

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0094b.3
(337 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
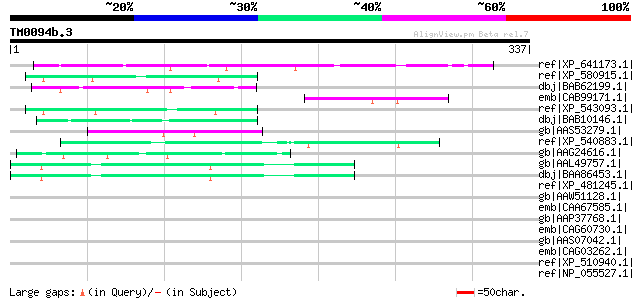
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_641173.1| dynactin 150 kDa subunit [Dictyostelium discoid... 50 1e-04
ref|XP_580915.1| PREDICTED: similar to YTH domain protein 1 (Der... 50 1e-04
dbj|BAB62199.1| hypothetical protein [Macaca fascicularis] 49 2e-04
emb|CAB99171.1| related to transport protein USO1 [Neurospora cr... 48 4e-04
ref|XP_543093.1| PREDICTED: similar to Dermatomyositis associate... 48 5e-04
dbj|BAB10146.1| unnamed protein product [Arabidopsis thaliana] g... 48 5e-04
gb|AAS53279.1| AFL095Wp [Ashbya gossypii ATCC 10895] gi|45198426... 48 5e-04
ref|XP_540883.1| PREDICTED: similar to Ribosomal protein S6 kina... 47 8e-04
gb|AAG24616.1| arabinogalactan protein [Nicotiana alata] 47 8e-04
gb|AAL49757.1| cask-interacting protein 2 [Homo sapiens] gi|4521... 47 8e-04
dbj|BAA86453.1| KIAA1139 protein [Homo sapiens] 47 8e-04
ref|XP_481245.1| putative diaphanous 1 [Oryza sativa (japonica c... 47 0.001
gb|AAW51128.1| minus agglutinin [Chlamydomonas incerta] 47 0.001
emb|CAA67585.1| arabinogalactan [Lycopersicon esculentum] gi|143... 46 0.002
gb|AAP37768.1| At3g24600 [Arabidopsis thaliana] gi|13877617|gb|A... 46 0.002
emb|CAG60730.1| unnamed protein product [Candida glabrata CBS138... 46 0.002
gb|AAS07042.1| minus agglutinin [Chlamydomonas reinhardtii] 46 0.002
emb|CAG03262.1| unnamed protein product [Tetraodon nigroviridis] 46 0.002
ref|XP_510940.1| PREDICTED: similar to KIAA0339 protein [Pan tro... 45 0.002
ref|NP_055527.1| hypothetical protein LOC9739 [Homo sapiens] 45 0.002
>ref|XP_641173.1| dynactin 150 kDa subunit [Dictyostelium discoideum]
gi|60469201|gb|EAL67196.1| dynactin 150 kDa subunit
[Dictyostelium discoideum]
Length = 1539
Score = 50.1 bits (118), Expect = 1e-04
Identities = 67/312 (21%), Positives = 133/312 (42%), Gaps = 29/312 (9%)
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTP 75
+SP+++ P++ P P TT ++ V P P+ S PS ++ T
Sbjct: 204 TSPLTSPPPEKKSEPSP-PTTTTISPPPPTVVEPTPIEQPQPTPSKISRPPSSASSRPT- 261
Query: 76 DAPRPVTLAEGQRSPSSIRPPIPSPVKV-----TLGPSAPNSPGVGGEGENLSAAQPTGP 130
P+P L E +P+ P P+P T P+ +P + + P P
Sbjct: 262 GIPKPSGLKEPTPTPTPTPTPTPTPTPTPTPTPTPTPTPTPTPIITPNPTITTTTTPI-P 320
Query: 131 SSSSPPYPF---SFLLSHPYLEKLKGAPSQNPQDVAQEAVLNLLRAGCLFAQVAWQV--- 184
+ S+P P S L+ P +K + Q + +E + +LL ++ V
Sbjct: 321 TPSTPLPPSTSNSITLTPPSTQKKQSKIIQGEDEEEEEEINDLLNKSLATERMNQAVQEN 380
Query: 185 --RPLTGIVGLRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEA 242
+ +T I L+++ Q ++++ + + ++ +A+KDK+I + ++++ + + +
Sbjct: 381 IDKMMTTINDLKEDKNRSQEEINSLKDRL---KEAVASKDKQIKENEKQIKENEKQIQQL 437
Query: 243 QSDLTFKCKLLADYEAQAATLKEKIRSEAAHAAAKERVRGSLIAKAQVKHLHPTINLDPL 302
+ + K K LK + R+E + E+ + SL+ Q+ +L+ NL+ L
Sbjct: 438 EKTIAIKEK------ENENLLKTQHRNEDKEKSEFEKEKQSLL--EQISNLNE--NLELL 487
Query: 303 GAFKRVTPAGLE 314
K VT LE
Sbjct: 488 TLDKEVTEEKLE 499
>ref|XP_580915.1| PREDICTED: similar to YTH domain protein 1 (Dermatomyositis
associated with cancer putative autoantigen-1 homolog)
(DACA-1 homolog), partial [Bos taurus]
Length = 533
Score = 49.7 bits (117), Expect = 1e-04
Identities = 47/168 (27%), Positives = 66/168 (38%), Gaps = 23/168 (13%)
Query: 11 GGKRQSSPVS---ACSPKRSKVGEPDPQTTPLTGTG-GGFVAPDPV-------VTDSACD 59
GG PVS + + SK +P P+ +G GG + P P+ D+
Sbjct: 202 GGTNVPLPVSKPTSWAAIASKPAKPQPKMKAKSGPAIGGALPPPPIKHNMDIGTWDNKGP 261
Query: 60 PLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEG 119
+ P +V AS P+P P +PP+P+P + T P + V
Sbjct: 262 VPKAPAPQQVPASSAAPQPQPAM------QPLPAQPPLPAPPQHTGPQQLPQTRWVAPRS 315
Query: 120 ENLSAAQPTGPSSSS------PPYPFSFLLSHPYLEKLKGAPSQNPQD 161
N + Q G S S P SHP LEKLK A S NP++
Sbjct: 316 RNAAFGQTGGLGSDSNSSGGAQPSTTPSAESHPVLEKLKAAHSYNPKE 363
>dbj|BAB62199.1| hypothetical protein [Macaca fascicularis]
Length = 599
Score = 48.9 bits (115), Expect = 2e-04
Identities = 52/175 (29%), Positives = 77/175 (43%), Gaps = 37/175 (21%)
Query: 15 QSSPVSACSPKRSKVGE----PDPQTTPLTGTGGGFVAPDPVVTDSACDP-LASDHPSEV 69
+ P+ A S + K+G P+ TPLT P VV+ P L S PS+
Sbjct: 80 EPKPIPALSMETQKLGSVLSPESPKPTPLTPLEPQ--KPGSVVSPELQTPSLPSPEPSKP 137
Query: 70 NASQTPDAPRPVTLAEGQR-------SPSSIRPPIPSPVKV---------------TLGP 107
+ +P+ P+PV + E Q+ P + P P PVK TLGP
Sbjct: 138 ASVSSPEPPKPVPVCESQKPAPVPSPEPQKLAPVSPEPVKATLPNPKPQKHSHFPETLGP 197
Query: 108 SAPNSPGVGGEGENLSAA-QPTGPS-SSSPPYPFSFLLSHPYLEKLKGAPSQNPQ 160
+ +SP E L+A+ +P GPS ++SP S + P E K +PS +P+
Sbjct: 198 PSASSP----ESPVLAASPEPWGPSPAASPESRKSARTTSP--EPRKPSPSGSPE 246
>emb|CAB99171.1| related to transport protein USO1 [Neurospora crassa]
gi|32415207|ref|XP_328083.1| hypothetical protein (
(AL390189) related to transport protein USO1 [Neurospora
