

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0100a.3
(244 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
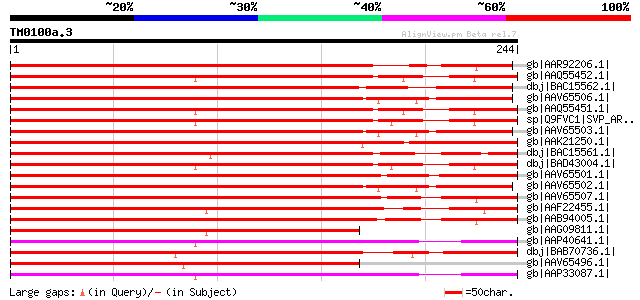
Score E
Sequences producing significant alignments: (bits) Value
gb|AAR92206.1| MADS box transcription factor [Populus tomentosa] 191 2e-47
gb|AAQ55452.1| short vegetative phase protein [Brassica rapa] 184 1e-45
dbj|BAC15562.1| IbMADS4 [Ipomoea batatas] gi|13448660|gb|AAK2715... 184 2e-45
gb|AAV65506.1| MPF2 [Physalis pubescens] gi|55792846|gb|AAV65505... 182 9e-45
gb|AAQ55451.1| short vegetative phase protein [Brassica rapa] 181 1e-44
sp|Q9FVC1|SVP_ARATH SHORT VEGETATIVE PHASE protein gi|30681743|r... 181 1e-44
gb|AAV65503.1| MPP4 [Physalis peruviana] 180 4e-44
gb|AAK21250.1| MADS-box transcription factor FBP13 [Petunia x hy... 179 5e-44
dbj|BAC15561.1| IbMADS3 [Ipomoea batatas] gi|13448658|gb|AAK2715... 179 6e-44
dbj|BAD43004.1| short vegegative phase protein (SVP) [Arabidopsi... 179 8e-44
gb|AAV65501.1| MSM2 [Solanum macrocarpon] 178 1e-43
gb|AAV65502.1| MPP3 [Physalis peruviana] 177 2e-43
gb|AAV65507.1| MADS16 [Solanum tuberosum] gi|55792844|gb|AAV6550... 177 3e-43
gb|AAF22455.1| MADS box protein [Paulownia kawakamii] 175 9e-43
gb|AAB94005.1| MADS transcriptional factor; STMADS16 [Solanum tu... 174 1e-42
gb|AAG09811.1| MADS-box transcription factor JOINTLESS [Lycopers... 174 3e-42
gb|AAP40641.1| SVP-like floral repressor [Eucalyptus occidentalis] 173 3e-42
dbj|BAB70736.1| putative MADS-domain transcription factor MpMADS... 173 4e-42
gb|AAV65496.1| MADS11 [Solanum tuberosum] gi|2735766|gb|AAB94006... 172 7e-42
gb|AAP33087.1| SVP-like floral repressor [Eucalyptus grandis] 171 2e-41
>gb|AAR92206.1| MADS box transcription factor [Populus tomentosa]
Length = 225
Score = 191 bits (485), Expect = 2e-47
Identities = 112/246 (45%), Positives = 159/246 (64%), Gaps = 27/246 (10%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++I+IKKIDN+++RQVTFSKRR+GLFKKA+ELS LCDA++A+I+FSAT KLFEY+SS
Sbjct: 1 MAREKIKIKKIDNVTARQVTFSKRRRGLFKKAEELSVLCDAEVAVIIFSATGKLFEYSSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
S G+L + +S DK PS +Q E+ ++ L K+V EK+H+LR++ GEDLQGL +
Sbjct: 61 SMKGVLARYNLHSNNLDKINQPSLELQLENSNHMRLSKEVSEKSHQLRRMRGEDLQGLNI 120
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFNE 180
+LQ+LE+ L+ L+ V K E+ M EISTL+RK V+L+EEN++LKQ + ++ +
Sbjct: 121 EELQQLEKALEVGLSRVLETKGERIMNEISTLERKGVQLLEENKQLKQKIATITR----- 175
Query: 181 YLCSIILSYGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTIS----SSSYLLEEDGSDT 236
+ L+ + S EST + SS +E+D SDT
Sbjct: 176 ------------GKRPALVDL------DTAVQEEGMSSESTTNVCSCSSGPPVEDDSSDT 217
Query: 237 SLKLGL 242
SLKLGL
Sbjct: 218 SLKLGL 223
>gb|AAQ55452.1| short vegetative phase protein [Brassica rapa]
Length = 241
Score = 184 bits (468), Expect = 1e-45
