

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0106.15
(120 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
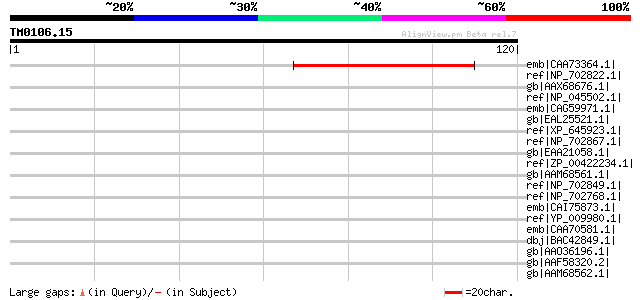
Score E
Sequences producing significant alignments: (bits) Value
emb|CAA73364.1| Pge1 protein [Lotus corniculatus var. japonicus] 61 7e-09
ref|NP_702822.1| hypothetical protein [Plasmodium falciparum 3D7... 35 0.55
gb|AAX68676.1| ABC transporter [Trichoderma atroviride] 34 0.94
ref|NP_045502.1| hypothetical protein BBH09 [Borrelia burgdorfer... 33 1.2
emb|CAG59971.1| unnamed protein product [Candida glabrata CBS138... 33 1.2
gb|EAL25521.1| GA14781-PA [Drosophila pseudoobscura] 33 2.1
ref|XP_645923.1| putative protein tyrosine kinase [Dictyostelium... 32 2.7
ref|NP_702867.1| erythrocyte membrane-associated antigen, putati... 32 3.6
gb|EAA21058.1| hypothetical protein [Plasmodium yoelii yoelii] 32 3.6
ref|ZP_00422234.1| hypothetical protein Bcep1808DRAFT_5011 [Burk... 32 4.7
gb|AAM68561.1| CG18076-PE, isoform E [Drosophila melanogaster] g... 31 6.1
ref|NP_702849.1| hypothetical protein [Plasmodium falciparum 3D7... 31 6.1
ref|NP_702768.1| hypothetical protein [Plasmodium falciparum 3D7... 31 6.1
emb|CAI75873.1| hypothetical protein [Theileria annulata] 31 6.1
ref|YP_009980.1| peptidase, M29 family [Desulfovibrio vulgaris s... 31 6.1
emb|CAA70581.1| Kakapo [Drosophila melanogaster] gi|7511874|pir|... 31 6.1
dbj|BAC42849.1| unknown protein [Arabidopsis thaliana] gi|289732... 31 6.1
gb|AAO36196.1| flagellar biosynthesis protein flhF [Clostridium ... 31 6.1
gb|AAF58320.2| CG18076-PH, isoform H [Drosophila melanogaster] g... 31 6.1
gb|AAM68562.1| CG18076-PC, isoform C [Drosophila melanogaster] g... 31 6.1
>emb|CAA73364.1| Pge1 protein [Lotus corniculatus var. japonicus]
Length = 210
Score = 60.8 bits (146), Expect = 7e-09
Identities = 26/43 (60%), Positives = 33/43 (76%)
Query: 68 VFDWAQLVGKNNDFVVIMIRSDYGTLNGKESLTLGCEHGGKFK 110
+ DWA+ VGK N +VVI+IRSDYG+ K +TLGCE GGK+K
Sbjct: 163 LIDWARCVGKENGYVVIVIRSDYGSAKRKPLITLGCERGGKYK 205
>ref|NP_702822.1| hypothetical protein [Plasmodium falciparum 3D7]
gi|23498238|emb|CAD49209.1| hypothetical protein
[Plasmodium falciparum 3D7]
Length = 802
Score = 34.7 bits (78), Expect = 0.55
Identities = 21/101 (20%), Positives = 50/101 (48%), Gaps = 6/101 (5%)
Query: 10 DNVAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVVMKFQNRQVF 69
+N E +H KV+++KE N+V+ +KE + + + ++ N ++ + + V + + +
Sbjct: 145 ENKIEEEHNKVNQLKEEHNKVNQLKEQHNKVNQLKEQHNKVNQLKEQHNKVNQLKEQHNI 204
Query: 70 DWAQLVGKNNDFVVIMIRSDYGTLNGKESLTLGCEHGGKFK 110
++ + +N I + G + + T+ +H GK K
Sbjct: 205 EFPATLKPSN------INNKKGVYHEEREETINKDHEGKIK 239
>gb|AAX68676.1| ABC transporter [Trichoderma atroviride]
Length = 1384
Score = 33.9 bits (76), Expect = 0.94
Identities = 18/58 (31%), Positives = 29/58 (49%), Gaps = 3/58 (5%)
