

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0101.12
(310 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
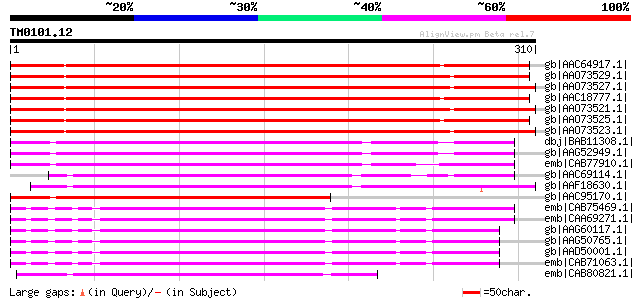
Score E
Sequences producing significant alignments: (bits) Value
gb|AAC64917.1| gag-pol polyprotein [Glycine max] 359 5e-98
gb|AAO73529.1| gag-pol polyprotein [Glycine max] 356 5e-97
gb|AAO73527.1| gag-pol polyprotein [Glycine max] 353 5e-96
gb|AAC18777.1| gag-protease polyprotein [Glycine max] gi|7488678... 352 6e-96
gb|AAO73521.1| gag-pol polyprotein [Glycine max] 352 8e-96
gb|AAO73525.1| gag-pol polyprotein [Glycine max] 348 9e-95
gb|AAO73523.1| gag-pol polyprotein [Glycine max] 348 9e-95
dbj|BAB11308.1| copia-like retroelement pol polyprotein [Arabido... 191 2e-47
gb|AAG52949.1| gag/pol polyprotein [Arabidopsis thaliana] 191 2e-47
emb|CAB77910.1| putative transposon protein [Arabidopsis thalian... 180 4e-44
gb|AAC69114.1| putative gag-protease polyprotein [Arabidopsis th... 163 7e-39
gb|AAF18630.1| F5J5.1 [Arabidopsis thaliana] gi|25403474|pir||C8... 151 2e-35
gb|AAC95170.1| copia-like retroelement pol polyprotein [Arabidop... 137 3e-31
emb|CAB75469.1| copia-type reverse transcriptase-like protein [A... 123 8e-27
emb|CAA69271.1| lectin receptor kinase [Arabidopsis thaliana] 123 8e-27
gb|AAG60117.1| copia-type polyprotein, putative [Arabidopsis tha... 122 1e-26
gb|AAG50765.1| copia-type polyprotein, putative [Arabidopsis tha... 122 1e-26
gb|AAD50001.1| Hypothetical protein [Arabidopsis thaliana] gi|25... 122 1e-26
emb|CAB71063.1| copia-type polyprotein [Arabidopsis thaliana] gi... 120 4e-26
emb|CAB80821.1| putative transposon protein [Arabidopsis thalian... 120 5e-26
>gb|AAC64917.1| gag-pol polyprotein [Glycine max]
Length = 1550
Score = 359 bits (922), Expect = 5e-98
Identities = 179/309 (57%), Positives = 242/309 (77%), Gaps = 5/309 (1%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M+AFLKS+D +TWKAV+KGW+HP EG T E KPE++ T E+E +L NSKA+NA
Sbjct: 1 MVAFLKSLDSRTWKAVIKGWEHPKMLDTEGKP-TNELKPEEDWTKEEDELALGNSKALNA 59
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
+FN VDKN+FRLINTCTVAKDAWEILKTTHEGT++V+MSRLQ+L T+FENL M E+E I
Sbjct: 60 LFNGVDKNIFRLINTCTVAKDAWEILKTTHEGTSKVKMSRLQLLATKFENLKMKEEECIH 119
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
E+HM + +++NA ALGE M+DEKLVRKILRS+ +F MKV AIEEAQDI +++VDELIG
Sbjct: 120 EFHMNILEIANACTALGERMTDEKLVRKILRSLPKRFDMKVTAIEEAQDICNMRVDELIG 179
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTT-EGDDADSES-ENLSEALALLARKFNRALRKID 238
SLQT++L L ++ EKK+K++AFVSN E D+ D ++ E L+ A+ LL ++FN+ L ++D
Sbjct: 180 SLQTFELGLSDRTEKKSKNLAFVSNDEGEEDEYDLDTDEGLTNAVVLLGKQFNKVLNRMD 239
Query: 239 RRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECATYLKKQKK 298
RR +P+V++I D K S+ + + ++K +KG+QCH CEGYGHI++EC T+LKKQ+K
Sbjct: 240 RRQKPHVRNIPFDIRK--GSEYQKRSDEKPSHSKGIQCHGCEGYGHIKAECPTHLKKQRK 297
Query: 299 GMVVTWSDE 307
G+ V SD+
Sbjct: 298 GLSVCRSDD 306
>gb|AAO73529.1| gag-pol polyprotein [Glycine max]
Length = 1577
Score = 356 bits (913), Expect = 5e-97
Identities = 178/309 (57%), Positives = 239/309 (76%), Gaps = 5/309 (1%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M+AFLKS+D +TWKAV+KGW+HP EG T E KPE++ T E+E +L NSKA+NA
Sbjct: 28 MVAFLKSLDSRTWKAVIKGWEHPKMLDTEGKP-TNELKPEEDWTKEEDELALGNSKALNA 86
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