crassa] ) gi|28917322|gb|EAA27030.1| hypothetical
protein ( (AL390189) related to transport protein USO1
[Neurospora crassa] )
Length = 1171
Score = 48.1 bits (113), Expect = 4e-04
Identities = 38/105 (36%), Positives = 54/105 (51%), Gaps = 11/105 (10%)
Query: 192 GLRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEG----QNEALAEAQSDLT 247
G+ +E EE +R L KE VA EK L K++E+ ++E LEG + EA+ +A ++
Sbjct: 952 GVIEEREEAERLLKEKEDAVAEVEKALKDKEREMVSMKERLEGMEKEKEEAVQKAVEEIR 1011
Query: 248 FKC-------KLLADYEAQAATLKEKIRSEAAHAAAKERVRGSLI 285
+ K A+ EA AAT EK + A AAAK S I
Sbjct: 1012 KEVETAKNNDKAEAEAEADAATGAEKAAAIEAEAAAKAAAAASEI 1056
>ref|XP_543093.1| PREDICTED: similar to Dermatomyositis associated with cancer
putative autoantigen-1 homolog (DACA-1 homolog) [Canis
familiaris]
Length = 1004
Score = 47.8 bits (112), Expect = 5e-04
Identities = 45/168 (26%), Positives = 67/168 (39%), Gaps = 23/168 (13%)
Query: 11 GGKRQSSPVS---ACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVT--------DSACD 59
GG + PVS + + SK +P P+ +G G P P + D+
Sbjct: 301 GGTNVNMPVSKPTSWAAIASKPAKPQPKMKAKSGPVIGAALPPPPIKHNMDIGTWDNQGP 360
Query: 60 PLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEG 119
+ P + A Q+ P+PV + P +P PSP + P + V
Sbjct: 361 VPKAPAPQQAPAPQSAPQPQPVVQPLPTQPPPLAQPQYPSPQQ------PPQTRWVAPRN 414
Query: 120 ENLSAAQPTGPSS------SSPPYPFSFLLSHPYLEKLKGAPSQNPQD 161
N + Q G SS ++ P + SHP LEKLK A S NP++
Sbjct: 415 RNAAFGQSGGTSSDGNSPGNTQPSSAPTVESHPVLEKLKAAHSYNPKE 462
>dbj|BAB10146.1| unnamed protein product [Arabidopsis thaliana]
gi|21360463|gb|AAM47347.1| AT5g38560/MBB18_10
[Arabidopsis thaliana] gi|18700153|gb|AAL77688.1|
AT5g38560/MBB18_10 [Arabidopsis thaliana]
gi|15240947|ref|NP_198672.1| protein kinase family
protein [Arabidopsis thaliana]
gi|15983497|gb|AAL11616.1| AT5g38560/MBB18_10
[Arabidopsis thaliana]
Length = 681
Score = 47.8 bits (112), Expect = 5e-04
Identities = 42/145 (28%), Positives = 56/145 (37%), Gaps = 7/145 (4%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
PV + SP V P P ++P + +P P V S P+ P + TP A
Sbjct: 50 PVVSSSPPPPVVSSPPPSSSP-PPSPPVITSPPPTVASSPPPPVVIASPPPSTPATTPPA 108
Query: 78 PRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPS-SSSPP 136
P P T++ +S PP P+ T P SP GE + P+ P S S P
Sbjct: 109 P-PQTVSPPPPPDASPSPPAPT----TTNPPPKPSPSPPGETPSPPGETPSPPKPSPSTP 163
Query: 137 YPFSFLLSHPYLEKLKGAPSQNPQD 161
P + P PS NP D
Sbjct: 164 TPTTTTSPPPPPATSASPPSSNPTD 188
Score = 34.3 bits (77), Expect = 5.4
Identities = 36/142 (25%), Positives = 50/142 (34%), Gaps = 15/142 (10%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
PV SP S P TTP P P + S P ++ P + + S +
Sbjct: 91 PVVIASPPPST-----PATTPPAPPQTVSPPPPPDASPSPPAPTTTNPPPKPSPSPPGET 145
Query: 78 PRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSS-SSPP 136
P P PS P P+P T P P + ++ PT PS+ + PP
Sbjct: 146 PSPPGETPSPPKPS---PSTPTPTTTTSPPPPP------ATSASPPSSNPTDPSTLAPPP 196
Query: 137 YPFSFLLSHPYLEKLKGAPSQN 158
P + + K G S N
Sbjct: 197 TPLPVVPREKPIAKPTGPASNN 218
>gb|AAS53279.1| AFL095Wp [Ashbya gossypii ATCC 10895] gi|45198426|ref|NP_985455.1|
AFL095Wp [Eremothecium gossypii]
Length = 2535
Score = 47.8 bits (112), Expect = 5e-04
Identities = 33/120 (27%), Positives = 51/120 (42%), Gaps = 6/120 (5%)
Query: 51 PVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIP----SPVKVTLG 106
P T C P A+ P++ ++S + P P T Q S S+ +P P P + +
Sbjct: 2228 PPATSQPCPPPATSQPAQSSSSASQPCPPPATSQPAQSSSSASQPCPPPATSQPAQSSSS 2287
Query: 107 PSAPNSPGVGGE--GENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQ 164
S P P + + SA+QP PS++S P S S P P+Q+ +Q
Sbjct: 2288 ASQPCPPAATSQPAQSSSSASQPCPPSATSQPAQSSSSASQPCPPSATSQPAQSSSSASQ 2347
Score = 40.8 bits (94), Expect = 0.058
Identities = 28/95 (29%), Positives = 44/95 (45%), Gaps = 6/95 (6%)
Query: 58 CDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPS----PVKVTLGPSAPNSP 113
C P A+ P++ ++S + P P T Q S S+ +P P+ P + + S P P
Sbjct: 2254 CPPPATSQPAQSSSSASQPCPPPATSQPAQSSSSASQPCPPAATSQPAQSSSSASQPCPP 2313
Query: 114 GVGGEG--ENLSAAQPTGPSSSSPPYPFSFLLSHP 146
+ + SA+QP PS++S P S S P
Sbjct: 2314 SATSQPAQSSSSASQPCPPSATSQPAQSSSSASQP 2348
>ref|XP_540883.1| PREDICTED: similar to Ribosomal protein S6 kinase alpha 4 (Nuclear
mitogen-and stress-activated protein kinase-2) (90 kDa
ribosomal protein S6 kinase 4) (Ribosomal protein kinase
B) (RSKB) [Canis familiaris]
Length = 2376
Score = 47.0 bits (110), Expect = 8e-04
Identities = 66/265 (24%), Positives = 102/265 (37%), Gaps = 38/265 (14%)
Query: 34 PQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSI 93
PQT+ LT G ++A P+ V + Q P L G+ S
Sbjct: 562 PQTSDLTLQESGPTVATQKCPENAGHGALLQIPTSVVSLQGPGLEIQAELLGGEAGESG- 620
Query: 94 RPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKG 153
K GP++ P L + G P LE + G
Sbjct: 621 -------PKAREGPASKPGPPEVSLRVELEEQETPGQGLELPKGQRDARGDEQKLEGMIG 673
