbjct: 652 SWVSKALSFAGRTQLISSVIFGLINFWMSTFLLPKGCIKKIESLCSKFLWAGSIDG 707
>gb|AAM94327.1| putative reverse transcriptase [Sorghum bicolor]
Length = 1998
Score = 354 bits (908), Expect = 1e-95
Identities = 239/837 (28%), Positives = 425/837 (50%), Gaps = 22/837 (2%)
Query: 6 WNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKLDSSRVKTVEAFANAVKMDYVSVDAVG 65
WNVRGL EK + + L +I+ +QE+K DS + + + + VG
Sbjct: 813 WNVRGLNSKEKWNPIRNKIVDLAVDIICLQETKKDSFDIAFLRNICPPDFDRFEYLPLVG 872
Query: 66 SAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYDFFNLL 125
++GG+++ W ++ E N I + + ++ ++ + NVY P D + F N
Sbjct: 873 ASGGILTAWKSKLFSGELIFSNEYAISIQMSSMHNNDSWLLTNVYAPCTDEGKRQFVNWF 932
Query: 126 SEALSLNNCFCKIMGGDFNAILNEGERIGIQMGVESS----FSDFVADSNLIDLPLQSDK 181
+ + ++G DFN + +R + G +SS F+ ++ L ++ LQ K
Sbjct: 933 KNIQTPHEVDWLVVG-DFNLMRKHEDRN--KEGGDSSEMFLFNSAISSLGLNEIILQGRK 989
Query: 182 FTWYSSRAGGLWSRIDRWLVSEDAIN-KFDGISQSVENWGL--SDRRPIILSLGAINFGP 238
+TW + ++ L +ID W + ++ N KF + SV+ + + SD P ++ +
Sbjct: 990 YTWSNMQSNPLLQKID-WTFTSNSWNIKF--LDTSVKGYDMTPSDHAPCLVKVSTKIPKT 1046
Query: 239 KPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVSRV 298
K F F N WL F++++SD W T A + K K L+ +KE Q +S++ +
Sbjct: 1047 KVFRFENCWLRLPGFQQIISDVWNVPTHHLDQAKSITAKFKKLRKALKEKQ---ASMAHL 1103
Query: 299 RINTIEADLQVTMDRLASEEPGIELRK--NRLTILEGLWQEYRKEEYRWRQKARVRWLRE 356
+ + L + L E + + + R + + L + +++ WRQ+ ++W++
Sbjct: 1104 KTVISKVKLVIQFLELLEEYRDLSMLEWNFRAILKKKLTELLEQQKIYWRQRGAIKWIKL 1163
Query: 357 GDRNTSFFHSISKIRKSKKLISQLVVGGVSIEDPNMIKDA-IFSHFQSFFTSENRLRPKI 415
GD T FFH+ + I+ L++QL SI + K+ I+ F+ ++ R I
Sbjct: 1164 GDATTKFFHANATIKMRGNLVNQLQKQDGSILTAHKDKEQHIWEEFKERLGTKEETRFGI 1223
Query: 416 RCANLSRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDVV 475
+ + S + +LE+ F+E EI V+K +++PG DGF+ F K W VK D+
Sbjct: 1224 NPDTILQASN-DLETLEEPFTETEIESVVKLLPNDKSPGPDGFSNEFMKASWQTVKVDIF 1282
Query: 476 KFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVV 535
FH+ ++ +N +FI+LIPK + P +V+D+RPISL+ ++ KLI+K+LA+RLQ+
Sbjct: 1283 NLCHAFHNHNVCLRSINTSFITLIPKSHNPVSVNDYRPISLLNTSMKLITKLLANRLQMK 1342
Query: 536 TPLLISENQFAFAQGRQISDCILIANEVVDFLNKKQGGGFLLKLDFAKDYDNIDWGFLFD 595
L+ +NQ+ F + R I DC+ + E + + + ++KLDF K +D ++ +
Sbjct: 1343 ITDLVHKNQYGFIRQRTIQDCLAWSFEYLHLCHHSRKAIVIIKLDFEKAFDKMNHKAMLT 1402
Query: 596 ILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGL 655
I+K FG +W+NW+++ ++ T ++L+NG+ +G+RQGDPLSP LF L + L
Sbjct: 1403 IMKAKGFGQKWLNWMKSIFTSGTSNVLLNGTPGKTIHCLRGVRQGDPLSPLLFVLAADFL 1462
Query: 656 SCLLNQL-LDDSNFSGVRIGPGLALNHLQFADDTLLFCENDENQIDLLCNTLLSFLFTSG 714
L+N+ D + + +Q+ADDTL+ E D Q+ +L + + +F +G
Sbjct: 1463 QSLINKAKCMDLLKLPIPMQTNQDFPIIQYADDTLIIAEGDTRQLLILKSIINTFSEATG 1522
Query: 715 LKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRRASFWEPMLDKL 774
LKVNL KS ++ N+D +A +G S G P YLG PLG ++P+++K
Sbjct: 1523 LKVNLQKSMMLPINMDEERLDTLARTFGCSKGSFPFTYLGLPLGITKPTIQDYQPLINKC 1582
Query: 775 RRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKILRSFLQGG 831
+ + + F+S +GRL L A ++P F M P V+ +++K + L G
Sbjct: 1583 EARLGS-VATFLSEAGRLELTNAVFTALPTFYMCTLAIPKSVIHKIDKFRKHCLWRG 1638
>gb|AAB82639.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25408936|pir||A84888 hypothetical protein
At2g45230 [imported] - Arabidopsis thaliana
Length = 1374
Score = 352 bits (904), Expect = 4e-95
Identities = 243/849 (28%), Positives = 420/849 (48%), Gaps = 32/849 (3%)
Query: 2 RWVSWNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKLDSSRVKTVEAFANAVKMDYVSV 61
R +SWN +G+G R + E PE++ + E+K + ++ V D +V
Sbjct: 2 RILSWNCQGVGNTPTVRHLREIRGLYFPEVIFLCETKKRRNYLENVVGHLGF--FDLHTV 59
Query: 62 DAVGSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYDF 121
+ +G +GGL +W S ++ + R I LL + ++ +YG V +ER +
Sbjct: 60 EPIGKSGGLALMWKD-SVQIKVLQSDKRLIDALL--IWQDKEFYLTCIYGEPVQAERGEL 116
Query: 122 FNLLSEALSLNNCFCKIMGGDFNAILNEGERIGIQMGVESS---FSDFVADSNLIDLPLQ 178
+ L+ L L+ ++ GDFN +++ E+IG ESS F + L ++
Sbjct: 117 WERLTR-LGLSRSGPWMLTGDFNELVDPSEKIGGPARKESSCLEFRQMLNSCGLWEVNHS 175
Query: 179 SDKFTWYSSRAGGLWS-RIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSLGAINFG 237
+F+WY +R L R+DR + ++ + F + SD P+I +L N+
Sbjct: 176 GYQFSWYGNRNDELVQCRLDRTVANQAWMELFPQAKATYLQKICSDHSPLINNLVGDNWR 235
Query: 238 P-KPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVS 296
F + W+ + F+ L+ ++W +T + ++ +K+ + I +W+ + S
Sbjct: 236 KWAGFKYDKRWVQREGFKDLLCNFWSQQSTK--TNALMMEKIASCRREISKWKRVSKPSS 293
Query: 297 RVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKARVRWLRE 356
VRI ++ L ++ + + K L+ QEY EE W++K+R+ W+R
Sbjct: 294 AVRIQELQFKLDAATKQIPFDRRELARLKKELS------QEYNNEEQFWQEKSRIMWMRN 347
Query: 357 GDRNTSFFHSISKIRKSKKLISQLVVGG----VSIEDPNMIKDAIFSHFQSFFTSENRLR 412
GDRNT +FH+ +K R+++ I +L+ S ED + +A +F+ F SE+
Sbjct: 348 GDRNTKYFHAATKNRRAQNRIQKLIDEEGREWTSDEDLGRVAEA---YFKKLFASEDVGY 404
Query: 413 PKIRCANLSRL-SQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVK 471
NL+ L S +L ++ E+ S + ++ PG DG N ++QFW +
Sbjct: 405 TVEELENLTPLVSDQMNNNLLAPITKEEVQRATFSINPHKCPGPDGMNGFLYQQFWETMG 464
Query: 472 EDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASR 531
+ + + + F +G + +G+N T I LIPK ++DFRPISL YK+I K++A+R
Sbjct: 465 DQITEMVQAFFRSGSIEEGMNKTNICLIPKILKAEKMTDFRPISLCNVIYKVIGKLMANR 524
Query: 532 LQVVTPLLISENQFAFAQGRQISDCILIANEVVDFL--NKKQGGGFL-LKLDFAKDYDNI 588
L+ + P LISE Q AF +GR ISD ILIA+E++ L N K F+ +K D +K YD +
Sbjct: 525 LKKILPSLISETQAAFVKGRLISDNILIAHELLHALSSNNKCSEEFIAIKTDISKAYDRV 584
Query: 589 DWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLF 648
+W FL ++ + F D WI I CV + +L+NG+ G +GLRQGDPLSP+LF
Sbjct: 585 EWPFLEKAMRGLGFADHWIRLIMECVKSVRYQVLINGTPHGEIIPSRGLRQGDPLSPYLF 644
Query: 649 NLCVNGLSCLLNQLLDDSNFSGVRIGPGL-ALNHLQFADDTLLFCENDENQIDLLCNTLL 707
+C L +L + +G+++ G ++HL FADD++ +C+ ++ + + +
Sbjct: 645 VICTEMLVKMLQSAEQKNQITGLKVARGAPPISHLLFADDSMFYCKVNDEALGQIIRIIE 704
Query: 708 SFLFTSGLKVNLHKSTI-IGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRRASF 766
+ SG +VN KS+I G ++ V G +YLG P + +
Sbjct: 705 EYSLASGQRVNYLKSSIYFGKHISEERRCLVKRKLGIEREGGEGVYLGLPESFQGSKVAT 764
Query: 767 WEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKILRS 826
+ D+L +K W S F+S G+ L+KA ++P + M+ FK P + ++E ++
Sbjct: 765 LSYLKDRLGKKVLGWQSNFLSPGGKEILLKAVAMALPTYTMSCFKIPKTICQQIESVMAE 824
Query: 827 FLQGGKLHG 835
F K G
Sbjct: 825 FWWKNKKEG 833
>gb|AAG50806.1| unknown protein [Arabidopsis thaliana] gi|25403016|pir||D86384
unknown protein [imported] - Arabidopsis thaliana
Length = 1213
Score = 352 bits (904), Expect = 4e-95
Identities = 272/862 (31%), Positives = 416/862 (47%), Gaps = 48/862 (5%)
Query: 6 WNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKLDSSRVKTVEAFANAVKMDYVSVD--A 63
WN+RG R + + KP V E+ + + + F NA+ + V+ A
Sbjct: 8 WNIRGFNNVSHRSGFKKWVKANKPIFGGVIETHVKQPKDRK---FINALLPGWSFVENYA 64
Query: 64 VGSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYDFFN 123
G + +W+P+ + A K+++ I + S + + VY N + R + +
Sbjct: 65 FSDLGKIWVMWDPSVQVVVVA-KSLQMITCEVLLPGSPSWIIVSVVYAANEVASRKELWI 123
