

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0094a.6
(990 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
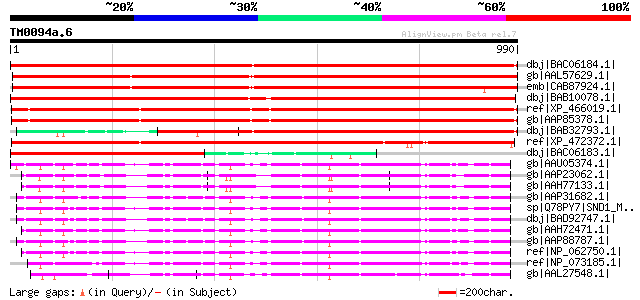
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAC06184.1| 110 kDa 4SNc-Tudor domain protein [Pisum sativum] 1642 0.0
gb|AAL57629.1| AT5g07350/T2I1_60 [Arabidopsis thaliana] gi|22326... 1431 0.0
emb|CAB87924.1| putative protein [Arabidopsis thaliana] gi|11358... 1424 0.0
dbj|BAB10078.1| transcription factor-like protein [Arabidopsis t... 1406 0.0
ref|XP_466019.1| RNA binding protein Rp120 [Oryza sativa (japoni... 1351 0.0
gb|AAP85378.1| RNA binding protein Rp120 [Oryza sativa (japonica... 1346 0.0
dbj|BAB32793.1| 110 kDa 4SNc-Tudor domain protein [Pisum sativum] 1169 0.0
ref|XP_472372.1| OSJNBb0012E08.11 [Oryza sativa (japonica cultiv... 1014 0.0
dbj|BAC06183.1| 110kDa protein HMP [Pisum sativum] 641 0.0
gb|AAU05374.1| SND p102 [Rattus norvegicus] gi|60415342|sp|Q66X9... 441 e-122
gb|AAP23062.1| p100 co-activator variant 1 [Danio rerio] gi|3350... 438 e-121
gb|AAH77133.1| Staphylococcal nuclease domain containing 1 [Dani... 438 e-121
gb|AAP31682.1| 100 kDa coactivator [Bos taurus] gi|60415927|sp|Q... 437 e-121
sp|Q78PY7|SND1_MOUSE Staphylococcal nuclease domain containing p... 437 e-121
dbj|BAD92747.1| EBNA-2 co-activator variant [Homo sapiens] 435 e-120
gb|AAH72471.1| Unknown (protein for MGC:91679) [Rattus norvegicus] 435 e-120
gb|AAP88787.1| EBNA-2 co-activator (100kD) [Homo sapiens] gi|613... 434 e-120
ref|NP_062750.1| staphylococcal nuclease domain containing 1 [Mu... 431 e-119
ref|NP_073185.1| p105 coactivator [Rattus norvegicus] gi|1800307... 411 e-113
gb|AAL27548.1| transcriptional coactivator p100 [Gallus gallus] 365 4e-99
>dbj|BAC06184.1| 110 kDa 4SNc-Tudor domain protein [Pisum sativum]
Length = 989
Score = 1642 bits (4252), Expect = 0.0
Identities = 826/992 (83%), Positives = 905/992 (90%), Gaps = 5/992 (0%)
Query: 1 MASAATGATGWYRGRVKAVPSGDCLVIVAVASS-KPGPLPEKSITLASLITPRLARRGGV 59
MA+ A G + WY+ +VKAV SGDC+V+V+VA++ K G LPEKSITL+SLI PRLARRGGV
Sbjct: 1 MATTAAGNSAWYKAKVKAVTSGDCVVVVSVAANAKSGVLPEKSITLSSLIAPRLARRGGV 60
Query: 60 DEPFAWESREYLRKLCIGKEVTFRVDYNVASINRDFGTVFLGEKNVGVLVVSQGWAKVRE 119
DE FAWESRE+LRKLCIG+E+TFR+DY V SINR+FGTVFLG+KNV +LVVSQGWAKVRE
Sbjct: 61 DEAFAWESREFLRKLCIGREITFRIDYTVPSINREFGTVFLGDKNVAMLVVSQGWAKVRE 120
Query: 120 QGQQKGEVSPYLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGL 179
QGQQKGEVSP+LAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSA+GDASNFDAMGL
Sbjct: 121 QGQQKGEVSPFLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSALGDASNFDAMGL 180
Query: 180 LAANKGSPMEAIVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELT 239
LA +KG PMEA+VEQVRDGSTLR+YLLPEFQFVQVFVAGIQSPQMGRRAAPETVVE E+T
Sbjct: 181 LAKSKGVPMEALVEQVRDGSTLRIYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVEPEVT 240
Query: 240 ADENDGDVPGEPRPPLTSAQRLAVSSST-ETAADPFGPDAKFFTEMRVLNRDVRIVLEGV 298
D +GD P EPR PLTSAQRLAVS+S ET+ADPFGPDAKFFTEMRVLNRDVRIVLEGV
Sbjct: 241 VDSTNGDAPAEPRAPLTSAQRLAVSASAAETSADPFGPDAKFFTEMRVLNRDVRIVLEGV 300
Query: 299 DKFSNLIGSVYYPDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSR 358
DKFSNLIGSVYYPDGESAKD LELVENG+AKYVEWSA+MMEE+AKR+LK+AELEAKKSR
Sbjct: 301 DKFSNLIGSVYYPDGESAKDWPLELVENGFAKYVEWSAHMMEEDAKRKLKSAELEAKKSR 360
Query: 359 LRMWTNYVPPASNSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRC 418
LR+WTNYVPP SNSKAIH+QN TGK+VEVVSGDC+IVADDSIPYGSP AERRVNLSSIRC
Sbjct: 361 LRIWTNYVPPVSNSKAIHDQNLTGKLVEVVSGDCVIVADDSIPYGSPQAERRVNLSSIRC 420
Query: 419 PKVGNPRRDEKPAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADSRVM 478
PK+GNPRRDEKPAPYAREAKEFLRTRL+GRQV+V+MEYSRK+ P D + P A D RVM
Sbjct: 421 PKMGNPRRDEKPAPYAREAKEFLRTRLIGRQVNVQMEYSRKVGPVDAAGAPLGAGD-RVM 479
Query: 479 DFGSVFLLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVIRHRDFEERSNYYD 538
DFGSVFL S+ KAD+D PS+ A S+ G+NVGELV+GRGFGTVIRHRDFEERSN+YD
Sbjct: 480 DFGSVFLSSSGKADNDQAPSAAAPASSK-LGLNVGELVIGRGFGTVIRHRDFEERSNFYD 538
Query: 539 ALLTAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEYVLS 598
ALL AESRA+SGRKGIHSAKDPPVMHITDLTT SAKKAKDF+PFLHRSRR+PAVVEYVLS
Sbjct: 539 ALLAAESRAISGRKGIHSAKDPPVMHITDLTTASAKKAKDFMPFLHRSRRVPAVVEYVLS 598
Query: 599 GHRFKLLIPKETCSIAFALSGVRCPGRGEPYSEEAIALMRRKIMQRDVEIEVETVDRNGT 658
GHRFKLLIPKETCSIAFA SGVRCPGR EPYS+EAIALMRR+IMQRDVEIEVETVDR GT
Sbjct: 599 GHRFKLLIPKETCSIAFAFSGVRCPGREEPYSDEAIALMRRRIMQRDVEIEVETVDRTGT 658
Query: 659 FLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKLKIWENFVEG 718
FLG LWES+TN A+ LLEAGLAKLQT+FGSDRIP L++ EQSAK +KLKIWENFVEG
Sbjct: 659 FLGPLWESKTNGAVALLEAGLAKLQTTFGSDRIPGSSCLEQPEQSAKSKKLKIWENFVEG 718
Query: 719 EEVSNGANVESKQQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALNLKEAPVLGA 778
E V +GANVE+KQQEVLKV VTEVLGGGKFYVQTVGDQKIASIQ QLA+LNLKEAPV+GA
Sbjct: 719 EVVPSGANVETKQQEVLKVTVTEVLGGGKFYVQTVGDQKIASIQNQLASLNLKEAPVIGA 778
Query: 779 FSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPL 838
F+PKKGDIVLCYF AD SWYRAMVVNTPRGPVES KD+FEVFY+DYGNQE+V YSQLRPL
Sbjct: 779 FNPKKGDIVLCYFRADTSWYRAMVVNTPRGPVESSKDVFEVFYLDYGNQEEVPYSQLRPL 838
Query: 839 DQSVSAAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTSGGK 898
D SVS APGLAQLCSLAYIK P+LEEDFGQEAAEYLSELTLSSGKEFRA VEERDT+GGK
Sbjct: 839 DPSVSLAPGLAQLCSLAYIKIPNLEEDFGQEAAEYLSELTLSSGKEFRAMVEERDTTGGK 898
Query: 899 VKGQGTGTILAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQDDA 958
VKGQGTG ++AVTLVAVDAEISVNAAMLQEGLARMEKRNRWD+ RK LD+L+ FQ +A
Sbjct: 899 VKGQGTGPVIAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDKSARKQALDNLEMFQGEA 958
Query: 959 RKERRGMWQYGDVESDDEDTAPPARKAGAGRR 990
R RRG+WQYGD++SDDEDTAPP + AG GRR
Sbjct: 959 RTSRRGIWQYGDIQSDDEDTAPPRKPAG-GRR 989
>gb|AAL57629.1| AT5g07350/T2I1_60 [Arabidopsis thaliana]
gi|22326646|ref|NP_196352.2| tudor domain-containing
protein / nuclease family protein [Arabidopsis thaliana]
Length = 991
Score = 1431 bits (3703), Expect = 0.0
Identities = 726/995 (72%), Positives = 843/995 (83%), Gaps = 14/995 (1%)
Query: 5 ATGATG-WYRGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLARRGGVDEPF 63
ATGA W +GRVKAV SGDCLVI A++ ++ GP PEK+IT +SL+ P++ARRGG+DEPF
Sbjct: 2 ATGAENQWLKGRVKAVTSGDCLVITALSHNRAGPPPEKTITFSSLMAPKMARRGGIDEPF 61
Query: 64 AWESREYLRKLCIGKEVTFRVDYNVASI-NRDFGTVFLGEKNVGVLVVSQGWAKVREQGQ 122
AWES+E+LRKLCIGKEV F+VDY V +I R+FG+VFLG +N+ LVV GWAKVRE GQ
Sbjct: 62 AWESKEFLRKLCIGKEVAFKVDYKVEAIAGREFGSVFLGNENLAKLVVKTGWAKVREPGQ 121
Query: 123 Q-KGEVSPYLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGLLA 181
Q + +VSPY+ ELL+LEE AKQEG GRWSKVPGAAEASIRNLPPSAIGD++ FDAMGLLA
Sbjct: 122 QNQDKVSPYIKELLQLEELAKQEGYGRWSKVPGAAEASIRNLPPSAIGDSAGFDAMGLLA 181
Query: 182 ANKGSPMEAIVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELTAD 241
ANKG PME IVEQVRDGST+RVYLLPEFQFVQVFVAG+Q+P MGRR +VVET D
Sbjct: 182 ANKGKPMEGIVEQVRDGSTIRVYLLPEFQFVQVFVAGVQAPSMGRRTTNGSVVET--VPD 239
Query: 242 ENDGDVPGEPRPPLTSAQRLAVS--SSTETAADPFGPDAKFFTEMRVLNRDVRIVLEGVD 299
E +GDV E R PLT+AQRLA S SS E ++DPF +AK+FTE RVL+RDVRIVLEGVD
Sbjct: 240 EPNGDVSAESRGPLTTAQRLAASAASSVEVSSDPFATEAKYFTEHRVLSRDVRIVLEGVD 299
Query: 300 KFSNLIGSVYYPDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRL 359
KF+NLIGSV+Y DGE+ KDL LELVENG AK+VEWSANMMEEEAK++LK AEL+ KK ++
Sbjct: 300 KFNNLIGSVHYSDGETVKDLGLELVENGLAKFVEWSANMMEEEAKKKLKAAELQCKKDKV 359
Query: 360 RMWTNYVPPASNSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCP 419
+MW NYVPPA+NSKAIH+QNFTGKVVEVVSGDC+IVADD++P+GSP AERRV LSSIR P
Sbjct: 360 KMWANYVPPATNSKAIHDQNFTGKVVEVVSGDCLIVADDAVPFGSPAAERRVCLSSIRSP 419
Query: 420 KVGNPRRDEKPAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADSRVMD 479
K+GNPRR+EKPAPYAREA+EFLR RL+G+QV V+MEYSRK+ DG S AAD R MD
Sbjct: 420 KMGNPRREEKPAPYAREAREFLRQRLIGKQVIVQMEYSRKVTQGDGPTT-SGAAD-RFMD 477
Query: 480 FGSVFLLSATKADSDDT--PSSIPSAGSQPTGVNVGELVVGRGFGTVIRHRDFEERSNYY 537
FGSVFL SA KADSD+ P + AGSQP GVN+ ELV+ RGFG V+RHRDFEERSN+Y
Sbjct: 478 FGSVFLPSAAKADSDEVTAPPAAAIAGSQPVGVNIAELVLVRGFGNVVRHRDFEERSNHY 537
Query: 538 DALLTAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEYVL 597
DALL AE+RAL+G+KGIHSAK+ P MHITDLT ++AKKAKDFLP L R RRIPAVVEYVL
Sbjct: 538 DALLAAEARALAGKKGIHSAKESPAMHITDLTVSAAKKAKDFLPSLQRIRRIPAVVEYVL 597
Query: 598 SGHRFKLLIPKETCSIAFALSGVRCPGRGEPYSEEAIALMRRKIMQRDVEIEVETVDRNG 657
SGHRFKL IPK TCSIAF+ SGVRCPGRGEPYSEEAI++MRR+IMQRDVEIEVETVDR G
Sbjct: 598 SGHRFKLYIPKITCSIAFSFSGVRCPGRGEPYSEEAISVMRRRIMQRDVEIEVETVDRTG 657
Query: 658 TFLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKLKIWENFVE 717
TFLGS+WESRTNVA LLEAGLAK+QTSFG+DRI E HLL++AE+SAK QKLKIWEN+VE
Sbjct: 658 TFLGSMWESRTNVATVLLEAGLAKMQTSFGADRIAEAHLLEQAERSAKNQKLKIWENYVE 717
Query: 718 GEEVSNGAN--VESKQQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALNLKEAPV 775
GEEVSNG VE++Q+E LKV+VTEVLGGG+FYVQ+ GDQKIASIQ QLA+L++K+AP+
Sbjct: 718 GEEVSNGNTNTVETRQKETLKVVVTEVLGGGRFYVQSAGDQKIASIQNQLASLSIKDAPI 777
Query: 776 LGAFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQL 835
+G+F+PK+GDIVL F D SW RAM+V PR V+SP + FEVFYIDYGNQE V YS +
Sbjct: 778 IGSFNPKRGDIVLAQFSLDNSWNRAMIVTAPRAAVQSPDEKFEVFYIDYGNQETVPYSAI 837
Query: 836 RPLDQSVSAAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTS 895
RP+D SVSAAPGLAQLC LAYIK PSLE+DFG EA EYL +TL SGKEF+A +EERDTS
Sbjct: 838 RPIDPSVSAAPGLAQLCRLAYIKVPSLEDDFGPEAGEYLHTVTLGSGKEFKAVIEERDTS 897
Query: 896 GGKVKGQGTGTILAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQ 955
GGKVKGQGTGT VTL+AVD EISVNAAMLQEG+ARMEKR +W K ++ LD+L+KFQ
Sbjct: 898 GGKVKGQGTGTEFVVTLIAVDDEISVNAAMLQEGIARMEKRQKWGHKGKQAALDALEKFQ 957
Query: 956 DDARKERRGMWQYGDVESDDEDTAPPARKAGAGRR 990
++ARK R G+WQYGD+ESDDEDT PARK GRR
Sbjct: 958 EEARKSRIGIWQYGDIESDDEDTG-PARKPAGGRR 991
>emb|CAB87924.1| putative protein [Arabidopsis thaliana] gi|11358274|pir||T49874
hypothetical protein T2I1.60 - Arabidopsis thaliana
Length = 1051
Score = 1424 bits (3687), Expect = 0.0
Identities = 726/1000 (72%), Positives = 843/1000 (83%), Gaps = 19/1000 (1%)
Query: 5 ATGATG-WYRGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLARRGGVDEPF 63
ATGA W +GRVKAV SGDCLVI A++ ++ GP PEK+IT +SL+ P++ARRGG+DEPF
Sbjct: 2 ATGAENQWLKGRVKAVTSGDCLVITALSHNRAGPPPEKTITFSSLMAPKMARRGGIDEPF 61
Query: 64 AWESREYLRKLCIGKEVTFRVDYNVASI-NRDFGTVFLGEKNVGVLVVSQGWAKVREQGQ 122
AWES+E+LRKLCIGKEV F+VDY V +I R+FG+VFLG +N+ LVV GWAKVRE GQ
Sbjct: 62 AWESKEFLRKLCIGKEVAFKVDYKVEAIAGREFGSVFLGNENLAKLVVKTGWAKVREPGQ 121