Identities = 112/253 (44%), Positives = 161/253 (63%), Gaps = 23/253 (9%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++IQI+KIDN ++RQVTFSKRR+GLFKKA+ELS LCDAD+ALI+FS+T KLFE+ SS
Sbjct: 1 MAREKIQIRKIDNATARQVTFSKRRRGLFKKAEELSVLCDADVALIIFSSTGKLFEFCSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQF-ESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
S +L + S +K PS +Q E+ + L K++ EK+H+LRQ+ GE+LQGL
Sbjct: 61 SMREVLERHNLQSKNLEKLDQPSLELQLVENSDHALLSKEIAEKSHQLRQMRGEELQGLN 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFN 179
+ +LQ+LE+ L+ L V K EK M EIS L+RK ++L++EN++L+Q L + N
Sbjct: 121 IEELQQLEKALEAGLTRVIETKSEKIMSEISDLQRKGMKLMDENKRLRQHGTQLTE--EN 178
Query: 180 EYLCSIILS-----YGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTI---SSSSYLLEE 231
E L I + YGG++S+ + Y + QS ES +S+ ++
Sbjct: 179 ERLGKQIYNNMHERYGGVESEKTAV------------YEEGQSSESITNAGNSTGAPVDS 226
Query: 232 DGSDTSLKLGLVY 244
+ SDTSL+LGL Y
Sbjct: 227 ESSDTSLRLGLPY 239
>dbj|BAC15562.1| IbMADS4 [Ipomoea batatas] gi|13448660|gb|AAK27151.1| MADS box
transcription factor [Ipomoea batatas]
Length = 229
Score = 184 bits (466), Expect = 2e-45
Identities = 108/242 (44%), Positives = 156/242 (63%), Gaps = 16/242 (6%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++I+IKKIDNI++RQVTFSKRR+GLFKKA+EL+ LCDAD+ALIVFSAT KLFE+ASS
Sbjct: 1 MAREKIKIKKIDNITARQVTFSKRRRGLFKKAEELAVLCDADVALIVFSATGKLFEFASS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
+ +L + +S+ D+ T PS +Q E+ + L K+V +KT ELRQ+ GE+LQGL+L
Sbjct: 61 NMKDILGKYELHSSNLDQATQPSRELQLENSLHVRLSKEVADKTRELRQMKGEELQGLSL 120
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFNE 180
+LQKLE+ L+ L V K E+ + EI+TL+RK EL++EN++LK+ + ++ +
Sbjct: 121 EELQKLEKRLENGLTRVLETKGERVVTEIATLQRKGAELMKENKQLKE---KMARVNGEK 177
Query: 181 YLCSIILSYGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTISSSSYLLEEDGSDTSLKL 240
+ + G+ +P Q + + +S E+D SDTSLKL
Sbjct: 178 FPVIADVEAAGL-------------IPEEGQSSESITTNVCSCNSGPPPEDDCSDTSLKL 224
Query: 241 GL 242
GL
Sbjct: 225 GL 226
>gb|AAV65506.1| MPF2 [Physalis pubescens] gi|55792846|gb|AAV65505.1| MPF2 [Physalis
pubescens]
Length = 249
Score = 182 bits (461), Expect = 9e-45
Identities = 111/250 (44%), Positives = 156/250 (62%), Gaps = 12/250 (4%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++I+IKKIDNI++RQVTFSKRR+GLFKKA+ELS LCDAD+ALI+FS+T KLF+++SS
Sbjct: 1 MAREKIKIKKIDNITARQVTFSKRRRGLFKKAEELSVLCDADVALIIFSSTGKLFDFSSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
S +L SA DK PS +Q E+ N LRK+V +KT ELRQ+ GE+L+GL+L
Sbjct: 61 SMKDILGKYKLQSANLDKVDQPSLDLQLENSLNVRLRKQVADKTRELRQMKGEELEGLSL 120
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKL---- 176
+LQ++E+ L+ V +K + M EI+ L+RK EL+EEN+KLKQ + + KL
Sbjct: 121 EELQQIEKRLEAGFNRVLEIKGTRIMDEIANLQRKGAELMEENKKLKQKM-EMMKLGKFP 179
Query: 177 LFNEYLCSIILSYGGIDS----QHILMKYM*F*VPSLTQYGQRQSLESTISSSSYLLEED 232
L + C +I DS ++ P SL+ ++ +++D
Sbjct: 180 LLTDMDCMVIEEGQSSDSIITTNNVCSSNS---GPPPEDDSSNASLKLGCNNGLAAVDDD 236
