

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
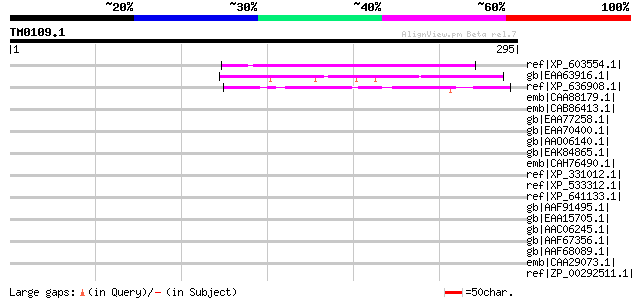
Query= TM0109.1
(295 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_603554.1| PREDICTED: similar to hypothetical protein FLJ3... 50 8e-05
gb|EAA63916.1| hypothetical protein AN2231.2 [Aspergillus nidula... 48 3e-04
ref|XP_636908.1| hypothetical protein DDB0219312 [Dictyostelium ... 47 7e-04
emb|CAA88179.1| gar2 [Schizosaccharomyces pombe] 46 0.001
emb|CAB86413.1| gar2 [Schizosaccharomyces pombe] gi|19114443|ref... 46 0.001
gb|EAA77258.1| hypothetical protein FG07399.1 [Gibberella zeae P... 46 0.001
gb|EAA70400.1| hypothetical protein FG10084.1 [Gibberella zeae P... 46 0.001
gb|AAO06140.1| PC10114w [Plasmodium chabaudi chabaudi] 46 0.001
gb|EAK84865.1| hypothetical protein UM03687.1 [Ustilago maydis 5... 46 0.001
emb|CAH76490.1| hypothetical protein PC102553.00.0 [Plasmodium c... 46 0.001
ref|XP_331012.1| predicted protein [Neurospora crassa] gi|289209... 45 0.002
ref|XP_533312.1| PREDICTED: similar to hypothetical protein [Can... 45 0.002
ref|XP_641133.1| hypothetical protein DDB0220644 [Dictyostelium ... 45 0.002
gb|AAF91495.1| PspA protein [Streptococcus pneumoniae] 45 0.003
gb|EAA15705.1| retinitis pigmentosa GTPase regulator-like protei... 45 0.003
gb|AAC06245.1| neurofilament medium subunit [Serinus canaria] 44 0.004
gb|AAF67356.1| PspA [Streptococcus pneumoniae] 44 0.004
gb|AAF68089.1| PspA [Streptococcus pneumoniae] 44 0.004
emb|CAA29073.1| NF-M c-terminus [Gallus gallus] 44 0.006
ref|ZP_00292511.1| hypothetical protein Tfus02001785 [Thermobifi... 44 0.006
>ref|XP_603554.1| PREDICTED: similar to hypothetical protein FLJ36144, partial [Bos
taurus]
Length = 1369
Score = 50.1 bits (118), Expect = 8e-05
Identities = 37/148 (25%), Positives = 70/148 (47%), Gaps = 2/148 (1%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGE 183
EEE++S +E+++ E E SL E + E ESL+++ E ESL E+ + S E+ G
Sbjct: 686 EEESRSEEESLSEEEE--SLSEEESLGEEEESLSEEESLSEEEESLREEESLSEKEEEGL 743
Query: 184 MEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTT 243
E+ ESL+ + LS + +L+ + K + + + +MEK E
Sbjct: 744 SEEEESLSEEEEGLSEKEKSLSEKEKHLSKKEKALHEKEKSLSEKEEEEEKMEKTEKKEE 803
Query: 244 DVASLSLGDDALASPPPQPKTRRKLAAP 271
+ + +++++ QP+ + P
Sbjct: 804 EKKARQKDEESMSKEKEQPRRLTEAMLP 831
>gb|EAA63916.1| hypothetical protein AN2231.2 [Aspergillus nidulans FGSC A4]
gi|67523551|ref|XP_659835.1| hypothetical protein
AN2231_2 [Aspergillus nidulans FGSC A4]
gi|49089276|ref|XP_406368.1| hypothetical protein
AN2231.2 [Aspergillus nidulans FGSC A4]
Length = 624
Score = 48.1 bits (113), Expect = 3e-04
Identities = 49/188 (26%), Positives = 83/188 (44%), Gaps = 26/188 (13%)
Query: 123 AEEENKSLKENVASETENKSLMEHVAAE--TENESLTKDVPEETENESLTEDVNES---- 176
A+E +S KE+ AS E ++ + AE E + +V +E++ E+L++ V ++
Sbjct: 140 ADESQQSNKESDASAKEPEAEIATQPAEEKAEEKETRTNVEDESQTEALSKPVEQAPGTT 199