Query: 123 Q-KGEVSPYLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGLLA 181
Q + +VSPY+ ELL+LEE AKQEG GRWSKVPGAAEASIRNLPPSAIGD++ FDAMGLLA
Sbjct: 122 QNQDKVSPYIKELLQLEELAKQEGYGRWSKVPGAAEASIRNLPPSAIGDSAGFDAMGLLA 181
Query: 182 ANKGSPMEAIVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELTAD 241
ANKG PME IVEQVRDGST+RVYLLPEFQFVQVFVAG+Q+P MGRR +VVET D
Sbjct: 182 ANKGKPMEGIVEQVRDGSTIRVYLLPEFQFVQVFVAGVQAPSMGRRTTNGSVVET--VPD 239
Query: 242 ENDGDVPGEPRPPLTSAQRLAVS--SSTETAADPFGPDAKFFTEMRVLNRDVRIVLEGVD 299
E +GDV E R PLT+AQRLA S SS E ++DPF +AK+FTE RVL+RDVRIVLEGVD
Sbjct: 240 EPNGDVSAESRGPLTTAQRLAASAASSVEVSSDPFATEAKYFTEHRVLSRDVRIVLEGVD 299
Query: 300 KFSNLIGSVYYPDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRL 359
KF+NLIGSV+Y DGE+ KDL LELVENG AK+VEWSANMMEEEAK++LK AEL+ KK ++
Sbjct: 300 KFNNLIGSVHYSDGETVKDLGLELVENGLAKFVEWSANMMEEEAKKKLKAAELQCKKDKV 359
Query: 360 RMWTNYVPPASNSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCP 419
+MW NYVPPA+NSKAIH+QNFTGKVVEVVSGDC+IVADD++P+GSP AERRV LSSIR P
Sbjct: 360 KMWANYVPPATNSKAIHDQNFTGKVVEVVSGDCLIVADDAVPFGSPAAERRVCLSSIRSP 419
Query: 420 KVGNPRRDEKPAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADSRVMD 479
K+GNPRR+EKPAPYAREA+EFLR RL+G+QV V+MEYSRK+ DG S AAD R MD
Sbjct: 420 KMGNPRREEKPAPYAREAREFLRQRLIGKQVIVQMEYSRKVTQGDGPTT-SGAAD-RFMD 477
Query: 480 FGSVFLLSATKADSDDT--PSSIPSAGSQPTGVNVGELVVGRGFGTVIRHRDFEERSNYY 537
FGSVFL SA KADSD+ P + AGSQP GVN+ ELV+ RGFG V+RHRDFEERSN+Y
Sbjct: 478 FGSVFLPSAAKADSDEVTAPPAAAIAGSQPVGVNIAELVLVRGFGNVVRHRDFEERSNHY 537
Query: 538 DALLTAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEYVL 597
DALL AE+RAL+G+KGIHSAK+ P MHITDLT ++AKKAKDFLP L R RRIPAVVEYVL
Sbjct: 538 DALLAAEARALAGKKGIHSAKESPAMHITDLTVSAAKKAKDFLPSLQRIRRIPAVVEYVL 597
Query: 598 SGHRFKLLIPKETCSIAFALSGVRCPGRGEPYSEEAIALMRRKIMQRDVEIEVETVDRNG 657
SGHRFKL IPK TCSIAF+ SGVRCPGRGEPYSEEAI++MRR+IMQRDVEIEVETVDR G
Sbjct: 598 SGHRFKLYIPKITCSIAFSFSGVRCPGRGEPYSEEAISVMRRRIMQRDVEIEVETVDRTG 657
Query: 658 TFLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKLKIWENFVE 717
TFLGS+WESRTNVA LLEAGLAK+QTSFG+DRI E HLL++AE+SAK QKLKIWEN+VE
Sbjct: 658 TFLGSMWESRTNVATVLLEAGLAKMQTSFGADRIAEAHLLEQAERSAKNQKLKIWENYVE 717
Query: 718 GEEVSNG--ANVESKQQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALNLKEAPV 775
GEEVSNG VE++Q+E LKV+VTEVLGGG+FYVQ+ GDQKIASIQ QLA+L++K+AP+
Sbjct: 718 GEEVSNGNTNTVETRQKETLKVVVTEVLGGGRFYVQSAGDQKIASIQNQLASLSIKDAPI 777
Query: 776 LGAFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQL 835
+G+F+PK+GDIVL F D SW RAM+V PR V+SP + FEVFYIDYGNQE V YS +
Sbjct: 778 IGSFNPKRGDIVLAQFSLDNSWNRAMIVTAPRAAVQSPDEKFEVFYIDYGNQETVPYSAI 837
Query: 836 RPLDQSVSAAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTS 895
RP+D SVSAAPGLAQLC LAYIK PSLE+DFG EA EYL +TL SGKEF+A +EERDTS
Sbjct: 838 RPIDPSVSAAPGLAQLCRLAYIKVPSLEDDFGPEAGEYLHTVTLGSGKEFKAVIEERDTS 897
Query: 896 GGKVKGQGTGTILAVTLVAVDAEISVNAAML-----QEGLARMEKRNRWDRKERKVGLDS 950
GGKVKGQGTGT VTL+AVD EISVNAAML QEG+ARMEKR +W K ++ LD+
Sbjct: 898 GGKVKGQGTGTEFVVTLIAVDDEISVNAAMLQDDDEQEGIARMEKRQKWGHKGKQAALDA 957
Query: 951 LQKFQDDARKERRGMWQYGDVESDDEDTAPPARKAGAGRR 990
L+KFQ++ARK R G+WQYGD+ESDDEDT PARK GRR
Sbjct: 958 LEKFQEEARKSRIGIWQYGDIESDDEDTG-PARKPAGGRR 996
Score = 39.7 bits (91), Expect = 0.50
Identities = 27/62 (43%), Positives = 33/62 (52%), Gaps = 4/62 (6%)
Query: 492 DSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVI----RHRDFEERSNYYDALLTAESRA 547
D D P+ P+ G + V LV+ T + DFEERSN YDALL AE+RA
Sbjct: 982 DEDTGPARKPAGGRRKDMVVGATLVLFSDEVTASSAGSQPADFEERSNLYDALLAAEARA 1041
Query: 548 LS 549
LS
Sbjct: 1042 LS 1043
>dbj|BAB10078.1| transcription factor-like protein [Arabidopsis thaliana]
gi|15240352|ref|NP_200986.1| tudor domain-containing
protein / nuclease family protein [Arabidopsis thaliana]
gi|25083258|gb|AAN72055.1| 100 kDa coactivator - like
protein [Arabidopsis thaliana]
Length = 985
Score = 1406 bits (3639), Expect = 0.0
Identities = 710/996 (71%), Positives = 833/996 (83%), Gaps = 19/996 (1%)
Query: 1 MASAATGATGWYRGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLARRGGVD 60
MA+ A W +GRVKAV SGDCLVI A+ ++ GP PEK+ITL+SL+ P++ARRGG+D
Sbjct: 1 MATGAATENQWLKGRVKAVTSGDCLVITALTHNRAGPPPEKTITLSSLMAPKMARRGGID 60
Query: 61 EPFAWESREYLRKLCIGKEVTFRVDYNVASI-NRDFGTVFLGEKNVGVLVVSQGWAKVRE 119
EPFAWESRE+LRKLCIGKEV F+VDY V +I R+FG+V+LG +N+ LVV GWAKVR
Sbjct: 61 EPFAWESREFLRKLCIGKEVAFKVDYKVEAIAGREFGSVYLGNENLAKLVVQNGWAKVRR 120
Query: 120 QGQQ-KGEVSPYLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMG 178
GQQ + +VSPY+AEL +LEEQA+QEG GRWSKVPGAAEASIRNLPPSA+GD+ NFDAMG
Sbjct: 121 PGQQNQDKVSPYIAELEQLEEQAQQEGFGRWSKVPGAAEASIRNLPPSAVGDSGNFDAMG 180
Query: 179 LLAANKGSPMEAIVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRR-AAPETVVETE 237
LLAA+KG PME IVEQVRDGST+RVYLLPEFQFVQVFVAG+Q+P MGRR + E VV+ +
Sbjct: 181 LLAASKGKPMEGIVEQVRDGSTIRVYLLPEFQFVQVFVAGLQAPSMGRRQSTQEAVVDPD 240
Query: 238 LTADENDGDVPGEPRPPLTSAQRLAVS--SSTETAADPFGPDAKFFTEMRVLNRDVRIVL 295
+TA N GD E R PLT+AQRLA S SS E ++DPF +AK+FTE+RVLNRDVRIVL
Sbjct: 241 VTATSN-GDASAETRGPLTTAQRLAASAASSVEVSSDPFAMEAKYFTELRVLNRDVRIVL 299
Query: 296 EGVDKFSNLIGSVYYPDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAK 355
EGVDKF+NLIGSVYY DG++ KDL LELVENG AKYVEWSANM++EEAK++LK EL+ K
Sbjct: 300 EGVDKFNNLIGSVYYSDGDTVKDLGLELVENGLAKYVEWSANMLDEEAKKKLKATELQCK 359
Query: 356 KSRLRMWTNYVPPASNSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSS 415
K+R++MW NYVPPASNSKAIH+QNFTGKVVEVVSGDC++VADDSIP+GSP+AERRV LSS
Sbjct: 360 KNRVKMWANYVPPASNSKAIHDQNFTGKVVEVVSGDCLVVADDSIPFGSPMAERRVCLSS 419
Query: 416 IRCPKVGNPRRDEKPAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADS 475
IR PK+GNPRR+EKPAPYAREAKEFLR +L+G +V V+MEYSRKI P DG V + A
Sbjct: 420 IRSPKMGNPRREEKPAPYAREAKEFLRQKLIGMEVIVQMEYSRKISPGDG--VTTSGAGD 477
Query: 476 RVMDFGSVFLLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVIRHRDFEERSN 535
RVMDFGSVFL S TK D+ ++ P G N+ EL++ RG GTV+RHRDFEERSN
Sbjct: 478 RVMDFGSVFLPSPTKGDTAVAAAATP-------GANIAELIISRGLGTVVRHRDFEERSN 530
Query: 536 YYDALLTAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEY 595
+YDALL AE+RA++G+K IHSAKD P +HI DLT SAKKAKDFLP L R +I AVVEY
Sbjct: 531 HYDALLAAEARAIAGKKNIHSAKDSPALHIADLTVASAKKAKDFLPSLQRINQISAVVEY 590
Query: 596 VLSGHRFKLLIPKETCSIAFALSGVRCPGRGEPYSEEAIALMRRKIMQRDVEIEVETVDR 655
VLSGHRFKL IPKE+CSIAFA SGVRCPGRGEPYSEEAIALMRRKIMQRDVEI VE VDR
Sbjct: 591 VLSGHRFKLYIPKESCSIAFAFSGVRCPGRGEPYSEEAIALMRRKIMQRDVEIVVENVDR 650
Query: 656 NGTFLGSLWE--SRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKLKIWE 713
GTFLGS+WE S+TN LLEAGLAK+QT FG+DRIPE H+L+ AE+SAK QKLKIWE
Sbjct: 651 TGTFLGSMWEKNSKTNAGTYLLEAGLAKMQTGFGADRIPEAHILEMAERSAKNQKLKIWE 710
Query: 714 NFVEGEEVSNGAN-VESKQQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALNLKE 772
N+VEGEEV NG++ VE++Q+E LKV+VTEVLGGG+FYVQTVGDQK+ASIQ QLAAL+LK+
Sbjct: 711 NYVEGEEVVNGSSKVETRQKETLKVVVTEVLGGGRFYVQTVGDQKVASIQNQLAALSLKD 770
Query: 773 APVLGAFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAY 832
AP++G+F+PKKGDIVL F D SW RAM+VN PRG V+SP++ FEVFYIDYGNQE V Y
Sbjct: 771 APIIGSFNPKKGDIVLAQFSLDNSWNRAMIVNGPRGAVQSPEEEFEVFYIDYGNQEIVPY 830
Query: 833 SQLRPLDQSVSAAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEER 892
S +RP+D SVS+APGLAQLC LAYIK P EEDFG++A EYL +TL SGKEFRA VEER
Sbjct: 831 SAIRPVDPSVSSAPGLAQLCRLAYIKVPGKEEDFGRDAGEYLHTVTLESGKEFRAVVEER 890
Query: 893 DTSGGKVKGQGTGTILAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQ 952
DTSGGKVKGQGTGT L VTL+AVD EISVNAAMLQEG+ARMEKR RW+ K+++ LD+L+
Sbjct: 891 DTSGGKVKGQGTGTELVVTLIAVDDEISVNAAMLQEGIARMEKRRRWEPKDKQAALDALE 950
Query: 953 KFQDDARKERRGMWQYGDVESDDEDTAPPARKAGAG 988
KFQD+ARK R G+W+YGD++SDDED P RK G G
Sbjct: 951 KFQDEARKSRTGIWEYGDIQSDDEDNV-PVRKPGRG 985
>ref|XP_466019.1| RNA binding protein Rp120 [Oryza sativa (japonica cultivar-group)]
gi|49388930|dbj|BAD26152.1| RNA binding protein Rp120
[Oryza sativa (japonica cultivar-group)]
gi|49388258|dbj|BAD25376.1| RNA binding protein Rp120
[Oryza sativa (japonica cultivar-group)]
Length = 986
Score = 1351 bits (3496), Expect = 0.0
Identities = 688/992 (69%), Positives = 816/992 (81%), Gaps = 11/992 (1%)
Query: 3 SAATGATGWYRGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLARRGGVDEP 62
++ATGA+GW RG+VK V SGDCL+I+ + P PEKSITL+ L+ PRLARRGGVDEP
Sbjct: 2 ASATGASGWLRGKVKGVTSGDCLLIMGSTKADVPP-PEKSITLSYLMAPRLARRGGVDEP 60
Query: 63 FAWESREYLRKLCIGKEVTFRVDYNVASINRDFGTVFLGEKNVGVLVVSQGWAKVREQGQ 122
FAWESRE+LRKLCIGKEVTFRVDY ++ R+FGTV+LG+KNV +++ GWA+V+EQG
Sbjct: 61 FAWESREFLRKLCIGKEVTFRVDYTAPNVGREFGTVYLGDKNVAYSIIAAGWARVKEQGP 120
Query: 123 QKGEVSPYLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGLLAA 182
+ GE SPYL ELLRLEE AKQ+GLGRWSK PGAAE SIR+LPPSAIG+AS FDA G A
Sbjct: 121 KGGEPSPYLTELLRLEEVAKQQGLGRWSKEPGAAEESIRDLPPSAIGEASGFDAKGFAVA 180
Query: 183 NKGSPMEAIVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVET-ELTAD 241
NKG +EAIVEQVRDGST+RVYLLP FQFVQ++VAG+QSP MGRR TVV E TAD
Sbjct: 181 NKGKSLEAIVEQVRDGSTVRVYLLPSFQFVQIYVAGVQSPSMGRRPPNPTVVAAAESTAD 240
Query: 242 --ENDGDVPGEPRPPLTSAQRLAVSS-STETAADPFGPDAKFFTEMRVLNRDVRIVLEGV 298
N GD P P LT+AQRLA ++ STE D FG +AK FTE RVLNRDVRIV+EG
Sbjct: 241 GATNGGDSEEAPAP-LTTAQRLAAAAVSTEIPPDRFGIEAKHFTETRVLNRDVRIVVEGT 299
Query: 299 DKFSNLIGSVYYPDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSR 358
D FSN+IGSVYY DG++ KDLALELVENG AKYVEWSANMM+ +AK +LK AEL+AKK +
Sbjct: 300 DSFSNIIGSVYYSDGDTLKDLALELVENGLAKYVEWSANMMDVDAKIKLKNAELQAKKDQ 359
Query: 359 LRMWTNYVPPASNSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRC 418
LR+WT + PP +NSK IH+Q FTGKVVEVVSGDCIIVADD+ PYGSP AERRVNLSSIR
Sbjct: 360 LRIWTGFKPPVTNSKPIHDQKFTGKVVEVVSGDCIIVADDAAPYGSPSAERRVNLSSIRA 419
Query: 419 PKVGNPRRDEKPAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADSRVM 478
PK+GNPRRDEKP +AREAKEFLRTRL+G+QV+VEMEYSR+I DG + AD+RV+
Sbjct: 420 PKMGNPRRDEKPDNFAREAKEFLRTRLIGKQVTVEMEYSRRISTVDGQPTTN-TADARVL 478
Query: 479 DFGSVFLLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVIRHRDFEERSNYYD 538
D+GSVFL S ++AD DD SSIPS+G+QP G+N+ E ++ RGF +HRD+EERS+Y+D
Sbjct: 479 DYGSVFLGSPSQADGDDV-SSIPSSGNQP-GINIAETLLSRGFARTSKHRDYEERSHYFD 536
Query: 539 ALLTAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEYVLS 598
LL AESRA +KG+HSAK+ PVMHITDLTT SAKKA+DFLPFL R+RR A+VEYV S
Sbjct: 537 LLLAAESRAEKAKKGVHSAKESPVMHITDLTTVSAKKARDFLPFLQRNRRHSAIVEYVFS 596
Query: 599 GHRFKLLIPKETCSIAFALSGVRCPGRGEPYSEEAIALMRRKIMQRDVEIEVETVDRNGT 658
