

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0106.12
(329 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
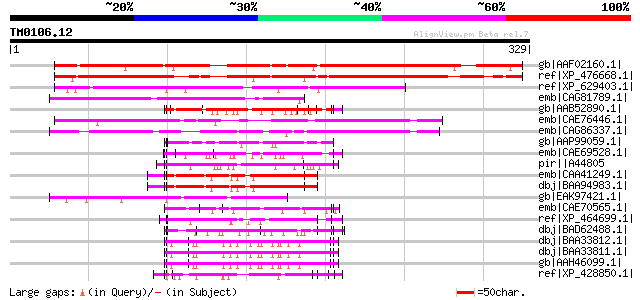
Score E
Sequences producing significant alignments: (bits) Value
gb|AAF02160.1| unknown protein [Arabidopsis thaliana] gi|2014731... 351 2e-95
ref|XP_476668.1| putative glycine-rich protein [Oryza sativa (ja... 301 2e-80
ref|XP_629403.1| hypothetical protein DDB0191662 [Dictyostelium ... 117 5e-25
emb|CAG81789.1| unnamed protein product [Yarrowia lipolytica CLI... 102 1e-20
gb|AAB52890.1| Hypothetical protein T20B6.3 [Caenorhabditis eleg... 102 2e-20
emb|CAE76446.1| related to peroxisomal membrane protein pex13 [N... 100 5e-20
emb|CAG86337.1| unnamed protein product [Debaryomyces hansenii C... 99 2e-19
gb|AAP99059.1| RNA-binding protein, RRM domain [Prochlorococcus ... 98 3e-19
emb|CAE69528.1| Hypothetical protein CBG15737 [Caenorhabditis br... 98 3e-19
pir||A44805 eggshell protein precursor - fluke (Schistosoma haem... 97 7e-19
emb|CAA41249.1| glycine rich protein [Arabidopsis thaliana] gi|7... 96 1e-18
dbj|BAA94983.1| unnamed protein product [Arabidopsis thaliana] g... 96 1e-18
gb|EAK97421.1| potential peroxisomal membrane protein Pex13 [Can... 95 3e-18
emb|CAE70565.1| Hypothetical protein CBG17212 [Caenorhabditis br... 95 3e-18
ref|XP_464699.1| putative heterogeneous nuclearribonucleoprotein... 94 4e-18
dbj|BAD62488.1| putative glycine-rich protein 1 [Oryza sativa (j... 94 6e-18
dbj|BAA33812.1| RBP56/hTAFII68 [Homo sapiens] gi|27501920|gb|AAO... 94 7e-18
dbj|BAA33811.1| RBP56/hTAFII68 [Homo sapiens] gi|21327701|ref|NP... 94 7e-18
gb|AAH46099.1| TAF15 protein [Homo sapiens] 94 7e-18
ref|XP_428850.1| PREDICTED: similar to type II keratin K6h [Gall... 93 1e-17
>gb|AAF02160.1| unknown protein [Arabidopsis thaliana] gi|20147311|gb|AAM10369.1|
AT3g07560/F21O3_27 [Arabidopsis thaliana]
gi|18700169|gb|AAL77696.1| AT3g07560/F21O3_27
[Arabidopsis thaliana] gi|15231481|ref|NP_187412.1|
glycine-rich protein [Arabidopsis thaliana]
Length = 304
Score = 351 bits (900), Expect = 2e-95
Identities = 191/319 (59%), Positives = 218/319 (67%), Gaps = 46/319 (14%)
Query: 29 PPKPWQQGGSSSGPTAPFKPPSAGSTSDVVEASGTARPGEIVSS--SDRSAAVNRNTLAR 86
PPKPW++ G++SGP PF+PPS ST+ VEASGTA PGE+V + + A N N+L+R
Sbjct: 10 PPKPWEKEGNTSGPN-PFRPPSNTSTAGSVEASGTANPGEVVPPPVNRPNTAANMNSLSR 68
Query: 87 PVPTRPWEQNTTYGGA--GGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGG 144
PVP RPWEQ YG GGYGS + SGYGSG YGS+ GG G YGGG
Sbjct: 69 PVPARPWEQQN-YGSTMGGGYGSNLGMTSGYGSGTYGSALGGY----------GSSYGGG 117
Query: 145 MYGSSMYNRGGYGGG-LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVG-----------P 192
MYG S RGGYGGG LYGSSG MYGGG GG GG MGGYG GMG G P
Sbjct: 118 MYGGSSMYRGGYGGGGLYGSSG-MYGGGAMGG-YGGTMGGYGMGMGTGMGMGMGMGMGGP 175
Query: 193 YGGDQDPNNPYGGPPSPPGFWISVMRVMQGVVNFFGRISILIDQNTQAFHMFMSALLQLF 252
YG QDPN+P+ PPSPPGFWIS +RVMQG VNFFGR+++LIDQNTQAFHMFMSALLQLF
Sbjct: 176 YGS-QDPNDPFNQPPSPPGFWISFLRVMQGAVNFFGRVAMLIDQNTQAFHMFMSALLQLF 234
Query: 253 DRSGLLYGELARFVLRLLGIKTKPNKVN--PQGPNGHMLGPNGHPLPGPHNSSANVNAIE 310
DR G+LYGELARFVLR+LG++T+P K+ PQGPNG LP PH N N +E
Sbjct: 235 DRGGMLYGELARFVLRMLGVRTRPRKMQQPPQGPNG---------LPLPHQPHGNQNYLE 285
Query: 311 GAKPAS----GGWDNVWGN 325
G K A+ GGWDNVWGN
Sbjct: 286 GPKTAAPGGGGGWDNVWGN 304
>ref|XP_476668.1| putative glycine-rich protein [Oryza sativa (japonica
cultivar-group)] gi|34395055|dbj|BAC84718.1| putative
glycine-rich protein [Oryza sativa (japonica
cultivar-group)]
Length = 280
Score = 301 bits (771), Expect = 2e-80
Identities = 168/313 (53%), Positives = 210/313 (66%), Gaps = 53/313 (16%)
Query: 29 PPKPWQQGGS--SSGPTAPFKPPSAGSTSDVVEASGTARPGEIVSSSDRSAAVN-RNTLA 85
PPKPW++ G+ +SGP APFKPPS G+TSDVVEASGTA+PGE V++++R+ + N N ++
Sbjct: 5 PPKPWERAGAEGTSGP-APFKPPSGGTTSDVVEASGTAKPGETVTATERNLSANVNNPVS 63
Query: 86 RPVPTRPWEQNTTYGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYG-GG 144
RP+P RPW+Q + YG G GYGS +Y SSYGG G YG GG
Sbjct: 64 RPMPQRPWQQTSGYGNTYG---------GYGSNMY-SSYGGF----------GNTYGSGG 103
Query: 145 MYGSSMYNR--GGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYG----------GGMGVGP 192
+YG+SMY+ GGYGG LYG SG MYGGGMY GLGG GGYG GGMG+GP
Sbjct: 104 LYGNSMYSSYGGGYGGSLYGGSG-MYGGGMYNSGLGGSYGGYGMGGMGGMGGMGGMGMGP 162
Query: 193 YGGDQDPNNPYGGPPSPPGFWISVMRVMQGVVNFFGRISILIDQNTQAFHMFMSALLQLF 252
YG +QDPN+ +G P PP W+S +RVM GVVNFFGR++ L++QNTQAF++F++A+LQLF
Sbjct: 163 YG-NQDPNS-FGPPAPPPSVWVSFLRVMHGVVNFFGRVAFLVEQNTQAFYLFITAMLQLF 220
Query: 253 DRSGLLYGELARFVLRLLGIKTKPNKVNPQGPNGHMLGPNGHPLPGPHNSSANVNAIEGA 312
DRSG+LYGELARFVLR+LGI+TK K G + GP+ GP IE
Sbjct: 221 DRSGMLYGELARFVLRMLGIRTKSKK-------GKVQGPDTPAFEGPAQ-----QFIEAP 268
Query: 313 KPASGGWDNVWGN 325
K + WDNVWGN
Sbjct: 269 K-GNNSWDNVWGN 280
>ref|XP_629403.1| hypothetical protein DDB0191662 [Dictyostelium discoideum]
gi|60462784|gb|EAL60984.1| hypothetical protein
DDB0191662 [Dictyostelium discoideum]
Length = 570
Score = 117 bits (293), Expect = 5e-25
Identities = 96/267 (35%), Positives = 120/267 (43%), Gaps = 51/267 (19%)
Query: 29 PPKPWQQ----GGSSS-----GPTAPFKPPSAGSTSDVVEASGTARPGEIVSSSDRSAAV 79
PPKPW+ GGS+S G + P P +TS +S T + + ++ S +V
Sbjct: 5 PPKPWENNSGGGGSTSMTGSGGISRPMSGPQR-TTSPGGGSSTTTQTSTLNNTGSPSTSV 63
Query: 80 NRNTLARP-VPTRPWEQNTT-----------YGGAGGYGSTMN--YNSGYGSGLYGSS-Y 124
+ RP P RPWE GG+ GY S+ Y Y SG YGSS Y
Sbjct: 64 GGPLVTRPRTPMRPWESGGAGSSSIDSFGGGVGGSSGYRSSYGGGYRDSYSSGGYGSSGY 123
Query: 125 GGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSS-GGMYG-------------- 169
G G G GG G +YGGG Y S GGYGG YGSS GG YG
Sbjct: 124 GSSYGSGGSGGYGSSLYGGGGYSS-----GGYGGSGYGSSYGGGYGSSYGSGYGSSYGGS 178
Query: 170 --GGMYGGGLGGPM-GGYGGGMGVGPYG--GDQDPNNPYGGPPSPPGFWISVMRVMQGVV 224
G YGGG G GGYGGG G G YG G + + G S S M + +V
Sbjct: 179 GYGSSYGGGYGSSYGGGYGGGYG-GGYGQRGYDNMDGKGGALQSVVSSGHSWMEALHSIV 237
Query: 225 NFFGRISILIDQNTQAFHMFMSALLQL 251
+ F R S L+D N A H S++++L
Sbjct: 238 DTFSRFSRLLDANFDAVHGSFSSIIRL 264
>emb|CAG81789.1| unnamed protein product [Yarrowia lipolytica CLIB99]
gi|50548037|ref|XP_501488.1| hypothetical protein
[Yarrowia lipolytica]
Length = 412
Score = 102 bits (255), Expect = 1e-20
Identities = 77/170 (45%), Positives = 89/170 (52%), Gaps = 19/170 (11%)
Query: 26 SNPPPKPWQQGGSSSGPTA-PFKPPSAGSTSDVVEASGTARPGEIVSSSDRSAAVNRNTL 84
S P PKPW+ SS TA P + ST V ++G+A + S + N L
Sbjct: 2 SVPRPKPWEGASGSSAATATPAATATPASTDAVSSSAGSATGAPELPSRPSAMGSTSNAL 61
Query: 85 ARPVPTRPWEQNTTYGGAG-GYGST-MNYNSGYGSGL-YGSSYGGGLGGGLGG-GLGGGM 140
+ P+ + N+ YGG GYG +Y SGYGS GSSYG G G GLGG G GGM
Sbjct: 62 SSPMGS---SMNSGYGGMNSGYGGMGSSYGSGYGSSYGMGSSYGSGYGSGLGGYGSYGGM 118
Query: 141 YG-GGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMG--GYGGG 187
G GGMYGS YGG YGS GGM G G YGG GGPMG G GG
Sbjct: 119 GGMGGMYGSR------YGG--YGSYGGMGGYGGYGGMGGGPMGQNGLAGG 160
>gb|AAB52890.1| Hypothetical protein T20B6.3 [Caenorhabditis elegans]
gi|17555156|ref|NP_497637.1| predicted CDS, putative