Query: 177 ------LTEDVGEMEKNESLTTDVASLSLG----DDALASPPPQP-------KTRRKLVA 219
+ +D G+ K E T VA+L + L++P P K +R+ +
Sbjct: 200 DEAEPKIVQDGGK--KTEPTDTQVAALGAQTPGVNTGLSTPLPAEELIDDVRKRKRRSQS 257
Query: 220 PQPAVRRSARIKTGEMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPA 279
P P + AR K EK+ +L T S+S D + PP Q T + PA
Sbjct: 258 PAPHLEEVARKKAKSAEKS-ALPTPDESISASHDDMQQPPVQSPTPEGKSPETPAKKNTP 316
Query: 280 VRRSARIK 287
++ R K
Sbjct: 317 QKQDVRFK 324
>ref|XP_636908.1| hypothetical protein DDB0219312 [Dictyostelium discoideum]
gi|60465317|gb|EAL63409.1| hypothetical protein
DDB0219312 [Dictyostelium discoideum]
Length = 912
Score = 47.0 bits (110), Expect = 7e-04
Identities = 40/176 (22%), Positives = 73/176 (40%), Gaps = 34/176 (19%)
Query: 125 EENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGEM 184
E+ + +KEN E E + E E ENE+ E ENE+ E+ NE++ E E
Sbjct: 143 EKGEKIKENKEKEKEKEKEKEK---ENENEN-----ENENENENENENENENVNEKEKEK 194
Query: 185 EKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTTD 244
EK + ++ + + P+ K ++++ ++ + +K + +K S
Sbjct: 195 EKEKEKENEIKEKEI---IIKKEEPKEKIKKEV-----RIKENEEVKEKQNKKLSSTLKQ 246
Query: 245 VASLSLGDDAL---------ASPPPQPKTRRKLAAPQPAAPQPAVRRSARIKTTPK 291
+ S+ +D L +PPP P P++P P+ + TPK
Sbjct: 247 LKRFSIHEDVLDYLSNSTLSGAPPP---------PPPPSSPSPSPSTQQKTPQTPK 293
Score = 33.1 bits (74), Expect = 9.8
Identities = 27/102 (26%), Positives = 41/102 (39%), Gaps = 15/102 (14%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDV------PEE--------TENESL 169
E EN++ EN K + E ENE K++ P+E ENE +
Sbjct: 173 ENENENENENENENVNEKEKEKEKEKEKENEIKEKEIIIKKEEPKEKIKKEVRIKENEEV 232
Query: 170 TEDVNESLTEDVGEMEKNESLTTDVASLSLGDDALASPPPQP 211
E N+ L+ + ++ K S+ DV +PPP P
Sbjct: 233 KEKQNKKLSSTLKQL-KRFSIHEDVLDYLSNSTLSGAPPPPP 273
>emb|CAA88179.1| gar2 [Schizosaccharomyces pombe]
Length = 500
Score = 46.2 bits (108), Expect = 0.001
Identities = 33/130 (25%), Positives = 58/130 (44%), Gaps = 2/130 (1%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGE 183
EEE +K E+ + S E ++E+E+ES + + EE E TE+ E +E +
Sbjct: 148 EEEEAVVKIEEKKESSSDSSSESSSSESESESSSSESEEEEEVVEKTEEKKEGSSESSSD 207
Query: 184 MEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTT 243
E + +++ D+ + + + +RK A + R A+I + NE+ T
Sbjct: 208 SESSSDSSSESGDSDSSSDSESESSSEDEKKRK--AEPASEERPAKITKPSQDSNETCTV 265
Query: 244 DVASLSLGDD 253
V LS D
Sbjct: 266 FVGRLSWNVD 275
>emb|CAB86413.1| gar2 [Schizosaccharomyces pombe] gi|19114443|ref|NP_593531.1|
hypothetical protein SPAC140.02 [Schizosaccharomyces
pombe 972h-] gi|6226864|sp|P41891|GAR2_SCHPO Protein
gar2
Length = 500
Score = 46.2 bits (108), Expect = 0.001
Identities = 33/130 (25%), Positives = 58/130 (44%), Gaps = 2/130 (1%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGE 183
EEE +K E+ + S E ++E+E+ES + + EE E TE+ E +E +
Sbjct: 148 EEEEAVVKIEEKKESSSDSSSESSSSESESESSSSESEEEEEVVEKTEEKKEGSSESSSD 207