Query: 124 LLSEALSLNNCFCKIMG-------GDFNAILNEGERIG-IQMGVESSFSDF---VADSNL 172
+ +N I+G GDFN +LN E + + V+ + DF + + L
Sbjct: 124 EI-----VNMVVSGIIGDRPWLVLGDFNQVLNPQEHSNPVSLNVDINMRDFRDCLLAAEL 178
Query: 173 IDLPLQSDKFTWYS-SRAGGLWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRP--IIL 229
DL + + FTW++ S + +IDR LV++ F + SD ++L
Sbjct: 179 SDLRYKGNTFTWWNKSHTTPVAKKIDRILVNDSWNALFPSSLGIFGSLDFSDHVSCGVVL 238
Query: 230 SLGAINFGPKPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQ 289
+I +PF F+N+ L + +F LV D W + VG S F + +KLK LK IK++
Sbjct: 239 EETSIK-AKRPFKFFNYLLKNLDFLNLVRDNWFTLNVVGSSMFRVSKKLKALKKPIKDFS 297
Query: 290 GYASSVSRVRINTIEADLQVTMDR-LASEEP---GIELRKNRLTILEGLWQEYRK-EEYR 344
S R L DR LA P EL R W EE
Sbjct: 298 RLNYSELEKRTKEAHDFLIGCQDRTLADPTPINASFELEAERK------WHILTAAEESF 351
Query: 345 WRQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVVG-GVSIEDPNMIKDAIFSHFQS 403
+RQK+R+ W EGD NT +FH ++ R S IS L G G ++ I D S+F S
Sbjct: 352 FRQKSRISWFAEGDGNTKYFHRMADARNSSNSISALYDGNGKLVDSQEGILDLCASYFGS 411
Query: 404 FFTSENRLRPKIRCAN-----LS-RLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDG 457
E + P + N LS R S A LE FS +I L S N++ G DG
Sbjct: 412 LLGDE--VDPYLMEQNDMNLLLSYRCSPAQVCELESTFSNEDIRAALFSLPRNKSCGPDG 469
Query: 458 FNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLI 517
F FF W IV +V ++F S+G L+K NAT I LIPK P+ SDFRPIS +
Sbjct: 470 FTAEFFIDSWSIVGAEVTDAIKEFFSSGCLLKQWNATTIVLIPKIVNPTCTSDFRPISCL 529
Query: 518 GSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLN-KKQGGGFL 576
+ YK+I+++L RLQ + +IS Q AF GR +++ +L+A ++V N +
Sbjct: 530 NTLYKVIARLLTDRLQRLLSGVISSAQSAFLPGRSLAENVLLATDLVHGYNWSNISPRGM 589
Query: 577 LKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKG 636
LK+D K +D++ W F+ L+ + +++INWI C+ST T ++ +NG + GFF+ KG
Sbjct: 590 LKVDLKKAFDSVRWEFVIAALRALAIPEKFINWISQCISTPTFTVSINGGNGGFFKSTKG 649
Query: 637 LRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGLALNHLQFADDTLLFCENDE 696
LRQGDPLSP+LF L + S LL+ + L+++HL FADD ++F +
Sbjct: 650 LRQGDPLSPYLFVLAMEAFSNLLHSRYESGLIHYHPKASNLSISHLMFADDVMIFFDGGS 709
Query: 697 NQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLPILYLGAP 756
+ +C TL F SGLKVN KS + ++ E+ A YG+ IG LPI YLG P
Sbjct: 710 FSLHGICETLDDFASWSGLKVNKDKSHLYLAGLNQLESNANAA-YGFPIGTLPIRYLGLP 768
Query: 757 LGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKV 816
L R + +EP+L+K+ + R+W +K +S +GR+ L+ + + F M+ F P
Sbjct: 769 LMNRKLRIAEYEPLLEKITARFRSWVNKCLSFAGRIQLISSVIFGSINFWMSTFLLPKGC 828
Query: 817 VGEVEKILRSFLQGGKLHGYHG 838
+ +E + FL G + G
Sbjct: 829 IKRIESLCSRFLWSGNIEQAKG 850
>emb|CAB81184.1| putative protein [Arabidopsis thaliana] gi|4539357|emb|CAB40051.1|
putative protein [Arabidopsis thaliana]
gi|7486147|pir||T04278 hypothetical protein F25I24.40 -
Arabidopsis thaliana
Length = 1294
Score = 343 bits (881), Expect = 2e-92
Identities = 242/781 (30%), Positives = 401/781 (50%), Gaps = 35/781 (4%)
Query: 65 GSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYDFFNL 124
G +GGL LW S + + ++ R I + + +++ ++ VYG SER+ +
Sbjct: 424 GHSGGLALLWKD-SVRLSNLYQDDRHIDVHIS--INNINFYLSRVYGHPCQSERHSLWTH 480
Query: 125 LSEALSLNNCFCKIMGGDFNAILNEGERIGIQMGVESSFSDF---VADSNLIDLPLQSDK 181
E LS I+ GDFN IL+ E+IG E +F F V+ +L D+ D+
Sbjct: 481 F-ENLSKTRNDPWILIGDFNEILSNNEKIGGPQRDEWTFRGFRNMVSTCDLKDIRSIGDR 539
Query: 182 FTWYSSR-AGGLWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSLGAINFGP-K 239
F+W R + + +DR ++ + F + SD +P+ LSL +
Sbjct: 540 FSWVGERHSHTVKCCLDRAFINSEGAFLFPFAELEFLEFTGSDHKPLFLSLEKTETRKMR 599
Query: 240 PFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVSRVR 299
PF F L F+ V W A + L +++ + + + + ++ SR+R
Sbjct: 600 PFRFDKRLLEVPHFKTYVKAGWNKA--INGQRKHLPDQVRTCRQAMAKLKHKSNLNSRIR 657
Query: 300 INTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKARVRWLREGDR 359
IN ++A L M + E R+ I L YR EE W+QK+R +W++EGDR
Sbjct: 658 INQLQAALDKAMSSVNRTE-----RRTISHIQRELTVAYRDEERYWQQKSRNQWMKEGDR 712
Query: 360 NTSFFHSISKIRKSKKLISQLVVGGVSIEDPNMIKD--AIFSHFQSFFTS--ENRLRPK- 414
NT FFH+ +K R S +++LV + E+ + + I H Q FFT E+ RP
Sbjct: 713 NTEFFHACTKTRFS---VNRLVT--IKDEEGMIYRGDKEIGVHAQEFFTKVYESNGRPVS 767
Query: 415 -IRCANLSRL--SQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVK 471
I A + Q N L K S+ EI+ + ++APG DG F+K W IV
Sbjct: 768 IIDFAGFKPIVTEQIND-DLTKDLSDLEIYNAICHIGDDKAPGPDGLTARFYKSCWEIVG 826
Query: 472 EDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASR 531
DV+K + F T + + +N T I +IPK P T+SD+RPI+L YK+ISK L R
Sbjct: 827 PDVIKEVKIFFRTSYMKQSINHTNICMIPKITNPETLSDYRPIALCNVLYKIISKCLVER 886
Query: 532 LQVVTPLLISENQFAFAQGRQISDCILIANEVVDFL--NKKQGGGFL-LKLDFAKDYDNI 588
L+ ++S++Q AF GR ++D ++IA+E++ L K+ ++ +K D +K YD +
Sbjct: 887 LKGHLDAIVSDSQAAFIPGRLVNDNVMIAHEMMHSLKTRKRVSQSYMAVKTDVSKAYDRV 946
Query: 589 DWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLF 648
+W FL ++ F + WI WI V + S+LVNG G + ++G+RQGDPLSP+LF
Sbjct: 947 EWNFLETTMRLFGFSETWIKWIMGAVKSVNYSVLVNGIPHGTIQPQRGIRQGDPLSPYLF 1006
Query: 649 NLCVNGLSCLLNQLLDDSNFSGVRIGPGL-ALNHLQFADDTLLFCENDENQIDLLCNTLL 707
LC + L+ L+ + + + G+RIG G+ + HLQFADD+L FC+++ L +
Sbjct: 1007 ILCADILNHLIKNRVAEGDIRGIRIGNGVPGVTHLQFADDSLFFCQSNVRNCQALKDVFD 1066
Query: 708 SFLFTSGLKVNLHKSTI-IGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRRASF 766
+ + SG K+N+ KS I G V R+ + G YLG P ++
Sbjct: 1067 VYEYYSGQKINMSKSMITFGSRVHGTTQNRLKNILGIQSHGGGGKYLGLPEQFGRKKRDM 1126
Query: 767 WEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKILRS 826
+ ++++++++ +W++K++S +G+ ++K+ S+P++ M+ FK PL +V E+E +L +
Sbjct: 1127 FNYIIERVKKRTSSWSAKYLSPAGKEIMLKSVAMSMPVYAMSCFKLPLNIVSEIEALLMN 1186
Query: 827 F 827
F
Sbjct: 1187 F 1187
>gb|AAC33961.1| contains similarity to reverse trancriptase (Pfam: rvt.hmm, score:
42.57) [Arabidopsis thaliana] gi|7486711|pir||T01893
hypothetical protein F8M12.22 - Arabidopsis thaliana
Length = 1662
Score = 343 bits (881), Expect = 2e-92
Identities = 242/781 (30%), Positives = 401/781 (50%), Gaps = 35/781 (4%)
Query: 65 GSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYDFFNL 124
G +GGL LW S + + ++ R I + + +++ ++ VYG SER+ +
Sbjct: 444 GHSGGLALLWKD-SVRLSNLYQDDRHIDVHIS--INNINFYLSRVYGHPCQSERHSLWTH 500
Query: 125 LSEALSLNNCFCKIMGGDFNAILNEGERIGIQMGVESSFSDF---VADSNLIDLPLQSDK 181
E LS I+ GDFN IL+ E+IG E +F F V+ +L D+ D+
Sbjct: 501 F-ENLSKTRNDPWILIGDFNEILSNNEKIGGPQRDEWTFRGFRNMVSTCDLKDIRSIGDR 559
Query: 182 FTWYSSR-AGGLWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSLGAINFGP-K 239
F+W R + + +DR ++ + F + SD +P+ LSL +
Sbjct: 560 FSWVGERHSHTVKCCLDRAFINSEGAFLFPFAELEFLEFTGSDHKPLFLSLEKTETRKMR 619
Query: 240 PFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVSRVR 299
PF F L F+ V W A + L +++ + + + + ++ SR+R
Sbjct: 620 PFRFDKRLLEVPHFKTYVKAGWNKA--INGQRKHLPDQVRTCRQAMAKLKHKSNLNSRIR 677
Query: 300 INTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKARVRWLREGDR 359
IN ++A L M + E R+ I L YR EE W+QK+R +W++EGDR
Sbjct: 678 INQLQAALDKAMSSVNRTE-----RRTISHIQRELTVAYRDEERYWQQKSRNQWMKEGDR 732
Query: 360 NTSFFHSISKIRKSKKLISQLVVGGVSIEDPNMIKD--AIFSHFQSFFTS--ENRLRPK- 414