protein (3D984) [Caenorhabditis elegans]
gi|7508036|pir||T15126 hypothetical protein T20B6.3 -
Caenorhabditis elegans
Length = 259
Score = 102 bits (254), Expect = 2e-20
Identities = 65/114 (57%), Positives = 69/114 (60%), Gaps = 14/114 (12%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGL-GGGLGGGMYGGGMYGSSMYNRGGYG 157
YGG+GG G+ +N GYG G YG GG GGG+ GGG GGG YGGG G Y GGYG
Sbjct: 79 YGGSGG-GAFGGFNGGYGQGGYGG--GGDFGGGMGGGGYGGGGYGGGGDGGGGY--GGYG 133
Query: 158 GGLYGSSG---GMYGGGMY--GGGLGGPMGGY-GGGMGVGPYGGDQDPNNPYGG 205
GG YG G G YG G Y GGG GG GGY GGGMG G YGG D YGG
Sbjct: 134 GGGYGGMGGGPGGYGMGGYGGGGGGGGDFGGYGGGGMGGGGYGGGGD--GGYGG 185
Score = 100 bits (250), Expect = 5e-20
Identities = 63/116 (54%), Positives = 68/116 (58%), Gaps = 13/116 (11%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGG--LGGGLGGGLGGGMYGGGMYGSSMYNRGGY 156
YGG GGYG GYG G YG GGG GG GGG+GGG YGGG G Y GG+
Sbjct: 132 YGG-GGYGGMGGGPGGYGMGGYGGGGGGGGDFGGYGGGGMGGGGYGGG--GDGGYGGGGF 188
Query: 157 GGG-LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGGPPSPPG 211
GGG + G GGM GGG GGG+GG GGYGGG G G YG P+ YGG P G
Sbjct: 189 GGGGMGGYGGGMGGGGYGGGGMGG--GGYGGG-GDGGYG----PSGGYGGGYGPGG 237
Score = 100 bits (250), Expect = 5e-20
Identities = 63/123 (51%), Positives = 65/123 (52%), Gaps = 20/123 (16%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGG-----GMYGSSMYNR 153
YGG G +G M GYG G YG GGG GGG GG GGG YGG G YG Y
Sbjct: 99 YGGGGDFGGGMG-GGGYGGGGYG---GGGDGGGGYGGYGGGGYGGMGGGPGGYGMGGYGG 154
Query: 154 GGYGGGLYGS-SGGMYGGGMYGGGL----------GGPMGGYGGGMGVGPYGGDQDPNNP 202
GG GGG +G GG GGG YGGG GG MGGYGGGMG G YGG
Sbjct: 155 GGGGGGDFGGYGGGGMGGGGYGGGGDGGYGGGGFGGGGMGGYGGGMGGGGYGGGGMGGGG 214
Query: 203 YGG 205
YGG
Sbjct: 215 YGG 217
Score = 89.4 bits (220), Expect = 1e-16
Identities = 55/106 (51%), Positives = 58/106 (53%), Gaps = 8/106 (7%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGG-LGGGLGGGMYGGGMYGSSMYNRGGYGG 158
GG GGYG G G G +G GGG+GGG GGG GG GGG G M GGYGG
Sbjct: 142 GGPGGYGMGGYGGGGGGGGDFGGYGGGGMGGGGYGGGGDGGYGGGGFGGGGM---GGYGG 198
Query: 159 GLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYG 204
G+ G GG GGGM GGG GG GG GG G YGG P YG
Sbjct: 199 GMGG--GGYGGGGMGGGGYGG--GGDGGYGPSGGYGGGYGPGGGYG 240
Score = 87.4 bits (215), Expect = 5e-16
Identities = 58/117 (49%), Positives = 59/117 (49%), Gaps = 15/117 (12%)
Query: 103 GGYGSTMNYNSGYGSGLYGSSYGGGLGG-----GLGGGLGGGMYGGGMYGSSMYNRGGYG 157
G G YG G YG S GG GG G GG GGG +GGGM G Y GGYG
Sbjct: 62 GQRGPFAAMTGSYGRGGYGGSGGGAFGGFNGGYGQGGYGGGGDFGGGM-GGGGYGGGGYG 120
Query: 158 GGLYGSSG-GMYGGGMYGGGLGGP----MGGYGGGMG----VGPYGGDQDPNNPYGG 205
GG G G G YGGG YGG GGP MGGYGGG G G YGG YGG
Sbjct: 121 GGGDGGGGYGGYGGGGYGGMGGGPGGYGMGGYGGGGGGGGDFGGYGGGGMGGGGYGG 177
Score = 84.3 bits (207), Expect = 4e-15
Identities = 53/93 (56%), Positives = 53/93 (56%), Gaps = 6/93 (6%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGG- 158
G GGYG GYG G G GGG GGG GG GGGM GGG YG GGYGG
Sbjct: 160 GDFGGYGGGGMGGGGYGGGGDGGYGGGGFGGGGMGGYGGGM-GGGGYGGGGMGGGGYGGG 218
Query: 159 --GLYGSSGGMYGGGMYGGGLGGPMGGYGGGMG 189
G YG SGG YGGG YG G G MGG GGG G
Sbjct: 219 GDGGYGPSGG-YGGG-YGPGGGYGMGGGGGGGG 249
Score = 82.0 bits (201), Expect = 2e-14
Identities = 57/103 (55%), Positives = 58/103 (55%), Gaps = 12/103 (11%)
Query: 99 YGGAGGYGSTMNYNSGYGSG-LYGSSYGGGLGGGLGGGL--GGGM--YGGGMYGSSMYNR 153
YGG GG G GYG G + G YGGG GG GGG GGGM YGGGM G Y
Sbjct: 152 YGGGGGGGGDFG---GYGGGGMGGGGYGGGGDGGYGGGGFGGGGMGGYGGGMGGGG-YGG 207
Query: 154 GGYGGGLYGSSG-GMYG-GGMYGGGLGGPMGGYGGGMGVGPYG 194
GG GGG YG G G YG G YGGG GP GGYG G G G G
Sbjct: 208 GGMGGGGYGGGGDGGYGPSGGYGGGY-GPGGGYGMGGGGGGGG 249
Score = 68.9 bits (167), Expect = 2e-10
Identities = 46/93 (49%), Positives = 48/93 (51%), Gaps = 24/93 (25%)
Query: 123 SYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMG 182
SYG G GG GGG GG GG Y +GGYGGG G +GGGM GGG GG G
Sbjct: 73 SYGRGGYGGSGGGAFGGFNGG-------YGQGGYGGG------GDFGGGMGGGGYGG--G 117
Query: 183 GYG----GGMGVGPYGGDQDPNNPYGGPPSPPG 211
GYG GG G G YGG YGG PG
Sbjct: 118 GYGGGGDGGGGYGGYGG-----GGYGGMGGGPG 145
Score = 65.1 bits (157), Expect = 3e-09
Identities = 46/85 (54%), Positives = 48/85 (56%), Gaps = 15/85 (17%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGL-GGGLGGGMYGGGMYGSSMYNRGGYG 157
YGG G G M GYG G+ G YGGG G+ GGG GGG G G YG S GGYG
Sbjct: 183 YGGGGFGGGGMG---GYGGGMGGGGYGGG---GMGGGGYGGG--GDGGYGPS----GGYG 230
Query: 158 GGLYGSSGGMYGGGMYGGGLGGPMG 182
GG YG GG YG G GGG G P G
Sbjct: 231 GG-YGPGGG-YGMGGGGGGGGCPNG 253
>emb|CAE76446.1| related to peroxisomal membrane protein pex13 [Neurospora crassa]
gi|32422747|ref|XP_331817.1| hypothetical protein
[Neurospora crassa] gi|28926821|gb|EAA35785.1|
hypothetical protein [Neurospora crassa]
Length = 457
Score = 100 bits (250), Expect = 5e-20
Identities = 78/252 (30%), Positives = 116/252 (45%), Gaps = 42/252 (16%)
Query: 29 PPKPWQQGGSSSGPTAPFKPPSAGST----SDVVEASGTARPGEIVSSSDRSAAVNRNTL 84
PPKPW++ G++SG +AP S+ +T S+ + + P + +A+VN+N
Sbjct: 4 PPKPWERQGATSGASAPAPAASSAATPSPESNTTTTTSGSTPAIPARPASLTASVNQNAA 63
Query: 85 ARPVPTRPWEQNTTYGGAGGYGSTMNYNSGYGSGLYGSSYGGGLG--GGLGGGLGGGMYG 142
P + G G YG+T Y S YGS Y S Y GG+GG
Sbjct: 64 NYSRPMGAMGTSPYGVGVGSYGTT--YGSSYGSA-YSSPYSSPYSRYGGMGG-------- 112
Query: 143 GGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNP 202
YGSSMY G YGG MYGGG+ G G YGGGMG G DPNNP
Sbjct: 113 ---YGSSMY--------------GGYGGSMYGGGMYGGGGMYGGGMG----GMPGDPNNP 151
Query: 203 YGGPPSPPGFWISVMRVMQGVVNFFGRISILIDQNTQAFHMFMSALLQLFDRSGLLYGEL 262
++ ++++G+V FG + +++ A H A++ + ++ +G L
Sbjct: 152 ESLTNRFNMSTMATFQMLEGIVGAFGGFAQMLESTYMATHSSFFAMVAVAEQ----FGNL 207
Query: 263 ARFVLRLLGIKT 274
+ + +LGI T
Sbjct: 208 RQTLGSILGIFT 219
>emb|CAG86337.1| unnamed protein product [Debaryomyces hansenii CBS767]
gi|50419467|ref|XP_458260.1| unnamed protein product
[Debaryomyces hansenii]
Length = 446
Score = 98.6 bits (244), Expect = 2e-19
Identities = 80/255 (31%), Positives = 112/255 (43%), Gaps = 38/255 (14%)
Query: 26 SNPPPKPWQQGGSSSGPTAPFKPPSAGSTSDVVEASGTARPGEIVSSSDRSAAVNRNTLA 85
S P KPW+ +S TA G + + T P SS+ +L
Sbjct: 2 SAPRTKPWEVSSGTSTATA------TGDNMNSINGGATTNPIGDNSSTQAQVPDRPTSLT 55
Query: 86 RPVPTRPWEQNTTYGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGL-GGGLGGGMYGGG 144
+ + NT+ G+ GYGST GS YGGG G + GGGLG MYG G
Sbjct: 56 SDLSSM---SNTSPYGSTGYGSTT-----------GSRYGGGYGSSMYGGGLGSSMYGDG 101
Query: 145 MYGSSMYNRGGYGGGLYGSSGGMYGGGMYG-------GGLGGPMGGYGGGMGVGPYGGDQ 197
YGSSMY GG G +Y GG YG MYG GG+GG MGGYG G G+G YGG
Sbjct: 102 -YGSSMYGSGGLGSSMY---GGGYGSSMYGMGSMGSMGGMGG-MGGYGMG-GMGGYGGGM 155
Query: 198 DPNNPYGGPPSPPGFWISVMRVMQGVVNFFGRISILIDQNTQAFHMFMSALLQLFDRSGL 257
G + ++++ ++ G + +++ A H ++ + ++
Sbjct: 156 YGQQRPGMGGGLAEGTQATFQLIESIIGAVGGFAQMLEATYMATHSSFFTMISVAEQ--- 212
Query: 258 LYGELARFVLRLLGI 272
+G L + LLGI
Sbjct: 213 -FGNLKNALGSLLGI 226
>gb|AAP99059.1| RNA-binding protein, RRM domain [Prochlorococcus marinus subsp.