Query: 20 VDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVVMK---FQNRQVFDWAQL 74
V+ V++VD E E GH+ ++D+ +NA D N+ + +N VF W L
Sbjct: 701 VNAVRQVDEEGQVSSESGHVSEKDDATVNAQSDNNSTDDTAAQGNLIRNSSVFTWKNL 758
>ref|NP_045502.1| hypothetical protein BBH09 [Borrelia burgdorferi B31]
gi|2690056|gb|AAC66000.1| B. burgdorferi predicted
coding region BBH09 [Borrelia burgdorferi B31]
gi|7463542|pir||B70236 hypothetical protein BBH09 - Lyme
disease spirochete plasmid H/lp28-3
Length = 1278
Score = 33.5 bits (75), Expect = 1.2
Identities = 21/62 (33%), Positives = 36/62 (57%), Gaps = 4/62 (6%)
Query: 3 HVLIEKVDNVAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDE--DLMNAIDDMNALSHVV 60
H LI +D + E ++D+ ++V E+D KE+G ILKE E D+ ++I L ++
Sbjct: 519 HFLISCLDYLTEKVWYELDKFEDVKKELD--KEYGIILKESEEYDIQDSISKELVLKRML 576
Query: 61 MK 62
+K
Sbjct: 577 LK 578
>emb|CAG59971.1| unnamed protein product [Candida glabrata CBS138]
gi|50289215|ref|XP_447038.1| unnamed protein product
[Candida glabrata]
Length = 2238
Score = 33.5 bits (75), Expect = 1.2
Identities = 21/57 (36%), Positives = 33/57 (57%), Gaps = 5/57 (8%)
Query: 5 LIEKVDNVAEGDHI-KVDRVKEVD--NEVDDMKEWGHILKEDEDLMNAIDDMNALSH 58
L+E+VD+V E DH+ +VD +E D EVD ++E H E+ D +D + + H
Sbjct: 265 LVEEVDHVEEVDHVEEVDHSEEEDLVEEVDHVEEVDH--SEEVDHSEEVDHVEEVDH 319
>gb|EAL25521.1| GA14781-PA [Drosophila pseudoobscura]
Length = 5405
Score = 32.7 bits (73), Expect = 2.1
Identities = 12/32 (37%), Positives = 20/32 (62%)
Query: 5 LIEKVDNVAEGDHIKVDRVKEVDNEVDDMKEW 36
++E++ AE H+ R KEV ++DD+ EW
Sbjct: 3563 IVEQIQRKAERLHLSNQRAKEVTGDIDDLLEW 3594
>ref|XP_645923.1| putative protein tyrosine kinase [Dictyostelium discoideum]
gi|60474038|gb|EAL71975.1| putative protein
serine/threonine kinase [Dictyostelium discoideum]
Length = 3184
Score = 32.3 bits (72), Expect = 2.7
Identities = 19/70 (27%), Positives = 34/70 (48%), Gaps = 7/70 (10%)
Query: 15 GDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVVMKFQ----NRQVFD 70
G+ + EV+N++DD+ G + K E L N+ + + +V FQ ++ D
Sbjct: 125 GEDKSAPEISEVENQIDDL---GILFKALEILTNSFNQNKDIEDIVSSFQYVGNESEIAD 181
Query: 71 WAQLVGKNND 80
++V NND
Sbjct: 182 LVKIVLSNND 191
>ref|NP_702867.1| erythrocyte membrane-associated antigen, putative [Plasmodium
falciparum 3D7] gi|23498284|emb|CAD49256.1| erythrocyte
membrane-associated antigen, putative [Plasmodium
falciparum 3D7]
Length = 4261
Score = 32.0 bits (71), Expect = 3.6
Identities = 22/61 (36%), Positives = 35/61 (57%), Gaps = 7/61 (11%)
Query: 6 IEKVDNVAEGDHIK-VDRVKEVDN--EVDDMKEWGHILKEDE----DLMNAIDDMNALSH 58
++ VDN+ D +K VD +K VD VD MK + D+ D MN+ID+MN++ +
Sbjct: 600 MKNVDNMKNVDKMKNVDDMKNVDKMKNVDKMKNVDKMKNVDKMNSVDKMNSIDNMNSMDN 659
Query: 59 V 59
+
Sbjct: 660 M 660
>gb|EAA21058.1| hypothetical protein [Plasmodium yoelii yoelii]
Length = 407
Score = 32.0 bits (71), Expect = 3.6
Identities = 15/46 (32%), Positives = 27/46 (58%), Gaps = 1/46 (2%)