+FN VDKN+FRLINTCTVAKDAWEILKTTHEGT++V+MSRLQ+L T+FENL M E+E I
Sbjct: 87 LFNGVDKNIFRLINTCTVAKDAWEILKTTHEGTSKVKMSRLQLLATKFENLKMKEEECIH 146
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
++HM + +++NA ALGE M+DEKLVRKILRS+ +F MKV AIEEAQDI +++VDELIG
Sbjct: 147 DFHMTILEIANACTALGERMTDEKLVRKILRSLPKRFDMKVTAIEEAQDICNMRVDELIG 206
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTT-EGDDADSES-ENLSEALALLARKFNRALRKID 238
SLQT++L L ++ EKK+K++AFVSN E D+ D ++ E L+ A+ L ++FN+ L ++D
Sbjct: 207 SLQTFELGLSDRTEKKSKNLAFVSNDEGEEDEYDLDTDEGLTNAVVFLGKQFNKVLNRMD 266
Query: 239 RRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECATYLKKQKK 298
RR +P+V++I D K QR K ++K +KG+QC CEGYGHI++EC T+LKKQ+K
Sbjct: 267 RRQKPHVRNISLDIRKGSEYQR--KSDEKPSHSKGIQCRGCEGYGHIKAECPTHLKKQRK 324
Query: 299 GMVVTWSDE 307
G+ V SD+
Sbjct: 325 GLSVCRSDD 333
>gb|AAO73527.1| gag-pol polyprotein [Glycine max]
Length = 1576
Score = 353 bits (905), Expect = 5e-96
Identities = 177/312 (56%), Positives = 239/312 (75%), Gaps = 5/312 (1%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M+AFLKS+D +TWKAV+KGW+HP EG T E KPE++ T E+E +L NSKA+NA
Sbjct: 28 MVAFLKSLDSRTWKAVIKGWEHPKMLDTEGKP-TDELKPEEDWTKEEDELALGNSKALNA 86
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
+FN VDKN+FRLINTCTVAKDAWEILK THEGT++V+MSRLQ+L T+FENL M E+E I
Sbjct: 87 LFNGVDKNIFRLINTCTVAKDAWEILKITHEGTSKVKMSRLQLLATKFENLKMKEEECIH 146
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
++HM + +++NA ALGE ++DEKLVRKILRS+ +F MKV AIEEAQDI +++VDELIG
Sbjct: 147 DFHMNILEIANACTALGERITDEKLVRKILRSLPKRFDMKVTAIEEAQDICNMRVDELIG 206
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTT-EGDDADSES-ENLSEALALLARKFNRALRKID 238
SLQT++L L ++ EKK+K++AFVSN E D+ D ++ E L+ A+ LL ++FN+ L ++D
Sbjct: 207 SLQTFELGLSDRAEKKSKNLAFVSNDEGEEDEYDLDTDEGLTNAVVLLGKQFNKVLNRMD 266
Query: 239 RRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECATYLKKQKK 298
+R +P+VQ+I D K Q+ + + K +KG+QCH CEGYGHI +EC T+LKK +K
Sbjct: 267 KRQKPHVQNIPFDIRKGSKYQK--RSDVKPSHSKGIQCHGCEGYGHIIAECPTHLKKHRK 324
Query: 299 GMVVTWSDEDSE 310
G+ V SD +SE
Sbjct: 325 GLSVCQSDTESE 336
>gb|AAC18777.1| gag-protease polyprotein [Glycine max] gi|7488678|pir||T06419
gag-proteinase polyprotein - soybean retrovirus-like
element
Length = 640
Score = 352 bits (904), Expect = 6e-96
Identities = 177/309 (57%), Positives = 239/309 (77%), Gaps = 5/309 (1%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M+AFLKS+D +TWKAV+K W+HP EG T KPE++ T E+E +L NSKA+NA
Sbjct: 28 MVAFLKSLDSRTWKAVIKDWEHPKMLDTEGKP-TDGLKPEEDWTKEEDELALGNSKALNA 86
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
+FN VDKN+FRLINTCTVAKDAWEILKTTHEGT++V+MSRLQ+L T+FENL M E+E I
Sbjct: 87 LFNGVDKNIFRLINTCTVAKDAWEILKTTHEGTSKVKMSRLQLLATKFENLKMKEEECIH 146
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
++HM + +++NA ALGE M+DEKLVRKILRS+ +F MKV AIEEAQDI +L+VDELIG
Sbjct: 147 DFHMNILEIANACTALGERMTDEKLVRKILRSLPKRFDMKVTAIEEAQDICNLRVDELIG 206
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTT-EGDDADSES-ENLSEALALLARKFNRALRKID 238
SLQT++L L ++ EKK+K++AFVSN E D+ D ++ E L+ A+ LL ++FN+ L ++D
Sbjct: 207 SLQTFELGLSDRTEKKSKNLAFVSNDEGEEDEYDLDTDEGLTNAVVLLGKQFNKVLNRMD 266
Query: 239 RRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECATYLKKQKK 298
RR +P+V++I D K S+ + + ++K +KG QCH CEGYGHI++EC T+LKKQ+K
Sbjct: 267 RRQKPHVRNIPFDIRK--GSEYQKRSDEKPSHSKGFQCHGCEGYGHIKAECPTHLKKQRK 324
