

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0108.13
(1262 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
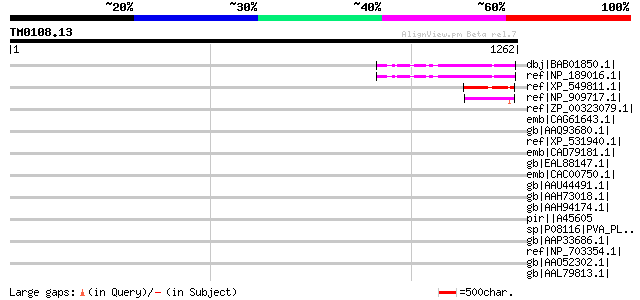
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAB01850.1| unnamed protein product [Arabidopsis thaliana] 152 9e-35
ref|NP_189016.1| expressed protein [Arabidopsis thaliana] 152 9e-35
ref|XP_549811.1| unknown protein [Oryza sativa (japonica cultiva... 111 2e-22
ref|NP_909717.1| hypothetical protein [Oryza sativa (japonica cu... 67 4e-09
ref|ZP_00323079.1| hypothetical protein PpenA01001008 [Pediococc... 48 0.002
emb|CAG61643.1| unnamed protein product [Candida glabrata CBS138... 46 0.009
gb|AAQ93680.1| starmaker [Danio rerio] gi|38488737|ref|NP_942112... 45 0.012
ref|XP_531940.1| PREDICTED: similar to myosin tail domain-contai... 44 0.045
emb|CAD79181.1| hypothetical protein-signal peptide prediction [... 43 0.059
gb|EAL88147.1| hypothetical protein Afu1g04370 [Aspergillus fumi... 43 0.077
emb|CAC00750.1| hypothetical protein [Arabidopsis thaliana] gi|1... 42 0.17
gb|AAU44491.1| hypothetical protein AT3G56870 [Arabidopsis thali... 42 0.17
gb|AAH73018.1| LOC443607 protein [Xenopus laevis] 42 0.17
gb|AAH94174.1| LOC443607 protein [Xenopus laevis] 42 0.17
pir||A45605 mature-parasite-infected erythrocyte surface antigen... 41 0.23
sp|P08116|PVA_PLAFA Processed variable antigen gi|552170|gb|AAA2... 41 0.23
gb|AAP33686.1| phoshoprotein 300 [Plasmodium falciparum] 41 0.23
ref|NP_703354.1| Mature parasite-infected erythrocyte surface an... 41 0.23
gb|AAO52302.1| similar to Homo sapiens (Human). Dentin sialophos... 41 0.23
gb|AAL79813.1| sialoprotein-phosphophoryn DSP-PP523 [Rattus norv... 41 0.23
>dbj|BAB01850.1| unnamed protein product [Arabidopsis thaliana]
Length = 566
Score = 152 bits (383), Expect = 9e-35
Identities = 117/350 (33%), Positives = 174/350 (49%), Gaps = 42/350 (12%)
Query: 914 RIKSLKHAGSPTYKRLLPLLSNAKKYNSCASVNRHDPKHQKFLDQTPLLPISDSDLRATS 973
R K K GS Y+R+LP L + ++ N P QK ++ IS L + S
Sbjct: 240 RGKLFKTPGSVNYRRMLPYLKDIQEDN---------PYPQKNTEEV----ISSPMLNSES 286
Query: 974 GGDSNDSVPMEHFTANSGTKQQKELNACELNDDSSSPRSRSPGFQSSNSKCKVMQMCDEQ 1033
+ V + T SGT + ++ P R P + K + +
Sbjct: 287 DNEGTQEVVTSNVTRESGTSSDE--------NEEPLPCERVPVNLEQSDPDKEQETQIKH 338
Query: 1034 VLLNGLCKPESSSGTPISVRRIDSPSTPLRPIVNEVMAREEATFALSESLINSKVKGNCS 1093
V+ + E++ G+ I + P+V + E + AL + +++ V G +
Sbjct: 339 VIPD----TENNLGSEIPLSS---------PLVGSRSSSEVNSSALHNTFVDNLV-GEEN 384
Query: 1094 FSISPDDKEKLS----EAHGCCLSLSQLELKEKLEVPA-VDFKKGILKRNPRGCRGLCTC 1148
+ + + K+S EAH + ++ L P+ KGILKR+ RGCRG+C+C
Sbjct: 385 MNGAEITEAKISAEELEAHSSDATAELVDPSVILATPSSFSPSKGILKRSMRGCRGICSC 444
Query: 1149 LNCASFRLHAERAFEYSRNQLLDAEEVAQDLLKELSHLRDMLGRPIDSVNDNPGFDRDQV 1208
LNC+SFRLHAERAFE+SRNQL D E + DL+ E+SHLRD+L + + + P + Q
Sbjct: 445 LNCSSFRLHAERAFEFSRNQLQDTEVMVLDLVGEISHLRDLLEKYNSADHSEP--YKSQA 502
Query: 1209 KEACRKAYAAEQLAKERLSMMKDDLSIHCRITNLQRPRVRFADHVQEKII 1258
EA ++A A +LAK RL M DDL IH RI N QR RV+FA ++ EK I
Sbjct: 503 GEASKRACEAAELAKSRLHQMNDDLQIHYRIPNEQRARVKFAHYIHEKTI 552
Score = 37.4 bits (85), Expect = 3.3
Identities = 44/155 (28%), Positives = 63/155 (40%), Gaps = 54/155 (34%)
Query: 492 NDDFILTTPPDAKI------YDNSAVNVDRS----KSV--------------PQDMKRLK 527
+DDF TTPPD+++ + S VN + KSV P KRL
Sbjct: 113 SDDFAQTTPPDSELLAISEEINGSVVNKSDTNLWRKSVLLPCSRPKIFKNTGPFSYKRLL 172
Query: 528 PMDLRTNTQESFSKGVCLKADS------------------------RKDSLLKSKSVRRP 563
P ++ + + S C K+ S R S LKS P
Sbjct: 173 PYLMQASDDGTSSSSRCSKSLSQNITKPVSQSMDSVYDKDSTGSFCRDTSPLKSVIASTP 232
Query: 564 H-----LPGKLFKTLGSVS-KRFLPFLMELAKDDP 592
+ GKLFKT GSV+ +R LP+L ++ +D+P
Sbjct: 233 NKNAAFSRGKLFKTPGSVNYRRMLPYLKDIQEDNP 267
>ref|NP_189016.1| expressed protein [Arabidopsis thaliana]
Length = 542
Score = 152 bits (383), Expect = 9e-35
Identities = 117/350 (33%), Positives = 174/350 (49%), Gaps = 42/350 (12%)
Query: 914 RIKSLKHAGSPTYKRLLPLLSNAKKYNSCASVNRHDPKHQKFLDQTPLLPISDSDLRATS 973
R K K GS Y+R+LP L + ++ N P QK ++ IS L + S
