

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0100a.12
(405 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
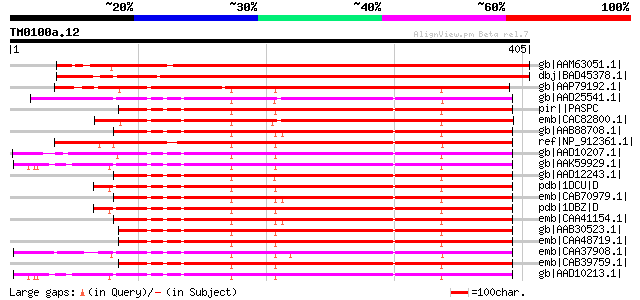
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM63051.1| fructose-bisphosphatase-like protein [Arabidopsis... 545 e-154
dbj|BAD45378.1| putative ructose 1,6-bisphosphatase [Oryza sativ... 534 e-150
gb|AAP79192.1| fructose-1,6 bisphosphatase [Bigelowiella natans] 293 5e-78
gb|AAD25541.1| fructose-1,6-bisphosphatase precursor [Solanum tu... 278 2e-73
pir||PASPC fructose-bisphosphatase (EC 3.1.3.11), chloroplast - ... 271 2e-71
emb|CAC82800.1| fructose 1,6-bisphosphatase [Galdieria sulphuraria] 271 3e-71
gb|AAB88708.1| fructose-1,6-bisphosphate [Brassica napus] gi|743... 270 4e-71
ref|NP_912361.1| fructose-1,6-bisphosphatase [Oryza sativa (japo... 270 4e-71
gb|AAD10207.1| fructose 1,6-bisphosphatase [Spinacia oleracea] g... 270 4e-71
gb|AAK59929.1| fructose-1,6-bisphosphatase [Pisum sativum] 268 2e-70
gb|AAD12243.1| fructose-1,6-bisphosphatase precursor [Brassica n... 267 4e-70
pdb|1DCU|D Chain D, Redox Signaling In The Chloroplast: Structur... 266 6e-70
emb|CAB70979.1| fructose-bisphosphatase precursor [Arabidopsis t... 266 8e-70
pdb|1DBZ|D Chain D, C153s Mutant Of Pea Fructose-1,6-Bisphosphat... 266 8e-70
emb|CAA41154.1| fructose-bisphosphatase [Arabidopsis thaliana] g... 266 1e-69
gb|AAB30523.1| fructose-1,6-biphosphatase, FBPase {EC 3.1.3.11} ... 266 1e-69
emb|CAA48719.1| fructose-bisphosphatase [Pisum sativum] gi|32271... 265 2e-69
emb|CAA37908.1| fructose-bisphosphatase [Triticum aestivum] gi|2... 265 2e-69
emb|CAB39759.1| fructose-1,6-bisphosphatase [Pisum sativum] 265 2e-69
gb|AAD10213.1| fructose-1,6-bisphosphatase [Pisum sativum] gi|74... 264 4e-69
>gb|AAM63051.1| fructose-bisphosphatase-like protein [Arabidopsis thaliana]
gi|21700925|gb|AAM70586.1| AT5g64380/MSJ1_22
[Arabidopsis thaliana] gi|9759414|dbj|BAB09869.1|
fructose-bisphosphatase-like protein [Arabidopsis
thaliana] gi|15237685|ref|NP_201243.1|
fructose-1,6-bisphosphatase family protein [Arabidopsis
thaliana] gi|17063187|gb|AAL32988.1|
fructose-bisphosphatase-like protein [Arabidopsis
thaliana]
Length = 404
Score = 545 bits (1404), Expect = e-154
Identities = 278/372 (74%), Positives = 318/372 (84%), Gaps = 13/372 (3%)
Query: 37 LASISGSAKMRPLRALGGSSSSAGGDGDDGDGVVTLIEYLGK---EGIDLKDDLVVLLDH 93
+AS S +PL A+G + + GDD DG TLI++ G EG ++ +DLVVLL H
Sbjct: 43 MASRHQSTSFKPL-AVGRNLT-----GDDDDGYCTLIDFAGSGGGEGKNVGEDLVVLLYH 96
Query: 94 IQYACKRIAAIVASPFSCSIGKQTGVGAGGGGGSSGRDAPKPLDIVSNEIILSSLKKSGK 153
+Q+ACKRIA++VASPF+ S+GK + + G S RDAPKPLDIVSN+I+LSSL+ SGK
Sbjct: 97 LQHACKRIASLVASPFNSSLGKLSVNSSSG----SDRDAPKPLDIVSNDIVLSSLRNSGK 152
Query: 154 VAVMASEENDAPTWIIDDGPYVVVTDPLDGSRNIDASIPTGTIFGIYNRLEELDNLPTEE 213
VAVMASEEND+PTWI DDGPYVVV DPLDGSRNIDASIPTGTIFGIYNRL ELD+LP EE
Sbjct: 153 VAVMASEENDSPTWIKDDGPYVVVVDPLDGSRNIDASIPTGTIFGIYNRLVELDHLPVEE 212
Query: 214 KAQLNSLQSGSKLIAAGYVLYSSATILCVSFGSGTQAFTLDHSTGDFILTNPSIKIPPRG 273
KA+LNSLQ GS+L+A+GYVLYSSATI CV+ GSGT AFTLDHSTG+F+LT+ +IKIP RG
Sbjct: 213 KAELNSLQRGSRLVASGYVLYSSATIFCVTLGSGTHAFTLDHSTGEFVLTHQNIKIPTRG 272
Query: 274 QIYSVNDARYFDWPEGLRQYIDTVRQGKGRYPKKYSARYICSLVADLHRTLMYGGVAMNP 333
QIYSVNDARYFDWPEGLR+YIDTVRQGKG+ PKKYSARYICSLVADLHRTL+YGGVAMNP
Sbjct: 273 QIYSVNDARYFDWPEGLRKYIDTVRQGKGQNPKKYSARYICSLVADLHRTLLYGGVAMNP 332
Query: 334 RDHLRLVYEANPLSFLVEQAGGRGSDGKNRILSLQPVKLHQRLPLFLGSLEDMEELESFG 393
RDHLRLVYE NPL+FLVEQAGG+ SDGK ILS+QPVKLHQRLPLFLGSLED+ ELES+G
Sbjct: 333 RDHLRLVYEGNPLAFLVEQAGGKSSDGKRGILSIQPVKLHQRLPLFLGSLEDVAELESYG 392
Query: 394 DVQQKVNPGYEV 405
DVQQ VNPGYEV
Sbjct: 393 DVQQTVNPGYEV 404
>dbj|BAD45378.1| putative ructose 1,6-bisphosphatase [Oryza sativa (japonica
cultivar-group)]
Length = 401
Score = 534 bits (1376), Expect = e-150
Identities = 267/369 (72%), Positives = 313/369 (84%), Gaps = 9/369 (2%)
Query: 37 LASISGSAKMRPLRALGGSSSSAGGDGDDGDGVVTLIEYLGKEGIDLKDDLVVLLDHIQY 96
LA S ++ R + A G ++++A + TL+EY+G+ G DDLVVL+ H+Q
Sbjct: 42 LARRSPVSRARAVAADGMAAATAAAETPP-----TLLEYMGQAGA--ADDLVVLVAHVQS 94
Query: 97 ACKRIAAIVASPFSCSIGKQTGVGAGGGGGSSGRDAPKPLDIVSNEIILSSLKKSGKVAV 156
ACKRIAA+VASP + + + G GG ++GRDAPKPLD +SNEIILSSL++SGKVAV
Sbjct: 95 ACKRIAALVASPGNAELSR--GKAGGGVAVAAGRDAPKPLDELSNEIILSSLRRSGKVAV 152
Query: 157 MASEENDAPTWIIDDGPYVVVTDPLDGSRNIDASIPTGTIFGIYNRLEELDNLPTEEKAQ 216
MASEEN P W+ +D PYVVVTDPLDGSRNI+ SIPTGTIFGIYNRL ELD+LP EE+AQ
Sbjct: 153 MASEENHLPIWVSNDSPYVVVTDPLDGSRNIEVSIPTGTIFGIYNRLAELDHLPEEERAQ 212