Query: 184 MEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTT 243
E + +++ D+ + + + +RK A + R A+I + NE+ T
Sbjct: 208 SESSSDSSSESGDSDSSSDSESESSSEDEKKRK--AEPASEERPAKITKPSQDSNETCTV 265
Query: 244 DVASLSLGDD 253
V LS D
Sbjct: 266 FVGRLSWNVD 275
>gb|EAA77258.1| hypothetical protein FG07399.1 [Gibberella zeae PH-1]
gi|46126043|ref|XP_387575.1| hypothetical protein
FG07399.1 [Gibberella zeae PH-1]
Length = 314
Score = 45.8 bits (107), Expect = 0.001
Identities = 43/159 (27%), Positives = 71/159 (44%), Gaps = 18/159 (11%)
Query: 137 ETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGEMEKNESLTTDVAS 196
E++N S E E E E + D +E E+E + +E +ED E E+NE D++S
Sbjct: 30 ESQNSSSEEDDDDEVEEEGIRGDSDDEDEDEDEEDQESEDGSEDESEPEQNEP-KIDLSS 88
Query: 197 LSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTTDVASLSLGDDALA 256
+S G ALA K A P+ R+S K+ + + ++ T + D
Sbjct: 89 VSFG--ALA----------KAQASLPSGRKSKNKKSTDEDTPKTETPAPRKPTRSKD--- 133
Query: 257 SPPPQPKTRRKLAAPQPAAPQPAVRRSARIKTTPKKFKD 295
P+PK K A + + +P RR I ++++D
Sbjct: 134 --DPKPKRSSKHAPQEQTSKKPVSRRREIIPENKRQYRD 170
>gb|EAA70400.1| hypothetical protein FG10084.1 [Gibberella zeae PH-1]
gi|46137137|ref|XP_390260.1| hypothetical protein
FG10084.1 [Gibberella zeae PH-1]
Length = 4221
Score = 45.8 bits (107), Expect = 0.001
Identities = 40/171 (23%), Positives = 73/171 (42%), Gaps = 18/171 (10%)
Query: 122 TAEEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEET--ENESLTE-DVNESLT 178
T + E+KS+ + +E VA + V ET ++E+ TE +V E+
Sbjct: 1113 TVDSESKSVTGPATEPEATEKAIESVAEPEDAAKPETVVKPETVIDSEAPTESEVVETAV 1172
Query: 179 EDVGEMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKN 238
E E E T + S + D++ A P+PK K + P + ++
Sbjct: 1173 EPTAEPEIAPEPETVIDSEAPSDESAAEATPEPKVVEKAIEP--------------VSES 1218
Query: 239 ESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPAVRRSARIKTT 289
E+ + A+ DD++ P +P+ K+A +PA +P + +A K +
Sbjct: 1219 EAAVENEAARPESDDSVVQPTTEPEVAEKVAVDEPAL-EPELETAAEAKAS 1268
Score = 33.9 bits (76), Expect = 5.7
Identities = 38/161 (23%), Positives = 57/161 (34%), Gaps = 11/161 (6%)
Query: 123 AEEENKSLKENVASETENKSLMEHVAAETENE-----SLTKDVPEETENESLTEDVNESL 177
AE E + +KE AS +E K E A+ + E + + PEE E + E +
Sbjct: 2229 AEAEVEPVKE--ASVSEEKLAEEEQTAQAQEEPQQGSAPADNTPEEEEKSAKDEHIEAPT 2286
Query: 178 TEDVGEMEKNESLTTDVASLSLGDDAL-ASPPPQPKTRRKLVAPQPAVRRSARIKTGEME 236
T D E +S AS + +D L SPP + K + +
Sbjct: 2287 TSDDKAGEDVKSAEEPPASAPIAEDELNQSPPTAAQEESKETTTAEETTEATPVAEPVSA 2346
Query: 237 KNESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQ 277
E D S DD P + A +P +P+
Sbjct: 2347 SEEPPAGDTVST---DDKAVDDPKNAEDVATPAVQEPCSPE 2384
>gb|AAO06140.1| PC10114w [Plasmodium chabaudi chabaudi]
Length = 577
Score = 45.8 bits (107), Expect = 0.001
Identities = 22/71 (30%), Positives = 44/71 (60%), Gaps = 5/71 (7%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDV-----NESLT 178
+EEN++ E + + EN++ E + + ENE+ +++ E+ ENE+ E++ NE+