Query: 299 GMVVTWSDE 307
G+ V SD+
Sbjct: 325 GLSVCRSDD 333
>gb|AAO73521.1| gag-pol polyprotein [Glycine max]
Length = 1574
Score = 352 bits (903), Expect = 8e-96
Identities = 177/312 (56%), Positives = 238/312 (75%), Gaps = 5/312 (1%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M+AFLKS+D +TWKAV+KGW+HP EG T E KPE++ T E+E +L NSKA+NA
Sbjct: 28 MVAFLKSLDSRTWKAVIKGWEHPKMLDTEGKP-TDELKPEEDWTKEEDELALGNSKALNA 86
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
+FN VDKN+FRLINTCTVAKDAWEILK THEGT++V++SRLQ+L T+FENL M E+E I
Sbjct: 87 LFNGVDKNIFRLINTCTVAKDAWEILKITHEGTSKVKISRLQLLATKFENLKMKEEECIH 146
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
++HM + +++NA ALGE ++DEKLVRKILRS+ +F MKV AIEEAQDI +++VDELIG
Sbjct: 147 DFHMNILEIANACTALGERITDEKLVRKILRSLPKRFDMKVTAIEEAQDICNMRVDELIG 206
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTT-EGDDAD-SESENLSEALALLARKFNRALRKID 238
SLQT++L L ++ EKK+K++AFVSN E D+ D + E L+ A+ LL ++FN+ L ++D
Sbjct: 207 SLQTFELGLSDRAEKKSKNLAFVSNDEGEEDEYDLNTDEGLTNAVVLLGKQFNKVLNRMD 266
Query: 239 RRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECATYLKKQKK 298
+R +P+VQ+I D K Q+ K + K +KG+QCH CEGYGHI +EC T+LKK +K
Sbjct: 267 KRQKPHVQNIPFDIRKGSKYQK--KSDVKPSHSKGIQCHGCEGYGHIIAECPTHLKKHRK 324
Query: 299 GMVVTWSDEDSE 310
G+ V SD +SE
Sbjct: 325 GLSVCQSDTESE 336
>gb|AAO73525.1| gag-pol polyprotein [Glycine max]
Length = 1576
Score = 348 bits (894), Expect = 9e-95
Identities = 178/310 (57%), Positives = 240/310 (77%), Gaps = 6/310 (1%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M+AFLKS+D +TWKAV+KGW+HP EG T E KPE++ T E+E +L NSKA+NA
Sbjct: 28 MVAFLKSLDSRTWKAVIKGWEHPKMLDTEGKP-TNELKPEEDWTKEEDELALGNSKALNA 86
Query: 61 IFNAVDKNMFRLINTCTVAKDAW-EILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERI 119
+FN VDKN+FRLINTCTVAKDA EILKTTHEGT++V+MSRLQ+L T+FENL M E+E I
Sbjct: 87 LFNGVDKNIFRLINTCTVAKDACGEILKTTHEGTSKVKMSRLQLLATKFENLKMKEEECI 146
Query: 120 SEYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELI 179
++HM + +++NA ALGE M+DEKLVRKILRS+ +F MKV AIEEAQDI +++VDELI
Sbjct: 147 HDFHMNILEIANACTALGERMTDEKLVRKILRSLPKRFDMKVTAIEEAQDICNMRVDELI 206
Query: 180 GSLQTYKLKLGEKPEKKTKSIAFVSNTT-EGDDADSES-ENLSEALALLARKFNRALRKI 237
GSLQT++L L ++ EKK+K++AFVSN E D+ D ++ E L+ A+ LL ++FN+ L ++
Sbjct: 207 GSLQTFELGLSDRNEKKSKNLAFVSNDEGEEDEYDLDTDEGLTNAVGLLGKQFNKVLNRM 266
Query: 238 DRRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECATYLKKQK 297
DRR +P+V++I D K S+ K ++K +KG+QCH CEGYGHI++EC T+LKKQ+
Sbjct: 267 DRRQKPHVRNIPFDIRK--GSEYHKKSDEKPSHSKGIQCHGCEGYGHIKAECPTHLKKQR 324
Query: 298 KGMVVTWSDE 307
KG+ V SD+
Sbjct: 325 KGLSVCRSDD 334
>gb|AAO73523.1| gag-pol polyprotein [Glycine max]
Length = 1576
Score = 348 bits (894), Expect = 9e-95
Identities = 176/312 (56%), Positives = 239/312 (76%), Gaps = 5/312 (1%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M+AFLKS+D +TWKAV+KGW+HP EG T E KPE++ T E+E +L NSKA+NA
Sbjct: 28 MVAFLKSLDSRTWKAVIKGWEHPKMLDTEGKP-TDELKPEEDWTKEEDELALGNSKALNA 86
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
+FN VDKN+FRLINTCTVAKDA EILK+THEGT++V+MSRLQ+L T+FENL M E+E I
Sbjct: 87 LFNGVDKNIFRLINTCTVAKDACEILKSTHEGTSKVKMSRLQLLATKFENLKMKEEECIH 146
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
++HM + +++NA ALGE ++DEKLVRKILRS+ +F MKV AIEEAQDI +++VDELIG