Query: 217 LNSLQSGSKLIAAGYVLYSSATILCVSFGSGTQAFTLDHSTGDFILTNPSIKIPPRGQIY 276
LNSLQSG+ L+A+GYVLYSSATI C+SFG+GT FTLDH TG+F+LT+PSI+IPPRGQIY
Sbjct: 213 LNSLQSGTHLVASGYVLYSSATIFCISFGAGTHGFTLDHLTGEFVLTHPSIQIPPRGQIY 272
Query: 277 SVNDARYFDWPEGLRQYIDTVRQGKGRYPKKYSARYICSLVADLHRTLMYGGVAMNPRDH 336
SVNDARYFDWPEGLR+YIDT+RQGKG++PKKYSARY+CSLVAD HRTL+YGGVAMNPRDH
Sbjct: 273 SVNDARYFDWPEGLRKYIDTIRQGKGQHPKKYSARYVCSLVADFHRTLIYGGVAMNPRDH 332
Query: 337 LRLVYEANPLSFLVEQAGGRGSDGKNRILSLQPVKLHQRLPLFLGSLEDMEELESFGDVQ 396
LRLVYEANPLSFL EQAGGRGSDGK+RILS+QPVKLHQRLPLFLG +EDM ELES+GDVQ
Sbjct: 333 LRLVYEANPLSFLAEQAGGRGSDGKSRILSIQPVKLHQRLPLFLGGMEDMLELESYGDVQ 392
Query: 397 QKVNPGYEV 405
QKVNPGYEV
Sbjct: 393 QKVNPGYEV 401
>gb|AAP79192.1| fructose-1,6 bisphosphatase [Bigelowiella natans]
Length = 420
Score = 293 bits (751), Expect = 5e-78
Identities = 169/375 (45%), Positives = 241/375 (64%), Gaps = 32/375 (8%)
Query: 36 RLASISGSAKMRPLRALGGSSSSAGGDGDDGDGVVTLIEYLGKEGIDLK----DDLVVLL 91
R A++ S P++ SAG DDG VTL ++ +E + K +D+V L+
Sbjct: 54 RRAAVRSSVAPSPVQ------DSAGTIVDDG--TVTLTRFMIEEAMKSKTPGQEDMVRLI 105
Query: 92 DHIQYACKRIAAIVASPFSCSIGKQTGVGAGGGGGSSGRDAPKPLDIVSNEIILSSLKKS 151
I ACKRIA++V + I TG+ GGG + + K LD++SN+++ S+L+ S
Sbjct: 106 SSISVACKRIASMVQT---AGISGSTGLAEGGGSVNVQGEEQKKLDVISNDVLKSALRPS 162
Query: 152 GKVAVMASEENDAPTWIIDD---GPYVVVTDPLDGSRNIDASIPTGTIFGIYNRLEEL-- 206
GK+ V+ASEE D P ++D+ G YV DPLDGS NIDA+I TGTIFG++ EE
Sbjct: 163 GKLGVIASEEEDNPV-VVDELYSGEYVATFDPLDGSSNIDAAISTGTIFGVFKAPEECLI 221
Query: 207 ---DNLPTEEKAQLNS-LQSGSKLIAAGYVLYSSATILCVSFGSGTQAFTLDHSTGDFIL 262
DNL E+ L + LQ G+ L+AAGY +YSS+TIL ++ G G FTLD S G+FIL
Sbjct: 222 GDSDNLSIAEQQCLEATLQPGTNLVAAGYCMYSSSTILVLTTGDGLNGFTLDPSIGEFIL 281
Query: 263 TNPSIKIPPRGQIYSVNDARYFDWPEGLRQYIDTVRQGKGRYPKKYSARYICSLVADLHR 322
T+P+I+IP RG+IYS+N+A YFDW L+ Y+D +++ +G+ +KYS+RYI S+V D+HR
Sbjct: 282 THPNIQIPSRGKIYSMNEANYFDWDPKLQTYVDNLKKAEGQTGEKYSSRYIGSMVGDVHR 341
Query: 323 TLMYGGVAMNPRD------HLRLVYEANPLSFLVEQAGGRGSDGK-NRILSLQPVKLHQR 375
TL+YGG+ P D LRL+YEA P+S + EQAGG+ G R+L L P K+HQR
Sbjct: 342 TLLYGGIFAYPGDKKNVNGKLRLLYEAAPMSLIFEQAGGKSITGPGGRVLDLVPDKVHQR 401
Query: 376 LPLFLGSLEDMEELE 390
P+F+GS +D++E+E
Sbjct: 402 CPVFIGSPDDVDEVE 416
>gb|AAD25541.1| fructose-1,6-bisphosphatase precursor [Solanum tuberosum]
Length = 408
Score = 278 bits (711), Expect = 2e-73
Identities = 159/394 (40%), Positives = 234/394 (59%), Gaps = 28/394 (7%)
Query: 17 PKFQTFQSKTPLTTPYCTPRLASISGSAKMRPLRALGGSSSSAGGDGDDGDGVVTLIEYL 76
P KTP P + + S +G K + G +++ G + + + +
Sbjct: 23 PFLFALDKKTPFLCPKSSTKRRSFNGGVKCMAIETTSGFTATKKRSGYELQTLTSWLLRQ 82
Query: 77 GKEGIDLKDDLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVGAGGGGGSSGRDAPKPL 136
+ G+ + +L +++ I ACK+IA++V I TGV G G D K L
Sbjct: 83 EQAGV-IDAELTIVISSISMACKQIASLVQR---AGISNLTGVQ--GAVNIQGEDQKK-L 135
Query: 137 DIVSNEIILSSLKKSGKVAVMASEENDAPTWIIDD--GPYVVVTDPLDGSRNIDASIPTG 194
D+VSNE+ + L+ SG+ ++ASEE D P + + G Y+VV DPLDGS NIDA++ TG
Sbjct: 136 DVVSNEVFSNCLRSSGRTGIIASEEEDVPVAVEESYSGNYIVVFDPLDGSSNIDAAVSTG 195
Query: 195 TIFGIYNRLEE----------LDNLPTEEKAQLNSLQSGSKLIAAGYVLYSSATILCVSF 244
+IFGIYN +E LDN+ E+K +N Q G+ L+AAGY +YSS+ I ++
Sbjct: 196 SIFGIYNPNDECLADHGDDSTLDNI--EQKCIVNVCQPGTNLLAAGYCMYSSSVIFVLTL 253
Query: 245 GSGTQAFTLDHSTGDFILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQYIDTVRQGKGRY 304
G+G +F LD G+F+LT +++IP G+IYS N+ Y W + L++YID ++ G
Sbjct: 254 GNGVFSFNLDPMYGEFVLTQENVQIPKSGKIYSFNEGNYQLWDDKLKKYIDDLKD-PGPS 312
Query: 305 PKKYSARYICSLVADLHRTLMYGGVAMNPRDH------LRLVYEANPLSFLVEQAGGRGS 358
K YSARYI SLV D HRTL+YGG+ PRD LRL+YE P+SF+VEQAGG+GS
Sbjct: 313 GKPYSARYIGSLVGDFHRTLLYGGIYGYPRDQKSKNGKLRLLYECAPMSFIVEQAGGKGS 372
Query: 359 DGKNRILSLQPVKLHQRLPLFLGSLEDMEELESF 392
DG R+L +QP ++HQR+PL++GS E++E+LE +
Sbjct: 373 DGHQRVLDIQPTEVHQRVPLYIGSTEEVEKLEKY 406
>pir||PASPC fructose-bisphosphatase (EC 3.1.3.11), chloroplast - spinach
gi|999624|pdb|1SPI|D Chain D,
Fructose-1,6-Bisphosphatase (D-Fructose-1,6-Bisphosphate
1-Phosphohydrolase) (E.C.3.1.3.11) gi|999623|pdb|1SPI|C
Chain C, Fructose-1,6-Bisphosphatase
(D-Fructose-1,6-Bisphosphate 1-Phosphohydrolase)
(E.C.3.1.3.11) gi|999622|pdb|1SPI|B Chain B,
Fructose-1,6-Bisphosphatase (D-Fructose-1,6-Bisphosphate
1-Phosphohydrolase) (E.C.3.1.3.11) gi|999621|pdb|1SPI|A
Chain A, Fructose-1,6-Bisphosphatase
(D-Fructose-1,6-Bisphosphate 1-Phosphohydrolase)
(E.C.3.1.3.11)
Length = 358
Score = 271 bits (694), Expect = 2e-71
Identities = 149/325 (45%), Positives = 208/325 (63%), Gaps = 25/325 (7%)
Query: 86 DLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVGAGGGGGSSGRDAPKPLDIVSNEIIL 145