marinus str. CCMP1375] gi|33239465|ref|NP_874407.1|
RNA-binding protein, RRM domain [Prochlorococcus marinus
subsp. marinus str. CCMP1375]
Length = 252
Score = 98.2 bits (243), Expect = 3e-19
Identities = 58/112 (51%), Positives = 59/112 (51%), Gaps = 6/112 (5%)
Query: 99 YGGAGGYGSTMNYNSG-YGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYG 157
YGG GGYG Y G YG G YGGG GG GGG GG GGG G +GGYG
Sbjct: 99 YGGGGGYGGGGGYGGGGYGGGGGQGGYGGGGQGGYGGGGQGGYGGGGQGGYGGGGQGGYG 158
Query: 158 GGLYGSSG----GMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
GG G G G YGGG GG GG GGYGGG G G YGG QD G
Sbjct: 159 GGGQGGYGGGGQGGYGGGGQGGYGGGGQGGYGGG-GQGGYGGGQDTEQRASG 209
Score = 85.1 bits (209), Expect = 3e-15
Identities = 59/114 (51%), Positives = 60/114 (51%), Gaps = 20/114 (17%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGM----YGSSMYNRGG 155
GG GG G GYG G G YGGG G G GGG GGG YGGG YG +GG
Sbjct: 82 GGGGGRGG------GYGGGGRGG-YGGGGGYGGGGGYGGGGYGGGGGQGGYGGG--GQGG 132
Query: 156 YGGGLYGSSGGM----YGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
YGGG G GG YGGG GG GG GGYGGG G G YGG YGG
Sbjct: 133 YGGGGQGGYGGGGQGGYGGGGQGGYGGGGQGGYGGG-GQGGYGG--GGQGGYGG 183
Score = 80.9 bits (198), Expect = 5e-14
Identities = 57/106 (53%), Positives = 57/106 (53%), Gaps = 12/106 (11%)
Query: 101 GAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGL 160
G GG G Y G G G YG GGG GG GGG GGG YGGG G Y GG GG
Sbjct: 81 GGGGGGRGGGYGGG-GRGGYGG--GGGYGG--GGGYGGGGYGGG-GGQGGYGGGGQGG-- 132
Query: 161 YGSSG-GMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
YG G G YGGG GG GG GGYGGG G G YGG YGG
Sbjct: 133 YGGGGQGGYGGGGQGGYGGGGQGGYGGG-GQGGYGG--GGQGGYGG 175
Score = 79.3 bits (194), Expect = 1e-13
Identities = 53/108 (49%), Positives = 54/108 (49%), Gaps = 15/108 (13%)
Query: 101 GAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGL-GGGMYGGGMYGSSMYNRGGYGGG 159
GA G + N G G GGG GGG GG GGG YGGG GGYGGG
Sbjct: 64 GAELMGRPLRINKAEPRGGGGGGRGGGYGGGGRGGYGGGGGYGGG---------GGYGGG 114
Query: 160 LYGSSGGM--YGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
YG GG YGGG GG GG GGYGGG G G YGG YGG
Sbjct: 115 GYGGGGGQGGYGGGGQGGYGGGGQGGYGGG-GQGGYGG--GGQGGYGG 159
Score = 39.3 bits (90), Expect = 0.16
Identities = 32/73 (43%), Positives = 34/73 (45%), Gaps = 11/73 (15%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGG------GLGG-GMYGGGMYGSSMYN 152
GG GGYG GYG G G YGGG GG GG G GG G YGGG +
Sbjct: 152 GGQGGYGG--GGQGGYGGGGQGG-YGGGGQGGYGGGGQGGYGGGGQGGYGGGQDTEQRAS 208
Query: 153 RG-GYGGGLYGSS 164
G+ YGSS
Sbjct: 209 GAQGWEDRSYGSS 221
>emb|CAE69528.1| Hypothetical protein CBG15737 [Caenorhabditis briggsae]
Length = 259
Score = 98.2 bits (243), Expect = 3e-19
Identities = 62/127 (48%), Positives = 65/127 (50%), Gaps = 17/127 (13%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGG----GMYGGGMYGSSMYNRGG 155
G GGYG G G G YG YGGG GG+GGG GG G GGG YG GG
Sbjct: 114 GPMGGYGGGYGNGGGDGGGGYGG-YGGGGYGGMGGGPGGYGMGGFGGGGDYGGYGGGMGG 172
Query: 156 YGGGLYGSSGGMYGGGM----------YGGGLGGPMGGYGGGMGVGPYGGD-QDPNNPYG 204
GGG G G YGGG YGGG+GG GGY GGMG G YGGD P+ YG
Sbjct: 173 GGGGYGGGGDGGYGGGQGGFGGGGMGGYGGGMGGGGGGY-GGMGGGGYGGDGYGPSGGYG 231
Query: 205 GPPSPPG 211
G P G
Sbjct: 232 GGYGPGG 238
Score = 93.2 bits (230), Expect = 9e-18
Identities = 60/123 (48%), Positives = 64/123 (51%), Gaps = 18/123 (14%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSS-----YGGGLGGGLGGGLGGGM---YGGGMYGSSM 150
YGG GGYG GYG G +G YGGG+GGG GGG GGG YGGG G
Sbjct: 137 YGG-GGYGGMGGGPGGYGMGGFGGGGDYGGYGGGMGGG-GGGYGGGGDGGYGGGQGGFGG 194
Query: 151 YNRGGYGGGLYGSSGGM--YGGGMYGGGLGGPMGGY------GGGMGVGPYGGDQDPNNP 202
GGYGGG+ G GG GGG YGG GP GGY GGG G+G GG PN
Sbjct: 195 GGMGGYGGGMGGGGGGYGGMGGGGYGGDGYGPSGGYGGGYGPGGGYGMGGGGGGGCPNGE 254
Query: 203 YGG 205
G
Sbjct: 255 CRG 257
Score = 89.0 bits (219), Expect = 2e-16
Identities = 61/123 (49%), Positives = 64/123 (51%), Gaps = 19/123 (15%)
Query: 98 TYGGAGGYGSTMN-------YNSGYGSGLYGSSYG--GGLGGGLGGGLG-GGMYGGGMYG 147
+YGG GGYG +N GYG G YG G GG GG GGG G GG GGG YG
Sbjct: 76 SYGGGGGYGGGNGGGGAFGGFNGGYGQGGYGGGGGDFGGPMGGYGGGYGNGGGDGGGGYG 135
Query: 148 SSMYNRGGYGG-----GLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNP 202
Y GGYGG G YG GG GGG YGG GG MGG GGG G G GG
Sbjct: 136 G--YGGGGYGGMGGGPGGYGM-GGFGGGGDYGG-YGGGMGGGGGGYGGGGDGGYGGGQGG 191
Query: 203 YGG 205
+GG
Sbjct: 192 FGG 194
Score = 80.5 bits (197), Expect = 6e-14
Identities = 53/109 (48%), Positives = 60/109 (54%), Gaps = 17/109 (15%)
Query: 106 GSTMNYNSGYGSGLYGSSYGGGLG-----GGLGGGLGGGMY--GGGMYGSSMYNRGGYGG 158
G +++ YGS G YGGG G GG GG G G Y GGG +G M GGYGG
Sbjct: 65 GPFASFSGNYGSYGGGGGYGGGNGGGGAFGGFNGGYGQGGYGGGGGDFGGPM---GGYGG 121
Query: 159 GLYGSSGGMYGGGM--YGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
G YG+ GG GGG YGGG G MGG GG G+G +GG D YGG
Sbjct: 122 G-YGNGGGDGGGGYGGYGGGGYGGMGGGPGGYGMGGFGGGGD----YGG 165
Score = 64.7 bits (156), Expect = 4e-09
Identities = 41/86 (47%), Positives = 43/86 (49%), Gaps = 11/86 (12%)
Query: 130 GGLGGGLGGGMYGGGMYGSSMYN--RGGYGGGLYGSSGGMYGG--GMYGGGLGGPMGGYG 185
G G GGG YGGG G + GGYG G YG GG +GG G YGGG G GG
Sbjct: 72 GNYGSYGGGGGYGGGNGGGGAFGGFNGGYGQGGYGGGGGDFGGPMGGYGGGYGN--GGGD 129
Query: 186 GGMGVGPYGGDQDPNNPYGGPPSPPG 211
GG G G YGG YGG PG
Sbjct: 130 GGGGYGGYGG-----GGYGGMGGGPG 150
Score = 43.9 bits (102), Expect = 0.007
Identities = 29/62 (46%), Positives = 32/62 (50%), Gaps = 8/62 (12%)
Query: 150 MYNRGGYGGGLYGSSG--GMYGGG-MYGGGL--GGPMGGYGGGMGVGPYGGDQDPNNPYG 204
M +RG G SG G YGGG YGGG GG GG+ GG G G YGG +G
Sbjct: 57 MRSRGARRGPFASFSGNYGSYGGGGGYGGGNGGGGAFGGFNGGYGQGGYGGG---GGDFG 113
Query: 205 GP 206
GP
Sbjct: 114 GP 115
>pir||A44805 eggshell protein precursor - fluke (Schistosoma haematobium)
Length = 220
Score = 97.1 bits (240), Expect = 7e-19
Identities = 59/116 (50%), Positives = 60/116 (50%), Gaps = 11/116 (9%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLG-------GGMYGGGMYGSSMY 151
YGG GG G N G G G G YGGG GG GGG G GG YGGG G
Sbjct: 52 YGGGGGGGYEGGGNGGGGGGYEGGRYGGGGGGYEGGGYGGGGGGYEGGGYGGG--GGGYE 109
Query: 152 NRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVG--PYGGDQDPNNPYGG 205
GGYGGG G G YGGG GGG GG GGYGGG G YG D+ PYGG
Sbjct: 110 GGGGYGGGCNGDDCGGYGGGGGGGGGGGGGGGYGGGCNGGGCGYGFDEAFPAPYGG 165
Score = 85.9 bits (211), Expect = 2e-15
Identities = 60/121 (49%), Positives = 61/121 (49%), Gaps = 15/121 (12%)
Query: 94 EQNTTYGGAGGYGSTMNYNSGYGSGLYGSS-YGGGLGGGL---GGGLGGGMYGGGMYGSS 149
E YGG YGS + GYG G YG YGGG GGG G G GGG Y GG YG
Sbjct: 24 ESPNDYGG-DCYGS--DCGGGYGDGGYGGGGYGGGGGGGYEGGGNGGGGGGYEGGRYGG- 79
Query: 150 MYNRGGYGGGLYGSSGGMYGGGMYGGGLGG--PMGGYGGGMG---VGPYGGDQDPNNPYG 204
GGY GG YG GG Y GG YGGG GG GGYGGG G YGG G
Sbjct: 80 --GGGGYEGGGYGGGGGGYEGGGYGGGGGGYEGGGGYGGGCNGDDCGGYGGGGGGGGGGG 137
Query: 205 G 205
G
Sbjct: 138 G 138
Score = 85.5 bits (210), Expect = 2e-15
Identities = 57/119 (47%), Positives = 60/119 (49%), Gaps = 17/119 (14%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGL------GGGLGGGLGGGMYGGGMYGSSMYNR 153
G A Y + + YG YGS GGG GGG GGG GGG GGG G
Sbjct: 14 GYATAYQISHESPNDYGGDCYGSDCGGGYGDGGYGGGGYGGGGGGGYEGGGNGGGG---- 69
Query: 154 GGYGGGLYGSSGGMYGGGMYGGGLGG-PMGGYGGG----MGVGPYGG--DQDPNNPYGG 205
GGY GG YG GG Y GG YGGG GG GGYGGG G G YGG + D YGG
Sbjct: 70 GGYEGGRYGGGGGGYEGGGYGGGGGGYEGGGYGGGGGGYEGGGGYGGGCNGDDCGGYGG 128
Score = 68.6 bits (166), Expect = 3e-10
Identities = 48/119 (40%), Positives = 49/119 (40%), Gaps = 15/119 (12%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GG GGY GYG G G GGG GGG G GG GGG G GGYGGG
Sbjct: 91 GGGGGYEG-----GGYGGGGGGYEGGGGYGGGCNGDDCGGYGGGGGGGGGGGGGGGYGGG 145
Query: 160 LYGSSGGM---------YGGGM-YGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGGPPS 208
G G YGG YGG G GG G G G GG GG P+
Sbjct: 146 CNGGGCGYGFDEAFPAPYGGNQGYGGNGHGKGGGKDNGHGKGHGGGKGGSKGGKGGKPN 204
Score = 67.4 bits (163), Expect = 6e-10
Identities = 51/113 (45%), Positives = 52/113 (45%), Gaps = 24/113 (21%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGG-----GLGGGLGGGMYGGGMYGSSMYNR 153
YGG GG Y G G G G YGGG GG G GGG G GG YG
Sbjct: 77 YGGGGGGYEGGGYGGG-GGGYEGGGYGGGGGGYEGGGGYGGGCNGDDCGG--YG------ 127
Query: 154 GGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGV-GPYGGDQDPNNPYGG 205
GG GGG G GG YGGG GGG GYG PYGG+Q YGG
Sbjct: 128 GGGGGGGGGGGGGGYGGGCNGGGC-----GYGFDEAFPAPYGGNQG----YGG 171
Score = 57.4 bits (137), Expect = 6e-07
Identities = 43/104 (41%), Positives = 47/104 (44%), Gaps = 26/104 (25%)
Query: 99 YGGAGGYGSTMNYNS-----GYGSGLYGSSYGGGLGGGL-GGGLGGGM-------YGGGM 145
Y G GGYG N + G G G G GGG GGG GGG G G YGG