NT FFH+ +K R S +++LV + E+ + + I H Q FFT E+ RP
Sbjct: 733 NTEFFHACTKTRFS---VNRLVT--IKDEEGMIYRGDKEIGVHAQEFFTKVYESNGRPVS 787
Query: 415 -IRCANLSRL--SQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVK 471
I A + Q N L K S+ EI+ + ++APG DG F+K W IV
Sbjct: 788 IIDFAGFKPIVTEQIND-DLTKDLSDLEIYNAICHIGDDKAPGPDGLTARFYKSCWEIVG 846
Query: 472 EDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASR 531
DV+K + F T + + +N T I +IPK P T+SD+RPI+L YK+ISK L R
Sbjct: 847 PDVIKEVKIFFRTSYMKQSINHTNICMIPKITNPETLSDYRPIALCNVLYKIISKCLVER 906
Query: 532 LQVVTPLLISENQFAFAQGRQISDCILIANEVVDFL--NKKQGGGFL-LKLDFAKDYDNI 588
L+ ++S++Q AF GR ++D ++IA+E++ L K+ ++ +K D +K YD +
Sbjct: 907 LKGHLDAIVSDSQAAFIPGRLVNDNVMIAHEMMHSLKTRKRVSQSYMAVKTDVSKAYDRV 966
Query: 589 DWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLF 648
+W FL ++ F + WI WI V + S+LVNG G + ++G+RQGDPLSP+LF
Sbjct: 967 EWNFLETTMRLFGFSETWIKWIMGAVKSVNYSVLVNGIPHGTIQPQRGIRQGDPLSPYLF 1026
Query: 649 NLCVNGLSCLLNQLLDDSNFSGVRIGPGL-ALNHLQFADDTLLFCENDENQIDLLCNTLL 707
LC + L+ L+ + + + G+RIG G+ + HLQFADD+L FC+++ L +
Sbjct: 1027 ILCADILNHLIKNRVAEGDIRGIRIGNGVPGVTHLQFADDSLFFCQSNVRNCQALKDVFD 1086
Query: 708 SFLFTSGLKVNLHKSTI-IGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRRASF 766
+ + SG K+N+ KS I G V R+ + G YLG P ++
Sbjct: 1087 VYEYYSGQKINMSKSMITFGSRVHGTTQNRLKNILGIQSHGGGGKYLGLPEQFGRKKRDM 1146
Query: 767 WEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKILRS 826
+ ++++++++ +W++K++S +G+ ++K+ S+P++ M+ FK PL +V E+E +L +
Sbjct: 1147 FNYIIERVKKRTSSWSAKYLSPAGKEIMLKSVAMSMPVYAMSCFKLPLNIVSEIEALLMN 1206
Query: 827 F 827
F
Sbjct: 1207 F 1207
>emb|CAA66812.1| non-ltr retrotransposon reverse transcriptase-like protein
[Arabidopsis thaliana]
Length = 893
Score = 343 bits (879), Expect = 3e-92
Identities = 273/865 (31%), Positives = 427/865 (48%), Gaps = 58/865 (6%)
Query: 6 WNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKLDSSRVKTVEAFANAVK-MDYVSVDAV 64
WNVRG + RR + KP + E+ + + K + +N + +V
Sbjct: 8 WNVRGFNISSHRRGFKKWFLLNKPLFGGLIETHVKQPKEK--KFISNLLPGWSFVENYEF 65
Query: 65 GSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMN-VYGPNVDSERYDFFN 123
G + LW+P S + ++++ I L L DS F+++ VY N + R + +N
Sbjct: 66 SVLGKIWVLWDP-SVKVVVIGRSLQMITCEL-LLPDSPSWFVVSIVYASNEEGTRKELWN 123
Query: 124 LLSEALSLNNCFCK---IMGGDFNAILNEGERIGIQMGVE-SSFSDFVADSNLIDLPLQS 179
L + L+L+ I+ GDFN ILN I +G + +F + DS+L DL +
Sbjct: 124 ELVQ-LALSPVVVGRSWIVLGDFNQILNPESAINANIGRKIRAFRSCLLDSDLYDLVYKG 182
Query: 180 DKFTWYSSRAGG-LWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSLG-AINFG 237
+TW++ + L +IDR LV++ F + SD + L A+
Sbjct: 183 SSYTWWNKCSSRPLAKKIDRILVNDHWNTLFPSAYANFGEPDFSDHSSCEVVLDPAVLKA 242
Query: 238 PKPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVSR 297
+PF F+N++L + +F +L+ + W S G + + + +KLK LK I SR
Sbjct: 243 KRPFRFFNYFLHNPDFLQLIRENWYSCNVSGSAMYRVSKKLKHLKLPI-------CCFSR 295
Query: 298 VRINTIEADLQVTMDRLASEEPGIELRKNRLTI-----------LEGL--WQEYRK-EEY 343
+ IE + SE I L + R+T+ LE WQ K EE
Sbjct: 296 ENYSDIE--------KRVSEAHAIVLHRQRITLTNPSVVHATLELEATRKWQILAKAEES 347
Query: 344 RWRQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVVG-GVSIEDPNMIKDAIFSHFQ 402
+ QK+ + WL EGD NT++FH ++ +RKS I+ L+ G IE IK+ I H
Sbjct: 348 FFCQKSSISWLYEGDNNTAYFHKMADMRKSINTINFLIDDFGERIETQQGIKEGIKEHSC 407
Query: 403 SFFTS-------ENRLRPKIRCANLS-RLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPG 454
+FF S EN L LS R S LE+ FS+ +I E S N+A G
Sbjct: 408 NFFESLLCGVEGENSLAQSDMNLLLSFRCSVDQINDLERSFSDLDIQEAFFSLPRNKASG 467
Query: 455 TDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPI 514
DG++ FFK W +V +V + ++F +G+L+K NAT + LIPK S ++DFRPI
Sbjct: 468 PDGYSSEFFKGVWFVVGPEVTEAVQEFFRSGQLLKQWNATTLVLIPKITNSSKMTDFRPI 527
Query: 515 SLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKKQ-GG 573
S + + YK+I+K+L SRL+ + +IS +Q AF GR +S+ +L+A E+V N K
Sbjct: 528 SCLNTLYKVIAKLLTSRLKKLLNEVISPSQSAFLPGRLLSENVLLATEIVHGYNTKNISS 587
Query: 574 GFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEM 633
+LK+D K +D++ W F+ + + ++++ WI C+ST S++VNGSS+GFF+
Sbjct: 588 RGMLKVDLRKAFDSVRWDFIISAFRALAVPEKFVCWINQCISTPYFSVMVNGSSSGFFKS 647
Query: 634 EKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGLALNHLQFADDTLLFCE 693
KGLRQGDPLSP+LF L + S LL D L+++HL FADD ++F +
Sbjct: 648 NKGLRQGDPLSPYLFVLAMEVFSSLLKARFDAGYIQYHPKTADLSISHLMFADDVMVFFD 707
Query: 694 NDENQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLPILYL 753
+ + + L F SGL VN K+ + D E ++ YG+ I LPI YL
Sbjct: 708 GGSSSLHGISEALDDFASWSGLHVNKDKTNLYLAGTDEVEALAIS-HYGFPISTLPIRYL 766
Query: 754 GAPLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAP 813
G PL + S +E L ++ R+W K +S +GR+ L+ + + + F M+ F
Sbjct: 767 GLPLMSRKLKISEYE-----LVKRFRSWAVKSLSFAGRVQLITSVITGLVNFWMSTFVLL 821
Query: 814 LKVVGEVEKILRSFLQGGKLHGYHG 838
L V ++E + FL G + G
Sbjct: 822 LGCVKKIESLCSRFLWSGSIDASKG 846
>dbj|BAB01845.1| non-LTR retroelement reverse transcriptase-like protein
[Arabidopsis thaliana]
Length = 893
Score = 342 bits (878), Expect = 4e-92
Identities = 273/865 (31%), Positives = 427/865 (48%), Gaps = 58/865 (6%)
Query: 6 WNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKLDSSRVKTVEAFANAVK-MDYVSVDAV 64
WNVRG + RR + KP + E+ + + K + +N + +V
Sbjct: 8 WNVRGFNISSHRRGFKKWFLLNKPLFGGLIETHVKQPKEK--KFISNLLPGWSFVENYEF 65
Query: 65 GSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMN-VYGPNVDSERYDFFN 123
G + LW+P S + ++++ I L L DS F+++ VY N + R + +N
Sbjct: 66 SVLGKIWVLWDP-SVKVVVIGRSLQMITCEL-LLPDSPSWFVVSIVYASNEEGTRKELWN 123
Query: 124 LLSEALSLNNCFCK---IMGGDFNAILNEGERIGIQMGVE-SSFSDFVADSNLIDLPLQS 179
L + L+L+ I+ GDFN ILN I +G + +F + DS+L DL +
Sbjct: 124 ELVQ-LALSPVVVGRSWIVLGDFNQILNPESAINANIGRKIRAFRSCLLDSDLYDLVYKG 182
Query: 180 DKFTWYSSRAGG-LWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSLG-AINFG 237
+TW++ + L +IDR LV++ F + SD + L A+
Sbjct: 183 SSYTWWNKCSSRPLAKKIDRILVNDHWNTLFPSAYANFGEPDFSDHSSCEVVLDPAVLKA 242
Query: 238 PKPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVSR 297
+PF F+N++L + +F +L+ + W S G + + + +KLK LK I SR
Sbjct: 243 KRPFRFFNYFLHNPDFLQLIRENWYSCNVSGSAMYRVSKKLKHLKLPI-------CCFSR 295
Query: 298 VRINTIEADLQVTMDRLASEEPGIELRKNRLTI-----------LEGL--WQEYRK-EEY 343
+ IE + SE I L + R+T+ LE WQ K EE
Sbjct: 296 ENYSDIE--------KRVSEAHAIVLHRQRITLTNPSVVHATLELEATRKWQILAKAEES 347
Query: 344 RWRQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVVG-GVSIEDPNMIKDAIFSHFQ 402
+ QK+ + WL EGD NT++FH ++ +RKS I+ L+ G IE IK+ I H
Sbjct: 348 FFCQKSSISWLYEGDNNTAYFHKMADMRKSINTINFLIDDFGERIETQQGIKEGIKEHSC 407
Query: 403 SFFTS-------ENRLRPKIRCANLS-RLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPG 454
+FF S EN L LS R S LE+ FS+ +I E S N+A G
Sbjct: 408 NFFESLLCGVEGENSLAQSDMNLLLSFRCSVDQINDLERSFSDLDIQEAFFSLPRNKASG 467
Query: 455 TDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPI 514
DG++ FFK W +V +V + ++F +G+L+K NAT + LIPK S ++DFRPI
Sbjct: 468 PDGYSSEFFKGVWFVVGPEVTEAVQEFFRSGQLLKQWNATTLVLIPKITNSSKMTDFRPI 527
Query: 515 SLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKKQ-GG 573