Sbjct: 216 RGKLFKTPGSVNYRRMLPYLKDIQEDN---------PYPQKNTEEV----ISSPMLNSES 262
Query: 974 GGDSNDSVPMEHFTANSGTKQQKELNACELNDDSSSPRSRSPGFQSSNSKCKVMQMCDEQ 1033
+ V + T SGT + ++ P R P + K + +
Sbjct: 263 DNEGTQEVVTSNVTRESGTSSDE--------NEEPLPCERVPVNLEQSDPDKEQETQIKH 314
Query: 1034 VLLNGLCKPESSSGTPISVRRIDSPSTPLRPIVNEVMAREEATFALSESLINSKVKGNCS 1093
V+ + E++ G+ I + P+V + E + AL + +++ V G +
Sbjct: 315 VIPD----TENNLGSEIPLSS---------PLVGSRSSSEVNSSALHNTFVDNLV-GEEN 360
Query: 1094 FSISPDDKEKLS----EAHGCCLSLSQLELKEKLEVPA-VDFKKGILKRNPRGCRGLCTC 1148
+ + + K+S EAH + ++ L P+ KGILKR+ RGCRG+C+C
Sbjct: 361 MNGAEITEAKISAEELEAHSSDATAELVDPSVILATPSSFSPSKGILKRSMRGCRGICSC 420
Query: 1149 LNCASFRLHAERAFEYSRNQLLDAEEVAQDLLKELSHLRDMLGRPIDSVNDNPGFDRDQV 1208
LNC+SFRLHAERAFE+SRNQL D E + DL+ E+SHLRD+L + + + P + Q
Sbjct: 421 LNCSSFRLHAERAFEFSRNQLQDTEVMVLDLVGEISHLRDLLEKYNSADHSEP--YKSQA 478
Query: 1209 KEACRKAYAAEQLAKERLSMMKDDLSIHCRITNLQRPRVRFADHVQEKII 1258
EA ++A A +LAK RL M DDL IH RI N QR RV+FA ++ EK I
Sbjct: 479 GEASKRACEAAELAKSRLHQMNDDLQIHYRIPNEQRARVKFAHYIHEKTI 528
Score = 37.0 bits (84), Expect = 4.2
Identities = 44/154 (28%), Positives = 62/154 (39%), Gaps = 54/154 (35%)
Query: 493 DDFILTTPPDAKI------YDNSAVNVDRS----KSV--------------PQDMKRLKP 528
DDF TTPPD+++ + S VN + KSV P KRL P
Sbjct: 90 DDFAQTTPPDSELLAISEEINGSVVNKSDTNLWRKSVLLPCSRPKIFKNTGPFSYKRLLP 149
Query: 529 MDLRTNTQESFSKGVCLKADS------------------------RKDSLLKSKSVRRPH 564
++ + + S C K+ S R S LKS P+
Sbjct: 150 YLMQASDDGTSSSSRCSKSLSQNITKPVSQSMDSVYDKDSTGSFCRDTSPLKSVIASTPN 209
Query: 565 -----LPGKLFKTLGSVS-KRFLPFLMELAKDDP 592
GKLFKT GSV+ +R LP+L ++ +D+P
Sbjct: 210 KNAAFSRGKLFKTPGSVNYRRMLPYLKDIQEDNP 243
>ref|XP_549811.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|52076600|dbj|BAD45502.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 676
Score = 111 bits (277), Expect = 2e-22
Identities = 54/127 (42%), Positives = 87/127 (67%), Gaps = 11/127 (8%)
Query: 1131 KKGILKRNPRGCRGLCTCLNCASFRLHAERAFEYSRNQLLDAEEVAQDLLKELSHLRDML 1190
KKGILKR+ RGC+G+C CL+C++FRL A+RAFE+SR Q+ +A+++ +LLKE+S LR+++
Sbjct: 548 KKGILKRHTRGCKGICMCLDCSTFRLRADRAFEFSRKQMQEADDIIDNLLKEVSSLRNLM 607
Query: 1191 GRPIDSVNDNPGFDRDQVKEACRKAYAAEQLAKERLSMMKDDLSIHCRITNLQRPRVRFA 1250
+ ++ + AC++A E +A+ER M +L+ HCRI PRV+FA
Sbjct: 608 --------EKSAGQQETKQTACQRASQVEVVARERRRQMLMELNSHCRIPG---PRVKFA 656
Query: 1251 DHVQEKI 1257
+V+E++
Sbjct: 657 QYVEERM 663
>ref|NP_909717.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|28269482|gb|AAO38025.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 863
Score = 67.0 bits (162), Expect = 4e-09
Identities = 42/143 (29%), Positives = 74/143 (51%), Gaps = 17/143 (11%)
Query: 1132 KGILK--RNPRGCRGLCTCLNCASFRLHAERAFEYSRNQLLDAEEVAQDLLKELSHLRDM 1189
KGILK +P + CTC+ AS L AE+A E+S+ Q+ D E +A L++ L+H+R +
Sbjct: 701 KGILKGTESPSPQKTTCTCMKAASVILDAEKAVEFSQRQMHDIENIASKLMRSLNHMRSI 760
Query: 1190 LGRPIDSVNDN--PGFDRDQVKEACRKAYAAEQLAKERLSMMKDDLSIHCRITNL----- 1242
+ + S + + P F+ +++ A A E+ ++ L++M D + C+I L
Sbjct: 761 VDGNLLSESHSLLPTFNTAEIRAASEDALEVERTTRKWLTIMNKDCNRFCKILRLAGKKA 820
Query: 1243 --------QRPRVRFADHVQEKI 1257
+R ++ FAD K+
Sbjct: 821 VSHSEVPRKRKKITFADETGGKL 843
>ref|ZP_00323079.1| hypothetical protein PpenA01001008 [Pediococcus pentosaceus ATCC
25745]
Length = 2334
Score = 47.8 bits (112), Expect = 0.002
Identities = 139/764 (18%), Positives = 271/764 (35%), Gaps = 88/764 (11%)
Query: 378 SQSEKPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNEREVSGECLSVPSGDDKCFN 437
S S S E +S S ++ ++ S+S + +S N + + + S +
Sbjct: 915 SDSNSKSTSESRSTSTSVSDSTSDSTSTSHSTSDSVSTSNS-DSNSKSTSESRSTSTSIS 973
Query: 438 ESKCDPAADSDGVKVVGNALHNDDFMQCSIHKNNLDYAKVTN---------DSGCTSVQL 488
+SK D A+ SD V + N S K++ +++ DS S
Sbjct: 974 DSKSDSASKSDSVSKSDSITSNSISESISTSKSDSSSKSMSDSRSASTSVSDSTSDSAST 1033
Query: 489 GVLNDDFILTTPPDAKIYDNSAVNVDR---SKSVPQDMKRLKPMDLRTNTQESFSKGVCL 545
D + T+ D+ +S D S SV + + T+T S SK
Sbjct: 1034 SHSKSDSVSTSNSDSSSKSDSVSTSDSRSTSTSVSDSISKSMSDSRSTSTSVSDSKS--- 1090