+L ++L I ACK+IA++V I TG+ G G D K LD+VSNE+
Sbjct: 39 ELTIVLSSISLACKQIASLVQR---AGISNLTGIQ--GAVNIQGEDQKK-LDVVSNEVFS 92
Query: 146 SSLKKSGKVAVMASEENDAPTWIIDD--GPYVVVTDPLDGSRNIDASIPTGTIFGIYNRL 203
S L+ SG+ ++ASEE D P + + G Y+VV DPLDGS NIDA++ TG+IFGIY+
Sbjct: 93 SCLRSSGRTGIIASEEEDVPVAVEESYSGNYIVVFDPLDGSSNIDAAVSTGSIFGIYSPN 152
Query: 204 EEL---------DNLPTEE-KAQLNSLQSGSKLIAAGYVLYSSATILCVSFGSGTQAFTL 253
+E L EE + +N Q G L+AAGY +YSS+ I ++ G G AFTL
Sbjct: 153 DECIVDSDHDDESQLSAEEQRCVVNVCQPGDNLLAAGYCMYSSSVIFVLTIGKGVYAFTL 212
Query: 254 DHSTGDFILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQYIDTVRQGKGRYPKKYSARYI 313
D G+F+LT+ I+IP G+IYS N+ Y WP+ L++Y+D +++ G K YS+RYI
Sbjct: 213 DPMYGEFVLTSEKIQIPKAGKIYSFNEGNYKMWPDKLKKYMDDLKE-PGESQKPYSSRYI 271
Query: 314 CSLVADLHRTLMYGGVAMNPRD------HLRLVYEANPLSFLVEQAGGRGSDGKNRILSL 367
SLV D HRTL+YGG+ PRD LRL+YE P+SF+VEQAGG+GSDG RIL +
Sbjct: 272 GSLVGDFHRTLLYGGIYGYPRDAKSKNGKLRLLYECAPMSFIVEQAGGKGSDGHQRILDI 331
Query: 368 QPVKLHQRLPLFLGSLEDMEELESF 392
QP ++HQR+PL++GS+E++E+LE +
Sbjct: 332 QPTEIHQRVPLYIGSVEEVEKLEKY 356
>emb|CAC82800.1| fructose 1,6-bisphosphatase [Galdieria sulphuraria]
Length = 422
Score = 271 bits (693), Expect = 3e-71
Identities = 152/347 (43%), Positives = 221/347 (62%), Gaps = 26/347 (7%)
Query: 67 DGVVTLIEYLGKEGIDLKD--DLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVGAGGG 124
D +L YL + KD D+V L++ IQ+ACK+IA++V + G+ G
Sbjct: 64 DTPTSLTRYLLEVAKQNKDMGDMVALINGIQFACKKIASLVGK---AGVTDLMGIYQQGI 120
Query: 125 GGSSGRDAPKPLDIVSNEIILSSLKKSGKVAVMASEENDAPTWIIDD--GPYVVVTDPLD 182
G + K LD++SNE++ ++LK SGK+AV+ASEE D P + + G YVVV DPLD
Sbjct: 121 VNVHGEEQKK-LDVLSNEVLKNALKYSGKMAVIASEEEDVPIMVEESYSGNYVVVFDPLD 179
Query: 183 GSRNIDASIPTGTIFGIYNRL---------EELDNLPTEEKAQLNSLQSGSKLIAAGYVL 233
GS N+DA +PTGTIFG++ + E +DN+ E N+LQ G +LIAAGY +
Sbjct: 180 GSSNLDAGLPTGTIFGVFQQQFSCLIHDYEESIDNM--ELACLQNTLQPGRRLIAAGYCI 237
Query: 234 YSSATILCVSFGSGTQAFTLDHSTGDFILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQY 293
YSS+T+L +S G+G FTLD G+F+LT +I+IP RG IYS N++ ++ W +G++ Y
Sbjct: 238 YSSSTMLVLSLGNGLHCFTLDTEVGEFVLTRANIQIPQRGNIYSFNESNFYQWDKGVQDY 297
Query: 294 IDTVRQGKGRYPKKYSARYICSLVADLHRTLMYGGVAMNPRD------HLRLVYEANPLS 347
I+ +++G + +YSARY+ S+VAD+HRT++YGG+ P D LRLVYE P++
Sbjct: 298 IERLKKGNNQTNCRYSARYVGSMVADVHRTILYGGIFGYPADKKNVSGKLRLVYECAPMA 357
Query: 348 FLVEQAGGRGSDGKNRILSLQPVKLHQRLPLFLGSLEDMEE-LESFG 393
+LVEQAGG+ + G IL L P +H+R PL LGS D+EE L+ +G
Sbjct: 358 YLVEQAGGKATTGIENILDLTPKDIHERKPLILGSPADIEEFLQVYG 404
>gb|AAB88708.1| fructose-1,6-bisphosphate [Brassica napus] gi|7435114|pir||T07987
fructose-bisphosphatase (EC 3.1.3.11) precursor,
chloroplast [validated] - rape
gi|585118|sp|Q07204|F16P_BRANA
FRUCTOSE-1,6-BISPHOSPHATASE, CHLOROPLAST PRECURSOR
(D-FRUCTOSE-1,6-BISPHOSPHATE 1-PHOSPHOHYDROLASE)
(FBPASE)
Length = 411
Score = 270 bits (691), Expect = 4e-71
Identities = 151/327 (46%), Positives = 211/327 (64%), Gaps = 23/327 (7%)
Query: 82 DLKDDLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVGAGGGGGSSGRDAPKPLDIVSN 141
++ +L +++ I ACK+IA++V I TGV G G D K LD+VSN
Sbjct: 90 EIDTELTIVMSSIAMACKQIASLVQR---AGISNLTGVQ--GAVNIQGEDQKK-LDVVSN 143
Query: 142 EIILSSLKKSGKVAVMASEENDAPTWIIDD--GPYVVVTDPLDGSRNIDASIPTGTIFGI 199
E+ + L+ SG+ ++ASEE D P + + G YVVV DPLDGS NIDA++ TG+IFGI
Sbjct: 144 EVFSNCLRSSGRTGIIASEEEDVPVAVEESYSGNYVVVFDPLDGSSNIDAAVSTGSIFGI 203
Query: 200 YNRLEEL--DNLPT------EEKAQLNSLQSGSKLIAAGYVLYSSATILCVSFGSGTQAF 251
Y+ +E D+ T EE+ +N Q G+ L+AAGY +YSS+ I ++ G G AF
Sbjct: 204 YSPNDECLPDSDDTSALGSEEERCIVNVCQPGNNLLAAGYCMYSSSVIFVLTLGKGVFAF 263
Query: 252 TLDHSTGDFILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQYIDTVRQGKGRYPKKYSAR 311
TLD G+F+LT +I+IP G+IYS N+ Y W E L++YID ++ G K YSAR
Sbjct: 264 TLDPMYGEFVLTQENIEIPKAGKIYSFNEGNYQMWDENLKKYIDDLKD-PGPSGKPYSAR 322
Query: 312 YICSLVADLHRTLMYGGVAMNPRD------HLRLVYEANPLSFLVEQAGGRGSDGKNRIL 365
YI SLV D HRTL+YGG+ PRD LRL+YE P+SF+VEQAGG+GSDG +R+L
Sbjct: 323 YIGSLVGDFHRTLLYGGIYGYPRDAKSKNGKLRLLYECAPMSFIVEQAGGKGSDGHHRVL 382
Query: 366 SLQPVKLHQRLPLFLGSLEDMEELESF 392
+QP ++HQR+PL++GS E++E+LE +
Sbjct: 383 DIQPTEIHQRVPLYIGSKEEVEKLEKY 409
>ref|NP_912361.1| fructose-1,6-bisphosphatase [Oryza sativa (japonica
cultivar-group)] gi|29893638|gb|AAP06892.1| putative
Fructose-1,6-Biphosphotase, chloroplast precursor [Oryza
sativa (japonica cultivar-group)]
gi|29893631|gb|AAP06885.1| fructose-1,6-bisphosphatase
[Oryza sativa (japonica cultivar-group)]
gi|3041777|dbj|BAA25423.1| fructose-1,6-bisphosphatase