S + + YK+I+K+L SRL+ + +IS +Q AF GR +S+ +L+A E+V N K
Sbjct: 528 SCLNTLYKVIAKLLTSRLKKLLNEVISPSQSAFLPGRLLSENVLLATEIVHGYNTKNISS 587
Query: 574 GFLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEM 633
+LK+D K +D++ W F+ + + ++++ WI C+ST S++VNGSS+GFF+
Sbjct: 588 RGMLKVDLRKAFDSVRWDFIISAFRALAVPEKFVCWINQCISTPYFSVMVNGSSSGFFKS 647
Query: 634 EKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGLALNHLQFADDTLLFCE 693
KGLRQGDPLSP+LF L + S LL D L+++HL FADD ++F +
Sbjct: 648 NKGLRQGDPLSPYLFVLAMEVFSSLLKARFDAGYIHYHPKTADLSISHLMFADDVMVFFD 707
Query: 694 NDENQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLPILYL 753
+ + + L F SGL VN K+ + D E ++ YG+ I LPI YL
Sbjct: 708 GGSSSLHGISEALDDFASWSGLHVNKDKTNLYLAGTDEVEALAIS-HYGFPISTLPIRYL 766
Query: 754 GAPLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAP 813
G PL + S +E L ++ R+W K +S +GR+ L+ + + + F M+ F
Sbjct: 767 GLPLMSRKLKISEYE-----LVKRFRSWAVKSLSFAGRVQLITSVITGLVNFWMSTFVLL 821
Query: 814 LKVVGEVEKILRSFLQGGKLHGYHG 838
L V ++E + FL G + G
Sbjct: 822 LGCVKKIESLCSRFLWSGSIDASKG 846
>emb|CAB78094.1| RNA-directed DNA polymerase-like protein [Arabidopsis thaliana]
gi|4538901|emb|CAB39638.1| RNA-directed DNA
polymerase-like protein [Arabidopsis thaliana]
gi|7485606|pir||T04018 hypothetical protein F17A8.60 -
Arabidopsis thaliana
Length = 1274
Score = 340 bits (873), Expect = 2e-91
Identities = 235/732 (32%), Positives = 364/732 (49%), Gaps = 32/732 (4%)
Query: 112 PNVDSERYDFFNLLSEALSLNNCFCKIMGGDFNAILNEGERIGIQMGVES---SFSDFVA 168
PN R F++ +S +L ++ GDFN IL+ E+ G + E +F FV+
Sbjct: 51 PNFIDNRSVFWDKIS-SLGAQRSSAWLLTGDFNDILDNSEKQGGPLRWEGFFLAFRSFVS 109
Query: 169 DSNLIDLPLQSDKFTWYSSRAGG-LWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPI 227
+ L D+ + +W +R + SR+DR L + F + SD RP+
Sbjct: 110 QNGLWDINHTGNSLSWRGTRYSHFIKSRLDRALGNCSWSELFPMSKCEYLRFEGSDHRPL 169
Query: 228 ILSLGAINFG-PKPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIK 286
+ GA KPF F +E LV + W A + K+ + I
Sbjct: 170 VTYFGAPPLKRSKPFRFDRRLREKEEIRALVKEVWELARQDS-----VLYKISRCRQSII 224
Query: 287 EWQGYASSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWR 346
+W +S S I + L+ L+++ P L + I + L YR+EE W+
Sbjct: 225 KWTKEQNSNSAKAIKKAQQALE---SALSADIPDPSLIGS---ITQELEAAYRQEELFWK 278
Query: 347 QKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVVG-GVSIEDPNMIKDAIFSHFQSFF 405
Q +RV+WL GDRN +FH+ ++ R+ +S + G G + I I S+FQ+ F
Sbjct: 279 QWSRVQWLNSGDRNKGYFHATTRTRRMLNNLSVIEDGSGQEFHEEEQIASTISSYFQNIF 338
Query: 406 TSENRLRPKIRCANLSRLSQAN-KMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFK 464
T+ N ++ LS + ++ L K S EI E L S ++APG DGF+ +FF
Sbjct: 339 TTSNNSDLQVVQEALSPIISSHCNEELIKISSLLEIKEALFSISADKAPGPDGFSASFFH 398
Query: 465 QFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLI 524
+W I++ DV + F L LN T ++LIPK + P VSD+RPI+L YK++
Sbjct: 399 AYWDIIEADVSRDIRSFFVDSCLSPRLNETHVTLIPKISAPRKVSDYRPIALCNVQYKIV 458
Query: 525 SKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFL---NKKQGGGFLLKLDF 581
+K+L RLQ LIS +Q AF GR I+D +LI +E++ FL K+ +K D
Sbjct: 459 AKILTRRLQPWLSELISLHQSAFVPGRAIADNVLITHEILHFLRVSGAKKYCSMAIKTDM 518
Query: 582 AKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGD 641
+K YD I W FL ++L + F D+WI W+ CV T + S L+NGS G +GLRQGD
Sbjct: 519 SKAYDRIKWNFLQEVLMRLGFHDKWIRWVMQCVCTVSYSFLINGSPQGSVVPSRGLRQGD 578
Query: 642 PLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGL-ALNHLQFADDTLLFCENDENQID 700
PLSP+LF LC LS L + + G+R+ G +NHL FADDT+ FC+ +
Sbjct: 579 PLSPYLFILCTEVLSGLCRKAQEKGVMVGIRVARGSPQVNHLLFADDTMFFCKTNPTCCG 638
Query: 701 LLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQRVAGM-----YGWSIGQLPILYLGA 755
L N L + SG +NL KS I + ++ +R + IG+ YLG
Sbjct: 639 ALSNILKKYELASGQSINLAKSAITFSSKTPQDIKRRVKLSLRIDNEGGIGK----YLGL 694
Query: 756 PLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLK 815
P R+ + ++D++R++ +W+ +F+S +G+ L+KA L S+P + M FK P
Sbjct: 695 PEHFGRRKRDIFSSIVDRIRQRSHSWSIRFLSSAGKQILLKAVLSSMPSYAMMCFKLPAS 754
Query: 816 VVGEVEKILRSF 827
+ +++ +L F
Sbjct: 755 LCKQIQSVLTRF 766
>gb|AAX96121.1| retrotransposon protein, putative, unclassified [Oryza sativa
(japonica cultivar-group)]
Length = 1556
Score = 335 bits (860), Expect = 5e-90
Identities = 234/762 (30%), Positives = 382/762 (49%), Gaps = 36/762 (4%)
Query: 65 GSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYDFFNL 124
G +GG++ N + + S + +I LR D+ ++ VYG D + DF
Sbjct: 790 GRSGGILLEINLELFDVTSIDEGEFYIKFHLRNKEDAFQWNLVAVYGAAQDEFKQDFLLE 849
Query: 125 LSEALSLNNCFCKIMGGDFNAILNEGERIGIQMGVESSFS-DFVADS-NLIDLPLQSDKF 182
L + S ++GGDFN I N E+ + F + V DS NL ++ L KF
Sbjct: 850 LVNSCS-KEALPMVIGGDFNIIRNPMEKNNERYNDRWPFLFNVVIDSLNLREILLSGCKF 908
Query: 183 TWYSSRAGGLWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSLG-AINFGPKP- 240
TW +S + ++DR LVS + K ++ LSD P++L G + +P
Sbjct: 909 TWANSMPNPTYEKLDRVLVSTEWETKNPLVTVHALCRDLSDHTPLVLDTGKGTHRNAQPL 968
Query: 241 FLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQ-----GYASSV 295
F F WLL +F+ L+ + W + Q K++ L+ ++ W Y
Sbjct: 969 FKFELGWLLRDDFQNLLVEIWSKDIQGATNIERWQNKIRRLRQFLRGWVKNLSGAYKKEK 1028
Query: 296 SRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKARVRWLR 355
R+ E D L S+E ++L+ + L Q R+EE +W Q+A+ + +
Sbjct: 1029 PRLIAKVDELDKLAETRILTSQE--LDLKNS---CKNALAQLLREEEVKWYQRAKTKRI- 1082
Query: 356 EGDRNTSFFHSISKIRKSKKLISQLVVGGVSIEDPNMIKDAIFSHFQSFFTSENRLRPKI 415
GDRNT ++H I+ + K I QL I+ +K + ++++ F + P
Sbjct: 1083 VGDRNTKYYHMITNGKHRKTKIFQLEQEEGVIKGDAQLKKYVTNYYRGLFGA-----PDG 1137
Query: 416 RCANLS--------RLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFW 467
C ++ ++S L K+F+E E+ + + N+APG DGF F++ FW
Sbjct: 1138 NCFSMCESQKDDIPQVSMEENTFLTKEFTEEEVKHAVFQMEHNKAPGPDGFPAEFYQVFW 1197
Query: 468 GIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKV 527
++KED++ F DFHS + LN I+L+PK+ + +R I L+ ++K+ +KV
Sbjct: 1198 EVIKEDLMAVFRDFHSGELPLHRLNFGIITLLPKNKEAKQIQQYRRICLLNVSFKVFTKV 1257
Query: 528 LASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKKQGGGFLLKLDFAKDYDN 587
+A+R+ +V +I +Q AF +GR I + +I +E + ++KK+ G +LKLDF K YD
Sbjct: 1258 MANRIALVAQKVIRPSQIAFLKGRNIIEGAIILHETLHEMHKKKSNGVILKLDFEKAYDK 1317
Query: 588 IDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFL 647
++W FL L+ F D W WI VS ++++ VN FF+ +KGLRQGDPLSP L
Sbjct: 1318 VNWNFLQQALRMKGFSDIWCKWIDQVVSGGSVAVKVNDEIGHFFQTKKGLRQGDPLSPIL 1377
Query: 648 FNLCVNGLSCLLNQLLDDSNFSGV---RIGPGLALNHLQFADDTLLFCENDENQIDLLCN 704
FNL + L+ L+N+ D +GV + GL++ LQ+ADDT+LF ++D + + L
Sbjct: 1378 FNLVADMLAILINRARDLETLNGVIPQLVDNGLSI--LQYADDTILFMDHDLVKANNLKL 1435
Query: 705 TLLSFLFTSGLKVNLHKSTIIGCNVDVRETQR-VAGMYGWSIGQLPILYLGAPLGGNPRR 763
L +F SGLK+N HKS + C +E ++ ++G +G P YLG P+
Sbjct: 1436 VLAAFERLSGLKINFHKSEMF-CFGKAKEVEKDYTTLFGCKMGTFPFRYLGLPMHFKRLS 1494
Query: 764 ASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLF 805
W+ + DK ++K W KF+S G+L L+ + L S+ F
Sbjct: 1495 NKDWKDLEDKFQKKLSCWRGKFLSSGGKLVLINSVLSSLATF 1536
>gb|AAD20714.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25412331|pir||G84649 hypothetical protein