GHRFKL IPKETCSIAF+ SGVRCPG+ EPYS EAIALMRR+I+QRDVEIEVE VDR GT
Sbjct: 597 GHRFKLTIPKETCSIAFSFSGVRCPGKDEPYSNEAIALMRRRILQRDVEIEVEAVDRTGT 656
Query: 659 FLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKLKIWENFVEG 718
FLGSLWES+TN+A LLEAGLAKL +SFG DRIP+ ++L RAEQSAK+QKLKIWEN+VEG
Sbjct: 657 FLGSLWESKTNMASVLLEAGLAKL-SSFGLDRIPDANVLMRAEQSAKQQKLKIWENYVEG 715
Query: 719 EEVSNGANVESKQQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALNLKEAPVLGA 778
EEVSNG+ ESKQ+E+LKV+VTEVLGGGKFYVQTVGD ++ASIQQQLA+L LK+APV+GA
Sbjct: 716 EEVSNGSASESKQKEILKVVVTEVLGGGKFYVQTVGDHRVASIQQQLASLKLKDAPVIGA 775
Query: 779 FSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPL 838
F+P KG+IVL F AD SW RAM+VN PRG V S D FEVFYIDYGNQE V YS++RP
Sbjct: 776 FNPVKGEIVLAQFSADNSWNRAMIVNGPRGAVSSQDDKFEVFYIDYGNQEVVPYSRIRPA 835
Query: 839 DQSVSAAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTSGGK 898
D S+S++P LAQLCSLA+IK P+LE+DFG EAA YL++ L+S K++RA +EERDTSGGK
Sbjct: 836 DPSISSSPALAQLCSLAFIKVPNLEDDFGHEAAVYLNDCLLNSQKQYRAMIEERDTSGGK 895
Query: 899 VKGQGTGTILAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQDDA 958
KGQGTGTIL VTLV + E S+NA ML+EGLAR+E+ RWD +ERK L +L++FQ+ A
Sbjct: 896 SKGQGTGTILIVTLVDAETETSINATMLEEGLARLERSKRWDTRERKAALQNLEQFQEKA 955
Query: 959 RKERRGMWQYGDVESDDEDTAPPARKAGAGRR 990
+KER +WQYGDVESD+E+ AP AR+ G GRR
Sbjct: 956 KKERLQIWQYGDVESDEEEQAPAARRTG-GRR 986
>gb|AAP85378.1| RNA binding protein Rp120 [Oryza sativa (japonica cultivar-group)]
gi|32492578|gb|AAP85377.1| RNA binding protein Rp120
[Oryza sativa (japonica cultivar-group)]
Length = 986
Score = 1346 bits (3484), Expect = 0.0
Identities = 686/992 (69%), Positives = 814/992 (81%), Gaps = 11/992 (1%)
Query: 3 SAATGATGWYRGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLARRGGVDEP 62
++ATGA+GW RG+VK V SGDCL+I+ + P PEKSITL+ L+ PRLARRGGVDEP
Sbjct: 2 ASATGASGWLRGKVKGVTSGDCLLIMGSTKADVPP-PEKSITLSYLMAPRLARRGGVDEP 60
Query: 63 FAWESREYLRKLCIGKEVTFRVDYNVASINRDFGTVFLGEKNVGVLVVSQGWAKVREQGQ 122
FAWESRE+LRKLCIGKEVTFRVDY ++ R+FGTV+LG+KNV +++ GWA+V+EQG
Sbjct: 61 FAWESREFLRKLCIGKEVTFRVDYTAPNVGREFGTVYLGDKNVAYSIIAAGWARVKEQGP 120
Query: 123 QKGEVSPYLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGLLAA 182
+ GE SPYL ELLRLEE AKQ+GLGRWSK PGAAE SIR+LPPSAIG+AS FDA G A
Sbjct: 121 KGGEPSPYLTELLRLEEVAKQQGLGRWSKEPGAAEESIRDLPPSAIGEASGFDAKGFAVA 180
Query: 183 NKGSPMEAIVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVET-ELTAD 241
NKG +EAIVEQVRDGST+RVYLLP FQFVQ++VAG+QSP MGRR TVV E TAD
Sbjct: 181 NKGKSLEAIVEQVRDGSTVRVYLLPSFQFVQIYVAGVQSPSMGRRPPNPTVVAAAESTAD 240
Query: 242 --ENDGDVPGEPRPPLTSAQRLAVSS-STETAADPFGPDAKFFTEMRVLNRDVRIVLEGV 298
N GD P P LT+AQRLA ++ STE D FG +AK FTE VLNRDVRIV+EG
Sbjct: 241 GATNGGDSEEAPAP-LTTAQRLAAAAVSTEIPPDRFGIEAKHFTETHVLNRDVRIVVEGT 299
Query: 299 DKFSNLIGSVYYPDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSR 358
D FSN+IGSVYY DG++ KDLALELVENG AKYVEWSANMM+ +AK +LK AEL+AKK +
Sbjct: 300 DSFSNIIGSVYYSDGDTLKDLALELVENGLAKYVEWSANMMDVDAKIKLKNAELQAKKDQ 359
Query: 359 LRMWTNYVPPASNSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRC 418
LR+WT + PP +NSK IH+Q FTGKVVEVVSGDCIIVADD+ PYGSP AERRVNLSSIR
Sbjct: 360 LRIWTGFKPPVTNSKPIHDQKFTGKVVEVVSGDCIIVADDAAPYGSPSAERRVNLSSIRA 419
Query: 419 PKVGNPRRDEKPAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADSRVM 478
PK+GNPRRDEKP +AREAKEFLRTRL+G+QV+VEMEYSR+I DG + AD+RV+
Sbjct: 420 PKMGNPRRDEKPDNFAREAKEFLRTRLIGKQVTVEMEYSRRISTVDGQPTTN-TADARVL 478
Query: 479 DFGSVFLLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVIRHRDFEERSNYYD 538
D+GSVFL S ++AD DD SSIPS+G+QP G+N+ E ++ RGF +HRD+E+RS+Y+D
Sbjct: 479 DYGSVFLGSPSQADGDDV-SSIPSSGNQP-GINIAETLLSRGFAKTSKHRDYEKRSHYFD 536
Query: 539 ALLTAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEYVLS 598
LL AESRA +KG+HSAK PVMHITDLTT SAKKA+DFLPFL R+RR A+VEYV S
Sbjct: 537 LLLAAESRAEKAKKGVHSAKKSPVMHITDLTTVSAKKARDFLPFLQRNRRHSAIVEYVFS 596
Query: 599 GHRFKLLIPKETCSIAFALSGVRCPGRGEPYSEEAIALMRRKIMQRDVEIEVETVDRNGT 658
GHRFKL IPKETCSIAF+ SGVRCPG+ EPYS EAIALMRR+I+QRDVEIEVE VDR GT
Sbjct: 597 GHRFKLTIPKETCSIAFSFSGVRCPGKDEPYSNEAIALMRRRILQRDVEIEVEAVDRTGT 656
Query: 659 FLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKLKIWENFVEG 718
FLGSLWES+TN+A LLEAGLAKL +SFG DRIP+ ++L RAEQSAK+QKLKIWEN+VEG
Sbjct: 657 FLGSLWESKTNMASVLLEAGLAKL-SSFGLDRIPDANVLMRAEQSAKQQKLKIWENYVEG 715
Query: 719 EEVSNGANVESKQQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALNLKEAPVLGA 778
EEVSNG+ ESKQ+E+LKV+VTEVLGGGKFYVQTVGD ++ASIQQQLA+L LK+APV+GA
Sbjct: 716 EEVSNGSASESKQKEILKVVVTEVLGGGKFYVQTVGDHRVASIQQQLASLKLKDAPVIGA 775
Query: 779 FSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPL 838
F+P KG+IVL F AD SW RAM+VN PRG V S D FEVFYIDYGNQE V YS++RP
Sbjct: 776 FNPVKGEIVLAQFSADNSWNRAMIVNGPRGAVSSQDDKFEVFYIDYGNQEVVPYSRIRPA 835
Query: 839 DQSVSAAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTSGGK 898
D S+S++P LAQLCSLA+IK P+LE+DFG EAA YL++ L+S K++RA +EERDTSGGK
Sbjct: 836 DPSISSSPALAQLCSLAFIKVPNLEDDFGHEAAVYLNDCLLNSQKQYRAMIEERDTSGGK 895
Query: 899 VKGQGTGTILAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQDDA 958
KGQGTGTIL VTLV + E S+NA ML+EGLAR+E+ RWD +ERK L +L++FQ+ A
Sbjct: 896 SKGQGTGTILIVTLVDAETETSINATMLEEGLARLERSKRWDTRERKAALQNLEQFQEKA 955
Query: 959 RKERRGMWQYGDVESDDEDTAPPARKAGAGRR 990
+KER +WQYGDVESD+E+ AP AR+ G GRR
Sbjct: 956 KKERLQIWQYGDVESDEEEQAPAARRTG-GRR 986
>dbj|BAB32793.1| 110 kDa 4SNc-Tudor domain protein [Pisum sativum]
Length = 699
Score = 1169 bits (3024), Expect = 0.0
Identities = 587/702 (83%), Positives = 640/702 (90%), Gaps = 3/702 (0%)
Query: 289 RDVRIVLEGVDKFSNLIGSVYYPDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLK 348
RDVRIVLEGVDKFSNLIGSVYYPDGESAKD LELVENG+AKYVEWSA+MMEE+AKR+LK
Sbjct: 1 RDVRIVLEGVDKFSNLIGSVYYPDGESAKDWPLELVENGFAKYVEWSAHMMEEDAKRKLK 60
Query: 349 TAELEAKKSRLRMWTNYVPPASNSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAE 408
+AELEAKKSRLR+WTNYVPP SNSKAIH+QN TGK+VEVVSGDC+IVADDSIPYGSP AE
Sbjct: 61 SAELEAKKSRLRIWTNYVPPVSNSKAIHDQNLTGKLVEVVSGDCVIVADDSIPYGSPQAE 120
Query: 409 RRVNLSSIRCPKVGNPRRDEKPAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAV 468
RRVNLSSIRCPK+GNPRRDEKPAPYAREAKEFLRTRL+GRQV+V+MEYSRK+ P D +
Sbjct: 121 RRVNLSSIRCPKMGNPRRDEKPAPYAREAKEFLRTRLIGRQVNVQMEYSRKVGPVDAAGA 180
Query: 469 PSPAADSRVMDFGSVFLLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVIRHR 528
P A D RVMDFGSVFL S+ KAD+D PS+ A S+ G+NVGELV+GRGFGTVIRHR
Sbjct: 181 PLGAGD-RVMDFGSVFLSSSGKADNDQAPSAAAPASSK-LGLNVGELVIGRGFGTVIRHR 238
Query: 529 DFEERSNYYDALLTAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRR 588
DFEERSN+YDALL AESRA+SGRKGIHSAKDPPVMHITDLTT SAKKAKDF+PFLHRSRR
Sbjct: 239 DFEERSNFYDALLAAESRAISGRKGIHSAKDPPVMHITDLTTASAKKAKDFMPFLHRSRR 298
Query: 589 IPAVVEYVLSGHRFKLLIPKETCSIAFALSGVRCPGRGEPYSEEAIALMRRKIMQRDVEI 648
+PAVVEYVLSGHRFKLLIPKETCSIAFA SGVRCPGR EPYS+EAIALMRR+IMQRDVEI
Sbjct: 299 VPAVVEYVLSGHRFKLLIPKETCSIAFAFSGVRCPGREEPYSDEAIALMRRRIMQRDVEI 358
Query: 649 EVETVDRNGTFLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQK 708
EVETVDR GTFLG LWES+TN A+ LLEAGLAKLQT+FGSDRIP L++ EQSAK +K
Sbjct: 359 EVETVDRTGTFLGPLWESKTNGAVALLEAGLAKLQTTFGSDRIPGSSCLEQPEQSAKSKK 418
Query: 709 LKIWENFVEGEEVSNGANVESKQQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAAL 768
LKIWENFVEGE V +GANVE+KQQEVLKV VTEVLGGGKFYVQTVGDQKIASIQ QLA+L
Sbjct: 419 LKIWENFVEGEVVPSGANVETKQQEVLKVTVTEVLGGGKFYVQTVGDQKIASIQNQLASL 478
Query: 769 NLKEAPVLGAFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQE 828
NLKEAPV+GAF+PKKGDIVLCYF AD SWYRAMVVNTPRGPVES KD+FEVFY+DYGNQE
Sbjct: 479 NLKEAPVIGAFNPKKGDIVLCYFRADTSWYRAMVVNTPRGPVESSKDVFEVFYLDYGNQE 538
Query: 829 QVAYSQLRPLDQSVSAAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQ 888
+V YSQLRPLD SVS APGLAQLCSLAYIK P+LEEDFGQEAAEYLSELTLSSGKEFRA
Sbjct: 539 EVPYSQLRPLDPSVSLAPGLAQLCSLAYIKIPNLEEDFGQEAAEYLSELTLSSGKEFRAM 598
Query: 889 VEERDTSGGKVKGQGTGTILAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKVGL 948
VEERDT+GGKVKGQGTG ++AVTLVAVDAEISVNAAMLQEGLARMEKRNRWD+ RK L
Sbjct: 599 VEERDTTGGKVKGQGTGPVIAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDKSARKQAL 658
Query: 949 DSLQKFQDDARKERRGMWQYGDVESDDEDTAPPARKAGAGRR 990
D+L+ FQ +AR RRG+WQYGD++SDDEDTAPP + AG GRR
Sbjct: 659 DNLEMFQGEARTSRRGIWQYGDIQSDDEDTAPPRKPAG-GRR 699
Score = 75.9 bits (185), Expect = 6e-12
Identities = 102/478 (21%), Positives = 185/478 (38%), Gaps = 103/478 (21%)
Query: 14 GRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLA--RRGGVDEPFAWESREYL 71
G++ V SGDC+++ + P E+ + L+S+ P++ RR P+A E++E+L
Sbjct: 94 GKLVEVVSGDCVIVADDSIPYGSPQAERRVNLSSIRCPKMGNPRRDEKPAPYAREAKEFL 153
Query: 72 RKLCIGKEVTFRVDYN--VASIN------------RDFGTVFLGEK-------------- 103
R IG++V +++Y+ V ++ DFG+VFL
Sbjct: 154 RTRLIGRQVNVQMEYSRKVGPVDAAGAPLGAGDRVMDFGSVFLSSSGKADNDQAPSAAAP 213
Query: 104 -------NVGVLVVSQGWAKVREQGQQKGEVSPYLAELLRLEEQAKQEGLGRWSKVPGAA 156
NVG LV+ +G+ V + E S + LL E +A G + A
Sbjct: 214 ASSKLGLNVGELVIGRGFGTVIRH-RDFEERSNFYDALLAAESRAISGRKG----IHSAK 268
Query: 157 EASIRNLPPSAIGDASNF-DAMGLLAANKGSPMEAIVEQVRDGSTLRVYLLPEFQFVQVF 215
+ + ++ A D M L ++ P A+VE V G ++ + E +
Sbjct: 269 DPPVMHITDLTTASAKKAKDFMPFLHRSRRVP--AVVEYVLSGHRFKLLIPKETCSIAFA 326
Query: 216 VAGIQSPQMGRRAAPETVVETELTADENDGDVPGEPRPPLTSAQRLAVSSSTETAADPFG 275
+G++ P GR +P+
Sbjct: 327 FSGVRCP--GRE--------------------------------------------EPYS 340
Query: 276 PDAKFFTEMRVLNRDVRIVLEGVDKFSNLIGSVYYPDGESAKDLALELVENGYAKYVEWS 335
+A R++ RDV I +E VD+ +G ++ ES + A+ L+E G AK ++ +
Sbjct: 341 DEAIALMRRRIMQRDVEIEVETVDRTGTFLGPLW----ESKTNGAVALLEAGLAK-LQTT 395
Query: 336 ANMMEEEAKRRLKTAELEAKKSRLRMWTNY-----VPPASNSKAIHNQNFTGKVVEVVSG 390
L+ E AK +L++W N+ VP +N + + V EV+ G
Sbjct: 396 FGSDRIPGSSCLEQPEQSAKSKKLKIWENFVEGEVVPSGANVETKQQEVLKVTVTEVLGG 455
Query: 391 DCIIVADDSIPYGSPLAERRVNLSSIRCPKVG--NPRRDEKPAPYAREAKEFLRTRLL 446
V + + + +L+ P +G NP++ + Y R + R ++
Sbjct: 456 GKFYVQTVGDQKIASIQNQLASLNLKEAPVIGAFNPKKGDIVLCYFRADTSWYRAMVV 513
>ref|XP_472372.1| OSJNBb0012E08.11 [Oryza sativa (japonica cultivar-group)]
gi|21740629|emb|CAD40787.1| OSJNBb0012E08.11 [Oryza
sativa (japonica cultivar-group)]
Length = 1056
Score = 1014 bits (2622), Expect = 0.0
Identities = 548/1051 (52%), Positives = 727/1051 (69%), Gaps = 80/1051 (7%)
Query: 4 AATGATGWYRGRVKAVPSGDCLVIVAVASSKPG-PLPEKSITLASLITPRLARRGGVDEP 62
A A ++G+VK+VPSGD +VI+ + ++ P PE S+TL+ +I P LARRGG+DEP
Sbjct: 6 AVPAAAPVWKGKVKSVPSGDTVVIMDTSKAEEVIPPPEMSVTLSCIIAPNLARRGGMDEP 65