Sbjct: 376 DEENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNENENKD 435
Query: 179 EDVGEMEKNES 189
E++GE +NE+
Sbjct: 436 EELGEDNENEN 446
Score = 43.9 bits (102), Expect = 0.006
Identities = 21/71 (29%), Positives = 43/71 (59%), Gaps = 5/71 (7%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDV-----NESLT 178
+ EN++ E + + EN++ E + + ENE+ +++ E+ ENE+ E++ NE+
Sbjct: 389 DNENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNENENKD 448
Query: 179 EDVGEMEKNES 189
E++GE +NE+
Sbjct: 449 EELGEDNENEN 459
Score = 43.9 bits (102), Expect = 0.006
Identities = 21/71 (29%), Positives = 43/71 (59%), Gaps = 5/71 (7%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDV-----NESLT 178
+ EN++ E + + EN++ E + + ENE+ +++ E+ ENE+ E++ NE+
Sbjct: 402 DNENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNENENKD 461
Query: 179 EDVGEMEKNES 189
E++GE +NE+
Sbjct: 462 EELGEDNENEN 472
Score = 43.5 bits (101), Expect = 0.007
Identities = 21/70 (30%), Positives = 42/70 (60%), Gaps = 5/70 (7%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDV-----NESLT 178
+ EN++ E + + EN++ E + + ENE+ +++ E+ ENE+ E++ NE+
Sbjct: 415 DNENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNENENKD 474
Query: 179 EDVGEMEKNE 188
E++GE KN+
Sbjct: 475 EELGEDNKNK 484
Score = 37.0 bits (84), Expect = 0.68
Identities = 13/58 (22%), Positives = 36/58 (61%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDV 181
+ EN++ E + + EN++ E + + ENE+ +++ E+ +N+ E++++ + E++
Sbjct: 441 DNENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNKNKDKGEEIDDEIDEEL 498
>gb|EAK84865.1| hypothetical protein UM03687.1 [Ustilago maydis 521]
gi|49074320|ref|XP_401302.1| hypothetical protein
UM03687.1 [Ustilago maydis 521]
Length = 460
Score = 45.8 bits (107), Expect = 0.001
Identities = 32/109 (29%), Positives = 43/109 (39%), Gaps = 3/109 (2%)
Query: 183 EMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQ-PAVRRSARIKTGEMEKNESL 241
++ +NES D G A A P P + K V P P +R A +
Sbjct: 200 DIRQNESFIKDFVGKRGGPGASARPTPAAAPKAKKVPPPAPTSKRKAPPPPPAPRSRHAA 259
Query: 242 TTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPAVRRSARIKTTP 290
T AS S A A PPP P++ A P P P P R+ + + P
Sbjct: 260 TASTASASSSAPAPAPPPPPPRSGTAAATPPPPPPPPL--RAPAVSSAP 306
>emb|CAH76490.1| hypothetical protein PC102553.00.0 [Plasmodium chabaudi]
Length = 526
Score = 45.8 bits (107), Expect = 0.001
Identities = 22/71 (30%), Positives = 44/71 (60%), Gaps = 5/71 (7%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDV-----NESLT 178
+EEN++ E + + EN++ E + + ENE+ +++ E+ ENE+ E++ NE+
Sbjct: 325 DEENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNENENKD 384
Query: 179 EDVGEMEKNES 189
E++GE +NE+
Sbjct: 385 EELGEDNENEN 395
Score = 43.9 bits (102), Expect = 0.006
Identities = 21/71 (29%), Positives = 43/71 (59%), Gaps = 5/71 (7%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDV-----NESLT 178