Query: 546 KADSRKDSLLKSKSVRRPHLPGKLFKTLGSVSKRFLPFLMELAKDDPGISKFGHGTPKKD 605
++S+ DS+ KS S+ T S+S+ + D
Sbjct: 1091 DSESKSDSISKSDSI-----------TSNSISESISTSNSDSISDS------------NS 1127
Query: 606 EADMHGKRFELTLSSQPEEASIDGHKIEKGPIHGTVESNALG-NILFNSANELSNGNLPK 664
++ + ++S +++ + K ++ S+++ +I ++++ +S+ N
Sbjct: 1128 KSTSDSRSTSTSISDSKSDSASKSDSVSKSD---SITSDSISESISTSNSDSISDSNSKS 1184
Query: 665 LTSSQDLPELPMQS-----DVKEVVQECLSAPSVEEPIEKAVVGSKDECSREPNFDLCSA 719
+ S+ S + +S + + + V + D SR + + +
Sbjct: 1185 TSDSRSTSTSVSDSKSDSASTSHSTSDSVSTSNSDSSSKSDSVSTSD--SRSTSTSISDS 1242
Query: 720 IVDFHSTK-----IVANDGHDVDIKHMQNSISSLQERGPSPKDQSMLYVNVGVSNQTLEH 774
D ST V+ D D K +S S+ S D + + S+ +
Sbjct: 1243 TSDSASTSHSTSDSVSTSNSDSDSKSTSDSRSASTSVSDSKSDSASKSDSTSKSDSITSN 1302
Query: 775 HSSEVKGFSVNNGEVKQLLNMEKRKYVSRYPSEGQSSSQLDTDMQDLEENERSNGAIRKS 834
SE S ++ K + K SR S S+S D++ + ++ ++ ++ S
Sbjct: 1303 SISESISTSNSDSSSK---SDSKSTSDSRSTSTSVSNSISDSNSKSTSDSRSTSTSVSDS 1359
Query: 835 -DDAVVLNHVGVSNEVPRPPPENIISKTQDMAAHGGNKEGSVQKGIAYGSKGNNGRKGCY 893
D+V +H +++ + D + + S ++ +N +
Sbjct: 1360 TSDSVSTSH---------STSDSVSTSNSDSDSKSASDSRSTSTSVSNSISDSNSKS--- 1407
Query: 894 AYEIENASESKITPVLNRCPRIKSLKHAGSPTYKRLLPLLSNAKKYNSCASVNRHDPKH- 952
+ S S T V + S H+ S + SN+ + AS +R
Sbjct: 1408 ----TSDSRSTSTSVSDSTSDSVSTSHSTSDSVST-----SNSDSDSKSASDSRSTSTSV 1458
Query: 953 QKFLDQTPLLPISDSDLRATSGGDS-NDSVPMEHFTANSGTKQQKELNACELNDDSSSPR 1011
+ + SDS +TS DS +DS H T++S + + ++ +D S+
Sbjct: 1459 SNSISDSNSKSTSDSRSTSTSVSDSTSDSASTSHSTSDSVSTSNSDSDSKSTSDSRSAST 1518
Query: 1012 SRSPGFQSSNSKCKVMQMCDEQVLLNGLCKPESSSGTPISVRRIDSPSTPLRPIVNEVMA 1071
S S S SK D + N + + S+S + S + + R V
Sbjct: 1519 SVSDSKSDSESKSDSTSKSD-SITSNSISESISTSNSDSSSKSDSKSISDSRSTSTSVSD 1577
Query: 1072 REEATFALSESLINSKVKGNCSFSISPDDKEKLSEAHGCCLSLS 1115
+ + S S +S S S S D + +SE+ S+S
Sbjct: 1578 STSDSASTSHSTSDS-----VSTSNSDSDSKSMSESRSTSTSVS 1616
Score = 45.4 bits (106), Expect = 0.012
Identities = 127/685 (18%), Positives = 247/685 (35%), Gaps = 55/685 (8%)
Query: 377 SSQSEKPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNEREVS-GECLSVPSGDDKC 435
+S S S+ S S + ++ S+S ++ +S + N + S S S D K
Sbjct: 801 TSTSHSTSDSVSTSNSNSDSKSKSESRSTSTSISDSISDSNSKSTSESRSTSTSSSDSKS 860
Query: 436 FNESKCDPAADSDGVKVVGNALHNDDFMQCSIHKNNLDYAKVTNDSGCTSVQLGVLNDDF 495
+ SK D + SD + N++ S + D +K T++S TS + D
Sbjct: 861 DSASKSDSISKSDSI--TSNSISESISTSNSDSSSKSD-SKSTSESRSTSTSVSDSISDS 917
Query: 496 ILTTPPDAK-----IYDNSAVNVDRSKSVPQDMKRLKPMDLRTNTQESFSKGVCLKADSR 550
+ +++ + D+++ + S S + +T ES S + +DS+
Sbjct: 918 NSKSTSESRSTSTSVSDSTSDSTSTSHSTSDSVSTSNSDSNSKSTSESRSTSTSI-SDSK 976
Query: 551 KDSLLKSKSVRRPHLPGKLFKTLGSVSKRFLPFLMELAKDDPGISKFGHGTPKKDEADM- 609
DS KS SV + T S+S+ + + S+ + +D
Sbjct: 977 SDSASKSDSVSKSDSI-----TSNSISESISTSKSDSSSKSMSDSRSASTSVSDSTSDSA 1031
Query: 610 ---HGKRFELTLSSQPEEASIDGHKIEKGPIHGTVESNALGNILFNSANELSNGNLPKLT 666
H K ++ S+ + D T S+++ + +S + ++ + K
Sbjct: 1032 STSHSKSDSVSTSNSDSSSKSDSVSTSDSRSTSTSVSDSISKSMSDSRSTSTSVSDSKSD 1091
Query: 667 SSQDLPELPMQSDV-KEVVQECLS---APSVEEPIEKAVVGSKDECSR--EPNFDLCSAI 720
S + + + E +S + S+ + K+ S+ + + D S
Sbjct: 1092 SESKSDSISKSDSITSNSISESISTSNSDSISDSNSKSTSDSRSTSTSISDSKSDSASKS 1151
Query: 721 VDFHSTKIVANDGHDVDIKHMQN-SISSLQERGPSPKDQSMLYVNVGVSNQTLEHHS--- 776
+ + +D I + SIS + S + V+ S+ HS
Sbjct: 1152 DSVSKSDSITSDSISESISTSNSDSISDSNSKSTSDSRSTSTSVSDSKSDSASTSHSTSD 1211
Query: 777 --------SEVKGFSVNNGEVKQLL-----NMEKRKYVSRYPSEGQSSSQLDTDMQDLEE 823
S K SV+ + + + S S+ S+S D+D + +
Sbjct: 1212 SVSTSNSDSSSKSDSVSTSDSRSTSTSISDSTSDSASTSHSTSDSVSTSNSDSDSKSTSD 1271
Query: 824 NERSNGAIR--KSDDAVVLNHVGVSNEVPRPP-PENIISKTQDMAAHGGNKEGSVQKGIA 880
+ ++ ++ KSD A + S+ + E+I + D ++ +K S + +
Sbjct: 1272 SRSASTSVSDSKSDSASKSDSTSKSDSITSNSISESISTSNSDSSSKSDSKSTSDSRSTS 1331
Query: 881 YGSKGNNGRKGCYAYEIENASESKITPVLNRCPRIKSLKHAGSPTYKRLLPLLSNAKKYN 940
+ + + S S T V + S H+ S + SN+ +
Sbjct: 1332 TSVSNSISDSNSKS---TSDSRSTSTSVSDSTSDSVSTSHSTSDSVST-----SNSDSDS 1383
Query: 941 SCASVNRHDPKH-QKFLDQTPLLPISDSDLRATSGGDS-NDSVPMEHFTANSGTKQQKEL 998