Query: 63 FAWESREYLRKLCIGKEVTFRVDYNVASINRDFGTVFLGEKNVGVLVVSQGWAKVREQGQ 122
FAWESREYLR+L IG++V FRV+Y + R FG VF EKNV +VV+ G AKV+EQGQ
Sbjct: 66 FAWESREYLRRLLIGQDVRFRVEYTASPSGRKFGMVFFAEKNVACMVVAAGLAKVKEQGQ 125
Query: 123 QKGEVSPYLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGLLAA 182
KGE+SPY+AELLRLE A+ +GLGRWSK+PGA E+SIR+LPPS IGD +FDA G +A
Sbjct: 126 -KGEISPYVAELLRLETIARDQGLGRWSKLPGALESSIRDLPPSTIGDGRSFDAKGFVAE 184
Query: 183 NKGSPMEAIVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELTADE 242
NKG +EAIVE VRDGST+RV+L+P F +VQV+VAG+Q+P MGRRA P + +
Sbjct: 185 NKGKSLEAIVEHVRDGSTIRVHLIPSFLYVQVYVAGVQAPSMGRRATPPPNAQAGVGNGA 244
Query: 243 NDGDVPGEPRPPLTSAQRLAVSSS--TETAADPFGPDAKFFTEMRVLNRDVRIVLEGVDK 300
+G+ P P + +AQ+L S+ +E D FG +AK FTE RVLNR+VRIV+EG D
Sbjct: 245 ANGEASTTPAP-MAAAQKLLASADIYSEVPPDRFGQEAKHFTETRVLNREVRIVMEGTDN 303
Query: 301 FSNLIGSVYYPDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRLR 360
F+N+ GSVYY DG+ KDLAL+LV+NG AKYVEWSAN+++ + K +L+ A+L+ KK +LR
Sbjct: 304 FNNIFGSVYYSDGDVVKDLALDLVQNGLAKYVEWSANVLDPQLKTKLRNADLQVKKEQLR 363
Query: 361 MWTNYVPPASNSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCPK 420
+WT + PP +N+K IHNQ FTGKV+EVV+G C+++ADD+ PYGSP AERRVNLSSIR PK
Sbjct: 364 IWTGFKPPVTNTKPIHNQKFTGKVIEVVNGYCLVIADDAEPYGSPSAERRVNLSSIRPPK 423
Query: 421 VGNPRRDEKPA-PYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDG--SAVPSPAADSRV 477
P + K + +AR AKEFLRTRL+G+QV+V MEYSR+I DG + + + ++RV
Sbjct: 424 FEKPSEENKSSEQFARTAKEFLRTRLIGKQVNVSMEYSRRINIADGQIAGPRTNSTETRV 483
Query: 478 MDFGSVFLLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVIRHRDFEERSNYY 537
+++GSVFL S++ AD + SS S+ +Q G+NV L+V RG + RHRD+E+RS++Y
Sbjct: 484 LEYGSVFLPSSSHADGETATSSSDSSNNQ-LGINVAALLVSRGLADITRHRDYEDRSHHY 542
Query: 538 DALLTAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEYVL 597
DAL+ A +RA +KG HS K+ P +H+TDLT KKAK+FL L RSRR A+VEYV
Sbjct: 543 DALIAAHARAEKTKKGYHSKKECPPIHMTDLTRV-PKKAKEFLHLLQRSRRHSAIVEYVF 601
Query: 598 SGHRFKLLIPKETCSIAFALSGVRCPGRGEPYSEEAIALMRRKIMQRDVEIEVETVDRNG 657
SGHRFK+ IPKETC+IAFALSGVRCPGR EPYS+EAI +MRR+I+QR+VEIE+ TVDR G
Sbjct: 602 SGHRFKVTIPKETCTIAFALSGVRCPGRDEPYSDEAITMMRRRILQRNVEIEINTVDRTG 661
Query: 658 TFLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKLKIWENFVE 717
TFLGSLWES NVA LLEAGLAK+ +SF D++P+ +L + E+ AK++KLK+WEN+ E
Sbjct: 662 TFLGSLWESNINVASVLLEAGLAKI-SSFAVDKMPDAQVLLKTEKIAKQKKLKVWENY-E 719
Query: 718 GEEVSNGANVESKQQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALNLKEAP--- 774
EVSN + ++K E LKVIVTEVLG G FYVQ + D+ + ++ QLA+L++K+ P
Sbjct: 720 EVEVSNVSLYDNK--ETLKVIVTEVLGAGMFYVQALADEHVEFVRHQLASLDIKDDPAEA 777
Query: 775 ---------------------VLGAFSPK-----------------------------KG 784
L A P KG
Sbjct: 778 LEVKELETSKEVATLTKDLPETLDAEDPSSDVAKDESVTSKDIDPLPDDSNTAPFTPMKG 837
Query: 785 DIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVSA 844
++VL F D SW RAM++ +G VE P+ FEVFYIDYGNQE V +S LRP++ S+S+
Sbjct: 838 EMVLALFRCDNSWNRAMIIGECQG-VEGPE--FEVFYIDYGNQELVPHSCLRPINLSISS 894
Query: 845 APGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEERD-TSGGKVKGQG 903
P LA+LCSLA++K PSL + GQEAA YL+ + L +G+EF A VEERD SGGK++GQG
Sbjct: 895 IPPLAKLCSLAFVKVPSLNDYLGQEAAMYLNSILLDNGREFEAIVEERDAASGGKLQGQG 954
Query: 904 TGTILAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQDDARKERR 963
TG IL VTL+ + + S+NA ML+ G ++E+R RWD +ER+ + L++FQ+ ARKE+
Sbjct: 955 TGEILGVTLLDSETDNSINAEMLERGYGQLERR-RWDSRERRAAIKKLEEFQEVARKEQL 1013
Query: 964 GMWQYGDVESDDED--------TAPPARKAG 986
G+W + D APP K G
Sbjct: 1014 GVWCPKNARKQGMDENEYPVLARAPPPPKKG 1044
>dbj|BAC06183.1| 110kDa protein HMP [Pisum sativum]
Length = 381
Score = 641 bits (1653), Expect = 0.0
Identities = 320/381 (83%), Positives = 353/381 (91%), Gaps = 2/381 (0%)
Query: 1 MASAATGATGWYRGRVKAVPSGDCLVIVAVASS-KPGPLPEKSITLASLITPRLARRGGV 59
MA+ A G + WY+ +VKAV SGDC+V+V+VA++ K G LPEKSITL+SLI PRLARRGGV
Sbjct: 1 MATTAAGNSAWYKAKVKAVTSGDCVVVVSVAANAKSGVLPEKSITLSSLIAPRLARRGGV 60
Query: 60 DEPFAWESREYLRKLCIGKEVTFRVDYNVASINRDFGTVFLGEKNVGVLVVSQGWAKVRE 119
DE FAWESRE+LRKLCIG+E+TFR+DY V SINR+FGTVFLG+KNV +LVVSQGWAKVRE
Sbjct: 61 DEAFAWESREFLRKLCIGREITFRIDYTVPSINREFGTVFLGDKNVAMLVVSQGWAKVRE 120
Query: 120 QGQQKGEVSPYLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGL 179
QGQQKGEVSP+LAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSA+GDASNFDAMGL
Sbjct: 121 QGQQKGEVSPFLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSALGDASNFDAMGL 180
Query: 180 LAANKGSPMEAIVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELT 239
LA +KG PMEA+VEQVRDGSTLR+YLLPEFQFVQVFVAGIQSPQMGRRAAPETVVE E+T
Sbjct: 181 LAKSKGVPMEALVEQVRDGSTLRIYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVEPEVT 240
Query: 240 ADENDGDVPGEPRPPLTSAQRLAVS-SSTETAADPFGPDAKFFTEMRVLNRDVRIVLEGV 298
D +GD P EPR PLTSAQRLAVS S+ ET+ADPFGPDAKFFTEMRVLNRDVRIVLEGV
Sbjct: 241 VDSTNGDAPAEPRAPLTSAQRLAVSASAAETSADPFGPDAKFFTEMRVLNRDVRIVLEGV 300
Query: 299 DKFSNLIGSVYYPDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSR 358
DKFSNLIGSVYYPDGESAKD LELVENG+AKYVEWSA+MMEE+AKR+LK+AELEAKKSR
Sbjct: 301 DKFSNLIGSVYYPDGESAKDWPLELVENGFAKYVEWSAHMMEEDAKRKLKSAELEAKKSR 360
Query: 359 LRMWTNYVPPASNSKAIHNQN 379
LR+WTNYVPP SNSKAIH+QN
Sbjct: 361 LRIWTNYVPPVSNSKAIHDQN 381
Score = 72.8 bits (177), Expect = 5e-11
Identities = 99/397 (24%), Positives = 161/397 (39%), Gaps = 100/397 (25%)
Query: 380 FTGKVVEVVSGDCIIVADDSIPYGSP-LAERRVNLSSIRCPKVGNPRRDEKPAPYAREAK 438
+ KV V SGDC++V + S L E+ + LSS+ P++ RR +A E++
Sbjct: 12 YKAKVKAVTSGDCVVVVSVAANAKSGVLPEKSITLSSLIAPRLA--RRGGVDEAFAWESR 69
Query: 439 EFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADSRVMDFGSVFLLSATKADSDDTPS 498
EFLR +GR+++ ++Y+ VPS +FG+VFL
Sbjct: 70 EFLRKLCIGREITFRIDYT----------VPSIN-----REFGTVFL------------- 101
Query: 499 SIPSAGSQPTGVNVGELVVGRGFGTVIRH-RDFEERSNYYDALLTAESRALSGRKGIHSA 557
G + NV LVV +G+ V + E S + LL E +A G S
Sbjct: 102 -----GDK----NVAMLVVSQGWAKVREQGQQKGEVSPFLAELLRLEEQAKQEGLGRWS- 151
Query: 558 KDPPVMH--ITDLTTTSAKKAKDF--LPFLHRSRRIP--AVVEYVLSGHRFKLLIPKETC 611
K P I +L ++ A +F + L +S+ +P A+VE V G ++ + E
Sbjct: 152 KVPGAAEASIRNLPPSALGDASNFDAMGLLAKSKGVPMEALVEQVRDGSTLRIYLLPEFQ 211
Query: 612 SIAFALSGVRCPGRG--------------------------------------------- 626
+ ++G++ P G
Sbjct: 212 FVQVFVAGIQSPQMGRRAAPETVVEPEVTVDSTNGDAPAEPRAPLTSAQRLAVSASAAET 271
Query: 627 --EPYSEEAIALMRRKIMQRDVEIEVETVDRNGTFLGSLW----ESRTNVALTLLEAGLA 680
+P+ +A +++ RDV I +E VD+ +GS++ ES + L L+E G A
Sbjct: 272 SADPFGPDAKFFTEMRVLNRDVRIVLEGVDKFSNLIGSVYYPDGESAKDWPLELVENGFA 331
Query: 681 K-LQTSFGSDRIPEFHLLDRAEQSAKKQKLKIWENFV 716
K ++ S L AE AKK +L+IW N+V
Sbjct: 332 KYVEWSAHMMEEDAKRKLKSAELEAKKSRLRIWTNYV 368
>gb|AAU05374.1| SND p102 [Rattus norvegicus] gi|60415342|sp|Q66X93|SND1_RAT
Staphylococcal nuclease domain containing protein 1
(p100 co-activator) (100 kDa coactivator) (SND p102)
(p105 coactivator)
Length = 909
Score = 441 bits (1134), Expect = e-122
Identities = 341/1021 (33%), Positives = 532/1021 (51%), Gaps = 162/1021 (15%)
Query: 1 MASA-ATGATGW------YRGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRL 53
MASA ++G++G RG VK V SG C +IV + GP PE+ I L+++ L
Sbjct: 1 MASAQSSGSSGGPAVPTVQRGIVKMVLSG-CAIIVR-GQPRGGPPPERQINLSNIRAGNL 58
Query: 54 ARRGGV---------DEPFAWESREYLRKLCIGKEVTFRVDYNVASINRDFGTVFLGE-- 102
ARR DEP+A+ +RE+LRK IGKEV F ++ N R++G ++LG+
Sbjct: 59 ARRAAATQPDGKDTPDEPWAFPAREFLRKKLIGKEVCFTIE-NKTPQGREYGMIYLGKDT 117
Query: 103 --KNVGVLVVSQGWAKVREQGQQKGEVSPYLAELLRLEEQAKQEGLGRWSKVPGAAEASI 160
+N+ +V++G A RE + +P L EEQAK G WS+ G +I
Sbjct: 118 NGENIAESLVAEGLASRREGMRAN---NPEQNRLSECEEQAKASKKGMWSE--GNGSHTI 172
Query: 161 RNLPPSAIGDASNFDAMGLLAANKGSPMEAIVEQVRDGSTLRVYLLPEFQFVQVFVAGIQ 220
R+L + I + +F + ++ P+ AI+E VRDGS +R LLP+ V V ++GI+
Sbjct: 173 RDLKYT-IENPRHF-----VDSHHQKPVNAIIEHVRDGSVVRALLLPDHYLVTVMLSGIK 226
Query: 221 SPQMGRRAAPETVVETELTADENDGDVPGEPRPPLTSAQRLAVSSSTETAADPFGPDAKF 280
P R E DG +ET +PF +AKF
Sbjct: 227 CPTFRR---------------ETDG---------------------SET-PEPFAAEAKF 249
Query: 281 FTEMRVLNRDVRIVLEGVDKFSNLIGSVYYPDGESAKDLALELVENGYAKYVEWSANMME 340
FTE R+L RDV+I+LE N++G++ +P+G ++ L++ G+A+ V+WS +
Sbjct: 250 FTESRLLQRDVQIILESCHN-QNILGTILHPNG----NITELLLKEGFARCVDWSIAVYT 304
Query: 341 EEAKRRLKTAELEAKKSRLRMWTNYVPPASNSKAIHNQNFTGKVVEVVSGDCIIVADDSI 400
A+ +L+ AE AK+ RLR+W +YVPP +N ++ F KV++V++ D I+V +S
Sbjct: 305 RGAE-KLRAAERFAKERRLRIWRDYVPPTANLDQ-KDKQFVAKVMQVLNADAIVVKLNSG 362
Query: 401 PYGSPLAERRVNLSSIRCPKVGNPRRDEK--------PAPYAREAKEFLRTRLLGRQVSV 452
Y + ++LSSIR P++ +K PY EA+EFLR +L+G++VSV
Sbjct: 363 DY------KTIHLSSIRPPRLEGDNIQDKNKKLRPLYDIPYMFEAREFLRKKLIGKKVSV 416
Query: 453 EMEYSRKIVPTDGSAVPSPAADSRVMDFGSVFLLSATKADSDDTPSSIPSAGSQPTGVNV 512
++Y R SPA ++ A S+ T +++ G+N+
Sbjct: 417 TVDYIRP---------ASPATET-------------VPAFSERTCATVTIG-----GINI 449
Query: 513 GELVVGRGFGTVIRHR-DFEERSNYYDALLTAESRALSGRKGIHSAKDPPVMHITDLTTT 571
E +V +G TVIR+R D ++RS++YD LL AE+RA+ KG+HS K+ P+ + D+ +
Sbjct: 450 AEALVSKGLATVIRYRQDDDQRSSHYDELLAAEARAIKNGKGLHSKKEVPIHRVADI-SG 508
Query: 572 SAKKAKDFLPFLHRSRRIPAVVEYVLSGHRFKLLIPKETCSIAFALSGVRCP-------- 623
+KAK FLPFL R+ R AVVEYV SG R KL +PKETC I F L+G+ CP
Sbjct: 509 DTQKAKQFLPFLQRAGRSEAVVEYVFSGSRLKLYLPKETCLITFLLAGIECPRGARNLPG 568
Query: 624 --GRGEPYSEEAIALMRRKIMQRDVEIEVETVDRNGTFLGSLWESRTNVALTLLEAGLAK 681
GEP+SEEA + ++QR+VE+EVE++D+ G F+G L N+++ L+E L+K
Sbjct: 569 LVQEGEPFSEEATLFTKELVLQREVEVEVESMDKAGNFIGWLHMDGANLSVLLVEHALSK 628
Query: 682 LQTSFGSDRIPEFHLLDRAEQSAKKQKLKIWENFVEG--EEVSNGANVESKQQEVLKVIV 739
+ F ++R + L AE++AK++K K+W ++ E EEV + + V V
Sbjct: 629 VH--FTAERSAYYKPLLSAEEAAKQRKEKVWAHYEEQPVEEVMPVLEEKERSASYKPVFV 686
Query: 740 TEVLGGGKFYVQTV-GDQKIASIQQQLAALNLKEAPVLGAFSPKKGDIVLCYFHADKSWY 798
TE+ FYVQ V ++ + + + + PV GA++P++G+ + F D WY
Sbjct: 687 TEITDDLHFYVQDVETGTQLEKLMENMRSDISSHPPVEGAYAPRRGEFCIAKF-VDGEWY 745
Query: 799 RAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLRPLDQSVS--AAPGLAQLCSLAY 856
RA V VESP + VFYIDYGN+E + ++L L + S P A + A+
Sbjct: 746 RARVEK-----VESPAKV-HVFYIDYGNREILPSTRLGALPPAFSTRVLPAQATEYAFAF 799
Query: 857 IKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTSGGKVKGQGTGTILAVTLVAVD 916
I+ P +ED +A + S + + VE S VTL D
Sbjct: 800 IQVPQ-DEDARTDAVD--SVVRDIQNTQCLLNVEHLSAS-----------CPHVTLQFAD 845
Query: 917 AEISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQDDARKERRGMWQYGDVESDDE 976
++ V +++EGL +E R +++ +KV + L Q+ A+ R +W+YGD +DD