Sbjct: 108 YEGGGGYGGGCNGDDCGGYGGGGGGGGGGGGGGGYGGGCNGGGCGYGFDEAFPAPYGGNQ 167
Query: 146 YGSSMYNRGGYGGGLYGSSGGM---YGGGMYGGGLGGPMGGYGG 186
GYGG +G GG +G G +GGG GG GG GG
Sbjct: 168 ---------GYGGNGHGKGGGKDNGHGKG-HGGGKGGSKGGKGG 201
>emb|CAA41249.1| glycine rich protein [Arabidopsis thaliana] gi|72320|pir||KNMU
glycine-rich cell wall protein precursor - Arabidopsis
thaliana
Length = 338
Score = 95.9 bits (237), Expect = 1e-18
Identities = 49/96 (51%), Positives = 56/96 (58%), Gaps = 1/96 (1%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GG GG G G+G G +G GGGLGGG GGG GGG GG G+ GG+GGG
Sbjct: 201 GGGGGAGGGGGLGGGHGGG-FGGGAGGGLGGGAGGGTGGGFGGGAGGGAGGGAGGGFGGG 259
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G +GG +GGG GG GG GG+GGG G G GG
Sbjct: 260 AGGGAGGGFGGGAGGGAGGGAGGGFGGGAGGGHGGG 295
Score = 91.3 bits (225), Expect = 4e-17
Identities = 50/96 (52%), Positives = 57/96 (59%), Gaps = 2/96 (2%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GGAGG G+ G+G G+ G + GGG GGGLGGG+GGG GG G GG GGG
Sbjct: 158 GGAGG-GAGGGLGGGHGGGIGGGA-GGGSGGGLGGGIGGGAGGGAGGGGGAGGGGGLGGG 215
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G GG GGG+ GG GG GG+GGG G G GG
Sbjct: 216 HGGGFGGGAGGGLGGGAGGGTGGGFGGGAGGGAGGG 251
Score = 90.9 bits (224), Expect = 5e-17
Identities = 56/117 (47%), Positives = 64/117 (53%), Gaps = 10/117 (8%)
Query: 88 VPTRPWEQNTTYGGAGG-YGSTMNYNSGYGSGL-YGSSYGGGLGGGLGGGLGGGMYGGGM 145
V R +E+ T GG GG G G+G G+ G +GGG GGG GGGLGGG GGG
Sbjct: 10 VECRRFEKETLGGGGGGGLGGGFGGGKGFGGGIGAGGGFGGGAGGGAGGGLGGGAGGGGG 69
Query: 146 YGSSMYNR------GGYGGGLYGSSGGMYGGGMYGGGLGGPM-GGYGGGMGVGPYGG 195
G GG GGGL G GG GGG GGG GG + GG+GGG+G G GG
Sbjct: 70 IGGGAGGGAGGGLGGGAGGGLGGGHGGGIGGGA-GGGAGGGLGGGHGGGIGGGAGGG 125
Score = 90.1 bits (222), Expect = 8e-17
Identities = 52/96 (54%), Positives = 56/96 (58%), Gaps = 2/96 (2%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GGAGG G + G G G G S GGGLGGG+GGG GGG GGG G GG+GGG
Sbjct: 162 GGAGG-GLGGGHGGGIGGGAGGGS-GGGLGGGIGGGAGGGAGGGGGAGGGGGLGGGHGGG 219
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G +GG GGG GG GG GG GGG G G GG
Sbjct: 220 FGGGAGGGLGGGAGGGTGGGFGGGAGGGAGGGAGGG 255
Score = 89.4 bits (220), Expect = 1e-16
Identities = 52/96 (54%), Positives = 58/96 (60%), Gaps = 8/96 (8%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GGAGG G + G G G G S GGGLGGG+GGG GGG GGG G GG+GGG
Sbjct: 104 GGAGG-GLGGGHGGGIGGGAGGGS-GGGLGGGIGGGAGGGAGGGGGLG------GGHGGG 155
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
+ G +GG GGG+ GG GG GG GGG G G GG
Sbjct: 156 IGGGAGGGAGGGLGGGHGGGIGGGAGGGSGGGLGGG 191
Score = 89.4 bits (220), Expect = 1e-16
Identities = 48/98 (48%), Positives = 57/98 (57%), Gaps = 2/98 (2%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGG 158
+GG G G+ G+G G G + GGG GGG GGG GGG GG G+ + GG GG
Sbjct: 240 FGGGAGGGAGGGAGGGFGGGAGGGA-GGGFGGGAGGGAGGGAGGGFGGGAGGGHGGGVGG 298
Query: 159 GLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGV-GPYGG 195
G G SGG +GGG GG GG GG+GGG G G +GG
Sbjct: 299 GFGGGSGGGFGGGAGGGAGGGAGGGFGGGGGAGGGFGG 336
Score = 88.6 bits (218), Expect = 2e-16
Identities = 51/96 (53%), Positives = 58/96 (60%), Gaps = 8/96 (8%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GGAGG G G+G G+ G + GGG GGGLGGG GGG+ GG GS GG GGG
Sbjct: 140 GGAGGGGGL---GGGHGGGIGGGA-GGGAGGGLGGGHGGGIGGGAGGGSG----GGLGGG 191
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
+ G +GG GGG GG GG GG+GGG G G GG
Sbjct: 192 IGGGAGGGAGGGGGAGGGGGLGGGHGGGFGGGAGGG 227
Score = 87.8 bits (216), Expect = 4e-16
Identities = 51/96 (53%), Positives = 55/96 (57%), Gaps = 2/96 (2%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GGAGG G G G G GGGLGGG GGG+GGG GG G + GG GGG
Sbjct: 62 GGAGGGGGIGGGAGGGAGGGLGGGAGGGLGGGHGGGIGGGAGGGAGGGLGGGHGGGIGGG 121
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G SGG GGG+ GGG GG GG GGG+G G GG
Sbjct: 122 AGGGSGGGLGGGI-GGGAGGGAGG-GGGLGGGHGGG 155
Score = 87.4 bits (215), Expect = 5e-16
Identities = 50/96 (52%), Positives = 59/96 (61%), Gaps = 6/96 (6%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GGAGG G+ + G G G G + GGG GGG GGG GGG +GGG G + GG GGG
Sbjct: 230 GGAGG-GTGGGFGGGAGGGAGGGA-GGGFGGGAGGGAGGG-FGGGAGGGA---GGGAGGG 283
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G +GG +GGG+ GG GG GG+GGG G G GG
Sbjct: 284 FGGGAGGGHGGGVGGGFGGGSGGGFGGGAGGGAGGG 319
Score = 87.0 bits (214), Expect = 7e-16
Identities = 48/97 (49%), Positives = 56/97 (57%), Gaps = 1/97 (1%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGG-LGGGLGGGLGGGMYGGGMYGSSMYNRGGYGG 158
G GG G + +G G+G G + GGG LGGG GGG GGG GG G+ GG+GG
Sbjct: 183 GSGGGLGGGIGGGAGGGAGGGGGAGGGGGLGGGHGGGFGGGAGGGLGGGAGGGTGGGFGG 242
Query: 159 GLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G G +GG GGG GG GG GG+GGG G G GG
Sbjct: 243 GAGGGAGGGAGGGFGGGAGGGAGGGFGGGAGGGAGGG 279
Score = 86.7 bits (213), Expect = 9e-16
Identities = 50/96 (52%), Positives = 57/96 (59%), Gaps = 3/96 (3%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GG G G G G GL G +GGG+GGG GGG GGG+ GGG+ G + GG GGG
Sbjct: 91 GGGHGGGIGGGAGGGAGGGL-GGGHGGGIGGGAGGGSGGGL-GGGIGGGAGGGAGG-GGG 147
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
L G GG GGG GG GG GG+GGG+G G GG
Sbjct: 148 LGGGHGGGIGGGAGGGAGGGLGGGHGGGIGGGAGGG 183
Score = 85.9 bits (211), Expect = 2e-15
Identities = 48/97 (49%), Positives = 54/97 (55%), Gaps = 1/97 (1%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGG 158
+GG G G+ G G G +G GGG GGG GGGLGGG+ GG G+ G GG
Sbjct: 152 HGGGIGGGAGGGAGGGLGGG-HGGGIGGGAGGGSGGGLGGGIGGGAGGGAGGGGGAGGGG 210
Query: 159 GLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
GL G GG +GGG GG GG GG GGG G G GG
Sbjct: 211 GLGGGHGGGFGGGAGGGLGGGAGGGTGGGFGGGAGGG 247
Score = 85.9 bits (211), Expect = 2e-15
Identities = 51/96 (53%), Positives = 57/96 (59%), Gaps = 4/96 (4%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GGAGG G+ G G GL G +GGG+GGG GGG GGG+ GG G GG GGG
Sbjct: 72 GGAGG-GAGGGLGGGAGGGL-GGGHGGGIGGGAGGGAGGGLGGGHGGGIGGGAGGGSGGG 129
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
L G GG GGG GG GG GG+GGG+G G GG
Sbjct: 130 LGGGIGGGAGGG--AGGGGGLGGGHGGGIGGGAGGG 163
Score = 85.9 bits (211), Expect = 2e-15
Identities = 48/97 (49%), Positives = 55/97 (56%), Gaps = 5/97 (5%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGG 158
+GG G G+ G G G G +GGG GGG GGG GGG +GGG G + GG+GG
Sbjct: 216 HGGGFGGGAGGGLGGGAGGGT-GGGFGGGAGGGAGGGAGGG-FGGGAGGGA---GGGFGG 270
Query: 159 GLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G G +GG GGG GG GG GG GGG G G GG
Sbjct: 271 GAGGGAGGGAGGGFGGGAGGGHGGGVGGGFGGGSGGG 307
Score = 85.9 bits (211), Expect = 2e-15
Identities = 52/97 (53%), Positives = 56/97 (57%), Gaps = 3/97 (3%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGG-LGGGLGGGLGGGMYGGGMYGSSMYNRGGYGG 158
GGAGG GS G G G G + GGG LGGG GGG+GGG GG G + GG GG
Sbjct: 120 GGAGG-GSGGGLGGGIGGGAGGGAGGGGGLGGGHGGGIGGGAGGGAGGGLGGGHGGGIGG 178
Query: 159 GLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G G SGG GGG+ GGG GG GG GG G G GG
Sbjct: 179 GAGGGSGGGLGGGI-GGGAGGGAGGGGGAGGGGGLGG 214
Score = 85.5 bits (210), Expect = 2e-15
Identities = 51/96 (53%), Positives = 57/96 (59%), Gaps = 6/96 (6%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GGAGG GS G G G G + GGG GG GGGLGGG +GGG G + GG GGG
Sbjct: 178 GGAGG-GSGGGLGGGIGGGAGGGAGGGGGAGG-GGGLGGG-HGGGFGGGA---GGGLGGG 231
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G +GG +GGG GG GG GG+GGG G G GG
Sbjct: 232 AGGGTGGGFGGGAGGGAGGGAGGGFGGGAGGGAGGG 267
Score = 85.1 bits (209), Expect = 3e-15
Identities = 47/102 (46%), Positives = 58/102 (56%), Gaps = 5/102 (4%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGG-----GLGGGLGGGLGGGMYGGGMYGSSMYNR 153
+GG G+G + G+G G G + GG G GGG+GGG GGG GG G+
Sbjct: 32 FGGGKGFGGGIGAGGGFGGGAGGGAGGGLGGGAGGGGGIGGGAGGGAGGGLGGGAGGGLG 91
Query: 154 GGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
GG+GGG+ G +GG GGG+ GG GG GG GGG G G GG
Sbjct: 92 GGHGGGIGGGAGGGAGGGLGGGHGGGIGGGAGGGSGGGLGGG 133
Score = 85.1 bits (209), Expect = 3e-15
Identities = 52/97 (53%), Positives = 57/97 (58%), Gaps = 7/97 (7%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
G GG G G G G G + GGGLGGG GGGLGGG +GGG+ G + GG GGG
Sbjct: 55 GAGGGLGGGAGGGGGIGGGAGGGA-GGGLGGGAGGGLGGG-HGGGIGGGA---GGGAGGG 109
Query: 160 LYGSSGGMYGGGMYGGGLGGPM-GGYGGGMGVGPYGG 195
L G GG GGG GGG GG + GG GGG G G GG
Sbjct: 110 LGGGHGGGIGGGA-GGGSGGGLGGGIGGGAGGGAGGG 145
Score = 84.3 bits (207), Expect = 4e-15
Identities = 51/98 (52%), Positives = 60/98 (61%), Gaps = 4/98 (4%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GG GG G+ G+G G+ G + GGG GGGLGGG GGG+ GG GS GG GGG