Query: 233 GSDTSLKLGL 242
S TSLKLGL
Sbjct: 237 CSITSLKLGL 246
>gb|AAQ55451.1| short vegetative phase protein [Brassica rapa]
Length = 241
Score = 181 bits (460), Expect = 1e-44
Identities = 111/253 (43%), Positives = 159/253 (61%), Gaps = 23/253 (9%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++IQI+KIDN ++RQVTFSKRR+GLFKKA+ELS LCDAD+ALI+FS+T KLFE+ SS
Sbjct: 1 MAREKIQIRKIDNATARQVTFSKRRRGLFKKAEELSVLCDADVALIIFSSTGKLFEFCSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQF-ESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
S +L + S +K PS +Q E+ + L K++ EK+H LRQ+ GE+LQGL
Sbjct: 61 SMREVLERHNLQSKNLEKLDQPSLELQLVENSDHALLSKEIAEKSHRLRQMRGEELQGLN 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFN 179
+ +LQ+LE+ L+ L V K EK M EIS L+RK ++L++EN++L+Q L + N
Sbjct: 121 IEELQQLEKALESGLTRVIETKSEKIMSEISDLQRKGMKLMDENKRLRQHGTQLTE--EN 178
Query: 180 EYLCSIILS-----YGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTI---SSSSYLLEE 231
E L I + YGG++S+ + Y + QS ES +S+ ++
Sbjct: 179 ERLGKQIYNNMHERYGGVESEKTAV------------YEEGQSSESITNAGNSTGAPVDS 226
Query: 232 DGSDTSLKLGLVY 244
+ SD SL+LGL Y
Sbjct: 227 ESSDISLRLGLPY 239
>sp|Q9FVC1|SVP_ARATH SHORT VEGETATIVE PHASE protein gi|30681743|ref|NP_179840.2| short
vegetative phase protein (SVP) [Arabidopsis thaliana]
gi|10944320|gb|AAG24508.1| short vegetative phase
protein [Arabidopsis thaliana]
Length = 240
Score = 181 bits (460), Expect = 1e-44
Identities = 107/252 (42%), Positives = 162/252 (63%), Gaps = 22/252 (8%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++IQI+KIDN ++RQVTFSKRR+GLFKKA+ELS LCDAD+ALI+FS+T KLFE+ SS
Sbjct: 1 MAREKIQIRKIDNATARQVTFSKRRRGLFKKAEELSVLCDADVALIIFSSTGKLFEFCSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQF-ESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
S +L + S +K PS +Q E+ + + K++ +K+H LRQ+ GE+LQGL
Sbjct: 61 SMKEVLERHNLQSKNLEKLDQPSLELQLVENSDHARMSKEIADKSHRLRQMRGEELQGLD 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFN 179
+ +LQ+LE+ L+ L V K +K M EIS L++K ++L++EN++L+Q L + N
Sbjct: 121 IEELQQLEKALETGLTRVIETKSDKIMSEISELQKKGMQLMDENKRLRQQGTQLTE--EN 178
Query: 180 EYL----CSIILSYGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTI---SSSSYLLEED 232
E L C+ + ++GG +S++ + Y + QS ES +S+ ++ +
Sbjct: 179 ERLGMQICNNVHAHGGAESENAAV------------YEEGQSSESITNAGNSTGAPVDSE 226
Query: 233 GSDTSLKLGLVY 244
SDTSL+LGL Y
Sbjct: 227 SSDTSLRLGLPY 238
>gb|AAV65503.1| MPP4 [Physalis peruviana]
Length = 247
Score = 180 bits (456), Expect = 4e-44
Identities = 110/250 (44%), Positives = 156/250 (62%), Gaps = 12/250 (4%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++I+IKKIDNI++RQVTFSKRR+GLFKKA+ELS LCDAD+ALI+FS+T KLF+++SS
Sbjct: 1 MAREKIKIKKIDNITARQVTFSKRRRGLFKKAEELSILCDADVALIIFSSTGKLFDFSSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
S +L SA DK P +Q E+ N LRK+V +KT ELRQ+ GE+L+GL+L
Sbjct: 61 SMKDILGKYKLQSANLDKVDQPFLDLQLENSLNVRLRKQVADKTRELRQMKGEELEGLSL 120