+ EN++ E + + EN++ E + + ENE+ +++ E+ ENE+ E++ NE+
Sbjct: 338 DNENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNENENKD 397
Query: 179 EDVGEMEKNES 189
E++GE +NE+
Sbjct: 398 EELGEDNENEN 408
Score = 43.9 bits (102), Expect = 0.006
Identities = 21/71 (29%), Positives = 43/71 (59%), Gaps = 5/71 (7%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDV-----NESLT 178
+ EN++ E + + EN++ E + + ENE+ +++ E+ ENE+ E++ NE+
Sbjct: 351 DNENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNENENKD 410
Query: 179 EDVGEMEKNES 189
E++GE +NE+
Sbjct: 411 EELGEDNENEN 421
Score = 43.5 bits (101), Expect = 0.007
Identities = 21/70 (30%), Positives = 42/70 (60%), Gaps = 5/70 (7%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDV-----NESLT 178
+ EN++ E + + EN++ E + + ENE+ +++ E+ ENE+ E++ NE+
Sbjct: 364 DNENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNENENKD 423
Query: 179 EDVGEMEKNE 188
E++GE KN+
Sbjct: 424 EELGEDNKNK 433
Score = 37.0 bits (84), Expect = 0.68
Identities = 13/58 (22%), Positives = 36/58 (61%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDV 181
+ EN++ E + + EN++ E + + ENE+ +++ E+ +N+ E++++ + E++
Sbjct: 390 DNENENKDEELGEDNENENKDEELGEDNENENKDEELGEDNKNKDKGEEIDDEIDEEL 447
>ref|XP_331012.1| predicted protein [Neurospora crassa] gi|28920929|gb|EAA30259.1|
predicted protein [Neurospora crassa]
Length = 577
Score = 45.4 bits (106), Expect = 0.002
Identities = 44/174 (25%), Positives = 68/174 (38%), Gaps = 15/174 (8%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGE 183
EE S ASE+E++S +E+E E K V + E + V ++ +D
Sbjct: 117 EESESSSDSESASESESES-----ESESEVEDKKKPVKKAEEKKPAAPAVKKAAKKDESS 171
Query: 184 MEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLTT 243
E + + S D P PK AP +++A+ T + ES
Sbjct: 172 SESSSEEESGSGSDESSSDDEEETKPAPKATTPKTAPAKTKQQTAKQPTPSSSEEES--- 228
Query: 244 DVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQ--PAVRRSARIKTTPKKFKD 295
+S S D+ P P+ T KLA P P+ P V +++ T K D
Sbjct: 229 --SSESESDE---EPAPKANTSAKLAKPASKTPEAKPVVNGTSKSNETVSKSDD 277
Score = 33.9 bits (76), Expect = 5.7
Identities = 39/171 (22%), Positives = 68/171 (38%), Gaps = 24/171 (14%)
Query: 123 AEEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVG 182
A ++++S E+ +SE E+ S + +++ E E TK P+ T ++ + +
Sbjct: 164 AAKKDESSSES-SSEEESGSGSDESSSDDEEE--TKPAPKATTPKTAPAKTKQQTAKQPT 220
Query: 183 EMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLT 242
E +++ S D+ P P+ T KL P + + G + NE+++
Sbjct: 221 PSSSEEESSSESES----DE---EPAPKANTSAKLAKPASKTPEAKPVVNGTSKSNETVS 273
Query: 243 -TDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPAVRRSARIKTTPKK 292
+D S S D+ S APQP KT PKK
Sbjct: 274 KSDDESSSEEDEESGSEEESSSDEEMADAPQP-------------KTAPKK 311
>ref|XP_533312.1| PREDICTED: similar to hypothetical protein [Canis familiaris]
Length = 1044
Score = 45.1 bits (105), Expect = 0.002
Identities = 30/140 (21%), Positives = 65/140 (46%), Gaps = 13/140 (9%)