Query: 154 APSQ-NPQDVAQEAVLNLLRAGCLFAQVAWQVRPLTGIVG---------LRQEVEELQRD 203
P+Q NPQ ++ G L AQ WQ P++G + LR+EV +L+R+
Sbjct: 674 DPAQQNPQQESE---------GALEAQT-WQ-GPISGEISSGGVPEQEALREEVAQLRRE 722
Query: 204 LDAKEITVASQEKFLAAKDKEIADLREELEGQNEALAEAQSDLTFKCK---------LLA 254
+A + +Q + L A+ E A L EEL + AEA ++ + + A
Sbjct: 723 AEALRAELEAQARRLEARGMEAARLSEELAQARKTEAEAHREVEAQAREQARLREAVEAA 782
Query: 255 DYEAQAATLKEKIRSEAAHAAAKER 279
E +AA+ + + +EA AA +ER
Sbjct: 783 GRELEAASQEREALAEALAAAGRER 807
>gb|AAG24616.1| arabinogalactan protein [Nicotiana alata]
Length = 228
Score = 47.0 bits (110), Expect = 8e-04
Identities = 57/207 (27%), Positives = 79/207 (37%), Gaps = 38/207 (18%)
Query: 5 PRVAEVGGKRQSSPVSACSPKRSKVGEPD------------------PQTTPLTGTGGGF 46
P A VG K + P +A P + K P P T P T
Sbjct: 28 PAKAPVGAKATTPPAAA--PTKPKTPAPATAPASAPPTAVSTPPAAAPATAPTTPVVTSP 85
Query: 47 VAPDPVVTDSACDPLA---SDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPV-- 101
V+ P T ++ P A S P V Q+P AP PV P++ PP P PV
Sbjct: 86 VSAPPAKTPASSPPAAVPVSSPPLAVTPVQSPPAPAPVAAT----PPAASAPPAPVPVSA 141
Query: 102 ----KVTLGPSAPNSPGVGGEGENLSAAQPT-GPSSSSPPYPFSFLLSHPYLEKLKGAPS 156
+ P+ S G G + A+ P P SPP P P +E +PS
Sbjct: 142 PAVSETVPAPAPSKSKGKGKKKGKKHASSPAPSPDMMSPPAP-PTEAPGPSMESDSPSPS 200
Query: 157 QNPQDVAQE-AVLNLLRAGCLFAQVAW 182
N + A++ +L L AG +A ++W
Sbjct: 201 LNDESGAEKLKMLGSLVAG--WAVMSW 225
>gb|AAL49757.1| cask-interacting protein 2 [Homo sapiens] gi|45219847|gb|AAH66643.1|
Cask-interacting protein 2 [Homo sapiens]
gi|61213006|sp|Q8WXE0|CSKI2_HUMAN Caskin-2
gi|24638431|ref|NP_065804.1| cask-interacting protein 2
[Homo sapiens]
Length = 1202
Score = 47.0 bits (110), Expect = 8e-04
Identities = 57/240 (23%), Positives = 85/240 (34%), Gaps = 41/240 (17%)
Query: 1 MSVKPRVAEVGGKRQSSPV----------SACSPKRSKVGEPDPQTTPLTGTGGGFVAPD 50
+++K R G + +PV S +R K E +P T L G P
Sbjct: 958 LTIKQRPKPAGPPPRETPVPPGLDFNLTESDTVKRRPKCREREPLQTALLAFGVASATPG 1017
Query: 51 PVVTDSACDPLASDHPSEVNASQTPDAPRPVTL-AEGQRSPSSIRPPIPSPVKVTLGPSA 109
P PL S P E + + P P +L A+G +P + P + PV GP
Sbjct: 1018 PAA------PLPSPTPGESPPASSLPQPEPSSLPAQGVPTPLAPSPAMQPPVPPCPGPGL 1071
Query: 110 PNSPGVGGEGENLSAAQPTG----PSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQE 165
+S GE A P P + + P P S + K AP P+ V
Sbjct: 1072 ESSAASRWNGETEPPAAPAALLKVPGAGTAPKPVSVACTQLAFSGPKLAPRLGPRPVPPP 1131
Query: 166 AVLNLLRAGCLFAQVAWQVRP-LTGIVGLRQEVEELQRDLDAKEITVASQEKFLAAKDKE 224
RP TG VG Q + L++ + + + EK + K++E
Sbjct: 1132 -------------------RPESTGTVGPGQAQQRLEQTSSSLAAALRAAEKSIGTKEQE 1172
>dbj|BAA86453.1| KIAA1139 protein [Homo sapiens]
Length = 1124
Score = 47.0 bits (110), Expect = 8e-04
Identities = 57/240 (23%), Positives = 85/240 (34%), Gaps = 41/240 (17%)
Query: 1 MSVKPRVAEVGGKRQSSPV----------SACSPKRSKVGEPDPQTTPLTGTGGGFVAPD 50
+++K R G + +PV S +R K E +P T L G P
Sbjct: 880 LTIKQRPKPAGPPPRETPVPPGLDFNLTESDTVKRRPKCREREPLQTALLAFGVASATPG 939
Query: 51 PVVTDSACDPLASDHPSEVNASQTPDAPRPVTL-AEGQRSPSSIRPPIPSPVKVTLGPSA 109
P PL S P E + + P P +L A+G +P + P + PV GP
Sbjct: 940 PAA------PLPSPTPGESPPASSLPQPEPSSLPAQGVPTPLAPSPAMQPPVPPCPGPGL 993
Query: 110 PNSPGVGGEGENLSAAQPTG----PSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQE 165
+S GE A P P + + P P S + K AP P+ V
Sbjct: 994 ESSAASRWNGETEPPAAPAALLKVPGAGTAPKPVSVACTQLAFSGPKLAPRLGPRPVPPP 1053
Query: 166 AVLNLLRAGCLFAQVAWQVRP-LTGIVGLRQEVEELQRDLDAKEITVASQEKFLAAKDKE 224
RP TG VG Q + L++ + + + EK + K++E
Sbjct: 1054 -------------------RPESTGTVGPGQAQQRLEQTSSSLAAALRAAEKSIGTKEQE 1094
>ref|XP_481245.1| putative diaphanous 1 [Oryza sativa (japonica cultivar-group)]
gi|37805990|dbj|BAC99403.1| putative diaphanous 1 [Oryza
sativa (japonica cultivar-group)]
gi|37806081|dbj|BAC99532.1| putative diaphanous 1 [Oryza
sativa (japonica cultivar-group)]
Length = 893
Score = 46.6 bits (109), Expect = 0.001
Identities = 44/140 (31%), Positives = 53/140 (37%), Gaps = 14/140 (10%)
Query: 15 QSSPVSACSPKRSKVGEPDPQ---TTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNA 71
QS CSP R+ P P ++P+ +G P P P P A
Sbjct: 272 QSPSTPRCSPVRTLASPPPPPAPTSSPVRMSGPPPPPPPPAPNSCPSRPAPPPPPPPPLA 331
Query: 72 SQT----PDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQP 127
S + P AP P L SP+ PP P P T+ SAP P + G S P
Sbjct: 332 STSSPPRPAAPSPCQLHTSTSSPARPVPP-PPPTLSTIRSSAPTPPLLPGATSAPSPPPP 390
Query: 128 TGPSSSS------PPYPFSF 141
P SSS PP P SF
Sbjct: 391 PPPCSSSNQLSAPPPPPPSF 410
>gb|AAW51128.1| minus agglutinin [Chlamydomonas incerta]
Length = 4027
Score = 46.6 bits (109), Expect = 0.001
Identities = 38/143 (26%), Positives = 52/143 (35%), Gaps = 7/143 (4%)
Query: 17 SPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPD 76
+P S P P TP +P P++ P + + PS S TP
Sbjct: 1275 APPSPEPPSPRPPSPEPPSPTPPIPPSPTPPSPQPLLPA----PPSPEPPSPAPPSPTPS 1330
Query: 77 APRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPP 136