Sbjct: 846 SKGDVGLGLVKEGLVMVEVRK--EKQFQKVITEYLNA-QESAKSARLNLWRYGDFRADDA 902
Query: 977 D 977
D
Sbjct: 903 D 903
>gb|AAP23062.1| p100 co-activator variant 1 [Danio rerio]
gi|33504513|ref|NP_878285.1| staphylococcal nuclease
domain containing 1 [Danio rerio]
Length = 888
Score = 438 bits (1126), Expect = e-121
Identities = 323/995 (32%), Positives = 518/995 (51%), Gaps = 159/995 (15%)
Query: 24 CLVIVAVASSKPGPLPEKSITLASLITPRLARRG---------GVDEPFAWESREYLRKL 74
C +IV + GP PE+ I L+++ LARR DEP+A+++RE++RK
Sbjct: 6 CAIIVR-GQPRGGPPPERQINLSNIRAGALARRAIQGQPDTKDTPDEPWAFQAREFMRKK 64
Query: 75 CIGKEVTFRVDYNVASINRDFGTVFLGE----KNVGVLVVSQGWAKVREQGQQKGEVSPY 130
IGKEV F V+ N R++G V+LG+ +N+ +V++G A VR +G + +P
Sbjct: 65 VIGKEVCFTVE-NKTPQGREYGMVYLGKDTSGENIAESLVAEGLAMVRREGIRGN--NPE 121
Query: 131 LAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGLLAANKGSPMEA 190
L LE+QAK G WS+ G +IR+L + + D++ P+ A
Sbjct: 122 QVRLCDLEDQAKSSKKGLWSE--GGGSHTIRDLKYTIENPRNFVDSL------HQKPVNA 173
Query: 191 IVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELTADENDGDVPGE 250
I+E VRDG +R LLP++ V V ++GI+SP R A DG
Sbjct: 174 IIEHVRDGCMVRALLLPDYYLVTVMLSGIKSPTFKREA---------------DG----- 213
Query: 251 PRPPLTSAQRLAVSSSTETAADPFGPDAKFFTEMRVLNRDVRIVLEGVDKFSNLIGSVYY 310
+ET +PF +AKFFTE R+L RDV+I+LE ++G++ +
Sbjct: 214 ----------------SETP-EPFAAEAKFFTESRLLQRDVQIILESCPN-QVILGTILH 255
Query: 311 PDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRLRMWTNYVPPAS 370
P+G ++ L++ G+A+ V+WS + + A+ +L+ AE AK+ ++R+W +YV P +
Sbjct: 256 PNG----NITELLLKEGFARCVDWSMAVYTQGAE-KLRAAERSAKERKVRIWKDYVAPTA 310
Query: 371 NSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCPKV-GNPRRDEK 429
N ++ F KV++VV+ D I+V +S Y + ++LSSIR P++ G + +K
Sbjct: 311 NLDQ-KDRQFVAKVMQVVNADAIVVKLNSGEY------KTIHLSSIRPPRLEGEEKNKDK 363
Query: 430 --------PAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPS-PAADSRVMDF 480
PY EA+EFLR +L+G++V+V ++Y R VP+ P +
Sbjct: 364 DKRFRPLYDIPYMFEAREFLRKKLIGKKVNVTVDYIRAATNAMEMGVPAFPERTCATVTI 423
Query: 481 GSVFLLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVIRHR-DFEERSNYYDA 539
G G+N+ E +V +G TVIR+R D ++RS++YD
Sbjct: 424 G---------------------------GINIAEALVSKGLATVIRYRQDDDQRSSHYDE 456
Query: 540 LLTAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEYVLSG 599
LL AE+RA+ KG+HS K+ P+ + D+ + +KAK F PFL R+ R AVVEYV SG
Sbjct: 457 LLAAEARAIKNGKGLHSKKEVPIHRVADI-SGETQKAKQFFPFLQRAGRSEAVVEYVFSG 515
Query: 600 HRFKLLIPKETCSIAFALSGVRCPGRG-----------EPYSEEAIALMRRKIMQRDVEI 648
R KL +PKETC I F L+G+ CP RG EPYSEEA+ + ++QR+VE+
Sbjct: 516 SRLKLYMPKETCLITFLLAGIECP-RGSRNMPGGMQVAEPYSEEAMLFTKELVLQREVEV 574
Query: 649 EVETVDRNGTFLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQK 708
EVE++D+ G F+G L N+++ L+E L+K+ F ++R + L AE+SA+++K
Sbjct: 575 EVESMDKAGNFIGWLHIEGVNLSVALVENALSKVH--FTAERSSYYKTLVSAEESARQRK 632
Query: 709 LKIWENFVE--GEEVSNGANVESKQQEVLKVIVTEVLGGGKFYVQTV-GDQKIASIQQQL 765
K+W N+ E EEV+ + + + V VTE+ G FY Q V K+ ++ + +
Sbjct: 633 EKLWANYEEKPKEEVAQVTEAKERVAKYRSVYVTEITDGLHFYAQDVETGTKLENLMESM 692
Query: 766 AALNLKEAPVLGAFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYG 825
+ PV G+F+P++G+ + F AD WYRA VESP + VFYIDYG
Sbjct: 693 RGEIAAQPPVEGSFAPRRGEFCIAKF-ADGEWYRARFEK-----VESPAKV-HVFYIDYG 745
Query: 826 NQEQVAYSQLRPLDQSVS--AAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGK 883
N+E ++ ++L L + S P A + AYI+ P +ED +A +
Sbjct: 746 NREVLSSTRLAALPPAFSTRTLPPQATEYAFAYIQVPQ-DEDARADAVD----------- 793
Query: 884 EFRAQVEERDTSGGKVKGQGTGTIL-AVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRK 942
+ V + + + + +G++ VTL D + V +++EG+ ++ R ++
Sbjct: 794 ---SVVRDIHNTQCLLNVEYSGSVCPQVTLQFADTKEDVGLGLVKEGMVMVDIRK--EKY 848
Query: 943 ERKVGLDSLQKFQDDARKERRGMWQYGDVESDDED 977
+K+ + L Q+ A+ R +W+YGD DD D
Sbjct: 849 LQKMVTEYLNA-QESAKSARLNIWRYGDFRDDDAD 882
Score = 94.4 bits (233), Expect = 2e-17
Identities = 93/372 (25%), Positives = 169/372 (45%), Gaps = 62/372 (16%)
Query: 387 VVSGDCIIVADDSIPYGSPLAERRVNLSSIRCPKV------GNPRRDEKP-APYAREAKE 439
V+SG IIV P G P ER++NLS+IR + G P + P P+A +A+E
Sbjct: 2 VLSGCAIIVRGQ--PRGGPPPERQINLSNIRAGALARRAIQGQPDTKDTPDEPWAFQARE 59
Query: 440 FLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADSRVMDFGSVFLLSATKADSDDTPSS 499
F+R +++G++V +E + ++G V+L
Sbjct: 60 FMRKKVIGKEVCFTVENK----------------TPQGREYGMVYL-------------- 89
Query: 500 IPSAGSQPTGVNVGELVVGRGFGTVIRHRDFEERSNYYDALLTAESRALSGRKGIHSAKD 559
G +G N+ E +V G ++R + L E +A S +KG+ S +
Sbjct: 90 ----GKDTSGENIAESLVAEGL-AMVRREGIRGNNPEQVRLCDLEDQAKSSKKGLWS-EG 143
Query: 560 PPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEYVLSGHRFKLLIPKETCSIAFALSG 619
I DL T + ++F+ LH+ + + A++E+V G + L+ + + LSG
Sbjct: 144 GGSHTIRDLKYT-IENPRNFVDSLHQ-KPVNAIIEHVRDGCMVRALLLPDYYLVTVMLSG 201
Query: 620 VRCP---------GRGEPYSEEAIALMRRKIMQRDVEIEVETVDRNGTFLGSLWESRTNV 670
++ P EP++ EA +++QRDV+I +E+ N LG++ N+
Sbjct: 202 IKSPTFKREADGSETPEPFAAEAKFFTESRLLQRDVQIILESCP-NQVILGTILHPNGNI 260
Query: 671 ALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKLKIWENFVEGEEVSNGANVESK 730
LL+ G A+ + L AE+SAK++K++IW+++ V+ AN++ K
Sbjct: 261 TELLLKEGFARCVDWSMAVYTQGAEKLRAAERSAKERKVRIWKDY-----VAPTANLDQK 315
Query: 731 QQEVLKVIVTEV 742
++ + ++ V
Sbjct: 316 DRQFVAKVMQVV 327
>gb|AAH77133.1| Staphylococcal nuclease domain containing 1 [Danio rerio]
Length = 888
Score = 438 bits (1126), Expect = e-121
Identities = 324/995 (32%), Positives = 518/995 (51%), Gaps = 159/995 (15%)
Query: 24 CLVIVAVASSKPGPLPEKSITLASLITPRLARRG---------GVDEPFAWESREYLRKL 74
C +IV + GP PE+ I L+++ LARR DEP+A+++RE++RK
Sbjct: 6 CAIIVR-GQPRGGPPPERQINLSNIRAGALARRAIQGQPDTKDTPDEPWAFQAREFMRKK 64
Query: 75 CIGKEVTFRVDYNVASINRDFGTVFLGE----KNVGVLVVSQGWAKVREQGQQKGEVSPY 130
IGKEV F V+ N R++G V+LG+ +N+ +V++G A VR +G + +P
Sbjct: 65 VIGKEVCFTVE-NKTPQGREYGMVYLGKDTSGENIAESLVAEGLAMVRREGIRGN--NPE 121
Query: 131 LAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGLLAANKGSPMEA 190
L LE+QAK G WS+ G +IR+L + + D++ P+ A
Sbjct: 122 QVRLCDLEDQAKSSKKGLWSE--GGGSHTIRDLKYTIENPRNFVDSL------HQKPVNA 173
Query: 191 IVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELTADENDGDVPGE 250
I+E VRDG +R LLP++ V V ++GI+SP R A DG
Sbjct: 174 IIEHVRDGCMVRALLLPDYYLVTVMLSGIKSPTFKREA---------------DG----- 213
Query: 251 PRPPLTSAQRLAVSSSTETAADPFGPDAKFFTEMRVLNRDVRIVLEGVDKFSNLIGSVYY 310
+ET +PF +AKFFTE R+L RDV+I+LE ++G++ +
Sbjct: 214 ----------------SETP-EPFAAEAKFFTESRLLQRDVQIILESCPN-QVILGTILH 255
Query: 311 PDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRLRMWTNYVPPAS 370
P+G ++ L++ G+A+ V+WS + + A+ +L+ AE AK+ ++R+W +YV P +
Sbjct: 256 PNG----NITELLLKEGFARCVDWSMAVYTQGAE-KLRAAERSAKERKVRIWKDYVAPTA 310
Query: 371 NSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCPKV-GNPRRDEK 429
N ++ F KV++VV+ D I+V +S Y + ++LSSIR P++ G + +K
Sbjct: 311 NLDQ-KDRQFVAKVMQVVNADAIVVKLNSGEY------KTIHLSSIRPPRLEGEEKNKDK 363
Query: 430 --------PAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPS-PAADSRVMDF 480
PY EA+EFLR +L+G++V+V ++Y R VP+ P +
Sbjct: 364 DKRFRPLYDIPYMFEAREFLRKKLIGKKVNVTVDYIRAATNAMEMGVPAFPERTCATVTI 423
Query: 481 GSVFLLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVIRHR-DFEERSNYYDA 539
G G+N+ E +V +G TVIR+R D ++RS++YD
Sbjct: 424 G---------------------------GINIAEALVSKGLATVIRYRQDDDQRSSHYDE 456
Query: 540 LLTAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEYVLSG 599
LL AE+RA+ KG+HS K+ P+ + D+ + +KAK F PFL R+ R AVVEYV SG
Sbjct: 457 LLAAEARAIKNGKGLHSKKEVPIHRVADI-SGETQKAKQFFPFLQRAGRSEAVVEYVFSG 515
Query: 600 HRFKLLIPKETCSIAFALSGVRCPGRG-----------EPYSEEAIALMRRKIMQRDVEI 648
R KL +PKETC I F L+G+ CP RG EPYSEEA+ + ++QR+VE+
Sbjct: 516 SRLKLYMPKETCLITFLLAGIECP-RGSRNMPGGMQVAEPYSEEAMLFTKELVLQREVEV 574
Query: 649 EVETVDRNGTFLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQK 708
EVE++D+ G F+G L N+++ L+E L+K+ F ++R L AE+SA+++K
Sbjct: 575 EVESMDKAGNFIGWLHIEGVNLSVALVENALSKVH--FTAERSSYCKTLVSAEESARQRK 632
Query: 709 LKIWENFVE--GEEVSNGANVESKQQEVLKVIVTEVLGGGKFYVQTV-GDQKIASIQQQL 765
K+W N+ E EEV+ + + + V VTE+ G FY Q V K+ ++ + +
Sbjct: 633 EKLWANYEEKPKEEVAQVTEAKERVAKYRSVYVTEITDGLHFYAQDVETGTKLENLMESM 692
Query: 766 AALNLKEAPVLGAFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYG 825
+ PV G+F+P++G+ + F AD WYRA V VESP + VFYIDYG
Sbjct: 693 RGEIAAQPPVEGSFAPRRGEFCIAKF-ADGEWYRARVEK-----VESPAKV-HVFYIDYG 745
Query: 826 NQEQVAYSQLRPLDQSVS--AAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGK 883
N+E ++ ++L L + S P A + AYI+ P +ED +A +
Sbjct: 746 NREVLSSTRLAALPPAFSTRTLPPQATEYAFAYIQVPQ-DEDARADAVD----------- 793
Query: 884 EFRAQVEERDTSGGKVKGQGTGTIL-AVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRK 942
+ V + + + + +G++ VTL D + V +++EG+ ++ R ++
Sbjct: 794 ---SVVRDIHNTQCLLNVEYSGSVCPQVTLQFADTKEDVGLGLVKEGMVMVDIRK--EKY 848
Query: 943 ERKVGLDSLQKFQDDARKERRGMWQYGDVESDDED 977
+K+ + L Q+ A+ R +W+YGD DD D
Sbjct: 849 LQKMVTEYLNA-QESAKSARLNIWRYGDFRDDDAD 882
Score = 94.4 bits (233), Expect = 2e-17
Identities = 93/372 (25%), Positives = 169/372 (45%), Gaps = 62/372 (16%)
Query: 387 VVSGDCIIVADDSIPYGSPLAERRVNLSSIRCPKV------GNPRRDEKP-APYAREAKE 439
V+SG IIV P G P ER++NLS+IR + G P + P P+A +A+E
Sbjct: 2 VLSGCAIIVRGQ--PRGGPPPERQINLSNIRAGALARRAIQGQPDTKDTPDEPWAFQARE 59
Query: 440 FLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADSRVMDFGSVFLLSATKADSDDTPSS 499
F+R +++G++V +E + ++G V+L
Sbjct: 60 FMRKKVIGKEVCFTVENK----------------TPQGREYGMVYL-------------- 89
Query: 500 IPSAGSQPTGVNVGELVVGRGFGTVIRHRDFEERSNYYDALLTAESRALSGRKGIHSAKD 559
G +G N+ E +V G ++R + L E +A S +KG+ S +
Sbjct: 90 ----GKDTSGENIAESLVAEGL-AMVRREGIRGNNPEQVRLCDLEDQAKSSKKGLWS-EG 143
Query: 560 PPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEYVLSGHRFKLLIPKETCSIAFALSG 619
I DL T + ++F+ LH+ + + A++E+V G + L+ + + LSG
Sbjct: 144 GGSHTIRDLKYT-IENPRNFVDSLHQ-KPVNAIIEHVRDGCMVRALLLPDYYLVTVMLSG 201
Query: 620 VRCP---------GRGEPYSEEAIALMRRKIMQRDVEIEVETVDRNGTFLGSLWESRTNV 670
++ P EP++ EA +++QRDV+I +E+ N LG++ N+
Sbjct: 202 IKSPTFKREADGSETPEPFAAEAKFFTESRLLQRDVQIILESCP-NQVILGTILHPNGNI 260
Query: 671 ALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKLKIWENFVEGEEVSNGANVESK 730
LL+ G A+ + L AE+SAK++K++IW+++ V+ AN++ K
Sbjct: 261 TELLLKEGFARCVDWSMAVYTQGAEKLRAAERSAKERKVRIWKDY-----VAPTANLDQK 315
Query: 731 QQEVLKVIVTEV 742
++ + ++ V
Sbjct: 316 DRQFVAKVMQVV 327
>gb|AAP31682.1| 100 kDa coactivator [Bos taurus] gi|60415927|sp|Q863B3|SND1_BOVIN
Staphylococcal nuclease domain containing protein 1