AS +R + + SDS +TS DS +DSV H T++S + +
Sbjct: 1384 KSASDSRSTSTSVSNSISDSNSKSTSDSRSTSTSVSDSTSDSVSTSHSTSDSVSTSNSDS 1443
Query: 999 NACELNDDSSSPRSRSPGFQSSNSK 1023
++ +D S+ S S SNSK
Sbjct: 1444 DSKSASDSRSTSTSVSNSISDSNSK 1468
Score = 35.8 bits (81), Expect = 9.5
Identities = 82/395 (20%), Positives = 144/395 (35%), Gaps = 52/395 (13%)
Query: 741 MQNSISSLQERGPSPKDQSMLYVNVGVSNQTLEHHSSEVKGFSVNNGEVKQLLNMEKRKY 800
+ NSIS+ S D + ++ S T S+ V + V Q ++ K
Sbjct: 589 ISNSISTSVSGSKSTSDSASQ--SISTSKSTSTSGSTSVSNSKSMSDSVSQSISTSKSTS 646
Query: 801 VSRYPSEGQSSSQLDTDMQDLEENERSNGAIRKSDDAVVLNHVGVSNEVPRPPPENIISK 860
S S S S D+ Q + ++ ++ S V N S+ V ++I +
Sbjct: 647 TSGSTSVSNSKSTSDSLSQSISASKSTS----TSGSTSVSNSKSTSDSVS----QSISTS 698
Query: 861 TQDMAAHGGNKEGSVQKGIAYGSKGNNGRKGCYAYEIENA---SESKITPVLNRCPRIKS 917
D + G+ SV K SK ++ + I + S+S V KS
Sbjct: 699 KSDSTSASGSVSDSVSK-----SKSDSITSDSISNSISTSVSGSKSTSDSVSQSISTSKS 753
Query: 918 LKHAGSPTYKRLLPLLSNAKKYNSCASVNRHDPKHQKFLDQTPLLPISDSDLRATSGGDS 977
+GS + +SN+K + S + K + ISDS + S
Sbjct: 754 TSTSGSTS-------VSNSKSTSDSVSQSISTSKSDSITSDS----ISDSISTSIS---- 798
Query: 978 NDSVPMEHFT----------ANSGTKQQKELNACELNDDSSSPRSRSPGFQSSNSKCKVM 1027
DS H T ++S +K + + ++D S S+S S S
Sbjct: 799 -DSTSTSHSTSDSVSTSNSNSDSKSKSESRSTSTSISDSISDSNSKSTSESRSTSTSSSD 857
Query: 1028 QMCDEQVLLNGLCKPESSSGTPI--SVRRIDSPSTPLRPIVNEVMAREEATFALSESLIN 1085
D + + K +S + I S+ +S S+ + +R +T ++S+S+ +
Sbjct: 858 SKSDSASKSDSISKSDSITSNSISESISTSNSDSSSKSDSKSTSESRSTST-SVSDSISD 916
Query: 1086 SKVKG-----NCSFSISPDDKEKLSEAHGCCLSLS 1115
S K + S S+S + S +H S+S
Sbjct: 917 SNSKSTSESRSTSTSVSDSTSDSTSTSHSTSDSVS 951
>emb|CAG61643.1| unnamed protein product [Candida glabrata CBS138]
gi|50292495|ref|XP_448680.1| unnamed protein product
[Candida glabrata]
Length = 1012
Score = 45.8 bits (107), Expect = 0.009
Identities = 113/539 (20%), Positives = 195/539 (35%), Gaps = 76/539 (14%)
Query: 389 KSESFTMRETNGSKLSSSQNLLESPMELNEREVSGECLSVPSG----------------- 431
K+ES + + S S +L ++ N+ V GE S+ G
Sbjct: 256 KNESISNELATEKEQSPSTAVLSDDLKKNQDNVQGEKQSIEKGSKADADILIKPETNSKL 315
Query: 432 --DDKCFNESKCDPAADSD-GVKVVGNALHNDDFMQCSIHKNNLDYAKVT----NDSGCT 484
D NES D + SD +K +G + Q + ++ + +T N+
Sbjct: 316 VKDAVIANESDADKKSISDTTIKEIGESNAEIKEKQTTQSRSEKEIISLTAEENNNKVTI 375
Query: 485 SVQLGVLNDDFILTTPPDAKIYDNSAVNVDRSKSVPQDMKRLKPMDLRT---NTQESFSK 541
S + LNDD + P N+D KSV Q+ K + ++ +E +K
Sbjct: 376 SDTIPDLNDDTMKAASP----------NIDEIKSVQQESYDSKSKETKSEPEEQEEDMNK 425
Query: 542 GVCLKAD--SRKDSLLKSKSVRRPHLPGK------------------LFKTLGSVSKRFL 581
G + S S K ++++ P L + L + L +VS
Sbjct: 426 GQLEPENKVSSSPSSSKKQALKEPALESETEQEHEGNLLPTGNKTDVLEENLETVS-AVK 484
Query: 582 PFLMELAKDDPGISKFGHG-TPKKDEADMHGKRF-ELTLSSQPEEASIDGHKIE-----K 634
P L ++ KD P HG T E K EL+ + P++ S+D K E
Sbjct: 485 PSLADVNKDLPLKEGVNHGKTVDVQELHTTSKPIEELSQAESPKKTSLDAMKKEDKLKDS 544
Query: 635 GPIHGTVESNALGNILFNSANELSNGNLPKLTSSQDLPELPMQSDVKEVVQECLSAPSVE 694
T+E++ I S E + P S+DL + P +S E V + +V
Sbjct: 545 RKTESTIEAHLDNRINEKSEAEEVGDDGPVDKKSEDLKDTPEKSLSTEAVTQSPKVHNVN 604
Query: 695 EPIEKAVVGSKDECSREPNFDLCSAIVDFHSTKIVANDGHDVDIKHMQNSISSLQERGPS 754
E V +D+ + D A + + + D + N +
Sbjct: 605 E--VSKVNEEEDKAAESLTVDKAGAENSHKPSNLKVLEAKDT-LSDTMNETKQDETMDKE 661
Query: 755 PKDQSMLYVNVGVSNQTLEHHSSEVKGFSVNNGEVKQLLNMEKRKYVSRYPSEGQSSSQL 814
PK + L + ++++ E S+ S E ++ + ++ Q+ S L
Sbjct: 662 PKSTAKL---IKLNDEVDEEASTLADDVSDLTEEKLKVKDNVMHTTAEIPNNDAQTDSDL 718
Query: 815 DTDMQDLEE-NERSNG--AIRKSDDAVVLNHVGVSNEVPRPPPENIISKTQDMAAHGGN 870
+ +DL E NE N ++K D V+ + +NE P +IS D+ + N
Sbjct: 719 PKEKEDLNEFNEDKNNEDPVKKGDGDVLSKIMTENNETPE--KRVVISILDDLPSRHDN 775
>gb|AAQ93680.1| starmaker [Danio rerio] gi|38488737|ref|NP_942112.1| starmaker
[Danio rerio]
Length = 613
Score = 45.4 bits (106), Expect = 0.012
Identities = 90/421 (21%), Positives = 145/421 (34%), Gaps = 53/421 (12%)
Query: 69 SSGNDEKREPPEED---DGDTLLVKYLRKKAKRDDDGGDLPRLLTKDLRARRVYSPQSSA 125
S +K E P+ D DGD+ + D+D KD + + S
Sbjct: 184 SPDTTDKPEGPDSDSAPDGDSASAEKTDSDHSPDEDANKSSTEADKDDTSDKDSSQTDEK 243