P P + A +P + PP PSP + P +P P + QP PS+ +PP
Sbjct: 1331 VPAPPSPAPPSPTPPQLPPPAPSPQPPSQPPPSPALPSPHPLAPFPPSPQPPSPSAPAPP 1390
Query: 137 YPFSFLLSHPYLEKLKGAPSQNP 159
P P G P Q P
Sbjct: 1391 SPAPL---EPAAPVPPGPPPQPP 1410
Score = 40.0 bits (92), Expect = 0.099
Identities = 35/123 (28%), Positives = 43/123 (34%), Gaps = 4/123 (3%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
P S P + P TP + P P P + PS A TP +
Sbjct: 879 PPSPVPPSPAPPSPEPPSPTPPSPAPPSPAPPSPEPPSPT--PPSPQPPSPAPALPTPPS 936
Query: 78 PRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPN--SPGVGGEGENLSAAQPTGPSSSSP 135
P P + A P S PP PSP + P +P SP L A P S +P
Sbjct: 937 PVPPSPAPPSPEPPSPFPPPPSPAPPSPEPPSPTPPSPQPPSPAPALPAPPSPVPPSPAP 996
Query: 136 PYP 138
P P
Sbjct: 997 PSP 999
Score = 39.3 bits (90), Expect = 0.17
Identities = 41/146 (28%), Positives = 51/146 (34%), Gaps = 27/146 (18%)
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGG-FVAPDPVVTDSACDPLASDHPSEVNASQT 74
+SP + +P R P P T+P G F AP P + S P P V +
Sbjct: 660 ASPRTPAAPPRP----PLPPTSPGKWAGAWPFRAPFPPIPPSPAPPQPPPLPPPVASLPP 715
Query: 75 PDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAP-----NSPGVGGEGEN-------- 121
P +P+P + SP P PSP T P +P N P G N
Sbjct: 716 PPSPKPPKPSPRPPSPRPPSPIPPSPKPPTPAPPSPAPPRPNPPSPGPPNLNPPSPEPPS 775
Query: 122 ---------LSAAQPTGPSSSSPPYP 138
A QP PS PP P
Sbjct: 776 PLPPSPAPPSPAPQPPSPSPPLPPSP 801
Score = 38.9 bits (89), Expect = 0.22
Identities = 31/100 (31%), Positives = 36/100 (36%), Gaps = 2/100 (2%)
Query: 60 PLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEG 119
P PS A P +P P + A P S RPP P P T P P SP
Sbjct: 1251 PQIPQPPSPAPALPAPPSPVPPSPAPPSPEPPSPRPPSPEPPSPT--PPIPPSPTPPSPQ 1308
Query: 120 ENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNP 159
L A P S +PP P + + P P Q P
Sbjct: 1309 PLLPAPPSPEPPSPAPPSPTPSVPAPPSPAPPSPTPPQLP 1348
Score = 38.5 bits (88), Expect = 0.29
Identities = 40/144 (27%), Positives = 51/144 (34%), Gaps = 14/144 (9%)
Query: 23 SPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVT 82
SP P P+ P +P P S P + PS + S P +P P +
Sbjct: 1422 SPAPPSPAPPSPEPRPPQPP-----SPVPPAPPSPAPP-SPVPPSPLPPSPEPPSPAPPS 1475
Query: 83 LAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPS------SSSPP 136
A SP S P P+PV + GP +P P G P PS + PP
Sbjct: 1476 PAPNAPSPPSSAP--PAPVPPSPGPPSPQPPSPGPAPPVQLPPSPRPPSLEPPSPAPRPP 1533
Query: 137 YPFSFLLSHPYLEKLKGAPSQNPQ 160
P S+P AP PQ
Sbjct: 1534 QPSPSPPSNPSPPSRPPAPPPAPQ 1557
Score = 38.1 bits (87), Expect = 0.38
Identities = 36/142 (25%), Positives = 47/142 (32%), Gaps = 5/142 (3%)
Query: 17 SPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTP- 75
+P S P + P P V P P +P + PS TP
Sbjct: 1185 APPSPEPPSPTPPSPQPPSPAPALPAPPSPVPPSPAPPSP--EPPSPFPPSPAPPRPTPP 1242
Query: 76 --DAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSS 133
+ P P SP+ P PSPV + P +P P + P P S
Sbjct: 1243 RPEPPSPTPQIPQPPSPAPALPAPPSPVPPSPAPPSPEPPSPRPPSPEPPSPTPPIPPSP 1302
Query: 134 SPPYPFSFLLSHPYLEKLKGAP 155
+PP P L + P E AP
Sbjct: 1303 TPPSPQPLLPAPPSPEPPSPAP 1324
Score = 37.4 bits (85), Expect = 0.64
Identities = 24/80 (30%), Positives = 28/80 (35%)
Query: 60 PLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEG 119
P + PS V S P +P P + +P S PP P P T P SP
Sbjct: 874 PPSPQPPSPVPPSPAPPSPEPPSPTPPSPAPPSPAPPSPEPPSPTPPSPQPPSPAPALPT 933
Query: 120 ENLSAAQPTGPSSSSPPYPF 139
P S PP PF
Sbjct: 934 PPSPVPPSPAPPSPEPPSPF 953
Score = 37.0 bits (84), Expect = 0.84
Identities = 30/88 (34%), Positives = 33/88 (37%), Gaps = 8/88 (9%)
Query: 60 PLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGP--SAPNSPGVGG 117
P + PS A P +P P A P S PP PSP + P AP SP
Sbjct: 1026 PPSPQPPSPAPALPAPPSPVPPRPAPPSPEPPSPFPPPPSPAPPSPEPPSPAPPSPQPPS 1085
Query: 118 EGENLSA------AQPTGPSSSSPPYPF 139
L A P PS S PP PF
Sbjct: 1086 PAPALPAPPSPVPPSPVPPSPSEPPSPF 1113
Score = 36.2 bits (82), Expect = 1.4
Identities = 31/121 (25%), Positives = 40/121 (32%), Gaps = 5/121 (4%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
P S P P P + P PV P Q P +
Sbjct: 1458 PPSPLPPSPEPPSPAPPSPAPNAPSPPSSAPPAPVPPSPGPPSPQPPSPGPAPPVQLPPS 1517
Query: 78 PRPVTLAEGQRSPSSIRPPIPSPVKVT--LGPSAPNSPGVGGEGENLSAAQPTGPSSSSP 135
PRP +L +P RPP PSP + PS P +P + + A P P +P
Sbjct: 1518 PRPPSLEPPSPAP---RPPQPSPSPPSNPSPPSRPPAPPPAPQPPSPPLAPPPAPPPPAP 1574
Query: 136 P 136
P
Sbjct: 1575 P 1575
Score = 36.2 bits (82), Expect = 1.4
Identities = 27/93 (29%), Positives = 38/93 (40%), Gaps = 3/93 (3%)
Query: 49 PDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPP---IPSPVKVTL 105
P P + P + PS + S P P P + A + +P S PP PSP +
Sbjct: 717 PSPKPPKPSPRPPSPRPPSPIPPSPKPPTPAPPSPAPPRPNPPSPGPPNLNPPSPEPPSP 776
Query: 106 GPSAPNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
P +P P + + S P P+ SSP P
Sbjct: 777 LPPSPAPPSPAPQPPSPSPPLPPSPAPSSPEPP 809
Score = 35.8 bits (81), Expect = 1.9
