

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0100a.11
(245 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
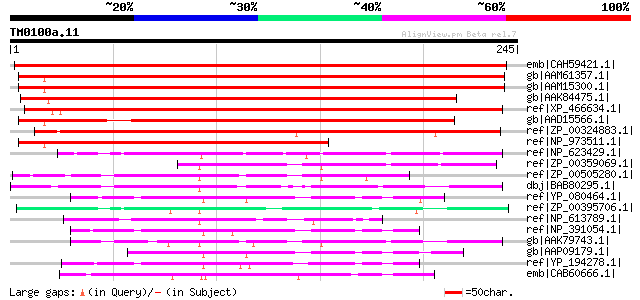
Score E
Sequences producing significant alignments: (bits) Value
emb|CAH59421.1| hypothetical protein [Plantago major] 349 4e-95
gb|AAM61357.1| unknown [Arabidopsis thaliana] 326 3e-88
gb|AAM15300.1| expressed protein [Arabidopsis thaliana] gi|26983... 324 2e-87
gb|AAK84475.1| unknown [Lycopersicon esculentum] 311 8e-84
ref|XP_466634.1| isochorismatase hydrolase-like protein [Oryza s... 307 1e-82
gb|AAD15566.1| unknown protein [Arabidopsis thaliana] gi|2541211... 273 4e-72
ref|ZP_00324883.1| COG1335: Amidases related to nicotinamidase [... 241 1e-62
ref|NP_973511.1| isochorismatase hydrolase family protein [Arabi... 203 4e-51
ref|NP_623429.1| Amidases related to nicotinamidase [Thermoanaer... 86 1e-15
ref|ZP_00359069.1| COG1335: Amidases related to nicotinamidase [... 77 6e-13
ref|ZP_00505280.1| Isochorismatase hydrolase [Clostridium thermo... 74 5e-12
dbj|BAB80295.1| probable pyrazinamidase/nicotinamidase [Clostrid... 69 1e-10
ref|YP_080464.1| Isochorismatase hydrolase [Bacillus licheniform... 63 8e-09
ref|ZP_00395706.1| Isochorismatase hydrolase [Deinococcus geothe... 59 9e-08
ref|NP_613789.1| Amidase related to nicotinamidase [Methanopyrus... 57 6e-07
ref|NP_391054.1| hypothetical protein BSU31760 [Bacillus subtili... 57 6e-07
gb|AAK79743.1| Amidase from nicotinamidase family [Clostridium a... 54 3e-06
gb|AAP09179.1| Pyrazinamidase [Bacillus cereus ATCC 14579] gi|30... 54 4e-06
ref|YP_194278.1| pyrazinamidase-nicotinamidase [Lactobacillus ac... 52 1e-05
emb|CAB60666.1| hypothetical protein [Bradyrhizobium japonicum] ... 51 3e-05
>emb|CAH59421.1| hypothetical protein [Plantago major]
Length = 242
Score = 349 bits (895), Expect = 4e-95
Identities = 159/238 (66%), Positives = 196/238 (81%)
Query: 3 SQIIELLKNEIPLEQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMINE 62
S + LLKNE+P++Q+ + L+ GLVLVDI+NGFCTVG+GNLAP+ ++QI M++E
Sbjct: 4 SDVFSLLKNELPVQQDSLSLSSHLKTGLVLVDIVNGFCTVGSGNLAPQAPDKQIQGMVDE 63
Query: 63 SARLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVTLR 122
S RLA+ FC ++ PV A LDSHHP+ PE PYPPHCIAGT+ES LVPAL WLE E N TLR
Sbjct: 64 SVRLAKEFCRRDWPVYALLDSHHPDIPEPPYPPHCIAGTEESELVPALHWLEKEPNATLR 123
Query: 123 RKECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGFLK 182
RK+C DG+IGS E+DGSN FVDWVK N+I+ +LVVGICTDICVLDFVCS +SA+NR L
Sbjct: 124 RKDCIDGFIGSLEKDGSNTFVDWVKANEIKAVLVVGICTDICVLDFVCSALSARNRRLLT 183
Query: 183 PLENVVVYSHGCATFDVPLEVARNTKGALAHPQEFLHHMGLYMAKERGAKIANEVLFG 240
PLE+V+VYSHGCATFD+P+ VA+N +GALAHPQE +HH+GLYMAK RGAK+ +EV FG
Sbjct: 184 PLEDVIVYSHGCATFDLPVHVAKNIEGALAHPQEIMHHVGLYMAKGRGAKVVSEVSFG 241
>gb|AAM61357.1| unknown [Arabidopsis thaliana]
Length = 244