[Oryza sativa] gi|3913641|sp|O64422|F16P_ORYSA
Fructose-1,6-bisphosphatase, chloroplast precursor
(D-fructose-1,6-bisphosphate 1-phosphohydrolase)
(FBPase)
Length = 406
Score = 270 bits (691), Expect = 4e-71
Identities = 157/378 (41%), Positives = 231/378 (60%), Gaps = 29/378 (7%)
Query: 36 RLASISGSAKMRPLRALGGSSSSAGGDGDDGDG--VVTLIEYLGKE--GIDLKDDLVVLL 91
RLA S + +R + A+ +S+ A + + TL +L K+ + ++ ++L
Sbjct: 35 RLAGGSSAPSVRCMAAVDTASAPAATEASKKSSYEITTLTTWLLKQEQAGTIDGEMTIVL 94
Query: 92 DHIQYACKRIAAIVA-SPFSCSIGKQTGVGAGGGGGSSGRDAPKPLDIVSNEIILSSLKK 150
I ACK+IA++V +P S G Q V G + K LD+VSNE+ + LK
Sbjct: 95 ASISTACKQIASLVQRAPISNLTGVQGAVNVQG-------EDQKKLDVVSNEVFSNCLKS 147
Query: 151 SGKVAVMASEENDAPTWIIDD--GPYVVVTDPLDGSRNIDASIPTGTIFGIYNRLEEL-- 206
SG+ V+ASEE D P + + G Y+VV DPLDGS NIDA++ TG+IFGIY+ +E
Sbjct: 148 SGRTGVIASEEEDVPVAVEESYSGNYIVVFDPLDGSSNIDAAVSTGSIFGIYSPNDECLA 207
Query: 207 -----DNLP-TEEKAQLNSLQSGSKLIAAGYVLYSSATILCVSFGSGTQAFTLDHSTGDF 260
NL E++ ++ Q GS L+AAGY +YSS+ I ++ G+G FTLD G+F
Sbjct: 208 DIADDQNLDQVEQRCIVSVCQPGSNLLAAGYCMYSSSVIFVLTIGTGVYVFTLDPMYGEF 267
Query: 261 ILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQYIDTVRQGKGRYPKKYSARYICSLVADL 320
+LT ++IP G+IY+ N+ Y W + L+ Y+D++++ G K YSARYI SLV D
Sbjct: 268 VLTQEKVQIPKAGKIYAFNEGNYALWDDKLKSYMDSLKE-PGPSGKPYSARYIGSLVGDF 326
Query: 321 HRTLMYGGVAMNPRDH------LRLVYEANPLSFLVEQAGGRGSDGKNRILSLQPVKLHQ 374
HRTL+YGG+ PRD LRL+YE P+SF+VEQAGG+GSDG RIL + P ++HQ
Sbjct: 327 HRTLLYGGIYGYPRDQKSKNGKLRLLYECAPMSFIVEQAGGKGSDGHQRILDIMPTEIHQ 386
Query: 375 RLPLFLGSLEDMEELESF 392
R+PL++GS+E++E++E F
Sbjct: 387 RVPLYIGSVEEVEKVEKF 404
>gb|AAD10207.1| fructose 1,6-bisphosphatase [Spinacia oleracea]
gi|7435113|pir||T09085 fructose-bisphosphatase (EC
3.1.3.11) precursor, chloroplast - spinach
gi|3915687|sp|P22418|F16P_SPIOL
Fructose-1,6-bisphosphatase, chloroplast precursor
(D-fructose-1,6-bisphosphate 1-phosphohydrolase)
(FBPase)
Length = 415
Score = 270 bits (691), Expect = 4e-71
Identities = 165/410 (40%), Positives = 239/410 (58%), Gaps = 39/410 (9%)
Query: 3 SAAASPFYQLVPFHPKFQTFQSKTPLTTPYCTPRLASISGSAKMRPLRALGGSSSSAGGD 62
S+ SP Q + F +F + T R SG +R + A+G +++
Sbjct: 23 SSTLSPSQQCITFTKSLHSFPTAT---------RHNVASG---VRCMAAVGEAATETKAR 70
Query: 63 GDDGDGVVTLIEYLGKEGID--LKDDLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVG 120
+ TL +L K+ + + +L ++L I ACK+IA++V I TG+
Sbjct: 71 TRSKYEIETLTGWLLKQEMAGVIDAELTIVLSSISLACKQIASLVQR---AGISNLTGIQ 127
Query: 121 AGGGGGSSGRDAPKPLDIVSNEIILSSLKKSGKVAVMASEENDAPTWIIDD--GPYVVVT 178
G G D K LD+VSNE+ S L+ SG+ ++ASEE D P + + G Y+VV
Sbjct: 128 --GAVNIQGEDQKK-LDVVSNEVFSSCLRSSGRTGIIASEEEDVPVAVEESYSGNYIVVF 184
Query: 179 DPLDGSRNIDASIPTGTIFGIYNRLEEL---------DNLPTEE-KAQLNSLQSGSKLIA 228
DPLDGS NIDA++ TG+IFGIY+ +E L EE + +N Q G L+A
Sbjct: 185 DPLDGSSNIDAAVSTGSIFGIYSPNDECIVDSDHDDESQLSAEEQRCVVNVCQPGDNLLA 244
Query: 229 AGYVLYSSATILCVSFGSGTQAFTLDHSTGDFILTNPSIKIPPRGQIYSVNDARYFDWPE 288
AGY +YSS+ I ++ G G AFTLD G+F+LT+ I+IP G+IYS N+ Y W +
Sbjct: 245 AGYCMYSSSVIFVLTIGKGVYAFTLDPMYGEFVLTSEKIQIPKAGKIYSFNEGNYKMWDD 304
Query: 289 GLRQYIDTVRQGKGRYPKKYSARYICSLVADLHRTLMYGGVAMNPRD------HLRLVYE 342
L++Y+D +++ G K YS+RYI SLV D HRTL+YGG+ PRD LRL+YE
Sbjct: 305 KLKKYMDDLKE-PGESQKPYSSRYIGSLVGDFHRTLLYGGIYGYPRDAKSKNGKLRLLYE 363
Query: 343 ANPLSFLVEQAGGRGSDGKNRILSLQPVKLHQRLPLFLGSLEDMEELESF 392
P+SF+VEQAGG+GSDG RIL +QP ++HQR+PL++GS+E++E+LE +
Sbjct: 364 CAPMSFIVEQAGGKGSDGHQRILDIQPTEIHQRVPLYIGSVEEVEKLEKY 413
>gb|AAK59929.1| fructose-1,6-bisphosphatase [Pisum sativum]
Length = 407
Score = 268 bits (686), Expect = 2e-70
Identities = 168/420 (40%), Positives = 250/420 (59%), Gaps = 47/420 (11%)
Query: 4 AAASPFYQLV---PFHPK----FQ--TFQSKTPLTTPYCTPRLASISGSAKMRPLRALGG 54
AAA+ QL+ P+ P FQ F +K+ L++ R ++GS +R +
Sbjct: 2 AAATASSQLIFSKPYSPSRLCPFQLCVFDAKSVLSSS----RRKHVNGSG----VRCMAV 53
Query: 55 SSSSAGGDGDDGDGVVTLIEYL---GKEGIDLKDDLVVLLDHIQYACKRIAAIVASPFSC 111
+++ G ++TL +L ++GI + +L ++L I ACK+IA++V
Sbjct: 54 KEATSETKKRSGYEIITLTSWLLQQEQKGI-IDAELTIVLSSISMACKQIASLVQR---A 109
Query: 112 SIGKQTGVGAGGGGGSSGRDAPKPLDIVSNEIILSSLKKSGKVAVMASEENDAPTWIIDD 171
+I TG G G D K LD++SNE+ + L+ SG+ ++ASEE D P + +
Sbjct: 110 NISNLTGTQ--GAVNIQGEDQKK-LDVISNEVFSNCLRSSGRTGIIASEEEDVPVAVEES 166
Query: 172 --GPYVVVTDPLDGSRNIDASIPTGTIFGIYNRLEEL----------DNLPTEE-KAQLN 218
G Y+VV DPLDGS N+DA++ TG+IFGIY+ +E + L TEE + +N
Sbjct: 167 YSGNYIVVFDPLDGSSNLDAAVSTGSIFGIYSPNDECLPDFGDDSDDNTLGTEEQRCIVN 226
Query: 219 SLQSGSKLIAAGYVLYSSATILCVSFGSGTQAFTLDHSTGDFILTNPSIKIPPRGQIYSV 278
Q GS L+AAGY +YSS+ I ++ G G FTLD G+F+LT +++IP G+IYS