(p100 co-activator) (100 kDa coactivator)
gi|45429977|ref|NP_991353.1| 100 kDa coactivator [Bos
taurus]
Length = 910
Score = 437 bits (1125), Expect = e-121
Identities = 333/1002 (33%), Positives = 523/1002 (51%), Gaps = 155/1002 (15%)
Query: 13 RGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLARRGGV---------DEPF 63
RG VK V SG C +IV + GP PE+ I L+++ LARR V DEP+
Sbjct: 21 RGIVKMVLSG-CAIIVR-GQPRGGPPPERQINLSNIRAGNLARRAAVAQPDAKDTPDEPW 78
Query: 64 AWESREYLRKLCIGKEVTFRVDYNVASINRDFGTVFLGE----KNVGVLVVSQGWAKVRE 119
A+ +RE+LRK IGKEV F ++ N R++G ++LG+ +N+ +V++G A RE
Sbjct: 79 AFPAREFLRKKLIGKEVCFTIE-NKTPQGREYGMIYLGKDTNGENIAESLVAEGLATRRE 137
Query: 120 QGQQKGEVSPYLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGL 179
+ LAE EEQAK G WS+ G +IR+L + I + +F
Sbjct: 138 GMRANNPEQNRLAEC---EEQAKASKKGMWSE--GNGSHTIRDLKYT-IENPRHF----- 186
Query: 180 LAANKGSPMEAIVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELT 239
+ ++ P+ AI+E VRDGS +R LLP++ V V ++GI+ P R A
Sbjct: 187 VDSHHQKPVNAIIEHVRDGSVVRALLLPDYYLVTVMLSGIKCPTFRREA----------- 235
Query: 240 ADENDGDVPGEPRPPLTSAQRLAVSSSTETAADPFGPDAKFFTEMRVLNRDVRIVLEGVD 299
DG +ET +PF +AKFFTE R+L RDV+I+LE
Sbjct: 236 ----DG---------------------SETP-EPFAAEAKFFTESRLLQRDVQIILESCH 269
Query: 300 KFSNLIGSVYYPDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRL 359
N++G++ +P+G ++ L++ G+A+ V+WS + A++ L+ AE AK+ RL
Sbjct: 270 N-QNILGTILHPNG----NITELLLKEGFARCVDWSIAVYTRGAEK-LRAAERFAKERRL 323
Query: 360 RMWTNYVPPASNSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCP 419
R+W +YV P +N Q F KV++V++ D I+V +S Y + ++LSSIR P
Sbjct: 324 RIWRDYVAPTANLDQKDKQ-FVAKVMQVLNADAIVVKLNSGDY------KTIHLSSIRPP 376
Query: 420 KVGNPRRDEK--------PAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSP 471
++ +K PY EA+EFLR +L+G++V+V ++Y R SP
Sbjct: 377 RLEGENTQDKNKKLRPLYDIPYMFEAREFLRKKLIGKKVNVTVDYIRPA---------SP 427
Query: 472 AADSRVMDFGSVFLLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVIRHR-DF 530
A D+ A S+ T +++ G +N+ E +V +G TVIR+R D
Sbjct: 428 ATDT-------------VPAFSERTCATVTIGG-----INIAEALVSKGLATVIRYRQDD 469
Query: 531 EERSNYYDALLTAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIP 590
++RS++YD LL AE+RA+ KG+HS K+ P+ + D++ + +KAK FLPFL R+ R
Sbjct: 470 DQRSSHYDELLAAEARAIKNGKGLHSKKEVPIHRVADISGDT-QKAKQFLPFLQRAGRSE 528
Query: 591 AVVEYVLSGHRFKLLIPKETCSIAFALSGVRCP----------GRGEPYSEEAIALMRRK 640
AVVEYV SG R KL +PKETC I F L+G+ CP GEP+SEEA +
Sbjct: 529 AVVEYVFSGSRLKLYLPKETCLITFLLAGIECPRGARNLPGLVQEGEPFSEEATLFTKEL 588
Query: 641 IMQRDVEIEVETVDRNGTFLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRA 700
++QR+VE+EVE++D+ G F+G L N+++ L+E L+K+ F ++R + L A
Sbjct: 589 VLQREVEVEVESMDKAGNFIGWLHIDGANLSVLLVEHALSKVH--FTAERSAYYKSLLSA 646
Query: 701 EQSAKKQKLKIWENFVEG--EEVSNGANVESKQQEVLKVIVTEVLGGGKFYVQTV-GDQK 757
E++AK++K K+W ++ E EE+ + + V VTE+ FYVQ V +
Sbjct: 647 EEAAKQKKEKVWAHYEEQPVEELMPVLEEKERSASYKPVFVTEITDDLHFYVQDVETGTQ 706
Query: 758 IASIQQQLAALNLKEAPVLGAFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIF 817
+ + + + PV G+++P++G+ + F D WYRA V VESP +
Sbjct: 707 LEKLMENMRNDIASHPPVEGSYAPRRGEFCIAKF-VDGEWYRARVEK-----VESPAKV- 759
Query: 818 EVFYIDYGNQEQVAYSQLRPLDQSVS--AAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLS 875
VFYIDYGN+E + ++L L + S P A + A+I+ P +ED +A + +
Sbjct: 760 HVFYIDYGNREILPSTRLGTLPPAFSTRVLPAQATEYAFAFIQVPQ-DEDARTDAVDSV- 817
Query: 876 ELTLSSGKEFRAQVEERDTSGGKVKGQGTGTILAVTLVAVDAEISVNAAMLQEGLARMEK 935
+ + + + S G VTL D++ V +++EGL +E
Sbjct: 818 ---VRDIQNTQCLLNVEHLSAG---------CPHVTLQFADSKGDVGLGLVKEGLVMVEV 865
Query: 936 RNRWDRKERKVGLDSLQKFQDDARKERRGMWQYGDVESDDED 977
R +++ +KV + L Q+ A+ R +W+YGD +DD D
Sbjct: 866 RK--EKQFQKVITEYLNA-QESAKSARLNLWRYGDFRADDAD 904
>sp|Q78PY7|SND1_MOUSE Staphylococcal nuclease domain containing protein 1 (p100
co-activator) (100 kDa coactivator)
gi|66840139|gb|AAH07126.3| AL033314 protein [Mus
musculus] gi|26352950|dbj|BAC40105.1| unnamed protein
product [Mus musculus]
Length = 910
Score = 437 bits (1124), Expect = e-121
Identities = 334/1002 (33%), Positives = 521/1002 (51%), Gaps = 155/1002 (15%)
Query: 13 RGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLARRGGV---------DEPF 63
RG VK V SG C +IV + GP PE+ I L+++ LARR DEP+
Sbjct: 21 RGIVKMVLSG-CAIIVR-GQPRGGPPPERQINLSNIRAGNLARRAAATQPDGKDTPDEPW 78
Query: 64 AWESREYLRKLCIGKEVTFRVDYNVASINRDFGTVFLGE----KNVGVLVVSQGWAKVRE 119
A+ +RE+LRK IGKEV F ++ N R++G ++LG+ +N+ +V++G A R
Sbjct: 79 AFPAREFLRKKLIGKEVCFTIE-NKTPQGREYGMIYLGKDTNGENIAESLVAEGLA-TRR 136
Query: 120 QGQQKGEVSPYLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGL 179
+G + +P L EEQAK G WS+ G +IR+L + I + +F
Sbjct: 137 EGMRAN--NPEQNRLSECEEQAKASKKGMWSE--GNGSHTIRDLKYT-IENPRHF----- 186
Query: 180 LAANKGSPMEAIVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELT 239
+ ++ P+ AI+E VRDGS +R LLP V V ++GI+ P R
Sbjct: 187 VDSHHQKPVNAIIEHVRDGSVVRALLLPGHHLVTVMLSGIKCPTFRR------------- 233
Query: 240 ADENDGDVPGEPRPPLTSAQRLAVSSSTETAADPFGPDAKFFTEMRVLNRDVRIVLEGVD 299
E DG +ET +PF +AKFFTE R+L RDV+I+LE
Sbjct: 234 --ETDG---------------------SET-PEPFAAEAKFFTESRLLQRDVQIILESCH 269
Query: 300 KFSNLIGSVYYPDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRL 359
NL+G++ +P+G ++ L++ G+A+ V+WS + A+ +L+ AE AK+ RL
Sbjct: 270 N-QNLLGTILHPNG----NITELLLKEGFARCVDWSIAVYTRGAE-KLRAAERFAKERRL 323
Query: 360 RMWTNYVPPASNSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCP 419
R+W +YVPP +N ++ F KV++V++ D I+V +S Y + ++LSSIR P
Sbjct: 324 RIWRDYVPPTANLDQ-KDKQFVAKVMQVLNADAIVVKLNSGDY------KTIHLSSIRPP 376
Query: 420 KVGNPRRDEK--------PAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSP 471
++ +K PY EA+EFLR +L+G++V+V ++Y R SP
Sbjct: 377 RLEGDNIQDKNKKLRPLYDIPYMFEAREFLRKKLIGKKVNVTVDYIRP---------ASP 427
Query: 472 AADSRVMDFGSVFLLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVIRHR-DF 530
A ++ A S+ T +++ G+N+ E +V +G TVIR+R D
Sbjct: 428 ATET-------------VPAFSERTCATVTIG-----GINIAEALVSKGLATVIRYRQDD 469
Query: 531 EERSNYYDALLTAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIP 590
++RS++YD LL AE+RA+ KG+HS K+ P+ + D+ + +KAK FLPFL R+ R
Sbjct: 470 DQRSSHYDELLAAEARAIKNGKGLHSKKEVPIHRVADI-SGDTQKAKQFLPFLQRAGRSE 528
Query: 591 AVVEYVLSGHRFKLLIPKETCSIAFALSGVRCP----------GRGEPYSEEAIALMRRK 640
AVVEYV SG R KL +PKETC I F L+G+ CP GEP+SEEA +
Sbjct: 529 AVVEYVFSGSRLKLYLPKETCLITFLLAGIECPRGARNLPGLVQEGEPFSEEATLFTKEL 588
Query: 641 IMQRDVEIEVETVDRNGTFLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRA 700
++QR+VE+EVE++D+ G F+G L N+++ L+E L+K+ F ++R + L A
Sbjct: 589 VLQREVEVEVESMDKAGNFIGWLHMDGANLSVLLVEQALSKVH--FTAERSAYYKPLLSA 646
Query: 701 EQSAKKQKLKIWENFVEG--EEVSNGANVESKQQEVLKVIVTEVLGGGKFYVQTV-GDQK 757
E++AK++K K+W ++ E EEV + + V VTE+ FYVQ V +
Sbjct: 647 EEAAKQRKEKVWAHYEERPVEEVMPVLEEKERSASYKPVFVTEITDDLHFYVQDVETGTQ 706
Query: 758 IASIQQQLAALNLKEAPVLGAFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIF 817
+ + + + PV G+++P++G+ + F D WYRA V VESP +
Sbjct: 707 LEKLMENMRNDISSHPPVEGSYAPRRGEFCIAKF-VDGEWYRARVEK-----VESPAKV- 759
Query: 818 EVFYIDYGNQEQVAYSQLRPLDQSVS--AAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLS 875
VFYIDYGN+E + ++L L + S P A + A+I+ P +ED +A + S
Sbjct: 760 HVFYIDYGNREILPSTRLGTLPPAFSTRVLPAQATEYAFAFIQVPQ-DEDARTDAVD--S 816
Query: 876 ELTLSSGKEFRAQVEERDTSGGKVKGQGTGTILAVTLVAVDAEISVNAAMLQEGLARMEK 935
+ + VE S VTL D++ V +++EGL +E
Sbjct: 817 VVRDIQNTQCLLNVEHLSAS-----------CPHVTLQFADSKGDVGLGLVKEGLVMVEV 865
Query: 936 RNRWDRKERKVGLDSLQKFQDDARKERRGMWQYGDVESDDED 977
R +++ +KV + L Q+ A+ R +W+YGD +DD D
Sbjct: 866 RK--EKQFQKVITEYLNA-QESAKSARLNLWRYGDFRADDAD 904
>dbj|BAD92747.1| EBNA-2 co-activator variant [Homo sapiens]
Length = 964
Score = 435 bits (1118), Expect = e-120
Identities = 330/1002 (32%), Positives = 523/1002 (51%), Gaps = 155/1002 (15%)
Query: 13 RGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLARRGGV---------DEPF 63
RG +K V SG C +IV + GP PE+ I L+++ LARR DEP+
Sbjct: 75 RGIIKMVLSG-CAIIVR-GQPRGGPPPERQINLSNIRAGNLARRAAATQPDAKDTPDEPW 132
Query: 64 AWESREYLRKLCIGKEVTFRVDYNVASINRDFGTVFLGE----KNVGVLVVSQGWAKVRE 119
A+ +RE+LRK IGKEV F ++ N R++G ++LG+ +N+ +V++G A R
Sbjct: 133 AFPAREFLRKKLIGKEVCFTIE-NKTPQGREYGMIYLGKDTNGENIAESLVAEGLA-TRR 190
Query: 120 QGQQKGEVSPYLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGL 179
+G + +P L EEQAK G WS+ G +IR+L + I + +F
Sbjct: 191 EGMRAN--NPEQNRLSECEEQAKAAKKGMWSE--GNGSHTIRDLKYT-IENPRHF----- 240
Query: 180 LAANKGSPMEAIVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELT 239
+ ++ P+ AI+E VRDGS +R LLP++ V V ++GI+ P R A
Sbjct: 241 VDSHHQKPVNAIIEHVRDGSVVRALLLPDYYLVTVMLSGIKCPTFRREA----------- 289
Query: 240 ADENDGDVPGEPRPPLTSAQRLAVSSSTETAADPFGPDAKFFTEMRVLNRDVRIVLEGVD 299
DG +ET +PF +AKFFTE R+L RDV+I+LE
Sbjct: 290 ----DG---------------------SETP-EPFAAEAKFFTESRLLQRDVQIILESCH 323
Query: 300 KFSNLIGSVYYPDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRL 359
N++G++ +P+G ++ L++ G+A+ V+WS + A++ L+ AE AK+ RL
Sbjct: 324 N-QNILGTILHPNG----NITELLLKEGFARCVDWSIAVYTRGAEK-LRAAERVAKERRL 377
Query: 360 RMWTNYVPPASNSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCP 419
R+W +YV P +N Q F KV++V++ D I+V +S Y + ++LSSIR P
Sbjct: 378 RIWRDYVAPTANLDQKDKQ-FVAKVMQVLNADAIVVKLNSGDY------KTIHLSSIRPP 430
Query: 420 KVGNPRRDEK--------PAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSP 471
++ +K PY EA+EFLR +L+G++V+V ++Y R SP
Sbjct: 431 RLEGENTQDKNKKLRPLYDIPYMFEAREFLRKKLIGKKVNVTVDYIRPA---------SP 481
Query: 472 AADSRVMDFGSVFLLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVIRHR-DF 530
A ++ A S+ T +++ G +N+ E +V +G TVIR+R D
Sbjct: 482 ATET-------------VPAFSERTCATVTIGG-----INIAEALVSKGLATVIRYRQDD 523
Query: 531 EERSNYYDALLTAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIP 590
++RS++YD LL AE+RA+ KG+HS K+ P+ + D++ + +KAK FLPFL R+ R
Sbjct: 524 DQRSSHYDELLAAEARAIKNGKGLHSKKEVPIHRVADISGDT-QKAKQFLPFLQRAGRSE 582
Query: 591 AVVEYVLSGHRFKLLIPKETCSIAFALSGVRCP----------GRGEPYSEEAIALMRRK 640
AVVEYV SG R KL +PKETC I F L+G+ CP GEP+SEEA +
Sbjct: 583 AVVEYVFSGSRLKLYLPKETCLITFLLAGIECPRGARNLPGLVQEGEPFSEEATLFTKEL 642
Query: 641 IMQRDVEIEVETVDRNGTFLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRA 700
++QR+VE+EVE++D+ G F+G L N+++ L+E L+K+ F ++R + L A
Sbjct: 643 VLQREVEVEVESMDKAGNFIGWLHIDGANLSVLLVEHALSKVH--FTAERSSYYKSLLSA 700
Query: 701 EQSAKKQKLKIWENFVEG--EEVSNGANVESKQQEVLKVIVTEVLGGGKFYVQTV-GDQK 757
E++AK++K K+W ++ E EEV + + V VTE+ FYVQ V +