Score = 326 bits (836), Expect = 3e-88
Identities = 146/236 (61%), Positives = 195/236 (81%), Gaps = 1/236 (0%)
Query: 5 IIELLKNEIPL-EQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMINES 63
I + LK +IP+ E+E ++L D+ GLV+VD++NGFCT+G+GN+AP + N QIS+M+ ES
Sbjct: 7 IFDQLKKQIPVDEEEPLILNRDSSVGLVIVDVVNGFCTIGSGNMAPTKHNEQISKMVEES 66
Query: 64 ARLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVTLRR 123
A+LAR FC++ PV+AF+DSHHP+ PE PYPPHCI GT+ES LVPAL+WLE+E TLRR
Sbjct: 67 AKLAREFCDRKWPVLAFIDSHHPDIPERPYPPHCIIGTEESELVPALKWLESEDCATLRR 126
Query: 124 KECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGFLKP 183
K+C +G++GS E DGSNVFVDWVK+N+I+ ++VVGICTDICV DFV + +SA+N G L P
Sbjct: 127 KDCINGFVGSMESDGSNVFVDWVKENQIKVIVVVGICTDICVFDFVATALSARNHGVLSP 186
Query: 184 LENVVVYSHGCATFDVPLEVARNTKGALAHPQEFLHHMGLYMAKERGAKIANEVLF 239
+E+VVVYS GCATFD+PL VA++ KGA AHPQE +HH+GLYMAK RGA++ +++ F
Sbjct: 187 VEDVVVYSRGCATFDLPLHVAKDIKGAQAHPQELMHHVGLYMAKGRGAQVVSKISF 242
>gb|AAM15300.1| expressed protein [Arabidopsis thaliana] gi|26983880|gb|AAN86192.1|
unknown protein [Arabidopsis thaliana]
gi|18400020|ref|NP_565539.1| isochorismatase hydrolase
family protein [Arabidopsis thaliana]
Length = 244
Score = 324 bits (830), Expect = 2e-87
Identities = 145/236 (61%), Positives = 194/236 (81%), Gaps = 1/236 (0%)
Query: 5 IIELLKNEIPL-EQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMINES 63
I + LK +IP+ E+E ++L D+ GLV+VD++NGFCT+G+GN+AP + N QIS+M+ ES
Sbjct: 7 IFDQLKKQIPVDEEEPLILNRDSSVGLVIVDVVNGFCTIGSGNMAPTKHNEQISKMVEES 66
Query: 64 ARLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVTLRR 123
A+LAR FC++ PV+AF+DSHHP+ PE PYPPHCI GT+ES LVPAL+WLE+E TLRR
Sbjct: 67 AKLAREFCDRKWPVLAFIDSHHPDIPERPYPPHCIIGTEESELVPALKWLESEDCATLRR 126
Query: 124 KECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGFLKP 183
K+C +G++GS E DGSNVFVDWVK+ +I+ ++VVGICTDICV DFV + +SA+N G L P
Sbjct: 127 KDCINGFVGSMESDGSNVFVDWVKEKQIKVIVVVGICTDICVFDFVATALSARNHGVLSP 186
Query: 184 LENVVVYSHGCATFDVPLEVARNTKGALAHPQEFLHHMGLYMAKERGAKIANEVLF 239
+E+VVVYS GCATFD+PL VA++ KGA AHPQE +HH+GLYMAK RGA++ +++ F
Sbjct: 187 VEDVVVYSRGCATFDLPLHVAKDIKGAQAHPQELMHHVGLYMAKGRGAQVVSKISF 242
>gb|AAK84475.1| unknown [Lycopersicon esculentum]
Length = 230
Score = 311 bits (798), Expect = 8e-84
Identities = 145/212 (68%), Positives = 179/212 (84%), Gaps = 1/212 (0%)
Query: 6 IELLKNEIPLEQ-ECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMINESA 64
I+LLK+EIP E+ E ++L D GLVLVDI+NGFCTVGAGNLAP NRQIS M++ES
Sbjct: 10 IDLLKSEIPAEEDEPLLLTGDVNTGLVLVDIVNGFCTVGAGNLAPVTPNRQISAMVDESV 69
Query: 65 RLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVTLRRK 124
+LA++FCEK P+ A DSHHP+ PE P PPHCIAGTDES LVPAL+WLENE NVT+R K
Sbjct: 70 KLAKVFCEKKWPIYALRDSHHPDVPEPPNPPHCIAGTDESELVPALQWLENEPNVTVRCK 129
Query: 125 ECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGFLKPL 184