Sbjct: 227 VCQPGSNLLAAGYCMYSSSVIFVLTIGKGVFVFTLDPLYGEFVLTQENLQIPKSGKIYSF 286
Query: 279 NDARYFDWPEGLRQYIDTVRQGKGRYPKKYSARYICSLVADLHRTLMYGGVAMNPRD--- 335
N+ Y W E L++YID +++ G K YSARYI SLV D HRTL+YGG+ PRD
Sbjct: 287 NEGNYKLWDENLKKYIDDLKE-PGPSGKPYSARYIGSLVGDFHRTLLYGGIYGYPRDKKS 345
Query: 336 ---HLRLVYEANPLSFLVEQAGGRGSDGKNRILSLQPVKLHQRLPLFLGSLEDMEELESF 392
LRL+YE P+SF+VEQAGG+GSDG R+L +QP ++HQR+PL++GS E++E++E +
Sbjct: 346 KNGKLRLLYECAPMSFIVEQAGGKGSDGHQRVLDIQPTEIHQRVPLYIGSTEEVEKVEKY 405
>gb|AAD12243.1| fructose-1,6-bisphosphatase precursor [Brassica napus]
Length = 417
Score = 267 bits (683), Expect = 4e-70
Identities = 148/328 (45%), Positives = 211/328 (64%), Gaps = 24/328 (7%)
Query: 82 DLKDDLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVGAGGGGGSSGRDAPKPLDIVSN 141
++ +L +++ I ACK+IA++V I TGV G G D K LD+VSN
Sbjct: 95 EIDTELTIVMSSIAMACKQIASLVQR---AGISNLTGVQ--GAVNIQGEDQKK-LDVVSN 148
Query: 142 EIILSSLKKSGKVAVMASEENDAPTWIIDD--GPYVVVTDPLDGSRNIDASIPTGTIFGI 199
E+ + L+ SG+ ++ASEE D P + + G Y+VV DPLDGS NIDA++ TG+IFGI
Sbjct: 149 EVFSNCLRSSGRTGIIASEEEDVPVAVEESYSGNYIVVFDPLDGSSNIDAAVSTGSIFGI 208
Query: 200 YNRLEE--------LDNLPTEE-KAQLNSLQSGSKLIAAGYVLYSSATILCVSFGSGTQA 250
Y+ +E + +L +EE + +N Q G+ L+AAGY +YSS+ I ++ G G +
Sbjct: 209 YSPNDECIVDDSDDISSLGSEEQRCIVNVCQPGNNLLAAGYCMYSSSVIFVLTLGKGVFS 268
Query: 251 FTLDHSTGDFILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQYIDTVRQGKGRYPKKYSA 310
FTLD G+F+LT +I+IP G+IYS N+ Y W E L++YID ++ G K YSA
Sbjct: 269 FTLDPMYGEFVLTQENIEIPKAGKIYSFNEGNYQMWDEKLKKYIDDLKD-PGPSGKPYSA 327
Query: 311 RYICSLVADLHRTLMYGGVAMNPRD------HLRLVYEANPLSFLVEQAGGRGSDGKNRI 364
RYI SLV D HRTL+YGG+ PRD LRL+YE P+SF+VEQAGG+GSDG R+
Sbjct: 328 RYIGSLVGDFHRTLLYGGIYGYPRDAKSKNGKLRLLYECAPMSFIVEQAGGKGSDGHQRV 387
Query: 365 LSLQPVKLHQRLPLFLGSLEDMEELESF 392
L +QP ++HQR+PL++GS E++E+LE +
Sbjct: 388 LDIQPTEIHQRVPLYIGSKEEVEKLEKY 415
>pdb|1DCU|D Chain D, Redox Signaling In The Chloroplast: Structure Of Oxidized
Pea Fructose-1,6-Bisphosphate Phosphatase
gi|6730313|pdb|1DCU|C Chain C, Redox Signaling In The
Chloroplast: Structure Of Oxidized Pea
Fructose-1,6-Bisphosphate Phosphatase
gi|6730312|pdb|1DCU|B Chain B, Redox Signaling In The
Chloroplast: Structure Of Oxidized Pea
Fructose-1,6-Bisphosphate Phosphatase
gi|6730311|pdb|1DCU|A Chain A, Redox Signaling In The
Chloroplast: Structure Of Oxidized Pea
Fructose-1,6-Bisphosphate Phosphatase
gi|6730299|pdb|1D9Q|D Chain D, Oxidized Pea
Fructose-1,6-Bisphosphatase Form 1 gi|6730298|pdb|1D9Q|C
Chain C, Oxidized Pea Fructose-1,6-Bisphosphatase Form 1
gi|6730297|pdb|1D9Q|B Chain B, Oxidized Pea
Fructose-1,6-Bisphosphatase Form 1 gi|6730296|pdb|1D9Q|A
Chain A, Oxidized Pea Fructose-1,6-Bisphosphatase Form 1
Length = 357
Score = 266 bits (681), Expect = 6e-70
Identities = 152/349 (43%), Positives = 220/349 (62%), Gaps = 30/349 (8%)
Query: 66 GDGVVTLIEYL---GKEGIDLKDDLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVGAG 122
G ++TL +L ++GI + +L ++L I ACK+IA++V +I TG
Sbjct: 15 GYEIITLTSWLLQQEQKGI-IDAELTIVLSSISMACKQIASLVQR---ANISNLTGTQ-- 68
Query: 123 GGGGSSGRDAPKPLDIVSNEIILSSLKKSGKVAVMASEENDAPTWIIDD--GPYVVVTDP 180
G G D K LD++SNE+ + L+ SG+ ++ASEE D P + + G Y+VV DP
Sbjct: 69 GAVNIQGEDQKK-LDVISNEVFSNCLRSSGRTGIIASEEEDVPVAVEESYSGNYIVVFDP 127
Query: 181 LDGSRNIDASIPTGTIFGIYNRLEEL----------DNLPTEE-KAQLNSLQSGSKLIAA 229
LDGS N+DA++ TG+IFGIY+ +E + L TEE + +N Q GS L+AA
Sbjct: 128 LDGSSNLDAAVSTGSIFGIYSPNDECLPDFGDDSDDNTLGTEEQRCIVNVCQPGSNLLAA 187
Query: 230 GYVLYSSATILCVSFGSGTQAFTLDHSTGDFILTNPSIKIPPRGQIYSVNDARYFDWPEG 289
GY +YSS+ I ++ G G FTLD G+F+LT +++IP G+IYS N+ Y W E
Sbjct: 188 GYCMYSSSVIFVLTIGKGVFVFTLDPLYGEFVLTQENLQIPKSGKIYSFNEGNYKLWDEN 247
Query: 290 LRQYIDTVRQGKGRYPKKYSARYICSLVADLHRTLMYGGVAMNPRD------HLRLVYEA 343
L++YID +++ G K YSARYI SLV D HRTL+YGG+ PRD LRL+YE
Sbjct: 248 LKKYIDDLKE-PGPSGKPYSARYIGSLVGDFHRTLLYGGIYGYPRDKKSKNGKLRLLYEC 306
Query: 344 NPLSFLVEQAGGRGSDGKNRILSLQPVKLHQRLPLFLGSLEDMEELESF 392
P+SF+VEQAGG+GSDG R+L +QP ++HQR+PL++GS E++E++E +
Sbjct: 307 APMSFIVEQAGGKGSDGHQRVLDIQPTEIHQRVPLYIGSTEEVEKVEKY 355
>emb|CAB70979.1| fructose-bisphosphatase precursor [Arabidopsis thaliana]
gi|23397203|gb|AAN31884.1| putative
fructose-bisphosphatase precursor [Arabidopsis thaliana]
gi|23297543|gb|AAN12891.1| putative
fructose-bisphosphatase precursor [Arabidopsis thaliana]
gi|14532620|gb|AAK64038.1| putative
fructose-bisphosphatase precursor [Arabidopsis thaliana]
gi|16226765|gb|AAL16256.1| AT3g54050/F24B22_10
[Arabidopsis thaliana] gi|15232408|ref|NP_190973.1|