Query: 123 AEEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENE-SLTEDVNESLTEDV 181
++ EN+ L AS++EN+ + +H A+++E E L K ++E E +L +++S +E+
Sbjct: 259 SDSENEELLNGHASDSENEDVRKHPASDSEVEELPKSPASDSETEDALKPQISDSESEEP 318
Query: 182 GEMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESL 241
+ + ++S ++ P P+ P+P V S + + ++S
Sbjct: 319 QQHQASDSENEEL------------PKPRISDSESEELPKPRVSDSESEEPQRTQASDSE 366
Query: 242 TTDVASLSLGDDALASPPPQ 261
++ + D PPP+
Sbjct: 367 NEELPKPRISDSESEDPPPR 386
>ref|XP_641133.1| hypothetical protein DDB0220644 [Dictyostelium discoideum]
gi|60469158|gb|EAL67154.1| CHD gene family protein
containing chromodomain, helicase domain, and
DNA-binding domain [Dictyostelium discoideum]
Length = 2373
Score = 45.1 bits (105), Expect = 0.002
Identities = 34/161 (21%), Positives = 70/161 (43%), Gaps = 4/161 (2%)
Query: 123 AEEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVG 182
++ N K++ S+ KS + +E ++ + KD E + E E+ E E++
Sbjct: 57 SDNNNNKNKKSTKSKNSKKSNLSGEDSENDDNDVEKDENENNDVEMKDEEEEEEEEEELE 116
Query: 183 EMEKNESL--TTDVASLSLGDD--ALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKN 238
E E+ + + D++ S G +++ PPP + K +A + S + EK+
Sbjct: 117 EEEEEDDSEDSDDLSKKSRGRPKASVSKPPPPKVPKSKPIAKTNNKKNSNTKEVIIKEKS 176
Query: 239 ESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPA 279
+ TT + + A +PPP + + P+ + + A
Sbjct: 177 KITTTTTTTTTATTTATITPPPPKPVSKPVGKPKGSVSKKA 217
>gb|AAF91495.1| PspA protein [Streptococcus pneumoniae]
Length = 227
Score = 44.7 bits (104), Expect = 0.003
Identities = 41/166 (24%), Positives = 67/166 (39%), Gaps = 10/166 (6%)
Query: 123 AEEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVG 182
AE+ N +N ++ ENK + E T +S KD E + + E ++++LT+
Sbjct: 52 AEDANIEALQNKVADLENK-VAELDKEVTRLQSDLKDAEENNVEDYVKEGLDKALTDKKV 110
Query: 183 EMEKNE--------SLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPA-VRRSARIKTG 233
E+ + +L T + L D +P P PK + P+PA + A
Sbjct: 111 ELNNTQKALDTAQKALDTALNELGPDGDEEETPAPAPKPEQPAEQPKPAPAPQPAPAPKP 170
Query: 234 EMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPA 279
E ++ D A S + PK + AP+P P PA
Sbjct: 171 EKTDDQQAEEDYARRSEEEYNRLPQQQPPKAEKPAPAPKPEQPVPA 216
>gb|EAA15705.1| retinitis pigmentosa GTPase regulator-like protein [Plasmodium
yoelii yoelii]
Length = 725
Score = 44.7 bits (104), Expect = 0.003
Identities = 32/89 (35%), Positives = 53/89 (58%), Gaps = 9/89 (10%)
Query: 125 EENKSLKENVASETENKSL-MEHVAAETENESL-TKDVPEETENESL-TEDVNESL---- 177
E N E+ + + ENKSL E + + EN+SL T+D+P++ EN+SL TED ++ +
Sbjct: 111 ENNFLDTEDTSQDIENKSLDTEDIPQDIENKSLDTEDIPQDIENKSLDTEDNSQDIENNF 170
Query: 178 --TEDVGEMEKNESLTTDVASLSLGDDAL 204
TED + +N+SL T+ S + + +L
Sbjct: 171 LDTEDTSQDIENKSLDTEDNSQDIENKSL 199
Score = 40.8 bits (94), Expect = 0.047
Identities = 41/139 (29%), Positives = 70/139 (49%), Gaps = 14/139 (10%)
Query: 126 ENKSLK-ENVASETENKSL-MEHVAAETENESL-TKDVPEETENESLTEDVNESLTEDVG 182