+C DG++GS E+DGSNVFV+WVK N+I+ +LVVGICTDICVLDFVCS +SA+NRGFL PL
Sbjct: 130 DCIDGFLGSIEKDGSNVFVNWVKANEIKIILVVGICTDICVLDFVCSVLSARNRGFLSPL 189
Query: 185 ENVVVYSHGCATFDVPLEVARNTKGALAHPQE 216
++V+VYS GCAT+D+P+++ARN KGAL HPQ+
Sbjct: 190 KDVIVYSPGCATYDLPVQIARNIKGALPHPQD 221
>ref|XP_466634.1| isochorismatase hydrolase-like protein [Oryza sativa (japonica
cultivar-group)] gi|47497929|dbj|BAD20134.1|
isochorismatase hydrolase-like protein [Oryza sativa
(japonica cultivar-group)]
Length = 252
Score = 307 bits (787), Expect = 1e-82
Identities = 141/238 (59%), Positives = 189/238 (79%), Gaps = 7/238 (2%)
Query: 8 LLKNEIPLEQEC-VVLA------EDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMI 60
+L+ +PL+ + +VLA V GLVLVD+ NGFCTVGAGNLAP N+QI +M+
Sbjct: 14 VLRAAVPLQADADLVLATGGGGERGQVVGLVLVDVSNGFCTVGAGNLAPVTPNKQIEKMV 73
Query: 61 NESARLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVT 120
+E+ARLA++FCE+N PV AFLD+H+P+KPE P+PPHCI G+ E N VPAL WLE + NVT
Sbjct: 74 DEAARLAKVFCERNWPVFAFLDTHYPDKPEPPFPPHCIIGSGEENFVPALEWLEKDPNVT 133
Query: 121 LRRKECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGF 180
+RRK+C DGY+G+ E+DGSNVF DWV K +I+T+LV+GICTD CVLDF S ++A+N G
Sbjct: 134 IRRKDCIDGYLGAFEKDGSNVFSDWVAKFQIKTVLVLGICTDFCVLDFASSALAARNIGR 193
Query: 181 LKPLENVVVYSHGCATFDVPLEVARNTKGALAHPQEFLHHMGLYMAKERGAKIANEVL 238
+ PLE+VV+YS GCAT+++P+EVAR+ +G LAHPQ+ +HHMGLYMAK RGAK+ + ++
Sbjct: 194 VPPLEDVVIYSEGCATYNLPVEVARSMQGTLAHPQDLMHHMGLYMAKSRGAKVVDRII 251
>gb|AAD15566.1| unknown protein [Arabidopsis thaliana] gi|25412119|pir||C84614
hypothetical protein At2g22570 [imported] - Arabidopsis
thaliana
Length = 226
Score = 273 bits (697), Expect = 4e-72
Identities = 126/212 (59%), Positives = 166/212 (77%), Gaps = 12/212 (5%)
Query: 5 IIELLKNEIPL-EQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMINES 63
I + LK +IP+ E+E ++L D+ GLV+VD++NGFCT+G+GN+ M+ ES
Sbjct: 7 IFDQLKKQIPVDEEEPLILNRDSSVGLVIVDVVNGFCTIGSGNM-----------MVEES 55
Query: 64 ARLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVTLRR 123
A+LAR FC++ PV+AF+DSHHP+ PE PYPPHCI GT+ES LVPAL+WLE+E TLRR
Sbjct: 56 AKLAREFCDRKWPVLAFIDSHHPDIPERPYPPHCIIGTEESELVPALKWLESEDCATLRR 115
Query: 124 KECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGFLKP 183
K+C +G++GS E DGSNVFVDWVK+ +I+ ++VVGICTDICV DFV + +SA+N G L P
Sbjct: 116 KDCINGFVGSMESDGSNVFVDWVKEKQIKVIVVVGICTDICVFDFVATALSARNHGVLSP 175
Query: 184 LENVVVYSHGCATFDVPLEVARNTKGALAHPQ 215
+E+VVVYS GCATFD+PL VA++ KGA AHPQ
Sbjct: 176 VEDVVVYSRGCATFDLPLHVAKDIKGAQAHPQ 207
>ref|ZP_00324883.1| COG1335: Amidases related to nicotinamidase [Trichodesmium
erythraeum IMS101]
Length = 247
Score = 241 bits (616), Expect = 1e-62
Identities = 116/228 (50%), Positives = 156/228 (67%), Gaps = 4/228 (1%)
Query: 13 IPLEQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMINESARLARLFCE 72