At2g25550 [imported] - Arabidopsis thaliana
Length = 1750
Score = 335 bits (859), Expect = 6e-90
Identities = 237/792 (29%), Positives = 406/792 (50%), Gaps = 57/792 (7%)
Query: 65 GSAGGLISLW-NPASYAMESAVKNVRFIGLLLRALVDSSLLF------IMNVYGPNVDSE 117
G +GGL +W N S ++ S + L+DS + F + VYG SE
Sbjct: 444 GRSGGLALMWKNNVSLSLISQDER----------LIDSHVTFNNKSFYLSCVYGHPTQSE 493
Query: 118 RYDFFNLLSEALSLNNCFCKIMGGDFNAILNEGERIGIQMGVESSFSDF---VADSNLID 174
R+ + L E +S N ++ GDFN IL+ E+IG M E +F +F V+ ++ D
Sbjct: 494 RHQLWQTL-EHISDNRNAEWLLVGDFNEILSNAEKIGGPMREEWTFRNFRNMVSHCDIED 552
Query: 175 LPLQSDKFTWYSSR-AGGLWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSLG- 232
+ + D+F+W R + +DR ++ F ++ SD +P+++
Sbjct: 553 MRSKGDRFSWVGERHTHTVKCCLDRVFINSAWTATFPYAEIEFLDFTGSDHKPVLVHFNE 612
Query: 233 AINFGPKPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYA 292
+ K F F N + F+++V W T + + +++ + + + +
Sbjct: 613 SFPRRSKLFRFDNRLIDIPTFKRIVQTSW--RTNRNSRSTPITERISSCRQAMARLKHAS 670
Query: 293 SSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKARVR 352
+ S RI +++ L M+ + R+ + E L + + EE W+QK+R +
Sbjct: 671 NLNSEQRIKKLQSSLNRAMESTRRVD-----RQLIPQLQESLAKAFSDEEIYWKQKSRNQ 725
Query: 353 WLREGDRNTSFFHSISKIRKSKKLISQLV--VGGVSIEDPNMIKDAIFSHFQSFFT---S 407
W++EGD+NT +FH+ +K R S+ ++ ++ G + D I +H Q FFT S
Sbjct: 726 WMKEGDQNTGYFHACTKTRYSQNRVNTIMDDQGRMFTGDKE-----IGNHAQDFFTNIFS 780
Query: 408 ENRLRPK-IRCANL-SRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQ 465
N ++ I A+ S ++ + L K+FS+ EI++ + ++APG DG F+K
Sbjct: 781 TNGIKVSPIDFADFKSTVTNTVNLDLTKEFSDTEIYDAICQIGDDKAPGPDGLTARFYKN 840
Query: 466 FWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLIS 525
W IV DV+ + F T + +N T I +IPK P+T+SD+RPI+L YK+IS
Sbjct: 841 CWDIVGYDVILEVKKFFETSFMKPSINHTNICMIPKITNPTTLSDYRPIALCNVLYKVIS 900
Query: 526 KVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKKQGGG---FLLKLDFA 582
K L +RL+ ++S++Q AF GR I+D ++IA+EV+ L ++ +K D +
Sbjct: 901 KCLVNRLKSHLNSIVSDSQAAFIPGRIINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVS 960
Query: 583 KDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDP 642
K YD ++W FL ++ F ++WI WI V + S+L+NGS G+ +G+RQGDP
Sbjct: 961 KAYDRVEWDFLETTMRLFGFCNKWIGWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDP 1020
Query: 643 LSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGL-ALNHLQFADDTLLFCENDENQIDL 701
LSP+LF LC + LS L+N + GVRIG G A+ HLQFADD+L FC+ +
Sbjct: 1021 LSPYLFILCGDILSHLINGRASSGDLRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQA 1080
Query: 702 LCNTLLSFLFTSGLKVNLHKSTI-IGCNVDVRETQRVAGMYGWSIGQLPI-----LYLGA 755
L + + + SG K+N+ KS I G V R+ I ++P YLG
Sbjct: 1081 LKDVFDVYEYYSGQKINVQKSMITFGSRVYGSTQSRLK-----QILEIPNQGGGGKYLGL 1135
Query: 756 PLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLK 815
P ++ +E ++D+++++ TW+++F+S +G+ ++K+ ++P++ M+ FK P
Sbjct: 1136 PEQFGRKKKEMFEYIIDRVKKRTSTWSARFLSPAGKEIMLKSVALAMPVYAMSCFKLPKG 1195
Query: 816 VVGEVEKILRSF 827
+V E+E +L +F
Sbjct: 1196 IVSEIESLLMNF 1207
>gb|AAD24831.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25408166|pir||G84721 hypothetical protein
At2g31520 [imported] - Arabidopsis thaliana
Length = 1524
Score = 335 bits (859), Expect = 6e-90
Identities = 235/793 (29%), Positives = 405/793 (50%), Gaps = 59/793 (7%)
Query: 65 GSAGGLISLW-NPASYAMESAVKNVRFIGLLLRALVDSSLLF------IMNVYGPNVDSE 117
G +GGL +W N S ++ S + L+DS + F + VYG SE
Sbjct: 218 GRSGGLALMWKNNVSLSLISQDER----------LIDSHVTFNNKSFYLSCVYGHPTQSE 267
Query: 118 RYDFFNLLSEALSLNNCFCKIMGGDFNAILNEGERIGIQMGVESSFSDF---VADSNLID 174
R+ + L E +S N ++ GDFN IL+ E+IG M E +F +F V+ ++ D
Sbjct: 268 RHQLWQTL-EHISDNRNAEWLLVGDFNEILSNAEKIGGPMREEWTFRNFRNMVSHCDIED 326
Query: 175 LPLQSDKFTWYSSR-AGGLWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSLG- 232
+ + D+F+W R + +DR ++ F ++ SD +P+++
Sbjct: 327 MRSKGDRFSWVGERHTHTVKCCLDRVFINSAWTATFPYAETEFLDFTGSDHKPVLVHFNE 386
Query: 233 AINFGPKPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYA 292
+ K F F N + F+++V W T + + +++ + + + +
Sbjct: 387 SFPRRSKLFRFDNRLIDIPTFKRIVQTSW--RTNRNSRSTPITERISSCRQAMARLKHAS 444
Query: 293 SSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKARVR 352
+ S RI +++ L M+ + R+ + E L + + EE W+QK+R +
Sbjct: 445 NLNSEQRIKKLQSSLNRAMESTRRVD-----RQLIPQLQESLAKAFSDEEIYWKQKSRNQ 499
Query: 353 WLREGDRNTSFFHSISKIRKSKKLISQLV--VGGVSIEDPNMIKDAIFSHFQSFFT---S 407
W++EGD+NT +FH+ +K R S+ ++ ++ G + D I +H Q FFT S
Sbjct: 500 WMKEGDQNTGYFHACTKTRYSQNRVNTIMDDQGRMFTGDKE-----IGNHAQDFFTNIFS 554
Query: 408 ENRLRPK-IRCANL-SRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQ 465
N ++ I A+ S ++ + L K+FS+ EI++ + ++APG DG F+K
Sbjct: 555 TNGIKVSPIDFADFKSTVTNTVNLDLTKEFSDTEIYDAICQIGDDKAPGPDGLTARFYKN 614
Query: 466 FWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLIS 525
W IV DV+ + F T + +N T I +IPK P+T+SD+RPI+L YK+IS
Sbjct: 615 CWDIVGYDVILEVKKFFETSFMKPSINHTNICMIPKITNPTTLSDYRPIALCNVLYKVIS 674
Query: 526 KVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKKQGGG---FLLKLDFA 582
K L +RL+ ++S++Q AF GR I+D ++IA+EV+ L ++ +K D +
Sbjct: 675 KCLVNRLKSHLNSIVSDSQAAFIPGRIINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVS 734
Query: 583 KDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDP 642
K YD ++W FL ++ F ++WI WI V + S+L+NGS G+ +G+RQGDP
Sbjct: 735 KAYDRVEWDFLETTMRLFGFCNKWIGWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDP 794
Query: 643 LSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGPGL-ALNHLQFADDTLLFCENDENQIDL 701
LSP+LF LC + LS L+N + GVRIG G A+ HLQFADD+L FC+ +
Sbjct: 795 LSPYLFILCGDILSHLINGRASSGDLRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQA 854
Query: 702 LCNTLLSFLFTSGLKVNLHKSTI-IGCNV------DVRETQRVAGMYGWSIGQLPILYLG 754
L + + + SG K+N+ KS I G V +++ + G YLG
Sbjct: 855 LKDVFDVYEYYSGQKINVQKSMITFGSRVYGSTQSKLKQILEIPNQGGGG------KYLG 908
Query: 755 APLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPL 814
P ++ +E ++D+++++ TW+++F+S +G+ ++K+ ++P++ M+ FK P
Sbjct: 909 LPEQFGRKKKEMFEYIIDRVKKRTSTWSARFLSPAGKEIMLKSVALAMPVYAMSCFKLPK 968
Query: 815 KVVGEVEKILRSF 827
+V E+E +L +F
Sbjct: 969 GIVSEIESLLMNF 981
>ref|XP_475426.1| putative non-LTR retroelement reverse transcriptase [Oryza sativa
(japonica cultivar-group)] gi|46576009|gb|AAT01370.1|
putative non-LTR retroelement reverse transcriptase
[Oryza sativa (japonica cultivar-group)]
Length = 1614
Score = 333 bits (854), Expect = 2e-89
Identities = 222/721 (30%), Positives = 372/721 (50%), Gaps = 22/721 (3%)
Query: 106 IMNVYGPNVDSERYDFFNLLSEALSLNNCFCKIMGGDFNA--ILNEGERIGIQMGVESSF 163
+++VYGP +S + F L +S ++GGDFN +E I + F
Sbjct: 725 VISVYGPVDNSLKNVFLEELRWHISGRE-HPVMLGGDFNLYKFASEKSNSNIDVRCMDMF 783
Query: 164 SDFVADSNLIDLPLQSDKFTWYSSRAGGLWSRIDRWLVSEDAINKF-DGISQSVENWGLS 222
+ F++D +L ++ + KFTW + + +DR LVS+D +++ + + SV G S
Sbjct: 784 NKFISDLDLREVHIIGPKFTWTNKQYCPTQEVLDRILVSDDWDDRYPNSLISSVLRVG-S 842
Query: 223 DRRPIILSLGAINFGP-KPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQ---QKL 278
D P++L + + F F + WL + F+ LV + +++L+ +K