Identities = 38/136 (27%), Positives = 48/136 (34%), Gaps = 16/136 (11%)
Query: 23 SPKRSKVGEPDPQT--------TPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQT 74
SP+ P PQ TP + P P P + PS S T
Sbjct: 911 SPEPPSPTPPSPQPPSPAPALPTPPSPVPPSPAPPSPEPPSPFPPPPSPAPPSPEPPSPT 970
Query: 75 PDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPS--- 131
P +P+P + A +P S PP P+P P SP E N + P PS
Sbjct: 971 PPSPQPPSPAPALPAPPSPVPPSPAPPSPEPPSPFPPSP----EPPNPAPPSPEPPSPIP 1026
Query: 132 -SSSPPYPFSFLLSHP 146
S PP P L + P
Sbjct: 1027 PSPQPPSPAPALPAPP 1042
Score = 34.7 bits (78), Expect = 4.2
Identities = 36/138 (26%), Positives = 47/138 (33%), Gaps = 17/138 (12%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNA--SQTP 75
P + +P + P P + AP V S P S+ PS + S P
Sbjct: 1062 PPPSPAPPSPEPPSPAPPSPQPPSPAPALPAPPSPVPPSPVPPSPSEPPSPFPSPPSPVP 1121
Query: 76 DAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENL---------SAAQ 126
+P P + A + P S PP P P AP SP S AQ
Sbjct: 1122 PSPEPPSPAPPRPEPPSPTPPSPQPPSPAPALPAPRSPVPPSPAPPSPEPPSPFPPSPAQ 1181
Query: 127 PTG------PSSSSPPYP 138
P+ P S +PP P
Sbjct: 1182 PSPAPPSPEPPSPTPPSP 1199
Score = 34.3 bits (77), Expect = 5.4
Identities = 33/125 (26%), Positives = 42/125 (33%), Gaps = 7/125 (5%)
Query: 14 RQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQ 73
R SP+ SPK P P G P + + +P + PS S
Sbjct: 732 RPPSPIPP-SPKPPTPAPPSPAPPRPNPPSPG----PPNLNPPSPEPPSPLPPSPAPPSP 786
Query: 74 TPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSS 133
P P P +PSS PP P+P + P +P P P P S
Sbjct: 787 APQPPSPSPPLPPSPAPSSPEPPSPAP--PSPDPPSPTPPSPQPPSPIPHLPPPPEPPSP 844
Query: 134 SPPYP 138
PP P
Sbjct: 845 EPPRP 849
Score = 34.3 bits (77), Expect = 5.4
Identities = 27/96 (28%), Positives = 30/96 (31%), Gaps = 2/96 (2%)
Query: 45 GFVAPDPVVTDSACDPLASDHPSEVNASQTPDA--PRPVTLAEGQRSPSSIRPPIPSPVK 102
G AP S P + E Q P P P + A P S PP P P
Sbjct: 1411 GAPAPPAPPPPSPAPPSPAPPSPEPRPPQPPSPVPPAPPSPAPPSPVPPSPLPPSPEPPS 1470
Query: 103 VTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
APN+P GP S PP P
Sbjct: 1471 PAPPSPAPNAPSPPSSAPPAPVPPSPGPPSPQPPSP 1506
Score = 33.9 bits (76), Expect = 7.1
Identities = 33/135 (24%), Positives = 44/135 (32%), Gaps = 5/135 (3%)
Query: 17 SPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVN-ASQTP 75
+P S P + P P V P P P N A +P
Sbjct: 960 APPSPEPPSPTPPSPQPPSPAPALPAPPSPVPPSPAPPSPEPPSPFPPSPEPPNPAPPSP 1019
Query: 76 DAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPS---- 131
+ P P+ + SP+ P PSPV P +P P + + P PS
Sbjct: 1020 EPPSPIPPSPQPPSPAPALPAPPSPVPPRPAPPSPEPPSPFPPPPSPAPPSPEPPSPAPP 1079
Query: 132 SSSPPYPFSFLLSHP 146
S PP P L + P
Sbjct: 1080 SPQPPSPAPALPAPP 1094
Score = 33.9 bits (76), Expect = 7.1
Identities = 34/131 (25%), Positives = 44/131 (32%), Gaps = 10/131 (7%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
P S P + P P + + +P P+ + P P A P
Sbjct: 1361 PPSPALPSPHPLAPFPPSPQPPSPSAPAPPSPAPLEPAAPVPPGPPPQPPGAPAPPAPPP 1420
Query: 78 PRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSP----GVGGEGENLSAAQPT----- 128
P P + SP P PSPV AP SP + E S A P+
Sbjct: 1421 PSPAPPSPAPPSPEPRPPQPPSPVPPAPPSPAPPSPVPPSPLPPSPEPPSPAPPSPAPNA 1480
Query: 129 -GPSSSSPPYP 138
P SS+PP P
Sbjct: 1481 PSPPSSAPPAP 1491
Score = 33.9 bits (76), Expect = 7.1
Identities = 29/99 (29%), Positives = 41/99 (41%), Gaps = 10/99 (10%)
Query: 48 APDPVVTDSACD-PLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLG 106
+P+P + + PL PS A +P P PV + SP P PSP +
Sbjct: 851 SPEPPIPEPPSPAPLIPAPPSP--APPSPQPPSPVPPSPAPPSPEPPSPTPPSPAPPSPA 908
Query: 107 PSAPNSPG---VGGEGENLSAAQPTGPS----SSSPPYP 138
P +P P + + + A PT PS S +PP P
Sbjct: 909 PPSPEPPSPTPPSPQPPSPAPALPTPPSPVPPSPAPPSP 947
Score = 33.5 bits (75), Expect = 9.3
Identities = 31/94 (32%), Positives = 39/94 (40%), Gaps = 5/94 (5%)
Query: 49 PDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPS 108
P P P + PS S P +P+P + A +P S PP P P + PS
Sbjct: 1052 PSPEPPSPFPPPPSPAPPSPEPPSPAPPSPQPPSPAPALPAPPSPVPPSPVPPSPSEPPS 1111
Query: 109 ---APNSPGVGGEGENLSAAQP-TGPSSSSPPYP 138
+P SP V E S A P P S +PP P
Sbjct: 1112 PFPSPPSP-VPPSPEPPSPAPPRPEPPSPTPPSP 1144
>emb|CAA67585.1| arabinogalactan [Lycopersicon esculentum]
gi|1430844|emb|CAA67584.1| arabinogalactan [Lycopersicon
esculentum] gi|872127|emb|CAA88023.1| putative
arabinogalactan-protein [Lycopersicon esculentum]
gi|1362097|pir||S55925 probable arabinogalactan protein
precursor - tomato
Length = 215
Score = 45.8 bits (107), Expect = 0.002
Identities = 49/192 (25%), Positives = 73/192 (37%), Gaps = 17/192 (8%)
Query: 5 PRVAEVGGKRQSSPVSACSPKRSKVGEP-------DPQTTPLTGTGGGFVAPDPVVTDSA 57
P A VG K ++P +A +P + K P P P+ AP V +
Sbjct: 24 PAAAPVGAKAGTTPPAA-APTKPKTPAPATAPASAPPTAVPVAPVTAPVTAPTTPVVAAP 82
Query: 58 CDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPV------KVTLGPSAPN 111
AS P + AS P P E P+ PP +PV + T P+