+P++ + ++A D GL++VD++NGFCTVG G LAP+E N QI+ M++ES RLAR F E
Sbjct: 19 LPIDPQPYIIA-DRPTGLIVVDVLNGFCTVGFGPLAPQEPNEQIATMVSESDRLARTFVE 77
Query: 73 KNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVTLRRKECFDGYIG 132
K PV+AFLD+H P KPE PYPPHC GT E LVP L+WL + TL K+C +G+IG
Sbjct: 78 KGWPVLAFLDTHEPGKPEPPYPPHCEKGTGEEELVPELQWLHDNPLATLVFKDCINGFIG 137
Query: 133 SAEED-GSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGFLKPLENVVVYS 191
S + D NV +DW+ KNK+ L+VVGICTDICV+DFV + +S +N G L++V VY
Sbjct: 138 SIDIDTQGNVLLDWINKNKLEALVVVGICTDICVMDFVVTILSVRNHGLAPTLKDVAVYD 197
Query: 192 HGCATFDVPLEVA--RNTKGALAHPQEFLHHMGLYMAKERGAKIANEV 237
GCATFD+ ++A + HPQ+ HH+GLY ERGA IA+ +
Sbjct: 198 QGCATFDMTAQMAAEKGLPKTAIHPQKISHHVGLYTMAERGAFIASTI 245
>ref|NP_973511.1| isochorismatase hydrolase family protein [Arabidopsis thaliana]
Length = 175
Score = 203 bits (516), Expect = 4e-51
Identities = 90/151 (59%), Positives = 123/151 (80%), Gaps = 1/151 (0%)
Query: 5 IIELLKNEIPL-EQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMINES 63
I + LK +IP+ E+E ++L D+ GLV+VD++NGFCT+G+GN+AP + N QIS+M+ ES
Sbjct: 7 IFDQLKKQIPVDEEEPLILNRDSSVGLVIVDVVNGFCTIGSGNMAPTKHNEQISKMVEES 66
Query: 64 ARLARLFCEKNLPVMAFLDSHHPNKPEDPYPPHCIAGTDESNLVPALRWLENETNVTLRR 123
A+LAR FC++ PV+AF+DSHHP+ PE PYPPHCI GT+ES LVPAL+WLE+E TLRR
Sbjct: 67 AKLAREFCDRKWPVLAFIDSHHPDIPERPYPPHCIIGTEESELVPALKWLESEDCATLRR 126
Query: 124 KECFDGYIGSAEEDGSNVFVDWVKKNKIRTL 154
K+C +G++GS E DGSNVFVDWVK+ +I+ +
Sbjct: 127 KDCINGFVGSMESDGSNVFVDWVKEKQIKVV 157
>ref|NP_623429.1| Amidases related to nicotinamidase [Thermoanaerobacter
tengcongensis MB4] gi|20516857|gb|AAM25033.1| Amidases
related to nicotinamidase [Thermoanaerobacter
tengcongensis MB4]
Length = 248
Score = 85.5 bits (210), Expect = 1e-15
Identities = 70/221 (31%), Positives = 108/221 (48%), Gaps = 19/221 (8%)
Query: 24 EDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMINESARLARLFCEKNLPVMAFLDS 83
ED V+ LV VD++NGFC G P S R ++ +I L + + + FL+
Sbjct: 39 EDKVSVLV-VDMLNGFCKSG-----PLASPR-VAGIIEPIKNLLKACYRMGIKNVFFLND 91
Query: 84 HHPNKPED--PYPPHCIAGTDESNLVPALRWLENETNVTLRRKECFDGYIGSAEEDGSNV 141
HP+ + +PPHC+ GT ES +V L+ + E + K + + G E +G N
Sbjct: 92 AHPSDAVEFGEFPPHCVKGTFESEIVDELKEI-IEGEPVIVEKNSLNVFFG-GELEGGNE 149
Query: 142 F----VDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGFLKPLENVVVYSHGCATF 197
F V+ +K+ K T +VVG CTD+CV S N LK NV+V + T+
Sbjct: 150 FLKKVVEMIKEGK-STFIVVGDCTDLCVYQTAMSIKMIANANNLK--VNVIVPENCVETY 206
Query: 198 DVPLEVARNTKGALAHPQEFLHHMGLYMAKERGAKIANEVL 238
D ++ A++ K + H +H M LY K G ++ E+L
Sbjct: 207 DTSVKTAQSLK-IMPHDGNLIHTMFLYHMKLNGIEVVKELL 246
>ref|ZP_00359069.1| COG1335: Amidases related to nicotinamidase [Chloroflexus
aurantiacus]
Length = 152
Score = 76.6 bits (187), Expect = 6e-13
Identities = 50/156 (32%), Positives = 76/156 (48%), Gaps = 12/156 (7%)