Sbjct: 701 EEAAKQKKEKVWAHYEEQPVEEVMPVLEEKERSASYKPVFVTEITDDLHFYVQDVETGTQ 760
Query: 758 IASIQQQLAALNLKEAPVLGAFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIF 817
+ + + + PV G+++P++G+ + F D WYRA V VESP I
Sbjct: 761 LEKLMENMRNDIASHPPVEGSYAPRRGEFCIAKF-VDGEWYRARVEK-----VESPAKI- 813
Query: 818 EVFYIDYGNQEQVAYSQLRPLDQSVS--AAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLS 875
VFYIDYGN+E + ++L L + S P A + A+I+ P ++D +A + +
Sbjct: 814 HVFYIDYGNREVLPSTRLGTLSPAFSTRVLPAQATEYAFAFIQVPQ-DDDARTDAVDSV- 871
Query: 876 ELTLSSGKEFRAQVEERDTSGGKVKGQGTGTILAVTLVAVDAEISVNAAMLQEGLARMEK 935
+ + + + S G VTL D++ V +++EGL +E
Sbjct: 872 ---VRDIQNTQCLLNVEHLSAG---------CPHVTLQFADSKGDVGLGLVKEGLVMVEV 919
Query: 936 RNRWDRKERKVGLDSLQKFQDDARKERRGMWQYGDVESDDED 977
R +++ +KV + L Q+ A+ R +W+YGD +DD D
Sbjct: 920 RK--EKQFQKVITEYLNA-QESAKSARLNLWRYGDFRADDAD 958
>gb|AAH72471.1| Unknown (protein for MGC:91679) [Rattus norvegicus]
Length = 885
Score = 435 bits (1118), Expect = e-120
Identities = 328/991 (33%), Positives = 515/991 (51%), Gaps = 154/991 (15%)
Query: 24 CLVIVAVASSKPGPLPEKSITLASLITPRLARRGGV---------DEPFAWESREYLRKL 74
C +IV + GP PE+ I L+++ LARR DEP+A+ +RE+LRK
Sbjct: 6 CAIIVR-GQPRGGPPPERQINLSNIRAGNLARRAAATQPDGKDTPDEPWAFPAREFLRKK 64
Query: 75 CIGKEVTFRVDYNVASINRDFGTVFLGE----KNVGVLVVSQGWAKVREQGQQKGEVSPY 130
IGKEV F ++ N R++G ++LG+ +N+ +V++G A RE + +P
Sbjct: 65 LIGKEVCFTIE-NKTPQGREYGMIYLGKDTNGENIAESLVAEGLASRREGMRAN---NPE 120
Query: 131 LAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGLLAANKGSPMEA 190
L EEQAK G WS+ G +IR+L + I + +F + ++ P+ A
Sbjct: 121 QNRLSECEEQAKASKKGMWSE--GNGSHTIRDLKYT-IENPRHF-----VDSHHQKPVNA 172
Query: 191 IVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELTADENDGDVPGE 250
I+E VRDGS +R LLP+ V V ++GI+ P R E DG
Sbjct: 173 IIEHVRDGSVVRALLLPDHYLVTVMLSGIKCPTFRR---------------ETDG----- 212
Query: 251 PRPPLTSAQRLAVSSSTETAADPFGPDAKFFTEMRVLNRDVRIVLEGVDKFSNLIGSVYY 310
+ET +PF +AKFFTE R+L RDV+I+LE N++G++ +
Sbjct: 213 ----------------SET-PEPFAAEAKFFTESRLLQRDVQIILESCHN-QNILGTILH 254
Query: 311 PDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRLRMWTNYVPPAS 370
P+G ++ L++ G+A+ V+WS + A+ +L+ AE AK+ RLR+W +YVPP +
Sbjct: 255 PNG----NITELLLKEGFARCVDWSIAVYTRGAE-KLRAAERFAKERRLRIWRDYVPPTA 309
Query: 371 NSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCPKVGNPRRDEK- 429
N ++ F KV++V++ D I+V +S Y + ++LSSIR P++ +K
Sbjct: 310 NLDQ-KDKQFVAKVMQVLNADAIVVKLNSGDY------KTIHLSSIRPPRLEGDNIQDKN 362
Query: 430 -------PAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADSRVMDFGS 482
PY EA+EFLR +L+G++VSV ++Y R SPA ++
Sbjct: 363 KKLRPLYDIPYMFEAREFLRKKLIGKKVSVTVDYIRP---------ASPATET------- 406
Query: 483 VFLLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVIRHR-DFEERSNYYDALL 541
A S+ T +++ G+N+ E +V +G TVIR+R D ++RS++YD LL
Sbjct: 407 ------VPAFSERTCATVTIG-----GINIAEALVSKGLATVIRYRQDDDQRSSHYDELL 455
Query: 542 TAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEYVLSGHR 601
AE+RA+ KG+HS K+ P+ + D+ + +KAK FLPFL R+ R AVVEYV SG R
Sbjct: 456 AAEARAIKNGKGLHSKKEVPIHRVADI-SGDTQKAKQFLPFLQRAGRSEAVVEYVFSGSR 514
Query: 602 FKLLIPKETCSIAFALSGVRCP----------GRGEPYSEEAIALMRRKIMQRDVEIEVE 651
KL +PKETC I F L+G+ CP GEP+SEEA + ++QR+VE+EVE
Sbjct: 515 LKLYLPKETCLITFLLAGIECPRGARNLPGLVQEGEPFSEEATLFTKELVLQREVEVEVE 574
Query: 652 TVDRNGTFLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKLKI 711
++D+ G F+G L N+++ L+E L+K+ F ++R + L AE++AK++K K+
Sbjct: 575 SMDKAGNFIGWLHMDGANLSVLLVEHALSKVH--FTAERSAYYKPLLSAEEAAKQRKEKV 632
Query: 712 WENFVEG--EEVSNGANVESKQQEVLKVIVTEVLGGGKFYVQTV-GDQKIASIQQQLAAL 768
W ++ E EEV + + V VTE+ FYVQ V ++ + + + +
Sbjct: 633 WAHYEEQPVEEVMPVLEEKERSASYKPVFVTEITDDLHFYVQDVETGTQLEKLMENMRSD 692
Query: 769 NLKEAPVLGAFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQE 828
PV GA++P++G+ + F D WYRA V VESP + VFYIDYGN+E
Sbjct: 693 ISSHPPVEGAYAPRRGEFCIAKF-VDGEWYRARVEK-----VESPAKV-HVFYIDYGNRE 745
Query: 829 QVAYSQLRPLDQSVS--AAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFR 886
+ ++L L + S P A + A+I+ P +ED +A + S + +
Sbjct: 746 ILPSTRLGALPPAFSTRVLPAQATEYAFAFIQVPQ-DEDARTDAVD--SVVRDIQNTQCL 802
Query: 887 AQVEERDTSGGKVKGQGTGTILAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKV 946
VE S VTL D++ V +++EGL +E R +++ +KV
Sbjct: 803 LNVEHLSAS-----------CPHVTLQFADSKGDVGLGLVKEGLVMVEVRK--EKQFQKV 849
Query: 947 GLDSLQKFQDDARKERRGMWQYGDVESDDED 977
+ L Q+ A+ R +W+YGD +DD D
Sbjct: 850 ITEYLNA-QESAKSARLNLWRYGDFRADDAD 879
>gb|AAP88787.1| EBNA-2 co-activator (100kD) [Homo sapiens]
gi|61362104|gb|AAX42161.1| staphylococcal nuclease
domain containing 1 [synthetic construct]
gi|61362100|gb|AAX42160.1| staphylococcal nuclease
domain containing 1 [synthetic construct]
gi|60415926|sp|Q7KZF4|SND1_HUMAN Staphylococcal nuclease
domain containing protein 1 (p100 co-activator) (100 kDa
coactivator) (EBNA2 coactivator p100)
Length = 910
Score = 434 bits (1117), Expect = e-120
Identities = 330/1002 (32%), Positives = 523/1002 (51%), Gaps = 155/1002 (15%)
Query: 13 RGRVKAVPSGDCLVIVAVASSKPGPLPEKSITLASLITPRLARRGGV---------DEPF 63
RG +K V SG C +IV + GP PE+ I L+++ LARR DEP+
Sbjct: 21 RGIIKMVLSG-CAIIVR-GQPRGGPPPERQINLSNIRAGNLARRAAATQPDAKDTPDEPW 78
Query: 64 AWESREYLRKLCIGKEVTFRVDYNVASINRDFGTVFLGE----KNVGVLVVSQGWAKVRE 119
A+ +RE+LRK IGKEV F ++ N R++G ++LG+ +N+ +V++G A R
Sbjct: 79 AFPAREFLRKKLIGKEVCFTIE-NKTPQGREYGMIYLGKDTNGENIAESLVAEGLA-TRR 136
Query: 120 QGQQKGEVSPYLAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGL 179
+G + +P L EEQAK G WS+ G +IR+L + I + +F
Sbjct: 137 EGMRAN--NPEQNRLSECEEQAKAAKKGMWSE--GNGSHTIRDLKYT-IENPRHF----- 186
Query: 180 LAANKGSPMEAIVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELT 239
+ ++ P+ AI+E VRDGS +R LLP++ V V ++GI+ P R A
Sbjct: 187 VDSHHQKPVNAIIEHVRDGSVVRALLLPDYYLVTVMLSGIKCPTFRREA----------- 235
Query: 240 ADENDGDVPGEPRPPLTSAQRLAVSSSTETAADPFGPDAKFFTEMRVLNRDVRIVLEGVD 299
DG +ET +PF +AKFFTE R+L RDV+I+LE
Sbjct: 236 ----DG---------------------SETP-EPFAAEAKFFTESRLLQRDVQIILESCH 269
Query: 300 KFSNLIGSVYYPDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRL 359
N++G++ +P+G ++ L++ G+A+ V+WS + A++ L+ AE AK+ RL
Sbjct: 270 N-QNILGTILHPNG----NITELLLKEGFARCVDWSIAVYTRGAEK-LRAAERFAKERRL 323
Query: 360 RMWTNYVPPASNSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCP 419
R+W +YV P +N Q F KV++V++ D I+V +S Y + ++LSSIR P
Sbjct: 324 RIWRDYVAPTANLDQKDKQ-FVAKVMQVLNADAIVVKLNSGDY------KTIHLSSIRPP 376
Query: 420 KVGNPRRDEK--------PAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSP 471
++ +K PY EA+EFLR +L+G++V+V ++Y R SP
Sbjct: 377 RLEGENTQDKNKKLRPLYDIPYMFEAREFLRKKLIGKKVNVTVDYIRPA---------SP 427
Query: 472 AADSRVMDFGSVFLLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVIRHR-DF 530
A ++ A S+ T +++ G +N+ E +V +G TVIR+R D
Sbjct: 428 ATET-------------VPAFSERTCATVTIGG-----INIAEALVSKGLATVIRYRQDD 469
Query: 531 EERSNYYDALLTAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIP 590
++RS++YD LL AE+RA+ KG+HS K+ P+ + D++ + +KAK FLPFL R+ R
Sbjct: 470 DQRSSHYDELLAAEARAIKNGKGLHSKKEVPIHRVADISGDT-QKAKQFLPFLQRAGRSE 528
Query: 591 AVVEYVLSGHRFKLLIPKETCSIAFALSGVRCP----------GRGEPYSEEAIALMRRK 640
AVVEYV SG R KL +PKETC I F L+G+ CP GEP+SEEA +
Sbjct: 529 AVVEYVFSGSRLKLYLPKETCLITFLLAGIECPRGARNLPGLVQEGEPFSEEATLFTKEL 588
Query: 641 IMQRDVEIEVETVDRNGTFLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRA 700
++QR+VE+EVE++D+ G F+G L N+++ L+E L+K+ F ++R + L A
Sbjct: 589 VLQREVEVEVESMDKAGNFIGWLHIDGANLSVLLVEHALSKVH--FTAERSSYYKSLLSA 646
Query: 701 EQSAKKQKLKIWENFVEG--EEVSNGANVESKQQEVLKVIVTEVLGGGKFYVQTV-GDQK 757
E++AK++K K+W ++ E EEV + + V VTE+ FYVQ V +
Sbjct: 647 EEAAKQKKEKVWAHYEEQPVEEVMPVLEEKERSASYKPVFVTEITDDLHFYVQDVETGTQ 706
Query: 758 IASIQQQLAALNLKEAPVLGAFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIF 817
+ + + + PV G+++P++G+ + F D WYRA V VESP I
Sbjct: 707 LEKLMENMRNDIASHPPVEGSYAPRRGEFCIAKF-VDGEWYRARVEK-----VESPAKI- 759
Query: 818 EVFYIDYGNQEQVAYSQLRPLDQSVS--AAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLS 875
VFYIDYGN+E + ++L L + S P A + A+I+ P ++D +A + +
Sbjct: 760 HVFYIDYGNREVLPSTRLGTLSPAFSTRVLPAQATEYAFAFIQVPQ-DDDARTDAVDSV- 817
Query: 876 ELTLSSGKEFRAQVEERDTSGGKVKGQGTGTILAVTLVAVDAEISVNAAMLQEGLARMEK 935
+ + + + S G VTL D++ V +++EGL +E
Sbjct: 818 ---VRDIQNTQCLLNVEHLSAG---------CPHVTLQFADSKGDVGLGLVKEGLVMVEV 865
Query: 936 RNRWDRKERKVGLDSLQKFQDDARKERRGMWQYGDVESDDED 977
R +++ +KV + L Q+ A+ R +W+YGD +DD D
Sbjct: 866 RK--EKQFQKVITEYLNA-QESAKSARLNLWRYGDFRADDAD 904
>ref|NP_062750.1| staphylococcal nuclease domain containing 1 [Mus musculus]
gi|6009521|dbj|BAA84944.1| p100 co-activator [Mus
musculus]
Length = 882
Score = 431 bits (1107), Expect = e-119
Identities = 327/991 (32%), Positives = 513/991 (50%), Gaps = 154/991 (15%)
Query: 24 CLVIVAVASSKPGPLPEKSITLASLITPRLARRGGV---------DEPFAWESREYLRKL 74
C +IV + GP PE+ I L+++ LARR DEP+A+ +RE+LRK
Sbjct: 6 CAIIVR-GQPRGGPPPERQINLSNIRAGNLARRAAATQPDGKDTPDEPWAFPAREFLRKK 64
Query: 75 CIGKEVTFRVDYNVASINRDFGTVFLGE----KNVGVLVVSQGWAKVREQGQQKGEVSPY 130
IGKEV F ++ N R++G ++LG+ +N+ +V++G A R +G + +P
Sbjct: 65 LIGKEVCFTIE-NKTPQGREYGMIYLGKDTNGENIAESLVAEGLA-TRREGMRAN--NPE 120
Query: 131 LAELLRLEEQAKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGLLAANKGSPMEA 190
L EEQAK G WS+ G +IR+L + I + +F + ++ P+ A
Sbjct: 121 QNRLSECEEQAKASKKGMWSE--GNGSHTIRDLKYT-IENPRHF-----VDSHHQKPVNA 172
Query: 191 IVEQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELTADENDGDVPGE 250
I+E VRDGS R LLP V V ++GI+ P R E DG
Sbjct: 173 IIEHVRDGSVARALLLPGHHLVTVMLSGIKCPTFRR---------------ETDG----- 212
Query: 251 PRPPLTSAQRLAVSSSTETAADPFGPDAKFFTEMRVLNRDVRIVLEGVDKFSNLIGSVYY 310
+ET +PF +AKFFTE R+L RDV+I+LE NL+G++ +
Sbjct: 213 ----------------SET-PEPFAAEAKFFTESRLLQRDVQIILESCHN-QNLLGTILH 254
Query: 311 PDGESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRLRMWTNYVPPAS 370
P+G ++ L++ G+A+ V+WS + A+ +L+ AE AK+ RLR+W +YVPP +
Sbjct: 255 PNG----NITELLLKEGFARCVDWSIAVYTRGAE-KLRAAERFAKERRLRIWRDYVPPTA 309
Query: 371 NSKAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCPKVGNPRRDEK- 429
N ++ F KV++V++ D I+V +S Y + ++LSSIR P++ +K
Sbjct: 310 NLDQ-KDKQFVAKVMQVLNADAIVVKLNSGDY------KTIHLSSIRPPRLEGDNIQDKN 362
Query: 430 -------PAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADSRVMDFGS 482
PY EA+EFLR +L+G++V+V ++Y R SPA ++
Sbjct: 363 KKLRPLYDIPYMFEAREFLRKKLIGKKVNVTVDYIRP---------ASPATET------- 406