Sbjct: 843 DHTPLLLDTYELVVEKCRYFRFESAWLAVEGFKDLVGKKFPDRD----GSYILEFWNKKQ 898
Query: 279 KGLKNRIKEWQGYASSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEY 338
++ + W S R +++ L+ + E + ++R + + L Y
Sbjct: 899 VEMRRFLTGWGINKQSEDRRAKLSLQKKLEEIDSKAVDAELNADEWRHRYELEDALEHIY 958
Query: 339 RKEEYRWRQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVVGGVSIEDPNMIKDAIF 398
EE W ++ +WL EGDRNT ++H I+ RK + I+ L+ G + +K I
Sbjct: 959 ELEELYWHKRCGEQWLLEGDRNTEYYHRIANGRKKRCTITSLMDGDRELTTKEDLKHHIV 1018
Query: 399 SHFQSFFTSEN----RLRPKIRCANLSRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPG 454
+++S F SE+ L + +L L + +K +L + F+ E+++VL+ N A G
Sbjct: 1019 EYYKSLFRSESSSSIHLSQGVWVDSLC-LKEEDKNTLMRPFTIEELYKVLREAKPNTAFG 1077
Query: 455 TDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPI 514
DGF++ F++ FW ++ D+ + ++ +K LN ISLIPK++ P+ + FRPI
Sbjct: 1078 PDGFSIPFYRAFWPQLRPDLFEMLLMLYNEELDLKRLNFGVISLIPKNSNPTDIKQFRPI 1137
Query: 515 SLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKKQGGG 574
++ +K ISK + +RL + +IS Q AF GR I + +I +EV+ +N+K G
Sbjct: 1138 CVLNDCFKFISKCVCNRLTEIARDVISPTQTAFIPGRFILEGCVIIHEVLHEMNRKNLEG 1197
Query: 575 FLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEME 634
+LK+DF K YD + W FL +++ F +W+NWI+TCV + I +NG T FF
Sbjct: 1198 IILKIDFEKAYDKVSWDFLIEVMVRKGFPSKWVNWIKTCVMGGRVCININGERTDFFRTF 1257
Query: 635 KGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGV--RIGPGLALNHLQFADDTLLFC 692
+GLRQGDPLSP LFNL + L+ +L+ + SG+ I PG + HLQ+ADDT+LF
Sbjct: 1258 RGLRQGDPLSPLLFNLISDALAAMLDSAKREGVLSGLVPDIFPG-GITHLQYADDTVLFV 1316
Query: 693 ENDENQIDLLCNTLLSFLFTSGLKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLPILY 752
ND+ QI L F + LKVN HKS I + +T RVA M+ +GQ P+ Y
Sbjct: 1317 ANDDKQIVATKFILYCFEEMAVLKVNYHKSEIFTLGLSDNDTNRVAMMFNCPVGQFPMKY 1376
Query: 753 LGAPLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKA 812
LG P+G + ++ + KL ++ +W + +S +GR + L S+P + M ++
Sbjct: 1377 LGLPIGPDKILNLGFDFLGQKLEKRLNSWGNN-LSHAGRAVQINTCLSSIPSYAMCFYQL 1435
Query: 813 P 813
P
Sbjct: 1436 P 1436
>gb|AAP54617.1| putative non-LTR retroelement reverse transcriptase [Oryza sativa
(japonica cultivar-group)] gi|37536056|ref|NP_922330.1|
putative non-LTR retroelement reverse transcriptase
[Oryza sativa (japonica cultivar-group)]
gi|10140689|gb|AAG13524.1| putative non-LTR retroelement
reverse transcriptase [Oryza sativa (japonica
cultivar-group)]
Length = 1382
Score = 326 bits (836), Expect = 3e-87
Identities = 261/856 (30%), Positives = 414/856 (47%), Gaps = 76/856 (8%)
Query: 4 VSW--NVRGLGRAEKRRAVVEALSGLKPEIVLVQESKLDSSRVKTVE---AFANAVKMDY 58
V+W N RGLG A + + L+P +V + E+K+ + + + F+ +
Sbjct: 7 VTWGRNCRGLGSAATVGELRWLVKSLRPSLVFLSETKMRDKQARNLMWSLGFSGSF---- 62
Query: 59 VSVDAVGSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSER 118
+V G +GGL W A Y + N FI +L+ + + I VYG R
Sbjct: 63 -AVSCEGLSGGLALFWTTA-YTVSLRGFNSHFIDVLV-STEELPPWRISFVYGEPKRELR 119
Query: 119 YDFFNLLSEALSLNNCFCK--IMGGDFNAILNEGERIGIQMGVESSFSDF---VADSNLI 173
+ F+NLL L++ + + GDFN +L E +G++ E F + D LI
Sbjct: 120 HFFWNLLRR---LHDQWRGPWLCCGDFNEVLCLDEHLGMRERSEPHMQHFRSCLDDCGLI 176
Query: 174 DLPLQSDKFTWYSSRAGGLWS--RIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSL 231
DL KFTW + + S R+DR + + + F+ SD I + L
Sbjct: 177 DLGFVGPKFTWSNKQDANSNSKVRLDRAVANGEFSRYFEDCLVENVITTSSDHYAISIDL 236
Query: 232 GAINFGPKP------FLFYNHWLLDKEFEKLVSDWW--CSATTVG----WSAFVLQQKLK 279
N G + F F WL +++ ++V + W SA VG WS VLQQ
Sbjct: 237 SRRNHGQRRIPIQQGFRFEAAWLRAEDYREVVENSWRISSAGCVGLRGVWS--VLQQVAV 294
Query: 280 GLKNRIKEWQGYASSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYR 339
LK +W + R +I +E L+ ++ + +++ +L I + L + +
Sbjct: 295 SLK----DWSKASFGSVRRKILKMERKLKSLRQSPVND---VVIQEEKL-IEQQLCELFE 346
Query: 340 KEEYRWRQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVVG-GVSIEDPNMIKDAIF 398
KEE RQ++RV WLREGDRNT+FFH+ + R+ I +LV G IK
Sbjct: 347 KEEIMARQRSRVDWLREGDRNTAFFHARASARRRTNRIKELVRDDGSRCISQEGIKRMAE 406
Query: 399 SHFQSFFTSENRLRPKIRCANLSRLSQA--NKMS------LEKQFSEGEIWEVLKSCDGN 450
+++ F+SE C ++ + A NK+ L KQ++ EI L
Sbjct: 407 VFYENLFSSEP-------CDSMEEVLDAIPNKVGDFINGELGKQYTNEEIKTALFQMGST 459
Query: 451 RAPGTDGFNLNFFKQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSD 510
+APG DGF F++ WGI++E + F ++ +GL + + LIPK N S +S
Sbjct: 460 KAPGPDGFPALFYQTHWGILEEHICNAVRGFLLGEEIPEGLCDSVVVLIPKVNNASHLSK 519
Query: 511 FRPISLIGSAYKLISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKK 570
FRPISL YK+ SKVLA+RL+ P ++SE Q AF GR I+D L+A E + + K+
Sbjct: 520 FRPISLCNVLYKIASKVLANRLKPFLPDIVSEFQSAFVPGRLITDSALVAYECLHTIRKQ 579
Query: 571 QGGG--FLLKLDFAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSST 628
F LK+D K YD ++W +L L ++ F WIN + CVS+ ++ +NG T
Sbjct: 580 HNKNPFFALKIDMMKAYDRVEWAYLSGCLSKLGFSQDWINTVMRCVSSVRYAVKINGELT 639
Query: 629 GFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIG-PGLALNHLQFADD 687
+G+RQGDP+SP+LF LC GLSCLL++ G++ G G ++HL FADD
Sbjct: 640 KPVVPSRGIRQGDPISPYLFLLCTEGLSCLLHKKEVAGELQGIKNGRHGPPISHLLFADD 699
Query: 688 TLLFCENDENQIDLLCNTLLSFLFTSGLKVNLHKSTII-------GCNVDVRETQRVAGM 740
++ F + D + L NTL S+ SG K+NLHKS+I + V+ +V
Sbjct: 700 SIFFAKADSRNVQALKNTLRSYCSASGQKINLHKSSIFFGKRCPDAVKISVKSCLQVDNE 759
Query: 741 YGWSIGQLPILYLGAPLGGNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLC 800
L YLG P +F++ + +++ ++ W + +S +G T++KA
Sbjct: 760 V------LQDSYLGMPTEIGLATTNFFKFLPERIWKRVNGWTDRPLSRAGMETMLKAVAQ 813
Query: 801 SVPLFLMAIFKAPLKV 816
++P ++M+ F+ P+ +
Sbjct: 814 AIPNYVMSCFRIPVSI 829
>gb|AAC28221.1| similar to reverse transcriptases (PFam: rvt.hmm, score: 60.13)
[Arabidopsis thaliana] gi|7488321|pir||T01871
RNA-directed DNA polymerase homolog T24M8.8 -
Arabidopsis thaliana
Length = 1164
Score = 326 bits (836), Expect = 3e-87
Identities = 228/713 (31%), Positives = 364/713 (50%), Gaps = 30/713 (4%)
Query: 141 GDFNAILNEGERI---GIQMGVESS-FSDFVADSNLIDLPLQSDKFTWYSSRAGG-LWSR 195
GDFN IL+ E G + + F + + ++L DL + + FTW++ R+ + +
Sbjct: 40 GDFNQILHPSEHSTSDGFNVDRPTRIFRETILLASLTDLSFRGNTFTWWNKRSRAPVAKK 99
Query: 196 IDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSL-GAINFGPKPFLFYNHWLLDKEFE 254
+DR LV++ F SD LSL A KPF F N L D+ F
Sbjct: 100 LDRILVNDKWTTTFPSSLGLFGEPDFSDHSSCELSLMSASPRSKKPFRFNNFLLKDENFL 159
Query: 255 KLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQGYASSVSRVRINTIEADLQVTMDR- 313
L+ W S + G + + + KLK LK I+++ + S + T EA + + +
Sbjct: 160 SLICLKWFSTSVTGSAMYRVSVKLKALKKVIRDFS--RDNYSDIEKRTKEAHDALLLAQS 217
Query: 314 --LASEEPG---IELRKNRLTILEGLWQEYRKEEYRW-RQKARVRWLREGDRNTSFFHSI 367
LAS P IE R W+ + E + Q++RV WLREGD N+S+FH +
Sbjct: 218 VLLASPCPSNAAIEAETQRK------WRILAEAEASFFYQRSRVNWLREGDMNSSYFHKM 271
Query: 368 SKIRKSKKLISQLVVG-GVSIEDPNMIKDAIFSHFQSFFTSENRLRPKIRCANLSRL--- 423
+ R+S I L G IE +++ +FQS SE L P A++S L
Sbjct: 272 ASARQSLNHIHFLSDPVGDRIEGQQNLENHCVEYFQSNLGSEQGL-PLFEQADISNLLSY 330
Query: 424 --SQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKEDVVKFFEDF 481
S A ++SL+ FS +I S N+A G DGF+ FF W I+ +V + +F
Sbjct: 331 RCSPAQQVSLDTPFSSEQIKNAFFSLPRNKASGPDGFSPEFFCACWPIIGGEVTEAIHEF 390