Sbjct: 147 DFHMNILEIANACTALGERITDEKLVRKILRSLPKRFDMKVTAIEEAQDICNMRVDELIG 206
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTT-EGDDADSES-ENLSEALALLARKFNRALRKID 238
SLQT++L L ++ EKK+K++AFVSN E D+ D ++ E L+ A+ LL ++FN+ L ++D
Sbjct: 207 SLQTFELGLSDRAEKKSKNLAFVSNDEGEEDEYDLDTDEGLTNAVVLLGKQFNKVLNRMD 266
Query: 239 RRTRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECATYLKKQKK 298
+R +P+VQ+I D K Q+ + + K +KG+QCH CEGYGHI +EC T+LKK +K
Sbjct: 267 KRQKPHVQNIPFDIRKGSKYQK--RSDVKPSHSKGIQCHGCEGYGHIIAECPTHLKKHRK 324
Query: 299 GMVVTWSDEDSE 310
G+ V SD +SE
Sbjct: 325 GLSVCQSDTESE 336
>dbj|BAB11308.1| copia-like retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1013
Score = 191 bits (486), Expect = 2e-47
Identities = 97/298 (32%), Positives = 174/298 (57%), Gaps = 17/298 (5%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M A ++ + + W A GWK PV +G K ED+ T AEE + NS+A++
Sbjct: 28 MRALIRGLGKEAWIATSVGWKAPV---VKGENGEDVLKTEDQWTDAEEAKATANSRALSL 84
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
IFN+V++N F+ I C AK+AW+ L +EGT+ V+ SR+ ML ++FENLTM E E I
Sbjct: 85 IFNSVNQNQFKRIQNCESAKEAWDKLAKAYEGTSSVKRSRIDMLASQFENLTMDESENIE 144
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
E+ ++ +++ + LG+ D+KLV+K+LR + S+F K A+ + D ++ +E++G
Sbjct: 145 EFSGKISAIASEAHNLGKKYKDKKLVKKLLRCLPSRFESKRTAMGTSLDTDTIDFEEVVG 204
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRR 240
LQ Y+L++ +K +A ++ + +E + L ++++++A+ F+RA++++++R
Sbjct: 205 MLQAYELEITSGKGGYSKGVALAVSSEK-----NEIQELKDSMSMMAKNFSRAMKRVEKR 259
Query: 241 TRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECATYLKKQKK 298
Q D ++ D++ + +QCHEC+GYGHI++EC + +K K
Sbjct: 260 GFARNQGSDRDRD---------RDRDRNSKRSEIQCHECQGYGHIKAECPSLKRKDLK 308
>gb|AAG52949.1| gag/pol polyprotein [Arabidopsis thaliana]
Length = 1643
Score = 191 bits (486), Expect = 2e-47
Identities = 97/298 (32%), Positives = 174/298 (57%), Gaps = 17/298 (5%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M A ++ + + W A GWK PV +G K ED+ T AEE + NS+A++
Sbjct: 28 MRALIRGLGKEAWIATSVGWKAPV---VKGENGEDVLKTEDQWTDAEEAKATANSRALSL 84
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
IFN+V++N F+ I C AK+AW+ L +EGT+ V+ SR+ ML ++FENLTM E E I
Sbjct: 85 IFNSVNQNQFKRIQNCESAKEAWDKLAKAYEGTSSVKRSRIDMLASQFENLTMDESENIE 144
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
E+ ++ +++ + LG+ D+KLV+K+LR + S+F K A+ + D ++ +E++G
Sbjct: 145 EFSGKISAIASEAHNLGKKYKDKKLVKKLLRCLPSRFESKRTAMGTSLDTDTIDFEEVVG 204
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRR 240
LQ Y+L++ +K +A ++ + +E + L ++++++A+ F+RA++++++R
Sbjct: 205 MLQAYELEITSGKGGYSKGVALAVSSEK-----NEIQELKDSMSMMAKNFSRAMKRVEKR 259
Query: 241 TRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECATYLKKQKK 298
Q D ++ D++ + +QCHEC+GYGHI++EC + +K K
Sbjct: 260 GFARNQGSDRDRD---------RDRDRNSKRSEIQCHECQGYGHIKAECPSLKRKDLK 308
>emb|CAB77910.1| putative transposon protein [Arabidopsis thaliana]
gi|4773881|gb|AAD29754.1| putative transposon protein
[Arabidopsis thaliana] gi|25407270|pir||H85055 probable
transposon protein [imported] - Arabidopsis thaliana
Length = 1008
Score = 180 bits (457), Expect = 4e-44
Identities = 95/298 (31%), Positives = 170/298 (56%), Gaps = 21/298 (7%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M A ++ + + W A GWK PV +G K +D+ AEE + NS+A++
Sbjct: 28 MRALIRGLGKEAWIATSIGWKAPV---IKGEDGEDVLKTKDQWNDAEEAKAKANSRALSL 84