Query: 126 RASLDLIDSDCADRKIAGLGFGSPSSEEKSGVHVDDVMIAVNSDCTDRKIADLGFGSSAE 185
DSD +D+ S E+ S +D + + +D+ +D G S A+
Sbjct: 244 H------DSDASDKDEKHEDKDEKSDEKDSSKDSEDK----SQEKSDK--SDDGSNSEAD 291
Query: 186 KGGLCVD--DATMKVTVSPDLGISSQARDFDRANTDSSDHKQFEEGNDGDCSGKINDFDE 243
+ V+ D S + D D+ ++DS+D K +E D D S + + D
Sbjct: 292 EQKESVESKDHDSDSQDSDSAEKKEKHDDKDQDSSDSADSKDSDEDKDKDHSEQKDSEDH 351
Query: 244 EIVVTTPPDAELLGDSKVDGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLK 303
E D E D DEGK +ED D + E S++ SK ++S +
Sbjct: 352 EHKEKHTKDKE----EHKDSDEGKDDEDKSKSD------EHDKDESESKEASKSDESEQE 401
Query: 304 SKYFSTAANHSFDLDLYAGARSQTAFPGQIVQGPGFEACGGTNWPDNFVGTKKLGHCDKD 363
K + + + D ++ + S + D+ + H D D
Sbjct: 402 EKKDDKSDSDNSSRDSHSDSDSD-------------------SHSDSDSDSHSDSHSDSD 442
Query: 364 EKGMGGQDFQLPLSSQSEKPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNEREVSG 423
D S S+ S+ + S +E K S S++ S + S
Sbjct: 443 SDSHSDSDSDSDSDSDSDSDSDSDSNSRD---KEEKKDKSSESRDEDSSDSDSKSNSESS 499
Query: 424 ECLSVPSGDDKCFNESKCDPAADSDGVKVVGNAL---HNDDFMQCSIHKNNLDYAKVTND 480
E + DDK + K D SD V N H+DD + + K D
Sbjct: 500 ETAEEDTNDDKDSSVEK-DKTDSSDSASVEANDSDDEHDDDSKDATPSSEDHTAEKTDED 558
Query: 481 S 481
S
Sbjct: 559 S 559
>ref|XP_531940.1| PREDICTED: similar to myosin tail domain-containing protein [Canis
familiaris]
Length = 1561
Score = 43.5 bits (101), Expect = 0.045
Identities = 42/169 (24%), Positives = 73/169 (42%), Gaps = 9/169 (5%)
Query: 722 DFHSTKIVANDGHDVDIKHMQNSISSLQERGPSPKDQSMLYVNVGVSNQTLEHHSSEVKG 781
+F +K++ ++ + + + N++ Q+ +Q+ + N LE HSSE
Sbjct: 1203 EFKQSKLITHEKFESACEELNNALLREQQAQMLLNEQAQQLQEL---NYRLELHSSEEAD 1259
Query: 782 FSVNNGEVKQLLN---MEKRKYVSRYPSEGQSSSQLDTDMQDLEEN-ERSNGAIRKS--D 835
+ GE + L+ ME R+ + +QL+ D + LEEN + A+R + D
Sbjct: 1260 KNQTLGEAVKSLSEAKMELRRKDQSLRQLNRHLTQLEQDKRRLEENIHDAESALRMAAKD 1319
Query: 836 DAVVLNHVGVSNEVPRPPPENIISKTQDMAAHGGNKEGSVQKGIAYGSK 884
V NH+ + E SK + + G +EG QK GSK
Sbjct: 1320 KECVANHMRTVENMLHKSKERRDSKKESVEVEGSAEEGMGQKKAGSGSK 1368
>emb|CAD79181.1| hypothetical protein-signal peptide prediction [Rhodopirellula
baltica SH 1] gi|32477034|ref|NP_870028.1| hypothetical
protein-signal peptide prediction [Rhodopirellula
baltica SH 1]
Length = 669
Score = 43.1 bits (100), Expect = 0.059
Identities = 87/471 (18%), Positives = 153/471 (32%), Gaps = 31/471 (6%)
Query: 5 PTLRESASAMEETLISRKRSNPSSSPPGALTRSRSQLFVHRNRSGQRRPD-SGPRQGGLY 63
P S+S+ ++ S S+PSSS + S S + SG S L
Sbjct: 11 PVAGSSSSSSSDSSSSSSSSSPSSSSSSGSSSSSSGSSSSSSMSGSSSSSMSSSSSSSLS 70
Query: 64 SGLGGSSGNDEKREPPEEDDGDTLLVKYLRKKAKRDDDGGDLPRLLTKDLRARRVYSPQS 123
S SSG+ + + G + ++ S S
Sbjct: 71 SSSSSSSGSSSSSSGSSSSSSGSPSSSSSSGSSSSSSSG-------SSSSSSKSSSSSDS 123
Query: 124 SARASLDLIDSDCADRKIAGLGFGSPSSEEKSGVHVDDVMIAVNSDCTDRKIADLGFGSS 183
S+ +S SD +D + S S + S V + +SD +D +
Sbjct: 124 SSNSS----SSDSSDSSHSSSASSSSDSSDSSTSSSSSVSSSQSSDSSDSSDSSTSTSLP 179
Query: 184 AEKGGLCVDDATMKVTVSPDLGISSQARDFDRANTDSSDHKQFEEGNDGDCSGKINDFDE 243
+ D ++ + S D SS ++ D +N+DSSD + + SG + D
Sbjct: 180 SVSSSSSSDGSSSSSSGSSD--SSSNSQSSDSSNSDSSDSS--DSTSLPSISGS-SSSDG 234
Query: 244 EIVVTTPPDAELLGDSKVDGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLK 303
+ D+ S + + + +G + ++G + DS + S +
Sbjct: 235 SSASSDSSDSSSNSQSSDSSNSESSDSSDSISLPSISGSSSSAGSSSASSDSSNSSSNSQ 294
Query: 304 SKYFSTAANH----SFDLDLYAG----------ARSQTAFPGQIVQGPGFEACGGTNWPD 349
S ST+ ++ S L +G + SQ++ G ++ + P
Sbjct: 295 SSDSSTSESNDSSASVSLPSVSGSSSSASDSSSSESQSSDSGSSSDSSTSQSSESASTPS 354
Query: 350 NFVGTKKLGHCDKDEKGMGGQDFQLPLSSQSEKPSEPELKSESFTMRETNGSKLSSSQNL 409
+ SS SE S E SES N S S S +
Sbjct: 355 VSSSQSSTSESASSTSESSNSESSQSESSTSESRSRSESDSESLPSYSWNSSTSSDSDSE 414
Query: 410 LESPMELNEREVSGECLSVPSGDDKCFNESKCDPAADSDGVKVVGNALHND 460
++ + SV +ES D ++DS +A ++D
Sbjct: 415 SSDSESESQSQSDSTSQSVSDSGSDSQSESSIDRSSDSQSYSASASASYSD 465
>gb|EAL88147.1| hypothetical protein Afu1g04370 [Aspergillus fumigatus Af293]
Length = 437
Score = 42.7 bits (99), Expect = 0.077
Identities = 52/232 (22%), Positives = 90/232 (38%), Gaps = 22/232 (9%)