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKL---- 176
+LQ++E+ L+ V +K + M EI+ L+RK EL+EEN+KLKQ + + KL
Sbjct: 121 EELQQIEKRLEAGFNRVLEIKGTRIMDEIANLQRKGAELMEENKKLKQKM-EMMKLGKLP 179
Query: 177 LFNEYLCSIILSYGGIDS----QHILMKYM*F*VPSLTQYGQRQSLESTISSSSYLLEED 232
L E C ++ DS ++ P SL+ + ++ +++D
Sbjct: 180 LITEMECMVMEEGQSSDSIITTNNVCSSNT---GPPPEDDSSNASLKLSCNNGLAPVDDD 236
Query: 233 GSDTSLKLGL 242
S TSLKLGL
Sbjct: 237 CSITSLKLGL 246
>gb|AAK21250.1| MADS-box transcription factor FBP13 [Petunia x hybrida]
Length = 245
Score = 179 bits (455), Expect = 5e-44
Identities = 107/246 (43%), Positives = 158/246 (63%), Gaps = 4/246 (1%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++I+IKKIDNI++RQVTFSKRR+GL KKA ELS LCDA++ALI+FSAT KLFEYASS
Sbjct: 1 MAREKIKIKKIDNITARQVTFSKRRRGLMKKAAELSVLCDAEVALIIFSATGKLFEYASS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
S +L +SA+ +K PS +Q E+ N L K++ +K ELRQ+ GE+L+GL+L
Sbjct: 61 SMEDILGKYKFHSASLEKDDQPSLDLQLENSLNMRLSKEIADKNRELRQMRGEELEGLSL 120
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ--VLYSLFKLLF 178
++LQK+E+ L+ L V ++K + M EI+ L++K +L+EEN++LKQ V+ S KL
Sbjct: 121 NELQKIEKKLEAGLTRVLQIKGTRIMDEITNLQKKGADLMEENKQLKQKMVIMSEGKLPL 180
Query: 179 NEYLCSIILSYGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTISSSSYLLEEDGSDTSL 238
+ L +++ G S + ++ + + S+ +E+D SDT L
Sbjct: 181 HSELECMVMEEG--QSSESITTHVCSCSSGPPEDDYSNASLKLGCSNGPTVEDDCSDTFL 238
Query: 239 KLGLVY 244
KLGL +
Sbjct: 239 KLGLPF 244
>dbj|BAC15561.1| IbMADS3 [Ipomoea batatas] gi|13448658|gb|AAK27150.1| MADS box
transcription factor [Ipomoea batatas]
Length = 227
Score = 179 bits (454), Expect = 6e-44
Identities = 105/245 (42%), Positives = 155/245 (62%), Gaps = 19/245 (7%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++I+IKKIDNI++RQVTFSKRR+GLFKKA+ELS LCDAD+ALI+FS+T KLF+YASS
Sbjct: 1 MAREKIKIKKIDNITARQVTFSKRRRGLFKKAEELSVLCDADVALIIFSSTGKLFDYASS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDT-LRKKVEEKTHELRQLNGEDLQGLT 119
S G+L + +S +K PS +Q ++N + L K++ + TH LRQ+ GEDLQG++
Sbjct: 61 SMKGILERRNLHSKNLEKMDQPSLELQLVENANHSRLSKEIADMTHRLRQMRGEDLQGMS 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFN 179
+ +LQ+LE L+ L+ V K EK M+EI+ L++K + L+EE ++L Q + ++
Sbjct: 121 IEELQQLERSLETGLSRVIEKKGEKIMKEINELQQKGMNLMEEKERLTQQVMAISN---G 177
Query: 180 EYLCSIILSYGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTISSSSYLLEEDGSDTSLK 239
+ + ++I S ++ + L+ ST Y +D SDTSLK
Sbjct: 178 QRVTAVINSDNMLNEE------------GLSSESITNVCNSTSPPQDY---DDSSDTSLK 222
Query: 240 LGLVY 244
LGL Y
Sbjct: 223 LGLPY 227
>dbj|BAD43004.1| short vegegative phase protein (SVP) [Arabidopsis thaliana]
Length = 240
Score = 179 bits (453), Expect = 8e-44
Identities = 106/252 (42%), Positives = 161/252 (63%), Gaps = 22/252 (8%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++IQI+KIDN ++RQVTFSKRR+GLFKKA+ELS LCDA +ALI+FS+T KLFE+ SS