ENKSL E+ + + ENKSL E + + EN+SL T+D ++ EN+SL + N E+
Sbjct: 223 ENKSLDTEDNSQDIENKSLDTEDNSQDIENKSLDTEDNSQDIENKSLDTEDNSQDIENNS 282
Query: 183 EMEKN-----ESLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPAVRRSARI--KTGEM 235
++ +N E L D+ + L + ++ Q + L A + ++ R R K G
Sbjct: 283 QVTENPPLSIEQLFLDIEKIFLDTERIS----QNTEKNSLHAKRRSLNRERRSLNKEGRS 338
Query: 236 EKNESLTTDVASLSLGDDA 254
ES + + S SL +++
Sbjct: 339 LNKESRSLNKESRSLNNES 357
Score = 37.7 bits (86), Expect = 0.40
Identities = 33/89 (37%), Positives = 52/89 (58%), Gaps = 10/89 (11%)
Query: 126 ENKSLK-ENVASETENKSL-MEHVAAETENESL-TKDVPEETENESL-TEDVNESL---- 177
ENKSL E+ + + ENKSL E + + EN L T+D ++ EN+SL TED ++ +
Sbjct: 181 ENKSLDTEDNSQDIENKSLDTEDNSQDIENNFLDTEDTSQDIENKSLDTEDNSQDIENKS 240
Query: 178 --TEDVGEMEKNESLTTDVASLSLGDDAL 204
TED + +N+SL T+ S + + +L
Sbjct: 241 LDTEDNSQDIENKSLDTEDNSQDIENKSL 269
Score = 37.7 bits (86), Expect = 0.40
Identities = 34/103 (33%), Positives = 54/103 (52%), Gaps = 24/103 (23%)
Query: 126 ENKSLK-ENVASETENKSL---------------MEHVAAETENESL-TKDVPEETENES 168
ENKSL E++ + ENKSL E + + EN+SL T+D ++ EN+S
Sbjct: 139 ENKSLDTEDIPQDIENKSLDTEDNSQDIENNFLDTEDTSQDIENKSLDTEDNSQDIENKS 198
Query: 169 L-TEDVNESL------TEDVGEMEKNESLTTDVASLSLGDDAL 204
L TED ++ + TED + +N+SL T+ S + + +L
Sbjct: 199 LDTEDNSQDIENNFLDTEDTSQDIENKSLDTEDNSQDIENKSL 241
>gb|AAC06245.1| neurofilament medium subunit [Serinus canaria]
Length = 487
Score = 44.3 bits (103), Expect = 0.004
Identities = 40/184 (21%), Positives = 71/184 (37%), Gaps = 17/184 (9%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGE 183
+EE K+ +E V E ++ E A E E E + E ++++ E +E E++ E
Sbjct: 115 QEEEKAEEEAVEEEAVSEKAAEQAAEEEEKEEEEAEEEEAAKSDAAEEGGSEK--EEIEE 172
Query: 184 MEKNESLTTDVASLS-----LGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKN 238
E+ E + A G P K+ K P ++ ++ E
Sbjct: 173 KEEGEEAEEEEAEAKGKAEEAGAKVEKVKSPPAKSPPKSPPKSPVTEQAKAVQKAAAEVG 232
Query: 239 ESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQ----------PAVRRSARIKT 288
+ + A+ + A+ P +P T + + +PA P+ P RS T
Sbjct: 233 KDQKAEKAAEKAAKEEKAASPEKPATPKVTSPEKPATPEKPPTPEKAITPEKVRSPEKPT 292
Query: 289 TPKK 292
TP+K
Sbjct: 293 TPEK 296
>gb|AAF67356.1| PspA [Streptococcus pneumoniae]
Length = 246
Score = 44.3 bits (103), Expect = 0.004
Identities = 41/157 (26%), Positives = 62/157 (39%), Gaps = 9/157 (5%)
Query: 131 KENVASETENK--SLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVG----EM 184
KE +E K +L VA E S +D ++ E ++ + + E L E + E+
Sbjct: 78 KEAAEAELNKKVEALQNQVAELEEELSKLEDNLKDAETNNVEDYIKEGLEEAIATKKAEL 137
Query: 185 EKNESLTTDVASLSLGDDA--LASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKNESLT 242
EK + D A LG D +P P P+ + AP P + A E ++
Sbjct: 138 EKTQK-ELDAALNELGPDGDEEETPAPAPQPEKPAPAPAPKPEQPAPAPKPEKSADQQAE 196
Query: 243 TDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPA 279
D A S + + PK + AP P QPA
Sbjct: 197 EDYARRSEEEYNRLTQQQPPKAEKPAPAPAPKPEQPA 233