Query: 22 RVKEVDNEVDDMKEWG-HILKEDEDLMNAIDDMNALSHVVMKFQNR 66
R+KE DN V +KE H+L+ ++++ N + LS ++ F N+
Sbjct: 53 RIKEEDNIVSKIKELNLHLLQNEDEINNMRQENETLSSQIINFSNK 98
>ref|ZP_00422234.1| hypothetical protein Bcep1808DRAFT_5011 [Burkholderia
vietnamiensis G4] gi|67534520|gb|EAM31264.1|
hypothetical protein Bcep1808DRAFT_5011 [Burkholderia
vietnamiensis G4]
Length = 272
Score = 31.6 bits (70), Expect = 4.7
Identities = 20/60 (33%), Positives = 33/60 (54%), Gaps = 3/60 (5%)
Query: 15 GDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVVMKFQNRQVFDWAQL 74
G+ + VDRV++V ++++DM G + D D + A D+ + V Q+RQ D A L
Sbjct: 22 GEDVSVDRVRQVQSKLEDM---GRLDGADVDSLRAAVDVARNARDVYPIQDRQRHDDAIL 78
>gb|AAM68561.1| CG18076-PE, isoform E [Drosophila melanogaster]
gi|24653491|ref|NP_725337.1| CG18076-PE, isoform E
[Drosophila melanogaster]
Length = 5201
Score = 31.2 bits (69), Expect = 6.1
Identities = 11/32 (34%), Positives = 20/32 (62%)
Query: 5 LIEKVDNVAEGDHIKVDRVKEVDNEVDDMKEW 36
++E++ AE H+ R KEV ++D++ EW
Sbjct: 3358 IVEQIQRKAERLHLSNQRAKEVTGDIDELLEW 3389
>ref|NP_702849.1| hypothetical protein [Plasmodium falciparum 3D7]
gi|23498265|emb|CAD49237.1| hypothetical protein
[Plasmodium falciparum 3D7]
Length = 3370
Score = 31.2 bits (69), Expect = 6.1
Identities = 18/86 (20%), Positives = 42/86 (47%)
Query: 7 EKVDNVAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVVMKFQNR 66
EK++NV E ++ +D +++ M E I+ E D+++ +++ + +M QN
Sbjct: 1453 EKINNVDEKKKKVDEQNNNIDEKINIMDEQNDIIDEQNDIIDEQNNIMDEQNNIMDEQND 1512
Query: 67 QVFDWAQLVGKNNDFVVIMIRSDYGT 92
+ + ++ + N+ +V D T
Sbjct: 1513 IIDEKINIMDQQNNIIVNNTYEDMNT 1538
>ref|NP_702768.1| hypothetical protein [Plasmodium falciparum 3D7]
gi|6562749|emb|CAB62888.1| Hypothetical protein
[Plasmodium falciparum 3D7]
Length = 722
Score = 31.2 bits (69), Expect = 6.1
Identities = 23/81 (28%), Positives = 40/81 (48%), Gaps = 8/81 (9%)
Query: 7 EKVDNVAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDED----LMNAIDDMNALSHVVM- 61
E + V D K +++ + DN DD+K +I+K DED + N DDMN L++ +
Sbjct: 449 ELYEKVISKDKNKEEKLVD-DNYYDDIKNECNIIKNDEDIHFNINNLDDDMNFLTYDLNY 507
Query: 62 --KFQNRQVFDWAQLVGKNND 80
N+ + ++ NN+
Sbjct: 508 NDSNNNKDIKKKENIISNNNE 528
>emb|CAI75873.1| hypothetical protein [Theileria annulata]
Length = 668
Score = 31.2 bits (69), Expect = 6.1
Identities = 20/88 (22%), Positives = 44/88 (49%), Gaps = 12/88 (13%)
Query: 5 LIEKVDNVAEGDHIKVDRVKEVDNEVDDM--------KEWGHILKEDEDLMNAIDDMNAL 56
L+ ++D + + K++++K + EV+D+ K+ H+LK+ DL N ++ +
Sbjct: 250 LVSELDELKSENQQKLEKIKSLQIEVEDLKSDLIVERKQNEHLLKDKVDLQNRLNLLTKE 309
Query: 57 SHVVMK----FQNRQVFDWAQLVGKNND 80
+ ++ QN + +L GK N+
Sbjct: 310 NKSLLSSQEIMQNMYKNEIEELKGKKNE 337
>ref|YP_009980.1| peptidase, M29 family [Desulfovibrio vulgaris subsp. vulgaris str.
Hildenborough] gi|46448585|gb|AAS95239.1| peptidase, M29
family [Desulfovibrio vulgaris subsp. vulgaris str.