Sbjct: 1 MAREKIQIRKIDNATARQVTFSKRRRGLFKKAEELSVLCDAGVALIIFSSTGKLFEFCSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQF-ESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
S +L + S +K PS +Q E+ + + K++ +K+H LRQ+ GE+LQGL
Sbjct: 61 SMKEVLERHNLQSKNLEKLDQPSLELQLVENSDHARMSKEIADKSHRLRQMRGEELQGLD 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFN 179
+ +LQ+LE+ L+ L V K +K M EIS L++K ++L++EN++L+Q L + N
Sbjct: 121 IEELQQLEKALETGLTRVIETKSDKIMSEISELQKKGMQLMDENKRLRQQGTQLTE--EN 178
Query: 180 EYL----CSIILSYGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTI---SSSSYLLEED 232
E L C+ + ++GG +S++ + Y + QS ES +S+ ++ +
Sbjct: 179 ERLGMQICNNVHAHGGAESENAAV------------YEEGQSSESITNAGNSTGAPVDSE 226
Query: 233 GSDTSLKLGLVY 244
SDTSL+LGL Y
Sbjct: 227 SSDTSLRLGLPY 238
>gb|AAV65501.1| MSM2 [Solanum macrocarpon]
Length = 239
Score = 178 bits (451), Expect = 1e-43
Identities = 105/244 (43%), Positives = 154/244 (63%), Gaps = 6/244 (2%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++I+IKKIDNI++RQVTFSKRR+GLFKKA+ELS LCDAD+ALI+FSAT KLF++AS+
Sbjct: 1 MAREKIKIKKIDNITARQVTFSKRRRGLFKKAEELSVLCDADVALIIFSATGKLFDFAST 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
S +L SA+ +K PS +Q E+ N L K++ +KT ELRQ+ GE+L+GL+L
Sbjct: 61 SMKDILGKYKLQSASLEKVEQPSLDLQLENSLNMRLNKEIADKTRELRQMRGEELEGLSL 120
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFNE 180
+LQ++E+ L+ V +K + M EI+ L+RK EL+EEN++LKQ + + K
Sbjct: 121 EELQQIEKKLEAGFNRVLEIKSTRIMGEITNLQRKGAELMEENKQLKQKMEIMKKGKLP- 179
Query: 181 YLCSIILSYGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTISSSSYLLEEDGSDTSLKL 240
L + ++ G S+ I+ + G + + +E+D S TSLKL
Sbjct: 180 -LVTEMVMEDGQSSESIITSNN----VCSSNSGPPPDQDDSSKIGGNAVEDDCSITSLKL 234
Query: 241 GLVY 244
GL +
Sbjct: 235 GLPF 238
>gb|AAV65502.1| MPP3 [Physalis peruviana]
Length = 249
Score = 177 bits (449), Expect = 2e-43
Identities = 109/250 (43%), Positives = 155/250 (61%), Gaps = 12/250 (4%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++I+IKKIDNI++RQVTFSKRR+GLFKKA+ELS LCDAD+ALI+FS+T KLF+++SS
Sbjct: 1 MAREKIKIKKIDNITARQVTFSKRRRGLFKKAEELSILCDADVALIIFSSTGKLFDFSSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
S +L SA DK P +Q E+ N LRK+V +KT ELRQ+ GE+L+GL+L
Sbjct: 61 SMKDILGKYKLQSANLDKVDQPFLDLQLENSLNVRLRKQVADKTRELRQMKGEELEGLSL 120
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKL---- 176
+LQ++E+ L+ V +K + M EI+ L+RK EL+EEN+KLKQ + + KL
Sbjct: 121 EELQQIEKRLEAGFNRVLEIKGTRIMDEIANLQRKGAELMEENKKLKQKM-EMMKLGKLP 179
Query: 177 LFNEYLCSIILSYGGIDS----QHILMKYM*F*VPSLTQYGQRQSLESTISSSSYLLEED 232
L + C ++ DS ++ P SL+ ++ +++D
Sbjct: 180 LLTDMDCMVMEEGQSSDSIITTNNVCSSNT---GPPPEDDSSNASLKLGCNNGLAPVDDD 236
Query: 233 GSDTSLKLGL 242
S TSLKLGL
Sbjct: 237 CSITSLKLGL 246
>gb|AAV65507.1| MADS16 [Solanum tuberosum] gi|55792844|gb|AAV65504.1| MADS16
[Solanum tuberosum]
Length = 235
Score = 177 bits (448), Expect = 3e-43