fructose-1,6-bisphosphatase, putative /
D-fructose-1,6-bisphosphate 1-phosphohydrolase, putative
/ FBPase, putative [Arabidopsis thaliana]
gi|11262867|pir||T47564 fructose-bisphosphatase
precursor - Arabidopsis thaliana
gi|21431766|sp|P25851|F16P_ARATH
Fructose-1,6-bisphosphatase, chloroplast precursor
(D-fructose-1,6-bisphosphate 1-phosphohydrolase)
(FBPase)
Length = 417
Score = 266 bits (680), Expect = 8e-70
Identities = 146/328 (44%), Positives = 211/328 (63%), Gaps = 24/328 (7%)
Query: 82 DLKDDLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVGAGGGGGSSGRDAPKPLDIVSN 141
++ +L +++ I ACK+IA++V I TGV G G D K LD++SN
Sbjct: 95 EIDAELTIVMSSISLACKQIASLVQR---AGISNLTGVQ--GAVNIQGEDQKK-LDVISN 148
Query: 142 EIILSSLKKSGKVAVMASEENDAPTWIIDD--GPYVVVTDPLDGSRNIDASIPTGTIFGI 199
E+ + L+ SG+ ++ASEE D P + + G YVVV DPLDGS NIDA++ TG+IFGI
Sbjct: 149 EVFSNCLRSSGRTGIIASEEEDVPVAVEESYSGNYVVVFDPLDGSSNIDAAVSTGSIFGI 208
Query: 200 YNRLEEL-----DNLPT----EEKAQLNSLQSGSKLIAAGYVLYSSATILCVSFGSGTQA 250
Y+ +E D++ E++ +N Q G+ L+AAGY +YSS+ I ++ G G +
Sbjct: 209 YSPNDECIVDDSDDISALGSEEQRCIVNVCQPGNNLLAAGYCMYSSSVIFVLTLGKGVFS 268
Query: 251 FTLDHSTGDFILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQYIDTVRQGKGRYPKKYSA 310
FTLD G+F+LT +I+IP G+IYS N+ Y W + L++YID ++ G K YSA
Sbjct: 269 FTLDPMYGEFVLTQENIEIPKAGRIYSFNEGNYQMWDDKLKKYIDDLKD-PGPTGKPYSA 327
Query: 311 RYICSLVADLHRTLMYGGVAMNPRD------HLRLVYEANPLSFLVEQAGGRGSDGKNRI 364
RYI SLV D HRTL+YGG+ PRD LRL+YE P+SF+VEQAGG+GSDG +R+
Sbjct: 328 RYIGSLVGDFHRTLLYGGIYGYPRDAKSKNGKLRLLYECAPMSFIVEQAGGKGSDGHSRV 387
Query: 365 LSLQPVKLHQRLPLFLGSLEDMEELESF 392
L +QP ++HQR+PL++GS E++E+LE +
Sbjct: 388 LDIQPTEIHQRVPLYIGSTEEVEKLEKY 415
>pdb|1DBZ|D Chain D, C153s Mutant Of Pea Fructose-1,6-Bisphosphatase
gi|6730305|pdb|1DBZ|C Chain C, C153s Mutant Of Pea
Fructose-1,6-Bisphosphatase gi|6730304|pdb|1DBZ|B Chain
B, C153s Mutant Of Pea Fructose-1,6-Bisphosphatase
gi|6730303|pdb|1DBZ|A Chain A, C153s Mutant Of Pea
Fructose-1,6-Bisphosphatase
Length = 357
Score = 266 bits (680), Expect = 8e-70
Identities = 152/349 (43%), Positives = 220/349 (62%), Gaps = 30/349 (8%)
Query: 66 GDGVVTLIEYL---GKEGIDLKDDLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVGAG 122
G ++TL +L ++GI + +L ++L I ACK+IA++V +I TG
Sbjct: 15 GYEIITLTSWLLQQEQKGI-IDAELTIVLSSISMACKQIASLVQR---ANISNLTGTQ-- 68
Query: 123 GGGGSSGRDAPKPLDIVSNEIILSSLKKSGKVAVMASEENDAPTWIIDD--GPYVVVTDP 180
G G D K LD++SNE+ + L+ SG+ ++ASEE D P + + G Y+VV DP
Sbjct: 69 GAVNIQGEDQKK-LDVISNEVFSNCLRSSGRTGIIASEEEDVPVAVEESYSGNYIVVFDP 127
Query: 181 LDGSRNIDASIPTGTIFGIYNRLEEL----------DNLPTEE-KAQLNSLQSGSKLIAA 229
LDGS N+DA++ TG+IFGIY+ +E + L TEE + +N Q GS L+AA
Sbjct: 128 LDGSSNLDAAVSTGSIFGIYSPNDESLPDFGDDSDDNTLGTEEQRCIVNVCQPGSNLLAA 187
Query: 230 GYVLYSSATILCVSFGSGTQAFTLDHSTGDFILTNPSIKIPPRGQIYSVNDARYFDWPEG 289
GY +YSS+ I ++ G G FTLD G+F+LT +++IP G+IYS N+ Y W E
Sbjct: 188 GYCMYSSSVIFVLTIGKGVFVFTLDPLYGEFVLTQENLQIPKSGKIYSFNEGNYKLWDEN 247
Query: 290 LRQYIDTVRQGKGRYPKKYSARYICSLVADLHRTLMYGGVAMNPRD------HLRLVYEA 343
L++YID +++ G K YSARYI SLV D HRTL+YGG+ PRD LRL+YE
Sbjct: 248 LKKYIDDLKE-PGPSGKPYSARYIGSLVGDFHRTLLYGGIYGYPRDKKSKNGKLRLLYEC 306
Query: 344 NPLSFLVEQAGGRGSDGKNRILSLQPVKLHQRLPLFLGSLEDMEELESF 392
P+SF+VEQAGG+GSDG R+L +QP ++HQR+PL++GS E++E++E +
Sbjct: 307 APMSFIVEQAGGKGSDGHQRVLDIQPTEIHQRVPLYIGSTEEVEKVEKY 355
>emb|CAA41154.1| fructose-bisphosphatase [Arabidopsis thaliana] gi|99693|pir||S16582
fructose-bisphosphatase (EC 3.1.3.11) precursor,
chloroplast - Arabidopsis thaliana
Length = 417
Score = 266 bits (679), Expect = 1e-69
Identities = 146/328 (44%), Positives = 211/328 (63%), Gaps = 24/328 (7%)
Query: 82 DLKDDLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVGAGGGGGSSGRDAPKPLDIVSN 141
++ +L +++ I ACK+IA++V I TGV G G D K LD++SN
Sbjct: 95 EIDAELTIVMSSISLACKQIASLVQR---AGISNLTGVQ--GAINIQGEDQKK-LDVISN 148
Query: 142 EIILSSLKKSGKVAVMASEENDAPTWIIDD--GPYVVVTDPLDGSRNIDASIPTGTIFGI 199
E+ + L+ SG+ ++ASEE D P + + G YVVV DPLDGS NIDA++ TG+IFGI
Sbjct: 149 EVFSNCLRSSGRTGIIASEEEDVPVAVEESYSGNYVVVFDPLDGSSNIDAAVSTGSIFGI 208
Query: 200 YNRLEEL-----DNLPT----EEKAQLNSLQSGSKLIAAGYVLYSSATILCVSFGSGTQA 250
Y+ +E D++ E++ +N Q G+ L+AAGY +YSS+ I ++ G G +
Sbjct: 209 YSPNDECIVDDSDDISALGSEEQRCIVNVCQPGNNLLAAGYCMYSSSVIFVLTLGKGVFS 268
Query: 251 FTLDHSTGDFILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQYIDTVRQGKGRYPKKYSA 310
FTLD G+F+LT +I+IP G+IYS N+ Y W + L++YID ++ G K YSA
Sbjct: 269 FTLDPMYGEFVLTQENIEIPKAGRIYSFNEGNYQMWDDKLKKYIDDLKD-PGPTGKPYSA 327