Query: 82 DSHHPNKPE-DPYPPHCIAGTDESNLVPALRWLENETNVTLRRKECFDGYIGSAEEDGSN 140
D+H P PE YPPHC+AGT ES + L L + + K ++G+
Sbjct: 5 DTHDPATPEFAAYPPHCLAGTAESQTIRELAELPFADQIVVIEKNSLSSHLGTR------ 58
Query: 141 VFVDWVKKN-KIRTLLVVGICTDICVLDFVCSTMSAKNRGFLKPLENVVVYSHGCATFDV 199
F W+ ++ ++ T ++VG CTD+CV N LK V+V ++ TFD
Sbjct: 59 -FGSWLAEHPQLDTFVLVGDCTDLCVYSAAMHLRLEANALNLK--RRVIVAANAVDTFDT 115
Query: 200 PLEVARNTKGALAHPQEFLHHMGLYMAKERGAKIAN 235
P+EVARN G AH + H + L+ + G ++ N
Sbjct: 116 PVEVARNL-GIYAHDGDLHHVLFLHHMAQNGVEVMN 150
>ref|ZP_00505280.1| Isochorismatase hydrolase [Clostridium thermocellum ATCC 27405]
gi|67850093|gb|EAM45679.1| Isochorismatase hydrolase
[Clostridium thermocellum ATCC 27405]
Length = 241
Score = 73.6 bits (179), Expect = 5e-12
Identities = 58/199 (29%), Positives = 95/199 (47%), Gaps = 26/199 (13%)
Query: 2 VSQIIELLKNEIPLEQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMIN 61
++ I+++L+ ++P+ + + A+ V LV++D+ NGF GA + + E+I
Sbjct: 34 LASILDMLE-KLPVVKFDEIEADKAV--LVIIDMTNGFAKEGA------LKSDAVKELIP 84
Query: 62 ESARLARLFCEKNLPVMAFLDSHHPNKPE-DPYPPHCIAGTDESNLVPALRWLENETNVT 120
L+ + + + +AF D H PE D YP HC+ GT ES +V ++ N T
Sbjct: 85 RICELSEICDRRKIRKIAFADCHTDESPEFDAYPKHCMKGTAESEIVDEIK---NIGGYT 141
Query: 121 LRRKECFDGYIGSAEEDGSNVFVDWVKKN-KIRTLLVVGICTDICVLDFVCS-----TMS 174
L K +G++ A F W+ +N I T ++ G CTDICV F + M+
Sbjct: 142 LIEKNSTNGFLEEA-------FRKWLLENPDINTFILTGDCTDICVQQFAITLKAYFNMN 194
Query: 175 AKNRGFLKPLENVVVYSHG 193
K + PL V Y G
Sbjct: 195 NKRARVIVPLNAVDTYDLG 213
>dbj|BAB80295.1| probable pyrazinamidase/nicotinamidase [Clostridium perfringens
str. 13] gi|18309571|ref|NP_561505.1| probable
pyrazinamidase/nicotinamidase [Clostridium perfringens
str. 13]
Length = 219
Score = 68.9 bits (167), Expect = 1e-10
Identities = 64/239 (26%), Positives = 109/239 (44%), Gaps = 36/239 (15%)
Query: 1 MVSQIIELLKNEIPLEQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMI 60
M+S I E L++ LE + + + L++VDI GF GA + +I ++I
Sbjct: 14 MISGIYERLQSLESLELKSL---DKNRTMLLIVDINKGFAKAGA------LYSDRIEKLI 64
Query: 61 NESARLARLFCEKNLPVMAFLDSHHPNKPE-DPYPPHCIAGTDESNLVPALRWLENETNV 119
N + LA+ + V AF D H + E YP HC+ TDE LV L N +
Sbjct: 65 NPISNLAKYALNNGIRVKAFTDYHTEDSIELKAYPKHCMKDTDEWELVEEL----NLEGI 120
Query: 120 TLRRKECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRG 179
+ +K +G++ EE+ F+ + K +I +++VG CTDIC+ S + NR
Sbjct: 121 EVIKKNSTNGFL---EEN----FI--LNKEEIDNVIIVGDCTDICIYQLAISLRAEFNR- 170
Query: 180 FLKPLENVVVYSHGCATFDVPLEVARNTKGALAHPQEFLHHMGLYMAKERGAKIANEVL 238
+ + V TFD+P+ H F++++ L + G K+ +++
Sbjct: 171 -VNKDGEIYVPKKLVDTFDMPM-----------HKANFMNYVFLNSMLDNGVKVIEDII 217
>ref|YP_080464.1| Isochorismatase hydrolase [Bacillus licheniformis ATCC 14580]
gi|52004884|gb|AAU24826.1| Isochorismatase hydrolase
[Bacillus licheniformis ATCC 14580]
gi|52787059|ref|YP_092888.1| YueJ [Bacillus
licheniformis ATCC 14580] gi|52349561|gb|AAU42195.1|