Identities = 106/245 (43%), Positives = 157/245 (63%), Gaps = 12/245 (4%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++I+IKKIDNI++RQVTFSKRR+GLFKKA+ELS LCDAD+ALI+FSAT KLF++AS+
Sbjct: 1 MAREKIKIKKIDNITARQVTFSKRRRGLFKKAEELSVLCDADVALIIFSATGKLFDFAST 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
S +L SA+ +K PS +Q E+ N L K+V +KT ELRQ+ GE+L+GL+L
Sbjct: 61 SMKDILGKYKLQSASLEKVDEPSLDLQLENSLNMRLSKQVADKTRELRQMRGEELEGLSL 120
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFNE 180
+LQ++E+ L+ V +K + M EI+ L+RK EL+EEN++LK + + K F
Sbjct: 121 EELQQIEKRLEAGFNRVLEIKGTRIMDEITNLQRKGAELMEENKQLKHKMEIMKKGKFP- 179
Query: 181 YLCSIILSYGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTIS-SSSYLLEEDGSDTSLK 239
L + ++ G S+ I+ + Q S +++ + +E++ S TSLK
Sbjct: 180 -LLTDMVMEEGQSSESII---------TTNNPDQDDSSNASLKLGGTTAVEDECSITSLK 229
Query: 240 LGLVY 244
LGL +
Sbjct: 230 LGLPF 234
>gb|AAF22455.1| MADS box protein [Paulownia kawakamii]
Length = 227
Score = 175 bits (444), Expect = 9e-43
Identities = 108/247 (43%), Positives = 152/247 (60%), Gaps = 25/247 (10%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++IQIKKIDN ++RQVTFSKRR+G+FKKA+ELS LCDAD+ LI+FS+T KLFEYASS
Sbjct: 1 MAREKIQIKKIDNATARQVTFSKRRRGIFKKAEELSVLCDADVGLIIFSSTGKLFEYASS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSN-DTLRKKVEEKTHELRQLNGEDLQGLT 119
S +L + +S DK PS +Q DSN L K+V E++H+LR++ GE+LQGL+
Sbjct: 61 SMKEILGRHNLHSKNLDKLEQPSLELQLVEDSNYSRLSKEVAERSHQLRRMRGEELQGLS 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFN 179
+ +LQ L++ L+ L+ V K EK M+ + RK +L+EEN++L+Q + + N
Sbjct: 121 IEKLQHLKKSLESGLSRVIEKKGEKIMKGDQSTSRKGKQLMEENERLRQQVADISNDCKN 180
Query: 180 EYLCSIILSYGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTISSSSY--LLEEDGSDTS 237
DS++I+ Y + QS ES + +S + D SDTS
Sbjct: 181 N---------AASDSENIV-------------YDEGQSSESVNACNSVGPPQDYDSSDTS 218
Query: 238 LKLGLVY 244
LKLGL Y
Sbjct: 219 LKLGLPY 225
>gb|AAB94005.1| MADS transcriptional factor; STMADS16 [Solanum tuberosum]
gi|7446547|pir||T06995 probable MADS box transcription
factor MADS16 - potato
Length = 234
Score = 174 bits (442), Expect = 1e-42
Identities = 105/245 (42%), Positives = 156/245 (62%), Gaps = 13/245 (5%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++I+IKKIDNI++RQVTFSKRR+GLFKKA+ELS LCDAD+ALI+FS+T KLF++AS+
Sbjct: 1 MAREKIKIKKIDNITARQVTFSKRRRGLFKKAEELSVLCDADVALIIFSSTGKLFDFAST 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLTL 120
S +L SA+ +K PS +Q E+ N L K+V +KT ELRQ+ GE+L+GL+L
Sbjct: 61 SMKDILGKYKLQSASLEKVDEPSLDLQLENSLNMRLSKQVADKTRELRQMRGEELEGLSL 120
Query: 121 HQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFNE 180
+LQ++E+ L+ V +K + M EI+ L+RK EL+EEN++LK + + L
Sbjct: 121 EELQQIEKRLEAGFNRVLEIKGTRIMDEITNLQRKGAELMEENKQLKHKMEIMKGKL--- 177
Query: 181 YLCSIILSYGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTIS-SSSYLLEEDGSDTSLK 239
L + ++ G S+ I+ + Q S +++ + +E+D S TSLK