Query: 483 VFLLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVIRHR-DFEERSNYYDALL 541
A S+ T +++ G+N+ E +V +G TVIR+R D ++RS++YD LL
Sbjct: 407 ------VPAFSERTCATVTIG-----GINIAEALVSKGLATVIRYRQDDDQRSSHYDELL 455
Query: 542 TAESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEYVLSGHR 601
AE+RA+ KG+HS K+ P+ + D+ + +KAK FLPFL R+ R AVVEYV SG R
Sbjct: 456 AAEARAIKNGKGLHSKKEVPIHRVADI-SGDTQKAKQFLPFLQRAGRSEAVVEYVFSGSR 514
Query: 602 FKLLIPKETCSIAFALSGVRCP----------GRGEPYSEEAIALMRRKIMQRDVEIEVE 651
KL +PKETC I F L+G+ CP GEP+SEEA + ++QR+VE+EVE
Sbjct: 515 LKLYLPKETCLITFLLAGIECPRGARNLPGLVQEGEPFSEEATLFTKELVLQREVEVEVE 574
Query: 652 TVDRNGTFLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKLKI 711
++D+ G F+G L N+++ L+E L+K+ F ++R + L AE++AK++K K+
Sbjct: 575 SMDKAGNFIGWLHMDGANLSVLLVEQALSKVH--FTAERSAYYKPLLSAEEAAKQRKEKV 632
Query: 712 WENFVEG--EEVSNGANVESKQQEVLKVIVTEVLGGGKFYVQTV-GDQKIASIQQQLAAL 768
W ++ E EEV + + V VTE+ FYVQ V ++ + + +
Sbjct: 633 WAHYEERPVEEVMPVLEEKERSASYKPVFVTEITDDLHFYVQDVETGTQLEKLMENMRND 692
Query: 769 NLKEAPVLGAFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQE 828
PV G+++P++G+ + F D WYRA V VESP + VFYIDYGN+E
Sbjct: 693 ISSHPPVEGSYAPRRGEFCIAKF-VDGEWYRARVEK-----VESPAKV-HVFYIDYGNRE 745
Query: 829 QVAYSQLRPLDQSVS--AAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFR 886
+ ++L L + S P A + A+I+ P +ED +A + S + +
Sbjct: 746 ILPSTRLGTLPPAFSTRVLPAQATEYAFAFIQVPQ-DEDARTDAVD--SVVRDIQNTQCL 802
Query: 887 AQVEERDTSGGKVKGQGTGTILAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKV 946
VE S VTL D++ V +++EGL +E R +++ +KV
Sbjct: 803 LNVEHLSAS-----------CPHVTLQFADSKGDVGLGLVKEGLVMVEVRK--EKQFQKV 849
Query: 947 GLDSLQKFQDDARKERRGMWQYGDVESDDED 977
+ L Q+ A+ R +W+YGD +DD D
Sbjct: 850 ITEYLNA-QESAKSARLNLWRYGDFRADDAD 879
>ref|NP_073185.1| p105 coactivator [Rattus norvegicus] gi|1800307|gb|AAB41439.1| p105
coactivator [Rattus norvegicus]
Length = 880
Score = 411 bits (1056), Expect = e-113
Identities = 311/974 (31%), Positives = 487/974 (49%), Gaps = 147/974 (15%)
Query: 36 GPLPEKSITLASLITPRLARRGGV----------DEPFAWESREYLRKLCIGKEVTFRVD 85
G PE+ I L+++ L R DEP+A+ +RE+LRK IGKEV F ++
Sbjct: 16 GTAPERQINLSNIRAGNLDTRRRAATQPDGKDTPDEPWAFPAREFLRKKLIGKEVCFTIE 75
Query: 86 YNVASINRDFGTVFLGEKNVGVLVVSQGWAKVREQGQQKGEVSPYLAELLRLEEQAKQEG 145
N R++G ++LG+ G + A+ G+ +P L EEQAK
Sbjct: 76 -NKTPQGREYGMIYLGKDTNGENIAESLVAEGLAPGESMRANNPEQNRLSECEEQAKASK 134
Query: 146 LGRWSKVPGAAEASIRNLPPSAIGDASNFDAMGLLAANKGSPMEAIVEQVRDGSTLRVYL 205
G WS+ G ++ I + +F + ++ P+ AI+E VRDGS +R L
Sbjct: 135 KGMWSEGTGHTHPDLKY----TIENPRHF-----VDSHHQKPVNAIIEHVRDGSVVRALL 185
Query: 206 LPEFQFVQVFVAGIQSPQMGRRAAPETVVETELTADENDGDVPGEPRPPLTSAQRLAVSS 265
LP+ V V ++GI+ P R E DG
Sbjct: 186 LPDHYLVTVMLSGIKCPTFRR---------------ETDG-------------------- 210
Query: 266 STETAADPFGPDAKFFTEMRVLNRDVRIVLEGVDKFSNLIGSVYYPDGESAKDLALELVE 325
+ET +PF +AKFFTE R+L RDV+I+LE N++G++ +P+G ++ L++
Sbjct: 211 -SET-PEPFAAEAKFFTESRLLQRDVQIILESCHN-QNILGTILHPNG----NITELLLK 263
Query: 326 NGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRLRMWTNYVPPASNSKAIHNQNFTGKVV 385
G+A+ V+WS + A+ +L+ AE AK+ RLR+W +YVPP +N ++ F KV+
Sbjct: 264 EGFARCVDWSIAVYTRGAE-KLRAAERFAKERRLRIWRDYVPPTANLDQ-KDKQFVAKVM 321
Query: 386 EVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCPKVGNPRRDEK--------PAPYAREA 437
+V++ D I+V S Y + ++LSSIR P++ +K PY EA
Sbjct: 322 QVLNADAIVVKLSSGDY------KTIHLSSIRPPRLEGDNIQDKNKKLRPLYDIPYMFEA 375
Query: 438 KEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADSRVMDFGSVFLLSATKADSDDTP 497
+EFLR +L+G++VSV ++Y R SPA ++ A S+ T
Sbjct: 376 REFLRKKLIGKKVSVTVDYIRP---------ASPATET-------------VPAFSERTC 413
Query: 498 SSIPSAGSQPTGVNVGELVVGRGFGTVIRHR-DFEERSNYYDALLTAESRALSGRKGIHS 556
+++ G+N+ E +V +G TVIR+R D ++RS++YD LL AE+RA+ KG+HS
Sbjct: 414 ATVTIG-----GINITEALVSKGLATVIRYRQDDDQRSSHYDELLAAEARAIKNGKGLHS 468
Query: 557 AKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEYVLSGHRFKLLIPKETCSIAFA 616
K+ P+ + D+ + +KAK FLPFL R+ R AVVEYV SG R KL +PKETC I F
Sbjct: 469 KKEVPIHRVADI-SGDTQKAKQFLPFLQRAGRSEAVVEYVFSGSRLKLYLPKETCLITFL 527
Query: 617 LSGVRCP----------GRGEPYSEEAIALMRRKIMQRDVEIEVETVDRNGTFLGSLWES 666
L+G+ CP GEP+SEEA + ++QR+VE+EVE++D+ G F+G L
Sbjct: 528 LAGIECPRGARNLPGLVQEGEPFSEEATLFTKELVLQREVEVEVESMDKAGNFIGWLHMD 587
Query: 667 RTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKLKIWENFVEG--EEVSNG 724
N+++ L+E L+K+ F ++R + L AE++AK++K K+W ++ E EEV
Sbjct: 588 GANLSVLLVEHALSKVH--FTAERSGYYKPLLSAEEAAKQRKEKVWAHYEEQPVEEVMPV 645
Query: 725 ANVESKQQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALNLKEAPVLGAFSPKKG 784
+ + V VTE+ FYVQ V + ++ P + P++G
Sbjct: 646 LEEKERSASYKPVFVTEITDDLHFYVQDVETGTQLEKLMENMRSDISSHPPVEGLRPRRG 705
Query: 785 DIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQEQVAYSQLR-PLDQSVS 843
+ + F D WYRA V ESP + VFYIDYGN+E + ++L P S
Sbjct: 706 EFCIAKF-VDGEWYRARVEKE-----ESPAKV-HVFYIDYGNREILPSTRLALPPAFSTR 758
Query: 844 AAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFRAQVEERDTSGGKVKGQG 903
P A + A+I+ P +ED +A +++ + VE S
Sbjct: 759 VLPAQATEYAFAFIQWPQ-DEDARTDAVTVCADI---QNTQCLLNVEHLSAS-------- 806
Query: 904 TGTILAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKVGLDSLQKFQDDARKERR 963
VTL D++ V +++EGL +E R +++ +KV + L Q+ A+ R
Sbjct: 807 ---CPHVTLQFADSKGDVGLGLVKEGLVMVEVRK--EKQFQKVITEYLNA-QESAKSARL 860
Query: 964 GMWQYGDVESDDED 977
+W+YGD +DD D
Sbjct: 861 NLWRYGDFRADDAD 874
>gb|AAL27548.1| transcriptional coactivator p100 [Gallus gallus]
Length = 714
Score = 365 bits (937), Expect = 4e-99
Identities = 267/811 (32%), Positives = 415/811 (50%), Gaps = 132/811 (16%)
Query: 193 EQVRDGSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELTADENDGDVPGEPR 252
E VRDGS +R LLP++ V V ++GI+ P R A V E
Sbjct: 4 EHVRDGSVVRALLLPDYYLVTVMLSGIKCPTFKREADATEVPE----------------- 46
Query: 253 PPLTSAQRLAVSSSTETAADPFGPDAKFFTEMRVLNRDVRIVLEGVDKFSNLIGSVYYPD 312
PF +AKFFTE R+L RDV+IVLE N++G++ +P+
Sbjct: 47 --------------------PFAAEAKFFTESRLLQRDVQIVLESCHN-QNILGTILHPN 85
Query: 313 GESAKDLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRLRMWTNYVPPASNS 372
G ++ L+ G+A+ V+WS + A+ +L+ AE AK+ +LR+W +YV P +N
Sbjct: 86 G----NITELLLREGFARCVDWSIAVYTRGAE-KLRAAERFAKERKLRIWRDYVAPTANL 140
Query: 373 KAIHNQNFTGKVVEVVSGDCIIVADDSIPYGSPLAERRVNLSSIRCPKVGNPRRDEK--- 429
+ ++ F KV++V++ D I+V +S + + ++LSSIR P++ +K
Sbjct: 141 EQ-KDKQFVAKVMQVLNADAIVVKLNSGDH------KTIHLSSIRPPRLEGEGAQDKNRK 193
Query: 430 -----PAPYAREAKEFLRTRLLGRQVSVEMEYSRKIVPTDGSAVPSPAADSRVMDFGSVF 484
PY EA+EFLR +L+G++V+V ++Y R P+ A V F
Sbjct: 194 LRPLYDIPYMFEAREFLRKKLIGKKVNVTVDYIR----------PASTATDTVPAF---- 239
Query: 485 LLSATKADSDDTPSSIPSAGSQPTGVNVGELVVGRGFGTVIRHR-DFEERSNYYDALLTA 543
S+ T +++ G+N+ E +V +G TVIR+R D ++RS++YD LL A
Sbjct: 240 --------SERTCATVCIG-----GINIAEALVSKGLATVIRYRQDDDQRSSHYDELLAA 286
Query: 544 ESRALSGRKGIHSAKDPPVMHITDLTTTSAKKAKDFLPFLHRSRRIPAVVEYVLSGHRFK 603
E+RA+ KG+HS K+ P+ + D+ + +KA +FLPFL R+ R AVVEYV SG R K
Sbjct: 287 EARAIKNGKGLHSKKEVPIHRVADI-SGDTQKANEFLPFLQRAGRSEAVVEYVFSGSRLK 345
Query: 604 LLIPKETCSIAFALSGVRCP----------GRGEPYSEEAIALMRRKIMQRDVEIEVETV 653
L +PKETC I F L+G+ CP GEP+SEEA + ++QR+VE+EVE +
Sbjct: 346 LFLPKETCLITFLLAGIECPRGARNLPGMVQEGEPFSEEATQFTKELVLQREVEVEVEAM 405
Query: 654 DRNGTFLGSLWESRTNVALTLLEAGLAKLQTSFGSDRIPEFHLLDRAEQSAKKQKLKIWE 713
D+ G F+G L N+++ L+E L+++ F ++R P L AE+SA++++ K+W
Sbjct: 406 DKAGNFIGWLHVDGLNLSVALVEHSLSRVH--FAAERSPYGKALLAAEESARQRRQKVWA 463
Query: 714 NFVE--GEEVSNGANVESKQQEVLKVIVTEVLGGGKFYVQTVGDQKIASIQQQLAALNLK 771
++ E EEV A + + V VTEV FYVQ V + A ++Q + +L +
Sbjct: 464 HYEESPSEEVVAVAEEKERSATYRPVFVTEVTDELHFYVQDV--ETGAQLEQLMDSLRAE 521
Query: 772 EA---PVLGAFSPKKGDIVLCYFHADKSWYRAMVVNTPRGPVESPKDIFEVFYIDYGNQE 828
A PV G++ P++GD + F D WYRA V VESP + +FYIDYGN+E
Sbjct: 522 VAAHPPVEGSYVPRRGDFCIAKF-VDGEWYRARVEK-----VESPTKV-HIFYIDYGNKE 574
Query: 829 QVAYSQLRPLDQSVS--AAPGLAQLCSLAYIKSPSLEEDFGQEAAEYLSELTLSSGKEFR 886
+ S+L L S P A + A+I+ P EE + ++ +
Sbjct: 575 TLPPSRLAALPPPFSPRTLPPQATEYAFAFIQVPQDEEARADAVDSAVRDI---QNTQCL 631
Query: 887 AQVEERDTSGGKVKGQGTGTILAVTLVAVDAEISVNAAMLQEGLARMEKRNRWDRKERKV 946
VE G G TL D + V +++EGL ++ R +R+ +KV
Sbjct: 632 LNVE-----------HGGGGCPHATLQLADTKGDVGLGLVREGLVMVQPRA--ERQFQKV 678
Query: 947 GLDSLQKFQDDARKERRGMWQYGDVESDDED 977
+ L Q+ A+ R +W+YGD +DD D
Sbjct: 679 MTEYLNA-QETAKSARLNLWRYGDFRADDAD 708
Score = 96.3 bits (238), Expect = 5e-18
Identities = 99/348 (28%), Positives = 150/348 (42%), Gaps = 75/348 (21%)
Query: 41 KSITLASLITPRLARRGGVDE----------PFAWESREYLRKLCIGKEVTFRVDY---- 86
K+I L+S+ PRL G D+ P+ +E+RE+LRK IGK+V VDY
Sbjct: 170 KTIHLSSIRPPRLEGEGAQDKNRKLRPLYDIPYMFEAREFLRKKLIGKKVNVTVDYIRPA 229
Query: 87 ------NVASINRDFGTVFLGEKNVGVLVVSQGWAKVREQGQQKGEVSPYLAELLRLEEQ 140
A R TV +G N+ +VS+G A V Q + S + ELL E +
Sbjct: 230 STATDTVPAFSERTCATVCIGGINIAEALVSKGLATVIRYRQDDDQRSSHYDELLAAEAR 289
Query: 141 AKQEGLGRWSKVPGAAEASIRNLPPSAIGDASNFDAMG---LLAANKGSPMEAIVEQVRD 197
A + G G SK + +P + D S L + EA+VE V
Sbjct: 290 AIKNGKGLHSK---------KEVPIHRVADISGDTQKANEFLPFLQRAGRSEAVVEYVFS 340
Query: 198 GSTLRVYLLPEFQFVQVFVAGIQSPQMGRRAAPETVVETELTADENDGDVPGEPRPPLTS 257
GS L+++L E + +AGI+ P+ G R P V E E
Sbjct: 341 GSRLKLFLPKETCLITFLLAGIECPR-GARNLPGMVQEGE-------------------- 379
Query: 258 AQRLAVSSSTETAADPFGPDAKFFTEMRVLNRDVRIVLEGVDKFSNLIGSVYYPDGESAK 317
PF +A FT+ VL R+V + +E +DK N IG ++ DG
Sbjct: 380 ---------------PFSEEATQFTKELVLQREVEVEVEAMDKAGNFIGWLHV-DG---L 420
Query: 318 DLALELVENGYAKYVEWSANMMEEEAKRRLKTAELEAKKSRLRMWTNY 365
+L++ LVE+ ++ V ++A + L AE A++ R ++W +Y
Sbjct: 421 NLSVALVEHSLSR-VHFAAE--RSPYGKALLAAEESARQRRQKVWAHY 465
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.315 0.133 0.378
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,672,278,818
Number of Sequences: 2540612
Number of extensions: 72967592
Number of successful extensions: 145991
Number of sequences better than 10.0: 290
Number of HSP's better than 10.0 without gapping: 89
Number of HSP's successfully gapped in prelim test: 201
Number of HSP's that attempted gapping in prelim test: 143878
Number of HSP's gapped (non-prelim): 1008
length of query: 990
length of database: 863,360,394
effective HSP length: 138
effective length of query: 852
effective length of database: 512,755,938
effective search space: 436868059176
effective search space used: 436868059176
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 80 (35.4 bits)
Lotus: description of TM0094a.6