YueJ [Bacillus licheniformis DSM 13]
Length = 183
Score = 62.8 bits (151), Expect = 8e-09
Identities = 59/194 (30%), Positives = 88/194 (44%), Gaps = 32/194 (16%)
Query: 30 LVLVDIINGFCTVGAGNLAPRESNRQISEMINESARLARLFCEKNLPVMAFLDSHHPNKP 89
L+ +D N F G L E R+I + + ARLA F + V+ +DSH
Sbjct: 5 LICIDYTNDFVAAD-GKLTCGEPGRKIEDAV---ARLADTFIQNGDYVVFAVDSHEAGDT 60
Query: 90 EDP----YPPHCIAGTDESNLVPALRWL----ENETNVTLRRKECFDGYIGSAEEDGSNV 141
P +PPH I GT +L L L E++ NV K + + G+ E
Sbjct: 61 LHPETRLFPPHNIKGTSGQDLYGKLEKLYRKHEHDQNVYYMEKTRYSAFAGTNLELK--- 117
Query: 142 FVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGFLKPLENVVVYSHGCATF---- 197
+++ I+ L +VG+CTDICVL + + A N+GF N+V++ + A+F
Sbjct: 118 ----LRERDIQELHLVGVCTDICVLH---TAVDAYNKGF-----NLVIHQNAVASFNPDG 165
Query: 198 -DVPLEVARNTKGA 210
D L NT GA
Sbjct: 166 HDWALSHFANTLGA 179
>ref|ZP_00395706.1| Isochorismatase hydrolase [Deinococcus geothermalis DSM 11300]
gi|66782607|gb|EAL83570.1| Isochorismatase hydrolase
[Deinococcus geothermalis DSM 11300]
Length = 255
Score = 59.3 bits (142), Expect = 9e-08
Identities = 60/243 (24%), Positives = 97/243 (39%), Gaps = 32/243 (13%)
Query: 4 QIIELLKNEIPLEQECVVLAEDTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMINES 63
Q +LL+ + E V+ +V VD++ GF G P S R ++E+I
Sbjct: 38 QQADLLQAWMSALPEWVLTTRPDRVAVVCVDLVEGFTREG-----PLASPR-VAEIIPRI 91
Query: 64 ARLARLFCEKNLP---VMAFLDSHHPNKPE-DPYPPHCIAGTDESNLVPALRWLENETNV 119
+L R ++ +P V+ DSH + E YPPHC+AGT E+ V LR L
Sbjct: 92 VQLLRRLLDRGVPAENVVLVQDSHPLDAKEFQAYPPHCVAGTAEAQAVAELRALPEFARF 151
Query: 120 TLRRKECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRG 179
+K S S F W+ + + ++ +G TD+C+ ++ G
Sbjct: 152 QHFQK-------NSIASHTSPAFQAWLAQAEFDVVIALGDVTDLCLYTLALHLVTF---G 201
Query: 180 FLKPLENVVVYSHGCA-TFDVPLEVARNTKGALAHPQEFLHHMGLYMAKERGAKIANEVL 238
+ VV C T+D P HP + H + L+ G ++ +
Sbjct: 202 MANQQDWTVVVPEECVQTWDAP-----------DHPGDLYHALFLHQLARNGVRVVRALS 250
Query: 239 FGA 241
GA
Sbjct: 251 VGA 253
>ref|NP_613789.1| Amidase related to nicotinamidase [Methanopyrus kandleri AV19]
gi|19886895|gb|AAM01719.1| Amidase related to
nicotinamidase [Methanopyrus kandleri AV19]
Length = 177
Score = 56.6 bits (135), Expect = 6e-07
Identities = 45/156 (28%), Positives = 78/156 (49%), Gaps = 21/156 (13%)
Query: 27 VNGLVLVDIINGFCTVGAGNLAPRESNRQISEMINESARLARLFCEKNLPVMAFLDSHHP 86
+ L++VD+I F GA P+ ++ ARLA F E+ V+ D H+P
Sbjct: 1 MRALLIVDMIRDFVEEGAPLEVPKARR-----LVPRIARLADEFRERGDLVVHVWDEHYP 55
Query: 87 NKPE-DPYPPHCIAGTDESNLVPALRWLENETNVTLRRKECFDGYIGSAEEDGSNVFVDW 145
+ PE + H +AGT+ + V L+ + + V RK + G+ G++ +D+
Sbjct: 56 DDPEFKVWGEHAVAGTEGAEPVEELKPEDGDLVV---RKRKYSGFYGTS--------LDY 104
Query: 146 -VKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGF 180
++ ++ + + G+CTDICVL F C+ A RG+
Sbjct: 105 DLRSRNVKEIYLTGVCTDICVL-FTCA--DALMRGY 137