Query: 74 EKREPPEEDDGDTLLVKYLRKKAKRDDDGGDLPRLLTKDLRARRVYSPQSSARASLDLID 133
+ R+ D ++ + + D + D P++ K R S SS+ +S D
Sbjct: 77 QDRKSESSSDSESSSSSSEDESSDSDVEMNDAPKMQKKQSRESSPSSSSSSSSSSSSSSD 136
Query: 134 SDCADRKIAGLGFGSPSSEEKSGVHVDDVMIAVNSDCTDRKIADLGFGSSAEKGGLCVDD 193
SD D + + + +G T RK SS+E D+
Sbjct: 137 SDADDEEDDDAAPAPAPAPKTTG--------------TKRKAESSSSESSSESES--EDE 180
Query: 194 ATM----KVTVSPDLGISSQARDFDRANTDSSDHKQFEEGNDGDC-SGKINDF-DEEIVV 247
A K++ D SS++ D + +DSSD + +D D SG +D D E V
Sbjct: 181 APKAKKAKLSTKSDSSSSSESSDSEDEESDSSDSESSASSSDSDSDSGSDSDSSDSESVS 240
Query: 248 TTPPDAELLGDSKVDGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRND 299
++ ++ DS+ + + K +++ A T E+S S DS +D
Sbjct: 241 SSDSSSDSDSDSESEKEPAKKAAKKILKTAAKTPLPESSSSSSSDSDSSSSD 292
>emb|CAC00750.1| hypothetical protein [Arabidopsis thaliana]
gi|15230148|ref|NP_191246.1| hypothetical protein
[Arabidopsis thaliana] gi|11358432|pir||T51275
hypothetical protein T8M16_200 - Arabidopsis thaliana
Length = 670
Score = 41.6 bits (96), Expect = 0.17
Identities = 30/128 (23%), Positives = 61/128 (47%), Gaps = 6/128 (4%)
Query: 1136 KRNPRGCRGLCTCLNCASFRLHAERAFEYSRNQLLDAEEVAQDLLKELSHLRDMLGRPID 1195
+RN + + S + +++A +S+ Q+ D + VA L KEL +R + R +
Sbjct: 510 RRNTQAAKAQPFSTGGTSIQGCSQKAIAFSQGQMRDFQNVAARLTKELKSMRQITKRCLQ 569
Query: 1196 SVNDNPGF---DRDQVKEACRKAYAAEQLAKERLSMMKDDLSIHCRITNLQR---PRVRF 1249
+ ++ + D+VK A E+ K+ LS+++ D + C++ ++ R P
Sbjct: 570 AESNTSNMSDCNLDEVKTVIGNAEKTEESCKKWLSIIERDCNRFCKLMSMVREDSPATEN 629
Query: 1250 ADHVQEKI 1257
H ++KI
Sbjct: 630 IVHKKKKI 637
>gb|AAU44491.1| hypothetical protein AT3G56870 [Arabidopsis thaliana]
Length = 700
Score = 41.6 bits (96), Expect = 0.17
Identities = 30/128 (23%), Positives = 61/128 (47%), Gaps = 6/128 (4%)
Query: 1136 KRNPRGCRGLCTCLNCASFRLHAERAFEYSRNQLLDAEEVAQDLLKELSHLRDMLGRPID 1195
+RN + + S + +++A +S+ Q+ D + VA L KEL +R + R +
Sbjct: 540 RRNTQAAKAQPFSTGGTSIQGCSQKAIAFSQGQMRDFQNVAARLTKELKSMRQITKRCLQ 599
Query: 1196 SVNDNPGF---DRDQVKEACRKAYAAEQLAKERLSMMKDDLSIHCRITNLQR---PRVRF 1249
+ ++ + D+VK A E+ K+ LS+++ D + C++ ++ R P
Sbjct: 600 AESNTSNMSDCNLDEVKTVIGNAEKTEESCKKWLSIIERDCNRFCKLMSMVREDSPATEN 659
Query: 1250 ADHVQEKI 1257
H ++KI
Sbjct: 660 IVHKKKKI 667
>gb|AAH73018.1| LOC443607 protein [Xenopus laevis]
Length = 325
Score = 41.6 bits (96), Expect = 0.17
Identities = 57/254 (22%), Positives = 93/254 (36%), Gaps = 30/254 (11%)
Query: 203 DLGISSQARDFDRANTDSSDHKQFEEGNDGDCSGKIN-----DFDEEIVVTTPPDAELLG 257
DL S + ++ +DS E GNDG S + D D+E P + EL G
Sbjct: 21 DLFGSDAESENEQKESDSGSGSDSEPGNDGSGSNRSGSDSDRDEDDEHSAVKPSNKELFG 80
Query: 258 DSKVD--GDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLKSKYFSTAANHSF 315
D D G + G + + ++ N+ E + D + ND ++ + A H
Sbjct: 81 DDSEDEVGSQHSGSYNR-------SERSYNASE-TAHSDQEVNDQSDAEQHSGSEALHDE 132
Query: 316 DLDLYAGARSQTAFPGQIVQGPGFEACGGTNWPDNFVGTKKLGHCDKDEKGMGGQDFQLP 375
D D G RS + P G +A N D G + +E+ M D +L
Sbjct: 133 DNDEDVGQRSDHSSPRSEADGSDKDA----NSDDEKWGREDKSDQSDEEEKMQNSDDELQ 188
Query: 376 LSSQSEKPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNEREVSGECLSVPSGDDKC 435
S + E E G + SS E+ +E + + + + DD+
Sbjct: 189 HSDRDEHRQSDE-----------EGQRPSSDAEEQETHQSDDEEQDNHHSDDLHNSDDEE 237
Query: 436 FNESKCDPAADSDG 449
++ + SDG
Sbjct: 238 NDQQHYNDRPHSDG 251
>gb|AAH94174.1| LOC443607 protein [Xenopus laevis]
Length = 703
Score = 41.6 bits (96), Expect = 0.17
Identities = 57/254 (22%), Positives = 93/254 (36%), Gaps = 30/254 (11%)
Query: 203 DLGISSQARDFDRANTDSSDHKQFEEGNDGDCSGKIN-----DFDEEIVVTTPPDAELLG 257
DL S + ++ +DS E GNDG S + D D+E P + EL G
Sbjct: 6 DLFGSDAESENEQKESDSGSGSDSEPGNDGSGSNRSGSDSDRDEDDEHSAVKPSNKELFG 65
Query: 258 DSKVD--GDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLKSKYFSTAANHSF 315
D D G + G + + ++ N+ E + D + ND ++ + A H
Sbjct: 66 DDSEDEVGSQHSGSYNR-------SERSYNASE-TAHSDQEVNDQSDAEQHSGSEALHDE 117
Query: 316 DLDLYAGARSQTAFPGQIVQGPGFEACGGTNWPDNFVGTKKLGHCDKDEKGMGGQDFQLP 375
D D G RS + P G +A N D G + +E+ M D +L
Sbjct: 118 DNDEDVGQRSDHSSPRSEADGSDKDA----NSDDEKWGREDKSDQSDEEEKMQNSDDELQ 173