Sbjct: 80 GGLGG-GAGGGLGGGHGGGIGGGA-GGGAGGGLGGGHGGGIGGGAGGGSGGGLGGGIGGG 137
Query: 160 LYGSSGGMYG-GGMYGGGLGGPM-GGYGGGMGVGPYGG 195
G +GG G GG +GGG+GG GG GGG+G G GG
Sbjct: 138 AGGGAGGGGGLGGGHGGGIGGGAGGGAGGGLGGGHGGG 175
Score = 82.8 bits (203), Expect = 1e-14
Identities = 53/99 (53%), Positives = 58/99 (58%), Gaps = 16/99 (16%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGG--GMYGSSMYNRGGYG 157
GGAGG G +G G GL G +GGG GGG GGGLGGG GG G +G GG G
Sbjct: 198 GGAGGGGG-----AGGGGGL-GGGHGGGFGGGAGGGLGGGAGGGTGGGFG------GGAG 245
Query: 158 GGLYGSSGGMYGGGMYGGGLGGPM-GGYGGGMGVGPYGG 195
GG G +GG +GGG GGG GG GG GGG G G GG
Sbjct: 246 GGAGGGAGGGFGGGA-GGGAGGGFGGGAGGGAGGGAGGG 283
Score = 74.7 bits (182), Expect = 3e-12
Identities = 44/92 (47%), Positives = 54/92 (57%), Gaps = 13/92 (14%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGG---MYGGGMYGSSMYNRGG 155
+GG G G+ + G G G G + GGG GGG GGG GGG +GGG + GG
Sbjct: 256 FGGGAGGGAGGGFGGGAGGGAGGGA-GGGFGGGAGGGHGGGVGGGFGGG-------SGGG 307
Query: 156 YGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGG 187
+GGG G +GG GGG +GGG GG GG+GGG
Sbjct: 308 FGGGAGGGAGGGAGGG-FGGG-GGAGGGFGGG 337
Score = 42.4 bits (98), Expect = 0.019
Identities = 30/78 (38%), Positives = 33/78 (41%), Gaps = 25/78 (32%)
Query: 128 LGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGG 187
LGGG GGGLGGG +GGG G+GGG+ G GG GG GG
Sbjct: 20 LGGGGGGGLGGG-FGGGK---------GFGGGI---------------GAGGGFGGGAGG 54
Query: 188 MGVGPYGGDQDPNNPYGG 205
G GG GG
Sbjct: 55 GAGGGLGGGAGGGGGIGG 72
>dbj|BAA94983.1| unnamed protein product [Arabidopsis thaliana]
gi|27735191|sp|P27483|GRP_ARATH Glycine-rich cell wall
structural protein precursor
Length = 349
Score = 95.9 bits (237), Expect = 1e-18
Identities = 49/96 (51%), Positives = 56/96 (58%), Gaps = 1/96 (1%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GG GG G G+G G +G GGGLGGG GGG GGG GG G+ GG+GGG
Sbjct: 212 GGGGGAGGGGGLGGGHGGG-FGGGAGGGLGGGAGGGTGGGFGGGAGGGAGGGAGGGFGGG 270
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G +GG +GGG GG GG GG+GGG G G GG
Sbjct: 271 AGGGAGGGFGGGAGGGAGGGAGGGFGGGAGGGHGGG 306
Score = 91.3 bits (225), Expect = 4e-17
Identities = 50/96 (52%), Positives = 57/96 (59%), Gaps = 2/96 (2%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GGAGG G+ G+G G+ G + GGG GGGLGGG+GGG GG G GG GGG
Sbjct: 169 GGAGG-GAGGGLGGGHGGGIGGGA-GGGSGGGLGGGIGGGAGGGAGGGGGAGGGGGLGGG 226
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G GG GGG+ GG GG GG+GGG G G GG
Sbjct: 227 HGGGFGGGAGGGLGGGAGGGTGGGFGGGAGGGAGGG 262
Score = 90.9 bits (224), Expect = 5e-17
Identities = 56/117 (47%), Positives = 64/117 (53%), Gaps = 10/117 (8%)
Query: 88 VPTRPWEQNTTYGGAGG-YGSTMNYNSGYGSGL-YGSSYGGGLGGGLGGGLGGGMYGGGM 145
V R +E+ T GG GG G G+G G+ G +GGG GGG GGGLGGG GGG
Sbjct: 21 VECRRFEKETLGGGGGGGLGGGFGGGKGFGGGIGAGGGFGGGAGGGAGGGLGGGAGGGGG 80
Query: 146 YGSSMYNR------GGYGGGLYGSSGGMYGGGMYGGGLGGPM-GGYGGGMGVGPYGG 195
G GG GGGL G GG GGG GGG GG + GG+GGG+G G GG
Sbjct: 81 IGGGAGGGAGGGLGGGAGGGLGGGHGGGIGGGA-GGGAGGGLGGGHGGGIGGGAGGG 136
Score = 90.1 bits (222), Expect = 8e-17
Identities = 52/96 (54%), Positives = 56/96 (58%), Gaps = 2/96 (2%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GGAGG G + G G G G S GGGLGGG+GGG GGG GGG G GG+GGG
Sbjct: 173 GGAGG-GLGGGHGGGIGGGAGGGS-GGGLGGGIGGGAGGGAGGGGGAGGGGGLGGGHGGG 230
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G +GG GGG GG GG GG GGG G G GG
Sbjct: 231 FGGGAGGGLGGGAGGGTGGGFGGGAGGGAGGGAGGG 266
Score = 89.4 bits (220), Expect = 1e-16
Identities = 52/96 (54%), Positives = 58/96 (60%), Gaps = 8/96 (8%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GGAGG G + G G G G S GGGLGGG+GGG GGG GGG G GG+GGG
Sbjct: 115 GGAGG-GLGGGHGGGIGGGAGGGS-GGGLGGGIGGGAGGGAGGGGGLG------GGHGGG 166
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
+ G +GG GGG+ GG GG GG GGG G G GG
Sbjct: 167 IGGGAGGGAGGGLGGGHGGGIGGGAGGGSGGGLGGG 202
Score = 89.4 bits (220), Expect = 1e-16
Identities = 48/98 (48%), Positives = 57/98 (57%), Gaps = 2/98 (2%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGG 158
+GG G G+ G+G G G + GGG GGG GGG GGG GG G+ + GG GG
Sbjct: 251 FGGGAGGGAGGGAGGGFGGGAGGGA-GGGFGGGAGGGAGGGAGGGFGGGAGGGHGGGVGG 309
Query: 159 GLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGV-GPYGG 195
G G SGG +GGG GG GG GG+GGG G G +GG
Sbjct: 310 GFGGGSGGGFGGGAGGGAGGGAGGGFGGGGGAGGGFGG 347
Score = 88.6 bits (218), Expect = 2e-16
Identities = 51/96 (53%), Positives = 58/96 (60%), Gaps = 8/96 (8%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GGAGG G G+G G+ G + GGG GGGLGGG GGG+ GG GS GG GGG
Sbjct: 151 GGAGGGGGL---GGGHGGGIGGGA-GGGAGGGLGGGHGGGIGGGAGGGSG----GGLGGG 202
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
+ G +GG GGG GG GG GG+GGG G G GG
Sbjct: 203 IGGGAGGGAGGGGGAGGGGGLGGGHGGGFGGGAGGG 238
Score = 87.8 bits (216), Expect = 4e-16
Identities = 51/96 (53%), Positives = 55/96 (57%), Gaps = 2/96 (2%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GGAGG G G G G GGGLGGG GGG+GGG GG G + GG GGG
Sbjct: 73 GGAGGGGGIGGGAGGGAGGGLGGGAGGGLGGGHGGGIGGGAGGGAGGGLGGGHGGGIGGG 132
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G SGG GGG+ GGG GG GG GGG+G G GG
Sbjct: 133 AGGGSGGGLGGGI-GGGAGGGAGG-GGGLGGGHGGG 166
Score = 87.4 bits (215), Expect = 5e-16
Identities = 50/96 (52%), Positives = 59/96 (61%), Gaps = 6/96 (6%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GGAGG G+ + G G G G + GGG GGG GGG GGG +GGG G + GG GGG
Sbjct: 241 GGAGG-GTGGGFGGGAGGGAGGGA-GGGFGGGAGGGAGGG-FGGGAGGGA---GGGAGGG 294
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G +GG +GGG+ GG GG GG+GGG G G GG
Sbjct: 295 FGGGAGGGHGGGVGGGFGGGSGGGFGGGAGGGAGGG 330
Score = 87.0 bits (214), Expect = 7e-16
Identities = 48/97 (49%), Positives = 56/97 (57%), Gaps = 1/97 (1%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGG-LGGGLGGGLGGGMYGGGMYGSSMYNRGGYGG 158
G GG G + +G G+G G + GGG LGGG GGG GGG GG G+ GG+GG
Sbjct: 194 GSGGGLGGGIGGGAGGGAGGGGGAGGGGGLGGGHGGGFGGGAGGGLGGGAGGGTGGGFGG 253
Query: 159 GLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G G +GG GGG GG GG GG+GGG G G GG
Sbjct: 254 GAGGGAGGGAGGGFGGGAGGGAGGGFGGGAGGGAGGG 290
Score = 86.7 bits (213), Expect = 9e-16
Identities = 50/96 (52%), Positives = 57/96 (59%), Gaps = 3/96 (3%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GG G G G G GL G +GGG+GGG GGG GGG+ GGG+ G + GG GGG
Sbjct: 102 GGGHGGGIGGGAGGGAGGGL-GGGHGGGIGGGAGGGSGGGL-GGGIGGGAGGGAGG-GGG 158
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
L G GG GGG GG GG GG+GGG+G G GG
Sbjct: 159 LGGGHGGGIGGGAGGGAGGGLGGGHGGGIGGGAGGG 194
Score = 85.9 bits (211), Expect = 2e-15
Identities = 48/97 (49%), Positives = 54/97 (55%), Gaps = 1/97 (1%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGG 158
+GG G G+ G G G +G GGG GGG GGGLGGG+ GG G+ G GG
Sbjct: 163 HGGGIGGGAGGGAGGGLGGG-HGGGIGGGAGGGSGGGLGGGIGGGAGGGAGGGGGAGGGG 221
Query: 159 GLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
GL G GG +GGG GG GG GG GGG G G GG
Sbjct: 222 GLGGGHGGGFGGGAGGGLGGGAGGGTGGGFGGGAGGG 258
Score = 85.9 bits (211), Expect = 2e-15
Identities = 51/96 (53%), Positives = 57/96 (59%), Gaps = 4/96 (4%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GGAGG G+ G G GL G +GGG+GGG GGG GGG+ GG G GG GGG
Sbjct: 83 GGAGG-GAGGGLGGGAGGGL-GGGHGGGIGGGAGGGAGGGLGGGHGGGIGGGAGGGSGGG 140
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
L G GG GGG GG GG GG+GGG+G G GG
Sbjct: 141 LGGGIGGGAGGG--AGGGGGLGGGHGGGIGGGAGGG 174
Score = 85.9 bits (211), Expect = 2e-15
Identities = 48/97 (49%), Positives = 55/97 (56%), Gaps = 5/97 (5%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGG 158
+GG G G+ G G G G +GGG GGG GGG GGG +GGG G + GG+GG
Sbjct: 227 HGGGFGGGAGGGLGGGAGGGT-GGGFGGGAGGGAGGGAGGG-FGGGAGGGA---GGGFGG 281
Query: 159 GLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G G +GG GGG GG GG GG GGG G G GG
Sbjct: 282 GAGGGAGGGAGGGFGGGAGGGHGGGVGGGFGGGSGGG 318
Score = 85.9 bits (211), Expect = 2e-15
Identities = 52/97 (53%), Positives = 56/97 (57%), Gaps = 3/97 (3%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGG-LGGGLGGGLGGGMYGGGMYGSSMYNRGGYGG 158
GGAGG GS G G G G + GGG LGGG GGG+GGG GG G + GG GG
Sbjct: 131 GGAGG-GSGGGLGGGIGGGAGGGAGGGGGLGGGHGGGIGGGAGGGAGGGLGGGHGGGIGG 189
Query: 159 GLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G G SGG GGG+ GGG GG GG GG G G GG
Sbjct: 190 GAGGGSGGGLGGGI-GGGAGGGAGGGGGAGGGGGLGG 225
Score = 85.5 bits (210), Expect = 2e-15
Identities = 51/96 (53%), Positives = 57/96 (59%), Gaps = 6/96 (6%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GGAGG GS G G G G + GGG GG GGGLGGG +GGG G + GG GGG