Query: 311 RYICSLVADLHRTLMYGGVAMNPRD------HLRLVYEANPLSFLVEQAGGRGSDGKNRI 364
RYI SLV D HRTL+YGG+ PRD LRL+YE P+SF+VEQAGG+GSDG +R+
Sbjct: 328 RYIGSLVGDFHRTLLYGGIYGYPRDAKSKNGKLRLLYECAPMSFIVEQAGGKGSDGHSRV 387
Query: 365 LSLQPVKLHQRLPLFLGSLEDMEELESF 392
L +QP ++HQR+PL++GS E++E+LE +
Sbjct: 388 LDIQPTEIHQRVPLYIGSTEEVEKLEKY 415
>gb|AAB30523.1| fructose-1,6-biphosphatase, FBPase {EC 3.1.3.11} [Pisum
sativum=peas, Lincoln, Peptide Chloroplast, 357 aa]
Length = 357
Score = 266 bits (679), Expect = 1e-69
Identities = 146/326 (44%), Positives = 209/326 (63%), Gaps = 26/326 (7%)
Query: 86 DLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVGAGGGGGSSGRDAPKPLDIVSNEIIL 145
+L ++L I ACK+IA++V +I TG G G D K LD++SNE+
Sbjct: 37 ELTIVLSSISMACKQIASLVQR---ANISNLTGTQ--GAVNIQGEDQKK-LDVISNEVFS 90
Query: 146 SSLKKSGKVAVMASEENDAPTWIIDD--GPYVVVTDPLDGSRNIDASIPTGTIFGIYNRL 203
+ L++SG+ ++ASEE D P + + G Y+VV DPLDGS N+DA++ TG+IFGIY+
Sbjct: 91 NCLRRSGRTGIIASEEEDVPVAVEESYSGNYIVVFDPLDGSSNLDAAVSTGSIFGIYSPN 150
Query: 204 EEL----------DNLPTEE-KAQLNSLQSGSKLIAAGYVLYSSATILCVSFGSGTQAFT 252
+E + L TEE + +N Q GS L+AAGY +YSS+ I ++ G G FT
Sbjct: 151 DECLPDFGDDSDDNTLGTEEQRCIVNVCQPGSNLLAAGYCMYSSSVIFVLTIGKGVFVFT 210
Query: 253 LDHSTGDFILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQYIDTVRQGKGRYPKKYSARY 312
LD G+F+LT +++IP G+IYS N+ Y W E L++YID +++ G K YSARY
Sbjct: 211 LDPLYGEFVLTQENLQIPKSGKIYSFNEGNYKLWDENLKKYIDDLKE-PGPSGKPYSARY 269
Query: 313 ICSLVADLHRTLMYGGVAMNPRD------HLRLVYEANPLSFLVEQAGGRGSDGKNRILS 366
I SLV D HRTL+YGG+ PRD LRL+YE P+SF+VEQAGG+GSDG R+L
Sbjct: 270 IGSLVGDFHRTLLYGGIYGYPRDKKSKNGKLRLLYECAPMSFIVEQAGGKGSDGHQRVLD 329
Query: 367 LQPVKLHQRLPLFLGSLEDMEELESF 392
+QP ++HQR+PL++GS E++E++E +
Sbjct: 330 IQPTEIHQRVPLYIGSTEEVEKVEKY 355
>emb|CAA48719.1| fructose-bisphosphatase [Pisum sativum] gi|322711|pir||S29560
fructose-bisphosphatase (EC 3.1.3.11) - garden pea
(fragment)
Length = 381
Score = 265 bits (677), Expect = 2e-69
Identities = 146/326 (44%), Positives = 208/326 (63%), Gaps = 26/326 (7%)
Query: 86 DLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVGAGGGGGSSGRDAPKPLDIVSNEIIL 145
+L ++L I ACK+IA++V +I TG G G D K LD++SNE+
Sbjct: 61 ELTIVLSSISMACKQIASLVQR---ANISNLTGTQ--GAVNIQGEDQKK-LDVISNEVFS 114
Query: 146 SSLKKSGKVAVMASEENDAPTWIIDD--GPYVVVTDPLDGSRNIDASIPTGTIFGIYNRL 203
+ L+ SG+ ++ASEE D P + + G Y+VV DPLDGS N+DA++ TG+IFGIY+
Sbjct: 115 NCLRSSGRTGIIASEEEDVPVAVEESYSGNYIVVFDPLDGSSNLDAAVSTGSIFGIYSPN 174
Query: 204 EEL----------DNLPTEE-KAQLNSLQSGSKLIAAGYVLYSSATILCVSFGSGTQAFT 252
+E + L TEE + +N Q GS L+AAGY +YSS+ I ++ G G FT
Sbjct: 175 DECLPDFGDDSDDNTLGTEEQRCIVNVCQPGSNLLAAGYCMYSSSVIFVLTIGKGVFVFT 234
Query: 253 LDHSTGDFILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQYIDTVRQGKGRYPKKYSARY 312
LD G+F+LT +++IP G+IYS N+ Y W E L++YID +++ G K YSARY
Sbjct: 235 LDPLYGEFVLTQENLQIPKSGKIYSFNEGNYKLWDENLKKYIDDLKE-PGPSGKPYSARY 293
Query: 313 ICSLVADLHRTLMYGGVAMNPRD------HLRLVYEANPLSFLVEQAGGRGSDGKNRILS 366
I SLV D HRTL+YGG+ PRD LRL+YE P+SF+VEQAGG+GSDG R+L
Sbjct: 294 IGSLVGDFHRTLLYGGIYGYPRDKKSKNGKLRLLYECAPMSFIVEQAGGKGSDGHQRVLD 353
Query: 367 LQPVKLHQRLPLFLGSLEDMEELESF 392
+QP ++HQR+PL++GS E++E++E +
Sbjct: 354 IQPTEIHQRVPLYIGSTEEVEKVEKY 379
>emb|CAA37908.1| fructose-bisphosphatase [Triticum aestivum]
gi|21737|emb|CAA30612.1| unnamed protein product
[Triticum aestivum] gi|67242|pir||PAWTF
fructose-bisphosphatase (EC 3.1.3.11) precursor,
chloroplast - wheat gi|119745|sp|P09195|F16P_WHEAT
FRUCTOSE-1,6-BISPHOSPHATASE, CHLOROPLAST PRECURSOR
(D-FRUCTOSE-1,6-BISPHOSPHATE 1-PHOSPHOHYDROLASE)
(FBPASE)
Length = 409
Score = 265 bits (677), Expect = 2e-69
Identities = 162/406 (39%), Positives = 238/406 (57%), Gaps = 36/406 (8%)
Query: 4 AAASPFYQLVPFHPKFQTFQSKTPLTTPYCTPRLASISGSAKMRPLRALGGSSSSAGGDG 63
AAAS +P P F + +TP +A + ++ P A SS
Sbjct: 19 AAASSLQCRLPRRPGSSLFAGQGQASTPNVRC-MAVVDTASAPAPAAARKRSSYD----- 72
Query: 64 DDGDGVVTLIEYLGK---EGIDLKDDLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVG 120
++TL +L K EG+ + +++ ++L I ACK+IA++V I TGV
Sbjct: 73 -----MITLTTWLLKQEQEGV-IDNEMTIVLSSISTACKQIASLVQR---APISNLTGVQ 123
Query: 121 AGGGGGSSGRDAPKPLDIVSNEIILSSLKKSGKVAVMASEENDAPTWIIDD--GPYVVVT 178
G G D K LD++SNE+ + L+ SG+ V+ASEE D P + + G Y+VV
Sbjct: 124 --GATNVQGEDQKK-LDVISNEVFSNCLRWSGRTGVIASEEEDVPVAVEESYSGNYIVVF 180
Query: 179 DPLDGSRNIDASIPTGTIFGIYNRLEEL---DNLPTEEKAQL---NSLQSGSKLIAAGYV 232
DPLDGS NIDA++ TG+IFGIY+ +E D+ +E Q+ N Q GS L+AAGY
Sbjct: 181 DPLDGSSNIDAAVSTGSIFGIYSPSDECHIGDDATLDEVTQMCIVNVCQPGSNLLAAGYC 240
Query: 233 LYSSATILCVSFGSGTQAFTLDHSTGDFILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQ 292