Query: 376 LSSQSEKPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNEREVSGECLSVPSGDDKC 435
S + E E G + SS E+ +E + + + + DD+
Sbjct: 174 HSDRDEHRQSDE-----------EGQRPSSDAEEQETHQSDDEEQDNHHSDDLHNSDDEE 222
Query: 436 FNESKCDPAADSDG 449
++ + SDG
Sbjct: 223 NDQQHYNDRPHSDG 236
>pir||A45605 mature-parasite-infected erythrocyte surface antigen MESA - malaria
parasite (Plasmodium falciparum)
Length = 1526
Score = 41.2 bits (95), Expect = 0.23
Identities = 41/168 (24%), Positives = 72/168 (42%), Gaps = 10/168 (5%)
Query: 257 GDSKVDGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLKSKYFSTAANHSFD 316
G+SK G+ + E ++ TG+++ +GE +SK ++KY +
Sbjct: 284 GESKETGESKETGESKETGESKETGESKETGESKETGESKETRIYEETKYNKITSEFRET 343
Query: 317 LDLYAGARSQTAFPGQIVQGPGFEACGGTNWPDNFVGTKKLGHCDKDEKGMGGQDFQLPL 376
++ S+ G V GP +E +N TKKL +++E G++
Sbjct: 344 ENVKITEESKDR-EGNKVSGP-YENSENSNVTSESEETKKLAEKEENEGEKLGENVNDGA 401
Query: 377 SSQSEKPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNEREVSGE 424
S SE P + T +E NG+K SS + + P E NE++ +
Sbjct: 402 SENSEDP-------KKLTEQEENGTKESSEETKDDKPEE-NEKKADNK 441
>sp|P08116|PVA_PLAFA Processed variable antigen gi|552170|gb|AAA29470.1| antigen
Length = 224
Score = 41.2 bits (95), Expect = 0.23
Identities = 41/168 (24%), Positives = 72/168 (42%), Gaps = 10/168 (5%)
Query: 257 GDSKVDGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLKSKYFSTAANHSFD 316
G+SK G+ + E ++ TG+++ +GE +SK ++KY +
Sbjct: 63 GESKETGESKETGESKETGESKETGESKETGESKETGESKETRIYEETKYNKITSEFRET 122
Query: 317 LDLYAGARSQTAFPGQIVQGPGFEACGGTNWPDNFVGTKKLGHCDKDEKGMGGQDFQLPL 376
++ S+ G V GP +E +N TKKL +++E G++
Sbjct: 123 ENVKITEESKDR-EGNKVSGP-YENSENSNVTSESEETKKLAEKEENEGEKLGENVNDGA 180
Query: 377 SSQSEKPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNEREVSGE 424
S SE P + T +E NG+K SS + + P E NE++ +
Sbjct: 181 SENSEDP-------KKLTEQEENGTKESSEETKDDKPEE-NEKKADNK 220
>gb|AAP33686.1| phoshoprotein 300 [Plasmodium falciparum]
Length = 273
Score = 41.2 bits (95), Expect = 0.23
Identities = 41/168 (24%), Positives = 72/168 (42%), Gaps = 10/168 (5%)
Query: 257 GDSKVDGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLKSKYFSTAANHSFD 316
G+SK G+ + E ++ TG+++ +GE +SK ++KY +
Sbjct: 15 GESKETGESKETGESKETGESKETGESKETGESKETGESKETRIYEETKYNKITSEFRET 74
Query: 317 LDLYAGARSQTAFPGQIVQGPGFEACGGTNWPDNFVGTKKLGHCDKDEKGMGGQDFQLPL 376
++ S+ G V GP +E +N TKKL +++E G++
Sbjct: 75 ENVKITEESKDR-EGNKVSGP-YENSENSNVTSESEETKKLAEKEENEGEKLGENVNDGA 132
Query: 377 SSQSEKPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNEREVSGE 424
S SE P + T +E NG+K SS + + P E NE++ +
Sbjct: 133 SENSEDP-------KKLTEQEENGTKESSEETKDDKPEE-NEKKADNK 172
>ref|NP_703354.1| Mature parasite-infected erythrocyte surface antigen (MESA) or
PfEMP2 [Plasmodium falciparum 3D7]
gi|23504494|emb|CAD51374.1| Mature parasite-infected
erythrocyte surface antigen (MESA) or PfEMP2 [Plasmodium
falciparum 3D7]
Length = 1434
Score = 41.2 bits (95), Expect = 0.23
Identities = 41/168 (24%), Positives = 72/168 (42%), Gaps = 10/168 (5%)
Query: 257 GDSKVDGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLKSKYFSTAANHSFD 316
G+SK G+ + E ++ TG+++ +GE +SK ++KY +
Sbjct: 265 GESKETGESKETGESKETGESKETGESKETGESKETGESKETRIYEETKYNKITSEFRET 324
Query: 317 LDLYAGARSQTAFPGQIVQGPGFEACGGTNWPDNFVGTKKLGHCDKDEKGMGGQDFQLPL 376
++ S+ G V GP +E +N TKKL +++E G++
Sbjct: 325 ENVKITEESKDR-EGNKVSGP-YENSENSNVTSESEETKKLAEKEENEGEKLGENVNDGA 382
Query: 377 SSQSEKPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNEREVSGE 424
S SE P + T +E NG+K SS + + P E NE++ +
Sbjct: 383 SENSEDP-------KKLTEQEENGTKESSEETKDDKPEE-NEKKADNK 422
>gb|AAO52302.1| similar to Homo sapiens (Human). Dentin sialophosphoprotein precursor
[Contains: Dentin phosphoprotein (Dentin phosphophoryn)
(DPP); Dentin sialoprotein (DSP)] [Dictyostelium
discoideum]
Length = 1206
Score = 41.2 bits (95), Expect = 0.23
Identities = 78/372 (20%), Positives = 151/372 (39%), Gaps = 39/372 (10%)