Query: 482 HSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQVVTPLLIS 541
++GKL+K NAT + LIPK S++SDFRPIS + + YK+ISK+L RL+ P IS
Sbjct: 391 FTSGKLLKQWNATNLVLIPKITNASSMSDFRPISCLNTVYKVISKLLTDRLKDFLPAAIS 450
Query: 542 ENQFAFAQGRQISDCILIANEVVDFLNKKQ-GGGFLLKLDFAKDYDNIDWGFLFDILKEM 600
+Q AF GR + +L+A E+V NKK +LK+D K +D++ W F+ L+ +
Sbjct: 451 HSQSAFMPGRLFLENVLLATELVHGYNKKNIAPSSMLKVDLRKAFDSVRWDFIVSALRAL 510
Query: 601 NFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCVNGLSCLLN 660
N +++ WI C+STA+ S+++NG S G F KGLRQGDP+SP+LF L + S LL
Sbjct: 511 NVPEKFTCWILECLSTASFSVILNGHSAGHFWSSKGLRQGDPMSPYLFVLAMEVFSGLLQ 570
Query: 661 QLLDDSNFSGVRIGPGLALNHLQFADDTLLFCENDENQIDLLCNTLLSFLFTSGLKVNLH 720
+ L ++HL FADD ++F + + + + +L F SGL +N +
Sbjct: 571 SRYTSGYIAYHPKTSQLEISHLMFADDVMIFFDGKSSSLHGIVESLEDFAGWSGLLMNTN 630
Query: 721 KSTIIGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRRASFWEPMLDKLRRKRRT 780
K+ + + E+ +A YG+ +G LP+ YLG PL + + P+++K+ + +
Sbjct: 631 KTQLYHAGLSQSESDSMAS-YGFKLGSLPVRYLGLPLMSRKLTIAEYAPLIEKITARFNS 689
Query: 781 WNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKILRSFLQGGKL 833
W + +S +GR+ L+ + + + F ++ F PL + ++E + FL ++
Sbjct: 690 WVVRLLSFAGRVQLLASVISGIVNFWISSFILPLGCIKKIESLCSRFLWSSRI 742
>gb|AAC13599.1| similar to reverse transcriptase (Pfam: transcript_fact.hmm, score:
72.31) [Arabidopsis thaliana] gi|7488318|pir||T01191
RNA-directed DNA polymerase homolog F21E10.5 -
Arabidopsis thaliana
Length = 928
Score = 325 bits (832), Expect = 9e-87
Identities = 221/719 (30%), Positives = 352/719 (48%), Gaps = 31/719 (4%)
Query: 138 IMGGDFNAILNEGERIGIQMG--VESSFSDF---VADSNLIDLPLQSDKFTWYSSRAGGL 192
I+ GDFN IL+ E + + DF V ++ DL FTW + R L
Sbjct: 26 IIFGDFNEILDMEEHSNSRENPVTTTGMRDFQMAVNHCSITDLAYHGPLFTWSNKRENDL 85
Query: 193 WSR-IDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSL----GAINFGPKPFLFYNHW 247
++ +DR LV++ + F E G SD ++L GA+ G +PF F N
Sbjct: 86 IAKKLDRVLVNDVWLQSFPRSYSVFEAGGCSDHLRCRINLNVGAGAVVKGKRPFKFVNVI 145
Query: 248 LLDKEFEKLVSDWWCSATTVGWSA---FVLQQKLKGLK----NRIKEWQGYASSVSRVRI 300
+ F V +W + S F +KLKGLK N KE G ++
Sbjct: 146 TEMEHFIPTVESYWNETEAIFMSTSSLFRFSKKLKGLKPLLRNLGKERLGNLVKQTKEAF 205
Query: 301 NTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRW-RQKARVRWLREGDR 359
T+ Q ++A+ P +N W E ++ +Q++++ WL GDR
Sbjct: 206 ETL---CQKQAMKMANPSPSSMQEENEAY---AKWDHIAVLEEKFLKQRSKLHWLDIGDR 259
Query: 360 NTSFFHSISKIRKSKKLISQLVVGGVSI-EDPNMIKDAIFSHFQSFFTSENRLRPKIRCA 418
N FH R+++ I +++ S+ IK HF+ F I
Sbjct: 260 NNKAFHRAVVAREAQNSIREIICHDGSVASQEEKIKTEAEHHFREFLQLIPNDFEGIAVE 319
Query: 419 NLS-----RLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWGIVKED 473
L R S ++K L S EI +V+ S +++PG DG+ F+K W I+ +
Sbjct: 320 ELQDLLPYRCSDSDKEMLTNHVSAEEIHKVVFSMPNDKSPGPDGYTAEFYKGAWNIIGAE 379
Query: 474 VVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVLASRLQ 533
+ + F + G L KG+N+T ++LIPK + D+RPIS YK+ISK++A+RL+
Sbjct: 380 FILAIQSFFAKGFLPKGINSTILALIPKKKEAKEMKDYRPISCCNVLYKVISKIIANRLK 439
Query: 534 VVTPLLISENQFAFAQGRQISDCILIANEVV-DFLNKKQGGGFLLKLDFAKDYDNIDWGF 592
+V P I NQ AF + R + + +L+A E+V D+ LK+D +K +D++ W F
Sbjct: 440 LVLPKFIVGNQSAFVKDRLLIENVLLATEIVKDYHKDSVSSRCALKIDISKAFDSVQWKF 499
Query: 593 LFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSPFLFNLCV 652
L ++L+ MNF + +WI C++TA+ S+ VNG G F + LRQG LSP+LF + +
Sbjct: 500 LINVLEAMNFPPEFTHWITLCITTASFSVQVNGELAGVFSSARELRQGCSLSPYLFVISM 559
Query: 653 NGLSCLLNQLLDDSNFSGVRIGPGLALNHLQFADDTLLFCENDENQIDLLCNTLLSFLFT 712
+ LS +L++ + F + L HL FADD ++ + ID + L F
Sbjct: 560 DVLSKMLDKAVGARQFGYHPKCRAIGLTHLSFADDLMILSDGKVRSIDGIVKVLYEFAKW 619
Query: 713 SGLKVNLHKSTIIGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRRASFWEPMLD 772
SGLK+++ KST+ V Q + + + +G+LP+ YLG PL AS P+++
Sbjct: 620 SGLKISMEKSTMYLAGVQASVYQEIVQKFSFDVGKLPVRYLGLPLVSKRLTASDCLPLIE 679
Query: 773 KLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKILRSFLQGG 831
+LR+K W S+F+S +GRL L+ +TL S+ F MA F+ P + E++K+ +FL G
Sbjct: 680 QLRKKIEAWTSRFLSFAGRLNLISSTLWSICNFWMAAFRLPRACIREIDKLCSAFLWSG 738
>gb|AAD21778.1| putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana] gi|25410938|pir||G84429 hypothetical protein
At2g01840 [imported] - Arabidopsis thaliana
Length = 1715
Score = 323 bits (829), Expect = 2e-86
Identities = 233/852 (27%), Positives = 408/852 (47%), Gaps = 46/852 (5%)
Query: 6 WNVRGLGRAEKRRAVVEALSGLKPEIVLVQESKLDSSRVKTVEAFANAVKM---DYVSVD 62
WN +GLG+ R + E +++ + E+K + + + VKM D +
Sbjct: 368 WNCQGLGQPLTVRRLEEVQRVYFLDMLFLIETKQQDNYTRDL-----GVKMGFEDMCIIS 422
Query: 63 AVGSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLFIMNVYGPNVDSERYDFF 122
G +GGL+ W +++ +VR + L + + ++ +YG + SER+ +
Sbjct: 423 PRGLSGGLVVYWKK-HLSIQVISHDVRLVDLYVE--YKNFNFYLSCIYGHPIPSERHHLW 479
Query: 123 NLLSEALSLNNCFCKIMGGDFNAILNEGERIGIQ---MGVESSFSDFVADSNLIDLPLQS 179
L + +S + +M GDFN ILN E+ G + +G +F++ + N+ DL +
Sbjct: 480 EKL-QRVSAHRSGPWMMCGDFNEILNLNEKKGGRRRSIGSLQNFTNMINCCNMKDLKSKG 538
Query: 180 DKFTWYSSRAGG-LWSRIDRWLVSEDAINKFDGISQSVENWGLSDRRPIILSL------- 231
+ ++W R + S +DR ++ D F SD P+I+ +
Sbjct: 539 NPYSWVGKRQNETIESCLDRVFINSDWQASFPAFETEFLPIAGSDHAPVIIDIAEEVCTK 598
Query: 232 -GAINFGPKPFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIKEWQG 290
G + + F F ++F V W + + +KL + + +W+
Sbjct: 599 RGQFRYDRRHFQF-------EDFVDSVQRGWNRGRSDSHGGYY--EKLHCCRQELAKWKR 649
Query: 291 YASSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILEGLWQEYRKEEYRWRQKAR 350
+ + +I T++ + A+E + L + + L Q YR EE W K+R
Sbjct: 650 RTKTNTAEKIETLKYRVD------AAERDHTLPHQTILRLRQDLNQAYRDEELYWHLKSR 703
Query: 351 VRWLREGDRNTSFFHSISKIRKSKKLISQLV--VGGVSIEDPNMIKDAIFSHFQSFFTSE 408
RW+ GDRNT FF++ +K+RKS+ I + G + D + K A F T++
Sbjct: 704 NRWMLLGDRNTMFFYASTKLRKSRNRIKAITDAQGIENFRDDTIGKVAENYFADLFTTTQ 763
Query: 409 NRLRPKIRCANLSRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFFKQFWG 468
+I ++++ L + ++ E+ + + + +RAPG DGF F+ W
Sbjct: 764 TSDWEEIISGIAPKVTEQMNHELLQSVTDQEVRDAVFAIGADRAPGFDGFTAAFYHHLWD 823
Query: 469 IVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKLISKVL 528
++ DV F + + +N T I LIPK P +SD+RPISL ++YK+ISK+L
Sbjct: 824 LIGNDVCLMVRHFFESDVMDNQINQTQICLIPKIIDPKHMSDYRPISLCTASYKIISKIL 883
Query: 529 ASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFLNKK---QGGGFLLKLDFAKDY 585
RL+ +IS++Q AF G+ ISD +L+A+E++ L + Q G +K D +K Y
Sbjct: 884 IKRLKQCLGDVISDSQAAFVPGQNISDNVLVAHELLHSLKSRRECQSGYVAVKTDISKAY 943
Query: 586 DNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQGDPLSP 645
D ++W FL ++ ++ F RW+ WI TCV++ + +L+NGS G +G+RQGDPLSP
Sbjct: 944 DRVEWNFLEKVMIQLGFAPRWVKWIMTCVTSVSYEVLINGSPYGKIFPSRGIRQGDPLSP 1003
Query: 646 FLFNLCVNGLSCLLNQLLDDSNFSGVRIGPG-LALNHLQFADDTLLFCENDENQIDLLCN 704
+LF C LS +L + + G++I LA++HL FADD+L FC I+ L
Sbjct: 1004 YLFLFCAEVLSNMLRKAEVNKQIHGMKITKDCLAISHLLFADDSLFFCRASNQNIEQLAL 1063
Query: 705 TLLSFLFTSGLKVNLHKSTII-GCNVDVRETQRVAGMYGWSIGQLPILYLGAPLGGNPRR 763
+ SG K+N KS+II G + QR+ + G + YLG P R+
Sbjct: 1064 IFKKYEEASGQKINYAKSSIIFGQKIPTMRRQRLHRLLGIDNVRGGGKYLGLPEQLGRRK 1123