>ref|NP_391054.1| hypothetical protein BSU31760 [Bacillus subtilis subsp. subtilis
str. 168] gi|2635671|emb|CAB15164.1| yueJ [Bacillus
subtilis subsp. subtilis str. 168]
gi|7429497|pir||C70008 pyrazinamidase/nicotinamidase
homolog yueJ - Bacillus subtilis
Length = 183
Score = 56.6 bits (135), Expect = 6e-07
Identities = 51/177 (28%), Positives = 78/177 (43%), Gaps = 27/177 (15%)
Query: 30 LVLVDIINGFCTVGAGNLAPRESNRQISEMINESARLARLFCEKNLPVMAFLDSHHPNKP 89
L+ +D N F G L E R I E I L + F V+ +DSH
Sbjct: 5 LICIDYTNDF-VASDGKLTCGEPGRMIEEAI---VNLTKEFITNGDYVVLAVDSHDEGDQ 60
Query: 90 EDP----YPPHCIAGTDESNL----VPALRWLENETNVTLRRKECFDGYIGSAEEDGSNV 141
P +PPH I GT+ +L +P + E+E NV K + + G+ E
Sbjct: 61 YHPETRLFPPHNIKGTEGKDLYGKLLPLYQKHEHEPNVYYMEKTRYSAFAGTDLELK--- 117
Query: 142 FVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGFLKPLENVVVYSHGCATFD 198
+++ +I L + G+CTDICVL + + A N+GF +VV+ A+F+
Sbjct: 118 ----LRERQIGELHLAGVCTDICVLH---TAVDAYNKGF-----RIVVHKQAVASFN 162
>gb|AAK79743.1| Amidase from nicotinamidase family [Clostridium acetobutylicum ATCC
824] gi|15895054|ref|NP_348403.1| Amidase from
nicotinamidase family [Clostridium acetobutylicum ATCC
824] gi|25498245|pir||D97119 amidase from nicotinamidase
family [imported] - Clostridium acetobutylicum
Length = 216
Score = 54.3 bits (129), Expect = 3e-06
Identities = 54/218 (24%), Positives = 98/218 (44%), Gaps = 42/218 (19%)
Query: 30 LVLVDIINGFCTVGAGNLAPRESNRQISEMINESARLARLFCEKNL--PVMAFLDSHHPN 87
+V+VD++NGF GA S+ ++ +++E R+ EK L + FLD H N
Sbjct: 32 IVIVDMVNGFVHEGA------LSSPRVEGIVDEIVRIN----EKTLGNKKIFFLDEHTNN 81
Query: 88 KPE-DPYPPHCIAGTDESNLVPALRWLENE----TNVTLRRKECFDGYIGSAEEDGSNVF 142
E Y HC+ G+ E+ L+P L+ NE +N + K +G+ F
Sbjct: 82 STEFKSYAKHCLEGSLEAELIPELK---NEALLDSNTVMIPKNSVNGFHAPG-------F 131
Query: 143 VDWVKKN--KIRTLLVVGICTDICVLDFVCSTMSAKNRGFLKPLENVVVYSHGCATFDVP 200
W+++N +I ++ G DICV +F + + N+ + + +++ S+ TFD
Sbjct: 132 KKWLEENESQIENYIICGCEVDICVSNFANTLKTYFNQKNMD--KRIIIPSNAVETFDFG 189
Query: 201 LEVARNTKGALAHPQEFLHHMGLYMAKERGAKIANEVL 238
H + + + L+ + G +I + VL
Sbjct: 190 -----------THDGDLMKIISLWEMQSNGIEIVDRVL 216
>gb|AAP09179.1| Pyrazinamidase [Bacillus cereus ATCC 14579]
gi|30020347|ref|NP_831978.1| Pyrazinamidase [Bacillus
cereus ATCC 14579]
Length = 184
Score = 53.9 bits (128), Expect = 4e-06
Identities = 48/170 (28%), Positives = 77/170 (45%), Gaps = 28/170 (16%)
Query: 58 EMINESARLARLFCEKNLPVMAFLDSHHPNKPEDP----YPPHCIAGTDESNLVPALRWL 113
E+ E + + + E V+ +D H N P +PPH IAGT +L L+ +
Sbjct: 31 EIEKEIVHITKQYIENGDYVVFAIDKHEENDVYHPESKLFPPHNIAGTSGRDLFGELQDV 90
Query: 114 ----ENETNVTLRRKECFDGYIGSAEEDGSNVFVDWVKKNKIRTLLVVGICTDICVLDFV 169
+N NV K + + G+ E +++ I + +VG+CTDICVL
Sbjct: 91 YETYKNAENVYYMDKTRYSAFAGTDLEMK-------LRERGIEEVHLVGVCTDICVLH-- 141
Query: 170 CSTMSAKNRGFLKPLENVVVYSHGCATFDVPLEVARNTKGALAHPQEFLH 219
+ + A N+GF +VVY A+F+ A+ + AL H + LH