Sbjct: 189 GGAGG-GSGGGLGGGIGGGAGGGAGGGGGAGG-GGGLGGG-HGGGFGGGA---GGGLGGG 242
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
G +GG +GGG GG GG GG+GGG G G GG
Sbjct: 243 AGGGTGGGFGGGAGGGAGGGAGGGFGGGAGGGAGGG 278
Score = 85.1 bits (209), Expect = 3e-15
Identities = 47/102 (46%), Positives = 58/102 (56%), Gaps = 5/102 (4%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGG-----GLGGGLGGGLGGGMYGGGMYGSSMYNR 153
+GG G+G + G+G G G + GG G GGG+GGG GGG GG G+
Sbjct: 43 FGGGKGFGGGIGAGGGFGGGAGGGAGGGLGGGAGGGGGIGGGAGGGAGGGLGGGAGGGLG 102
Query: 154 GGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
GG+GGG+ G +GG GGG+ GG GG GG GGG G G GG
Sbjct: 103 GGHGGGIGGGAGGGAGGGLGGGHGGGIGGGAGGGSGGGLGGG 144
Score = 85.1 bits (209), Expect = 3e-15
Identities = 52/97 (53%), Positives = 57/97 (58%), Gaps = 7/97 (7%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
G GG G G G G G + GGGLGGG GGGLGGG +GGG+ G + GG GGG
Sbjct: 66 GAGGGLGGGAGGGGGIGGGAGGGA-GGGLGGGAGGGLGGG-HGGGIGGGA---GGGAGGG 120
Query: 160 LYGSSGGMYGGGMYGGGLGGPM-GGYGGGMGVGPYGG 195
L G GG GGG GGG GG + GG GGG G G GG
Sbjct: 121 LGGGHGGGIGGGA-GGGSGGGLGGGIGGGAGGGAGGG 156
Score = 84.3 bits (207), Expect = 4e-15
Identities = 51/98 (52%), Positives = 60/98 (61%), Gaps = 4/98 (4%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
GG GG G+ G+G G+ G + GGG GGGLGGG GGG+ GG GS GG GGG
Sbjct: 91 GGLGG-GAGGGLGGGHGGGIGGGA-GGGAGGGLGGGHGGGIGGGAGGGSGGGLGGGIGGG 148
Query: 160 LYGSSGGMYG-GGMYGGGLGGPM-GGYGGGMGVGPYGG 195
G +GG G GG +GGG+GG GG GGG+G G GG
Sbjct: 149 AGGGAGGGGGLGGGHGGGIGGGAGGGAGGGLGGGHGGG 186
Score = 82.8 bits (203), Expect = 1e-14
Identities = 53/99 (53%), Positives = 58/99 (58%), Gaps = 16/99 (16%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGG--GMYGSSMYNRGGYG 157
GGAGG G +G G GL G +GGG GGG GGGLGGG GG G +G GG G
Sbjct: 209 GGAGGGGG-----AGGGGGL-GGGHGGGFGGGAGGGLGGGAGGGTGGGFG------GGAG 256
Query: 158 GGLYGSSGGMYGGGMYGGGLGGPM-GGYGGGMGVGPYGG 195
GG G +GG +GGG GGG GG GG GGG G G GG
Sbjct: 257 GGAGGGAGGGFGGGA-GGGAGGGFGGGAGGGAGGGAGGG 294
Score = 74.7 bits (182), Expect = 3e-12
Identities = 44/92 (47%), Positives = 54/92 (57%), Gaps = 13/92 (14%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGG---MYGGGMYGSSMYNRGG 155
+GG G G+ + G G G G + GGG GGG GGG GGG +GGG + GG
Sbjct: 267 FGGGAGGGAGGGFGGGAGGGAGGGA-GGGFGGGAGGGHGGGVGGGFGGG-------SGGG 318
Query: 156 YGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGG 187
+GGG G +GG GGG +GGG GG GG+GGG
Sbjct: 319 FGGGAGGGAGGGAGGG-FGGG-GGAGGGFGGG 348
Score = 42.4 bits (98), Expect = 0.019
Identities = 30/78 (38%), Positives = 33/78 (41%), Gaps = 25/78 (32%)
Query: 128 LGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGG 187
LGGG GGGLGGG +GGG G+GGG+ G GG GG GG
Sbjct: 31 LGGGGGGGLGGG-FGGGK---------GFGGGI---------------GAGGGFGGGAGG 65
Query: 188 MGVGPYGGDQDPNNPYGG 205
G GG GG
Sbjct: 66 GAGGGLGGGAGGGGGIGG 83
>gb|EAK97421.1| potential peroxisomal membrane protein Pex13 [Candida albicans
SC5314]
Length = 371
Score = 94.7 bits (234), Expect = 3e-18
Identities = 65/164 (39%), Positives = 84/164 (50%), Gaps = 16/164 (9%)
Query: 26 SNPPPKPWQ--QGGSSSGPTAPFK---PPSAGSTSDVVEASGTARPGEI----VSSSDRS 76
S P KPW+ QG S++ T P PPS+ S S + +I V S
Sbjct: 4 SQPRTKPWEVSQGKSAASTTEPLTSNIPPSSLSNSSTTATAQGQPQSQIQPPSVPELPTS 63
Query: 77 AAVNRNTLARPVPTRPWEQNTT--YGGAGGYGSTMNYNSGYGSGLYGSS-YGGGLGGGL- 132
+ + NT+A P + ++ YG GGYGS+M Y GYGS +YG+S YGGG G +
Sbjct: 64 LSSDLNTMAESSPYGGYGNSSMNRYGTTGGYGSSM-YGGGYGSSMYGNSMYGGGYGSSMY 122
Query: 133 GGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGG 176
GGG GG YG G G Y G YGGG+YG++ M G G G G
Sbjct: 123 GGGYGG--YGSGYGGYGSYGSGMYGGGMYGNNPSMMGQGGLGEG 164
>emb|CAE70565.1| Hypothetical protein CBG17212 [Caenorhabditis briggsae]
Length = 161
Score = 94.7 bits (234), Expect = 3e-18
Identities = 63/130 (48%), Positives = 67/130 (51%), Gaps = 23/130 (17%)
Query: 99 YGGAGGYGSTMNYNSGYGSGL-YGSSYGGG--LGGGLGGGLGGGM--YGGGMYGSSMYNR 153
YGG GGYG + GYG G G YGGG +GGG GGG GG YGGG YG+
Sbjct: 35 YGG-GGYGGGGDMGGGYGGGGDMGGGYGGGGDMGGGYGGGGDGGYGGYGGGGYGAGGDMG 93
Query: 154 GGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDP--------NNPYGG 205
GGYG G G GG YGGG YGG +GG G GGG G G Y G P PYGG
Sbjct: 94 GGYGMGGGGGDGG-YGGGGYGGNMGG--GNMGGGGGYGSYQGGPQPGGGQGGYGGQPYGG 150
Query: 206 ------PPSP 209
PP P
Sbjct: 151 GGQGGWPPMP 160
Score = 84.7 bits (208), Expect = 3e-15
Identities = 55/114 (48%), Positives = 59/114 (51%), Gaps = 16/114 (14%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGG 158
+GG GGYG + GYG G YG GG +GGG GGG GG YG GGYGG
Sbjct: 20 WGGGGGYGGGGDMG-GYGGGGYGG------GGDMGGGYGGGGDMGGGYGGGGDMGGGYGG 72
Query: 159 GLYGSSGGMYGGGMYGGGLGGPMGGYG-------GGMGVGPYGGDQDPNNPYGG 205
G G GG YGGG YG G G GGYG GG G G YGG+ N GG
Sbjct: 73 GGDGGYGG-YGGGGYGAG-GDMGGGYGMGGGGGDGGYGGGGYGGNMGGGNMGGG 124
Score = 69.7 bits (169), Expect = 1e-10
Identities = 47/86 (54%), Positives = 48/86 (55%), Gaps = 7/86 (8%)
Query: 121 GSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGG--MYGGGLG 178
G +GGG G G GGG GG YGGG YG GGYGGG G GG YGGG M GG G
Sbjct: 17 GQDWGGGGGYG-GGGDMGG-YGGGGYGGGGDMGGGYGGG--GDMGGGYGGGGDMGGGYGG 72
Query: 179 GPMGGYGGGMGVGPYGGDQDPNNPYG 204
G GGY GG G G YG D YG
Sbjct: 73 GGDGGY-GGYGGGGYGAGGDMGGGYG 97
Score = 57.0 bits (136), Expect = 8e-07
Identities = 35/70 (50%), Positives = 37/70 (52%), Gaps = 2/70 (2%)
Query: 136 LGGGMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
LG GGG YG + GGYGGG YG G M GG GG +GG GG GG MG G GG
Sbjct: 16 LGQDWGGGGGYGGG-GDMGGYGGGGYGGGGDMGGGYGGGGDMGGGYGG-GGDMGGGYGGG 73
Query: 196 DQDPNNPYGG 205
YGG
Sbjct: 74 GDGGYGGYGG 83
>ref|XP_464699.1| putative heterogeneous nuclearribonucleoprotein A2 [Oryza sativa
(japonica cultivar-group)] gi|46806519|dbj|BAD17632.1|
putative heterogeneous nuclearribonucleoprotein A2
[Oryza sativa (japonica cultivar-group)]
gi|46806500|dbj|BAD17624.1| putative heterogeneous
nuclearribonucleoprotein A2 [Oryza sativa (japonica
cultivar-group)]
Length = 397
Score = 94.4 bits (233), Expect = 4e-18
Identities = 54/108 (50%), Positives = 57/108 (52%), Gaps = 19/108 (17%)
Query: 100 GGAGGYGSTMNYNS-----------GYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGS 148
G +GGYG Y S GYG G YG G G G GGG GG +YGG YG+
Sbjct: 278 GSSGGYGYGAGYRSAGGGYYDSTAYGYGRGGYGYGGNAGFGSGYGGGYGGSLYGGA-YGA 336
Query: 149 SMYNRGGYGGGLYGSSGGMYGGGMYGGG-LGGPMGGYGGGMGVGPYGG 195
G YGGG YG GG YGGG YGGG GG G YGG G G YGG
Sbjct: 337 ----YGAYGGGAYG--GGAYGGGAYGGGAYGGAPGAYGGAGGYGSYGG 378
Score = 87.8 bits (216), Expect = 4e-16
Identities = 53/112 (47%), Positives = 58/112 (51%), Gaps = 17/112 (15%)
Query: 96 NTTYG-GAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGG--GMYGGGMYGSSMYN 152
+T YG G GGYG N+G+GSG YGGG GG L GG G G YGGG YG Y
Sbjct: 299 STAYGYGRGGYG--YGGNAGFGSG-----YGGGYGGSLYGGAYGAYGAYGGGAYGGGAYG 351
Query: 153 RGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYG 204
G YGGG YG + G YGG G GG G GG G Y +PYG
Sbjct: 352 GGAYGGGAYGGAPGAYGGAGGYGSYGGAGAGGAGGRGSSRY-------HPYG 396
Score = 73.9 bits (180), Expect = 6e-12
Identities = 53/132 (40%), Positives = 56/132 (42%), Gaps = 28/132 (21%)
Query: 101 GAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGG--------------MYGGGMY 146
G +GS Y S Y SG G GG GGG GGG GGG Y
Sbjct: 244 GRSSHGSGGGYRSSYRSG------GAAASGGGGGGGGGGSGSSGGYGYGAGYRSAGGGYY 297
Query: 147 GSSM--YNRGGYG----GGLYGSSGGMYGGGMYGGGLGGPMGGYGGG-MGVGPYGGDQDP 199
S+ Y RGGYG G GG YGG +YGG G G YGGG G G YGG
Sbjct: 298 DSTAYGYGRGGYGYGGNAGFGSGYGGGYGGSLYGGAYGA-YGAYGGGAYGGGAYGGGAYG 356
Query: 200 NNPYGGPPSPPG 211
YGG P G
Sbjct: 357 GGAYGGAPGAYG 368
Score = 36.2 bits (82), Expect = 1.4
Identities = 27/69 (39%), Positives = 31/69 (44%), Gaps = 9/69 (13%)
Query: 143 GGMYGSSMYNRGGYGGGL---YGSSGGMYGGGMYGGGLGG----PMGGYGGGMGVGPYGG 195
GG + S+ + G GGG Y S G GG GGG GG GGYG G G GG
Sbjct: 237 GGDHSSNGRSSHGSGGGYRSSYRSGGAAASGG--GGGGGGGGSGSSGGYGYGAGYRSAGG 294
Query: 196 DQDPNNPYG 204
+ YG
Sbjct: 295 GYYDSTAYG 303
>dbj|BAD62488.1| putative glycine-rich protein 1 [Oryza sativa (japonica
cultivar-group)] gi|54291363|dbj|BAD62129.1| putative
glycine-rich protein 1 [Oryza sativa (japonica
cultivar-group)]
Length = 246
Score = 94.0 bits (232), Expect = 6e-18
Identities = 56/118 (47%), Positives = 59/118 (49%), Gaps = 7/118 (5%)
Query: 101 GAGGYGSTMNYNSGYG---SGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYG 157
G GGYG Y SGYG G+ G YGGG G G GGG G G YG G YG N GG G
Sbjct: 46 GNGGYGHGYGYGSGYGFGNGGVGGGGYGGGGGYGSGGGEGNGAYGQG-YGYGSGNGGGGG 104
Query: 158 GGLYGSSGGMYGGGMYGGGLGGPMGG---YGGGMGVGPYGGDQDPNNPYGGPPSPPGF 212