>gb|AAF68089.1| PspA [Streptococcus pneumoniae]
Length = 256
Score = 44.3 bits (103), Expect = 0.004
Identities = 41/166 (24%), Positives = 66/166 (39%), Gaps = 10/166 (6%)
Query: 123 AEEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVG 182
AE+ N +N ++ ENK + E T +S KD E + + E + ++LT+
Sbjct: 81 AEDANIEALQNKVADLENK-VAELDKEVTRLQSDLKDAEENNVEDYVKEGLEKALTDKKV 139
Query: 183 EMEKNE--------SLTTDVASLSLGDDALASPPPQPKTRRKLVAPQPA-VRRSARIKTG 233
E+ + +L T + L D +P P PK + P+PA + A
Sbjct: 140 ELNNTQKALDTAQKALDTALNELGPDGDEEETPAPAPKPEQPAEQPKPAPAPQPAPAPKP 199
Query: 234 EMEKNESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQPA 279
E ++ D A S + PK + AP+P P PA
Sbjct: 200 EKTDDQQAEEDYARRSEEEYNRLPQQQPPKAEKPAPAPKPEQPVPA 245
>emb|CAA29073.1| NF-M c-terminus [Gallus gallus]
Length = 509
Score = 43.9 bits (102), Expect = 0.006
Identities = 41/184 (22%), Positives = 70/184 (37%), Gaps = 17/184 (9%)
Query: 124 EEENKSLKENVASETENKSLMEHVAAETENESLTKDVPEETENESLTEDVNESLTEDVGE 183
+EE K+ +E V E ++ E A E E E ++ EE +S + S E++ E
Sbjct: 137 QEEEKAEEEAVEEEAVSEKAAEQAAEEEEKEE--EEAEEEEAAKSDAAEEGGSKKEEIEE 194
Query: 184 MEKNESLTTDVASLS-----LGDDALASPPPQPKTRRKLVAPQPAVRRSARIKTGEMEKN 238
E+ E + A G P K+ K P ++ ++ E
Sbjct: 195 KEEREEAEEEEAEAKGKAEEAGAKVEKVKSPPAKSPPKSPPKSPVTEQAKAVQKAAAEVG 254
Query: 239 ESLTTDVASLSLGDDALASPPPQPKTRRKLAAPQPAAPQ----------PAVRRSARIKT 288
+ + A+ + A+ P +P T + + +PA P+ P RS T
Sbjct: 255 KDQKAEKAAEKAAKEEKAASPEKPATPKVTSPEKPATPEKPPTPEKAITPEKVRSPEKPT 314
Query: 289 TPKK 292
TP+K
Sbjct: 315 TPEK 318
>ref|ZP_00292511.1| hypothetical protein Tfus02001785 [Thermobifida fusca]
Length = 406
Score = 43.9 bits (102), Expect = 0.006
Identities = 32/109 (29%), Positives = 51/109 (46%), Gaps = 11/109 (10%)
Query: 188 ESLTTDVASLSLGDDALASPPPQPKTRRK----LVAPQPAVRRSARIKTGEMEKNESLTT 243
ES+T GD LA P + +T + APQ A R SA + + E+
Sbjct: 298 ESITDAEDDWETGDTPLADDPSEDETHSEPAEVAAAPQAAPRSSAAGAPHPVSREEAAAD 357
Query: 244 DVASLSLGDDALASPPPQPKTRRK--LAAPQPAAPQPAVRRSARIKTTP 290
D AS+ SPPP P+ RR+ + P+ ++P P S+ ++++P
Sbjct: 358 DAASVPE-----QSPPPAPRLRRRPIVLRPRRSSPPPLRPPSSEVRSSP 401
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.309 0.127 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 506,328,938
Number of Sequences: 2540612
Number of extensions: 22385625
Number of successful extensions: 138686
Number of sequences better than 10.0: 1369
Number of HSP's better than 10.0 without gapping: 96
Number of HSP's successfully gapped in prelim test: 1349
Number of HSP's that attempted gapping in prelim test: 134785
Number of HSP's gapped (non-prelim): 3943
length of query: 295
length of database: 863,360,394
effective HSP length: 127
effective length of query: 168
effective length of database: 540,702,670
effective search space: 90838048560
effective search space used: 90838048560
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0109.1