+YSS+ I ++ G+G FTLD G+F+LT ++IP G+IYS N+ Y W + L++
Sbjct: 241 MYSSSVIFVLTIGTGVYVFTLDPMYGEFVLTQEKVQIPKSGKIYSFNEGNYALWDDKLKK 300
Query: 293 YIDTVRQGKGRYPKKYSARYICSLVADLHRTLMYGGVAMNPRDH------LRLVYEANPL 346
Y+D++++ G K YSARYI SLV D HRT++YGG+ P D LRL+YE P+
Sbjct: 301 YMDSLKE-PGTSGKPYSARYIGSLVGDFHRTMLYGGIYGYPSDQKSKNGKLRLLYECAPM 359
Query: 347 SFLVEQAGGRGSDGKNRILSLQPVKLHQRLPLFLGSLEDMEELESF 392
SF+ EQAGG+GSDG R+L + P +HQR+PL++GS+E++E++E F
Sbjct: 360 SFIAEQAGGKGSDGHQRVLDIMPTAVHQRVPLYVGSVEEVEKVEKF 405
>emb|CAB39759.1| fructose-1,6-bisphosphatase [Pisum sativum]
Length = 357
Score = 265 bits (677), Expect = 2e-69
Identities = 146/326 (44%), Positives = 208/326 (63%), Gaps = 26/326 (7%)
Query: 86 DLVVLLDHIQYACKRIAAIVASPFSCSIGKQTGVGAGGGGGSSGRDAPKPLDIVSNEIIL 145
+L ++L I ACK+IA++V +I TG G G D K LD++SNE+
Sbjct: 37 ELTIVLSSISMACKQIASLVQR---ANISNLTGTQ--GAVNIQGEDQKK-LDVISNEVFS 90
Query: 146 SSLKKSGKVAVMASEENDAPTWIIDD--GPYVVVTDPLDGSRNIDASIPTGTIFGIYNRL 203
+ L+ SG+ ++ASEE D P + + G Y+VV DPLDGS N+DA++ TG+IFGIY+
Sbjct: 91 NCLRSSGRTGIIASEEEDVPVAVEESYSGNYIVVFDPLDGSSNLDAAVSTGSIFGIYSPN 150
Query: 204 EEL----------DNLPTEE-KAQLNSLQSGSKLIAAGYVLYSSATILCVSFGSGTQAFT 252
+E + L TEE + +N Q GS L+AAGY +YSS+ I ++ G G FT
Sbjct: 151 DECLPDFGDDSDDNTLGTEEQRCIVNVCQPGSNLLAAGYCMYSSSVIFVLTIGKGVFVFT 210
Query: 253 LDHSTGDFILTNPSIKIPPRGQIYSVNDARYFDWPEGLRQYIDTVRQGKGRYPKKYSARY 312
LD G+F+LT +++IP G+IYS N+ Y W E L++YID +++ G K YSARY
Sbjct: 211 LDPLYGEFVLTQENLQIPKSGKIYSFNEGNYKLWDENLKKYIDDLKE-PGPSGKPYSARY 269
Query: 313 ICSLVADLHRTLMYGGVAMNPRD------HLRLVYEANPLSFLVEQAGGRGSDGKNRILS 366
I SLV D HRTL+YGG+ PRD LRL+YE P+SF+VEQAGG+GSDG R+L
Sbjct: 270 IGSLVGDFHRTLLYGGIYGYPRDKKSKNGKLRLLYECAPMSFIVEQAGGKGSDGHQRVLD 329
Query: 367 LQPVKLHQRLPLFLGSLEDMEELESF 392
+QP ++HQR+PL++GS E++E++E +
Sbjct: 330 IQPTEIHQRVPLYIGSTEEVEKVEKY 355
>gb|AAD10213.1| fructose-1,6-bisphosphatase [Pisum sativum] gi|7435117|pir||T06408
probable fructose-bisphosphatase (EC 3.1.3.11) precursor
- garden pea chloroplast gi|1094867|prf||2106425A
fructose bisphosphatase gi|2506391|sp|P46275|F16P_PEA
Fructose-1,6-bisphosphatase, chloroplast precursor
(D-fructose-1,6-bisphosphate 1-phosphohydrolase)
(FBPase)
Length = 407
Score = 264 bits (674), Expect = 4e-69
Identities = 166/420 (39%), Positives = 248/420 (58%), Gaps = 47/420 (11%)
Query: 4 AAASPFYQLV---PFHPK----FQ--TFQSKTPLTTPYCTPRLASISGSAKMRPLRALGG 54
AAA+ QL+ P+ P FQ F +K+ L++ R ++GS +R +
Sbjct: 2 AAATASSQLIFSKPYSPSRLCPFQLCVFDAKSVLSSS----RRKHVNGSG----VRCMAV 53
Query: 55 SSSSAGGDGDDGDGVVTLIEYL---GKEGIDLKDDLVVLLDHIQYACKRIAAIVASPFSC 111
+++ G ++TL +L ++GI + +L ++L I ACK+IA++V
Sbjct: 54 KEATSETKKRSGYEIITLTSWLLQQEQKGI-IDAELTIVLSSISMACKQIASLVQR---A 109
Query: 112 SIGKQTGVGAGGGGGSSGRDAPKPLDIVSNEIILSSLKKSGKVAVMASEENDAPTWIIDD 171
+I TG G G D K LD++SNE+ + L+ SG+ ++ASEE D + +
Sbjct: 110 NISNLTGTQ--GAVNIQGEDQKK-LDVISNEVFSNCLRSSGRTGIIASEEEDVAVAVEES 166
Query: 172 --GPYVVVTDPLDGSRNIDASIPTGTIFGIYNRLEEL----------DNLPTEE-KAQLN 218
G Y+VV DPLDGS N+DA++ TG+IFGIY+ +E + L TEE + +N
Sbjct: 167 YSGNYIVVFDPLDGSSNLDAAVSTGSIFGIYSPNDECLPDFGDDSDDNTLGTEEQRCIVN 226
Query: 219 SLQSGSKLIAAGYVLYSSATILCVSFGSGTQAFTLDHSTGDFILTNPSIKIPPRGQIYSV 278
Q GS L+AAGY +YSS+ ++ G G FTLD G+F+LT +++IP G+IYS
Sbjct: 227 VCQPGSNLLAAGYCMYSSSVAFVLTIGKGVFVFTLDPLYGEFVLTQENLQIPKSGEIYSF 286
Query: 279 NDARYFDWPEGLRQYIDTVRQGKGRYPKKYSARYICSLVADLHRTLMYGGVAMNPRD--- 335
N+ Y W E L++YID +++ G K YSARYI SLV D HRTL+YGG+ PRD
Sbjct: 287 NEGNYKLWDENLKKYIDDLKE-PGPSGKPYSARYIGSLVGDFHRTLLYGGIYGYPRDKKS 345
Query: 336 ---HLRLVYEANPLSFLVEQAGGRGSDGKNRILSLQPVKLHQRLPLFLGSLEDMEELESF 392
LRL+YE P+SF+VEQAGG+GSDG R+L +QP ++HQR+PL++GS E++E++E +
Sbjct: 346 KNGKLRLLYECAPMSFIVEQAGGKGSDGHQRVLDIQPTEIHQRVPLYIGSTEEVEKVEKY 405
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.137 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 761,393,399
Number of Sequences: 2540612
Number of extensions: 35504870
Number of successful extensions: 147722
Number of sequences better than 10.0: 335
Number of HSP's better than 10.0 without gapping: 297
Number of HSP's successfully gapped in prelim test: 41
Number of HSP's that attempted gapping in prelim test: 146399
Number of HSP's gapped (non-prelim): 381
length of query: 405
length of database: 863,360,394
effective HSP length: 130
effective length of query: 275
effective length of database: 533,080,834
effective search space: 146597229350
effective search space used: 146597229350
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0100a.12