GG G GG YG G G G GG G YGG G G +GG + YG GF
Sbjct: 105 GGYGGGGGGSYGSGGMGSGYGGGYGSGYDYGGQGGGGGHGGGGGGGSGYGNGGYGSGF 162
Score = 92.8 bits (229), Expect = 1e-17
Identities = 57/117 (48%), Positives = 59/117 (49%), Gaps = 12/117 (10%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGG--LGGGLGGGMYGGGMYGSSMYNRGGYG 157
GG GGYG YGSG GS YGGG G G GG GGG +GGG G S Y GGYG
Sbjct: 102 GGGGGYGG--GGGGSYGSGGMGSGYGGGYGSGYDYGGQGGGGGHGGGGGGGSGYGNGGYG 159
Query: 158 GGL---YGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGGPPSPPG 211
G YGS GG+ GGG GGG GG GGG YG YGG P G
Sbjct: 160 SGFGEGYGSGGGVNGGGGSGGG-----GGGGGGASGYRYGAGYGKGYGYGGGPGGGG 211
Score = 88.6 bits (218), Expect = 2e-16
Identities = 58/113 (51%), Positives = 60/113 (52%), Gaps = 14/113 (12%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGG 158
YGG GGYGS G G+G YG YG G G G GGG G G GGG YGS GYGG
Sbjct: 72 YGGGGGYGS----GGGEGNGAYGQGYGYGSGNGGGGGGGYGGGGGGSYGSGGMG-SGYGG 126
Query: 159 GLYGSS---GGMYGGGMYGGGLGGPM----GGYGGGMGVGPYGGDQDPNNPYG 204
G YGS GG GGG +GGG GG GGYG G G G YG N G
Sbjct: 127 G-YGSGYDYGGQGGGGGHGGGGGGGSGYGNGGYGSGFGEG-YGSGGGVNGGGG 177
Score = 88.6 bits (218), Expect = 2e-16
Identities = 59/110 (53%), Positives = 61/110 (54%), Gaps = 11/110 (10%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLG-GGLGGGMYG-GGMYGS-SMYNRGGY 156
GG GGYGS G G+G YG YG G G G G GG+GGG YG GG YGS G Y
Sbjct: 34 GGGGGYGS----GGGEGNGGYGHGYGYGSGYGFGNGGVGGGGYGGGGGYGSGGGEGNGAY 89
Query: 157 GGGL-YGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
G G YGS G GGG YGGG GG G GGMG G YGG YGG
Sbjct: 90 GQGYGYGSGNGGGGGGGYGGGGGGSYG--SGGMGSG-YGGGYGSGYDYGG 136
Score = 84.0 bits (206), Expect = 6e-15
Identities = 52/105 (49%), Positives = 56/105 (52%), Gaps = 11/105 (10%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLG----GGLGGGMYGGG-----MYGSSM 150
GG GG+G SGYG+G YGS +G G G G G GG GGG GGG YG+
Sbjct: 138 GGGGGHGGGGGGGSGYGNGGYGSGFGEGYGSGGGVNGGGGSGGGGGGGGGASGYRYGAGY 197
Query: 151 YNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
GYGGG G GG GGG GG G GGYG G G G YGG
Sbjct: 198 GKGYGYGGGPGGGGGGA-GGGGGGGSYNGGTGGYGEGHGSG-YGG 240
Score = 82.4 bits (202), Expect = 2e-14
Identities = 58/118 (49%), Positives = 59/118 (49%), Gaps = 24/118 (20%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSY---GGGLGGGLGGGLGGGM-YGGGMYGSSM----- 150
GG G YGS SGYG G YGS Y G G GGG GGG GGG YG G YGS
Sbjct: 110 GGGGSYGSG-GMGSGYGGG-YGSGYDYGGQGGGGGHGGGGGGGSGYGNGGYGSGFGEGYG 167
Query: 151 --------YNRGGYGGGLYGSSGGMYGGGM-----YGGGLGGPMGGYGGGMGVGPYGG 195
GG GGG G+SG YG G YGGG GG GG GGG G G Y G
Sbjct: 168 SGGGVNGGGGSGGGGGGGGGASGYRYGAGYGKGYGYGGGPGGGGGGAGGGGGGGSYNG 225
Score = 65.1 bits (157), Expect = 3e-09
Identities = 45/99 (45%), Positives = 50/99 (50%), Gaps = 6/99 (6%)
Query: 119 LYGSSYGGGLGG-GLGGGLGGGMYGGGM-YGSSM-YNRGGYGGGLYGSSGGMYGGGMYGG 175
++ + GGG GG G GGG G G YG G YGS + GG GGG YG GG GG G
Sbjct: 27 IFVTGIGGGGGGYGSGGGEGNGGYGHGYGYGSGYGFGNGGVGGGGYGGGGGYGSGGGEGN 86
Query: 176 GLGGPMGGYG---GGMGVGPYGGDQDPNNPYGGPPSPPG 211
G G GYG GG G G YGG + GG S G
Sbjct: 87 GAYGQGYGYGSGNGGGGGGGYGGGGGGSYGSGGMGSGYG 125
Score = 47.4 bits (111), Expect = 6e-04
Identities = 26/61 (42%), Positives = 27/61 (43%), Gaps = 10/61 (16%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGG 158
Y GYG Y G G G GGG GGG GGG Y GG G + GYGG
Sbjct: 191 YRYGAGYGKGYGYGGGPGGG----------GGGAGGGGGGGSYNGGTGGYGEGHGSGYGG 240
Query: 159 G 159
G
Sbjct: 241 G 241
Score = 46.6 bits (109), Expect = 0.001
Identities = 29/65 (44%), Positives = 31/65 (47%), Gaps = 18/65 (27%)
Query: 100 GGAGGYGSTMNYNSGYGSGL-YGSSYGGGLGGGLGGGLGGGMY----------------G 142
GG GG S Y +GYG G YG GGG GGG GGG GGG Y G
Sbjct: 182 GGGGGGASGYRYGAGYGKGYGYGGGPGGG-GGGAGGGGGGGSYNGGTGGYGEGHGSGYGG 240
Query: 143 GGMYG 147
GG +G
Sbjct: 241 GGGHG 245
>dbj|BAA33812.1| RBP56/hTAFII68 [Homo sapiens] gi|27501920|gb|AAO13485.1| TAF15 RNA
polymerase II, TATA box binding protein (TBP)-associated
factor, 68kDa [Homo sapiens] gi|4507353|ref|NP_003478.1|
TBP-associated factor 15 isoform 2 [Homo sapiens]
gi|1628403|emb|CAA67398.1| hTAFII68 [Homo sapiens]
Length = 589
Score = 93.6 bits (231), Expect = 7e-18
Identities = 61/129 (47%), Positives = 65/129 (50%), Gaps = 19/129 (14%)
Query: 99 YGG---AGGYGSTMNYNSGY-----GSGLYGSSYGGGLGGGLGGGLGG---GMYGGGMYG 147
YGG GGYG + GY G G G GGG GG GGG GG G YGG G
Sbjct: 415 YGGDRSGGGYGGDRSSGGGYSGDRSGGGYGGDRSGGGYGGDRGGGYGGDRGGGYGGDRGG 474
Query: 148 SSMYNRGGYGG---GLYGSSGGMYGG--GMYGG---GLGGPMGGYGGGMGVGPYGGDQDP 199
+RGGYGG G YG G YGG G YGG G GG GGYGG G YGGD+
Sbjct: 475 GYGGDRGGYGGDRGGGYGGDRGGYGGDRGGYGGDRGGYGGDRGGYGGDRSRGGYGGDRGG 534
Query: 200 NNPYGGPPS 208
+ YGG S
Sbjct: 535 GSGYGGDRS 543
Score = 75.9 bits (185), Expect = 2e-12
Identities = 57/124 (45%), Positives = 58/124 (45%), Gaps = 22/124 (17%)
Query: 99 YGG--AGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGG------GLGG--GMYGG--GMY 146
YGG GGYG G G YG GGG GG GG G GG G YGG G Y
Sbjct: 460 YGGDRGGGYGGDRGGGYGGDRGGYGGDRGGGYGGDRGGYGGDRGGYGGDRGGYGGDRGGY 519
Query: 147 GSSMYNRGGYGGGLYGSSG------GMYGGGMYGGGLGGPM-GGYGGGMGVGPYGGDQDP 199
G +RGGYGG G SG G YGG GGG GG GGYGG G YGG
Sbjct: 520 GGDR-SRGGYGGDRGGGSGYGGDRSGGYGGDRSGGGYGGDRGGGYGGDR--GGYGGKMGG 576
Query: 200 NNPY 203
N Y
Sbjct: 577 RNDY 580
Score = 74.7 bits (182), Expect = 3e-12
Identities = 58/137 (42%), Positives = 60/137 (43%), Gaps = 36/137 (26%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGG--GMYGGGMYGSSMYNRGGYG 157
GG GG S Y G G YG GGG GG GGG GG G YGG G +RGGYG
Sbjct: 441 GGYGGDRSGGGYGGDRGGG-YGGDRGGGYGGDRGGGYGGDRGGYGGDRGGGYGGDRGGYG 499
Query: 158 G---------GLYGSSGGMYGG----GMYGGGLGGPMG----------------GYGGGM 188
G G YG G YGG G YGG GG G GYGG
Sbjct: 500 GDRGGYGGDRGGYGGDRGGYGGDRSRGGYGGDRGGGSGYGGDRSGGYGGDRSGGGYGGDR 559
Query: 189 GVGPYGGDQDPNNPYGG 205
G G YGGD+ YGG
Sbjct: 560 G-GGYGGDR---GGYGG 572
Score = 67.4 bits (163), Expect = 6e-10
Identities = 49/110 (44%), Positives = 51/110 (45%), Gaps = 22/110 (20%)
Query: 114 GYGS--GLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMY----NRGGY-----GGGLYG 162
GYG G G GG GG GG GG YGG Y + GGY GGG G
Sbjct: 395 GYGGERGYRGRGGRGGDRGGYGGDRSGGGYGGDRSSGGGYSGDRSGGGYGGDRSGGGYGG 454
Query: 163 SSGGMYG-------GGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
GG YG GG GGG GG GGYGG G G YGGD+ YGG
Sbjct: 455 DRGGGYGGDRGGGYGGDRGGGYGGDRGGYGGDRG-GGYGGDR---GGYGG 500
Score = 43.5 bits (101), Expect = 0.009
Identities = 24/57 (42%), Positives = 27/57 (47%), Gaps = 8/57 (14%)
Query: 164 SGGMYGGGMYGG--------GLGGPMGGYGGGMGVGPYGGDQDPNNPYGGPPSPPGF 212
SGG + G YGG G GG GGYGG G YGGD+ Y G S G+
Sbjct: 387 SGGDFRGRGYGGERGYRGRGGRGGDRGGYGGDRSGGGYGGDRSSGGGYSGDRSGGGY 443
Score = 35.4 bits (80), Expect = 2.3
Identities = 29/72 (40%), Positives = 31/72 (42%), Gaps = 8/72 (11%)
Query: 142 GGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGV-GPYGGDQDPN 200
GG G GY G G GG GG YGG G GGYGG G Y GD+
Sbjct: 388 GGDFRGRGYGGERGYRGR--GGRGGDRGG--YGGDRSG--GGYGGDRSSGGGYSGDRS-G 440
Query: 201 NPYGGPPSPPGF 212
YGG S G+
Sbjct: 441 GGYGGDRSGGGY 452
>dbj|BAA33811.1| RBP56/hTAFII68 [Homo sapiens] gi|21327701|ref|NP_631961.1|
TBP-associated factor 15 isoform 1 [Homo sapiens]
gi|8928305|sp|Q92804|RBP56_HUMAN TATA-binding protein
associated factor 2N (RNA-binding protein 56) (TAFII68)
(TAF(II)68) gi|1613775|gb|AAC50932.1| putative RNA
binding protein RBP56
Length = 592
Score = 93.6 bits (231), Expect = 7e-18
Identities = 61/129 (47%), Positives = 65/129 (50%), Gaps = 19/129 (14%)
Query: 99 YGG---AGGYGSTMNYNSGY-----GSGLYGSSYGGGLGGGLGGGLGG---GMYGGGMYG 147
YGG GGYG + GY G G G GGG GG GGG GG G YGG G
Sbjct: 418 YGGDRSGGGYGGDRSSGGGYSGDRSGGGYGGDRSGGGYGGDRGGGYGGDRGGGYGGDRGG 477
Query: 148 SSMYNRGGYGG---GLYGSSGGMYGG--GMYGG---GLGGPMGGYGGGMGVGPYGGDQDP 199
+RGGYGG G YG G YGG G YGG G GG GGYGG G YGGD+
Sbjct: 478 GYGGDRGGYGGDRGGGYGGDRGGYGGDRGGYGGDRGGYGGDRGGYGGDRSRGGYGGDRGG 537
Query: 200 NNPYGGPPS 208
+ YGG S
Sbjct: 538 GSGYGGDRS 546
Score = 75.9 bits (185), Expect = 2e-12
Identities = 57/124 (45%), Positives = 58/124 (45%), Gaps = 22/124 (17%)
Query: 99 YGG--AGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGG------GLGG--GMYGG--GMY 146
YGG GGYG G G YG GGG GG GG G GG G YGG G Y
Sbjct: 463 YGGDRGGGYGGDRGGGYGGDRGGYGGDRGGGYGGDRGGYGGDRGGYGGDRGGYGGDRGGY 522
Query: 147 GSSMYNRGGYGGGLYGSSG------GMYGGGMYGGGLGGPM-GGYGGGMGVGPYGGDQDP 199
G +RGGYGG G SG G YGG GGG GG GGYGG G YGG
Sbjct: 523 GGDR-SRGGYGGDRGGGSGYGGDRSGGYGGDRSGGGYGGDRGGGYGGDR--GGYGGKMGG 579
Query: 200 NNPY 203
N Y
Sbjct: 580 RNDY 583
Score = 74.7 bits (182), Expect = 3e-12