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
IFN V++N F+ I C AK+AW+ L +EGT+ V+ SR+ ML ++FENL+M E E I
Sbjct: 85 IFNFVNQNQFKRIQNCESAKEAWDKLAKAYEGTSSVKRSRIDMLASQFENLSMEETENIE 144
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
E+ ++ +++ + LG+ D+KLV+K+LR + S+F K A+ + D S+ +E++G
Sbjct: 145 EFSGKISAIASEAHNLGKKYKDKKLVKKLLRCLPSRFESKRTAMGTSLDTDSIDFEEVVG 204
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRR 240
LQ Y+L++ +K +A ++ + +E + L + ++++A+ F+RA+R+++++
Sbjct: 205 MLQAYELEITSGKGGYSKGLALAASAKK-----NEIQELKDTMSMMAKDFSRAMRRVEKK 259
Query: 241 TRPNVQDIKSDNSKSYNSQRKVKEEDKSGQNKGVQCHECEGYGHIRSECATYLKKQKK 298
+Q + D+S + +QCHEC+GYGHI++EC + +K K
Sbjct: 260 -------------GFGRNQGTDRYRDRSSKRDEIQCHECQGYGHIKAECPSLKRKDLK 304
>gb|AAC69114.1| putative gag-protease polyprotein [Arabidopsis thaliana]
gi|25411268|pir||B84482 probable gag-proteinase
polyprotein [imported] - Arabidopsis thaliana
Length = 627
Score = 163 bits (412), Expect = 7e-39
Identities = 91/275 (33%), Positives = 154/275 (55%), Gaps = 20/275 (7%)
Query: 24 VKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNAIFNAVDKNMFRLINTCTVAKDAW 83
++ Q +G T KP+ T+ E+ S N++AM AIFN VD++ F+LI C AK AW
Sbjct: 32 IRGQEDGFKIT---KPKANWTAEEKLQSKFNARAMKAIFNGVDEDEFKLIQGCKSAKQAW 88
Query: 84 EILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERISEYHMRVRDLSNASFALGEPMSDE 143
+ L+ +HEGT+ V+ +RL + T+FE L M DE+I ++ ++ L+N + +G+ D+
Sbjct: 89 DTLQKSHEGTSSVKRTRLDHIATQFEYLKMEPDEKIVKFSSKISALANEAEVMGKTYKDQ 148
Query: 144 KLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIGSLQTYKLKLGEKPEKKTKSIAFV 203
KLV+K+LR + KFA + A + + +L+G L+ ++K + K +K+IAF
Sbjct: 149 KLVKKLLRCLPPKFAAHKAVMRVAGNTDKISFVDLVGMLKLEEMKADQDKVKPSKNIAF- 207
Query: 204 SNTTEGDDADSESENLSEALALLARKFNRALRKIDRRTRPNVQDIKSDNSKSYNSQRKVK 263
D + + + + +ALLAR F +AL+++ +I + S+ S+
Sbjct: 208 ----NADQGSEQFQEIKDGMALLARNFGKALKRV---------EIDGERSRGRFSR---S 251
Query: 264 EEDKSGQNKGVQCHECEGYGHIRSECATYLKKQKK 298
E D + K +QC+EC G+GHI+ EC +K+ K
Sbjct: 252 ENDDLRKKKEIQCYECGGFGHIKPECPITKRKEMK 286
>gb|AAF18630.1| F5J5.1 [Arabidopsis thaliana] gi|25403474|pir||C86482 protein
F5J5.1 [imported] - Arabidopsis thaliana
Length = 1463
Score = 151 bits (382), Expect = 2e-35
Identities = 87/301 (28%), Positives = 157/301 (51%), Gaps = 11/301 (3%)
Query: 13 WKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNAIFNAVDKNMFRL 72
W AV +GW+ P +G T KP+ T+ E+ S N++AMNAI N +D++ F+L
Sbjct: 40 WTAVEEGWEPPFDLTEDGFKIT---KPKANWTAEEKLQSKFNARAMNAIVNGIDEDEFKL 96
Query: 73 INTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERISEYHMRVRDLSNA 132
I C AK AW+ L+ +HEGT+ V+ +RL + T+FE L M E I ++ ++ L+N
Sbjct: 97 IQGCKSAKQAWDTLQKSHEGTSSVKRTRLDHIATQFEYLKMEPYETIVKFSSKISALANE 156
Query: 133 SFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIGSLQTYKLKLGEK 192
+ LG+ D+KLV+K+LR + KF + A + + +L+G L++ +++ +
Sbjct: 157 AEVLGKTYKDQKLVKKLLRCLPPKFPAHKAVMRVAGNTDKISFVDLVGMLKSEEMEPDQD 216
Query: 193 PEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRRTRPNVQDIKSDN 252
K +K+IAF D + + + + +ALLAR F +AL++++R + + +
Sbjct: 217 KVKPSKNIAF-----NADQGSEQFQQIKDGMALLARNFGKALKRVERGQNRDSTSWSNKD 271
Query: 253 SKSYNSQRKVKEEDKSGQNKGVQCH---ECEGYGHIRSECATYLKKQKKGMVVTWSDEDS 309
++ + E D G+ K +QC+ E + G ++ V++ +D D
Sbjct: 272 GETSRGRFSRSENDDLGKKKEIQCYDDPESDDEGEELLNFVAFMASSDSSKVMSDTDSDC 331
Query: 310 E 310