Sbjct: 178 PLLTDMVMEEGQSSESII---------TTNNPDQDDSSNASLKLGGTTAVEDDCSITSLK 228
Query: 240 LGLVY 244
LGL +
Sbjct: 229 LGLPF 233
>gb|AAG09811.1| MADS-box transcription factor JOINTLESS [Lycopersicon esculentum]
gi|17433048|sp|Q9FUY6|JOIN_LYCES MADS-box JOINTLESS
protein (LeMADS)
Length = 265
Score = 174 bits (440), Expect = 3e-42
Identities = 90/169 (53%), Positives = 125/169 (73%), Gaps = 1/169 (0%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++IQIKKIDN ++RQVTFSKRR+GLFKKA+ELS LCDAD+ALI+FS+T KLF+Y+SS
Sbjct: 1 MAREKIQIKKIDNSTARQVTFSKRRRGLFKKAEELSVLCDADVALIIFSSTGKLFDYSSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQFESDSN-DTLRKKVEEKTHELRQLNGEDLQGLT 119
S +L D +S +K PS +Q +SN L K++ EK+H LRQ+ GE+LQGL
Sbjct: 61 SMKQILERRDLHSKNLEKLDQPSLELQLVENSNYSRLSKEISEKSHRLRQMRGEELQGLN 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
+ +LQ+LE L+ L+ V K +K M+EI+ L++K + L+EEN+KL+Q
Sbjct: 121 IEELQQLERSLETGLSRVIERKGDKIMREINQLQQKGMHLMEENEKLRQ 169
>gb|AAP40641.1| SVP-like floral repressor [Eucalyptus occidentalis]
Length = 227
Score = 173 bits (439), Expect = 3e-42
Identities = 108/245 (44%), Positives = 146/245 (59%), Gaps = 21/245 (8%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++IQIKKI N ++RQVTFSKRR+GLFKKA+ELS LCDAD+ALIVFS++ KLFEY SS
Sbjct: 1 MAREKIQIKKITNATARQVTFSKRRRGLFKKAEELSVLCDADVALIVFSSSGKLFEYCSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQF-ESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
S +L ++S K PS +Q E+ L K+V EK H+LRQ+ GE+LQGL
Sbjct: 61 SMKEILERHHSHSENLGKLDQPSLKLQLVENGDYSRLSKEVAEKGHQLRQMRGEELQGLN 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFN 179
+ +LQ+LE+ L+ L V K EK M+EI+ L++K +L+EEN++LKQ + +
Sbjct: 121 IDELQQLEKSLEAGLNRVIEKKGEKIMKEITDLQQKGAKLMEENKRLKQQVTEISGRKTT 180
Query: 180 EYLCSIILSYGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTISSSSYLLEEDGSDTSLK 239
I++ G+ S+ I SSS E+D SD SLK
Sbjct: 181 ATDSETIINEEGLSSESI--------------------TNVCSSSSGPPQEDDSSDISLK 220
Query: 240 LGLVY 244
LGL Y
Sbjct: 221 LGLPY 225
>dbj|BAB70736.1| putative MADS-domain transcription factor MpMADS1 [Magnolia
praecocissima]
Length = 229
Score = 173 bits (438), Expect = 4e-42
Identities = 108/247 (43%), Positives = 154/247 (61%), Gaps = 23/247 (9%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R +IQIKKIDN+++RQVTFSKRR+GLFKKA+ELS LCDA++ALI+FSAT KLFEY+SS
Sbjct: 1 MARGKIQIKKIDNVTARQVTFSKRRRGLFKKAEELSILCDAEVALIIFSATGKLFEYSSS 60
Query: 61 SRGGLLPLPDANSAT*DK-TTPPSPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
S ++ +S K PS +Q E+ + + L K+V EK+H +RQ+ GED+QGLT
Sbjct: 61 SMKEIIERHTMHSKNLQKLDQQPSLELQLENSNYNRLSKQVAEKSHLIRQMRGEDIQGLT 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFN 179
+ +LQKLE+ L+ L+ V K E+ M+EIS L+ K V+L+EEN +L+Q +
Sbjct: 121 VEELQKLEKTLETGLSRVMERKAEQIMKEISGLQIKGVKLMEENMRLRQRI--------- 171
Query: 180 EYLCSIILSYGGI--DSQHILMKYM*F*VPSLTQYGQRQSLESTISSSSYLLEEDGSDTS 237