Identities = 58/137 (42%), Positives = 60/137 (43%), Gaps = 36/137 (26%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGG--GMYGGGMYGSSMYNRGGYG 157
GG GG S Y G G YG GGG GG GGG GG G YGG G +RGGYG
Sbjct: 444 GGYGGDRSGGGYGGDRGGG-YGGDRGGGYGGDRGGGYGGDRGGYGGDRGGGYGGDRGGYG 502
Query: 158 G---------GLYGSSGGMYGG----GMYGGGLGGPMG----------------GYGGGM 188
G G YG G YGG G YGG GG G GYGG
Sbjct: 503 GDRGGYGGDRGGYGGDRGGYGGDRSRGGYGGDRGGGSGYGGDRSGGYGGDRSGGGYGGDR 562
Query: 189 GVGPYGGDQDPNNPYGG 205
G G YGGD+ YGG
Sbjct: 563 G-GGYGGDR---GGYGG 575
Score = 67.4 bits (163), Expect = 6e-10
Identities = 49/110 (44%), Positives = 51/110 (45%), Gaps = 22/110 (20%)
Query: 114 GYGS--GLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMY----NRGGY-----GGGLYG 162
GYG G G GG GG GG GG YGG Y + GGY GGG G
Sbjct: 398 GYGGERGYRGRGGRGGDRGGYGGDRSGGGYGGDRSSGGGYSGDRSGGGYGGDRSGGGYGG 457
Query: 163 SSGGMYG-------GGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
GG YG GG GGG GG GGYGG G G YGGD+ YGG
Sbjct: 458 DRGGGYGGDRGGGYGGDRGGGYGGDRGGYGGDRG-GGYGGDR---GGYGG 503
Score = 43.5 bits (101), Expect = 0.009
Identities = 24/57 (42%), Positives = 27/57 (47%), Gaps = 8/57 (14%)
Query: 164 SGGMYGGGMYGG--------GLGGPMGGYGGGMGVGPYGGDQDPNNPYGGPPSPPGF 212
SGG + G YGG G GG GGYGG G YGGD+ Y G S G+
Sbjct: 390 SGGDFRGRGYGGERGYRGRGGRGGDRGGYGGDRSGGGYGGDRSSGGGYSGDRSGGGY 446
Score = 35.4 bits (80), Expect = 2.3
Identities = 29/72 (40%), Positives = 31/72 (42%), Gaps = 8/72 (11%)
Query: 142 GGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGV-GPYGGDQDPN 200
GG G GY G G GG GG YGG G GGYGG G Y GD+
Sbjct: 391 GGDFRGRGYGGERGYRGR--GGRGGDRGG--YGGDRSG--GGYGGDRSSGGGYSGDRS-G 443
Query: 201 NPYGGPPSPPGF 212
YGG S G+
Sbjct: 444 GGYGGDRSGGGY 455
>gb|AAH46099.1| TAF15 protein [Homo sapiens]
Length = 501
Score = 93.6 bits (231), Expect = 7e-18
Identities = 61/129 (47%), Positives = 65/129 (50%), Gaps = 19/129 (14%)
Query: 99 YGG---AGGYGSTMNYNSGY-----GSGLYGSSYGGGLGGGLGGGLGG---GMYGGGMYG 147
YGG GGYG + GY G G G GGG GG GGG GG G YGG G
Sbjct: 327 YGGDRSGGGYGGDRSSGGGYSGDRSGGGYGGDRSGGGYGGDRGGGYGGDRGGGYGGDRGG 386
Query: 148 SSMYNRGGYGG---GLYGSSGGMYGG--GMYGG---GLGGPMGGYGGGMGVGPYGGDQDP 199
+RGGYGG G YG G YGG G YGG G GG GGYGG G YGGD+
Sbjct: 387 GYGGDRGGYGGDRGGGYGGDRGGYGGDRGGYGGDRGGYGGDRGGYGGDRSRGGYGGDRGG 446
Query: 200 NNPYGGPPS 208
+ YGG S
Sbjct: 447 GSGYGGDRS 455
Score = 75.9 bits (185), Expect = 2e-12
Identities = 57/124 (45%), Positives = 58/124 (45%), Gaps = 22/124 (17%)
Query: 99 YGG--AGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGG------GLGG--GMYGG--GMY 146
YGG GGYG G G YG GGG GG GG G GG G YGG G Y
Sbjct: 372 YGGDRGGGYGGDRGGGYGGDRGGYGGDRGGGYGGDRGGYGGDRGGYGGDRGGYGGDRGGY 431
Query: 147 GSSMYNRGGYGGGLYGSSG------GMYGGGMYGGGLGGPM-GGYGGGMGVGPYGGDQDP 199
G +RGGYGG G SG G YGG GGG GG GGYGG G YGG
Sbjct: 432 GGDR-SRGGYGGDRGGGSGYGGDRSGGYGGDRSGGGYGGDRGGGYGGDR--GGYGGKMGG 488
Query: 200 NNPY 203
N Y
Sbjct: 489 RNDY 492
Score = 74.7 bits (182), Expect = 3e-12
Identities = 58/137 (42%), Positives = 60/137 (43%), Gaps = 36/137 (26%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGG--GMYGGGMYGSSMYNRGGYG 157
GG GG S Y G G YG GGG GG GGG GG G YGG G +RGGYG
Sbjct: 353 GGYGGDRSGGGYGGDRGGG-YGGDRGGGYGGDRGGGYGGDRGGYGGDRGGGYGGDRGGYG 411
Query: 158 G---------GLYGSSGGMYGG----GMYGGGLGGPMG----------------GYGGGM 188
G G YG G YGG G YGG GG G GYGG
Sbjct: 412 GDRGGYGGDRGGYGGDRGGYGGDRSRGGYGGDRGGGSGYGGDRSGGYGGDRSGGGYGGDR 471
Query: 189 GVGPYGGDQDPNNPYGG 205
G G YGGD+ YGG
Sbjct: 472 G-GGYGGDR---GGYGG 484
Score = 67.4 bits (163), Expect = 6e-10
Identities = 49/110 (44%), Positives = 51/110 (45%), Gaps = 22/110 (20%)
Query: 114 GYGS--GLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMY----NRGGY-----GGGLYG 162
GYG G G GG GG GG GG YGG Y + GGY GGG G
Sbjct: 307 GYGGERGYRGRGGRGGDRGGYGGDRSGGGYGGDRSSGGGYSGDRSGGGYGGDRSGGGYGG 366
Query: 163 SSGGMYG-------GGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYGG 205
GG YG GG GGG GG GGYGG G G YGGD+ YGG
Sbjct: 367 DRGGGYGGDRGGGYGGDRGGGYGGDRGGYGGDRG-GGYGGDR---GGYGG 412
Score = 43.5 bits (101), Expect = 0.009
Identities = 24/57 (42%), Positives = 27/57 (47%), Gaps = 8/57 (14%)
Query: 164 SGGMYGGGMYGG--------GLGGPMGGYGGGMGVGPYGGDQDPNNPYGGPPSPPGF 212
SGG + G YGG G GG GGYGG G YGGD+ Y G S G+
Sbjct: 299 SGGDFRGRGYGGERGYRGRGGRGGDRGGYGGDRSGGGYGGDRSSGGGYSGDRSGGGY 355
Score = 35.4 bits (80), Expect = 2.3
Identities = 29/72 (40%), Positives = 31/72 (42%), Gaps = 8/72 (11%)
Query: 142 GGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGV-GPYGGDQDPN 200
GG G GY G G GG GG YGG G GGYGG G Y GD+
Sbjct: 300 GGDFRGRGYGGERGYRGR--GGRGGDRGG--YGGDRSG--GGYGGDRSSGGGYSGDRS-G 352
Query: 201 NPYGGPPSPPGF 212
YGG S G+
Sbjct: 353 GGYGGDRSGGGY 364
>ref|XP_428850.1| PREDICTED: similar to type II keratin K6h [Gallus gallus]
Length = 1659
Score = 92.8 bits (229), Expect = 1e-17
Identities = 60/129 (46%), Positives = 67/129 (51%), Gaps = 25/129 (19%)
Query: 92 PWEQNTT-----YGGAG---GYGSTMNYNSGYGS-----GLYGSSYGGGLGGGLGGGLGG 138
PW+ + YGG G G+GS +N G G Y GG GG GGG GG
Sbjct: 531 PWDHDMLQWPRGYGGGGRCGGFGSRSLHNIGSSGRIAMGGCYSGGGYGGRMGGFGGGYGG 590
Query: 139 -GMYGGGMYGSSMYNRGGYGGGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGGDQ 197
G YGGGM G M GG+GGG+ G GGGM GG GG MGGYGGGMG G GG
Sbjct: 591 LGSYGGGMGGGGM---GGFGGGMGG------GGGM--GGFGGGMGGYGGGMGGGGMGGYG 639
Query: 198 DPNNPYGGP 206
P +GGP
Sbjct: 640 GPGFGFGGP 648
Score = 84.7 bits (208), Expect = 3e-15
Identities = 51/95 (53%), Positives = 56/95 (58%), Gaps = 14/95 (14%)
Query: 99 YGGAGGYGSTMNYNSGYGSGLYG-SSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYG 157
Y G GGYG M G+G G G SYGGG+GGG GG GGGM GGG GG+G
Sbjct: 572 YSG-GGYGGRMG---GFGGGYGGLGSYGGGMGGGGMGGFGGGMGGGG-------GMGGFG 620
Query: 158 GGLYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGP 192
GG+ G GGM GGGM GG GGP G+GG GP
Sbjct: 621 GGMGGYGGGMGGGGM--GGYGGPGFGFGGPGRGGP 653
Score = 59.3 bits (142), Expect = 2e-07
Identities = 50/127 (39%), Positives = 56/127 (43%), Gaps = 33/127 (25%)
Query: 104 GYGSTMNYNSGYGSGLYGSSYGGGLGGGLGG------------GLGGGMYGGGMYGSSMY 151
G GS Y+S S+ GGG GG G G GGG GG S++
Sbjct: 1314 GGGSRRGYSSC-------SAVGGGFGGSGGRSRISYSSFSTSRGTGGGGRCGGFSSRSLH 1366
Query: 152 NRGGYGG-GLYGSSGGMYG------GGMYGGGLGGPMGGYGGGMGVGPYGGDQDPNNPYG 204
N GG G + GS GG YG GG YGGG G GG GG GVG +GG P
Sbjct: 1367 NMGGSGRISMGGSYGGGYGCRIVGLGGGYGGGFGSIGGGVIGG-GVGSFGG------PVR 1419
Query: 205 GPPSPPG 211
G P PG
Sbjct: 1420 GGPGFPG 1426
Score = 52.4 bits (124), Expect = 2e-05
Identities = 45/119 (37%), Positives = 58/119 (47%), Gaps = 24/119 (20%)
Query: 99 YGGAGGYG--STMNYNSGYGSGLYGSSYGGGLGGGLG-------GGLG----GGMYGGGM 145
+GG+GG S ++++ G+G GGG GG GG G GG YGGG
Sbjct: 1331 FGGSGGRSRISYSSFSTSRGTG------GGGRCGGFSSRSLHNMGGSGRISMGGSYGGG- 1383
Query: 146 YGSSMYNRGGYGGGLYGS-SGGMYGGGMYGGGLGGPM-GGYGGGMGVGPYGGDQDPNNP 202
YG + GG GG +GS GG+ GGG+ G GGP+ GG G G+ P D P
Sbjct: 1384 YGCRIVGLGGGYGGGFGSIGGGVIGGGV--GSFGGPVRGGPGFPGGIQPVQVDSSLLRP 1440
Score = 48.9 bits (115), Expect = 2e-04
Identities = 36/96 (37%), Positives = 46/96 (47%), Gaps = 12/96 (12%)
Query: 100 GGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRGGYGGG 159
G AGG+ S Y+ G G ++ GG GG+ G G G G +G + N G+GGG
Sbjct: 232 GNAGGFSSHSLYSLG---GCKSTALGGFGAGGICRGFGAGGQG---FGYGVGNGIGFGGG 285
Query: 160 LYGSSGGMYGGGMYGGGLGGPMGGYGGGMGVGPYGG 195
+G +GGG GG GG G GG P GG
Sbjct: 286 ----AGCGFGGGF--GGFGGGFDGGVGGFAQCPPGG 315
Score = 43.9 bits (102), Expect = 0.007
Identities = 29/72 (40%), Positives = 35/72 (48%), Gaps = 13/72 (18%)
Query: 95 QNTTYGGAGGYGSTMNYNSGYGSGLYGSSYGGGLGGGLGGGLGGGMYGGGMYGSSMYNRG 154
++T GG G G G+G+G G YG G G G GGG G +GGG G
Sbjct: 249 KSTALGGFGAGGICR----GFGAGGQGFGYGVGNGIGFGGG-AGCGFGGGF--------G 295
Query: 155 GYGGGLYGSSGG 166
G+GGG G GG
Sbjct: 296 GFGGGFDGGVGG 307
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.312 0.139 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 772,363,391
Number of Sequences: 2540612
Number of extensions: 47566425
Number of successful extensions: 761469
Number of sequences better than 10.0: 10694
Number of HSP's better than 10.0 without gapping: 5185
Number of HSP's successfully gapped in prelim test: 5965
Number of HSP's that attempted gapping in prelim test: 263464
Number of HSP's gapped (non-prelim): 111577
length of query: 329
length of database: 863,360,394
effective HSP length: 128
effective length of query: 201
effective length of database: 538,162,058
effective search space: 108170573658
effective search space used: 108170573658
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0106.12