+
Sbjct: 332 D 332
>gb|AAC95170.1| copia-like retroelement pol polyprotein [Arabidopsis thaliana]
gi|25411196|pir||B84473 copia-like retroelement pol
polyprotein [imported] - Arabidopsis thaliana
Length = 916
Score = 137 bits (346), Expect = 3e-31
Identities = 70/189 (37%), Positives = 115/189 (60%), Gaps = 3/189 (1%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M A ++ + + W A GWK PV +G K ED+ AEE + NS+A++
Sbjct: 40 MRALIRGLGKEAWIATSIGWKAPV---IKGEDGEDVLKTEDQWNDAEEAKATANSRALSL 96
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
IFN+V++N F+ I C AK+AW+ L +EGT+ V+ SR+ ML ++FENLTM E E I
Sbjct: 97 IFNSVNQNQFKQIQNCESAKEAWDKLAKAYEGTSSVKRSRIDMLASQFENLTMEETENIE 156
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
E+ ++ +++ + LG+ D+KLV+K+LR + S+F K A+ + D +S+ +E++G
Sbjct: 157 EFSGKISAIASEAHNLGKKYKDKKLVKKLLRCLPSRFESKRTAMGTSLDTNSIDFEEVVG 216
Query: 181 SLQTYKLKL 189
Q Y+L++
Sbjct: 217 MFQAYELEI 225
>emb|CAB75469.1| copia-type reverse transcriptase-like protein [Arabidopsis
thaliana] gi|11278363|pir||T49313 copia-type reverse
transcriptase-like protein - Arabidopsis thaliana
Length = 1272
Score = 123 bits (308), Expect = 8e-27
Identities = 95/300 (31%), Positives = 152/300 (50%), Gaps = 23/300 (7%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M A L + D+ W+ V KG+ P + EGS S ++ D L + + + KA+
Sbjct: 25 MKAILGAHDV--WEIVEKGFIEP---ENEGSLSQTQK---DGLRDSRKR----DKKALCL 72
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
I+ +D++ F + T AK+AWE L+T+++G +V+ RLQ L FE L M E E +S
Sbjct: 73 IYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVS 132
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
+Y RV ++N GE + D +++ K+LRS+ KF V IEE +D+ ++ +++L+G
Sbjct: 133 DYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLG 192
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRR 240
SLQ Y+ EK +KK + V N + + +S + R R R
Sbjct: 193 SLQAYE----EKKKKKEDIVEQVLNMQITKEENGQSYQRRGGGQVRGR--GRGGYGNGRG 246
Query: 241 TRPNVQDIKSDNSKSYNSQR-KVKEEDKSGQNK-GVQCHECEGYGHIRSECATYLKKQKK 298
RP+ + N + NS R + K KS +K V+C+ C +GH SEC K+ K
Sbjct: 247 WRPHED---NTNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKFK 303
>emb|CAA69271.1| lectin receptor kinase [Arabidopsis thaliana]
Length = 544
Score = 123 bits (308), Expect = 8e-27
Identities = 95/300 (31%), Positives = 152/300 (50%), Gaps = 23/300 (7%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M A L + D+ W+ V KG+ P + EGS S ++ D L + + + KA+
Sbjct: 41 MKAILGAHDV--WEIVEKGFIEP---ENEGSLSQTQK---DGLRDSRKR----DKKALCL 88
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
I+ +D++ F + T AK+AWE L+T+++G +V+ RLQ L FE L M E E +S
Sbjct: 89 IYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVS 148
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
+Y RV ++N GE + D +++ K+LRS+ KF V IEE +D+ ++ +++L+G
Sbjct: 149 DYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLG 208
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRR 240
SLQ Y+ EK +KK + V N + + +S + R R R
Sbjct: 209 SLQAYE----EKKKKKEDIVEQVLNMQITKEENGQSYQRRGGGQVRGR--GRGGYGNGRG 262
Query: 241 TRPNVQDIKSDNSKSYNSQR-KVKEEDKSGQNK-GVQCHECEGYGHIRSECATYLKKQKK 298
RP+ + N + NS R + K KS +K V+C+ C +GH SEC K+ K
Sbjct: 263 WRPHED---NTNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASECKAPSNKKFK 319
>gb|AAG60117.1| copia-type polyprotein, putative [Arabidopsis thaliana]