Query: 764 ASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVGEVEKI 823
+E ++ K++ + W ++S +G+ ++KA ++P++ M F P + E+ +
Sbjct: 1124 VELFEYIVTKVKERTEGWAYNYLSPAGKEIVIKAIAMALPVYSMNCFLLPTLICNEINSL 1183
Query: 824 LRSFLQGGKLHG 835
+ +F G + G
Sbjct: 1184 ITAFWWGKENEG 1195
>pir||G86379 protein F5A9.24 [imported] - Arabidopsis thaliana
gi|9945082|gb|AAG03119.1| F5A9.24 [Arabidopsis thaliana]
Length = 1254
Score = 323 bits (829), Expect = 2e-86
Identities = 237/789 (30%), Positives = 398/789 (50%), Gaps = 42/789 (5%)
Query: 60 SVDAVGSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLF------IMNVYGPN 113
+V+ VG GGL LW S +++F+ + L+D+ + F + VYG
Sbjct: 24 TVEPVGKCGGLALLWK------SSVQVDLKFVD---KNLMDAQVQFGAVNFCVSCVYGDP 74
Query: 114 VDSERYDFFNLLSE-ALSLNNCFCKIMGGDFNAILNEGERIGIQMGVE---SSFSDFVAD 169
S+R + +S + + +C M GDFN IL+ GE+ G + +F++ +
Sbjct: 75 DRSKRSQAWERISRIGVGRRDKWC--MFGDFNDILHNGEKNGGPRRSDLDCKAFNEMIKG 132
Query: 170 SNLIDLPLQSDKFTWYSSRAGGLW--SRIDRWLVSEDAINKFDGISQSVENWGLSDRRPI 227
+L+++P + FTW + R G W R+DR +++ F +Q+ ++ SD RP+
Sbjct: 133 CDLVEMPAHGNGFTW-AGRRGDHWIQCRLDRAFGNKEWFCFFPVSNQTFLDFRGSDHRPV 191
Query: 228 ILSLGAINFGPK-PFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIK 286
++ L + + F F +L ++ ++ + W S G + V +L+ + +
Sbjct: 192 LIKLMSSQDSYRGQFRFDKRFLFKEDVKEAIIRTW-SRGKHGTNISVAD-RLRACRKSLS 249
Query: 287 EWQGYASSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILE-GLWQEYRKEEYRW 345
W+ + S +IN +EA L+ E+ + R+++L+ L + YR+EE W
Sbjct: 250 SWKKQNNLNSLDKINQLEAALE-------KEQSLVWPIFQRVSVLKKDLAKAYREEEAYW 302
Query: 346 RQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVVGGVSIEDPNMIKDAIFS-HFQSF 404
+QK+R +WLR G+RN+ +FH+ K + +K I +L +++ K + + +F +
Sbjct: 303 KQKSRQKWLRSGNRNSKYFHAAVKQNRQRKRIEKLKDVNGNMQTSEAAKGEVAAAYFGNL 362
Query: 405 FTSENRLRPKIRCANL-SRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFF 463
F S N + L R+S+ SL + S EI E + S APG DG + FF
Sbjct: 363 FKSSNPSGFTDWFSGLVPRVSEVMNESLVGEVSAQEIKEAVFSIKPASAPGPDGMSALFF 422
Query: 464 KQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKL 523
+ +W V V + F + G + N T + LIPK P+ + D RPISL YK+
Sbjct: 423 QHYWSTVGNQVTSEVKKFFADGIMPAEWNYTHLCLIPKTQHPTEMVDLRPISLCSVLYKI 482
Query: 524 ISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFL--NKKQGGGFL-LKLD 580
ISK++A RLQ P ++S+ Q AF R I+D IL+A+E+V L + + F+ +K D
Sbjct: 483 ISKIMAKRLQPWLPEIVSDTQSAFVSERLITDNILVAHELVHSLKVHPRISSEFMAVKSD 542
Query: 581 FAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQG 640
+K YD ++W +L +L + F +W+NWI CVS+ T S+L+N G +++GLRQG
Sbjct: 543 MSKAYDRVEWSYLRSLLLSLGFHLKWVNWIMVCVSSVTYSVLINDCPFGLIILQRGLRQG 602
Query: 641 DPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGP-GLALNHLQFADDTLLFCENDENQI 699
DPLSPFLF LC GL+ LLN+ + G++ G ++HL FADD+L C+ Q
Sbjct: 603 DPLSPFLFVLCTEGLTHLLNKAQWEGALEGIQFSENGPMVHHLLFADDSLFLCKASREQS 662
Query: 700 DLLCNTLLSFLFTSGLKVNLHKSTI-IGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLG 758
+L L + +G +NL+KS+I G VD + + G YLG P
Sbjct: 663 LVLQKILKVYGNATGQTINLNKSSITFGEKVDEQLKGTIRTCLGIFTEGGAGTYLGLPEC 722
Query: 759 GNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVG 818
+ + + D+L+ K W ++ +S G+ L+K+ ++P+F M+ FK P+
Sbjct: 723 FSGSKVDMLHYLKDRLKEKLDVWFTRCLSQGGKEVLLKSVALAMPVFAMSCFKLPITTCE 782
Query: 819 EVEKILRSF 827
+E + SF
Sbjct: 783 NLESAMASF 791
>gb|AAF97969.1| F21J9.30 [Arabidopsis thaliana]
Length = 1270
Score = 323 bits (829), Expect = 2e-86
Identities = 237/789 (30%), Positives = 398/789 (50%), Gaps = 42/789 (5%)
Query: 60 SVDAVGSAGGLISLWNPASYAMESAVKNVRFIGLLLRALVDSSLLF------IMNVYGPN 113
+V+ VG GGL LW S +++F+ + L+D+ + F + VYG
Sbjct: 21 TVEPVGKCGGLALLWK------SSVQVDLKFVD---KNLMDAQVQFGAVNFCVSCVYGDP 71
Query: 114 VDSERYDFFNLLSE-ALSLNNCFCKIMGGDFNAILNEGERIGIQMGVE---SSFSDFVAD 169
S+R + +S + + +C M GDFN IL+ GE+ G + +F++ +
Sbjct: 72 DRSKRSQAWERISRIGVGRRDKWC--MFGDFNDILHNGEKNGGPRRSDLDCKAFNEMIKG 129
Query: 170 SNLIDLPLQSDKFTWYSSRAGGLW--SRIDRWLVSEDAINKFDGISQSVENWGLSDRRPI 227
+L+++P + FTW + R G W R+DR +++ F +Q+ ++ SD RP+
Sbjct: 130 CDLVEMPAHGNGFTW-AGRRGDHWIQCRLDRAFGNKEWFCFFPVSNQTFLDFRGSDHRPV 188
Query: 228 ILSLGAINFGPK-PFLFYNHWLLDKEFEKLVSDWWCSATTVGWSAFVLQQKLKGLKNRIK 286
++ L + + F F +L ++ ++ + W S G + V +L+ + +
Sbjct: 189 LIKLMSSQDSYRGQFRFDKRFLFKEDVKEAIIRTW-SRGKHGTNISVAD-RLRACRKSLS 246
Query: 287 EWQGYASSVSRVRINTIEADLQVTMDRLASEEPGIELRKNRLTILE-GLWQEYRKEEYRW 345
W+ + S +IN +EA L+ E+ + R+++L+ L + YR+EE W
Sbjct: 247 SWKKQNNLNSLDKINQLEAALE-------KEQSLVWPIFQRVSVLKKDLAKAYREEEAYW 299
Query: 346 RQKARVRWLREGDRNTSFFHSISKIRKSKKLISQLVVGGVSIEDPNMIKDAIFS-HFQSF 404
+QK+R +WLR G+RN+ +FH+ K + +K I +L +++ K + + +F +
Sbjct: 300 KQKSRQKWLRSGNRNSKYFHAAVKQNRQRKRIEKLKDVNGNMQTSEAAKGEVAAAYFGNL 359
Query: 405 FTSENRLRPKIRCANL-SRLSQANKMSLEKQFSEGEIWEVLKSCDGNRAPGTDGFNLNFF 463
F S N + L R+S+ SL + S EI E + S APG DG + FF
Sbjct: 360 FKSSNPSGFTDWFSGLVPRVSEVMNESLVGEVSAQEIKEAVFSIKPASAPGPDGMSALFF 419
Query: 464 KQFWGIVKEDVVKFFEDFHSTGKLVKGLNATFISLIPKHNCPSTVSDFRPISLIGSAYKL 523
+ +W V V + F + G + N T + LIPK P+ + D RPISL YK+
Sbjct: 420 QHYWSTVGNQVTSEVKKFFADGIMPAEWNYTHLCLIPKTQHPTEMVDLRPISLCSVLYKI 479
Query: 524 ISKVLASRLQVVTPLLISENQFAFAQGRQISDCILIANEVVDFL--NKKQGGGFL-LKLD 580
ISK++A RLQ P ++S+ Q AF R I+D IL+A+E+V L + + F+ +K D
Sbjct: 480 ISKIMAKRLQPWLPEIVSDTQSAFVSERLITDNILVAHELVHSLKVHPRISSEFMAVKSD 539
Query: 581 FAKDYDNIDWGFLFDILKEMNFGDRWINWIRTCVSTATLSILVNGSSTGFFEMEKGLRQG 640
+K YD ++W +L +L + F +W+NWI CVS+ T S+L+N G +++GLRQG
Sbjct: 540 MSKAYDRVEWSYLRSLLLSLGFHLKWVNWIMVCVSSVTYSVLINDCPFGLIILQRGLRQG 599
Query: 641 DPLSPFLFNLCVNGLSCLLNQLLDDSNFSGVRIGP-GLALNHLQFADDTLLFCENDENQI 699
DPLSPFLF LC GL+ LLN+ + G++ G ++HL FADD+L C+ Q
Sbjct: 600 DPLSPFLFVLCTEGLTHLLNKAQWEGALEGIQFSENGPMVHHLLFADDSLFLCKASREQS 659
Query: 700 DLLCNTLLSFLFTSGLKVNLHKSTI-IGCNVDVRETQRVAGMYGWSIGQLPILYLGAPLG 758
+L L + +G +NL+KS+I G VD + + G YLG P
Sbjct: 660 LVLQKILKVYGNATGQTINLNKSSITFGEKVDEQLKGTIRTCLGIFTEGGAGTYLGLPEC 719
Query: 759 GNPRRASFWEPMLDKLRRKRRTWNSKFISLSGRLTLMKATLCSVPLFLMAIFKAPLKVVG 818
+ + + D+L+ K W ++ +S G+ L+K+ ++P+F M+ FK P+
Sbjct: 720 FSGSKVDMLHYLKDRLKEKLDVWFTRCLSQGGKEVLLKSVALAMPVFAMSCFKLPITTCE 779
Query: 819 EVEKILRSF 827
+E + SF
Sbjct: 780 NLESAMASF 788
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.334 0.147 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,256,620,161
Number of Sequences: 2540612
Number of extensions: 93096300
Number of successful extensions: 304968
Number of sequences better than 10.0: 1316
Number of HSP's better than 10.0 without gapping: 809
Number of HSP's successfully gapped in prelim test: 507
Number of HSP's that attempted gapping in prelim test: 301715
Number of HSP's gapped (non-prelim): 1866
length of query: 1389
length of database: 863,360,394
effective HSP length: 141
effective length of query: 1248
effective length of database: 505,134,102
effective search space: 630407359296
effective search space used: 630407359296
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 82 (36.2 bits)
Lotus: description of TM0110a.13