Hildenborough]
Length = 401
Score = 31.2 bits (69), Expect = 6.1
Identities = 22/77 (28%), Positives = 33/77 (42%), Gaps = 13/77 (16%)
Query: 29 EVDDMKEWGHILKEDEDLMNAIDDMNALSHVVMKFQNRQVFD----------WAQLVGKN 78
E D + W I KE E L A+D AL VV+ ++ D WA + G+N
Sbjct: 176 EADPVARWRRIAKEAEALRTALD---ALGDVVLHVESDDGTDLRLSVGQQRRWAGVTGRN 232
Query: 79 NDFVVIMIRSDYGTLNG 95
+ + D+ T+ G
Sbjct: 233 IPSFELYVSPDWRTVEG 249
>emb|CAA70581.1| Kakapo [Drosophila melanogaster] gi|7511874|pir||T13734 groovin gene
protein - fruit fly (Drosophila melanogaster) (fragment)
Length = 4151
Score = 31.2 bits (69), Expect = 6.1
Identities = 11/32 (34%), Positives = 20/32 (62%)
Query: 5 LIEKVDNVAEGDHIKVDRVKEVDNEVDDMKEW 36
++E++ AE H+ R KEV ++D++ EW
Sbjct: 2312 IVEQIQRKAERLHLSNQRAKEVTGDIDELLEW 2343
>dbj|BAC42849.1| unknown protein [Arabidopsis thaliana] gi|28973251|gb|AAO63950.1|
unknown protein [Arabidopsis thaliana]
gi|15239396|ref|NP_197916.1| expressed protein
[Arabidopsis thaliana]
Length = 208
Score = 31.2 bits (69), Expect = 6.1
Identities = 11/41 (26%), Positives = 29/41 (69%), Gaps = 1/41 (2%)
Query: 14 EGDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMN 54
+GD + ++R++E+D+ DD++ G ++ +D+D+ + ++ N
Sbjct: 131 DGDEV-LNRIEEIDDSWDDLERGGLVMYKDDDMETSSNESN 170
>gb|AAO36196.1| flagellar biosynthesis protein flhF [Clostridium tetani E88]
gi|28211315|ref|NP_782259.1| flagellar biosynthesis
protein flhF [Clostridium tetani E88]
Length = 377
Score = 31.2 bits (69), Expect = 6.1
Identities = 26/92 (28%), Positives = 42/92 (45%), Gaps = 2/92 (2%)
Query: 20 VDRVKE--VDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVVMKFQNRQVFDWAQLVGK 77
+DR KE V+N +D+ ILKE E++ NA ++ + RQ + + K
Sbjct: 77 IDRKKERPVENTLDNFNGDKRILKEIEEMKNAFTSFVDNTYKGNIEEERQEIKEIKSILK 136
Query: 78 NNDFVVIMIRSDYGTLNGKESLTLGCEHGGKF 109
+ND +I LN E++T E +F
Sbjct: 137 DNDISEDIIEKILDELNEVENITEALEKTKEF 168
>gb|AAF58320.2| CG18076-PH, isoform H [Drosophila melanogaster]
gi|24653497|ref|NP_725339.1| CG18076-PH, isoform H
[Drosophila melanogaster]
Length = 8805
Score = 31.2 bits (69), Expect = 6.1
Identities = 11/32 (34%), Positives = 20/32 (62%)
Query: 5 LIEKVDNVAEGDHIKVDRVKEVDNEVDDMKEW 36
++E++ AE H+ R KEV ++D++ EW
Sbjct: 6962 IVEQIQRKAERLHLSNQRAKEVTGDIDELLEW 6993
>gb|AAM68562.1| CG18076-PC, isoform C [Drosophila melanogaster]
gi|24653495|ref|NP_725338.1| CG18076-PC, isoform C
[Drosophila melanogaster]
Length = 5160
Score = 31.2 bits (69), Expect = 6.1
Identities = 11/32 (34%), Positives = 20/32 (62%)
Query: 5 LIEKVDNVAEGDHIKVDRVKEVDNEVDDMKEW 36
++E++ AE H+ R KEV ++D++ EW
Sbjct: 3317 IVEQIQRKAERLHLSNQRAKEVTGDIDELLEW 3348
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.139 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 209,529,907
Number of Sequences: 2540612
Number of extensions: 8033492
Number of successful extensions: 21167
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 32
Number of HSP's that attempted gapping in prelim test: 21138
Number of HSP's gapped (non-prelim): 68
length of query: 120
length of database: 863,360,394
effective HSP length: 96
effective length of query: 24
effective length of database: 619,461,642
effective search space: 14867079408
effective search space used: 14867079408
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0106.15