Length = 1352
Score = 122 bits (307), Expect = 1e-26
Identities = 94/291 (32%), Positives = 149/291 (50%), Gaps = 23/291 (7%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M A L + D+ W+ V KG+ P + EGS S ++ D L + + + KA+
Sbjct: 25 MKAILGAHDV--WEIVEKGFIEP---ENEGSLSQTQK---DGLRDSRKR----DKKALCL 72
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
I+ +D++ F + T AK+AWE L+T+++G +V+ RLQ L FE L M E E +S
Sbjct: 73 IYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVS 132
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
+Y RV ++N GE + D +++ K+LRS+ KF V IEE +D+ ++ +++L+G
Sbjct: 133 DYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLG 192
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRR 240
SLQ Y+ EK +KK I V N + + +S + R R R
Sbjct: 193 SLQAYE----EKKKKKEDIIEQVLNMQITKEENGQSYQRRGGGQVRGR--GRGGYGNGRG 246
Query: 241 TRPNVQDIKSDNSKSYNSQR-KVKEEDKSGQNK-GVQCHECEGYGHIRSEC 289
RP+ + N + NS R + K KS +K V+C+ C +GH SEC
Sbjct: 247 WRPHED---NTNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASEC 294
>gb|AAG50765.1| copia-type polyprotein, putative [Arabidopsis thaliana]
gi|12321254|gb|AAG50698.1| copia-type polyprotein,
putative [Arabidopsis thaliana] gi|25301687|pir||F96614
probable copia-type polyprotein T18I24.5 [imported] -
Arabidopsis thaliana
Length = 1320
Score = 122 bits (306), Expect = 1e-26
Identities = 93/291 (31%), Positives = 149/291 (50%), Gaps = 23/291 (7%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M A L + D+ W+ V KG+ P + EGS S ++ D L + + + KA+
Sbjct: 25 MKAILGAHDV--WEIVEKGFIEP---ENEGSLSQTQK---DGLRDSRKR----DKKALCL 72
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
I+ +D++ F + T AK+AWE L+T+++G +V+ RLQ L FE L M E E +S
Sbjct: 73 IYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVS 132
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
+Y RV ++N GE + D +++ K+LRS+ KF V IEE +D+ ++ +++L+G
Sbjct: 133 DYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLG 192
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRR 240
SLQ Y+ EK +KK + V N + + +S + R R R
Sbjct: 193 SLQAYE----EKKKKKEDIVEQVLNMQITKEENGQSYQRRGGGQVRGR--GRGGYGNGRG 246
Query: 241 TRPNVQDIKSDNSKSYNSQR-KVKEEDKSGQNK-GVQCHECEGYGHIRSEC 289
RP+ + N + NS R + K KS +K V+C+ C +GH SEC
Sbjct: 247 WRPHED---NTNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASEC 294
>gb|AAD50001.1| Hypothetical protein [Arabidopsis thaliana] gi|25301681|pir||F86246
hypothetical protein [imported] - Arabidopsis thaliana
Length = 1352
Score = 122 bits (306), Expect = 1e-26
Identities = 93/291 (31%), Positives = 149/291 (50%), Gaps = 23/291 (7%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M A L + D+ W+ V KG+ P + EGS S ++ D L + + + KA+
Sbjct: 25 MKAILGAHDV--WEIVEKGFIEP---ENEGSLSQTQK---DGLRDSRKR----DKKALCL 72
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
I+ +D++ F + T AK+AWE L+T+++G +V+ RLQ L FE L M E E +S
Sbjct: 73 IYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVS 132
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
+Y RV ++N GE + D +++ K+LRS+ KF V IEE +D+ ++ +++L+G
Sbjct: 133 DYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLG 192
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRR 240
SLQ Y+ EK +KK + V N + + +S + R R R
Sbjct: 193 SLQAYE----EKKKKKEDIVEQVLNMQITKEENGQSYQRRGGGQVRGR--GRGGYGNGRG 246