I +S G D Q I + + + + GQ + +S + + SDTS
Sbjct: 172 -----IEMSRGDSKGDRQIIESEIV------VNEDGQSSDSVTNACNSGAPQDYESSDTS 220
Query: 238 LKLGLVY 244
LKLG+ +
Sbjct: 221 LKLGVPF 227
>gb|AAV65496.1| MADS11 [Solanum tuberosum] gi|2735766|gb|AAB94006.1| MADS
transcriptional factor; STMADS11 [Solanum tuberosum]
gi|7446550|pir||T06996 MADS-box transcription factor
MADS11 - potato
Length = 221
Score = 172 bits (436), Expect = 7e-42
Identities = 95/171 (55%), Positives = 122/171 (70%), Gaps = 3/171 (1%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++IQIKKIDN+++RQVTFSKRR+GLFKKAQELSTLCDADI LIVFSAT KLFEY+SS
Sbjct: 1 MVRQKIQIKKIDNLTARQVTFSKRRRGLFKKAQELSTLCDADIGLIVFSATGKLFEYSSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPP---SPTMQFESDSNDTLRKKVEEKTHELRQLNGEDLQG 117
S L+ S P S + E ++ L + EK ELRQL+GE+LQG
Sbjct: 61 SMMQLIEKHKMQSERDSMDNPEQLHSSNLLSEKKTHAMLSRDFVEKNRELRQLHGEELQG 120
Query: 118 LTLHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQ 168
L L L KLE++++ ++ V R+K +KFM+EIS+LK+KE +L EEN +LKQ
Sbjct: 121 LGLDDLMKLEKLVEGGISRVLRIKGDKFMKEISSLKKKEAQLQEENSQLKQ 171
>gb|AAP33087.1| SVP-like floral repressor [Eucalyptus grandis]
Length = 227
Score = 171 bits (433), Expect = 2e-41
Identities = 107/245 (43%), Positives = 145/245 (58%), Gaps = 21/245 (8%)
Query: 1 MTRKRIQIKKIDNISSRQVTFSKRRKGLFKKAQELSTLCDADIALIVFSATNKLFEYASS 60
M R++IQIKKI N ++RQVTFSKRR+GLFKKA+ELS LCDAD+ALIVFS++ KLFEY SS
Sbjct: 1 MAREKIQIKKITNATARQVTFSKRRRGLFKKAEELSVLCDADVALIVFSSSGKLFEYCSS 60
Query: 61 SRGGLLPLPDANSAT*DKTTPPSPTMQF-ESDSNDTLRKKVEEKTHELRQLNGEDLQGLT 119
S +L ++S K PS +Q E+ L K+V EK H+LRQ+ GE+LQGL
Sbjct: 61 SMKEILERHHSHSENLGKLDQPSLKLQLVENGDYSRLSKEVAEKGHQLRQMRGEELQGLN 120
Query: 120 LHQLQKLEEVLKRSLASVSRVKDEKFMQEISTLKRKEVELIEENQKLKQVLYSLFKLLFN 179
+ +LQ+LE+ L+ L V K EK M+EI+ L++K +L+EE ++LKQ + +
Sbjct: 121 IDELQQLEKSLEAGLNRVIEKKGEKIMKEITDLQQKGAKLMEETKRLKQQVTEISGRKTT 180
Query: 180 EYLCSIILSYGGIDSQHILMKYM*F*VPSLTQYGQRQSLESTISSSSYLLEEDGSDTSLK 239
I++ G+ S+ I SSS E+D SD SLK
Sbjct: 181 ATDSETIINEEGLSSESI--------------------TNVCSSSSGPPQEDDSSDISLK 220
Query: 240 LGLVY 244
LGL Y
Sbjct: 221 LGLPY 225
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.322 0.136 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 350,023,168
Number of Sequences: 2540612
Number of extensions: 13379939
Number of successful extensions: 58228
Number of sequences better than 10.0: 2152
Number of HSP's better than 10.0 without gapping: 1863
Number of HSP's successfully gapped in prelim test: 290
Number of HSP's that attempted gapping in prelim test: 54782
Number of HSP's gapped (non-prelim): 2656
length of query: 244
length of database: 863,360,394
effective HSP length: 125
effective length of query: 119
effective length of database: 545,783,894
effective search space: 64948283386
effective search space used: 64948283386
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0100a.3