Sbjct: 83 VSAPASSPPLKAPASSPPVQSPPAPAPEVATPPAVSTPPAAAPVAAPVASETTPAPAPSK 142
Query: 112 SPGVGGEGENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQE-AVLNL 170
G +G+ +A+ P SPP P S +PS N + A++ +L
Sbjct: 143 GKVKGKKGKKHNASPAPSPDMMSPPAPPSEAPGPSMDSDSAPSPSLNDESGAEKLKMLGS 202
Query: 171 LRAGCLFAQVAW 182
L AG +A ++W
Sbjct: 203 LVAG--WAVMSW 212
>gb|AAP37768.1| At3g24600 [Arabidopsis thaliana] gi|13877617|gb|AAK43886.1| protein
kinase-like protein [Arabidopsis thaliana]
Length = 652
Score = 45.8 bits (107), Expect = 0.002
Identities = 37/134 (27%), Positives = 53/134 (38%), Gaps = 10/134 (7%)
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTP 75
++P +P S P TT +P P T S+ P PS + S P
Sbjct: 3 TAPSPGTTPSPSPPSPPTNSTTTTPPPAAS--SPPPTTTPSSPPP----SPSTNSTSPPP 56
Query: 76 DAPRPVTLAEGQRSPSSIRPPIPSP---VKVTLGPSAPNSPGVGGEGENLSAAQPTGPSS 132
+P P +L P S+ PPIP P +T P +P +P + + P PS
Sbjct: 57 SSPLPPSLPPPS-PPGSLTPPIPQPSPSAPITPSPPSPTTPSNPRSPPSPNQGPPNTPSG 115
Query: 133 SSPPYPFSFLLSHP 146
S+P P + S P
Sbjct: 116 STPRTPSNTKPSPP 129
>emb|CAG60730.1| unnamed protein product [Candida glabrata CBS138]
gi|50290701|ref|XP_447783.1| unnamed protein product
[Candida glabrata]
Length = 979
Score = 45.8 bits (107), Expect = 0.002
Identities = 31/114 (27%), Positives = 50/114 (43%), Gaps = 9/114 (7%)
Query: 54 TDSACDPLASDHPSEVN-ASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNS 112
TDS C+ + PS V +S P + P ++ +PSS+ P +P + P+S
Sbjct: 320 TDSKCEDIPPIEPSSVKPSSMNPSSVNPSSMNPSSMNPSSMNPSSMNPSSMNPSSMNPSS 379
Query: 113 PGVGGEGENLSAAQPTGPSSSSPPYPFSFLLS----HPYLEKLKGAPSQNPQDV 162
P N S+ P+ PSS +PP+ S ++ +P S NP +
Sbjct: 380 P----SSINPSSMNPSSPSSMNPPFANSSSMNPSSMNPSSMNPSSPSSMNPSSI 429
Score = 43.1 bits (100), Expect = 0.012
Identities = 43/169 (25%), Positives = 69/169 (40%), Gaps = 14/169 (8%)
Query: 2 SVKPRVAEVGGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPL 61
S+ P SSP S+ +P +P + + + P V S+ +P
Sbjct: 405 SMNPSSMNPSSMNPSSP-SSMNPSSINPSSMNPSSINPSSMNPSSMNPSSV-NPSSMNP- 461
Query: 62 ASDHPSEVN------ASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVT-LGPSAPNSPG 114
+S +PS VN +S P + P ++ +PSS+ P PS + + + PS+P+S
Sbjct: 462 SSVNPSSVNPSSMNPSSMNPSSMNPSSMNPSSVNPSSMNPSSPSSMNPSSMNPSSPSSMN 521
Query: 115 VGGEGE----NLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNP 159
+ S+ P+ PSS +P P S S P S NP
Sbjct: 522 PSSPSSMNPSSPSSMNPSSPSSMNPSSPSSMNPSSPSSMNPSSPSSMNP 570
Score = 35.8 bits (81), Expect = 1.9
Identities = 36/169 (21%), Positives = 66/169 (38%), Gaps = 6/169 (3%)
Query: 2 SVKPRVAEVGGKRQSSPVSACSPKRSKVGEPDPQT-TPLTGTGGGFVAPDPVVTDSACDP 60
S+ P +P S S S +P + + + + + P + + P
Sbjct: 506 SMNPSSMNPSSPSSMNPSSPSSMNPSSPSSMNPSSPSSMNPSSPSSMNPSSPSSMNPSSP 565
Query: 61 LASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGE 120
+ + P ++S P + P ++ +PSS+ P +P ++ PS+ N +
Sbjct: 566 SSMNPPFANSSSMNPSSVNPSSMNPSSMNPSSMNPSSMNPS--SMNPSSMNPSSMNPSSM 623
Query: 121 NLSAAQPTG--PSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQEAV 167
N S+ P+ PSS +P P S S + + S NP V +V
Sbjct: 624 NPSSMNPSSMNPSSMNPSSPSSMNPSSINPSSMNPS-SMNPSSVNPSSV 671
>gb|AAS07042.1| minus agglutinin [Chlamydomonas reinhardtii]
Length = 3889
Score = 45.8 bits (107), Expect = 0.002
Identities = 35/118 (29%), Positives = 49/118 (40%), Gaps = 16/118 (13%)
Query: 48 APDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIP------SPV 101
+P+P + P + + PS V S P +P P + A +P PP P SPV
Sbjct: 1148 SPEPPSPAPSPPPPSPEPPSPVPPSPPPPSPEPPSPAPPSPAPPLPLPPSPHTQSPPSPV 1207
Query: 102 KVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNP 159
+ PSAP+ P + QP P + SPP P S P+ AP+ P
Sbjct: 1208 PPSPAPSAPSPP----------SPQPPSPLAPSPPSPAPQAPSSPFPPPQPTAPTAPP 1255
Score = 44.7 bits (104), Expect = 0.004
Identities = 33/122 (27%), Positives = 45/122 (36%), Gaps = 9/122 (7%)
Query: 17 SPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPD 76
+P SA P P P + V P P+ P + + PS V S P
Sbjct: 797 APPSAAPPSPMPPSPAPPSPDPPSPKPPSPVPPSPL-------PPSPEPPSPVPPSPPPA 849
Query: 77 APRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPP 136
+P P + A P S PP P+P P +P P L + +P P+ PP
Sbjct: 850 SPEPTSPAPPSPPPPSPEPPSPAPPSPP--PPSPEPPSPAPPSPPLPSPEPPSPAPPLPP 907
Query: 137 YP 138
P
Sbjct: 908 PP 909
Score = 44.3 bits (103), Expect = 0.005
Identities = 27/90 (30%), Positives = 40/90 (44%), Gaps = 2/90 (2%)
Query: 49 PDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPS 108
P P + + P + + PS S P +P P + A P S +PP SPV + P
Sbjct: 777 PSPSIPPPSPGPPSPEPPSPAPPSAAPPSPMPPSPAPPSPDPPSPKPP--SPVPPSPLPP 834
Query: 109 APNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
+P P ++ +PT P+ SPP P
Sbjct: 835 SPEPPSPVPPSPPPASPEPTSPAPPSPPPP 864
Score = 42.4 bits (98), Expect = 0.020
Identities = 35/125 (28%), Positives = 49/125 (39%), Gaps = 10/125 (8%)
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNA-SQT 74