Sbjct: 142 -TAVDAYNKGF-----KIVVYEKAVASFN-----AQGHEYALGHFKSCLH 180
>ref|YP_194278.1| pyrazinamidase-nicotinamidase [Lactobacillus acidophilus NCFM]
gi|58255010|gb|AAV43247.1| pyrazinamidase-nicotinamidase
[Lactobacillus acidophilus NCFM]
Length = 184
Score = 52.4 bits (124), Expect = 1e-05
Identities = 50/181 (27%), Positives = 85/181 (46%), Gaps = 27/181 (14%)
Query: 26 TVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMINESARLARLFCEKNLPVMAFLDSHH 85
T L+++D N F G L + +++ + I A LA F +++ V+ D H
Sbjct: 2 TNEALLIIDYTNDFVD-DKGALTCGKPAQELDDTI---ANLADKFLKEDKWVIFPTDKHF 57
Query: 86 PNKPEDP----YPPHCIAGTDESNLVPAL-RWLE---NETNVTLRRKECFDGYIGSAEED 137
+ P P +PPH + T L + +W E + +V L K + + G++ +
Sbjct: 58 KDNPYHPETKLFPPHNLPNTWGRELYGKVGKWYEAHKDNNHVILMDKTRYSAFAGTSLDL 117
Query: 138 GSNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGFLKPLENVVVYSHGCATF 197
+++ KI TL + G+CTDICVL + M A N + N+VV+ +G A+F
Sbjct: 118 -------LLRERKIDTLHLTGVCTDICVLH---TAMDAYNHCY-----NLVVHENGVASF 162
Query: 198 D 198
D
Sbjct: 163 D 163
>emb|CAB60666.1| hypothetical protein [Bradyrhizobium japonicum]
gi|27375788|ref|NP_767317.1| bifunctional
pyrazinamidase/nicotinamidase [Bradyrhizobium japonicum
USDA 110] gi|27348926|dbj|BAC45942.1| bifunctional
pyrazinamidase/nicotinamidase [Bradyrhizobium japonicum
USDA 110]
Length = 240
Score = 51.2 bits (121), Expect = 3e-05
Identities = 48/196 (24%), Positives = 81/196 (40%), Gaps = 25/196 (12%)
Query: 25 DTVNGLVLVDIINGFCTVGAGNLAPRESNRQISEMINESARLARLFCEKNLPV---MAFL 81
D + L+++D+ N C + G+LA +E + + + S + + ++ ++F
Sbjct: 33 DDASALLVIDVQN--CFLPGGSLAVKEGEQVVPVINKISKAFSNVVMTQDWHTPGHVSFA 90
Query: 82 DSHHPNKPED----PY------PPHCIAGTDESNLVPALRWLENETNVTLRRKECFDGYI 131
H KP + PY P HC+ GTD ++L L E + + D Y
Sbjct: 91 SVHSGKKPFETIDLPYGKQVLWPDHCVQGTDGASLSKDLAIPHAELIIRKGFHKDVDSYS 150
Query: 132 GSAEEDG--SNVFVDWVKKNKIRTLLVVGICTDICVLDFVCSTMSAKNRGFLKPLENVVV 189
E DG S ++K KI+ + V G+ TD CV + + A+ GF V V
Sbjct: 151 AFLEADGKTSTGLAGYLKGRKIKRVFVAGLATDFCV---AWTALDARKAGF-----EVYV 202
Query: 190 YSHGCATFDVPLEVAR 205
C D +A+
Sbjct: 203 VEDACRGIDTQGSLAK 218
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.138 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 419,187,342
Number of Sequences: 2540612
Number of extensions: 17263444
Number of successful extensions: 36673
Number of sequences better than 10.0: 159
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 150
Number of HSP's that attempted gapping in prelim test: 36496
Number of HSP's gapped (non-prelim): 164
length of query: 245
length of database: 863,360,394
effective HSP length: 125
effective length of query: 120
effective length of database: 545,783,894
effective search space: 65494067280
effective search space used: 65494067280
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0100a.11