Query: 241 TRPNVQDIKSDNSKSYNSQR-KVKEEDKSGQNK-GVQCHECEGYGHIRSEC 289
RP+ + N + NS R + K KS +K V+C+ C +GH SEC
Sbjct: 247 WRPHED---NTNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASEC 294
>emb|CAB71063.1| copia-type polyprotein [Arabidopsis thaliana]
gi|11278364|pir||T47925 copia-type polyprotein -
Arabidopsis thaliana
Length = 1352
Score = 120 bits (302), Expect = 4e-26
Identities = 93/291 (31%), Positives = 148/291 (49%), Gaps = 23/291 (7%)
Query: 1 MIAFLKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNA 60
M A L + D+ W+ V KG+ P + EGS S ++ D L + + + KA+
Sbjct: 25 MKAILGAHDV--WEIVEKGFIEP---ENEGSLSQTQK---DGLRDSRKR----DKKALCL 72
Query: 61 IFNAVDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERIS 120
I+ +D++ F + T AK+AWE L+T+++G +V+ RLQ L FE L M E E +S
Sbjct: 73 IYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLRGEFEALQMKEGELVS 132
Query: 121 EYHMRVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIG 180
+Y RV ++N GE + D +++ K+LRS+ KF V IEE +D+ ++ +++L+G
Sbjct: 133 DYFSRVLTVTNNLKRNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEETKDLEAMTIEQLLG 192
Query: 181 SLQTYKLKLGEKPEKKTKSIAFVSNTTEGDDADSESENLSEALALLARKFNRALRKIDRR 240
SLQ Y+ EK +KK V N + + +S + R R R
Sbjct: 193 SLQAYE----EKKKKKEDIAEQVLNMQITKEENGQSYQRRGGGQVRGR--GRGGYGNGRG 246
Query: 241 TRPNVQDIKSDNSKSYNSQR-KVKEEDKSGQNK-GVQCHECEGYGHIRSEC 289
RP+ + N + NS R + K KS +K V+C+ C +GH SEC
Sbjct: 247 WRPHED---NTNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYASEC 294
>emb|CAB80821.1| putative transposon protein [Arabidopsis thaliana]
gi|4773900|gb|AAD29770.1| putative transposon protein
[Arabidopsis thaliana] gi|25407277|pir||E85057 probable
transposon protein [imported] - Arabidopsis thaliana
Length = 590
Score = 120 bits (301), Expect = 5e-26
Identities = 68/213 (31%), Positives = 113/213 (52%), Gaps = 7/213 (3%)
Query: 5 LKSIDIKTWKAVVKGWKHPVKAQAEGSTSTPEEKPEDELTSAEEEASLENSKAMNAIFNA 64
++SID+ W AV GW P A+ + K E + E+ A+ NS+A++ IF +
Sbjct: 364 IQSIDMDAWFAVEDGWMPPTTKDAKRDIVS---KSRTEWIADEKTAANHNSQALSVIFGS 420
Query: 65 VDKNMFRLINTCTVAKDAWEILKTTHEGTTRVRMSRLQMLTTRFENLTMTEDERISEYHM 124
+ +N F + C AK+ WEIL+ + E T V+ +RL ML + FENLTM +E + +++
Sbjct: 421 LLRNKFTQVQGCLSAKEVWEILQVSFECTNNVKRTRLDMLASEFENLTMEAEESVDDFNG 480
Query: 125 RVRDLSNASFALGEPMSDEKLVRKILRSVTSKFAMKVIAIEEAQDISSLKVDELIGSLQT 184
++ ++ + LG+ D+K+V+K LRS+ KF AI+ + + LK D+++G +Q
Sbjct: 481 KLSSITQEAVVLGKTYKDKKMVKKFLRSLPDKFQSHKSAIDVSLNSDQLKFDQVVGMMQA 540
Query: 185 YKLKLGEKPEKKTKSIAFVSNTTEGDDADSESE 217
Y E+ S A E DD E +
Sbjct: 541 Y----DTDKEEILNSYATYFGAIEDDDHTVEED 569
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.310 0.126 0.341
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 468,031,886
Number of Sequences: 2540612
Number of extensions: 17697931
Number of successful extensions: 68691
Number of sequences better than 10.0: 618
Number of HSP's better than 10.0 without gapping: 275
Number of HSP's successfully gapped in prelim test: 345
Number of HSP's that attempted gapping in prelim test: 67902
Number of HSP's gapped (non-prelim): 866
length of query: 310
length of database: 863,360,394
effective HSP length: 127
effective length of query: 183
effective length of database: 540,702,670
effective search space: 98948588610
effective search space used: 98948588610
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0101.12