S P + P P P P + P P A A P + +Q+
Sbjct: 1143 SPPPPSPEPPSPAPSPPPPSPEPPSPVPPSPPPPSPEPPSPAPPSPAPPLPLPPSPHTQS 1202
Query: 75 PDAPRPVTLAEGQRSPSSIRPPIP-SPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSS 133
P +P P + A SP S +PP P +P + P AP+SP QPT P++
Sbjct: 1203 PPSPVPPSPAPSAPSPPSPQPPSPLAPSPPSPAPQAPSSP--------FPPPQPTAPTAP 1254
Query: 134 SPPYP 138
PP+P
Sbjct: 1255 PPPFP 1259
Score = 40.8 bits (94), Expect = 0.058
Identities = 38/137 (27%), Positives = 55/137 (39%), Gaps = 16/137 (11%)
Query: 17 SPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPL--ASDHPSE---VNA 71
+P S P + + P TP + +P+P V PL + PS V
Sbjct: 1326 APPSPMPPSPAPLAPQPPSPTPPSPAPPVPPSPEPPVPPGPDPPLPPSPTPPSPQPPVPP 1385
Query: 72 SQTPDAPRPVTLAEGQRSPS----------SIRPPIPSPVKVTLGPSAPNSPGVGGEGEN 121
S TP +P+P + A +PS S +PP P+P PS P++P
Sbjct: 1386 SPTPPSPQPPSPAPPSPAPSAPLQPSPDPPSPQPPSPAPGPPPSPPSPPSTPSPPSPAP- 1444
Query: 122 LSAAQPTGPSSSSPPYP 138
L+ A P P + PP P
Sbjct: 1445 LAPAPPVPPMAPQPPSP 1461
Score = 37.7 bits (86), Expect = 0.49
Identities = 31/121 (25%), Positives = 41/121 (33%), Gaps = 5/121 (4%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
P S P + P P + P P P + + PS V S P +
Sbjct: 1061 PPSPAPPSPAPPSPEPPSPAPPSPAPPSPEPPSPAPPSPR--PPSPEPPSPVPPSPPPPS 1118
Query: 78 PRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPY 137
P P + A P S+ PP P+P P SP + +P P SPP
Sbjct: 1119 PEPPSPAPPSPPPPSLEPPSPAPPSPPPPSPEPPSP---APSPPPPSPEPPSPVPPSPPP 1175
Query: 138 P 138
P
Sbjct: 1176 P 1176
Score = 37.7 bits (86), Expect = 0.49
Identities = 38/143 (26%), Positives = 46/143 (31%), Gaps = 1/143 (0%)
Query: 17 SPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPD 76
SP P + P P + P AP P + +P + PS S P
Sbjct: 989 SPAPLPPPPSPEPPSPAPPSPPPPSPEPPSPAP-PSPPPPSPEPPSPAPPSPPPPSPEPP 1047
Query: 77 APRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPP 136
+P P + A P S PP P+P AP SP A P S PP
Sbjct: 1048 SPAPPSPAPPSPEPPSPAPPSPAPPSPEPPSPAPPSPAPPSPEPPSPAPPSPRPPSPEPP 1107
Query: 137 YPFSFLLSHPYLEKLKGAPSQNP 159
P P E AP P
Sbjct: 1108 SPVPPSPPPPSPEPPSPAPPSPP 1130
Score = 37.0 bits (84), Expect = 0.84
Identities = 32/126 (25%), Positives = 46/126 (36%), Gaps = 5/126 (3%)
Query: 17 SPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPD 76
SP + P + P P + P AP P+ + +P + PS S P
Sbjct: 930 SPAPSPPPPSPEPPSPAPPSPPPPSPEPPSPAP-PLPPPPSPEPPSPAPPSPPPPSPEPP 988
Query: 77 APRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPS----S 132
+P P+ PS P P P P+ P+ P E + + P PS S
Sbjct: 989 SPAPLPPPPSPEPPSPAPPSPPPPSPEPPSPAPPSPPPPSPEPPSPAPPSPPPPSPEPPS 1048
Query: 133 SSPPYP 138
+PP P
Sbjct: 1049 PAPPSP 1054
Score = 36.6 bits (83), Expect = 1.1
Identities = 27/122 (22%), Positives = 38/122 (31%)
Query: 17 SPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPD 76
S S SP+ P P + F P P + P +P+
Sbjct: 1214 SAPSPPSPQPPSPLAPSPPSPAPQAPSSPFPPPQPTAPTAPPPPFPPSPAPPSPTPPSPE 1273
Query: 77 APRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPP 136
P P + +P S PP P+P P SP ++ + P S PP
Sbjct: 1274 PPAPQPPSPTPHAPPSPEPPSPTPPSPLPPSPEPPSPSPPSPAPSVPSPPSPAPPSPMPP 1333
Query: 137 YP 138
P
Sbjct: 1334 SP 1335
Score = 35.4 bits (80), Expect = 2.4
Identities = 26/91 (28%), Positives = 37/91 (40%), Gaps = 3/91 (3%)
Query: 48 APDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGP 107
+P+P + P + + PS S P +P P + A P S PP P+P P
Sbjct: 925 SPEPPSPAPSPPPPSPEPPSPAPPSPPPPSPEPPSPAPPLPPPPSPEPPSPAPPSPP--P 982
Query: 108 SAPNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
+P P S +P P+ SPP P
Sbjct: 983 PSPEPPSPAPLPPPPS-PEPPSPAPPSPPPP 1012
Score = 35.4 bits (80), Expect = 2.4
Identities = 40/144 (27%), Positives = 49/144 (33%), Gaps = 3/144 (2%)
Query: 17 SPVSACSPKRS-KVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTP 75
SPV P S + P P + P AP P + +P + PS S P
Sbjct: 840 SPVPPSPPPASPEPTSPAPPSPPPPSPEPPSPAP-PSPPPPSPEPPSPAPPSPPLPSPEP 898
Query: 76 DAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSP 135
+P P P S PP P P AP+ P E + + P P S P
Sbjct: 899 PSPAPPLPPPPSPEPPSPAPPSPPPPSPEPPSPAPSPPPPSPEPPSPAPPSPP-PPSPEP 957
Query: 136 PYPFSFLLSHPYLEKLKGAPSQNP 159
P P L P E AP P
Sbjct: 958 PSPAPPLPPPPSPEPPSPAPPSPP 981
Score = 35.0 bits (79), Expect = 3.2
Identities = 25/83 (30%), Positives = 33/83 (39%), Gaps = 13/83 (15%)
Query: 59 DPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPI---PSPVKVTLGPSAPNSPGV 115
+P + PS + S P +P P + A SP S PP PSP + P +P P
Sbjct: 1291 EPPSPTPPSPLPPSPEPPSPSPPSPAPSVPSPPSPAPPSPMPPSPAPLAPQPPSPTPP-- 1348
Query: 116 GGEGENLSAAQPTGPSSSSPPYP 138
+ P P S PP P
Sbjct: 1349 --------SPAPPVPPSPEPPVP 1363
Score = 35.0 bits (79), Expect = 3.2
Identities = 35/126 (27%), Positives = 46/126 (35%), Gaps = 23/126 (18%)
Query: 23 SPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSE-----VNASQTPDA 77
SP+ PDP P P T + P + PS + S P +