Query: 698 EKAVVGSKDECSREPNFDLCSAIVDFHSTKIVANDGHDVDIKHMQNSISSLQERGPSPKD 757
E + GS +E S + + S H+ +N G + + H N SS S +
Sbjct: 666 ESSNQGSNNESSNNESSNQGSNNESSHNE--TSNQGSNNESSH--NESSSQGSNNESAHN 721
Query: 758 QSMLYVNVGVSNQTLEHHSSEVKGFSVNNGEVKQLLNMEKRKYVSRYPSEGQSSSQLDTD 817
+S N G +N++ H+ S +G S N N S S QSS+ L ++
Sbjct: 722 ESS---NQGSNNES-SHNESSSQG-SNNESSNNDSNNQNSNNEPSNNESSNQSSNNLSSN 776
Query: 818 MQDLEENERSNGAIRKSDDAVVLNHVG----VSNEVPRPPPENIISKTQ----------- 862
+ + G+ +S + N+ +++E P + S++
Sbjct: 777 NESSNNESSNQGSNNESSHSESSNNESSNQSLNSEYNNEPSQGSNSESSNESSNQSLNSE 836
Query: 863 ---DMAAHGGNKEGSVQKGIAYGSKGNNGRKGCYAYEIENASESKITPVLNRCPRIKSLK 919
D ++HG N E S + ++GS + + + N S + + + + +
Sbjct: 837 SNNDSSSHGSNSESSNNESSSHGSNNESSNNESSSQDSSNYSSNNGSSNNESSSQGSNNE 896
Query: 920 HAGSPTYKRLLPLLSNAKKYNSCASVNRHDPKHQKFLDQTPLLPISD---SDLRATSGGD 976
+ + S+++ N+ +S N +P +Q +++ S+ S+ ++S G
Sbjct: 897 SSNQGSNNESSNNESSSQGSNNESSHN--EPSNQSSNNESSNNESSNNESSNNESSSKGS 954
Query: 977 SNDSVPMEHFTANSGTKQQKELNACELNDDSSSPRSRSPGF--QSSNSKCKVMQMCDEQV 1034
+N+S E ++N G+ + N E N++SS+ S S G +SSN++ + Q
Sbjct: 955 NNESSNNE--SSNQGSNNESSNN--ESNNESSNNESSSQGSNNESSNNESSNNE-SSSQS 1009
Query: 1035 LLNGLCKPESSS 1046
NG ESS+
Sbjct: 1010 SNNGSSNNESSN 1021
>gb|AAL79813.1| sialoprotein-phosphophoryn DSP-PP523 [Rattus norvegicus]
gi|17224418|gb|AAL36971.1| dentin sialophosphoprotein
precursor [Rattus norvegicus]
gi|18077391|emb|CAC81983.1| dentin
sialoprotein-phosphophoryn [Rattus norvegicus]
gi|18076245|emb|CAC81980.1| phosphophoryn [Rattus
norvegicus] gi|19923672|ref|NP_036922.2| dentin
sialophosphoprotein [Rattus norvegicus]
Length = 970
Score = 41.2 bits (95), Expect = 0.23
Identities = 83/444 (18%), Positives = 145/444 (31%), Gaps = 38/444 (8%)
Query: 70 SGNDEKREPPEEDDGDTLLVKYLRKKAKRDDDGGDLPRLLTKDLRARRVYSPQSSARASL 129
+G+D E+D D D +G D KD + S + +
Sbjct: 489 NGSDSDSHAGEDDSSDDT-----SDTDDSDSNGDDDSESKDKDESDNSNHDNDSDSESKS 543
Query: 130 DLIDSDCADRKIAGLGFGSPSSEEKSGVHVDDVMIAVNSDCTDRKIADLGFGSSAEKGGL 189
D DSD + S SSE D +SD +D + SS
Sbjct: 544 DSSDSDSDSSDSSDSSDSSDSSESSDSSDSSD-----SSDSSDSSDSSDSSDSSDSSDSD 598
Query: 190 CVDDATMKVTVSPDLGISSQARDF----------DRANTDSSDHKQFEEGNDGDCSGKIN 239
D + + S D SS + D D +N+DSSD + +D S +
Sbjct: 599 SSDSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSNSDSSDSSDSSDSSDSSDSSDSS 658
Query: 240 DFDEEIVVTTPPDAELLGDS-----KVDGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKD 294
D + + D+ DS D D + D+ + + +S + D
Sbjct: 659 DSSDSSDSSDSSDSSESSDSSDSSDSSDSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSD 718
Query: 295 SKRNDSVLKSKYFSTAANHSFDLDLYAGA-----RSQTAFPGQIVQGPGFEACGGTNWPD 349
S +DS S+ ++ S D + + S ++ ++ ++ D
Sbjct: 719 SSDSDSSDSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSNSDSSDSSDSSDSSDSSD 778
Query: 350 NFVGTKKLGHCDKDEKGMGGQDFQLPLSSQSEKPSEPELKSESFTMRETNGSKLSSSQNL 409
+ + D + SS S S+ S+S +++ S SS N
Sbjct: 779 SSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSNS 838
Query: 410 LESPMELNEREVSGECLSVPSGD--------DKCFNESKCDPAADSDGVKVVGNALHNDD 461
+S + + S S S D D + D + SDG G++ +D
Sbjct: 839 SDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDGDSSDGDSSDSDS 898
Query: 462 FMQCSIHKNNLDYAKVTNDSGCTS 485
S + ++ D + ++ S S
Sbjct: 899 SDSDSSNSSDSDSSDSSDSSSSDS 922
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.311 0.131 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,226,485,845
Number of Sequences: 2540612
Number of extensions: 102140237
Number of successful extensions: 214758
Number of sequences better than 10.0: 129
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 127
Number of HSP's that attempted gapping in prelim test: 214363
Number of HSP's gapped (non-prelim): 419
length of query: 1262
length of database: 863,360,394
effective HSP length: 140
effective length of query: 1122
effective length of database: 507,674,714
effective search space: 569611029108
effective search space used: 569611029108
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 81 (35.8 bits)
Lotus: description of TM0108.13