Sbjct: 1357 SPEPPVPPGPDPPLPPSPTPPSPQPPVPPSPTPPSPQPPSPAPPSPAPSAPLQPSPDPPS 1416
Query: 78 PRPVTLAEG-----QRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSS 132
P+P + A G PS+ PP P+P L P+ P P A QP P
Sbjct: 1417 PQPPSPAPGPPPSPPSPPSTPSPPSPAP----LAPAPPVPP---------MAPQPPSPPL 1463
Query: 133 SSPPYP 138
SPP+P
Sbjct: 1464 PSPPFP 1469
Score = 34.7 bits (78), Expect = 4.2
Identities = 34/131 (25%), Positives = 46/131 (34%), Gaps = 19/131 (14%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
P S P + P P P + P P +P + PS A +P+
Sbjct: 1026 PPSPEPPSPAPPSPPPPSPEPPSPAPPSPAPPSP-------EPPSPAPPSP--APPSPEP 1076
Query: 78 PRPVTLAEGQRSPS-------SIRPPI---PSPVKVTLGPSAPNSPGVGGEGENLSAAQP 127
P P + SP S RPP PSPV + P +P P + +P
Sbjct: 1077 PSPAPPSPAPPSPEPPSPAPPSPRPPSPEPPSPVPPSPPPPSPEPPSPAPPSPPPPSLEP 1136
Query: 128 TGPSSSSPPYP 138
P+ SPP P
Sbjct: 1137 PSPAPPSPPPP 1147
Score = 34.3 bits (77), Expect = 5.4
Identities = 31/117 (26%), Positives = 42/117 (35%), Gaps = 13/117 (11%)
Query: 24 PKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTL 83
P + P P + PL AP P+ + +P + PS S P +P P
Sbjct: 878 PPSPEPPSPAPPSPPLPSPEPPSPAP-PLPPPPSPEPPSPAPPSPPPPSPEPPSPAPSPP 936
Query: 84 AEGQRSPSSIRP--PIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
PS P P PSP + P P P + +P P+ SPP P
Sbjct: 937 PPSPEPPSPAPPSPPPPSPEPPSPAPPLPPPP----------SPEPPSPAPPSPPPP 983
Score = 33.9 bits (76), Expect = 7.1
Identities = 22/73 (30%), Positives = 28/73 (38%), Gaps = 2/73 (2%)
Query: 66 PSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAA 125
PS S TP +P P + SP S P +PSP P +P P +
Sbjct: 1288 PSPEPPSPTPPSPLPPSPEPPSPSPPSPAPSVPSPPSPA--PPSPMPPSPAPLAPQPPSP 1345
Query: 126 QPTGPSSSSPPYP 138
P P+ PP P
Sbjct: 1346 TPPSPAPPVPPSP 1358
Score = 33.9 bits (76), Expect = 7.1
Identities = 23/91 (25%), Positives = 32/91 (34%), Gaps = 2/91 (2%)
Query: 48 APDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGP 107
+P+P P S P P +P P + A P S PP P+P P
Sbjct: 969 SPEPPSPAPPSPPPPSPEPPSPAPLPPPPSPEPPSPAPPSPPPPSPEPPSPAPPSPP--P 1026
Query: 108 SAPNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
+P P + +P P+ SP P
Sbjct: 1027 PSPEPPSPAPPSPPPPSPEPPSPAPPSPAPP 1057
>emb|CAG03262.1| unnamed protein product [Tetraodon nigroviridis]
Length = 4006
Score = 45.8 bits (107), Expect = 0.002
Identities = 43/134 (32%), Positives = 54/134 (40%), Gaps = 17/134 (12%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
P+S+C+ + E +PQ L G+G A L S P A P A
Sbjct: 3864 PLSSCASPPPPISE-EPQQEQLPGSG------------QARGRLDSGPPCRYPAQAVPQA 3910
Query: 78 PRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLS-AAQPTGPSSSSPP 136
P LA R PSS I S + + + P V G G N S AA P P SPP
Sbjct: 3911 PGSNPLA---RQPSSEHLDILSSILASFNSTVLTPPAVSGPGPNGSPAALPPPPPHLSPP 3967
Query: 137 YPFSFLLSHPYLEK 150
P ++ S P L K
Sbjct: 3968 PPTAYPRSPPTLSK 3981
>ref|XP_510940.1| PREDICTED: similar to KIAA0339 protein [Pan troglodytes]
Length = 920
Score = 45.4 bits (106), Expect = 0.002
Identities = 31/126 (24%), Positives = 49/126 (38%), Gaps = 4/126 (3%)
Query: 11 GGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVN 70
G +S S CS GE D + + + + + S+ +S SE +
Sbjct: 203 GEDEESDSSSKCSLYADSDGENDSTSDSESSSSSSSSSS----SSSSSSSSSSSSSSESS 258
Query: 71 ASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGP 130
+ + RP L P + P P+PV+V + SP + S A+P GP
Sbjct: 259 SEDEEEEERPAALPSASPPPREVPVPTPAPVEVPVPERVAGSPVTPLPEQEASPARPAGP 318
Query: 131 SSSSPP 136
+ SPP
Sbjct: 319 TEESPP 324
>ref|NP_055527.1| hypothetical protein LOC9739 [Homo sapiens]
Length = 1707
Score = 45.4 bits (106), Expect = 0.002
Identities = 31/126 (24%), Positives = 49/126 (38%), Gaps = 4/126 (3%)
Query: 11 GGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVN 70
G +S S CS GE D + + + + + S+ +S SE +
Sbjct: 1006 GEDEESDSSSKCSLYADSDGENDSTSDSESSSSSSSSSS----SSSSSSSSSSSSSSESS 1061
Query: 71 ASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGP 130
+ + RP L P + P P+PV+V + SP + S A+P GP
Sbjct: 1062 SEDEEEEERPAALPSASPPPREVPVPTPAPVEVPVPERVAGSPVTPLPEQEASPARPAGP 1121
Query: 131 SSSSPP 136
+ SPP
Sbjct: 1122 TEESPP 1127
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.310 0.129 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 625,169,682
Number of Sequences: 2540612
Number of extensions: 30208365
Number of successful extensions: 157028
Number of sequences better than 10.0: 3447
Number of HSP's better than 10.0 without gapping: 622
Number of HSP's successfully gapped in prelim test: 2967
Number of HSP's that attempted gapping in prelim test: 143890
Number of HSP's gapped (non-prelim): 11523
length of query: 337
length of database: 863,360,394
effective HSP length: 128
effective length of query: 209
effective length of database: 538,162,058
effective search space: 112475870122
effective search space used: 112475870122
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0094b.3


